Abstract
The solute carrier (SLC) protein family, traditionally recognized for transporting molecules across cell membranes, is gaining attention for its broader roles, including signaling. Among SLC proteins, the ascorbate transporter SVCT2 remains poorly understood, particularly in relation to estrogen receptor alpha (ERα), a key regulator in breast cancer cells. Here, we investigate how ERα regulates SVCT2 and its implications for chemoresistance. Our results demonstrate that ERα knockdown significantly reduces SVCT2 protein levels, impairing cellular ascorbic acid uptake. Mechanistically, ERα directly interacts with SVCT2. We show that X-linked inhibitor of apoptosis protein (XIAP), an E3 ubiquitin ligase, targets SVCT2 for ubiquitination and subsequent proteasomal degradation in ERα-deficient conditions. Notably, silencing XIAP restored SVCT2 stability, underscoring its regulatory role. Functionally, ERα or SVCT2 knockdown decreases doxorubicin-induced cytotoxicity, accompanied by increased expression of ATP-binding cassette (ABC) transporter genes, which mediate drug efflux and contribute to chemoresistance. These findings uncover a novel regulatory axis between ERα and SVCT2, mediated by XIAP, and establish SVCT2 as a critical factor in maintaining cellular ascorbic acid levels and drug sensitivity. Targeting XIAP or modulating SVCT2 may represent promising therapeutic strategies for overcoming resistance in ERα-positive breast cancer. This study advances our understanding of the interplay between nutrient transport and cancer therapy, offering new avenues for intervention.
Similar content being viewed by others
Introduction
Breast cancer, a heterogeneous and complex disease, continues to be a major global health concern affecting millions of individuals worldwide1,2. The complexity of breast cancer progression involves numerous molecular factors that contribute to its initiation, growth, and metastasis. Among these factors, estrogen receptor alpha (ERα) plays a critical role in regulating essential cellular processes in breast tissue3,4.
As a ligand-activated transcription factor, ERα regulates the expression of numerous genes involved in cell proliferation, survival, and differentiation5. In normal breast tissue, ERα signaling is tightly regulated and contributes to tissue homeostasis6. However, dysregulation of ERα signaling is characteristic of hormone receptor-positive breast cancer, which comprises approximately 70% of all breast cancer cases7. In hormone receptor-positive breast cancer, aberrant activation of ERα signaling promotes tumor growth and progression8. The binding of estrogen to ERα leads to receptor dimerization and translocation to the nucleus, where it interacts with specific DNA sequences known as estrogen response elements (EREs) to regulate gene transcription9,10. Consequently, ERα target genes involved in cell cycle progression (e.g., cyclin D1), anti-apoptosis (e.g., Bcl-2), and estrogen biosynthesis (e.g., aromatase) are upregulated, fueling tumor proliferation and survival11,12,13. In addition to its canonical role as a transcriptional regulator, ERα can also exert non-genomic effects that influence various signaling pathways involved in cell growth and survival14. These rapid, non-genomic actions of ERα are mediated through interactions with cytoplasmic signaling proteins, such as kinases and G protein-coupled receptors, leading to the activation of downstream signaling cascades, including the PI3K/Akt and MAPK pathways15. Importantly, ERα status serves as a critical biomarker for breast cancer treatment decisions16. Hormone receptor-positive breast cancers are generally sensitive to endocrine therapies targeting ERα signaling, such as selective estrogen receptor modulators (e.g., tamoxifen), aromatase inhibitors, and selective estrogen receptor degraders (e.g., fulvestrant)17,18. However, acquired resistance to endocrine therapy remains a substantial clinical challenge, underscoring the necessity for a deeper understanding of the mechanisms underlying ERα signaling in breast cancer progression and therapy resistance19,20. Given ERα's central role in hormone receptor-positive breast cancer, it is crucial to explore its regulatory interactions with other signaling pathways and cellular processes to identify novel therapeutic targets and enhance patient outcomes21,22.
SVCT2, a sodium-dependent vitamin C transporter encoded by the SLC23A2 gene, has recently garnered interest in cancer research due to its multifaceted roles in cellular physiology and its implications for cancer biology and therapy23,24,25. Facilitating the cellular uptake of ascorbic acid (vitamin C), an essential nutrient with antioxidant properties26, SVCT2 has been implicated in various facets of cancer progression across multiple cancer types27. Notably, aberrant SVCT2 expression has been associated with tumor growth, metastasis, and resistance to therapy in several malignancies including breast cancer, prostate cancer, lung cancer, and colorectal cancer, underscoring its potential as a prognostic biomarker and therapeutic target in these diseases28,29,30,31. Moreover, emerging evidence suggests that SVCT2-mediated vitamin C uptake may have multifaceted effects on cancer cells beyond its antioxidant capabilities32,33. Vitamin C has been implicated in regulating epigenetic modifications, immune responses, and metabolism, thereby influencing various hallmarks of cancer progression34,35. Consequently, targeting SVCT2-mediated ascorbic acid transport offers a promising approach for developing novel therapeutic strategies aimed at impeding cancer cell viability and overcoming resistance to therapy36,37. Previous studies have elucidated the mechanisms by which ascorbic acid induces cancer cell death23,24,25,26. Notably, colorectal cancer cells harboring KRAS or BRAF mutations have been reported to exhibit heightened sensitivity to oxidative stress-induced apoptosis triggered by ascorbic acid38. This effect is mediated by the sodium-dependent vitamin C transporter 2 (SVCT2), which facilitates the uptake of ascorbic acid into cells. Building on this, our previous studies investigated the cytotoxic effects of ascorbic acid on various types of breast cancer cells, a subtype characterized by the presence or absence of hormone receptors, to explore the role of SVCT2 expression in mediating these effects. Given the importance of SVCT2 in ascorbic acid uptake, we hypothesized that mechanisms might extend to breast cancer subtypes, particularly those involving hormone receptor pathways. Therefore, this study focuses on the potential role of estrogen receptor alpha (ERα) in regulating SVCT2 expression in breast cancer, aiming to provide insights into therapeutic strategies for overcoming treatment resistance, particularly in ERα-positive cases.
