Abstract
Overcoming resistance to epidermal growth factor receptor tyrosine kinase inhibitors, including osimertinib, is urgent to improve lung cancer treatment outcomes. Extracellular vesicle (EV)-derived microRNAs (EV-miRNAs) play important roles in drug resistance and serve as promising biomarkers. In this study, we aimed to identify EV-miRNAs associated with osimertinib resistance and investigate their clinical relevance. The release of excess EVs was confirmed in the osimertinib-resistant lung adenocarcinoma cell line PC9OR. The exposure of PC9OR-derived EVs and EV-miRNAs to PC9 cells increased cell viability after osimertinib treatment. Microarray analysis revealed that miR-130a-3p was upregulated in EVs derived from PC9OR cells and another osimertinib-resistant cell line (H1975OR). Transfection with miR-130a-3p attenuated osimertinib-induced cytotoxicity and apoptosis in both PC9 and H1975 cells, whereas osimertinib resistance in PC9OR cells was reversed after miR-130a-3p inhibition. Bioinformatics analysis revealed that runt-related transcription factor 3 is a target gene of miR-130a-3p, and it induced osimertinib resistance in PC9 cells. Patients with lower baseline serum miR-130a-3p concentrations had longer progression-free survival. miR-130a-3p is a potential therapeutic target and a predictive biomarker of osimertinib resistance in adenocarcinomas.
Similar content being viewed by others
Introduction
The principal cause of mortality associated with malignancy is lung cancer. It accounts for approximately 1.8 million deaths globally each year and has a 5-year survival rate of 4–17%1,2,3. Approximately 85% of patients with lung cancer are diagnosed with non-small-cell lung cancers (NSCLCs) such as adenocarcinoma4. Activating mutations in the epidermal growth factor receptor (EGFR) gene is among the most prevalent and targetable genetic aberrations. These mutations are mainly observed in patients with adenocarcinoma, those who have never smoked, and those of Eastern Asian ethnicity5. EGFR tyrosine kinase inhibitors (EGFR-TKIs) are the main treatments for advanced NSCLC and are used for postoperative adjuvant therapy for patients with NSCLC with EGFR mutations6,7. Osimertinib, a third-generation EGFR-TKI, has been approved for the treatment of NSCLC with both EGFR-sensitizing and T790M resistance mutations8. Osimertinib, compared with first and second-generation EGFR-TKIs, clinically improved the progression-free survival (PFS) of patients with EGFR-mutant NSCLC in a phase III clinical trial with acceptable toxicity9,10. Despite these clinical advantages, several patients eventually develop resistance to osimertinib, resulting in disease progression and poor outcomes11. Therapies for osimertinib-resistant NSCLC, however, remain to be established. Therefore, elucidating the mechanisms underlying resistance and developing effective strategies are crucial for the success of osimertinib therapy.
Extracellular vesicles (EVs) are small membrane-bound vesicles released by various cells that transport bioactive molecules such as proteins, lipids, and nucleic acids12,13. They are important mediators of intercellular communication, especially in cancer14. There has been increasing interest in the contribution of EV-derived microRNAs (EV-miRNAs) to tumor development and associated drug resistance15,16. miRNAs are key regulators with diverse roles in various physiological and developmental processes, including cell development, differentiation, apoptosis, and proliferation17,18. Several studies have demonstrated that certain EV-miRNAs are associated with resistance to first and second-generation EGFR-TKIs19,20,21,22. However, the involvement of EVs and EV-miRNAs in the development of resistance to third-generation EGFR-TKIs has not been established.
In the present study, we aimed to establish novel osimertinib-resistant lung adenocarcinoma cell lines and assess the functions of EVs and EV-miRNAs derived from them; identify a promising miRNA involved in the development of osimertinib resistance and its target genes; and elucidate the relationship between serum miRNA levels and clinical outcomes in patients with NSCLC treated with osimertinib.
Results
Establishment of osimertinib-resistant lung adenocarcinoma cell lines
The osimertinib-resistant cell lines PC9OR and H1975OR were established from the lung adenocarcinoma cell lines PC9 and H1975, respectively. These were used to investigate the effect of EVs on osimertinib resistance in lung adenocarcinoma cell lines. The IC50 values for osimertinib in the PC9 and PC9OR cells were 0.04 µM (95% confidence interval [CI]: 0.02–0.08) and 2.07 µM (95% CI: 1.43–3.00), and those for the H1975 and H1975OR cells were 0.91 µM (95% CI: 0.43–1.62) and 4.30 µM (95% CI: 3.21–5.79), respectively (Supplementary Figs. S1a and S1b). Furthermore, apoptosis was less extensive at 48 and 72 h after osimertinib treatment in the PC9OR than in the PC9 cells (Supplementary Fig. S1c). Next, we identified the key factors associated with osimertinib resistance in PC9OR cells. We did not observe any mutations in EGFR exons 18–21 (including T790M and C797S) and Met overexpression, but the levels of expression of phospho-Akt (P-Akt) were significantly higher in the PC9OR than in the PC9 cells (Supplementary Figs. S1d-S1f).
Evaluation of the characteristics of EVs secreted from PC9 and PC9OR cells in the medium
We confirmed the characteristics of EVs secreted from PC9 and PC9OR cells to the culture medium using transmission electron microscopy (TEM). Representative micrographs of the EVs derived from the PC9 and PC9OR cells are shown in Fig. 1a. The isolated EVs were round cup-shaped and had a lipid bilayer. Their approximate diameter was 100 nm, which was within the reported small EV size range of 50–130 nm12. In addition, the expressions of the exosomal markers CD9 and CD81 in EVs isolated from the PC9 and PC9OR cells were validated through immunoblotting (Fig. 1b). The particle diameters of the EVs derived from the PC9 and PC9OR cells peaked at approximately 100 nm, and the particle concentration in the PC9OR cells was higher than that in the PC9 cells (Fig. 1c). The protein concentration and number of CD63- and CD9-positive EVs secreted by the PC9OR cells were significantly higher than those of the EVs secreted by the PC9 cells (Fig. 1d and e).
