Abstract
Breast Cancer (BC) is the most common cause of cancer-associated deaths in females worldwide. Despite advancements in BC treatment driven by extensive characterization of its molecular hallmarks, challenges such as drug resistance, tumor relapse, and metastasis persist. Therefore, there is an urgent need for alternative treatment approaches with multi-modal efficacy to overcome these hurdles. In this context, natural bioactives are increasingly recognized for their pivotal role as anti-cancer compounds. This study focuses on predicting molecular targets for key herbal phytoconstituents—gallic acid, piperine, quercetin, resveratrol, and beta-sitosterol—present in the polyherbal formulation, Krishnadi Churna. Using an in-silico network pharmacology model, key genes were identified and docked against these marker compounds and controls. Mammary carcinoma emerged as the most significant phenotype of the putative targets. Analysis of an online database revealed that out of 135 predicted targets, 134 were mutated in breast cancer patients. Notably, ESR1, CYP19A1, and EGFR were identified as key genes which are known to regulate the BC progression. Docking studies demonstrated that the herbal phytoconstituents had similar or better docking scores than positive controls for these key genes, with convincing protein-ligand interactions confirmed by molecular dynamics simulations, MM/GBSA and free energy landscape (FEL) analysis. Overall, this study highlights the predictive potential of herbal phytoconstituents in targeting BC genes, suggesting their promise as a basis for developing new therapeutic formulations for BC.
Similar content being viewed by others
Introduction
Breast cancer (BC) is the most prominent malignancy among females worldwide and as per GLOBOCAN 2022 reports, it accounts for 23.8% of the total cancer cases and shares the highest cancer mortality rate (15.4% of the total cancer deaths)1. BC is a heterogeneous disease at the molecular level, and is clinically categorized as luminal estrogen receptor-positive (ER+) and progesterone receptor positive (PR+); human epidermal growth factor receptor 2 positive (HER2+); and triple-negative breast cancer (TNBC)2. Locoregional therapies (surgery and radiotherapy) are major modalities for BC management, but de-escalation schemes have become the standard of care today. Over the decade, the treatment options have been chosen by considering the molecular heterogeneity of breast cancer, and importance is given to biologically directed therapies or targeted therapies, especially in HER2 + and TNBC3. Several systemic therapies including endocrine therapy, chemotherapy, immunotherapy, etc. have been adapted as part of the treatment regime4.
Despite significant improvement in current therapy and appreciable progress in scientific research for targeting breast cancer and extending a patient’s life; there exists a gap in the complete prevention or cure of the disease either because the treatment response is often short-term, or mostly prone to resistance like endocrine resistance or chemotherapy resistance, or due to high metastasis of breast cancer cells4,5. Hence, there is an urgent need to explore molecular causal factors and an appropriate treatment regime, which can overcome the hurdles of existing therapy.
In this context, natural bioactive compounds from Indian Traditional Medicine (Ayurveda) have shown positive outcomes in preventing cancer or suppressing its progression6. Notably, many modern chemotherapy drugs also have roots in one or another traditional medicine. To date, many preclinical studies on Indian botanicals have come showcasing their tumoricidal activities against BC7,8,9. Indian traditional medicine has the potential for repurposing as the Ayurveda philosophy recommends using a group of phytoconstituents that can prevent, manage or cure a disease by acting on multiple drug targets10,11,12. In this study, we have studied the marker compounds present in a less explored polyherbal Ayurvedic formulation, Krishnadi Churna(KC), which is known for the management of bronchial asthma, cough, fever, and others13,14,15,16. Additionally, the herbal phytoconstituents and individual herbs (dried fruits of Piper longum, Vitis vinifera, and Scindapsus Officinalis, whole plant parts of Fagonia cretica, and galls of Pistacia integerrima) of this polyherbal formulation are reported to have antioxidant, anti-inflammatory and anticancer activity17.
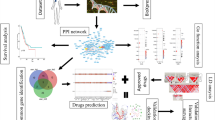
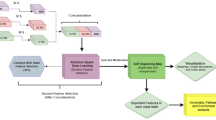
One drug one target approach to treat cancer faces many challenges of efficacy and sustainability. Currently, the integration of network biology and poly-pharmacology as network pharmacology has gained much recognition as an approach for multi-target drug development. These approaches can serve as useful tools to understand the putative actions and mechanisms of complex poly-herbal formulations11. Hence, in this study, we studied the marker compounds- gallic acid, piperine, quercetin, resveratrol, and beta-sitosterol present in the polyherbal formulation, Krishnadi Churna17. Molecular targets of all five marker compounds were predicted and network pharmacology was built. Interestingly, mammary carcinoma was identified as the most significant phenotype of the targeted genes. Further, key genes were identified, and molecular docking was performed, followed by molecular simulation studies (Fig. 1). The overall results showed that the studied botanicals can target the genes which are known to be involved in BC progression; hence can be possibly intervened to be developed as a new therapeutic drug for BC targeting.
A graphical abstract depicting the summary of the study.
Materials and methods
Bioinformatics tools and databases
Molecular docking was performed using the Schrodinger Glide software (Schrödinger Release 2024-3: Glide, Schrödinger, LLC, New York, NY, 2024.) Molecular dynamics was performed by Schrodinger Desmond software (Schrödinger Release 2024-3: Desmond Molecular Dynamics System, D. E. Shaw Research, New York, NY, 2024. Maestro-Desmond Interoperability Tools, Schrödinger, New York, NY, 2024.) Network construction was done through Cytoscape_v3.9.1. The Human Protein Atlas (HPA) database (The Human Protein Atlas) was used to investigate the association between the key gene expression and overall survival of BC patients18. cBioportal (cBioPortal for Cancer Genomics) was used to assess the mutation frequency of BC genes in patients19,20.
ADME screening
A simplified molecular-input line-entry system (SMILES) was searched for marker/bio-active compounds, followed by screening for Oral bioavailability, Drug-likeness, and pharmacokinetic analysis using SwissADME (http://www.swissadme.ch/) software21. Three or more models among five drug-likeness models (Lipinski, Ghose, Veber, Egan, and Muegge) were selected as bioactive with good bioavailability if met “yes”.
Network construction and pathway enrichment analyses
The compound–target network was constructed with persuasive representation using Cytoscape 3.9.1 (https://cytoscape.org/) software22. GeneCodis4 (https://genecodis.genyo.es/)23 was used to elucidate the phenotype of the targeted genes. Simultaneously, ShinyGO 0.76 (http://bioinformatics.sdstate.edu/go76/)24 was used to perform Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis with FDR cut-off 0.05, pathway size of minimum 2 to maximum 2000 genes and 20 pathways to be shown in the graph25,26,27,28. Gene Ontology (GO) pathway enrichment analysis was performed using the online functional annotation and enrichment tool g-profiler (https://biit.cs.ut.ee/gprofiler/gost)29. Protein-protein interactions were studied using the STRING database (https://string-db.org/cgi/input?sessionId=bqt6h0oqpCNwinput_page_show_search=on)30 with the highest confidence (0.9) of interaction score.