This study examines the regulatory relationship between ERα and SVCT2 in breast cancer cells, focusing on its potential impact on chemotherapeutic resistance. Our results reveal a significant correlation between ERα and SVCT2 expression levels, indicating that ERα knockdown leads to a substantial decrease in SVCT2 protein levels. This implies that ERα is essential for maintaining SVCT2 expression and stability. Further investigation demonstrated that while ERα knockdown does not directly affect SVCT2 transcription, it significantly impacts SVCT2 protein stability. Specifically, ERα regulates SVCT2 through the ubiquitin–proteasome degradation pathway, with the E3 ligase XIAP identified as a crucial mediator for SVCT2 ubiquitination and subsequent degradation. This underscores the post-transcriptional regulation of SVCT2 by ERα, highlighting the complexity of protein stability in breast cancer cells. Moreover, we observed that reduced SVCT2 levels following ERα knockdown are linked with increased resistance to the chemotherapeutic agent doxorubicin. This resistance is particularly pronounced in the presence of ascorbic acid, suggesting that SVCT2, as a facilitator of vitamin C uptake, may affect cancer cell sensitivity to chemotherapy. Additionally, our data suggest that SVCT2 silencing leads to the upregulation of ATP-binding cassette (ABC) transporter genes, further implicating SVCT2 in the development of multidrug resistance (MDR). ABC transporters are known to facilitate drug efflux, thereby decreasing intracellular drug accumulation and contributing to chemotherapy failure39,40,41.
Overall, this study clarifies the intricate relationship between ERα and SVCT2 in breast cancer, suggesting that ERα-mediated SVCT2 regulation could play a role in the development of drug resistance. By exploring these molecular interactions, we aim to offer new insights into potential therapeutic strategies that could enhance treatment outcomes for patients with hormone receptor-positive breast cancer.
Results
ERα knockdown leads to a decrease in SVCT2 level
We examined the endogenous expression levels of ERα and SVCT2 in various breast cancer cell lines, including MCF7, T47D, MDA-MB-231, and others (Fig. 1A, Supplementary Figure S1A, B ). A significant correlation was observed between ERα and SVCT2 expression levels across these cell lines. To further clarify the relationship between ERα and SVCT2 expression, we conducted knockdown experiments targeting ERα (Fig. 1B). Using type I siRNA against ERα in a concentration-dependent manner in MCF7 and T47D cells, we observed a corresponding decrease in SVCT2 expression, suggesting that SVCT2 expression depends on ERα levels (Fig. 1C, D, Supplementary Figure S1C, D). SVCT2 was primarily localized in the cytoplasm, where its levels decreased upon ERα inhibition, while ERα levels decreased in both the cytoplasmic and nuclear fractions. To further validate these findings, we introduced His-tagged ERα and Myc-tagged SVCT2 constructs into COS7 cells, which resulted in a similar decrease in SVCT2 expression as ERα expression declined (Fig. 1E). These observations suggest a significant association between ERα and SVCT2 expression in breast cancer cells, supporting a potential regulatory role of ERα in maintaining SVCT2 expression.
ERα knockdown decreases SVCT2 expression. (A) The expression levels of SVCT2 and ERα in breast cancer cells. GAPDH was used as a loading control. Data represent the mean ± SD of triplicate assays and were analyzed using Student’s t test. *p < 0.05, **p < 0.01 compared with MCF7. (B) MCF7 cells were transfected with scrambled siRNA, ERα-siRNA type I and ERα-siRNA type II. The protein and transcript expression levels of ERα were measured by western blotting and RT-PCR, respectively. Relative protein expression was quantified using densitometry. (C, D) MCF7 and T47D cells were transfected with increasing doses of si- ERα. The expression levels of SVCT2 and ERα were analyzed by western blotting. Relative protein expression was quantified using densitometry. (E) COS7 cells were transfected with Myc-DDK-SVCT2, His- ERα and increasing doses of si- ERα. The expression levels of SVCT2 and ERα were determined by western blotting using anti-DDK and anti-ERα antibodies. GAPDH was used as a loading control. Data represent the mean ± SD of triplicate assays and were analyzed using Student’s t test. *p < 0.05, **p < 0.01 compared with control.
ERα knockdown increases SVCT2 transcriptional activity
To elucidate the mechanism by which ERα regulates SVCT2, we investigated SVCT2 mRNA expression and transcriptional activity following ERα knockdown using real-time PCR and a luciferase assay, respectively. While ERα knockdown led to modest decrease in SVCT2 mRNA expression, these changes were not as pronounced as those observed at the protein level in MCF7 and T47D cells (Fig. 2A). In MCF7 cells, SVCT2 transcriptional activity showed a concentration-dependent increase in response to ERα knockdown, in contrast to the decrease observed in protein levels (Fig. 2B). Prior research suggests that p53 acts as a transcriptional repressor of SVCT2 and is also a downstream target of ERα regulation42. Consistent with this, we observed a reduction in p53 expression following ERα knockdown in both MCF7 and T47D cells (Fig. 2C, F). Additionally, to investigate the role of p53 as a transcriptional repressor of SVCT2, we conducted p53 knockdown experiments at varying concentrations. In both MCF7 (Fig. 2D, E) and T47D (Fig. 2G, H) cells, p53 knockdown led to increased SVCT2 expression and transcriptional activity. ERα overexpression did not significantly affect SVCT2 expression in a concentration-dependent manner across several cell lines (Supplementary Figure S2A–I), indicating that while ERα may influence SVCT2 transcriptional activity, its overexpression alone does not directly alter SVCT2 expression levels. These results suggest that ERα influences SVCT2 protein stability rather than its transcription, as shown by the mismatch between mRNA and protein levels following ERα knockdown. Additionally, p53 appears to act as a transcriptional repressor of SVCT2, potentially mediating ERα's regulatory effects on SVCT2 expression.
