Abstract
Pancreatic ductal adenocarcinoma (PDAC) presents challenges in detecting somatic mutations due to its complex cellular composition. This study investigated the utility of patient-derived organoids (PDOs) to overcome these obstacles and enhance somatic mutation identification. Surgically resected PDAC tumors and their paired PDOs from 21 patients were examined. Whole-exome sequencing (WES) of tumor tissue, organoids, and peripheral blood mononuclear cells was performed to identify somatic mutations. Our findings demonstrate that PDOs retained about 80% of the somatic mutations from the original tumors, showing high concordance in mutation types. PDOs exhibited increased tumor purity and uncovered key driver mutations, aiding in identifying clinically relevant genomic alterations. Moreover, eight cycles of FOLFIRINOX treatment did not significantly alter the mutational landscape at the DNA level, indicating the stability of the mutational profile after therapeutic pressure in patients. In conclusion, PDOs are potentially important tools for exploring the somatic mutational landscape of PDAC. While they can reveal mutations that may be challenging to detect through traditional biopsy sequencing due to the inherently low tumor purity of PDAC, it is important to note that PDOs may not always fully recapitulate all mutations found in primary tumors. Despite this limitation, PDOs can still offer critical insights into the genomic complexities of PDAC, which is crucial for the development of personalized vaccines and therapies for this disease.
Similar content being viewed by others
Introduction
Pancreatic ductal adenocarcinoma (PDAC) is an aggressive cancer with a poor prognosis and the lowest 5-year survival rate of 12% among all other cancers1. This poor outcome is primarily due to its aggressive nature, late diagnosis, and limited treatment options. PDAC is often diagnosed at an advanced stage when it has already spread to other organs (> 80% cases), making it ineligible for surgical resection, the most effective treatment for pancreatic cancer2. The inherent difficulty in diagnosing PDAC is compounded by the dense fibrotic stroma surrounding the tumor, which acts as a barrier that obscures neoplastic cells. This stroma shields tumor cells from therapeutic agents and dilutes the percentage of neoplastic cells in biopsy samples, impeding accurate genomic profiling and hindering therapeutic targeting3,4,5.
The mutation landscape of PDAC is marked by frequent alternations in several key driver genes, including KRAS, CDKN2A, TP53, SMAD4, and ARID1A6. Understanding the effect of these mutations can provide insights into disease progression, potential therapeutic targets, and prognosis7. Despite the critical role of these mutations, PDAC is characterized by a low tumor mutational load (TMB)8, a significant factor contributing to PDAC’s limited response to immune checkpoint inhibitors (ICI)9,10,11,12. The scarcity of mutation-derived neoantigens in PDAC, due to its low TMB, leads to a less robust antigen-specific T-cell response, making immunotherapy less effective.
Considering the challenges of genomic profiling of PDAC, patient-derived organoids (PDOs) offer a promising approach to better represent the mutational landscape of primary tumors. PDOs can help address the limitations imposed by low tumor purity and the fibrotic stroma, providing a more accurate model of the primary tumor environment13. They offer a valuable platform for studying tumor biology, testing therapeutic interventions, and analyzing neoantigen profiles14,15. The goal of this study was to evaluate the utility of PDOs in reflecting the mutational landscape of PDAC by using whole-exome sequencing (WES). We compared the somatic mutations present in surgically resected tumors and their corresponding PDOs to determine how well PDOs capture and predict the genetic features of PDAC. This comparison aims to highlight the potential of PDOs as a tool for overcoming the challenges associated with traditional biopsy methods and improving our understanding of PDAC genomics.
Materials and methods
Study design
This study aimed to evaluate the accuracy of pancreatic ductal adenocarcinoma (PDAC) organoids in capturing the mutational landscape of primary tumors. We enrolled 21 patients with PDAC at Erasmus Medical Center (EMC), categorizing them based on their treatment status: 11 were untreated and 10 had undergone eight cycles of neoadjuvant FOLFIRINOX chemotherapy before the operation. Organoids were generated from resected tumor tissues, and whole-exome sequencing (WES) was performed on tumor samples, organoids, and matched blood samples to compare somatic mutations between tumor and organoid samples. In select cases, targeted panel sequencing (target-seq) was undertaken using DNA extracted from formalin-fixed paraffin-embedded (FFPE) samples using tumor regions annotated by an expert pathologist.
Patient and sample collection
Fresh tumor tissues and blood samples were collected from PDAC patients who underwent standard-of-care pancreas resection at EMC. All experimental methods, including collecting and processing tumor specimens to generate organoid cultures, followed relevant guidelines and regulations, adhering to the principles of the Declaration of Helsinki. The study protocols for tissue collection (MEC-2015-085) and organoid culture generation (MEC-2021-0325) were approved by the local medical ethics committee at EMC. All patients provided written informed consent for sample collection and use. Inclusion criteria required patients to either be treatment-naïve or have completed eight cycles of FOLFIRINOX. Patients were enrolled from 2019 to 2022, with a WHO performance status of zero or one at the time of surgery. Blood samples were mandatory for inclusion. The data cutoff date for overall survival (OS) and progression-free survival (PFS) analysis was February 23, 2024.
Following tumor resection, tissue underwent pathological evaluation, with part of each tumor embedded in paraffin for histological analysis. The remaining tissue was divided into two portions: one for snap-freezing and DNA extraction, and the other for organoid culture.
Organoid establishment
Fresh tumor specimens were collected and sectioned into fragments of approximately 2–3 mm3 in a basic media consisting of AdDMEM/F12 (Gibco), 1x Hepes (Gibco, #15630), 1x Glutamax (Gibco, #35050061), 100 µg/mL Pyromycin (InvivoGen, #ant- pm-1). These fragments underwent enzymatic dissociation using a mixture of 1 mg/mL liberase (TH) (Roche, #5401135001), 10 µg/mL DNAase I (Sigma, #4263), and 10 µM ROCK inhibitor (Tocris, #1254), for 30 min in 37 °C. Digested and isolated pancreatic cells were filtered and embedded in Matrigel (Corning, #356231) and expanded in an organoid-specific serum-free growth medium. The organoid-specific medium consists of basic media enriched with a cocktail of growth factors and supplements that included: 1mM N-Acetyl L cysteine (Sigma, #A9165), 1mM Nicotinamide (Sigma, #N0630), 1x B27 (Thermofisher, # 17504044), 100 ng/ml mouse recombinant NOGGIN (Peprotec, #250 − 38), 500 ng/mL recombinant RSPONDIN (Peprotec, #120 − 38), 30% serum-free WNT conditioned media16, 50 ng/mL mouse recombinant EGF (Gibco, #PMG8041), 100 ng/mL FGF10 ( Peprotec, #100 − 26), 10 nM Gastrin I (Sigma, #G9020), 500 nM A83-01 (Tocris, #2939), and 10 µM Y-27,632 dihydrochloride (Tocris, #1254).