Characteristics of EVs secreted by PC9 and PC9OR cells in the medium (a) TEM images of isolated EVs from PC9 and PC9OR cells. The black sphere image shows 40-nm gold colloids bound to the CD9 antibody. Scale bars:100 nm. (b) Protein concentrations of CD9 and CD81 (exosomal markers) in isolated EVs from PC9 and PC9OR cells. Original blots are presented in Supplementary Figure S4a. These images show that EVs were isolated (a, b). (c) Size distribution of isolated EVs from PC9 and PC9OR cells. (d) Protein concentrations of isolated total EVs from PC9 and PC9OR cells. (e) Number of CD63- and CD9-positive EVs from PC9 and PC9OR cells. The release of EVs differed in the PC9 and PC9OR cells (c–e). Data are presented as mean ± SEM (n = 3–4) (d, e). *P < 0.05, compared with PC9 cells. P-values were determined using Student’s t-test. EVs, extracellular vesicles; TEM, transmission electron microscopy.
Effect of EVs and EV-miRNAs on osimertinib resistance in PC9 cells
We visualized the incorporation and internalization of EVs isolated from PC9OR cells (PC9OR-EVs) in PC9 cells following their incubation with labeled PC9OR-EVs for 24 h. Fluorescence microscopy showed intracellular red fluorescence (Fig. 2a). The uptake and internalization of PC9OR-EVs into PC9 cells were similar to those of EVs isolated from PC9 cells (PC9-EVs). The effect of PC9OR-EVs on osimertinib resistance was ascertained after incubating PC9 cells with PC9-EVs or PC9OR-EVs for 24 h. As shown in Fig. 2b, exposure to PC9OR-EVs, but not PC9-EVs, significantly enhanced osimertinib resistance in PC9 cells, whereas the viability of PC9 cells in the absence of osimertinib was not affected by exposure to PC9OR-EVs. Following the extraction of miRNAs from PC9-EVs and PC9OR-EVs to elucidate their role in osimertinib resistance, we found that the total concentration of miRNA in PC9OR-EVs was markedly higher than that in PC9-EVs (Fig. 2c). The transfection of miRNAs derived from PC9OR-EVs did not affect the viability of PC9 cells in the absence of osimertinib but increased it in the presence of osimertinib (Fig. 2d).
Effects of PC9OR-EV- and PC9OR-EV-derived miRNAs on osimertinib resistance in PC9 cells (a) Fluorescence images of PC9 cells after exposure to red-labeled EVs from PC9 (PC9-EVs) and PC9OR cells (PC9OR-EVs). Scale bars: 50 μm. (b) Viability of PC9 cells treated with 5 µg of PC9-EVs and PC9OR-EVs in the absence or presence of 1 µM of osimertinib for 72 h. The controls are PC9 cells treated with PBS (vehicle). (c) The total concentration of miRNA derived from PC9-EVs and PC9OR-EVs. (d) Viability of PC9 cells transfected with 6–12 nM of miRNAs derived from PC9OR-EVs in the absence or presence of 1 µM osimertinib for 72 h. Data are presented as mean ± SEM (n = 3–6) (b–d). *P < 0.05, compared with control or PC9 cells. P-values were determined using Student’s t-test (c) or ANOVA with Tukey–Kramer (b, d). ns, no significant difference. EVs, extracellular vesicles; ANOVA, analysis of variance.
Identification of EV-miRNAs associated with osimertinib resistance in lung adenocarcinoma cell lines
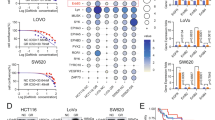
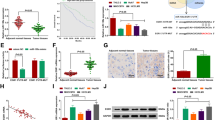
We evaluated the differential expression of EV-miRNAs in PC9 and PC9OR cells via miRNA microarray analysis and identified the candidate EV-miRNAs affecting osimertinib resistance. A total of 6,599 miRNAs were detected in each sample (Fig. 3a). Figure 3b shows a heat map of the expression of the top 22 EV-miRNAs, demonstrating a 3.5-fold change between PC9 and PC9OR cells. Of these 22 miRNAs, the expression levels of 12 were validated via reverse transcription-quantitative polymerase chain reaction (RT-qPCR). The expression levels of six miRNAs were significantly upregulated in PC9OR-EVs relative to PC9-EVs (Fig. 3c). They included miR-378a-3p, miR-130a-3p, miR-424-3p, miR-18a-5p, miR-378f, and miR-20b-5p. Among these, only miR-130a-3p demonstrated increased expression in the EVs isolated from H1975OR cells (Fig. 3d). In addition, the intracellular concentrations of miR-130a-3p were significantly higher in PC9OR than in PC9 cells (Fig. 3e).
Identification of EV-miRNAs associated with osimertinib resistance in lung adenocarcinoma cell lines (a) Scatter plot for the miRNA array analysis of isolated EVs from PC9 and PC9OR cells. (b) A heat map illustrating the expression of 22 EV-miRNAs and demonstrating a 3.5-fold change between PC9 and PC9OR cells. (c) Relative EV-miRNA expression levels in PC9 and PC9OR cells. (d) Relative EV-miRNA expression levels in H1975 and H1975OR cells. (e) Relative intracellular miRNA levels in PC9 and PC9OR cells. miRNA levels were normalized using U6 levels determined by RT-qPCR (c–e). Data are presented as mean ± SEM (n = 5–6). *P < 0.05, compared with PC9 or H1975 cells. P-values were determined using Student’s t-test. EVs, extracellular vesicles; RT-qPCR, reverse transcription-quantitative polymerase chain reaction.
Effect of miR-130a-3p on osimertinib resistance in lung adenocarcinoma cell lines
The relative miR-130a-3p concentrations were markedly higher in PC9 and H1975 cells after transfection with an miR-130a-3p mimic than those in cells transfected with the negative control (NC) (Fig. 4a and b). The cell viability assay showed that osimertinib resistance in PC9 and H1975 cells significantly increased following transfection with miR-130a-3p mimics (Fig. 4c and d). In contrast, PC9 cells transfected with an miR-18a-5p or miR-20b-5p mimic did not show enhanced osimertinib resistance (Supplementary Fig. S2). In addition, a significant reduction in apoptosis was observed in PC9 and H1975 cells after transfection with miR-130a-3p mimics at 48 and 72 h in the presence of osimertinib (Fig. 4e and f).
