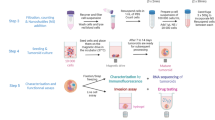
Abstract
Kidney cancer may result from different gene mutations, each contributing to different histological subtypes and prognoses. RAD52 motif-containing 1 (RDM1) regulates multiple cancer pathways, but its role in ccRCC is unknown to date. According to our results, RDM1 expression increased in ccRCC cells, which was correlated to poor survival in ccRCC patients. Knockdown of RDM1 arrested the cell cycle, promoted cell apoptosis, and apparently suppressed ccRCC cell growth in vitro and in vivo. From the mechanism perspective, RDM1 drove MCM2 to modulate ccRCC cell cycle. Thus, RDM1 inhibition blocks cell cycle progression, suppresses ccRCC cell growth, and is a promising approach for treating ccRCC.
Similar content being viewed by others
Introduction
Renal cell carcinoma (RCC), the renal tubular epithelial cell cancer, is also known as kidney cancer1. Kidney cancer encompasses over 10 histological subtypes, among which clear cell RCC (ccRCC) has the highest prevalence, occurring in about 70% of the cases of RCCs2,3. Obesity, hypertension, and smoking are recognized as risk factors for developing RCC4. Surgical resection of RCC is curative for patients with localized disease; however, in the case of metastasis, its 5-year survival rate decreases to 12%5. Systemic treatment is used for advanced RCC, and drugs based on immune checkpoint inhibitors (ICIs) and tyrosine kinase inhibitors (TKIs) are available6. Nonetheless, many patients present original or acquired resistance to these agents and eventually die from the disease7.
The RAD52 motif containing 1 (RDM1), the protein-coding gene, is on chromosome 17q11.2. The human RDM1 protein is a 284-aa polypeptide that is especially abundant in the human testis tissue8. RDM1 binds to both RNA and DNA and has been associated with DNA double-strand break repair and homologous recombination9. Recently, the relationship between RDM1 and cancer development has attracted interest from the research community. RDM1 have apparently elevated mRNA and protein levels in human lung adenocarcinoma cells, whereas RDM1 knockdown apparently inhibited lung tumor growth10. In human ovarian cancer, RDM1 could inhibit the p53/RAD51/RAD52 signaling pathway and displayed an oncogenic function, indicating that targeting RDM1 inhibits ovarian carcinoma cell proliferation in vitro and in vivo11. Similarly, RDM1 expression was reportedly upregulated in human breast cancer tissue compared to its levels in the non-carcinoma counterpart and may also exert a tumorigenic effect on breast cancer12. However, in contrast to the above results, RDM1 downregulation in hepatocellular carcinoma tissues is reportedly correlated to increased tumor size and poor survival, suggesting the complex functions of RDM1 in various circumstances. Mechanistically, RDM1 has been reported to increase the p53 stability and inhibit ERK and Raf phosphorylation13. Nonetheless, the effect of RDM1 on ccRCC development is still largely unclear.
According to our findings, RDM1 expression is increased in human ccRCC tissue and is related to poor survival. Further analysis revealed that RDM1 knockdown arrested cell cycle in G0/G1 stage and promoted ccRCC cell apoptosis. Besides, inhibiting RDM1 expression suppressed xenograft ccRCC tumor growth in vivo. Furthermore, MCM2 was potentially regulated by RDM1 signaling. Accordingly, it was inferred that targeting RDM1 could be a unique and promising approach for treating ccRCC.
Materials and methods
Bioinformatic analysis
RNA-sequencing data and clinicopathological features were obtained in TCGA database (https://cancergenome.nih.gov/), and bioinformatics results were visualized using R-project version 3.6.1 (https://www.r-project.org/).
Patients and specimens
Tissue samples were collected from surgical ccRCC patients at Liaoning Cancer Hospital and Institute (Shenyang, China). Our study protocol gained approval from Ethics Committee of Liaoning Cancer Hospital and Institute, and all methods were carried out in accordance with relevant guidelines and regulations. This work followed the Declaration of Helsinki. Informed consents were obtained from all subjects and/or their legal guardian(s) for ccRCC tissue use in this study. A total of 36 paired ccRCC and 36 matched non-carcinoma kidney tissues were obtained for quantitative real-time PCR (RT-qPCR). Additionally, a tissue microarray (TMA) cohort comprising 183 ccRCC and 27 matched non-carcinoma kidney tissues was established. The clinicopathological data of all cases were acquired based on hospital medical records (Table S1).
Cell culture
The 786-O cells were cultivated within RPMI-1640 (BasalMedia, China) supplemented with 1% penicillin-streptomycin (PS) (BasalMedia, China) and 10% fetal bovine serum (FBS) (Thermo Fisher Scientific, USA). The Caki-1 and 293T cells were cultivated within DMEM (BasalMedia, China) that contained 1% PS and 10% FBS. These cells later underwent incubation under 37 °C with 5% CO2. All cell lines were purchased from the Cell Bank of the Chinese Academy of Sciences.
RNA interference
The siRNAs targeting RDM1 (si-RDM1#1, si-RDM1#2) and the non-targeting control siRNAs were provided by Genechem (Shanghai, China). Lipo8000™ transfection reagent (C0533, Beyotime) was used to transfect the siRNAs into the cells following specific instructions. At 48–72 h post-transfection, we conducted RT-qPCR and western blotting to determine gene knockdown efficiency.
Lentivirus Preparation and infection
Lentiviral vectors carrying the RDM1 plasmids or shRNAs were purchased from Genechem (Shanghai, China). For lentivirus production, 293T cells were used to package the lentivirus. Briefly, 293T cells were subjected to co-transfection using 5 µg of Gagpol, 1 µg of VSVG, and 5 µg of target virus by a calcium phosphate method14. The virus particles were obtained at 36, 48, and 72 h post-transfection, followed by filtering with the 0.45-µm filter (Millipore, USA). Thereafter, the viruses were added to infect target cells during transfection using 8 µg/mL polybrene (ST1380, Beyotime) and then subjected to 60 min of centrifugation (800 rpm) at ambient temperature. After inoculation into the 6-well plastic plates for culture at 37 °C with 5% CO2, 2 µg/mL puromycin (P8833, Sigma-Aldrich) was used for selecting the positively transfected cells for 1 week. After selection, these cells were cultivated with the maintenance medium that contained 0.2 µg/mL puromycin. We conducted RT-qPCR and western blotting to analyze transfection efficiency.