Ligand selection and preparation
The marker compounds present in the polyherbal formulation, Krishnadi Churna were used for computational simulation, where gallic acid (phenol), piperine (alkaloid), Resveratrol (phenol), Quercetin (flavonol), and beta-sitosterol (phytosterol) have been chosen as ligands17 along with Raloxifene core, Erlotinib, and Letrozole, which have been used as positive controls. The 2D SDF files for all compounds were obtained by querying the PubChem database31. To ensure the highest level of accuracy, all small molecule ligands were refined per molecular modelling tools using LigPrep to convert 2D to 3D. Moreover, the tool is capable of handling tautomeric, and neutral ionization pH states, as well as energy minimization using OPLS4. Therefore, we obtained molecules with low-energy 3D structures, which were appropriately ionized and tautomerised to be used for computational chemistry studies.
Target protein retrieval and preparation
Putative targets were analyzed using BindingDB (similarity score of > 0.7)32. Further, a Protein-protein interaction (PPI) network was constructed using STRING, with an interaction score of 0.9. Further filtered common targets (3 genes) from the above two steps were taken as the key genes. Cytohubba, a Cytoscape plugin was also used to identify hub genes, two out of three key genes were found to be the top nodes in the majority of algorithms, hence considered as super crucial genes in this study. However, molecular docking and molecular dynamics were done on all three key genes/targets (ESR1, CYP19A1, and EGFR; ESR1 and EGFR being the super crucial genes). Subunits of the above target genes having previously reported ligands33,34,35 were derived from the PDB structure database. In any molecular modelling study, the X-ray PDB ID artifacts have to be fixed before use for any computational studies. Therefore, we used Schrodinger protein preparation for all protein PDB to ensure protein compliance. All water molecules were deleted except those forming any bond near the target pocket. ProPKa software maintained the required pH of protein. Prime software was utilised for filling any missing amino acid or loop-in side chains. The protein minimization used force field-based energy optimization utilizing OPLS 4 with energy convergence with a cutoff of 0.3 Armstrong to maintain structural X-ray coordinate integrity. Finally, we got optimal energetically and structurally relevant protein coordinates in maegz format which was used for in-silico calculations.
Molecular docking
A molecular docking procedure requires three major inputs, the prepared ligand, protein grid centroid, and docking mode. The ligands for small molecules and phytochemicals were taken from previous steps. Targets were ESR1 (5T97), CYP19A1 (4GL7), and EGFR (4LRM) for this study. Based on ligands bound to receptors or binding pockets in targets, the centroid of receptors was identified using Schrodinger SiteMap. We generated the grid files using the Receptor Grid Generation module. We analyzed the target-ligand complexes with Schrodinger glide software to be applied for molecular docking. Molecular docking using rigid receptors and flexible ligand conformations is performed using SP precision mode with default settings. The output was a pose viewer file showcasing the protein-ligand complex. Thus, phytochemical molecules were placed in identified binding pockets, and resulting in affinity scores and interactions.
Molecular dynamics
Only the receptor-ligand complex with the highest binding affinity was simulated to determine its thermostability. Molecular docking was enhanced further by induced fit docking of protein-ligand couples. All molecular dynamics calculations were conducted using Schrodinger-Desmond software. The thermodynamic calculations were carried out in three stages. To simulate the biological system, the TIP3P solvent system was utilized to dissolve the ligand in the first protein-ligand complex. In the next step, we conducted molecular dynamics with eight phase stabilizations, and finally, we retrieved NPT 100 nanosecond production trajectory data incorporating velocities, energies, and coordinates. The simulation was conducted using the Nose-Hover thermostat and the Martyna tobias klein barostat to maintain an atmospheric pressure of 1 bar and a temperature of 300 K. Based on the above data, experimental thermodynamic stability, affinity dynamics, and interaction and nonbonding interactions have been evaluated.
Molecular mechanics generalized born surface area (MM/GBSA)
The molecular mechanic-generalized born surface area (MM/GBSA) approach was used to estimate the post-MD binding free energies using a set of predetermined parameters for all MD complexes using Schrodinger suite, LLC, New York, NY, USA, 2024-3. At around the same time, the binding free energy was estimated at every 10 ns frame using the OPLS 2005 force field, the VSGB solvent model and rotamer search techniques36.
Results
Five marker compounds-gallic acid (phenol), piperine (alkaloid), resveratrol (phenol), quercetin (flavonol), and beta-sitosterol (phytosterol), were checked and quantified earlier for their presence in Krishnadi Churna through HPTLC method17. This study aims to find out the putative human targets and associated phenotype of these five marker compounds. All five bio-actives were screened for Oral bioavailability, Drug-likeness, and pharmacokinetic analysis using SwissADME (Table 1 and Supplementary Fig. 1); except Beta-sitosterol, all qualified for Drug likeness based on predefined criteria, and among all only Piperine qualified for oral bioavailability. A detailed description of ADME screening and pharmacokinetic analysis is shown in Supplementary File 2. Since all five bio-actives are the marker compounds of polyherbal formulation, these predictions may change with the presence of multiple phytoconstituents together. Hence, all five biomarkers were considered for further analysis.
Targets and network analysis of polyherbal marker compounds
Putative targets for each bioactive were predicted using BindingDB with a similarity score of 0.7.; where 29 targets for Resveratrol, 14 for Gallic Acid, 8 for Piperine, 90 for Quercetin, and 27 for Beta-sitosterol were found. In the group of any two bio-actives; Resveratrol and Galic Acid had 5, Resveratrol and Quercetin had 15, Resveratrol and Beta-sitosterol had 2, Gallic Acid and Quercetin had 6, Piperine and Quercetin had 3, and Quercetin and Beta-sitosterol had 7 common targets. Among the group of any three bio-actives, only Resveratrol, Gallic Acid and Quercetin had 3, and Resveratrol, Quercetin and Beta-sitosterol had 2 common targets. No common target was found for any four or five bioactive groups. All the bioactives along with their target list are shown in Table 2.
A total of 168 compound targets were found and after removing duplicates the remaining 135 targets were eloquently represented along with their bio-actives using Cytoscape (Fig. 2). A Cytoscape network was also constructed for individual bioactive and their targets (Supplementary Fig. 2). Further, a Protein-protein interaction (PPI) network was constructed using STRING, and the targets with an interaction score of 0.9, the highest confidence is shown in Fig. 3. Near about 67.4% of the total putative targets had the highest confidence of interaction, a few of these proteins are HDAC1, ESR1, VEGFA, RELA, EGFR, etc.
Network of marker compounds (herbal phytoconstituents) present in polyherbal formulation and their putative targets.
Protein–protein interaction of putative targets of herbal phytoconstituents with an interaction score of 0.9.