Effect of miR-130a-3p on osimertinib resistance in lung adenocarcinoma cell lines (a, b) Relative miR-130a-3p levels in PC9 and H1975 cells transfected with a negative control (NC) and an miR-130a-3p mimic (130a mimic). The miR-130a-3p concentrations were normalized using U6 levels relative to the levels in the cells transfected with the NC. (c, d) Viability of PC9 and H1975 cells transfected with the NC and miR-130a-3p mimic after exposure to various concentrations of osimertinib for 72 h. (e, f) Apoptotic rate in PC9 and H1975 cells after transfection with the NC and miR-130a-3p mimic at 24, 48, and 72 h in the absence or presence of 1 µM osimertinib. (g) Relative miR-130a-3p concentrations in PC9 and PC9OR cells transfected with the NC and miR-130a-3p inhibitor (130a inhibitor). The miR-130a-3p concentrations were normalized using U6 levels relative to the levels in PC9 cells transfected with the NC. (h) Cell viability in PC9 and PC9OR cells transfected with the NC and 130a inhibitor in the absence or presence of 1 µM osimertinib for 72 h. Cell viability is the percentage of viable cells relative to PC9 cells transfected with the NC in the absence of osimertinib. When the standard errors of the means are small, they are contained within the symbols. Data are presented as mean ± SEM (n = 3–12). *P < 0.05, compared with NC. P-values were determined using Student’s t-test (a, b) or ANOVA with Tukey–Kramer (c–h). RLUs, relative luminescence units; ANOVA, analysis of variance.
Further, the relative concentrations of miR-130a-3p in PC9 and PC9OR cells transfected with an miR-130a-3p inhibitor were significantly lower than those in cells transfected with NC (Fig. 4g). The downregulation of miR-130a-3p did not affect the viability of PC9 or PC9OR cells in the absence of osimertinib. However, the viability of PC9OR cells transfected with the 130a-3p inhibitor decreased in the presence of osimertinib (Fig. 4h).
Identification of miR-130a-3p target genes
Of the top 300 predicted genes in the miRDB and TargetScan databases ranked based on computational methods, 148 targets of miR-130a-3p were overlapping. We narrowed down the candidate target genes of miR-130a-3p via further bioinformatics analyses using the miRTarBase database (experimentally validated microRNA-target interaction database), and four overlapping targets (ATG16L1, PPARG, runt-related transcription factor 3 [RUNX3], and BMPR2) were selected (Fig. 5a). After manually searching and reviewing the available literature articles on these four target genes, we identified RUNX3 as the only recognized tumor suppressor gene in several cancers23. We confirmed that the protein concentrations of RUNX3 were significantly lower in PC9OR than in PC9 cells (Fig. 5b and c). As shown in Fig. 5d, the protein concentration of RUNX3 decreased after transfection with the miR-130a-3p mimic. The RUNX3 mRNA concentrations were significantly higher in PC9 cells transfected with the miR-130a-3p inhibitor than in those transfected with the NC (Fig. 5e). The protein concentrations of RUNX3 in both PC9 and PC9OR cells increased with the transfection of miR-130a-3p inhibitor (Fig. 5f). Further, the protein concentration of RUNX3 decreased following its knockdown (Fig. 5h), as confirmed by RT-qPCR (Fig. 5g). The cell viability assay also showed that RUNX3 knockdown increased osimertinib resistance in PC9 cells (Fig. 5i).
Effect of RUNX3 on osimertinib resistance in PC9 cells (a) Identification of the target genes of miR-130a-3p using miRDB, TargetScan, and miRTarBase databases. (b, c) Protein concentrations of RUNX3 in PC9 and PC9OR cells normalized with β-actin. (d) Protein concentrations of RUNX3 and β-actin in PC9 cells transfected with the negative control (NC) and the miR-130a-3p mimic (130a mimic). (e) Relative mRNA expressions of RUNX3 in PC9 cells transfected with NC and miR-130a-3p inhibitor (130a inhibitor). (f) Protein concentrations of RUNX3 and β-actin in PC9 and PC9OR cells transfected with the NC and the 130a inhibitor. (g) Relative mRNA expression of RUNX3 in PC9 cells transfected with NC and si-RUNX3. The RUNX3 mRNA levels were normalized using the glyceraldehyde-3-phosphate dehydrogenase concentrations relative to the levels in PC9 cells transfected with the NC (e, g). (h) Protein concentrations of RUNX3 and β-actin in PC9 cells transfected with si-RUNX3. (i) Viability of PC9 cells transfected with si-RUNX3 in the absence or presence of 1 µM osimertinib for 72 h. Data are presented as mean ± SEM (n = 3–6). *P < 0.05, compared with PC9 cells or NC. P-values were determined using Student’s t-test (c, e, g) or ANOVA with Tukey–Kramer (i). Original blots are presented in Supplementary Figures S4b-d. ANOVA, analysis of variance.
Analysis of the relationship between serum miR-130a-3p concentrations and clinical outcomes in patients with NSCLC treated with osimertinib
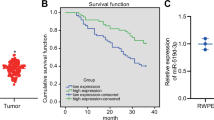
The characteristics of the 12 enrolled patients are summarized in Table 1. Their median age was 70 years (range, 38–84 years). The pathological type was lung adenocarcinoma for all patients. The performance status for the initial treatment ranged from 0 to 2. Six patients had an EGFR L858R mutation, five had an exon 19 deletion, and one had an L861Q mutation. Seven patients had never smoked, and five were former smokers. Figure 6a and b show the relationship between the pre-treatment serum miR-130a-3p concentration and PFS of patients with NSCLC receiving osimertinib. The PFS and time to treatment failure were longer for patients with low (n = 6) than for those with high (n = 6) serum miR-130a-3p concentrations (Supplementary Figs. S3a and S3b).