ERα is involved in SVCT2 transcription activity. (A) MCF7 and T47D cells were transfected with scrambled siRNA or si-ERα. ERα and SVCT2 transcript levels were measured by quantitative real-time PCR. (B) Knockdown of ERα affects SVCT2 luciferase activity. MCF7 cells were transfected with different type of si-ERα together with SVCT2-luciferase reporter plasmid. Luciferase activity was normalized by β‐galactosidase activity. (C, F) MCF7 and T47D cells were transfected with scrambled siRNA or si-ERα in different independent experiments. The expression levels of SVCT2, ERα and p53 were determined by western blotting. (D, G) MCF7 and T47D cells were transfected with increasing doses of si-p53. The expression levels of SVCT2 and p53 were analyzed by western blotting. GAPDH was used as a loading control. (E, H) MCF7 and T47D cells were transfected with increasing doses of si-p53 together with SVCT2-luciferse reporter plasmid. Luciferase activity was normalized by β‐galactosidase activity. Data represent the mean ± SD of triplicate assays and were analyzed using Student’s t test. *p < 0.05, **p < 0.01 compared with control (scrambled si-RNA).
ERα knockdown regulates the SVCT2 stability
Our previous results demonstrated that ERα knockdown significantly reduced SVCT2 protein levels, with no notable changes in SVCT2 mRNA levels, indicating that this reduction is likely regulated through protein degradation mechanisms. To explore whether ERα knockdown impacts SVCT2 protein stability, we conducted a stability assay using cycloheximide, a protein synthesis inhibitor. The results confirmed a modest reduction in SVCT2 protein stability under ERα knockdown conditions compared to the control group in both MCF7 and T47D cells (Fig. 3A, C). To determine if the observed decrease in SVCT2 with ERα knockdown was due to proteasomal degradation, we applied the proteasome inhibitor MG132. This treatment reversed the decrease in SVCT2 levels, while the lysosome inhibitor leupeptin had no effect (Fig. 3B, D). These findings indicate that ERα knockdown promotes SVCT2 degradation through the ubiquitin–proteasome pathway, suggesting a regulatory role for ERα in maintaining SVCT2 protein stability.
ERα knockdown led to a reduction in the stability of SVCT2. (A, C) MCF7 and T47D Cells were transfected with scrambled siRNA or si-ERα and exposed to CHX (40 μM) for indicated times. Cell lysates were prepared for western blot analysis. (B, D) Cells were transfected with scrambled siRNA or si-ERα and exposed to MG132 (10 μM) or Leupeptin (10 μM) for 24 h, and whole cell lysates were immunoblotted as indicated. GAPDH was used as an internal control.
ERα knockdown promotes ubiquitin-mediated proteasomal degradation of SVCT2
To investigate the mechanism underlying ERα knockdown-induced destabilization of SVCT2, we examined the interaction between ERα and SVCT2. Immunoprecipitation (IP) assays revealed that ERα binds directly to SVCT2 (Supplementary Figure S3A- C). To identify the specific E3 ligase responsible for targeting SVCT2 for ubiquitination under ERα-deficient condition, we conducted QuantSeq 3’mRNA analysis in cells treated with scrambled si-RNA and si-SVCT2 (Fig. 4A). We focused on E3 ligases that displayed at least a 1.2-fold increase in expression compared to the scramble control under SVCT2 knockdown conditions. Real-time PCR confirmed significant upregulation of several E3 ligases, including XIAP, NEURL1, NEDD4, MDM2, and TRIM71. Among these, XIAP was identified as the primary E3 ligase involved in SVCT2 ubiquitination post-ERα knockdown (Fig. 4B–F). Additionally, XIAP protein levels increased proportionally with ERα knockdown (Fig. 4G, H). In order to further investigate the intracellular localization of XIAP, SVCT2, and ERα, we performed subcellular fractionation in the breast cancer cell lines MCF7 and T47D. ERα was present in both cytoplasmic and nuclear fractions, while SVCT2 and XIAP were primarily cytoplasmic (Supplementary Figure S4A, B). We also investigated the interactions between XIAP and ERα, as well as XIAP and SVCT2, through additional IP assays. No interaction was observed between XIAP and ERα; however, a direct interaction between XIAP and SVCT2 was confirmed (Supplementary Figure S5A, B). Notably, XIAP knockdown restored SVCT2 protein levels in cells with reduced ERα, indicating that XIAP functions as an E3 ligase for SVCT2 (Fig. 4I, J). Further tests demonstrated that XIAP facilitated SVCT2 ubiquitination, as indicated by experiments using either Flag-XIAP or si-XIAP (Fig. 4K, L). Collectively, these findings establish XIAP as a key regulator of ubiquitin-dependent SVCT2 degradation following ERα knockdown.