Polyherbal marker compounds target cancer-related pathways
To elucidate the phenotype of target genes, a list of target genes was uploaded as an input in GeneCodis4, annotation for the phenotype (DisGeNET, HPO, OMIM) was selected, and analysis was launched. As shown in Fig. 4a., most genes showed the phenotype for mammary carcinoma, followed by prostate cancer and Alzheimer’s Disease. Figure 4b. represents the annotations and the genes associated with them in the form of a network where genes like RELA, ABCB1, CYP19A1, SERPINE1, AKR1C3, MMP9, TERT, etc., belonged to mammary and prostate cancer, and genes like APP, BACE1, MOAB, etc. belonged to Alzheimer’s disease; while ESR1, EGFR, VEGFA, GSK3B, ACHE, etc. are crucial for cancer and Alzheimer’s both. Further, KEGG pathway enrichment analysis25,26,27,28 was performed and as shown in Fig. 4c, Pathways in cancer, NF-kappa B signaling pathway, Ras signaling pathway, MAPK signaling pathway, and PI3K-Akt signaling pathway were among the highly enriched pathway. Interestingly, these are the key pathways for breast cancer survival, proliferation, metastasis, and drug resistance. A myriad of reports is available showcasing their crucial role in breast cancer progression and how different strategies are being adapted to target these pathways37,38,39,40,41,42. Gene Ontology (GO) analysis was also performed on target genes and as shown in Fig. 4d and Supplementary Fig. 3, these potential targets of polyherbal marker compounds were mainly located in the cytoplasm (GO:0005737), receptor complex (GO:0043235), cell surface (GO:0009986), nucleoplasm (GO:0005654), protein kinase complex (GO: 1902911), chromosome (GO: 0005694); involved in the biological process like cell death (GO:0008219), apoptotic process (GO:0006915), peptidyl-serine modification (GO:0018209), cell population proliferation (GO:0008283), endocytosis (GO:0006897), cell cycle (GO:0007049), cell junction assembly (GO:0034329); and regulates molecular functions like identical protein binding (GO:0042802), nuclear receptor activity (GO:0004879), signaling receptor activity (GO:0038023), enzyme binding (GO:0019899), protein-containing complex binding (GO:0044877), chromatin binding (GO:0003682), non-membrane spanning protein tyrosine phosphatase activity (GO:0004726), etc.
(a) Bar graph showing the most significant phenotype for herbal phytoconstituents’ targets. (b) Network of annotations and genes associated with the gene targets’ phenotype. (c) KEGG Pathway enrichment analysis of BC genes targeted by herbal phytoconstituents. This graph was generated using ShinyGO 0.7624,25,26,27,28. (d) Gene Ontology analysis showing Molecular Function, Biological Process and Cellular Component of gene targets.
Key genes identified are crucial for BC progression
Further, key genes were identified among the various targets of marker compounds. To start with, 28 targets with ≥ 5 interacting proteins in the PPI network were chosen. Simultaneously, proteins targeted by the majority of bio-actives were selected, here, as evident from Table 2, five proteins-ESR1, CA12, CYP19A1, EGFR, and TTR were targeted by 3 bioactive compounds and hence selected for analysis. Further, the common targets from the two selections were considered as the key genes (Fig. 5a). Three key genes selected were, ESR1, CYP19A1, and EGFR; plenty of reports based on clinical and pre-clinical studies have shown the important role of these genes in poor clinical outcomes, relapse of disease, therapy resistance and metastasis of BC43,44,45,46. Also, based on the calculation of cytoHubba, the top 10 nodes ranked by all twelve algorithms were evaluated. Out of 12 algorithms, ESR1 was ranked as the top 10 nodes in 9 algorithms and EGFR in 10 algorithms (Fig. 5b) hence considered as the super crucial gene in this study.
Further, the Human Protein Atlas was explored to check the 5-year survival of Breast cancer patients with varied expression of the key genes- ESR1, CYP19A1, and EGFR18. The five-year survival rate for patients with high CYP19A1 expression was 76%, compared to 83% for those with low expression, with a p-value of 0.094. Similarly, for ESR1, the survival rates were 80% and 82% respectively, with a p-value of 0.083. For EGFR, the rates were 85% and 80% with a p-value of 0.026. Interestingly, a positive correlation in their expression was observed with poor survival of BC patients (Fig. 5c–e). However, EGFR showed an opposite trend, which could be because of its conflicting role in BC47. Additionally, we checked the mutation frequency of the target gene in BC patients using the program and tools made available online at cBioportal (cBioPortal for Cancer Genomics)19,20. Interestingly, out of 135 putative targets, 134 genes were found mutated in breast cancer patients (Fig. 5f). Further, the mutation frequency of key genes was identified in breast cancer patients, and as shown in Table 3, 7.3% mutation frequency was observed for ESR1, 1.5% for EGFR and 0.30% for CYP19A1. Overall, the published manuscripts43,44,45,46 and database results highlight the critical role of these genes in breast cancer progression and prognosis, underscoring the importance of developing therapeutic strategies that target them.
(a) Venn Diagram depicting the key genes identified among the putative targets of herbal phytoconstituents. (b) Identified Hub gene network of herbal phytoconstituents for different algorithms of CytoHubba. Survival Plot with high and low expression of (c) ESR1, (d) CYP19A1, (e) EGFR in BC patients from Human Protein Atlas online database. (f) Venn Diagram depicting mutation status of target genes in BC from cBioportal.
Molecular docking
Marker compounds were predicted to possess a good affinity for key genes (targets). Possible regions in the target receptor ESR1, CYP19A1, and EGFR were examined to find the potential binding poses of bioactive, using subunits of the protein structure ESR1(5T97), CYP19A1 (4GL7), and EGFR (4LRM). Gallic acid (ID 370), Piperine (ID 638024), Quercetin (ID 5280343), and Resveratrol (ID 445154) were docked with the above targets. Raloxifene Core (ID 445920), Letrozole (ID 3902), and Erlotinib were the positive controls for ESR1, CYP19A1, and EGFR respectively. Results obtained after docking revealed strong interactions of bioactive against the targets in terms of binding energy (Kcal/mol) shown in Table 4, least binding energy (more negative binding energy) is interpreted as more stability.
The binding affinity of ESR1 (5T97) for positive control (Raloxifene Core) was − 8.819 Kcal/mol. Piperine expressed − 8.288 Kcal/mol binding affinity with 5T97. It was nearest to the positive control. Resveratrol (-7.95 Kcal/mol), Gallic acid (-6.984 Kcal/mol), and Quercetin (-6.755 Kcal/mol) also exhibited a fair binding affinity. Resveratrol showed hydrogen bonding with LEU387, ARG394, LEU346 and pi-pi bond with PHE404 (Fig. 6a). Gallic acid showed Hydrogen bonding with LEU346, GLU353, ARG394, and pi-pi bonding with PHE404 (Fig. 6b). With dual hydrogen bonding with ASP351, the Quercetin molecule has no other bond in molecular docking. Raloxifene core exhibits hydrogen bonding with GLU353, and ARG394 while pi-pi bonding with PHE404 (Fig. 6c).
The binding affinity of CYP19A1 (4GL7) for positive control (Letrozole) was − 6.771 kcal/mol. Letrozole exhibits hydrogen bonding with VAL373 and MET374 (Fig. 6d). Among bioactive, Quercetin exhibits the best docking score (-8.628 Kcal/mol) and hydrogen bonds ASP309, MET374, and LEU477 and a pi-pi bonding HEM600(Fig. 6e). Resveratrol (-7.089 Kcal/mol), Piperine (-5.878 Kcal/mol), and gallic acid (-5.516 Kcal/mol) also reflected a moderate binding affinity. Piperine has a hydrogen bond with GLN218, pi-cation bond with ARG192 and pi-pi Bonding with HIE480 and PHE221 (Fig. 6f).