Relationship between serum miR-130a-3p concentration and progression-free survival of patients with NSCLC treated with osimertinib (a) Relative serum miR-130a-3p concentrations in patients with NSCLC. The miR-130a-3p expression indicates the values calculated using the 2−ΔΔCT method and normalized using U6 levels. (b) Progression-free survival of patients with NSCLC treated with first-line osimertinib. Arrows indicate ongoing treatment. * transfer to different institution; # death; AE, discontinuation of treatment due to adverse event; PD, progressive disease; NSCLC, non-small-cell lung cancer.
Discussion
The role of EV-miRNAs in the development of osimertinib resistance in lung adenocarcinoma cells has not been established. In the present study, we analyzed the function of EVs released from osimertinib-resistant lung adenocarcinoma cell lines and demonstrated that EV-miRNAs, particularly miR-130a-3p, induce osimertinib resistance by suppressing RUNX3. Furthermore, serum miR-130a-3p concentrations in patients with NSCLC treated with osimertinib were negatively correlated with PFS. To the best of our knowledge, this is the first study to report the involvement of miR-130a-3p in osimertinib resistance in lung adenocarcinoma cells and patients with NSCLC. Our findings suggest that miR-130a-3p may serve as a promising biomarker and novel therapeutic target for osimertinib resistance.
Previous studies have reported that miR-130a-3p has complex functions in cell proliferation, apoptosis, and susceptibility to drugs, including cisplatin and gemcitabine24,25,26,27. Moreover, the expression of miR-130a-3p is aberrant in several types of cancers, including NSCLC28,29. High expression of miR-130a-3p has been associated with tumor migration, invasion, and proliferation, and it is a potential diagnostic biomarker for several cancers30,31. Despite the significance of miR-130a-3p in tumorigenesis, its role in osimertinib resistance in patients with lung adenocarcinoma has not been fully investigated. In this study, we demonstrated that EV-derived miR-130a-3p was markedly upregulated in PC9OR and H1975OR cells relative to that in PC9 and H1975 cells (Fig. 3c and d). Increased osimertinib resistance in PC9 and H1975 cells was verified by transfection with an miR-130a-3p mimic (Fig. 4c and d). The sensitivity of PC9OR cells to osimertinib was restored upon transfection with an miR-130a-3p inhibitor (Fig. 4h). However, multiple mechanisms are associated with osimertinib resistance32. Osimertinib resistance in PC9OR cells was not completely suppressed by the miR-130a-3p inhibitor (Fig. 4h). This suggests that other mechanisms, such as the activation of bypass signaling associated with phosphorylated Akt (Supplementary Figs. S1e and S1f), may be involved in the development of osimertinib resistance. Nevertheless, our findings suggest that miR-130a-3p may contribute to the development of osimertinib resistance in PC9 and H1975 cells.
A recent study has revealed that miRNAs significantly influence lung cancer prognosis by targeting tumor suppressor genes (TSGs)33. RUNX3, which is recognized as a TSG, demonstrates pleiotropic anti-tumorigenic effects by inhibiting cell proliferation, migration, and invasion and inducing cell cycle arrest and apoptosis23,34. Lee et al. demonstrated that RUNX3 expression is significantly downregulated in NSCLC tissues, indicating that RUNX3 may be involved in NSCLC progression35. The previous study reported that miR-130a-3p promoted the proliferation and inhibited the apoptosis of cervical cancer cells by directly targeting the 3’-untranslated regions of RUNX336. miR-130a has also been reported to directly inhibit RUNX3 and induce cisplatin resistance by activating WNT/β-catenin signaling in hepatocellular carcinoma37. Based on these reports, we focused on RUNX3 as the target gene of miR-130a-3p. The protein concentrations of RUNX3 were significantly lower in the PC9OR than in the PC9 cells (Fig. 5b and c). They were also decreased by the transfection of the miR-130a-3p mimic in PC9 cells (Fig. 5d) and increased by the transfection of the miR-130a-3p inhibitor in PC9OR cells (Fig. 5f), respectively. In addition, osimertinib resistance in PC9 cells was significantly increased by transfection with the miR-130a-3p mimic or si-RUNX3 (Figs. 4c and 5i). These findings suggest that miR-130a-3p induces osimertinib resistance in lung adenocarcinoma cell lines by suppressing RUNX3. A limitation of the present in vitro study is the lack of RUNX3 overexpression experiments. Nevertheless, this is the first report demonstrating an association between miR-130a-3p/RUNX3 signaling and EGFR-TKI resistance, to the best of our knowledge. In contrast, several previous reports have indicated a negative correlation between RUNX3 and Akt signaling38,39,40. In the present study, decreased RUNX3 (Fig. 5b and c) and increased P-Akt (Supplementary Figs. S1e and S1f) expression levels in PC9OR cells were observed. The reduction of RUNX3 expression may contribute to osimertinib resistance in lung adenocarcinoma cell lines through the upregulation of P-Akt. Our preliminary experiments demonstrated that the miR-130a-3p mimic or si-RUNX3 did not affect P-Akt expression in PC9 cells, but further investigation is needed to elucidate the mechanisms regarding the impact of RUNX3 on EGFR-TKI resistance. Regarding target genes other than RUNX3, it has been reported that miR-130a-3p regulates the mammalian target of the rapamycin (mTOR) pathway by suppressing tuberous sclerosis 1 (TSC1)41. The bioinformatics analyses in the present study indicated that peroxisome proliferator-activated receptor gamma (PPARG), which inhibits the proliferation of EGFR-TKI-resistant lung adenocarcinoma cells via the PPARG/PTEN/Akt pathway, was another candidate target gene of miR-130a-3p42. Further studies are required to evaluate the impact of other target genes of miR-130a-3p on osimertinib resistance.