Cell proliferation and cell viability assay
In order to measure cell growth, we transfected 786-O cells, and Caki-1 cells with shNC lentivirus (non-targeting) or the shRDM1 lentivirus. The cells were inoculated in 96-well plastic plates (1500 cells/well), each with 3 technical replicate wells, followed by incubation under 37 °C with 5% CO2. A Celigo® image cytometer (Nexcelom) was used to calculate cell quantity daily for 5 days.
For the cell viability assay, CCK8 reagent (10 µL, DOJINDO, Japan) was introduced into every well before 1.5 h of incubation under 37 °C. The absorbance (OD) was read at 450 and 600 nm by FLUOstar Omega system (BMG LABTECH GmbH, Germany).
Flow cytometric analysis
In order to assess apoptosis, the cells were rinsed with cold PBS twice. Thereafter, a suspension of 105 cells was made in 100 µL of PBS solution and later added into the 5-mL tube. Next, after adding FITC Annexin V (556547, BD biosciences) and PI (BD biosciences) (5 µL each) in this tube, the mixture was gently vortexed, followed by 15 min of incubation under ambient temperature away from light. Then, 1× binding buffer (400 µL) was introduced into every tube. After staining, cells were detected with FACSCelesta™ flow cytometer (BD Biosciences). FlowJo V10 software (FlowJo, USA) was adopted for data analysis.
For identifying the cell cycle arrest, the cells were rinsed before incubation using 70% ethanol for 12 h under 4 °C before PI staining (550825, BD Biosciences). Later, the FACSCelesta™ flow cytometer (BD Biosciences) was applied in determining cell cycle.
Colony formation assay
At 14 days following cell inoculation in the wells of 6-well plates (200 cells/well), all plates were rinsed by PBS before 15 min of fixation with 4% paraformaldehyde. When the colonies were counted, the cells were subjected to 10 min of 0.1% crystal violet staining, after which the colony number was quantified with ImageJ software (NIH, USA). Each assay was performed using three biological replicates.
Quantitative real-time PCR
TRIzol reagent (Invitrogen, USA) and Total RNA Isolation Kit (RC101-01, Vazyme) were adopted for extracting total tissue and cellular RNA separately following specific instructions. A NanoDrop 2000 (Thermo Fisher Scientific) was adopted for quantifying total RNA. Total RNA (1000 ng) was subsequently used for reverse transcription using HiScript® III All-in-one RT SuperMix (R333-01, Vazyme). Then, quantitative RT-PCR was completed with ChamQ SYBR Color qPCR Master Mix (Q411-02, Vazyme). The PCR procedure involved initial denaturation for 3 min at 95 °C, denaturation for 15 s for 30 cycles at 95 °C, annealing for 15 s at 60 °C, and extension for 45 s at 72 °C. ACTB served as the endogenous control. The 2(–ΔΔCT) approach was employed for calculating target gene expression. The primers utilized included: ACTB-F, 5´-CATGTACGTTGCTATCCAGGC-3´; ACTB-R, 5´-CTCCTTAATGTCACGCACGAT-3´; RDM1-F, 5´-GCCCATCCTGGTTTCTATGCC-3´; RDM1-R, 5´-AGACGAACCTTGACTGGAGAT-3´; MCM2-F, 5´-ATGGCGGAATCATCGGAATCC-3´; and MCM2-R, 5´-GGTGAGGGCATCAGTACGC-3´.
Western blotting
RIPA buffer (P0013K, Beyotime Biotechnology) was added for 10 min of cell lysis on ice, followed by brief sonication on a water bath using a sonicator (SONICS, USA). In order to clear the lysates, the cells were subjected to 30 min of centrifugation at 4 °C at 12,000 rpm. The BCA protein assay kit (P0010, Beyotime Biotechnology) was employed in detecting protein contents. Proteins were loaded onto 10% SDS-PAGE gels (PG112, Epizyme Biotech) and separated through electrophoresis before transfer on 0.45-µm PVDF membranes (Millipore, USA). Following 1 h of 5% non-fat milk blocking at room temperature, cells underwent overnight primary antibody probing under 4 °C: anti-RDM1 (20156–1-AP, Proteintech, 1:500), anti-MCM2 (3619, CST, 1:1000), anti-FLAG (14793 S, CST, 1:1000), anti-PRKAA1 (2603, CST, 1:1000), anti-SKP2 (ab183039, Abcam, 1:200), anti-EPS15 (ab174291, Abcam, 1:1000), anti-RALB (ab156799, Abcam, 1:200), anti-Actin (HRP-60008, Proteintech, 1:5000), anti-Cyclin D1 (2922 S, CST, 1:1000), anti-CDK6 (13331 S, CST, 1:1000), anti-CDK4 (12790 S, CST, 1:1000), anti-Bax (14796 S, CST, 1:1000), anti-Caspase 3 (9662 S, CST, 1:1000), anti-Bcl-2 (ab182858, Abcam, 1:2000), and anti-Ki67 (12202). On following day, these membranes were rinsed thrice using TBST (5 min each) and subsequently probed using a goat anti-rabbit IgG secondary antibody (1:5000) under ambient temperature for a 1-h duration. After washing thrice by TBST thrice (5 min each), ECL (S6009, YuHeng) was used for immunoblotting visualization. The quantification of the immunoblotting results was completed using ImageJ (NIH, USA).