The binding affinity of EGFR (4LRM) with positive control (Erlotinib) was − 6.285 Kcal/mol. In comparison, the binding affinity for bioactive was Quercetin (-8.629 Kcal/mol), Resveratrol (-7.772 Kcal/mol), Gallic acid (-6.501 Kcal/mol), and Piperine (-6.464 Kcal/mol). All ligands exhibit better docking scores than the positive control. Erlotinib has hydrogen bond interaction at MET796 and LYS745 (Fig. 6g). Quercetin has hydrogen bonds with MET796 and GLU762 (Fig. 6h). Gallic acid showed 2 converging hydrogen bonds with GLU762 (Fig. 6i). With the highest interaction, Resveratrol exhibits 3 hydrogen bonding with MET796, THR793, and GLU762 (Fig. 6j).
The mutant variants of ESR1 and EGFR were also analyzed to identify the potential binding poses of the bioactive compounds using the protein subunits 4Q50 ESR1 (D538G) and 4I20 EGFR (L858R). Docking results revealed strong interactions between the bioactives and the target proteins, as indicated by their binding energies (Kcal/mol) presented in Table 5.
The binding affinity of Raloxifene core, the positive control, with the mutant ESR1 (D538G) was recorded at -10.116 Kcal/mol. In comparison, Gallic acid (-6.331 Kcal/mol), Piperine (-8.769 Kcal/mol), Quercetin (-6.894 Kcal/mol), and Resveratrol (-8.149 Kcal/mol) also showed significant binding affinities. In terms of molecular interactions, Raloxifene core showed a hydrogen bond with GLU353 and a pi-pi bond with PHE404 (Fig. 7a). Gallic acid formed hydrogen bonds with GLU353, LEU387, and a pi-pi bond with PHE404 (Fig. 7b). Piperine and Quercetin exhibited hydrogen bonding with HIS524 and ACE532, respectively, while Resveratrol formed hydrogen bonds with LEU387 and GLU353, along with a pi-pi bond with PHE404 (Fig. 7c-e).
The binding affinity of mutant EGFR (L858R) with the positive control, Erlotinib, was found to be -4.456 Kcal/mol. Notably, all the tested bioactives demonstrated superior docking scores compared to the positive control: Gallic acid (-5.177 Kcal/mol), Piperine (-4.673 Kcal/mol), Quercetin (-5.686 Kcal/mol), and Resveratrol (-5.915 Kcal/mol). In terms of specific interactions, Erlotinib showed hydrogen bonding with CYS797 and LYS745. Gallic acid formed hydrogen bonds with ASP800 and CYS797, while Resveratrol showed hydrogen bonding with LYS745, MET793, and ASP800. Quercetin established hydrogen bonds with ASN842 and ASP800. Piperine exhibited a hydrogen bond with CYS797(Fig. 7f-j).
Docking results showing H-bonds and other interactions between protein-ligand complex of (a) ESR1-resveratrol, (b) ESR1-gallic acid, (c) ESR1-raloxifene core, (d) CYP19A1-letrozole, (e) CYP19A1-quercetin, (f) CYP19A1-piperine, (g) EGFR-Erlotinib, (h) EGFR-quercetin, (i) EGFR-gallic acid, (j) EGFR-resveratrol.
Docking results showing H-bonds and other interactions between protein-ligand complex of (a) ESR1 (D538G)-raloxifene core, (b) ESR1 (D538G)-gallic acid, (c) ESR1 (D538G)-piperine, (d) ESR1 (D538G)-quercetin, (e) ESR1 (D538G)-resveratrol, (f) EGFR (L858R)-erlotinib, (g) EGFR(L858R)-gallic acid, (h) EGFR(L858R)-piperine, (i) EGFR(L858R)-resveratrol, (j) EGFR (L858R)-quercetin.
Molecular dynamics
Molecular dynamics simulations were performed to analyze the protein-ligand complex behavior in explicit biological solutions. Such behavior is predicted using RMSD (root mean square deviation), ligand affinity, and non-bonding interaction in the protein-ligand complex.
The best dock score for CYP19A1 was Quercetin, hence CYP19A1-Quercetin complex was selected for Molecular docking simulation. Complex simulation was performed at a standard temperature (300 K) for a stipulated time of 100 nanoseconds. Protein was positively charged with total residue of 454. Initial protein RMSD fluctuation was nearly at 1 A. Ligand was stable between 2 and 2.7 A throughout the simulation period. Both the ligand and the protein were stable in a frame of 1–2.7 A. Complex interaction was stable throughout except at around the 50th ns frame. Here it deviates for more than 0.8 A. Figure 8a shows the significant interaction between the ligand and protein. A total of 25 contacts were noted as shown in Fig. 8b. The complex showed Hydrogen, Hydrophobic, and ionic bonds, having the highest interaction fraction of 1. Hydrogen bond was noted at ARG_115, ALA_306, THR_310 (39%), MET_374 (87%), SER_478. CYS 437 had interaction with Ferrous ions for 94% of the duration of the simulation. Protein SSE was 54.54% with 47.86% of alpha helix and 6.68% of beta strands (Fig. 8c).
Complex EGFR-Quercetin expressed the best dock score and was further subjected to simulation at standard temperature (300 K) for 100 nanoseconds. The interaction had positively charged protein of 309 total residues. Initial protein RMSD fluctuation was nearly at 1 A. Protein was stable throughout the simulation time at 2–2.5 A. Ligand was stable except at around 20–40 nanoseconds. Complex interaction was not found during this timeframe as shown in Fig. 8d. The complex had nearly 24 contacts as depicted in Fig. 8e. Complex shows H-bond and Hydrophobic bonds, having the highest interaction fraction around 1.6. H bond was seen at LYS_745, GLU_762 (75%), THR_793, GLN_794 (91%), MET_796 (98%), and ASP_858 (52%). Total protein secondary structures were 43.93%, with 28.54% alpha helix and 15.38% beta strands (Fig. 8f).
The ESR1-Piperine protein-ligand complex expressed the best dock score and was further subjected to simulation for 100 nanoseconds. Protein was negatively charged with 185 total residues. Initial protein RMSD fluctuation was nearly at 1 A. Both protein and ligand were unstable during this framework. Ligand RMSD fluctuated between 6 and 14 A. The complex had interaction twice during the 100 nanoseconds frame, at around 20ns and 60-80ns (Fig. 8g). A total of 32 contacts was seen between the protein-ligand as shown in Fig. 8h Complex shows H-bond and Hydrophobic bonds, having the highest interaction fraction around 0.25. No ionic bonds were noted. Hydrogen bond was noted at GLY_344, THR_347, TRP_383, GLY_420, MET_421, NMA_525. None of the interactions occurred for more than 30.0% of the simulation time in the selected trajectory. Total protein secondary structures were 71.32%, with 69.82% alpha helix and 1.50% beta strands (Fig. 8i).