DNA extraction
AllPrep DNA/RNA mini kit (QIAGEN) was used to extract DNA from organoid cultures, snap-frozen tissue samples, and peripheral blood mononuclear cells (PBMCs). Before adding lysis buffer, organoids were extracted from matrigel domes using Cell Recovery Solution (Corning, #354253). The samples were then incubated on ice for 15 min to ensure complete removal of the Matrigel. Snap-frozen tumor samples were homogenized in lysis buffer using an Ultra-Turrax T25 homogenizer (Janke & Kunkel).
Whole exome sequencing and analysis
WES was performed using the SureSelectXT reagent kit and SureSelect Clinical Research Exome V2 baits kit (Agilent Technologies) to capture targeted exome regions. DNA libraries were prepared and hybridized on the SureSelectXT Enrichment System, followed by sequencing on a NovaSeq 6000 platform with 2 × 150 bp paired-end reads.
Raw sequencing reads were processed using the nf-core Sarek pipeline (3.0.1)17,18. Low-quality bases and adapters were trimmed with fastp (0.23.2)19, followed by quality checks with FastQC (0.11.9). Reads were aligned to the GRCh38 human reference genome using the bwa-mem2 (2.2.1)20, with further processing, including duplicate marking and base quality recalibration, handled by GATK4 (4.2.6.1)21 and samtools (1.15.1)22. Sequencing quality metrics were evaluated using GATK’s HsMetric, and only patients with duplicated read percentages below 50% in all three samples (tumor, organoid, PBMCs) were included in the subsequent analyses. The analyses were conducted using in-house R scripts. Alignment and variant calling results were checked using the integrative genomics viewer (IGV, 2.15.2)23.
Calling, annotating and filtering somatic variants
Somatic variants, including single nucleotide variants (SNVs), small insertions/deletions (InDels), and multi-nucleotide variants (MNVs), were called by Mutect2 (GATK4) using the GATK best practice workflow24. Contamination and orientation bias were estimated by GetPileupSummaries, CalculateContamination, and LearnReadOrientationModel in GATK4.
Small somatic mutations were further annotated using Ensembl Variant Effect Predictor (VEP, V104)25 with plugins for functional annotations (SingleletterAA, gnomAD, and ClinVar). Only the most severe mutational consequence among all overlapping transcripts was remained in annotation results by choosing –pick when running VEP. Variants were filtered according to the following criteria: (1) pass all Mutect2 default filters (PASS-only), (2) locate in the library kit’s covered and targeted regions, (3) gnomAD exome allele frequency \(\:\le\:\) 0.001, (4) mutated position depth \(\:\ge\:\) 30, (5) mutated allele depth \(\:\ge\:\) 3, (6) variant allele frequency \(\:\ge\:\) 0.05, and (7) variant located in exon, or splice site intronic region. In addition, protein-altering small somatic mutations were further categorized into frameshift, missense, splicing, inframe insertion/deletion, stop/gain variants, and multiple independent coding mutations. Exome-wise tumor mutational burden (TMB) was calculated as the total number of small somatic mutations per mega-base (Mb) pair of the targeted exome size (38,514,517 bp).
Copy number alternations (CNAs) were called using CNVkit (0.9.9)26. Sequence-accessible regions were determined using a 100-kmer mapability file (GCA_000001405.15_GRCh38_no_alt_analysis_set_100.bw, download from PureCN), and a male reference was built from PBMC bam files. Segmentation was performed using the circular binary segmentation algorithm27, with very-low-coverage bins and outlier bins dropped, and germline mutations incorporated to improve accuracy. Finally, an absolute integer copy number (CN) was derived for each segment by call command in CNVkit with the following setting: `-m threshold -t=-1.2,-0.25,0.3,0.7 -y –center biweight –ploidy 2 --drop-low-coverage -x $gender –filter cn`. CNA burden was then calculated as the number of CNA segments with CN not equal to 2. Furthermore, the CNA status of PDAC-related genes was assigned. The thresholds were: (1) Deep deletion: genes with CN less than or equal to 0, (2) Shallow deletion: genes with CN equal to 1, (3) Shallow gain: genes with CN equal to 3, (4) Deep gain: genes with CN more than 4. Heatmaps showing CNA calls in log2 ratio were plotted using the heatmap command in CNVkit.
Estimation of tumor purity and exome ploidy
The in-silico tumor purity and ploidy levels were estimated using PureCN (2.4.0)28. The process involved utilizing the CNAs obtained from CNVkit and the somatic mutation data obtained from Mutect2 as input for the analysis.
Targeted panel sequencing and processing
Target-seq was performed on DNA extracted from FFPE samples, in which mutations in driver genes were identified in organoid cultures but not in the corresponding original tissue DNA. The targeted gene panel includes 69 cancer-associated genes29, including those frequently mutated in PDAC (Supplementary Table 1).
For targeted panel sequencing, paraffin-embedded primary tumor tissue blocks were used for DNA extraction. From these blocks, at least five 10-µm‐thick FFPE tissue sections were prepared. These tissue sections were then deparaffinized and stained with hematoxylin. An expert pathologist microscopically identified and annotated specific tumor areas, which were then manually microdissected into a solution of 5% Chelex 100 Resin (Bio‐Rad, Hercules, CA, USA) and Cell lysis solution (Promega, Madison, WI, USA). DNA extraction was achieved through proteinase K (Roch, Mannheim, Germany) digestion, involving an overnight incubation at 56 °C. DNA was then used for library creation, using the Ion AmpliSeq library Kit Plus (Thermofisher scientific), involving two multiplex PCR processes (utilizing 10ng of DNA each) according to the Ion AmpliSeq kit guidelines. The sequencing was conducted on the Ion S5XL Semiconductor sequencer (Thermofisher scientific).