Human gene expression microarray analysis
Affymetrix® gene expression microarrays (901838) were applied in identifying differential gene levels in shRDM1-treated versus shNC-treated 786-O cells. A GeneChip™ 3’IVT PLUS kit (902416, Thermo Fisher Scientific) was used for amplification and labeling. Briefly, TRIzol (Invitrogen, USA) was applied in extracting total RNA. A NanoDrop 2000 and Agilent Bioanalyzer 2100 were then used to measure the total RNA quality. Next, both first-strand and second-strand cDNA were prepared to obtain the double-stranded DNA. Through in vitro transcriptional reactions, biotin-labeled amplified RNA was acquired. After amplification, the RNA was subjected to purification, fragmentation, and hybridization using chip probes. After washing and dyeing, the scanning machine (GeneChip Scanner 3000) automatically scanned the results. Fold change > 1.5 and P value < 0.05 were used to detect differentially expressed genes (DEGs) in target versus control groups. IPA and Metascape® (http://metascape.org/) were used in data analysis and visualization. STRING (https://string-db.org/) was adopted in assessing protein-protein interactions (PPIs).
Co-immunoprecipitation (Co-IP)
The 293T cell clones overexpressing RDM1-FLAG, MCM2-MYC, or RDM1-MCM2 were generated through lentivirus infection. The Co-IP assay was performed using immunomagnetic beads (B23202, Bimake). Briefly, the cells were washed using pre-chilled PBS twice, prior to 15 min of lysis on ice using cell lysis buffer (P0013, Beyotime) and additional 5 min of centrifugation (14000 ×g, 4 °C). Supernatants were collected and incubated using 2 µL of anti-Myc tag (100029-MM08, SinoBiological) and 1 µL of anti-IgG (sc-2025, Santa Cruz) onto a rotator under 4 °C for a 4-h period. The mixtures were subsequently incubated with immunomagnetic beads onto the rotator for 1 h under 4 °C. After collection, all beads were gently rinsed 5 times by IP wash buffer. After that, 1× loading buffer was introduced into these beads to elute antigens, before 7 min of boiling under 95 °C and analysis by western blotting. In immunoblotting, antibodies utilized included: anti-MCM2 (3619 S, CST) and anti-FLAG (14793 S, CST).
Immunohistochemistry (IHC) analysis
For IHC, the paraffin-embedded sample was prepared in 4-µm sections, followed by dewaxing using toluene and rehydration with gradient alcohol. Tris-EDTA (pH 9.0) was then added to achieve 3 min of antigen retrieval in a microwave. After 15 min of blocking with 3% H2O2 at ambient temperature and 10 min of blocking using 5% goat serum under ambient temperature, sections were later probed using appropriate primary antibodies overnight under 4 °C. On following day, tissue sections were further probed using an appropriate secondary antibody (PR30011, Proteintech) for 25 min under ambient temperature, after which they were subjected to DAB staining and hematoxylin.
For analysis, the staining intensity scores were defined below: 0–3, negative, weak, medium and strong separately. The positive cell percentage was 1, 0–25%; 2, 26–50%; 3, 51–75%; 4, > 75%. In addition, IHC scores were determined through multiplying staining intensity by positive cell percentage, yielding scores of 0–12. The expression levels were divided into two categories according to their IHC scores: low (0–3) or high (4–12) expression.
TUNEL assay
Through using the TUNEL Assay Kit-FITC (ab66108, Abcam), the TUNEL assay was conducted for detecting the late apoptotic cells following specific protocols. To be specific, 1% paraformaldehyde was added to fix 1 × 106 cells on ice for a 15-min duration and subsequently permeabilized using pre-chilled 70% ethanol for a 30-min period. The fixed cells subsequently underwent incubation using the staining solution that contained the TdT enzyme, reaction buffer, and FITC-dUTP under 37 °C for 1 h. All cells were later rinsed twice using rinsing buffer, whereas the FITC-positive cell percentage was determined by flow cytometry.
Hoechst staining
Apoptotic cells were evaluated using Hoechst 33,258 staining. Specifically, 786-O cells underwent transfection using the lentivirus and then inoculation into the wells of 12-well plates, followed by washing and staining with Hoechst 33,258 for a 30-min period at 37 °C. Finally, a Nikon fluorescence microscope with a UV filter was used to examine and photograph these apoptotic cells.
Immunofluorescence (IF) analysis
After stable transfection with Vector-FLAG or RDM1-FLAG, the 786-O cells were inoculated in the 12-well plates. Following culture for 24 h, cells that reached about 60% confluence were rinsed using PBS thrice (2 min each), followed by 15 min of fixation using 4% paraformaldehyde at ambient temperature, and later clearing using 0.3% Triton X-100 solution (ST797, Beyotime) for an additional 20 min under ambient temperature. All slides were washed thrice using PBS (5 min each), and 2% BSA (ST025, Beyotime) was subsequently added for 1 h of antigen blocking under ambient temperature. Thereafter, these slides received overnight primary antibody probing in dark under 4 °C, including anti-FLAG (F3165, Sigma, 1:800) and anti-MCM2 (3619 S, CST, 1:800). The next day, all slides were washed thrice using PBS, followed by 1 h of secondary antibody (Jackson, 1:500) incubation under ambient temperature away from light, including Alexa Fluor® 594 AffiniPure goat anti-mouse IgG (115–585–003) and fluorescein (FITC) AffiniPure goat anti-rabbit IgG (115–095–003). Next, these slides were incubated with DAPI (P0131, Beyotime) for 5 min away from light before observation with laser confocal microscopy (A1R, Leica).