XIAP mediates proteasomal degradation of SVCT2. (A) QuantSeq 3′mRNA analysis was conducted to assess gene expression in cells treated with either scrambled siRNA or si-SVCT2. The expression of E3 ligase-associated genes was compared between these two conditions. (B, D) MCF7 and T47D cells were transfected with scrambled siRNA or si-SVCT2. (C, E) MCF7 and T47D cells were transfected with scrambled siRNA or si-ERα. Total RNA was extracted, and the expression of endogenous E3 ligase genes—XIAP, NEURL1, NEDD4, MDM2, and TRIM71—was quantified using real-time PCR. (F) The expression of XIAP in MCF7 and T47D cells following transfection with scrambled siRNA or si-ERα was measured by real-time PCR. Statistical significance was analyzed using a Student’s t-test (*p < 0.05, **p < 0.01 compared with control (scrambled si-RNA). (G, H) MCF7 and T47D cells were transfected with increasing doses of si-ERα, and the expression levels of XIAP and ERα were evaluated via Western blotting. (I, J) XIAP and ERα were silenced individually or in combination in MCF7 and T47D cells using siRNA, followed by analysis of SVCT2 levels through Western blotting. (K) MCF7 cells were transiently transfected with various combinations of vectors encoding GFP-SVCT2, Flag-XIAP, HA-Ub, and si-ERα. Cells were treated with 10 μM MG132 for 4 h before lysate preparation. After 48 h post-transfection, cell lysates were immunoprecipitated with an anti-HA antibody, and SVCT2 polyubiquitination was detected by immunoblotting with an anti-GFP antibody. (L) MCF7 cells were transiently transfected with various combinations of vectors encoding Myc-DDK-SVCT2 and si-XIAP and/or si-ERα. Cells were treated with 10 μM MG132 for 4 h before lysate preparation. After 48 h post-transfection, cell lysates were immunoprecipitated with an anti-DDK antibody, and polyubiquitination was detected by immunoblotting with an anti-ubiquitin antibody.
Reduction of SVCT2 by ERα knockdown contributes to doxorubicin resistance
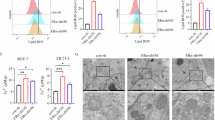
Ascorbic acid has been widely used as an adjunctive treatment in cancer, with SVCT2 recognized as a key transporter in its cellular uptake25. A recent study provided insights into ascorbic acid’s therapeutic potential, especially in cancers with KRAS or BRAF mutations38. We assessed ascorbic acid uptake following ERα knockdown and found a dose-dependent decrease in uptake in both MCF7 and T47D cells (Fig. 5A, C). Similarly, SVCT2 knockdown produced a comparable dose-dependent reduction in ascorbic acid uptake across these cell lines (Fig. 5B, D). These findings suggest that the diminished ascorbic acid uptake observed after ERα knockdown is likely due to decreased SVCT2 expression. To further understand the interaction between ascorbic acid and doxorubicin, we evaluated cell viability in MCF7 and T47D cells treated with each agent separately and in combination (Fig. 6A-D). Both ascorbic acid and doxorubicin independently reduced cell viability in a dose-dependent manner. Notably, combined treatment with ascorbic acid and doxorubicin enhanced doxorubicin’s cytotoxic effects (Fig. 6E, F). However, under ERα knockdown conditions, the co-treatment effect was diminished, showing that ERα knockdown reduced doxorubicin-induced cell death, whether in the presence or absence of ascorbic acid (Fig. 6G, H). This finding suggests that ERα knockdown promotes resistance to doxorubicin in breast cancer cells. To determine if this resistance is due to decreased SVCT2 expression, we examined cell viability and apoptosis under SVCT2 knockdown conditions. Post-doxorubicin treatment, cells with SVCT2 knockdown demonstrated lower cell death rates compared to controls (Fig. 7A, B). These results indicate that SVCT2 suppression may contribute to resistance against doxorubicin-induced apoptosis, highlighting a possible mechanism of drug resistance in breast cancer cells.
ERα knockdown induces inhibition of ascorbic acid uptake. (A-D) Cells were transfected with scrambled siRNA, si-ERα and si-SVCT2, followed by exposure to ascorbic acid at the indicated doses for 24 h. Ascorbic acid uptake was assessed using the EZ-Ascorbic Acid Assay kit. The data are presented as the fold ratio of ascorbic acid uptake in cells transfected with si-ERα relative to the control. Data represent the mean ± SD of triplicate assays and were analyzed using Student’s t test. *p < 0.05, **p < 0.01 compared with control.
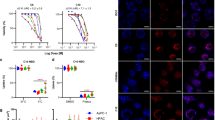
ERα knockdown induces resistance to doxorubicin. (A, B) MCF7 and T47D were treated with the indicated concentrations of ascorbic acid for 24 h, and cell viability was measured using the MTS method. (C, D) MCF7 and T47D cells were treated with the indicated concentrations of doxorubicin for 24 h. Cell viability was measured using the MTS method. (E, F) MCF7 and T47D cells were treated with or without 0.5 mg/ml ascorbic acid and 200 μM doxorubicin for 24 h, and cell viability was measured using the MTS method. MCF7 and T47D cells were transfected with scrambled siRNA or si-ERα and treated with or without 0.5 mg/ml ascorbic acid and 200 μM doxorubicin for 24 h. Statistical significance was determined by Student’s t- test. (*p < 0.05, **p < 0.01 compared with the untreated group; #p < 0.05, ##p < 0.01 compared with single-drug treatment) (G, H) Cell viability was measured by the MTS method. The results are representative of three independent experiments. Statistical significance was determined by Student’s t- test. (*p < 0.05, **p < 0.01 compared with the untreated group; #p < 0.05, ##p < 0.01 compared with single-drug treatment).