For variant identification and assessment, VARIANT CALLER V7.10.0.18 (Thermofisher scientific) and human reference genome hg19 were employed. Exonic or splice site variants with a minimum 10% frequency were considered in the analysis. In the filtering process, synonymous point mutations, and mutations noted at frequencies above 1% in the 1000 genome database, were excluded as typical polymorphisms. VCF files generated by VARIANT CALLER were lifted over from hg19 to hg38 using the GATK LiftoverVcf command to compare the variant calling results with WES data.
Tumor organoid assessment
Organoid samples harboring KRAS somatic mutations encoding G12N or Q61R and those bearing identical mutations in PDAC driver genes (CDKN2A, TP53, SMAD4, and ARID1A)6 (Supplementary Table 1) in the paired tumor samples were assessed as tumor organoids. When mutations in driver genes were detected exclusively in the WES organoid data, the presence of these mutations in driver genes was validated in both tumor and organoid samples through target-seq data. The variant allele frequency (VAF) and clonal structure analyses were performed on the patients exhibiting consistent mutations in driver genes in both tumor and paired tumor organoid samples.
Clonal evolution analysis
Clonal evolution was inferred using somatic mutations from Mutect2 and CNAs from CNVkit as input for PyClone-VI (0.1.1)30. Clones with less than five mutations were excluded from clonal evolution inferring analysis. Next, a consensus clonal evolution model for each tumor and tumor organoid pair was inferred and visualized using ClonEvol (0.99.11)31.
Statistical analysis
Clinical, genomic characteristics, and VAFs of tumor and organoid samples were compared using the Fisher’s exact test, Wilcoxon rank-sum test, and paired Wilcoxon test when applicable. The concordance of mutations between tumor and organoid samples was assessed using the Jaccard index, applied across different mutation types. All statistical analyses were performed in R (4.3.1). Of note, the number of protein-altering mutations and TMB comparison analyses in Supplementary Figs. 4 and 5 were separately performed for tumors and organoids. We conducted all tests two-sided, considering a P value below 0.05 as indicative of statistical significance.
Code availability
The parameter configurations and scripts for running the nfcore Sarek pipeline to process raw sequencing data and additional R code to process, analyze and visualize the data in this study, are currently accessible exclusively upon request.
Results
Patient-derived tumor organoids provide a reliable genetic testing platform for PDAC
Tissue samples were obtained from surgical resections of 21 confirmed PDAC patients, with 10 undergoing FOLFIRINOX treatment and 11 being treatment-naïve, as detailed in Table 1 and Supplementary Table 2. Pure epithelial cancer cell cultures were established within approximately two weeks after initial tissue processing. These cultures were maintained for around four weeks, reaching passage 3 or 4, to ensure their viability and stability before undergoing genomic evaluation.
To assess whether the cultured organoids accurately represent the mutational landscape of the original tumors, we conducted whole-exome sequencing (WES) on the organoids, parental tumor tissues, and matched PBMCs from all 21 patients (as detailed in the schematic overview, Fig. 1). An initial quality assessment was performed on WES data from 63 samples, comprising 21 tumor tissues, their corresponding 21 organoids, and 21 matched PBMC samples. Of these, 95% of patients (20 out of 21) met our quality criteria across all three sample types (tissue, organoid, and PBMCs) and were included in subsequent analyses (Fig. S1). We compared the mutational status of PDAC driver genes in tumor-organoid pairs (Fig. 2A), finding KRAS mutations in 12 tumors (60%) and 11 organoids (55%), and TP53 mutations in 10 tumors (50%) and 7 organoids (35%). All missense and frameshift variants in KRAS and TP53 were classified as (likely) pathogenic by ClinVar (Supplementary Table 2). Other oncogenic genes, including KDM6A, BCOR, ARID1A, and RNF43, were shared less frequently (Fig. 2B).
Study design of mutational landscape comparison between pancreatic ductal adenocarcinoma (PDAC) tumors and their derived organoids. PDAC tumors were resected from patients after (n = 10) or without FOLFIRINOX (n = 11) neo-adjuvant therapy. Organoids were cultured and their DNA with matched tumor tissue and peripheral blood mononuclear cells (PBMCs) were subject to whole-exome sequence (WES). The histological features of the tumors were assessed, and the WES data on somatic mutational changes in the DNA was analyzed. Representative bright field images of organoid cultures from patients S14 and S16 are shown in the top panel. The middle row panel displays H&E staining of fixed and sectioned organoids, while the bottom panel illustrates H&E staining of sectioned parental primary PDAC tissues. Scale bars are 100 μm for the top bright field images and 250 μm for the middle and bottom H&E stained sections (Created with http://biorender.com/). H&E = hematoxylin and eosin stain.
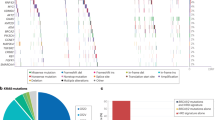
Preservation of oncogenic profile in pancreatic ductal adenocarcinoma (PDAC) organoids. Organoid models preserve the oncogenic somatic profile of PDAC tumors with genetic concordance in 50% of cases (S2, S4, S5, S6, S11, S13, S14, S16, S17, and S20). (A) Oncogenic profile of PDAC driver and PDAC-related genes showing small coding mutations and copy number alternations (CNA) status in each sample. Frequently mutated driver genes are listed on the y-axis, with the number of detected changes and their percentages in both the tumor and organoid groups. The colored inner squares indicate the type of small somatic mutations, and the colored outer frames indicate the status of CNA. For each sample and each gene, the mutational status of the gene is shown including: (1) Non-synonymous mutations; (2) Non-synonymous mutations and shallow deletion/gain; (3) Copy number deep deletion/gain. The sample names on the x-axis denote the tumor and organoid from each patient proceeded by the patient ID (i.e. S2: patient S2). The color of sample names indicates whether the paired samples show a similar oncogenic profile (in black), only tumors show PDAC oncogenes (in orange), or only organoids show PDAC oncogenes (in blue), or none of the paired samples show expected PDAC oncogenes (in purple). (B) Bar plots demonstrate the preservation of the somatic mutations. Each bar represents a gene, with the height indicating the frequency of mutations observed. The color coding presents the preservation status of the mutation, categorized as ‘Shared’ (present in both organoid and tumor), ‘Organoid’ (exclusive to organoid), or ‘Tumor’ (exclusive to the tumor). (C) The plot summarizes the concordance of oncogenic driver events at the patient level. Ten patients (50%) had the same driver mutations found in both tumors and corresponding organoids; four patients (20%) had driver mutations exclusively found in tumors, and three patients (15%) had driver mutations exclusively found in organoids. Additionally, there were three patients (15%) with no driver mutations detected in either the tumors or the organoids.