Given that EV-derived miRNAs are encapsulated within EVs and escape degradation by nucleases through their conjugation with the Argonaute protein, miRNAs exhibit stability and can be detected in plasma, blood, urine, saliva, and various other body fluids13. Thus, miRNAs are promising diagnostic and prognostic biomarkers for human diseases43,44,45. Several studies have demonstrated that some miRNAs (miR-19a, miR-153-3p, miR-206, and miR-214) are associated with acquired resistance to first- and second-generation EGFR-TKIs19,20,21,22, but their involvement in resistance to third-generation EGFR-TKIs has not been established. In the present study, we evaluated the clinical utility of miR-130a-3p as a biomarker in 12 patients with NSCLC receiving osimertinib. As shown in Fig. 6, PFS was longer for patients with low than for those with high serum miR-130a-3p concentrations. This indicates that serum miR-130a-3p concentrations are potential biomarkers for osimertinib resistance in lung adenocarcinoma.
This clinical study had some limitations. First, the rate of response to osimertinib was not evaluated in the present study. Second, the sample was small. Third, the PFS data for many cases were censored, and it was not possible to analyze serum samples after the development of osimertinib resistance (i.e., after disease progression). Therefore, large-scale studies should be conducted to assess the relationship between serum miR-130a-3p concentrations and osimertinib efficacy in patients with NSCLC.
In conclusion, the findings of this study suggest that miR-130a-3p induces osimertinib resistance in lung adenocarcinoma cells, at least partly, by negatively regulating the expression of its direct target gene RUNX3. In addition, miR-130a-3p may serve as a biomarker and novel therapeutic strategy for osimertinib resistance in lung adenocarcinoma.
Methods
Ethical considerations
The research protocols for this study were approved by the Ethics Committee of Osaka University Hospital (approval number: 22152). The study was conducted in accordance with the Declaration of Helsinki. All patients provided written informed consent for a comprehensive study utilizing the serum bank of the Department of Respiratory Medicine and Clinical Immunology at the Graduate School of Medicine, Osaka University (approval number: 11122-9). This study utilized frozen serum samples from patients who consented to the secondary use of the serum bank. Information about this study was made publicly available on the website (https://www.med.osaka-u.ac.jp/pub/hp-drug/pharm/research/), and all patients were guaranteed the opportunity to opt out at any time.
Materials
Osimertinib was obtained from Selleck Chemicals (Houston, TX, USA). The primary antibody used for the immunoblotting analyses was RUNX3 (R3-5G4) obtained from Santa Cruz Biotechnology (Santa Cruz, CA, USA). Antibodies against Akt (#9272), phospho-Akt (Ser473) (#9271), Met (#8198), EGFR (#4267), and β-actin (#4970) were purchased from Cell Signaling Technology (Danvers, MA, USA). CD9 (12A12) and CD81 (12C4) antibodies were purchased from Cosmo Bio (Tokyo, Japan). All other chemicals used were of the highest available purity.
Cell culture
PC9 cells (EGFR exon 19 deletion/E746-A750) and H1975 cells (EGFR L858R/T790M), which are human lung adenocarcinoma cell lines, were purchased from the Riken BioResource Center (Tsukuba, Japan) and American Type Culture Collection (ATCC, VA), respectively. Both cell lines were cultured in RPMI 1640 medium (FUJIFILM WAKO Chemicals, Osaka, Japan) supplemented with 10% fetal bovine serum (FBS) (Bovogen Biologicals, Australia) and incubated at 37 °C in humidified air with 5% CO2.
Establishment of osimertinib-resistant lung adenocarcinoma cell lines
The osimertinib-resistant PC9OR and H1975OR cells were established by exposing PC9 and H1975 cells to increasing concentrations of osimertinib. In brief, PC9 and H1975 cells were exposed to osimertinib with a stepwise escalation from 10 nM to 1 µM over 6 months. The PC9OR and H1975OR cells were routinely cultured with a medium containing 1 µM osimertinib. Osimertinib resistance was determined by cell viability assay every month.
EV isolation
The PC9 and PC9OR cells were seeded in 100-mm dishes and cultured in RPMI 1640 medium with exosome-depleted FBS (SBI; System Biosciences, CA) for 7 days. The EVs were isolated from the culture medium using ExoQuick precipitation solution according to the manufacturer’s protocol. EV pellets were diluted in phosphate-buffered saline (PBS) or other suitable diluent for the subsequent experiments and stored at -80 °C. The EV protein concentrations were determined using the Pierce BCA Protein Assay (Thermo Fisher Scientific).
Evaluation of the characteristics of EVs using TEM
EVs were identified using TEM according to previously reported methods46. In brief, EV samples (1 µg) isolated from PC9 and PC9OR cells were placed onto formvar/carbon-coated nickel grids for 1 h and fixed in 2% paraformaldehyde in 0.1 M phosphate buffer (pH 7.4) for 10 min after removing the solution on the nickel grids. The grids were incubated with a CD9 primary antibody for 1 h, followed by incubation with 40-nm colloidal gold-affinipure goat anti-mouse IgG (Jackson ImmunoResearch, PA, USA) as a secondary antibody for 1 h. The grids were fixed with 2.5% glutaraldehyde in 0.1 M phosphate buffer for 10 min and negatively stained with 2% uranyl acetate for 15 min. Finally, the grids were embedded in 0.13% methylcellulose with 0.4% uranyl acetate for 10 min and examined at 80 kV using an HT7800 transmission electron microscope (Hitachi, Tokyo, Japan).
Nanoparticle measurement
The size and concentration of EVs obtained from the culture medium of PC9 and PC9OR cells were determined using NanoSight LM10 (Quantum Design Japan, Tokyo, Japan). Data were analyzed using the Nanoparticle Tracking Analysis software (NanoSight Ltd., Malvern, UK; Version 2.3, Build 0034). CD63/CD9-positive EVs were assayed using the ExoCounter CD63/CD9 Disc Kit (JVCKENWOOD, Yokohama, Japan), according to the manufacturer’s protocol.
Fluorescence images of PC9 cells after exposure to red-labeled EVs from PC9OR cells
EVs obtained from the culture medium of PC9OR cells were fluorescently stained using the ExoGlow-Membrane EV Labeling Kit (System Biosciences, CA, USA) according to the manufacturer’s protocol. PC9 cells were incubated with red-labeled PC9OR-EVs (5 µg) for 24 h. The samples were observed under a fluorescence microscope (BZ-X800; KEYENCE, Osaka, Japan).