SVCT2 is involved in resistance to doxorubicin. MCF7 and T47D cells were transfected with scrambled siRNA or si-SVCT2 and treated with or without 0.5 mg/ml ascorbic acid and 200 μM doxorubicin for 24 h. (A, B) Cell viability was measured by the MTS. The results are representative of three independent experiments. Statistical significance was determined by Student’s t- test. (*p < 0.05, **p < 0.01 compared with the untreated group; #p < 0.05, ##p < 0.01 compared with single-drug treatment).
SVCT2 silencing modulates the expression of ABC transporter genes
Drug resistance in cancer cells is frequently linked to the overexpression of ABC transporter genes. These genes encode proteins that function as efflux pumps, actively transporting chemotherapeutic agents out of the cells, thereby reducing intracellular drug concentrations and diminishing their cytotoxic effects. Consequently, the overexpression of ABC transporters can lead to decreased drug accumulation in cancer cells, thereby reducing treatment efficacy and promoting multidrug resistance. Recognizing the role of ABC transporter genes in drug resistance is crucial for developing strategies to overcome this challenge and improve therapeutic outcomes. Building on prior data (Figs. 5–7) suggesting that SVCT2 knockdown may contribute to drug resistance, we performed QuantSeq 3’ mRNA analysis under SVCT2 knockdown conditions (Fig. 8A). This analysis revealed a notable upregulation of ABC transporter genes when SVCT2 expression was reduced. Real-time PCR further confirmed the increased expression of ABC transporter genes in MCF7 and T47D breast cancer cell lines, specifically under conditions of reduced SVCT2 levels due to ERα knockdown (Fig. 8B, C). Consistent with this, ABC transporter gene expression was also elevated under direct SVCT2 knockdown (Fig. 8D, E). These findings demonstrate that reduced SVCT2 levels correlate with increased expression of ABC transporters in both ERα and SVCT2 knockdown conditions. Among these genes, ABCA12 expression exhibited more than a two-fold increase. ABCA12, part of the ABC transporter family’s subfamily A, primarily facilitates lipid transport across cell membranes, contributing to the formation of the skin’s lipid barrier. In cancer, ABCA12 is linked to drug resistance by exporting chemotherapeutic agents from cells, thereby lowering their intracellular levels and efficacy. Additionally, ERα knockdown led to a dose-dependent increase in ABCC1 protein levels in both MCF7 and T47D cells (Supplementary Figure S6A, B). ABCC1, also known as multidrug resistance protein 1 (MRP1), is a key ABC transporter involved in the efflux of various substrates, including chemotherapeutic drugs, out of cells. Its overexpression is closely associated with multidrug resistance, contributing significantly to chemotherapy failure in cancer treatment. In summary, these results suggest that SVCT2 knockdown may enhance drug resistance in breast cancer cells by upregulating specific ABC transporter genes, highlighting a potential mechanism for resistance to chemotherapy.
SVCT2 silencing affects the expression of ABC transporter genes. (A) QuantSeq 3’mRNA analysis was performed to compare gene expression in cells treated with scrambled siRNA and si-SVCT2. The expression of genes associated with the ABC transporter was compared between the two conditions. (B, C) MCF7 and T47D were transfected with scrambled siRNA or si-ERα. (D, E) MCF7 and T47D were transfected with scrambled siRNA or si-SVCT2. Total RNA was prepared and the expression of the endogenous ABC transporter genes, ABCA12, ABCB8, ABCC1, ABCC10, ABCG1, ABCG2 and ABCG4 were determined by real-time PCR. Statistical significance was determined by Student’s t- test with *p < 0.05, **p < 0.01.
Discussion
The results of our study provide valuable insights into the mechanisms by which ERα knockdown influences SVCT2 expression and its subsequent effects on drug resistance in breast cancer cells. One of the primary objectives of this study was to assess how ERα knockdown influences SVCT2 expression. We observed a strong correlation between ERα and SVCT2 protein levels, particularly in breast cancer cell lines MCF7 and T47D, where ERα knockdown led to a concurrent decrease in SVCT2 protein levels. Notably, while the reduction in SVCT2 protein levels was substantial, changes in SVCT2 mRNA levels were modest. This discrepancy suggests that ERα likely regulates SVCT2 primarily at the post-translational level rather than through direct transcriptional control. We hypothesized that ERα might be involved in stabilizing SVCT2 protein rather than directly modulating its transcription. To further elucidate the mechanisms underlying this regulation, we examined potential pathways associated with SVCT2 destabilization following ERα knockdown. Using cycloheximide, a protein synthesis inhibitor, we found that SVCT2 protein stability significantly decreased in the absence of ERα, supporting the role of ERα in stabilizing SVCT2 at the protein level. This aligns with our observation that ERα overexpression did not lead to significant changes in SVCT2 protein levels, suggesting a more complex regulatory relationship beyond transcriptional control. A key finding in this study was the involvement of the ubiquitin–proteasome pathway in SVCT2 degradation following ERα knockdown. Proteasome inhibition via MG132 restored SVCT2 levels, implying that ERα knockdown induces SVCT2 degradation through this pathway. Further investigation into the specific E3 ligases responsible for tagging SVCT2 for degradation revealed XIAP as a critical regulator. QuantSeq 3’mRNA analysis and subsequent real-time PCR experiments indicated that XIAP expression increased in response to ERα knockdown. Immunoprecipitation analysis showed an interaction between XIAP and SVCT2, suggesting that XIAP, regulated by ERα signaling, directly contributes to SVCT2 degradation. The identification of XIAP as the E3 ligase responsible for SVCT2 ubiquitination is particularly significant because XIAP has well-established roles in regulating cell survival and apoptosis, making it a potentially attractive target for cancer therapies43,44. Our findings confirmed that XIAP knockdown restored SVCT2 protein levels, highlighting XIAP’s role in facilitating SVCT2’s ubiquitination and proteasomal degradation. Interestingly, although ERα knockdown reduced SVCT2 protein levels, it led to increased SVCT2 promoter activity in our luciferase assay (Fig. 2A, B), suggesting a dual and context-dependent role of ERα in regulating SVCT2. Mechanistically, ERα promotes SVCT2 protein stability by suppressing XIAP-mediated ubiquitination and proteasomal degradation, as demonstrated in our study. At the transcriptional level, however, ERα may act indirectly through p53. Previous studies have shown that p53 functions as a transcriptional repressor of SVCT2 and is also a downstream target negatively regulated by ERα42. Consistent with this, we observed decreased p53 expression following ERα knockdown in both MCF7 and T47D cells (Fig. 2C, F), which may relieve repression on the SVCT2 promoter and account for the enhanced luciferase activity. These findings suggest that ERα modulates SVCT2 expression through distinct mechanisms at the transcriptional and post-translational levels, and that compensatory transcriptional upregulation may occur in response to reduced SVCT2 protein stability. This regulatory axis among ERα, XIAP, and SVCT2 underscores the importance of post-translational regulation in breast cancer and suggests a potential therapeutic target for stabilizing SVCT2 in ERα-positive cases.