For 10 of the 20 patients, organoids and tumors had similar mutations in key driver genes. Notably, three patients (S4, S13, and S17) displayed matched ‘two-hit’ patterns in tumor suppressor genes, with consistent small mutations and copy number alternations. Patient S4 had frameshift variants and CNA deletions in ARID1A, while patients S13 and S17 had missense variants and CNA deletions in TP53, demonstrating that organoids can reflect genetic concordance and loss-of-function (LOF) mutations.
However, discrepancies were observed in the remaining 7 patients (Fig. 2A, highlighted in orange and blue on the x-axis). Three patients (S7, S9, and S15, representing 15% of the cases) had oncogenic events that were exclusively detected in the organoids. Conversely, four patients (S1, S3, S10, and S21) exhibited mutated KRAS and TP53 detected in the tumors but not in their organoid counterparts. Overall, as summarized in Fig. 2C, 65% (13 out of 20) of the PDOs were classified as tumor organoids based on WES data, with 76.9% of these showing concordance in oncogenic events with tumors. In 20% of cases (4 out of 20), organoids did not fully capture the tumor’s mutational profile, suggesting potential outgrowth of normal cells32. Additionally, for three patients, neither the tumor tissue nor the organoids exhibited detectable driver mutations, raising questions about the neoplastic nature of the original tissue materials and, consequently the derived organoids.
Mutations hidden by low tumor purity or mutation frequency are unmasked by organoids
In three patients (S7, S9, and S15), mutations in key driver genes were detected exclusively in the organoids through WES but not in their parental tumor tissues. We, therefore, questioned whether the organoids revealed mutations below the detection threshold value in the bulk tumor samples. Reviewing the alignment results in the integrative genomics viewer (IGV), no or only a few reads covered the mutated KRAS or TP53 in the parental tumor tissues (Fig. S2). Considering that DNA extraction for WES was from fresh specimens selected macroscopically by pathologists—where mutation detection might be less sensitive—we explored if using a more targeted DNA extraction method from annotated macrodissected tumor regions on formalin-fixed paraffin-embedded (FFPE) slides could confirm these mutations observed in their organoid counterpart. Thus, we extracted DNA from these annotated areas and performed targeted panel sequencing (target-seq) focusing on 69 cancer-associated genes29 (Supplementary Tables 1 and Fig. 3A). Our refined method revealed that tissue samples S9 and S15 indeed harbored the mutations in key driver genes (KRAS, TP53, and SMAD4) of PDAC, consistent with what was observed in their derived organoids through both target-seq and WES. However, for sample S7, the primary tissue lacked the PDAC mutations in driver genes even with the tumor-targeted extraction approach. In contrast, its organoid counterpart consistently manifested these mutations, as identified by both target-seq and WES (Fig. 3B).
Validation of driver mutations captured by organoids but missed by bulk tumor WES. Targeted panel sequencing (target-seq) was performed to verify the presence of driver mutations that were detected in organoid WES but not tumor WES in three patients. (A) Procedure of sample preparation for target-seq. (B) Validation of driver mutations detected in organoids but missed by tumor using target-seq. Out of the seven driver mutations detected in organoids WES, target-seq confirmed four mutations as true somatic mutations. These mutations are shown with gene SYMBOL, ENSEMBL gene ID, and protein sequence name. VAF = Variant allele frequency.
Overall, of the seven mutations found in the organoids but not in bulk WES tumors, target-seq confirmed the presence of four mutations. This underscores the value of organoid cultures in capturing and potentially amplifying key mutations that may not be detected in primary tumor samples using WES due to challenges such as tumor purity or low mutation frequency33.
Organoids provide enhanced mutational insight into the tumor genomic landscape
Our investigation delved deeper into the genomic landscape of PDAC using WES to assess the potential of PDAC PDOs in refining personalized cancer treatment and drug development. This analysis focused on the 10 patients who exhibited identical oncogenic profiles in their tumor tissues and derived organoids (Fig. 2A, highlighted in black on the x-axis).
We first profiled the total count of small mutations, including single nucleotide variants (SNVs), small insertions/deletions (InDels), and multi-nucleotide variants (MNVs) (Fig. 4A), revealing no significant difference between tumors and organoids (P = 0.053, paired Wilcoxon test; Fig. 4B). Tumor mutational burden (TMB) showed similar median values between the primary tumors (1.70 Muts/MB) and organoids (2.41 Muts/MB), with organoids displaying slightly higher TMB than tumors (P = 0.053; Fig. 4C and D). This generally low TMB in our cohort was in line with previous studies that rank pancreatic cancer among the cancer types with the lowest mutational burden34. Additionally, to evaluate mutations potentially generating neoantigens, we profiled and compared the total counts of protein-altering mutations between tumors and organoids. Organoids did not show significant differences from tumors concerning the count of protein-altering mutations (P = 0.052; Fig. 4E and F).
Enhanced sensitivity of somatic genomic alterations detection in PDAC organoids compared to tumor samples. WES data was used to explore the genomic landscape of PDAC, revealing organoid cultures’ enhanced sensitivity in capturing the mutational landscape compared to tumor samples. (A) Number of small somatic mutations, including single nucleotide variants (SNVs), small insertions/deletions (InDels), and multi-nucleotide variants (MNVs) detected in the exome region for each sample. The y-axis is depicted logarithmically, and sample types (T for tumor, O for organoid) are indicated on the top. (B) Boxplot comparing the distribution of small somatic mutations between tumor tissues (dark green) and organoid samples (yellow), corresponding to the data in panel (A). (C) Number of small somatic mutations per megabase over the targeted exome region (tumor mutational burden, TMB). (D) Boxplot comparing TMB between tumor and organoid samples, based on (C). (E) Number of small somatic mutations leading to amino acid changes (Pr-coding). (F) Boxplot comparing Pr-coding mutations between tumor and organoid samples, based on (E). (G) In-silico estimated ploidy for each sample. (H) Boxplot comparing ploidy between tumor and organoid samples, based on (G). (I) In-silico estimated tumor purity. (J) Boxplot comparing tumor purity between groups, based on (I). (K) Number of copy number alteration (CNA) segments (CNA burden). Patient IDs are shown at the bottom. (L) Boxplot comparing CNA burden between tumor and organoid samples, based on (K). (M) Summary of neoadjuvant treatments administered. Median values for both groups are displayed next to the boxplots. P-values for all boxplots are calculated using the paired Wilcoxon test.