Microarray analysis
Biotin-labeled samples for miRNA microarray analysis were prepared with EV-miRNAs (400 ng) obtained from the culture medium of PC9 and PC9OR cells using the FlashTag Biotin HSR RNA Labeling Kit (Thermo Fisher Scientific). Hybridization was performed using GeneChip miRNA 4.0 Array (Thermo Fisher Scientific) according to the manufacturer’s protocol. Array scanning was performed using the GeneChip Scanner 3000 7G (Thermo Fisher Scientific).
Small RNA transfection
Transfection of an miR-130a-3p mimic (mirVana miRNA mimic, Invitrogen), an miR-130a-3p inhibitor (mirVana miRNA inhibitor, Invitrogen), si-RUNX3 (Silencer Select siRNAs, Invitrogen), or an NC (mirVana miRNA mimic Negative Control #1, Invitrogen or Silencer Select Negative Control #1 siRNA, Invitrogen) was performed using the Lipofectamine RNAi-MAX transfection reagent (Invitrogen) and Opti-MEM Reduced Serum Medium (Gibco, Grand Island, NY) according to the manufacturer’s instructions. Transfected cells were harvested after the indicated incubation periods for RT-qPCR, cell viability assays, apoptosis assays, and immunoblotting.
mRNA quantification
Total RNA was extracted from the PC9 cells transfected with si-RUNX3 using the TRIzol reagent (Invitrogen) according to the manufacturer’s protocol. The RNA concentration was measured using a NanoDrop ND-1000 spectrophotometer (Thermo Fisher Scientific). Reverse transcription was performed using PrimeScript RT Master Mix (Takara Bio), and RT-qPCR was performed using TB Green Premix Ex Taq II (Takara Bio). The reaction was conducted on a QuantStudio 5 Real-Time PCR System (Thermo Fisher Scientific) using primers purchased from Invitrogen (Supplementary Table S1). The target gene expression was normalized to that of glyceraldehyde-3-phosphate dehydrogenase, which was used as an internal control. Data were analyzed using the comparative Ct method (also referred to as the 2 − ΔΔCT method).
miRNA quantification
EV-miRNAs in PC9, PC9OR, H1975, and H1975OR cells; intracellular miRNAs in PC9 and PC9OR cells; and miRNAs in PC9, PC9OR, and H1975 cells transfected with an miR-130a-3p mimic or miR-130a-3p inhibitor were isolated using the mirVana miRNA Isolation Kit (Invitrogen) according to the manufacturer’s protocols. The miRNA concentration was measured using a Qubit microRNA assay kit (Invitrogen). Reverse transcription and qPCR were performed using the Mir-X miRNA qRT-PCR TB Green Kit (Takara Bio, Kusatsu, Shiga) according to the manufacturer’s protocol. The reaction was conducted on a QuantStudio 5 Real-Time PCR System (Thermo Fisher Scientific) with specific primers purchased from Invitrogen (Supplementary Table S2). The target miRNA expression was normalized to that of the internal control U6 (Takara Bio). Data were analyzed according to the comparative Ct method (also referred to as the 2 − ΔΔCT method).
DNA sequencing
DNA was isolated from the cell lines using the Purelink Genomic DNA Mini Kit (Thermo Fisher Scientific). EGFR tyrosine kinase domains (exons 18–21) in PC9 and PC9OR cells were amplified via PCR using specific primers purchased from Invitrogen (Supplementary Table S3), and mutations were detected through Sanger sequencing.
Immunoblotting
Immunoblotting was performed as previously described47 with some modifications. EVs isolated from PC9 and PC9OR cells or PC9 cells transfected with the miR-130a-3p mimic and si-RUNX3 were washed in ice-cold PBS and lysed using RIPA protein extraction reagent (Thermo Fisher Scientific) containing protease and phosphatase inhibitor cocktails (Nacalai Tesque, Kyoto, Japan). Proteins (10 µg) were separated on NuPAGE Novex 4–12% Bis-Tris Gel (Invitrogen, Grand Island, NY, USA) and transferred onto polyvinylidene fluoride membranes (Invitrogen). After blocking with TBS containing 0.1% Tween 20 and 1% non-fat dry skimmed milk for 1 h at room temperature, the membranes were incubated with primary antibodies overnight at 4 °C, followed by incubation with goat anti-rabbit horseradish peroxidase (HRP)-conjugated secondary antibody (#7074, Cell Signaling Technology) or anti-mouse HRP-conjugated secondary antibody (#7076, Cell Signaling Technology) for 1 h at room temperature. The immunoreactive bands were visualized using SuperSignal West Pico Plus Chemiluminescent Substrate (Thermo Fisher Scientific) and a ChemiDoc Imaging System (Bio-Rad Laboratories, Hercules, CA, USA).
Cell viability assay
PC9, PC9OR, and H1975 cells treated with isolated EVs and EV-miRNAs or transfected with the miR-130a-3p mimic or inhibitor or si-RUNX3 were seeded at a density of 5,000 cells/well in 96well flat-bottomed tissue culture plates and cultured for 72 h. One hundred microliters of CellTiter Glo 2.0 (Promega Corporation, WI) were added to the medium and mixed for 2 min on an orbital shaker. Luminescence was determined using a Centro XS3 LB960 (Berthold Technologies, Bad Wildbad, Germany) after incubating the plate at room temperature for 10 min.
Apoptosis assay
The RealTime-Glo Annexin V Apoptosis Assay (Promega Corporation) was performed according to the manufacturer’s protocol. Briefly, PC9 and H1975 cells were cultured in white-bottomed 96-well plates after transfection with miR-130a-3p, and the prepared Detection Reagent was added to each well. Luminescence was measured at different time points (24, 48, and 72 h) after osimertinib treatment using the CentroXS3 LB960 system.
miRNA target prediction
Prediction of target genes and miRNA binding sites was performed using three bioinformatics programs: TargetScan (http://targetscan.org/), miRDB (https://mirdb.org/), and miRTarBase (https://mirtarbase.cuhk.edu.cn/).