The highest docking score for mutant EGFR-L858R was observed with Resveratrol, making the EGFR (L858R)-Resveratrol complex the prime candidate for molecular docking simulation. The simulation was conducted at a standard temperature of 300 K over 100 nanoseconds. The protein, consisting of 289 residues, carried a positive charge. The initial RMSD fluctuation for the protein was around 1Å, while the ligand remained stable within a range of 1–2Å throughout the simulation. Both the ligand and protein exhibited stability between 1-2.4Å. Notably, the complex interaction remained stable at the 10th and 70th nanosecond frames. As shown in Fig. 9a, significant interactions occurred between the ligand and protein, with a total of 19 contacts identified (Fig. 9b). The complex featured hydrogen, hydrophobic, and ionic bonds, with the highest interaction fraction being 1.4. Hydrogen bonds were observed at LYS_745 (55%), GLN_791 (76%), MET_793 (95%), and ASP_800 (40%). The protein secondary structure elements (SSE) consisted of 48.20%, with 30.68% alpha helices and 17.52% beta strands (Fig. 9c).
Similarly, the best docking score for mutant ESR1-D538G was achieved with Piperine, prompting molecular dynamics analysis of the ESR1-Piperine complex. The simulation, performed at 300 K for 100 nanoseconds. The protein, composed of 222 residues, was positively charged. The initial RMSD fluctuation for the protein was about 1Å, while the ligand remained stable between 4.8–5.6Å throughout the simulation. The complex interaction remained stable at various intervals, particularly at the 10th, 30th, and 70th nanosecond frames. As illustrated in Fig. 9d, significant interactions were observed between the ligand and protein, with 25 contacts noted (Fig. 9e). Hydrogen, hydrophobic, and ionic bonds were present, with the highest interaction fraction recorded at 0.7. Hydrogen bonds were observed at GLY_420, HIS_524 (74%), and SER_527. The protein SSE was 67.65%, comprising 64.61% alpha helices and 3.04% beta strands (Fig. 9f).
Molecular docking simulation results of (a) protein ligand RMSD, (b) protein ligand contacts and (c) protein secondary structure analysis of CYP19A1-Quercetin complex. RMSD, Protein Ligand Contacts and Protein secondary structure analysis of (d–f) EGFR-Quercetin and (g–i) ESR1-Piperine complex.
Molecular docking simulation results of (a) protein ligand RMSD (b) protein ligand contacts and (c) protein secondary structure analysis of EGFR (L858R)-Resveratrol complex. RMSD, Protein Ligand Contacts and Protein secondary structure analysis of (d-f) ESR1(D538G)-Piperine complex.
MM/GBSA analysis highlights a good ligand-binding affinity
Prime MM/GBSA (Molecular Mechanics Generalized Born Surface Area), also referred to as Gibbs binding free energy, is a computational chemistry method widely used to estimate the binding free energy of small molecules with their target proteins. This approach combines molecular mechanics calculations with the Generalized Born model for solvation, incorporating a surface area term to account for nonpolar solvation effects within the protein binding site. By evaluating Gibbs binding free energy, Prime MM/GBSA provides critical insights into ligand-receptor interaction strength, helping identify promising candidates for drug design.
In this study, the binding energy analysis of molecular dynamics (MD) trajectories, taken at fixed intervals of 10 frames across 1000 frames for three complexes—EGFR-Quercetin (5280343), ESR1-Piperine (638024), and CYP19A1-Quercetin (5280343)—yielded the significant observations. The EGFR-Quercetin (5280343) complex exhibited the binding affinity fluctuations within a range of -67 kcal/mol at 14 ns to -45 kcal/mol at 65 ns, with intermediate values oscillating between − 56 kcal/mol and − 52 kcal/mol throughout the 0-100 ns simulation frame (Fig. 10a). This indicates relatively stable binding interactions with moderate energy variations over time. In the ESR1-Piperine (638024) complex (Fig. 10b), the lowest binding energy was − 124 kcal/mol at 10 ns, and the highest was − 72 kcal/mol at 72 ns. The average binding energy, represented by the best-fit line, fluctuated between − 111 kcal/mol and − 90 kcal/mol over the 0-100 ns frame, reflecting strong and consistent binding affinity, with significant energy fluctuations highlighting dynamic ligand-receptor interactions. For the CYP19A1-Quercetin (5280343) complex (Fig. 10c), the lowest binding energy observed was − 37 kcal/mol at 20 ns, while the highest was − 9 kcal/mol at 71 ns. The energy varied between − 25 kcal/mol and − 20 kcal/mol throughout the 0-100 ns simulation, indicating a relatively weaker interaction compared to the other complexes.
Overall, the study reveals that all three ligands bind to their respective targets, with the ESR1-Piperine (638024) complex demonstrating the strongest and most stable binding affinity. The EGFR-Quercetin (5280343) complex also shows moderate stability, suggesting potential, though further refinement may be required for optimization.
MM/GBSA analysis of (a) EGFR-Quercetin (b) ESR1-Piperine and (c) CYP19A1-Quercetin complex.
Post molecular dynamics structural analysis of protein-ligand complex
An in-depth molecular dynamics trajectory analysis using principal component analysis (PCA) of the EGFR in complex with the ligand Quercetin (528034) has revealed key computational insights into this interaction. The study focused on the first two principal modes, which drive the collective motion of the protein-ligand complex during the molecular simulations. Pronounced anticorrelated fluctuations, particularly in the loop regions of the binding site residues, were observed, depicted by pink vectors in the image shown as Fig. 11a. These fluctuations suggest that Quercetin’s binding induces coordinated movements within the EGFR structure, which could be crucial for the receptor’s functional dynamics. The EGFR-Quercetin interaction exhibited remarkable stability, with restrained motion of EGFR in the presence of Quercetin, signaling a robust binding interaction. This stability was further supported by the free energy landscape (FEL) analysis, where the kernel distribution function served as the third dimension. The FEL plot revealed three distinct energy minima, representing stable conformational states of the EGFR-Quercetin complex (Fig. 11b). Importantly, the structural frames corresponding to these minima did not exhibit significant conformational changes, implying that the ligand binding interactions remained largely intact and were not disrupted by any major protein structural reconfigurations. Further analysis showed that the root mean square deviation (RMSD) of Quercetin in the EGFR-Quercetin (528034) complex ranged around 1.2 Å, with the radius of gyration (rGyr) spanning from 3.90 to 4.08 Å. The ligand’s torsion, corresponding to its six rotatable bonds, is shown in Fig. 11c. These results underscore Quercetin’s strong binding capability to EGFR across various energy states without inducing destabilizing structural alterations.
Previous studies utilizing principal component analysis (PCA) on protein-ligand molecular dynamics trajectories have provided valuable insights into the structural determinants that govern ligand-induced effects on molecular targets. This analysis identified significant alterations in binding site affinity, marked by pronounced vector variations within the binding site. These fluctuations are vividly illustrated in the protein’s secondary structure, with pink-highlighted vectors indicating the maximum fluctuations in the binding site (Fig. 11d). Furthermore, an energetic analysis of mode 1 versus mode 2 revealed three distinct energy minima, each carefully assessed for stability (Fig. 11e). In the ESR1-Piperine (638024) complex, the root mean square deviation (RMSD) ranged around 1.0 Å, while the radius of gyration (rGyr) extended between 4.75 and 5.25 Å. The torsion of the ligand, corresponding to its four rotatable bonds, is displayed in Fig. 11f. Despite these structural fluctuations, non-bonding interactions remained consistent throughout the simulation, highlighting the sustained binding efficacy of the ligand molecules.