Xenograft tumor model
The 4-6-week-old BALB/c female nude mice (14–16 g) were obtained from Genechem Co. Ltd. (Shanghai, China) and housed in a pathogen-free environment. Our experimental protocols gained approval from the China Medical University Ethics Committee (CMU2021587). Each approach was conducted following the ARRIVE guidelines. All methods were performed in accordance with the relevant guidelines and regulations. Briefly, 786-O cells were infected with shNC or shRDM1. Next, the same number of cells (1 × 107) were given in every mouse through the right flank. The tumors were palpable at 8 weeks. Mouse weight and tumor size (long and short diameters, mm) were determined 2 times every week. Tumor volume was computed as π/6 × L (length) × W2 (width). The mice were imaged weekly using the IVIS® Lumina LT system (PerkinElmer) to monitor tumor growth and metastasis in vivo. After 2 weeks, the mice were euthanized with carbon dioxide. CO2 injection was administered inside a euthanasia box at a rate of 10–30% per min to replace the euthanasia box volume, with no movement or breathing of the mice or dilation of the pupils. When the CO2 injection was stopped, samples were monitored for 2 min to confirm mouse death. Tumor samples were obtained, photos were taken, and measurements were taken after the mice were euthanized.
Statistical analysis
GraphPad Prism (version 8.0.1) was employed in statistical analysis. Each in vitro experiment was carried out three times or more. Two-tailed Student’s t-test was employed in analyzing between-group differences. Results were represented by means ± SDs. Intra-group differences were compared using ANOVA. Chi-squared test was conducted to examine categorical data. Correlation was analyzed through Spearman’s correlation analysis. Additionally, log-rank test was utilized in survival studies. P < 0.05 represented significant differences.
Results
RDM1 was overexpressed and related to unfavorable prognostic outcome in ccRCC patients
In a previous study, whole gene mRNA expression data of 69 ccRCC and adjacent tissues from the TCGA database were analyzed. According to the high ccRCC expression of the tumor samples and the results reported in the literature, 22 genes that may be related to ccRCC were identified in this study. The shRNAs corresponding to these genes were then transfected in 786-O cells (shRNA) for knocking down these target genes. Celigo analysis revealed that the knockdown of ERCC6L, PLEKHO1, GLYATL2, OIP5, and RDM1 significantly inhibited tumor cell proliferation (Figure S1). Related studies on the involvement of ERCC6L15PLEKHO116, and OIP517 in renal cancer have been reported. Therefore, the impact of RDM1 on ccRCC genesis and progression was explored further.
In order to assess RDM1 mRNA expression in ccRCC, TCGA-derived RNA-sequencing data were screened. Compared with non-carcinoma kidney samples, the ccRCC samples exhibited RDM1 overexpression (Fig. 1A). When the RDM1 expression levels in samples with different Fuhrman nuclear grades were compared, RDM1 expression increased in Grade 3–4 tumors relative to Grade 1–2 counterparts (Fig. 1B). Similarly, stage III-IV tumors presented higher RDM1 expression levels than stage I-II tumors (Fig. 1C). Moreover, RDM1 overexpression was related to relatively short overall survival (OS) in ccRCC patients, according to the TCGA dataset (Fig. 1D). These data suggested that RDM1 might affect ccRCC progression. In order to validate these TCGA results, RDM1 mRNA expression was detected in the tumor tissues and paired adjacent normal kidney samples obtained in 36 ccRCC patients using real-time PCR. Compared to those in normal controls, RDM1 mRNA levels were increased in the ccRCC samples (Fig. 1E). Furthermore, a TMA cohort comprising 183 ccRCC samples and 27 adjacent normal kidney tissues was established. The immunohistochemical staining of RDM1 in this TMA confirmed that the RDM1 protein levels elevated in tumor tissues in comparison with non-carcinoma controls (Fig. 1F and G). When tumors were divided in RDM1-High or RDM1-Low group according to IHC scores, patients in the RDM1-Low groups had longer survival times than those in the RDM1-High group (Fig. 1H). Furthermore, univariate analysis revealed that the RDM1 protein expression level in the TMA was associated with the clinical stage and vital status (Table 1). The above findings revealed that RDM1 overexpression in human ccRCC tissues is related to a dismal OS.
RDM1 is overexpressed in ccRCC cells and is related to poor patient survival. (A-C) TCGA database showing RDM1 mRNA levels in normal and ccRCC tissues (A), as well as in different grades (B) and different stages (C). Student´s t‑test. Error bars, mean ± SD. *p < 0.05. (D) TCGA database analysis revealed that increased expression of RDM1 mRNA is related to unfavorable patient survival. Long-rank test. (E) Tissue samples were obtained from the surgical resections of ccRCC patients and subjected to RT-qPCR to quantify the RDM1 mRNA expression (n = 36). Student´s t‑test. *p < 0.05. (F) Typical images showing RDM1 IHC staining for human normal renal tissues and ccRCC tissues. Scale bars, 100 μm. (G) Quantification of RDM1 expression based on IHC scores. Student´s t‑test. Error bars, mean ± SD. *p < 0.05. (H) Human ccRCC tissues were classified into RDM1-High or RDM1-Low-expression groups on the basis of RDM1 IHC scores, and patient overall survival was subsequently determined using the log-rank test.
RDM1 downregulation suppresses ccRCC cell proliferation
In order to analyze RDM1’s role in ccRCC, RDM1 mRNA expression in RCC cell lines was decreased through transfection with short hairpin RNAs (shRNAs) (Fig. 2A and C). Cell viability was evaluated by the CCK8 assay after shRNA treatment. In comparison with the non-targeting shRNAs-transfected cells, both 786-O and Caki-1 cells presented significantly decreased cell viability following shRDM1 transfection (Fig. 2D). Moreover, a colony formation assay demonstrated that the knockdown of RDM1 mRNA expression in the tested RCC cells significantly suppressed their colony formation ability (Fig. 2E and F). In line with these findings, the Celigo analysis revealed that pretreatment with shRDM1 significantly inhibited cell proliferation compared to that in the control groups (Fig. 2G and H, and 2I). These results suggested that the loss of RDM1 mRNA expression repressed ccRCC cell proliferation.
Nonetheless, RDM1 overexpression made no difference to the viability or colony formation ability of ccRCC cells (Figure S2). Furthermore, RDM1 overexpression did not affect the sensitivity of either 786-O or Caki-1 cells to cisplatin (Figure S2).