Analysis of the relationship between serum miR-130a-3p concentration and clinical outcomes in patients with NSCLC receiving osimertinib
All patients included in this study were diagnosed with advanced NSCLC with EGFR mutations and treated with first-line osimertinib. Twelve frozen serum samples from patients with NSCLC before starting osimertinib treatment between January 2016 and July 2022 were retrieved from the serum bank of the Department of Respiratory Medicine and Clinical Immunology, Graduate School of Medicine, Osaka University. The Maxwell RSC miRNA Plasma and Serum kit (Promega Corporation) was used to extract total miRNA from serum samples according to the manufacturer’s instructions. Comprehensive demographic, clinical, pharmacological, and laboratory data were collected manually from the electronic medical records. Patients who did not experience any events were censored on September 19, 2023, which was the date of the last follow-up update.
Statistical analysis
All data are expressed as mean ± standard error of the mean (SEM). JMP Pro version 17.0.0 (SAS, Tokyo, Japan) and GraphPad Prism version 9.4.0 (GraphPad Software, San Diego, CA, USA) were used for the statistical analysis. Two-tailed Student’s t-test was used to compare the mean values of the two groups. One-way analysis of variance was used to compare three or more groups. The Kaplan–Meier method with the log-rank test was used to analyze the PFS and time to treatment failure. Statistical significance was set at P < 0.05.
Data availability
All data used during the current study are available from the corresponding author upon reasonable request.
References
Torre, L. A. et al. Global cancer statistics, 2012. CA Cancer J. Clin. 65, 87–108 (2015).
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Hirsch, F. R. et al. Lung cancer: current therapies and new targeted treatments. Lancet. 389, 299–311 (2017).
Thai, A. A., Solomon, B. J., Sequist, L. V., Gainor, J. F. & Heist, R. S. Lung cancer. Lancet. 398, 535–554 (2021).
Zhou, W. & Christiani, D. C. East meets West: ethnic differences in epidemiology and clinical behaviors of lung cancer between East asians and caucasians. Chin. J. Cancer. 30, 287–292 (2011).
Duma, N., Santana-Davila, R. & Molina, J. R. Non-small cell lung cancer: epidemiology, screening, diagnosis, and treatment. Mayo Clin. Proc. 94, 1623–1640 (2019).
Wu, Y. L. et al. Osimertinib in resected EGFR-mutated non-small-cell lung cancer. N Engl. J. Med. 383, 1711–1723 (2020).
Wang, S., Cang, S. & Liu, D. Third-generation inhibitors targeting EGFR T790M mutation in advanced non-small cell lung cancer. J. Hematol. Oncol. 9, 34 (2016).
Soria, J. C. et al. Osimertinib in untreated EGFR-mutated advanced non-small-cell lung cancer. N Engl. J. Med. 378, 113–125 (2018).
Cho, B. C. et al. Osimertinib versus standard of care EGFR TKI as first-line treatment in patients with EGFRm advanced NSCLC: FLAURA Asian subset. J. Thorac. Oncol. 14, 99–106 (2019).
Laface, C. et al. The resistance to EGFR-TKIs in non-small cell lung cancer: from molecular mechanisms to clinical application of new therapeutic strategies. Pharmaceutics. 15, 1604 (2023).
Malkin, E. Z. & Bratman, S. V. Bioactive DNA from extracellular vesicles and particles. Cell. Death Dis. 11, 584 (2020).
Kumar, M. A. et al. Extracellular vesicles as tools and targets in therapy for diseases. Signal. Transduct. Target. Ther. 9, 27 (2024).
Kalluri, R. & McAndrews, K. M. The role of extracellular vesicles in cancer. Cell. 186, 1610–1626 (2023).
Saito, M. et al. The association of microRNA expression with prognosis and progression in early-stage, non-small cell lung adenocarcinoma: a retrospective analysis of three cohorts. Clin. Cancer Res. 17, 1875–1882 (2011).
Fontana, F., Carollo, E., Melling, G. E. & Carter, D. R. F. Extracellular vesicles: emerging modulators of cancer drug resistance. Cancers (Basel). 13, 749 (2021).
van Kouwenhove, M., Kedde, M. & Agami, R. MicroRNA regulation by RNA-binding proteins and its implications for cancer. Nat. Rev. Cancer. 11, 644–656 (2011).
Calin, G. A. & Croce, C. M. MicroRNA signatures in human cancers. Nat. Rev. Cancer. 6, 857–866 (2006).
Cao, X. et al. miR-19a contributes to gefitinib resistance and epithelial mesenchymal transition in non-small cell lung cancer cells by targeting c-Met. Sci. Rep. 7, 2939 (2017).
Zhang, W., Dong, Y. Z., Du, X., Peng, X. N. & Shen, Q. M. MiRNA-153-3p promotes gefitinib-sensitivity in non-small cell lung cancer by inhibiting ATG5 expression and autophagy. Eur. Rev. Med. Pharmacol. Sci. 23, 2444–2452 (2019).
Yang, Y. et al. Reciprocal regulation of miR-206 and IL-6/STAT3 pathway mediates IL6-induced gefitinib resistance in EGFR-mutant lung cancer cells. J. Cell. Mol. Med. 23, 7331–7341 (2019).
Liao, J. et al. Down-regulation of miR-214 reverses erlotinib resistance in non-small-cell lung cancer through up-regulating LHX6 expression. Sci. Rep. 7, 781 (2017).
Chuang, L. S. H., Matsuo, J., Douchi, D., Bte Mawan, N. A. & Ito, Y. RUNX3 in stem cell and cancer biology. Cells. 12, 408 (2023).
Jia, J. et al. LncRNA H19 interacted with miR-130a-3p and mir-17-5p to modify radio-resistance and chemo-sensitivity of cardiac carcinoma cells. Cancer Med. 8, 1604–1618 (2019).
Liu, Y. et al. MiR-130a-3p regulates cell migration and invasion via inhibition of Smad4 in gemcitabine resistant hepatoma cells. J. Exp. Clin. Cancer Res. 35, 19 (2016).