The collective motion of the CYP19A1-Quercetin (5280343) complex is visualized in Fig. 11g-i, where the protein vectors at the interaction site are oriented in a specific direction, indicating a stable interaction (Fig. 11g). The overall protein movement also follows a unidirectional pattern, reinforcing the stability of the complex. The free energy landscape (FEL) plot of the CYP19A1-Quercetin (5280343) complex reveals three local energy minima (Fig. 11h), reflecting a limited number of energetically favoured conformations, similar to those observed in the EGFR-Quercetin complex. The Quercetin within the CYP19A1-Quercetin (5280343) complex exhibits an RMSD range of 0.8 Å, a radius of gyration (rGyr) between 3.90 and 4.08 Å, and torsion data for its six rotatable bonds (Fig. 11i). These findings collectively support the stability of the ligand within the complex.
Post molecular dynamics structural analysis of protein-ligand complex depicting (a) EGFR-Quercetin Protein Vectors, (b) EGFR-Quercetin PCA1 vs. PCA2 analysis, (c) EGFR-Quercetin heatmap representing RMSD, (d) ESR1-Piperine Protein Vectors, (e) ESR1-Piperine PCA1 vs. PCA2 analysis, (f) ESR1-Piperine heatmap representing RMSD, (g) CYP19A1-Quercetin Protein Vectors, (h) CYP19A1-Quercetin PCA1 vs. PCA2 analysis, (i) CYP19A1-Quercetin heatmap representing RMSD.
Discussion
BC is a global concern, associated with a maximum number of cancer-related deaths in females worldwide. Unfortunately, most cancer registries only record the incidence and mortality rate of cancer, but not the cancer relapse; hence, a clear picture of survivors and the ones with metastatic disease is still not available. Despite cornerstone BC treatment, late detection, therapy resistance, metastasis, and relapse are some of the significant hurdles in BC treatment4,5. Major emphasis needs to be put on multimodal treatment and novel drugs for the multidisciplinary management of this heterogeneous disease.
Currently, herbal medicines are gaining public interest, especially for the treatment of chronic and lifestyle disorders. According to the World Health Organization (WHO), 88% of all countries rely on traditional medicine, including herbal medicines, acupuncture, yoga, indigenous therapies, and others for primary health care48. Herbal formulations are under evaluation to treat different cancers including breast using modern technologies and have shown positive results as therapeutic agents49. Withania somnifera, Butea monosperma, Eclipta alba, Piper nigrum, Vernonia cinerea, Curcuma longa, etc. are among the explored herbs for the treatment of BC50,51,52,53,54.
In this study, we used the marker compounds of a polyherbal formulation from Indian Traditional Medicine, which has not yet been reported as an anticancer drug but possesses anti-oxidant and anti-inflammatory activity; nevertheless, the individual herbs (dried fruits of Piper longum, Vitis vinifera, and Scindapsus Officinalis, whole plant parts of Fagonia cretica, and galls of Pistacia integerrima) of this formulation are known to possess anti-cancer activity17. Piperine, the active component of Piper longum is known to induce apoptosis via increasing caspase 3 activity, upregulating Bax expression and downregulating Bcl-2 expression; reduce proliferation through the reduction in cyclin B1 expression and cell cycle arrest in the G2/M phase; reduce cell migration, and inhibit the expression of MMPb-9 and MMP-13 in BC cells55,56. Subcritical water extract and dried extract of vine shoots (Vitis vinifera) after pruning, rich in trans-resveratrol; showed preferential high cytotoxic effects on malignant cell lines compared to non-malignant cells in culture. Also, these extracts are reported to induce apoptosis in BC cells and necrosis (possibly ferroptosis) in cervical cancer cells and lead to the activation of NF-kB (p65 translocation)57. Entelo (Vitis vinifera Seed Extract) is reported to inhibit the MEK/ERK and PI3K/AKT signaling pathway and downregulate the expression of Interleukin-1 alpha, suppressing the lung metastasis of Triple-negative BC cells58. The ethyl acetate and methanol extract of the fruits of Scindapsus officinalis is reported to show the antitumor effect by increasing the mean survival time (MST) lifespan, reducing the solid tumor volume, and reversing the altered hematological parameters almost equal to normal in mice model and inducing cytotoxicity and anti-oxidant effect on human cancer cell lines59. Fagonia cretica (aqueous extract) is known to induce DNA damage and p53-associated G0/G1 phase cell cycle arrest, and FOXO3a-mediated apoptosis in BC cells60. The crude extract from Pistacia integerrima is reported to induce cytotoxicity in MCF-7 cells. Also, the compounds extracted from its chloroform extract including β-sitosterol were found to exhibit significant dose-dependent MDR reversing effect in cancer cells; supported by molecular docking study indicating their interaction with P-glycoprotein (P-gp)61,62.
Since very few reports are available on marker compounds of Krishnadi Churna, we explored the potential of these herbal phytoconstituents in targeting BC using in silico approaches and proposed the marker phytoconstituents with immense potential to be developed as phytopharmaceuticals. Prediction of gene targets of polyherbal maker compound - gallic acid, piperine, quercetin, resveratrol, and beta-sitosterol was done using in silico analysis. Collectively, 135 targets were predicted for the marker compounds, and most of them showed the phenotype either for mammary and prostate cancer or Alzheimer’s disease. Interestingly, KEGG pathway enrichment analysis also showed Pathways in cancer, NF-kappa B signaling pathway, Ras signaling pathway, MAPK signaling pathway, and PI3K-Akt signaling pathway as the highly enriched pathway. To be noted, all these are crucial pathways of BC progression37,38,40,41,42.
The current BC therapy is complex due to the molecular heterogeneity of the cancer. ER-positive tumors are mostly treated with endocrine therapy and a targeted approach is used for metastatic BC, such as CDK4 and CDK6 inhibitors, PI3K inhibitors, PARP inhibitors, and anti-PD-L1 immunotherapy63,64,65. Strikingly, the putative targets of these marker compounds also include ESR1, CDK6, PIK3CG, and PARP1 which suggests the substantial potential of this herbal formulation to target multiple molecular markers of BC and reduce the burden of multiple drugs.
Interestingly, in this study, ESR1, CYP19A1, and EGFR were identified as the key genes of polyherbal biomarkers, where ESR1, encodes an estrogen receptor and ligand-activated transcription factor. About 70% of breast cancers are ER-positive and are majorly targeted by Endocrine therapy (ET). However, the cancer relapses or progresses to the incurable metastatic stage in more than half of the patients either due to loss of ER expression, ER crosstalk with growth factor receptors, or acquisition of ESR1 mutations43. CYP19A1, also called aromatase, is involved in catalyzing the last steps of estrogen biosynthesis; associated with major regulatory events for intratumoral production of estrogens in breast malignant tissues hence been studied as a prognostic marker of breast cancer44,66. EGFR also known as ErbB1 and HER1, belongs to the family of most notable cancer molecular targets identified to date. EGFR overexpression is seen in almost half of the triple-negative breast cancer (TNBC) and inflammatory breast cancer (IBC) cases and is associated with large tumor size, poor differentiation, and poor clinical outcomes45,46.