Suppressing RDM1 expression in renal cell carcinoma inhibits ccRCC cell proliferation. (A and B) RCC cells underwent transfection using non-targeting shRNA (shNC) or shRDM1 lentivirus, following which RT-qPCR was performed to measure the mRNA knockdown efficiency. Student´s t‑test. Error bars, mean ± SD. ***p < 0.001. (C) Western blotting results demonstrating RDM1 protein levels before and after the shRNA transfection. (D) Human RCC cells were treated with shNC or shRDM1, followed by a CCK8 assay to determine the absorbance at 450 nm at different time points. One‑way ANOVA. Error bars, mean ± SD. n.s., not significant. ***p < 0.001. (E) Representative images of the colony formation assay results for the indicated cells. (F) Colony formation assays. Student´s t‑test. Error bars, mean ± SD. ***p < 0.001. (G) RCC cells were pretreated with shNC or shRDM1, after which the cell counts were measured using the Celigo assay at different time points. (H-I) Quantification of the cell counts in (G). Student´s t‑test. Error bars, mean ± SD. ***p < 0.001. (J) Cell viability, as determined by a CCK8 assay. Student´s t‑test plus Welch´s correction was utilized. Error bars, mean ± SD. ***p < 0.001.
Knocking down RDM1 mRNA expression blocks the cell cycle, promotes apoptosis in vitro, and restrains tumor development in vivo
To determine why RDM1 mRNA silencing suppressed kidney cancer cell growth, we conducted flow cytometry to analyze cell cycle distribution and cell apoptosis in ccRCC cells. From Fig. 3A and D, a decrease in the mRNA expression of RDM1 arrested cell cycle at G0/G1 phase. According to western blotting, RDM1 silencing downregulated the cyclines related to the G0/G1 phase (Fig. 3E). Moreover, RDM1 knockdown significantly induced the apoptosis of ccRCC cells compared to that observed from control cells (Fig. 3F and G). According to Western blotting, RDM1 silencing upregulated Bax and Caspase 3 but downregulated Bcl-2 (Fig. 3H). Furthermore, to determine whether RDM1 expression impacts ccRCC tumor development in vivo, we established a xenograft tumor nude mouse model by transplanting 786-O ccRCC cells with or without the shRDM1 lentivirus (Figure S3A). Compared to the vehicle control, shRDM1 transfection significantly inhibited tumor growth in these ccRCC xenograft tumor models (Fig. 3I and K, Figure S3B and S3C, Table S2). In addition, as suggested by TUNEL assay, RDM1 silencing remarkably induced tumor cell apoptosis in vivo (Fig. 3L and M). Furthermore, a decreased percentage of Bcl-2-positive cells, accompanied by the upregulation of Bax and Caspase 3, was observed from shRDM1 group (Fig. 3N). Therefore, lowering RDM1 expression in ccRCC cells arrested cell cycle at G0/G1 phase, promoted apoptosis in vitro, and markedly suppressed tumor growth in vivo.
RDM1 loss suppresses cell cycle, promotes apoptosis, and suppresses tumor development in vivo. (A–D) After shNC or shRDM1 treatment, the cell cycle progression of 786-O and Caki-1 cells could be observed through assays. Student´s t‑test. Error bars, mean ± SD. ***p < 0.001. (E) Western blotting results showing Cyclin D1, CDK4, and CDK6 protein levels before and after shRNA transfection. (F and G) PI and Annexin V staining of shNC- or shRDM1-treated human ccRCC cells, and results analyzed using flow cytometry. Student´s t‑test. Error bars, mean ± SD. ***p < 0.001. (H) Western blotting results showing Bax, Bcl-2, and Caspase 3 protein levels before and after shRNA transfection. (I) Excision and imaging of the mice xenotransplanted with human 786-O cells transfected with shNC or shRDM1. (J) Tumor volume and weight (K) at the end points of subcutaneous xenograft tumors (n = 10 for every group). Student´s t-test. Error bars, mean ± SD. **p < 0.01. (L-M) Representative images of the TUNEL assay. Scale bars, 20 μm. Student´s t‑test. Error bars, mean ± SD. ***p < 0.001. (N) Representative images of IHC. Scale bars, 100 μm.
MCM2 is speculated to be the downstream gene of RDM1
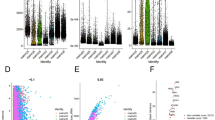
Next, an mRNA microarray analysis was conducted to investigate the potential mechanism through which RDM1 regulates RCC cell survival. After RDM1 mRNA expression in RCC 786-O cells was silenced using shRNA (Figure S4), there were 433 DEGs detected, including 260 upregulated, whereas 173 downregulated ones (Fig. 4A and B). The GSEA plot analysis revealed that the apoptosis pathway was markedly activated after RDM1 was knocked down in these cells (Fig. 4G). Next, a Metascape database assay was carried out to enrich all DEGs. The major downstream regulatory genes after RDM1 knockdown were correlated to multiple biological processes, including homophilic cell adhesion, regulation of mRNA stability, and DNA replication (Fig. 4H). Furthermore, the potential interacting genes and proteins of RDM1 were explored by searching the IPA and STRING databases (Fig. 4C and D). Notably, MCM2 is among the listed genes in both potential interacting gene network and protein network. In order to verify the results of the mRNA microarray, the protein expressions of selected genes in the same network as RDM1 were determined through immunoblotting after RDM1 mRNA expression was knocked down in RCC cells (Fig. 4E). As shown in Fig. 4F, MCM2 expression dramatically decreased in RDM1-knockdown cells. This result prompted the subsequent exploration of the functional association between RDM1 and MCM2 in this study.
DEGs associated with shRDM1 in RCC cells. (A) Volcano plot and heatmap (B) showing the upregulated genes (n = 260) and downregulated genes (n = 173) in the 786-O cells infected with shRDM1. (C) IPA showing RDM1-related genes. (D) PPI analysis of RDM1-related proteins using STRING. (E) Immunoblot analysis of the indicated proteins in RDM1-KD 786-O cells and quantification using ImageJ (F). Student´s t-test. Error bars, mean ± SD. n.s., not significant. ***p < 0.001. (G) GSEA plot showing the signaling pathways associated with RDM1 knockdown. (H) Enrichment heatmap for DEGs, generated using Metascape.