Next, we assessed the ploidy, which indicates the number of genome copies. Ploidy did not significantly differ between tumors and organoids (median = 2.06, range 1.69–3.5 and median = 2.67, range 2.01–3.52, for tumors and organoids respectively; P = 0.26) (Fig. 4G and H). However, organoids exhibited significantly higher tumor purity compared to tumor specimens (63% tumor purity in organoids versus 30% in tumor specimens; P = 0.019; Fig. 4I and J), reflecting the predominance of cancer cells in organoid cultures. In addition, genome-wide DNA CNAs were profiled between paired samples, demonstrating a concordant pattern in the segmentation level. However, organoids displayed more pronounced CNA signals compared to tumors, which was evidenced by higher log2 ratios for amplifications and lower log2 ratios for deletions (Fig. S3). Similarly, the CNA burden, which was the count of abnormal CN segments (CN not equal to two), increased in organoids compared to tumors (P = 0.014; Fig. 4K and L).
DNA-damaging agents, like chemotherapy drugs, have the potential to trigger novel mutations and cause instability in the genome. FOLFIRINOX is a potent regimen for pancreatic cancer treatment. Thus, elucidating its effects on the mutational landscape is of paramount importance. To address this, we examined the impact of FOLFIRINOX treatment on protein-altering mutations and TMB in tumors and organoids, respectively, by comparing samples of treatment-naïve and FOLFIRINOX-treated patients (Fig. 4M). The total number of protein-altering mutations and the associated mutational burdens for tumor and organoid cohorts did not differ between the treatment-naïve and FOLFIRINOX-treated groups (Fig. S4). Additionally, we expanded the analysis to include 14 tumor samples (8 treatment-naïve, 6 FOLFIRINOX-treated) and 13 tumor organoid samples (7 treatment-naïve, 6 FOLFIRINOX-treated) with confirmed PDAC driver mutations, irrespective of whether they had matching counterparts in tumor organoids or tumor specimens. Despite this expansion, no significant shift in the number of protein-altering mutations was detected when comparing treated and non-treated samples (Fig. S5). While these findings suggest FOLFIRINOX may have minimal impact on the PDAC mutational landscape, the limited sample size and absence of matched pre- and post-treatment samples prevent a definitive conclusion. Nevertheless, our analysis showed that tumor-derived organoid cultures can provide mutational insight into the tumor genomic profile.
Organoids outperform tumors in neoantigen prediction for PDAC
We continued our assessment to focus on overall small mutations, which have the potential to generate entirely novel neoantigens35 in the same cohort of 10 patients. Comparing tumor specimens to their organoid counterparts (comparing mutations of tumors found in organoids), we observed a substantial overlap in small mutations, with a median frequency of shared mutations at 80%, ranging from 39 to 98% (Fig. 5A). Conversely, when assessing from the perspective of the organoids (mutations of organoids found in tumors), the median frequency of shared mutations dropped to 53%, ranging between 38% and 68% (Fig. 5B). The concordance between tumor and organoid mutations was shared across different mutation types, including frameshift variants (with the highest mean Jaccard index), missense variants, and stop/gain variants (Fig. S6). We hypothesized the mutations identified only in the organoids were due to the increased variant allele frequency (VAF), reflecting the proportion of DNA molecules carrying the mutations. Indeed, the VAF of both total detected (Fig. 5C) and shared mutations (Fig. 5D) between tumors and organoids significantly increased in all patients (except for total mutations in patient S17). Overall, we observed the high mutation concordance and increased VAF in organoids which is instrumental for reliable identification of potential targets for immunotherapy. This suggests that organoids provide a more accurate representation of the tumor’s mutation profile.
Intratumor heterogeneity is captured by organoids
We performed a comprehensive clonal analysis for each of the ten patients to explore the clonal dynamics in matching tumors and organoids. This analysis involved constructing clonal architectures and their trajectories based on all detected small somatic mutations and CNAs in organoids and matching tumors. We visualized the clonal trajectories using scatter plots of VAF distributions, bell plots (Fig. S7 and S8), and branch-based phylogeny trees (Fig. 6) to capture the changes in clonal architecture over time, leading up to the moment of sample collection. Our analyses revealed a consistent presence of shared ‘trunk’ mutations enriched in driver genes in both tumors and organoids. However, there were distinct ‘branch’ mutations that were unique to either tumors or their organoid counterparts (Fig. 6). This pattern of divergence from a common ancestor was consistently observed across all patients. Visual inspection of the phylogenetic trees showed similar clonal trajectories between treated and treatment-naïve patients (Fig. 6). To further quantify this observation, we used the Jaccard index to assess the similarity of trunk mutations between the two groups. The analysis showed no significant differences in the proportion of shared mutations between treated and treatment-naïve patients (Wilcoxon rank-sum test, p-value = 0.1425). These observations suggest that FOLFIRINOX treatment might not significantly affect the clonal dynamics captured by tumor organoids.
Organoids exhibit genomic concordance with tumors and unveil additional mutations. Assessing small somatic mutations in PDAC using WES revealed substantial overlap between tumors and organoids, indicating organoids’ ability to capture the majority of tumor mutations and uncover additional hidden mutations, enhancing their utility for neoantigen prediction. (A) Relative proportions of shared (pink) and tumor-specific (dark green) mutations in tumor samples. The number and percentage of shared mutations are shown on the left, with total mutations per tumor sample on the right. (B) Relative proportions of shared (pink) and organoid-specific (yellow) mutations in organoid samples. The number and percentage of shared mutations are shown on the left, with total mutations per organoid sample on the right. (C) Variant allele frequency distribution of all small somatic mutations in tumor tissue and organoid for each patient. P-values are calculated using the Wilcoxon rank sum test. (D) Paired analysis of variant allele frequency for shared mutations between each tumor and organoid pair. P-values are calculated using the paired Wilcoxon test. None = treatment-naïve; FFX = FOLFIRINOX-treated. The patients shown in each panel are the same in Fig. 4.