Additionally, cBioportal was explored and out of 135 putative targets identified, 134 were found to be prone to a mutation in BC patients, where 7.3% mutation frequency was observed for ESR1, 1.5% for EGFR, and 0.30% for CYP19A1. Also, the tendency of mutations to co-occur was observed for ESR1-EGFR (pvalue = 0.041), ESR1-CYP19A1 (pvalue = 0.155), and most significantly for EGFR-CYP19A1 (pvalue = 0.007)19,20. All these results implicate the important role of putative targets of marker compounds present in Krishnadi Churna in BC.
Molecular docking of ESR1 with piperine demonstrated higher affinity compared to the positive control (raloxifene core), with interactions similar to those of raloxifene, which bonded with GLU353, ARG394, and PHE504. Bioactives showed similar molecular interactions with these amino acids as of positive control. Mutant ESR1 was also targeted through similar interactions. The docking score for EGFR with all the bioactives was higher than that of the positive control (erlotinib), with MET796 being a common interaction site for both bioactives and erlotinib. In mutant EGFR, while different molecular interactions formed, bioactives covered key interactions made by mutant-EGFR-erlotinib, including CYS797 and LYS745. Additionally, CYP19A1-letrozole’s docking score was surpassed by quercetin and resveratrol, with the bioactives mimicking all interactions except VAL373. The shared molecular interactions between the positive control and bioactives suggest a higher prediction of biological activity for the bioactives.
While the chemical nature of the other constituents in Krishnadi Churna remains unreported, it is recognized that even minor components can substantially impact biological activity through synergistic or opposing effects. Therefore, a future study is warranted to evaluate the comprehensive multitarget biological activity of Krishnadi Churna. Our study has shown a good chance of effective pharmacotherapeutic targeting of breast cancer; however, the drug concentration, absorption, distribution, metabolism and excretion play an important role in both in-vitro and in vivo studies, therefore a wet lab validation of the obtained data is warranted. Secondly, our research results cannot explain which chemical component from Krishnadi Churna exerts the most significant anti-BC effect. Finally, the molecular mechanism of these five marker compounds individually, in combination, or as a complete formulation in breast cancer cells still needs more clinical and experimental verification. Nevertheless, we have shown that the herbal phytoconstituents discussed in this study have the potential to modulate key molecular markers of BC, supported by Molecular Docking and Dynamics studies including MM-GBSA analysis and post Molecular Dynamics Structural Analysis of Protein-Ligand Complex; where phytoconstituents showed a similar or better docking score compared to positive controls of targets and good protein-ligand interactions under the explicit biological environment of Molecular Dynamic simulations.
Conclusion
Herbal phytoconstituents have the predictive potential to target BC genes. We hence postulate that herbal phytoconstituents should be explored more to design a better and more efficient treatment to curb BC growth.
Data availability
All the data generated have been submitted with this manuscript.
Abbreviations
- BC:
-
Breast cancer
- TNBC:
-
Triple-negative breast cancer
- ER:
-
Estrogen-receptor
- PR:
-
Progesterone-receptor
- HER 2:
-
Human epidermal growth factor receptor
- CYP19A1:
-
Cytochrome p450 family 19 subfamily a member 1
- EGFR:
-
Epidermal growth factor receptor
- MAPK:
-
Mitogen-activated protein kinases
- MDR:
-
Multi-drug resistance
- RMSD:
-
Root mean square deviation
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 74(3), 229–263 (2024).
Denkert, C. et al. Molecular alterations in triple-negative breast cancer—the road to new treatment strategies. Lancet 389(10087), 2430–2442 (2017).
Loibl, S., Morrow, P. P., Denkert, M. & Curigliano, C. Breast cancer. Lancet 397, 1750–1769 (2021).
Harbeck, N. et al. Breast cancer. Nat. Rev. Dis. Primers 5, 66 (2019).
Jin, X. & Mu, P. Targeting breast cancer metastasis. Breast Cancer Basic Clin. Res. 9, 25460 (2015).
Desai, A. G. et al. Medicinal plants and cancer chemoprevention. Curr. Drug Metab. 9(7), 581–591 (2008).
Vashi, R., Patel, B. M. & Goyal, R. K. Keeping abreast about Ashwagandha in breast cancer. J. Ethnopharmacol. 269, 113759 (2021).
Henamayee, S. et al. Therapeutic emergence of rhein as a potential anticancer drug: a review of its molecular targets and anticancer properties. Molecules 25(10), 2278 (2020).
Rafieian-Kopaei, M. & Movahedi, M. Breast cancer chemopreventive and chemotherapeutic effects of Camellia Sinensis (green tea): an updated review. Electron. Phys. 9(2), 3838 (2017).
Roy, A. Reigniting pharmaceutical innovation through holistic drug targeting. Drug Discov. 45 (2016).
Chandran, U. et al. Network pharmacology. Innovative Approaches Drug Discovery 127 (2017).
Borse, S. et al. Ayurveda botanicals in COVID-19 management: an in silico multi-target approach. PLoS ONE 16(6), e0248479 (2021).
Mathur, B. Brihad Nighantu Ratnakar, Mumbai. Khemraj Shrikrishnadass Prakashan, vol. 5/6, 495 (2012).
Srivastava, D. S. Sharangdhara Samhita of Acharya Sharangdhara. Madhyam Khanda, Ch. 6 Verse 16 Chaukhambha Orientalia Varanasi 175 (2011).
Mishra, P. S. N. Bhaishajya Ratnavali of Kaviraj Govind Das Sen. Ch. 16, Verse 20. Chaukhambha Surbharti Prakashan 460 (2012).
Tripathi, D. B. Charaka Samhita of Agnivesha. Chaukhambha Surbharti Prakashan, vol. 2, 883 (2013).
Patel, S. et al. High-performance thin-layer chromatographic standardization and quantification of marker compounds in an ayurvedic polyherbal formulation: Krishnadi Churna. J. Planar Chromatogr. 34(6), 493–502 (2021).
Uhlen, M. et al. A pathology atlas of the human cancer transcriptome. Science 357(6352), 2507 (2017).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2(5), 401–404 (2012).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6(269), pl1–pl1 (2013).
Daina, A., Michielin, O. & Zoete, V. SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 7(1), 42717 (2017).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13(11), 2498–2504 (2003).
Garcia-Moreno, A. et al. Functional enrichment analysis of regulatory elements. Biomedicines 10(3), 590 (2022).
Ge, S. X., Jung, D. & Yao, R. ShinyGO: a graphical gene-set enrichment tool for animals and plants. Bioinformatics 36(8), 2628–2629 (2020).
Kanehisa, M. et al. KEGG: integrating viruses and cellular organisms. Nucleic Acids Res. 49(D1), D545–D551 (2021).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28(1), 27–30 (2000).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28(11), 1947–1951 (2019).
Kanehisa, M. et al. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51(D1), D587–D592 (2023).
Raudvere, U. et al. G: profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 47(W1), W191–W198 (2019).
Szklarczyk, D. et al. The STRING database in 2023: protein–protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51(D1), D638–D646 (2023).
Kim, S. et al. PubChem 2023 update. Nucleic Acids Res. 51(D1), D1373–D1380 (2023).
Liu, T. et al. BindingDB: a web-accessible database of experimentally determined protein–ligand binding affinities. Nucleic Acids Res. 35(suppl_1), D198–D201 (2007).
Burks, H. E. et al. Discovery of an acrylic acid based tetrahydroisoquinoline as an orally bioavailable selective estrogen receptor degrader for ERα + breast cancer. J. Med. Chem. 60(7), 2790–2818 (2017).