Another significant aspect of our study was the functional link between ERα, SVCT2, and ascorbic acid uptake. SVCT2 is crucial for ascorbic acid transport, and its downregulation impacts cellular uptake of this nutrient, which plays essential roles in antioxidant defense and other metabolic processes within cells45,46. ERα knockdown decreased both SVCT2 protein levels and ascorbic acid uptake in a dose-dependent manner, further supporting ERα’s role in maintaining cellular ascorbic acid homeostasis through SVCT2. Interestingly, although ERα knockdown significantly reduced SVCT2 protein levels and ascorbic acid uptake, ERα overexpression did not have a similar effect, suggesting a complex regulation of SVCT2 by ERα that might involve indirect signaling pathways or interactions with other regulatory proteins. Additionally, p53, a known transcriptional repressor of SVCT242, showed decreased expression following ERα knockdown, further contributing to the observed changes in SVCT2 expression and activity. This highlights the interaction of multiple regulatory factors, including ERα, XIAP, and p53, in modulating SVCT2 stability. The reduction in ascorbic acid uptake observed with SVCT2 knockdown has broader implications for cancer cell metabolism and drug resistance. Previous research has indicated that ascorbic acid supplementation can enhance the efficacy of certain chemotherapeutic agents, including doxorubicin47,48. In our study, we found that knockdown of either SVCT2 or ERα led to increased resistance to doxorubicin treatment in breast cancer cells, as evidenced by decreased cell death and enhanced cell viability following doxorubicin exposure. This suggests that SVCT2 depletion may reduce doxorubicin efficacy, potentially by altering cellular redox states and impeding the accumulation of oxidative stress, one of the mechanisms by which doxorubicin induces cancer cell death.
Our findings also propose a mechanism by which SVCT2 knockdown confers drug resistance through the upregulation of ABC transporter genes. ABC transporters are well-known for their role in mediating multidrug resistance in cancer cells, actively expelling chemotherapeutic agents from cells, thereby reducing their intracellular concentration and effectiveness49,50,51. Notably, previous studies have reported that the expression of certain ABC transporters is influenced by ERα. For example, ABCG2 (breast cancer resistance protein, BCRP) expression is known to be regulated by ERα in hormone-responsive breast cancer cells, with some studies indicating ERα-mediated downregulation. Similarly, ABCB1 (MDR1), a well-known efflux transporter, is also affected by estrogen signaling in various cancer models52. These findings, which highlight the intricate relationship between ERα and ABC transporter regulation, support the possibility that the upregulation of specific ABC transporter genes observed in our ERα knockdown models may result from compensatory transcriptional or post-transcriptional mechanisms in response to the loss of ERα signaling. Through QuantSeq 3’mRNA analysis, we observed a significant increase in the expression of several ABC transporter genes, particularly ABCA12 and ABCC1, following SVCT2 knockdown. These genes, associated with the efflux of various substrates including chemotherapeutics, have been linked to resistance to several treatments53,54,55. The upregulation of ABC transporter genes in response to SVCT2 knockdown suggests a previously unrecognized role of SVCT2 in regulating transporter gene expression, which may reduce breast cancer cell sensitivity to doxorubicin as these transporters decrease the drug’s intracellular concentration. This finding provides new insight into the relationship between SVCT2 and drug resistance, underscoring the potential to target SVCT2 or the associated ABC transporter pathway to improve chemotherapy efficacy.
In conclusion, this study provides a comprehensive view of the molecular mechanisms by which ERα knockdown modulates SVCT2 expression and its relevance to drug resistance in breast cancer. Identifying XIAP as the E3 ligase responsible for SVCT2 degradation opens new avenues for therapeutic intervention, notably in ERα-positive breast cancers with impaired SVCT2 stability. Furthermore, the association of SVCT2 downregulation with reduced ascorbic acid uptake and increased drug resistance via ABC transporter gene upregulation underscores the complex regulatory networks that contribute to the development of chemoresistance in breast cancer.
Future studies should explore the therapeutic potential of targeting the ERα–XIAP–SVCT2 pathway, either by inhibiting XIAP-mediated SVCT2 degradation or by restoring SVCT2 levels to sensitize cancer cells to chemotherapeutic agents. Additionally, examining the role of ABC transporter genes in mediating drug resistance in the context of SVCT2 downregulation may provide deeper insights into overcoming multidrug resistance in breast cancer treatment. These findings suggest opportunities for enhancing therapeutic strategies and highlight the importance of understanding complex molecular interactions in cancer progression and treatment resistance.