Clonal architectures and evolutionary paths are largely shared by matching tumors and organoids. Clonal analysis of organoids and tumors revealed that mutations were frequently shared between them but some appeared to be organoid or tumor-specific. For each patient, a colored bar represents the tumor and organoid trajectories, and the number of mutations was independently scaled by its length. The founding clone names are marked with asterisks and driver genes mutated with non-synonymous small mutations are labeled next to the bar. Individual patients are indicated by S and number while Organoid or Tumor are indicated by O or T respectively (i.e. S2_O: organoid of patient S2). (A) Patient S2. (B) Patient S4. (C) Patient S5. (D) Patient S6. (E) Patient S11. (F) Patient S13 (FOLFIRINOX-treated). (G) Patient S14 (FOLFIRINOX-treated). (H) Patient S16 (FOLFIRINOX-treated). (I) Patient S17 (FOLFIRINOX-treated). (J) Patient S20 (FOLFIRINOX-treated).
Discussion
The cellular complexity of primary PDAC tumors, especially its pronounced fibrotic stromal environment, complicates the accurate detection of somatic mutations and hinders therapeutic interventions. Historically, this has often required focusing on tumors with high neoplastic cellularity or depending on cultivated cell lines to enrich cancer cells4,5. However, our study showed that pancreatic tumor organoids retained a significant proportion of somatic mutations detected in the originating tumor tissue, with a median retention rate of 80.5%. Importantly, these organoids accurately reflected both mutation types and copy number alternations (CNAs) patterns and exhibited increased tumor purity and variant allele frequency (VAF).
In our investigation, a clear distinction between CNA burden was observed between primary tumor tissues and their corresponding organoids. While the concordance at both gene and segmentation levels was appreciable, most organoids exhibited more noticeable CNA calling results, likely due to heightened tumor purity, which enables more robust detection of CNAs. This finding positions organoids as a superior platform for comprehensive genomic profiling in pancreatic cancer. In contrast, tumor samples with low tumor purity often mask significant CNAs, limiting our ability to comprehensively profile the genomic landscape in tumors36,37. Meanwhile, signatures derived from CNAs are of great clinical potential for PDAC patients38. The increased VAF, along with improved CNA calling in tumor organoids, is particularly advantageous as it can help distinguish clonal from subclonal neoantigens.
Moreover, we report that organoids were able to identify key mutations undetectable in bulk tumor analyses using WES, in 15% of our study cases. We employed macrodissection of neoplastic cells followed by targeted panel sequencing (target-seq) to confirm these mutations in driver genes in the primary tumors. Specifically, organoids from three patients revealed hidden KRAS missense mutations, which can be used as the neoantigens targeted by modified T cell receptors, bringing continuous tumor regression39. The hidden key mutations revealed by organoids may allow researchers to develop personalized treatment strategies more accurately than traditional cell cultures or animal models. Importantly, our study highlights a critical clinical implication: negative results for known driver mutations in PDAC reported from bulk sequencing may be due to the limitations of this method. As a result, patients initially reported as negative might actually harbor actionable genomic alternations. Additionally, patients who are reported negative for key driver mutations, such as KRAS, may receive treatments to which their tumor is resistant due to an undetected KRAS mutation40. This finding underscores the necessity for comprehensive genomic profiling in PDAC to ensure accurate diagnosis and optimal treatment strategies. While our study employed WES, most hospitals typically use target-seq for PDAC diagnostics. The targeted approach focuses on a preselected panel of genes or regions known to be relevant in PDAC, allowing for deeper coverage and more sensitive detection of mutations in those areas. However, it is less effective in discovering new or rare mutations that might be important for understanding the full genetic profile of PDAC in a patient. In contrast, WES can be reanalyzed for different purposes if new information becomes relevant. Thus, the target-seq used in routine clinical practice can reveal a higher rate of known actionable mutations, but only captures a limited overall genetic landscape.
While advanced sequencing technologies offer a partial solution5, patient-derived organoids (PDOs) provide added value. They do not only facilitate the identification of somatic mutations and tumor-specific neoantigens but also serve as an in vitro platform to test the immunogenicity of these neoantigens in a co-culture system41. However, the impact of biopsy methods and culture conditions on organoids is an important consideration. Biopsy biases could affect the representation of certain cell types, and influence tumor heterogeneity. In our study, four patients had oncogenic events detectable only in tumor specimens, suggesting that their organoids might be outgrowths of normal cells. Additionally, three patients exhibited no driver mutations in either tumor tissue or organoids, raising questions about the neoplastic nature of their biopsy samples. These findings highlight the limitations of organoid models in faithfully replicating the entire genetic landscape of the original tumors. Moreover, tumor organoids require specific culture conditions that differ from the native tumor environment, which can potentially influence tumor cell expansion and alter the mutational landscape42. Further investigation is needed to understand the potential for tumor organoids to favor the outgrowth of rare subclones or accumulate mutations over time.
Assessing clonal diversity is pivotal for understanding PDAC development, heterogeneity, and treatment response. Our phylogeny analysis demonstrates that organoids and tumors share common trunk mutations reflecting key cancer-driving events, offering insights into tumorigenesis. These organoids allow researchers to study tumor evolution in a more controlled environment. However, they also exhibit unique branches, which might indicate higher intra-tumor heterogeneity in their parental tumors. These additional subclones may contribute to treatment resistance and invasiveness43. It’s important to consider that the observed higher intra-tumor heterogeneity might also result from the culturing process itself. This possibility underscores the need for careful interpretation of organoid data and the importance of validating findings with matched tumor samples whenever possible.
DNA-damaging agents, such as the FOLFIRINOX regimen are used as primary treatment for PDAC patients both in metastatic stages and as neoadjuvant therapy for those with borderline resectable or locally advanced tumors44,45. FOLFIRINOX, however, can induce genomic instability46, and such instability may increase the number of mutations, thus reshaping the neoantigen landscape47. Consequently, assessing the neoantigen landscape in tumors treated with prior chemotherapy is essential for our understanding of the genomic effect of such therapies, while this can be a challenge due to the reduced size of these treated tumors. We assessed the FOLFIRINOX regimen’s impact on the number of protein-altering mutations and the overall mutational burden and compared these results with the treatment-naïve cohort. In our previous research15, RNA sequencing revealed that organoids derived from FOLFIRINOX-treated patients exhibited a chemo-resistant pattern and heightened genomic instability, mainly due to the upregulated expression of DNA repair pathways. Such changes would theoretically result in an elevated mutational burden. However, despite administering eight cycles of FOLFIRINOX treatment, our current data do not support increased protein-altering mutations or overall mutational burden. This suggests that the observed chemo-resistance in these organoids and the primary effect of FOLFIRINOX on PDAC might not be purely genomic. Instead, they could be epigenetically driven or mediated by more intricate processes at the transcriptional or translational levels48. However, it is important to note that these findings are based on limited sample size and the absence of matched pre- and post-treatment samples, which restricts the generalizability of our conclusions. Future studies with larger, matched cohorts will be necessary for drawing more definitive conclusions.