Ghosh, D. et al. Novel aromatase inhibitors by structure-guided design. J. Med. Chem. 55(19), 8464–8476 (2012).
Yasuda, H. et al. Structural, biochemical, and clinical characterization of epidermal growth factor receptor (EGFR) exon 20 insertion mutations in lung cancer. Sci. Transl. Med. 5(216), 216 (2013).
Ylilauri, M. & Pentikäinen, O. T. MMGBSA as a tool to understand the binding affinities of filamin–peptide interactions. J. Chem. Inf. Model. 53(10), 2626–2633 (2013).
Wang, W. et al. MAPK4 promotes triple negative breast cancer growth and reduces tumor sensitivity to PI3K blockade. Nat. Commun. 13(1), 245 (2022).
Jiang, W. et al. Expression and clinical significance of MAPK and EGFR in triple–negative breast cancer. Oncol. Lett. 19(3), 1842–1848 (2020).
Aziz, S. W. & Aziz, M. H. Major signaling pathways involved in breast cancer. In Breast Cancer Metastasis Drug Resistance: Progress Prospects 47–64 (2013).
Galiè, M. RAS as supporting actor in breast cancer. Front. Oncol. 9, 1199 (2019).
Abdin, S. M. et al. Nuclear factor-κB signaling inhibitors revert multidrug-resistance in breast cancer cells. Chemico-Biol. Interact. 340, 109450 (2021).
Miricescu, D. et al. PI3K/AKT/mTOR signaling pathway in breast cancer: from molecular landscape to clinical aspects. Int. J. Mol. Sci. 22(1), 173 (2020).
Dustin, D., Gu, G. & Fuqua, S. A. ESR1 mutations in breast cancer. Cancer 125(21), 3714–3728 (2019).
Barros-Oliveira, M. C. et al. Influence of CYP19A1 gene expression levels in women with breast cancer: a systematic review of the literature. Clinics 76, e2846 (2021).
Burness, M. L., Grushko, T. A. & Olopade, O. I. Epidermal growth factor receptor in triple-negative and basal-like breast cancer: promising clinical target or only a marker? Cancer J. 16(1), 23–32 (2010).
Masuda, H. et al. Role of epidermal growth factor receptor in breast cancer. Breast Cancer Res. Treat. 136, 331–345 (2012).
Oshi, M. et al. Conflicting roles of EGFR expression by subtypes in breast cancer. Am. J. Cancer Res. 11(10), 5094 (2021).
World Health Organisation. https://www.who.int/initiatives/who-global-centre-for-traditional-medicine/ (2023).
Horneber, M. et al. How many cancer patients use complementary and alternative medicine: a systematic review and metaanalysis. Integr. Cancer Therap. 11(3), 187–203 (2012).
Yang, Z. et al. Withania somnifera root extract inhibits mammary cancer metastasis and epithelial to mesenchymal transition. PLoS ONE 8(9), e75069 (2013).
Hahm, E. R., Lee, J. & Singh, S. V. Role of mitogen-activated protein kinases and Mcl‐1 in apoptosis induction by withaferin A in human breast cancer cells. Mol. Carcinogenes. 53(11), 907–916 (2014).
Cao, X. et al. Curcumin suppresses tumorigenesis by ferroptosis in breast cancer. PLoS ONE 17(1), e0261370 (2022).
Karia, P., Patel, K. V. & Rathod, S. S. Breast cancer amelioration by Butea monosperma in-vitro and in-vivo. J. Ethnopharmacol. 217, 54–62 (2018).
Arya, R. K. et al. Anti-breast tumor activity of Eclipta extract in-vitro and in-vivo: novel evidence of endoplasmic reticulum specific localization of Hsp60 during apoptosis. Sci. Rep. 5(1), 18457 (2015).
Lai, L. et al. Piperine suppresses tumor growth and metastasis in vitro and in vivo in a 4T1 murine breast cancer model. Acta Pharmacol. Sin. 33(4), 523–530 (2012).
Waks, A. G. & Winer, E. P. J. J. Breast cancer treatment: a review. JAMA 321(3), 288–300 (2019).
Jovanović Galović, A. et al. The effects of resveratrol-rich extracts of Vitis vinifera pruning waste on HeLa, MCF-7 and MRC-5 cells: apoptosis, autophagia and necrosis interplay. Pharmaceutics 14(10), 2017–p (2022).
You, D. et al. Entelon® (Vitis vinifera seed extract) prevents cancer metastasis via the downregulation of Interleukin-1 alpha in triple-negative breast cancer cells. Molecules 26(12), 3644 (2021).
Shivhare, S. C. et al. Antioxidant and anticancer evaluation of Scindapsus Officinalis (Roxb.) Schott fruits. AYU 32(3), 388 (2011).
Matt Lam, K. W. et al. Correction: an aqueous extract of Fagonia Cretica induces DNA damage, cell cycle arrest and apoptosis in breast cancer cells via FOXO3a and p53 expression. PLoS ONE 9(7), e102655 (2014).
Bibi, Y. et al. The study of anticancer and antifungal activities of Pistacia integerrima extract in vitro. Indian J. Pharm. Sci. 74(4), 375 (2012).
Bawazeer, S. et al. Isolation of bioactive compounds from Pistacia integerrima with promising effects on reverse cancer multidrug resistance. Russ J. Bioorg. Chem. 47, 997–1003 (2021).
Blok, E. J. et al. Optimal duration of extended adjuvant endocrine therapy for early breast cancer; results of the IDEAL trial (BOOG 2006-05). J. Natl. Cancer Inst. 110(1), 40–48 (2018).
Robson, M. et al. Olaparib for metastatic breast cancer in patients with a germline BRCA mutation. N Engl. J. Med. 377(6), 523–533 (2017).
André, F. et al. Alpelisib for PIK3CA-mutated, hormone receptor–positive advanced breast cancer. N Engl. J. Med. 380(20), 1929–1940 (2019).
Zhao, H. et al. Aromatase expression and regulation in breast and endometrial cancer. J. Mol. Endocrinol. 57(1), R19–R33 (2016).
Acknowledgements
Authors thank All India Institute of Ayurveda, New Delhi, India for providing infrastructural and library support. Authors acknowledge Dr Vinod Devaraji, Schrodinger India for his support for structural analysis studies.
Author information
Authors and Affiliations
Contributions
Conceptualization: H.S, T.N.; Methodology: H.S, P.K.G; Software: H.S, P.K.G, A.K.M; Formal analysis: H.S, P.K.G; Investigation: H.S; Writing –original draft preparation: H.S, P.K.G; Writing – review, and editing.: H.S, P.K.G, R.T; Visualisation.: H.S; Supervision.: T.N., S.K.R.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
*Correspondence.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Saini, H., Gupta, P.K., Mahapatra, A.K. et al. Deciphering the multi-scale mechanism of herbal phytoconstituents in targeting breast cancer: a computational pharmacological perspective. Sci Rep 14, 23795 (2024). https://doi.org/10.1038/s41598-024-75059-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-75059-z
Keywords
This article is cited by
-
Exploring the role of pomalidomide in androgen-dependent prostate cancer: a computational analysis
Molecular Diversity (2025)