Materials and methods
Cell culture and materials
MCF7, T47D, MDA-MB-231, COS7, Hela and HEK293 cells were purchased from Korea Cell Line Bank (Seoul, Korea). The cells were maintained in DMEM with high glucose or RPMI1640 medium (Cytiva, Marlborough, MA, USA) supplemented with 10% FBS and antibiotics (Gibco, Waltham, MA, USA) under 5% CO2 at 37 °C. L-Ascorbic acid (A4544), Cycloheximide (C1988), MG132 (M8699), leupeptin (L2884) and DAPI (D9564) were purchased from Sigma-Aldrich (St. Louis, MO, USA). Doxorubicin was purchased from FutureChem Co., Ltd. (Seoul, Korea). Primary antibodies used in immunoprecipitation (IP) and western blotting (WB) are as follows. Anti-ERα (SC-8002), anti-p53 (SC-126), anti-GAPDH (SC-47724), anti-Ub (SC-8017), anti-HA (SC-7392), anti-Lamin A (SC-518013) and β-actin (SC-47778) were from Santa Cruz Biotechnology (Santa Cruz, CA, USA). Anti-DDK (TA50011-100) and turbo GFP (TA150041) were from Origene (Rockville, MD, USA). Anti-SVCT2 (NBP2-13,319) was from Novus (Centennial, CO, USA). Anti-XIAP (#14,334), anti-Flag (#14792S) and anti-ABCC1 (#72,202) were from Cell Signaling Technology (Boston, MA, USA).
Expression plasmids and siRNAs
Myc-DDK-SVCT2 and Myc-DDK-Mock were purchased from Origene (Rockville, MD, USA). His-ERα plasmids were kindly provided by Dr. Hee Min Yoo (Korea Research Institute of Standards and Science, Korea). HA-ERα and HA-Ub plasmids were kindly provided by Prof. Jong-Ho Kim (Kyung Hee University, Korea). Flag-XIAP plasmids were generously provided by Prof. Young J. Oh (Yonsei University, Korea). siRNA duplexes against ERα (2099), p53 (7157), SVCT2 (9962), XIAP (331) and control siRNA duplex served as a negative control were synthesized by Bioneer Inc. (Daejeon, Korea). Transfection was performed using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocol.
Western blotting and immunoprecipitation
The cells were washed with ice cold PBS and lysed in RIPA buffer containing 1% NP40, 0.5% sodium deoxycholate, 0.1% SDS and a protease inhibitor cocktail. The lysates were subjected to SDS-PAGE and transferred to a nitrocellulose (NC) membrane (Cytiva, Pittsburgh, PA, USA). The membranes were blocked with TBS buffer containing 0.1% Tween 20 and 5% skim milk and then incubated with the primary antibody. The immunoreactive bands were detected by using enhanced chemiluminescence (ECL) (Bio-Rad, Hercules, CA, USA) after incubation with an appropriate secondary antibody. For immunoprecipitation, the cells were washed thoroughly with ice-cold PBS to remove any residual media and debris. Subsequently, they were lysed using a buffer composed of 10 mM Tris–HCl (pH 7.4), 150 mM NaCl, 1 mM EDTA, 1 mM EGTA, 0.1% Nonidet P-40, 1% TX-100, 0.2 mM PMSF, and a 100X protease inhibitor mixture (Cell Signaling Technology, Boston, MA, USA) to ensure protein preservation. The resulting cell lysates were centrifuged at 13,000 rpm for 20 min at 4 °C to pellet the cell debris. The supernatants, containing the soluble proteins, were carefully collected and their protein concentration was measured using a protein assay reagent (Bio-Rad, Hercules, CA, USA). These lysates were then incubated overnight at 4 °C with specific antibodies, gently rotating to allow proper binding. To capture the antigen–antibody complexes, 50 µL of protein A/G-PLUS agarose beads (Santa Cruz Biotechnology) were added. The beads were washed five times with the lysis buffer, each wash involving centrifugation at 3000 rpm for 2 min at 4 °C to remove non-specifically bound proteins. Finally, the proteins bound to the beads were separated by SDS-PAGE and analyzed by immunoblotting to detect the proteins of interest. The data were visualized using ImageQuant™ 800 system (Cytiva). All western blot images were processed uniformly. Cropping was performed to focus on relevant bands, without altering the data. Full-length blots are available upon request. In some experiments, membranes were cut prior to hybridization with antibodies to allow probing for multiple proteins on a single blot. As a result, full-length blots may not be available in all cases.
Luciferase assay
Cells were transiently co-transfected with 0.5 μg SVCT2 luciferase plasmids56 and 0.5 μg of the pSV-β-galactosidase reporter vector, utilizing Lipofectamine 2000 (Invitrogen) as the transfection agent. Following transfection, luciferase activity was assessed using the Luciferase Assay System (Promega, Madison, WI, USA) and measured with a TECAN Spark microplate reader (TECAN, Männedorf, Switzerland). To ensure accurate normalization, β-galactosidase activity was quantified using the β-galactosidase enzyme assay system (Promega).
Quantitative real-time PCR and semi-quantitative reverse transcription-PCR (RT-PCR).