MCM2 is involved in regulating RDM1 in ccRCC cells
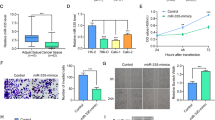
To verify the relevance of RDM1 and MCM2 in ccRCC cells, the mRNA levels of MCM2 were determined in RDM1-knockdown cells. As expected, suppressing RDM1 mRNA expression inside 786-O and Caki-1 cells apparently decreased MCM2 mRNA and protein expression (Fig. 5A and B). In contrast, RDM1 overexpression increased MCM2 protein level (Fig. 5C and D). A similar effect was also observed in the immunofluorescence (IF) analysis, in which RDM1 overexpression increased the tumor-intrinsic MCM2 levels (Fig. 5E and F). In addition, the RDM1 protein accumulated and colocalized with the MCM2 protein in the nucleus (Fig. 5E), highlighting the potential relevance of these two proteins in terms of function. However, a co-immunoprecipitation (co-IP) study did not reveal a direct binding between RDM1 and MCM2, suggesting a likely indirect interaction between these two proteins (Figure S5). Given that downregulating RDM1 expression robustly arrested cell cycle at G0/G1 phase, it was explored whether MCM2 is involved in this biological process. To this end, the 786-O cells in which RDM1 mRNA expression was decreased using shRNA were supplemented with exogenous MCM2 (Fig. 5G). As expected, RDM1 KD-induced G0/G1 phase arrest in 786-O cells was largely reversed upon MCM2 overexpression (Fig. 5H and Figure S6). This finding was supported by the results of a colony formation assay, in which the MCM2 overexpression in ccRCC cells in which RDM1 expression was silenced using shRNA significantly increased colony formation capacity of cells compared to control cells that had been transfected with only shRDM1 but without MCM2 overexpression (Fig. 5I and J). Apoptosis was assessed based on the Hoechst staining results, which again indicated that overexpression of MCM2 could rescue the apoptosis caused by shRDM1 (Fig. 5K and L). Taken together, these results suggested that MCM2 is involved in the regulatory effect of RDM1 on cell survival.
Overexpression of MCM2 can rescue the RDM1 knockdown phenotype. (A and B) RT-qPCR and immunoblotting of the renal cancer cells that expressed RDM1 and MCM2. The cells received shNC or shRDM1 pretreatment. Student´s t-test. Error bars, mean ± SD. ***p < 0.001. (C and D) RT-qPCR and immunoblot analysis of the renal cancer cells that expressed RDM1 and MCM2. The cells received vector-FLAG or RDM1-FLAG pretreatment. Student´s t-test. Error bars, mean ± SD. ***p < 0.001. (E) The 786-O cells were overexpressed with control (Vector-FLAG) or RDM1-FLAG, and the representative images of IF for RDM1 (red), MCM2 (green), and DAPI (blue) are shown. Scale bars, 10 μm. (F) Quantification of the MCM2 fluorescence intensity. Student´s t-test. Error bars, mean ± SD. *p < 0.05. (G and H) The 786-O cells were stably transfected with an empty vector or MCM2, after which the cells were transfected with shNC or shRDM1, and the cell cycle stage of the indicated cells was quantified. One-way ANOVA plus Tukey’s multiple comparisons test was used. Error bars, mean ± SD. n.s., not significant. *p < 0.05. (I) Colony formation ability analysis was conducted for the 786-O cells of the indicated genotypes. (J) Quantification of colony formation ability. One-way ANOVA. Error bars, mean ± SD. n.s., not significant. **p < 0.01, ***p < 0.001. (K) The apoptosis degree in 786-O cells of the indicated genotypes determined based on Hoechst staining. (L) Quantification of apoptosis. One-way ANOVA. Error bars, mean ± SD. n.s., not significant, ***p < 0.001.
RDM1 expression is positively correlated to MCM2 expression in ccRCC tissue samples
To validate the functional relevance of RDM1 and MCM2, RDM1 and MCM2 mRNA and protein levels inside ccRCC tissue specimens were determined. As shown in Fig. 6A, relative to that in adjacent non-carcinoma samples, MCM2 mRNA expression was marginally upregulated in 36 human ccRCC tissues, with no significant difference, probably associated with our small sample size. However, a likely positive relationship between RDM1 mRNA expression and MCM2 expression was observed in these samples (Fig. 6B). Furthermore, the relation of MCM2 and RDM1 protein levels in a TMA cohort was explored using an IHC assay. Indeed, more samples expressed a high MCM2 protein in RDM1-high group relative to RDM1-low group. On the contrary, there were more samples expressing a low MCM2 protein level in RDM1-low group compared with RDM1-high group, suggesting the potential relationship between the expressions of these two proteins (Fig. 6C and D). More interestingly, patients showing high RDM1 and MCM2 expression presented the worst OS, whereas those displaying low expression presented the best survival rate (Fig. 6E), highlighting the important role of the coexistence of these two proteins in the progression of ccRCC.
RDM1 expression is positively associated with MCM2 in ccRCC patients. (A) Tissue samples were obtained from surgical resections for ccRCC, and RT-qPCR was used to quantify MCM2 mRNA expression (n = 36). Student´s t‑test. n.s., not significant. (B) RT-qPCR analysis to determine the correlation between RDM1 and MCM2. Spearman´s correlation analysis. (C) Representative images of the RDM1 and MCM2 IHC staining of human ccRCC tissues from different patients. Scale bars, 100 μm. (D) Quantification of RDM1 and MCM2 protein levels. Chi-squared test. *p < 0.05. (E) RDM1 and MCM2 IHC scores were used, and patient OS was examined through log-rank test. *p < 0.05.