In summary, our study underscores the advantage of PDOs in deciphering the genetic complexities of PDAC. These organoids help address issues like low tumor purity and mutation frequency, thus aiding in personalized treatment approaches even with small amounts of tumor material. Additionally, our findings indicate that FOLFIRINOX treatment does not necessarily increase the mutational burden, suggesting its effects might be mediated through non-genomic mechanisms.
Data availability
The somatic mutation data for this study have been deposited in the European Variation Archive (EVA) at EMBL-EBI under accession number PRJEB76301. The processed datasets supporting the findings are provided in this paper (Supplementary Table 2).
References
Siegel, R. L., Miller, K. D., Wagle, N. S. & Jemal, A. Cancer statistics, 2023. CA Cancer J. Clin. 73, 17–48. https://doi.org/10.3322/caac.21763 (2023).
Park, W., Chawla, A. & O’Reilly, E. M. Pancreatic cancer: A review. JAMA 326, 851–862. https://doi.org/10.1001/jama.2021.13027 (2021).
Iacobuzio-Donahue, C. A., Ryu, B., Hruban, R. H. & Kern, S. E. Exploring the host desmoplastic response to pancreatic carcinoma - Gene expression of stromal and neoplastic cells at the site of primary invasion. Am. J. Pathol. 160, 91–99. https://doi.org/10.1016/S0002-9440(10)64353-2 (2002).
Wood, L. D. & Hruban, R. H. Pathology and molecular genetics of pancreatic neoplasms. Cancer J. 18, 492–501. https://doi.org/10.1097/PPO.0b013e31827459b6 (2012).
Aguirre, A. J., Hruban, R. H., Raphael, B. J. & Network, C. G. A. R. Integrated genomic characterization of pancreatic ductal adenocarcinoma. Cancer Cell 32, 185-203.e113. https://doi.org/10.1016/j.ccell.2017.07.007 (2017).
Waddell, N. et al. Whole genomes redefine the mutational landscape of pancreatic cancer. Nature 518, 495–501. https://doi.org/10.1038/nature14169 (2015).
Hosein, A. N., Dougan, S. K., Aguirre, A. J. & Maitra, A. Translational advances in pancreatic ductal adenocarcinoma therapy. Nat. Cancer 3, 272–286. https://doi.org/10.1038/s43018-022-00349-2 (2022).
Aaltonen, L. A. et al. Pan-cancer analysis of whole genomes. Nature 578, 82–93. https://doi.org/10.1038/s41586-020-1969-6 (2020).
Robert, C. A decade of immune-checkpoint inhibitors in cancer therapy. Nat. Commun. 11, 3801. https://doi.org/10.1038/s41467-020-17670-y (2020).
Reck, M. et al. Pembrolizumab versus chemotherapy for PD-L1-positive non-small-cell lung cancer. N. Engl. J. Med. 375, 1823–1833. https://doi.org/10.1056/NEJMoa1606774 (2016).
Robert, C. et al. Pembrolizumab versus ipilimumab in advanced melanoma. N. Engl. J. Med. 372, 2521–2532. https://doi.org/10.1056/NEJMoa1503093 (2015).
Royal, R. E. et al. Phase 2 trial of single agent ipilimumab (Anti-CTLA-4) for locally advanced or metastatic pancreatic adenocarcinoma. J. Immunother. 33, 828–833. https://doi.org/10.1097/CJI.0b013e3181eec14c (2010).
Fujii, M. & Sato, T. Somatic cell-derived organoids as prototypes of human epithelial tissues and diseases. Nat. Mater. 20, 156–169. https://doi.org/10.1038/s41563-020-0754-0 (2021).
Grönholm, M. et al. Patient-derived organoids for precision cancer immunotherapy. Cancer Res. 81, 3149–3155. https://doi.org/10.1158/0008-5472.Can-20-4026 (2021).
Farshadi, E. A. et al. Organoids derived from neoadjuvant FOLFIRINOX patients recapitulate therapy resistance in pancreatic ductal adenocarcinoma. Clin. Cancer Res. 27, 6602–6612. https://doi.org/10.1158/1078-0432.Ccr-21-1681 (2021).
Mihara, E. et al. Active and water-soluble form of lipidated Wnt protein is maintained by a serum glycoprotein afamin/α-albumin. Elife 5, e11621. https://doi.org/10.7554/eLife.11621 (2016).
Garcia, M. et al. Sarek: A portable workflow for whole-genome sequencing analysis of germline and somatic variants. F1000Res 9, 63. https://doi.org/10.12688/f1000research.16665.2 (2020).
Friederike, H. et al. Scalable and efficient DNA sequencing analysis on different compute infrastructures aiding variant discovery. bioRxiv, https://doi.org/10.1101/2023.07.19.549462 (2023).
Chen, S. Ultrafast one-pass FASTQ data preprocessing, quality control, and deduplication using fastp. iMeta 2, e107. https://doi.org/10.1002/imt2.107 (2023).
Vasimuddin, M., Misra, S., Li, H. & Aluru, S. Efficient architecture-aware acceleration of BWA-MEM for multicore systems. In 2019 IEEE 33rd International Parallel and Distributed Processing Symposium (Ipdps 2019), 314–324. https://doi.org/10.1109/Ipdps.2019.00041 (2019).
McKenna, A. et al. The genome analysis toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303. https://doi.org/10.1101/gr.107524.110 (2010).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079. https://doi.org/10.1093/bioinformatics/btp352 (2009).
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative genomics viewer (IGV): High-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192. https://doi.org/10.1093/bib/bbs017 (2013).
Van der Auwera, G. A. et al. From FastQ data to high confidence variant calls: The genome analysis toolkit best practices pipeline. Curr. Protoc. Bioinform. 43, 11.10.11-11.10.33. https://doi.org/10.1002/0471250953.bi1110s43 (2013).
McLaren, W. et al. The ensembl variant effect predictor. Genome Biol. 17, 122. https://doi.org/10.1186/s13059-016-0974-4 (2016).