Eichelmann, A. K. et al. Complex role of miR-130a-3p and miR-148a-3p balance on drug resistance and tumor biology in esophageal squamous cell carcinoma. Sci. Rep. 8, 17553 (2018).
Zhang, H. D., Jiang, L. H., Sun, D. W., Li, J. & Ji, Z. L. The role of miR-130a in cancer. Breast Cancer. 24, 521–527 (2017).
Wang, X. C. et al. Expression of miRNA-130a in nonsmall cell lung cancer. Am. J. Med. Sci. 340, 385–388 (2010).
Shi, J. et al. MicroRNA-130a targeting hypoxia-inducible factor 1 alpha suppresses cell metastasis and Warburg effect of NSCLC cells under hypoxia. Life Sci. 255, 117826 (2020).
Jiang, H., Yu, W. W., Wang, L. L. & Peng, Y. miR-130a acts as a potential diagnostic biomarker and promotes gastric cancer migration, invasion and proliferation by targeting RUNX3. Oncol. Rep. 34, 1153–1161 (2015).
Chen, C. et al. [Expression of MiR-130a in serum samples of patients with epithelial ovarian cancer and its association with platinum resistance]. Sichuan Da Xue Xue Bao Yi Xue Ban. 47, 60–63 (2016).
Cooper, A. J., Sequist, L. V. & Lin, J. J. Third-generation EGFR and ALK inhibitors: mechanisms of resistance and management. Nat. Rev. Clin. Oncol. 19, 499–514 (2022).
Sadeghi, M. S. et al. Recent advances on high-efficiency of microRNAs in different types of lung cancer: a comprehensive review. Cancer Cell. Int. 23, 284 (2023).
Wang, Y., Li, Y., Wu, B., Shi, C. & Li, C. MicroRNA-661 promotes non-small cell lung cancer progression by directly targeting RUNX3. Mol. Med. Rep. 16, 2113–2120 (2017).
Lee, Y. S. et al. Runx3 inactivation is a crucial early event in the development of lung adenocarcinoma. Cancer Cell. 24, 603–616 (2013).
Wang, M., Wang, X. & Liu, W. MicroRNA-130a-3p promotes the proliferation and inhibits the apoptosis of cervical cancer cells via negative regulation of RUNX3. Mol. Med. Rep. 22, 2990–3000 (2020).
Xu, N. et al. Upregulated miR-130a increases drug resistance by regulating RUNX3 and wnt signaling in cisplatin-treated HCC cell. Biochem. Biophys. Res. Commun. 425, 468–472 (2012).
Zheng, Y. et al. Epigenetic downregulation of RUNX3 by DNA methylation induces docetaxel chemoresistance in human lung adenocarcinoma cells by activation of the AKT pathway. Int. J. Biochem. Cell. Biol. 45, 2369–2378 (2013).
Lin, F. C. et al. RUNX3-mediated transcriptional inhibition of akt suppresses tumorigenesis of human gastric cancer cells. Oncogene. 31, 4302–4316 (2012).
Li, D. J., Shi, M. & Wang, Z. RUNX3 reverses cisplatin resistance in esophageal squamous cell carcinoma via suppression of the protein kinase B pathway. Thorac. Cancer. 7, 570–580 (2016).
Wang, Y. et al. miR-130a upregulates mTOR pathway by targeting TSC1 and is transactivated by NF-κB in high-grade serous ovarian carcinoma. Cell. Death Differ. 24, 2089–2100 (2017).
Ni, J. et al. PPARγ agonist efatutazone and gefitinib synergistically inhibit the proliferation of EGFR-TKI-resistant lung adenocarcinoma cells via the PPARγ/PTEN/Akt pathway. Exp. Cell. Res. 361, 246–256 (2017).
Mori, M. A., Ludwig, R. G., Garcia-Martin, R., Brandão, B. B. & Kahn, C. R. Extracellular miRNAs: from biomarkers to mediators of physiology and disease. Cell. Metab. 30, 656–673 (2019).
Kim, K. M., Abdelmohsen, K., Mustapic, M., Kapogiannis, D. & Gorospe M. RNA in extracellular vesicles. Wiley Interdiscip Rev. RNA. 8https://doi.org/10.1002/wrna.1413 (2017).
Parashar, D., Mukherjee, T., Gupta, S., Kumar, U. & Das, K. MicroRNAs in extracellular vesicles: a potential role in cancer progression. Cell. Signal. 121, 111263 (2024).
Lässer, C., Eldh, M. & Lötvall, J. Isolation and characterization of RNA-containing exosomes. J. Vis. Exp. 59, e3037 (2012).
Shintani, T. et al. Eukaryotic translation initiation factor 3 subunit C is associated with acquired resistance to erlotinib in non-small cell lung cancer. Oncotarget. 9, 37520–37533 (2018).
Acknowledgements
This study was supported by the Center for Medical Research and Education, Graduate School of Medicine, Osaka University, Japan. The authors thank Tomoaki Mizuno from the Center for Medical Research and Education, Graduate School of Medicine, Osaka University for his exceptional support and expertise in facilitating the transmission electron microscopy aspect of our work. We thank Masaya Hamada from the Graduate School of Pharmaceutical Sciences, Osaka University for the establishment of osimertinib-resistant cell lines. We would like to thank Editage (www.editage.jp) for English language editing.
Funding
This work was supported by JSPS KAKENHI through grants JP20K16385 and JP24K18331 and The Research Foundation for Pharmaceutical Sciences.
Author information
Authors and Affiliations
Contributions
T.S. designed and conducted the study. Y.S. and Y.T. contributed to the in vitro experiments. K.J. contributed to the EVs analysis. T.S., I.N., Y.T., and A.K. collected human serum samples and performed clinical analysis. T.S., Y.S., K.I., and M.O. analyzed and interpreted the data and wrote the manuscript. K.I. and M.O. critically revised the manuscript. All authors have read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shintani, T., Shun, YT., Toyozumi, Y. et al. MicroRNA-130a-3p regulates osimertinib resistance by targeting runt-related transcription factor 3 in lung adenocarcinoma. Sci Rep 14, 24429 (2024). https://doi.org/10.1038/s41598-024-76196-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-76196-1