Discussion
As new therapeutic agents and methods develop, the treatment for advanced RCC patients has changed dramatically over the past decade. However, owing to inherent tumor heterogeneity and the development of drug resistance, therapeutic efficacy remains limited to date, and individualized therapy for metastatic RCC is difficult to achieve18,19. In RCC, differences in the gene expression profiles allow for the separation of tumors into distinct histological subtypes, which are associated with diverse overall survival rates20. In this study, it was revealed that RDM1 mRNA expression in ccRCC tissues increased in comparison with non-carcinoma kidney tissues, which was correlated to advanced-stage disease and poor survival. Knockdown of RDM1 in ccRCC cells arrested cell cycle at G0/G1 phase, accelerated apoptosis in vitro, and inhibited xenograft tumor growth in vivo. Furthermore, RDM1 may function through the regulation of MCM2 expression. Thus, targeting RDM1 has the potential to provide a promising approach for treating ccRCC.
RDM1 is previously suggested to enhance tumor progression. RDM1 mRNA and protein levels were significantly increased in human lung adenocarcinoma cells, whereas RDM1 knockdown apparently inhibited lung tumor growth10. Similarly, in human ovarian cancer or breast cancer, RDM1 was reported to perform an oncogenic function by inhibiting the p53/RAD51/RAD52 signaling pathway11,12. These findings are consistent with this study, in which RDM1 overactivation was observed to promote cell proliferation or tumor growth in ccRCC. In another study, RDM1 mRNA expression increased in the hepatocellular carcinoma tissues, which was related to short-term OS and progression-free survival. Interestingly, in this study, the elevated RDM1 mRNA expression strongly associated with immune cell infiltration, probably involved in T-cell exhaustion21. In recent years, ICIs have gained FDA approval for treating advanced RCC as monotherapies or in combination with other strategies7,22,23. From this point of view, RDM1 might be involved in the immunoregulation of the RCC microenvironment, which could be the future direction of the research to explore the regulatory effect of RDM1 on ICI treatment.
In a previous study, knocking down RDM1 mRNA expression in chicken B cells DT40 enhanced the cisplatin sensitivity8. However, in this study, RDM1 overexpression did not affect the sensitivity of either 786-O or Caki-1 cell line to cisplatin. Given that ccRCC cells are largely resistant to chemotherapy in the clinic, it is speculated that there are robust signaling pathways regulating the chemosensitivity of RCC cells, which may not be evident in other cancer cells. It is likely that RDM1 is not involved in the major signaling pathway of chemoresistance in RCC, although RDM1 was found to strongly impact cell survival in this work.
Although the RDM1 regulatory mechanism on cell survival in ccRCC has not been clearly characterized, the functional relevance between RDM1 and MCM2 was demonstrated in the present study. Notably, MCM2 is a key protein in the mini-chromosome maintenance (MCM) complex, and it serves as the replicative DNA helicase unwinding duplex DNA while initiating DNA replication24,25. In addition, MCM2 can promote symmetric inheritance of parental histones to sister chromatids during DNA replication26. Therefore, MCM2 has an important effect on maintaining normal cell division and DNA replication. MCM2 was previously suggested to be related to tumor development and prognosis. For instance, NF-κB upregulates MCM2, resulting in increased cell viability and maintenance of colon cancer cell stemness27. In addition, MCM2, MCM4, and MCM6 expressions increase in triple-negative breast cancer, and these biomarkers exhibit prognostic value in predicting relapse-free survival28. The findings of this study revealed that the RDM1 and MCM2 mRNA expression levels were positively correlated in 36 paired ccRCC tissues. Notably, it was demonstrated that these two proteins might be co-expressed inside ccRCC cells, suggesting a potential functional association between these two proteins. More importantly, the patients showing high RDM1 and MCM2 expression showed the worst OS in the TMA cohort of ccRCC patients.
However, how RDM1 regulates the expression of MCM2 remains elusive. Notably, RDM1 contains an RNA recognition motif (RRM), which has been previously shown to participate in important molecular processes, such as RNA splicing, processing, editing, and transcriptional modulation29,30. Therefore, RDM1 directly regulates the expression of MCM2, although more intensive work is needed in the future. In addition, RDM1 has multiple protein isoforms, among which full-length RDM1 protein is found predominantly in the cytoplasm, whereas mild heat shock and proteotoxic stress induce RDM1 accumulation in nucleolus31. In this study, overexpressed RDM1-FLAG (full-length RDM1) directly accumulated in the nucleus, suggesting its potential involvement in processes that occur in the nucleus, including DNA replication and DNA unwinding, which are regulated by MCM2. Indeed, the key effect of MCM2 during the regulation of RDM1 on the survival of ccRCC cells was revealed. In this context, the present results suggested that the overexpression of the RDM1-MCM2 axis in ccRCC patients may be important for cancer development and progression and is thus worthy of further research.
Conclusion
In conclusion, RDM1 is overexpressed in ccRCC tissue samples and is related to poor survival in these patients. Functionally, RDM1 affects ccRCC cell proliferation and promotes apoptosis, potentially by regulating MCM2. Further investigations are, nonetheless, necessary to analyze the potential of targeting the RDM1-MCM2 axis in treating ccRCC.
Limitations of the study
Although this study demonstrated the crucial role of RDM1 in ccRCC, further work is needed to define the mechanism and function of RDM1 in various kidney cancer histological subtypes. How RDM1 is distributed in the cytoplasm and the nucleus and the mechanism and functions of RDM1 remain unknown. In addition, while the result data from this study suggest that RDM1 positively regulates MCM2 expression, the underlying molecular mechanism is still unknown and needs more investigations.
Data availability
Data is provided within the manuscript or supplementary information files.
References
Hsieh, J. J. et al. Renal cell carcinoma. Nat. Rev. Dis. Primers. 3, 17009 (2017).
Barata, P. C. & Rini, B. I. Treatment of renal cell carcinoma: Current status and future directions. Cancer J. Clin. 67 (6), 507–524 (2017).
Jonasch, E. et al. Clear cell renal cell carcinoma ontogeny and mechanisms of lethality. Nat. Rev. Nephrol. 17 (4), 245–261 (2021).