Talevich, E., Shain, A. H., Botton, T. & Bastian, B. C. CNVkit: Genome-wide copy number detection and visualization from targeted dna sequencing. PLoS Comput. Biol. 12, e1004873. https://doi.org/10.1371/journal.pcbi.1004873 (2016).
Olshen, A. B. et al. Parent-specific copy number in paired tumor-normal studies using circular binary segmentation. Bioinformatics 27, 2038–2046. https://doi.org/10.1093/bioinformatics/btr329 (2011).
Riester, M. et al. PureCN: Copy number calling and SNV classification using targeted short read sequencing. Source Code Biol. Med. 11, 13. https://doi.org/10.1186/s13029-016-0060-z (2016).
Van Bockstal, M. R. et al. Somatic mutations and copy number variations in breast cancers with heterogeneous HER2 amplification. Mol. Oncol. 14, 671–685. https://doi.org/10.1002/1878-0261.12650 (2020).
Gillis, S. & Roth, A. PyClone-VI: Scalable inference of clonal population structures using whole genome data. BMC Bioinform. 21, 571. https://doi.org/10.1186/s12859-020-03919-2 (2020).
Dang, H. X. et al. ClonEvol: Clonal ordering and visualization in cancer sequencing. Ann. Oncol. 28, 3076–3082. https://doi.org/10.1093/annonc/mdx517 (2017).
Berg, H. F. et al. Patient-derived organoids reflect the genetic profile of endometrial tumors and predict patient prognosis. Commun. Med. (Lond) 1, 20. https://doi.org/10.1038/s43856-021-00019-x (2021).
Cheng, J. et al. Biased influences of low tumor purity on mutation detection in cancer. Front. Mol. Biosci. 7, 533196. https://doi.org/10.3389/fmolb.2020.533196 (2020).
Chalmers, Z. R. et al. Analysis of 100,000 human cancer genomes reveals the landscape of tumor mutational burden. Genome Med. 9, 34. https://doi.org/10.1186/s13073-017-0424-2 (2017).
Katsikis, P. D., Ishii, K. J. & Schliehe, C. Challenges in developing personalized neoantigen cancer vaccines. Nat. Rev. Immunol. https://doi.org/10.1038/s41577-023-00937-y (2023).
Tiriac, H. et al. Organoid profiling identifies common responders to chemotherapy in pancreatic cancer. Cancer Discov. 8, 1112–1129. https://doi.org/10.1158/2159-8290.Cd-18-0349 (2018).
Martínez-Jiménez, F. et al. Genetic immune escape landscape in primary and metastatic cancer. Nat. Genet. 55, 820–831. https://doi.org/10.1038/s41588-023-01367-1 (2023).
Yang, J. et al. Integrated genomic and transcriptomic analysis reveals unique characteristics of hepatic metastases and pro-metastatic role of complement C1q in pancreatic ductal adenocarcinoma. Genome Biol. 22, 4. https://doi.org/10.1186/s13059-020-02222-w (2021).
Leidner, R. et al. Neoantigen T-cell receptor gene therapy in pancreatic cancer. N. Engl. J. Med. 386, 2112–2119. https://doi.org/10.1056/NEJMoa2119662 (2022).
Principe, D. R. et al. Calcium channel blockers potentiate gemcitabine chemotherapy in pancreatic cancer. Proc. Natl. Acad. Sci. USA 119, e2200143119. https://doi.org/10.1073/pnas.2200143119 (2022).
Dijkstra, K. K. et al. Generation of tumor-reactive T cells by co-culture of peripheral blood lymphocytes and tumor organoids. Cell 174, 1586-1598.e1512. https://doi.org/10.1016/j.cell.2018.07.009 (2018).
Weeber, F., Ooft, S. N., Dijkstra, K. K. & Voest, E. E. Tumor organoids as a pre-clinical cancer model for drug discovery. Cell Chem. Biol. 24, 1092–1100. https://doi.org/10.1016/j.chembiol.2017.06.012 (2017).
Le Compte, M. et al. Single-organoid analysis reveals clinically relevant treatment-resistant and invasive subclones in pancreatic cancer. NPJ Precis. Oncol. 7, 128. https://doi.org/10.1038/s41698-023-00480-y (2023).
Conroy, T. et al. FOLFIRINOX versus gemcitabine for metastatic pancreatic cancer. N. Engl. J. Med. 364, 1817–1825. https://doi.org/10.1056/NEJMoa1011923 (2011).
Conroy, T. et al. FOLFIRINOX or gemcitabine as adjuvant therapy for pancreatic cancer. N. Engl. J. Med. 379, 2395–2406. https://doi.org/10.1056/NEJMoa1809775 (2018).
Wiegmans, A. P. et al. Genome instability and pressure on non-homologous end joining drives chemotherapy resistance via a DNA repair crisis switch in triple negative breast cancer. Nar Cancer 3, zcab022. https://doi.org/10.1093/narcan/zcab022 (2021).
Mouw, K. W., Goldberg, M. S., Konstantinopoulos, P. A. & D’Andrea, A. D. DNA damage and repair biomarkers of immunotherapy response. Cancer Discov. 7, 675–693. https://doi.org/10.1158/2159-8290.Cd-17-0226 (2017).
Swayden, M., Iovanna, J. & Soubeyran, P. Pancreatic cancer chemo-resistance is driven by tumor phenotype rather than tumor genotype. Heliyon 4, e01055. https://doi.org/10.1016/j.heliyon.2018.e01055 (2018).
Acknowledgements
We acknowledge the Department of Pathology Laboratory for Molecular Diagnostics for performing targeted panel sequencing.
Author information
Authors and Affiliations
Contributions
EAF and WW designed and performed all experiments, analyzed datasets, and wrote the manuscript; FM and EO contributed with technical assistance during the study; MD contributed with all the pathological assessments and supported the collection of patient materials; CHJE, HJGW, and PDK supervised the study, wrote and reviewed the manuscript. All the co-authors commented and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests. The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Farshadi, E.A., Wang, W., Mohammad, F. et al. Tumor organoids improve mutation detection of pancreatic ductal adenocarcinoma. Sci Rep 14, 25468 (2024). https://doi.org/10.1038/s41598-024-75888-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-75888-y
Keywords
This article is cited by
-
Organoids in translation: a bench-to-bedside framework for pancreatic cancer precision medicine
Journal of Translational Medicine (2026)
-
Organoid analytical toolkits
Nature Reviews Bioengineering (2026)