Capitanio, U. et al. Epidemiology of renal cell carcinoma. Eur. Urol. 75 (1), 74–84 (2019).
Choueiri, T. K. & Motzer, R. J. Systemic therapy for metastatic Renal-Cell carcinoma. N. Engl. J. Med. 376 (4), 354–366 (2017).
Deleuze, A. et al. Immunotherapy in renal cell carcinoma: the future is now. International J. Mol. Sciences 21(7), (2020).
Braun, D. A. et al. Beyond conventional immune-checkpoint inhibition - novel immunotherapies for renal cell carcinoma. Nat. Rev. Clin. Oncol. 18 (4), 199–214 (2021).
Hamimes, S. et al. RDM1, a novel RNA recognition motif (RRM)-containing protein involved in the cell response to cisplatin in vertebrates. J. Biol. Chem. 280 (10), 9225–9235 (2005).
Hamimes, S. et al. Nucleic acid-binding properties of the RRM-containing protein RDM1. Biochem. Biophys. Res. Commun. 344 (1), 87–94 (2006).
Tong, L. et al. RDM1 plays an oncogenic role in human lung adenocarcinoma cells. Sci. Rep. 8 (1), 11525 (2018).
Tong, L. et al. RDM1 plays an oncogenic role in human ovarian carcinoma cells. Artif. Cells Nanomed. Biotechnol. 48 (1), 885–892 (2020).
Chen, Y. et al. RDM1 promotes critical processes in breast cancer tumorigenesis. J. Cell. Mol. Med. 23 (8), 5432–5439 (2019).
Chen, S. L. et al. Loss of RDM1 enhances hepatocellular carcinoma progression via p53 and ras/raf/erk pathways. Mol. Oncol. 14 (2), 373–386 (2020).
Kwon, M. & Firestein, B. L. DNA transfection: calcium phosphate method. Methods Mol. Biol. 1018, 107–110 (2013).
Zhang, G. et al. ERCC6L that is upregulated in high grade of renal cell carcinoma enhances cell viability in vitro and promotes tumor growth in vivo potentially through modulating MAPK signalling pathway. Cancer Gene Ther. 26 (9–10), 323–333 (2019).
Yu, Z. et al. PLEKHO1 knockdown inhibits RCC cell viability in vitro and in vivo, potentially by the Hippo and MAPK/JNK pathways. Int. J. Oncol. 55 (1), 81–92 (2019).
Gong, M. et al. OIP5 is a novel prognostic biomarker in clear cell renal cell Cancer correlating with immune infiltrates. Front. Immunol. 13, 805552 (2022).
Posadas, E. M. et al. Targeted therapies for renal cell carcinoma. Nat. Rev. Nephrol. 13 (8), 496–511 (2017).
Kotecha, R. R. et al. Towards individualized therapy for metastatic renal cell carcinoma. Nat. Rev. Clin. Oncol. 16 (10), 621–633 (2019).
Ricketts, C. J. et al. The Cancer genome atlas comprehensive molecular characterization of renal cell carcinoma. Cell. Rep. 23 (1), 313–326e315 (2018).
Qiu, C. et al. Correlation analysis of RDM1 gene with immune infiltration and clinical prognosis of hepatocellular carcinoma. Biosci Rep 41(9), (2021).
Díaz-Montero, C. M. et al. The immunology of renal cell carcinoma. Nat. Rev. Nephrol. 16 (12), 721–735 (2020).
Hu, J. et al. Single-Cell transcriptome analysis reveals intratumoral heterogeneity in ccrcc, which results in different clinical outcomes. Mol. Ther. 28 (7), 1658–1672 (2020).
Li, Z. & Xu, X. Post-Translational modifications of the Mini-Chromosome maintenance proteins in DNA replication. Genes (Basel) 10(5), (2019).
Sedlackova, H. et al. Equilibrium between nascent and parental MCM proteins protects replicating genomes. Nature 587 (7833), 297–302 (2020).
Petryk, N. et al. MCM2 promotes symmetric inheritance of modified histones during DNA replication. Sci. (New York N Y). 361 (6409), 1389–1392 (2018).
Wang, L. et al. NF-κB maintains the stemness of colon cancer cells by downregulating miR-195-5p/497-5p and upregulating MCM2. J. Experimental Clin. cancer Research: CR. 39 (1), 225 (2020).
Issac, M. S. M. et al. MCM2, MCM4, and MCM6 in Breast Cancer: Clinical Utility in Diagnosis and Prognosis. Neoplasia 21(10), 1015–1035, (2019).
Bhattacharya, S. et al. Structural basis of the interaction between SETD2 methyltransferase and HnRNP L paralogs for governing co-transcriptional splicing. Nat. Commun. 12 (1), 6452 (2021).
Kang, H. S. et al. An autoinhibitory intramolecular interaction proof-reads RNA recognition by the essential splicing factor U2AF2. Proc. Natl. Acad. Sci. U S A. 117 (13), 7140–7149 (2020).
Messaoudi, L. et al. Subcellular distribution of human RDM1 protein isoforms and their nucleolar accumulation in response to heat shock and proteotoxic stress. Nucleic Acids Res. 35 (19), 6571–6587 (2007).
Acknowledgements
Not applicable.
Funding
Natural Science Foundation of Hebei Province (Grant No. H2021406054 and H2023406033). Science and Technology Research Program of the Hebei Provincial Department of Education (Grant No. QN2022114).
Author information
Authors and Affiliations
Contributions
X.L. wrote the main manuscript. H.L. performed main analysis. Y.W., A.W. and C.L. performed the main experiments. Y.Z. designed the study and confirmed the authenticity of all the raw data. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The present study was approved by the Ethics Committee of the China Medical University Ethics Committee (Shenyang, China).
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, X., Liu, H., Wei, Y. et al. RDM1 plays an oncogenic role in clear cell renal cell carcinoma potentially by modulating MCM2. Sci Rep 15, 22649 (2025). https://doi.org/10.1038/s41598-025-07230-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-07230-z