Total RNA was extracted with Trizol (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocol. Then, 1 μg of total RNA was used for cDNA synthesis using the iScript cDNA Synthesis Kit (Bio-Rad). Gene expression was then measured by iQ SYBR green Supermix (Bio-Rad). The relative expression levels of genes were calculated based on glyceraldehyde phosphate dehydrogenase (GAPDH) using 2−△△CT equation. The primers of genes were designed and purchased from Macrogen Inc. (Seoul, Korea). SVCT2 F: 5’- CGATCAAGTGTTGAACGTCCT-3’, R: 5’- CGATCAAGTGTTGAACGTCCT-3’; ERα F: 5’- TGATGAAAGGTGGGATACGA-3’, R: 5’- AAGGTTGGCAGCTCTCATGT −3’; ABCA12 F: 5’- CTCACAGCATGGAAGAATGTG −3’, R: 5’- AGAGTGGTCTGACTCACTAAG −3’; ABCB8 F: 5’- CTGGAAGCTTCCGATGAAGAG −3’, R: 5’- TTCAGGAGCTCTTCATGTGTC −3’; ABCC1 F: 5’- GACACAGTGGACTCCATGATC −3’, R: F’- CCACCAAGCCAGCACTGAGGC −3’; ABCC10 F: 5’- TCCCTGTTGTTGGTGCTCTTC −3’, R: 5’- TCTGAGTTCAGGATCGTGTTG −3’; ABCG1 F: 5’- TTCAGATCATGTTCCCAGTGG −3’, R: 5’- GAGGACAAAATAGGCAATGAG −3’; ABCG2 F: 5’- TCAGGAAGACTTATGTTCCAC −3’, R: 5’- AGCTCTGTTCTGGATTCCAGT −3’; ABCG4 F: 5’- TATGGCTGAGAAGAAGAGCAG −3’, R: 5’- AAGGTGAGCACAGTTGGCATG −3’; GAPDH F: 5’- GCAAATTCCATGGCACCGT-3’, R: 5’- TCGCCCCACTTGATTTTGG-3’; XIAP F: 5’- TGGCAGATTATGAAGCACGGATC −3’, R: 5’- AGTTAGCCCTCCTCCACAGTGA −3’; RFFL F: 5’- TGGAACCAAGCTGCAAGTCCTG −3’, R 5’- GAAACCGTTGGCAGAGAAGGCA −3’; NEURL1 F: 5’- CCTCATCTACGAGCAAGTCAGG −3’, R: 5’- AGAAGCCACTCTGGGACACCAG −3’; NEDD4 5’- CAGAAGAGGCAGCTTACAAGCC −3’, R: 5’- CTTCCCAACCTGGTGGTAATCC −3’; MDM2 F: 5’- TGTTTGGCGTGCCAAGCTTCTC −3’, R: 5’- CACAGATGTACCTGAGTCCGATG −3’; TRIM71 F: 5’- CGAGGCATAAGAAAGCCCTGGA −3’, R: 5’- GCTTGTTGAGGTTTTGCCGCAG −3’. The target genes were amplified by reverse transcription using specific primers: ERα sense 5’- CGACGCCAGGGTGGCAGAGAAAGATT −3’ and antisense 5’- GGCCAAAGGTTGG-CAGCTCTCATGTC −3’; GAPDH sense 5’- ATCTTCCAGGAGCGA-GATCCC −3’ and antisense 5’- AGTGAGCTTCCCGTTCAGCTC −3’. GAPDH was amplified as a normalization control. The PCR bands were detected following electrophoresis in a 2% agarose gel in 1 × Tris–acetate-EDTA (TAE) buffer and stained with Midori Green Direct (Nippon Genetic, Tokyo, Japan). The resulting data were integrated and analyzed using Quantity One software (Bio-Rad).
Immunofluorescence
Cells were fixed with 4% paraformaldehyde in phosphate-buffered saline (PBS) for 15 min at room temperature, followed by permeabilization with 0.2% Triton X-100 in PBS for 10 min. They were blocked with 5% bovine serum albumin (BSA) in PBS for 1 h at room temperature. Subsequently, the cells were incubated overnight at 4 °C with anti-ERα antibody (SC-8002). After washing, the cells were stained with appropriate secondary antibodies. Nuclei were counterstained with DAPI for 5 min. Fluorescence images were acquired using an Axio Vert.A1 by Carl Zeiss (Germany).
Subcellular fractionation
Cells were washed with ice-cold PBS and then incubated on ice in cytosol buffer (10 mM NaCl, 10 mM HEPES, 0.1 mM EDTA, 0.1 mM EGTA, 1% NP-40) for 10 min, followed by lysis using Dounce homogenization. The lysates were centrifuged for 7 min at 500 × g, and the supernatants were collected as the cytoplasmic fraction. The nuclear pellets were washed multiple times with cytosol buffer and then resuspended in nuclear buffer (20 mM HEPES, 25% glycerol, 0.4 M NaCl, 1 mM EDTA, 1 mM EGTA) for 15 min. Subsequently, the samples were centrifuged at 20,000 × g for 15 min, and the supernatant was collected as the nuclear fraction. Lamin A and β-actin were used as markers for nuclear and cytoplasmic proteins, respectively.
Library preparation and sequencing
Total RNA was isolated using Trizol reagent (Invitrogen). RNA quality was assessed by Agilent TapeStation 4000 system (Agilent Technologies, Amstelveen, The Netherlands), and RNA quantification was performed using ND-2000 Spectrophotometer (Thermo Inc., DE, USA).
For library preparation of control and test samples, the QuantSeq 3’ mRNA-Seq Library Prep Kit (Lexogen, Inc., Austria) was employed according to the manufacturer’s protocol. Briefly, total RNA was combined with an oligo-dT primer that included an Illumina-compatible sequence at its 5’ end, followed by reverse transcription. After RNA template degradation, second-strand synthesis was initiated with a random primer carrying an Illumina-compatible linker sequence at the 5’ end. The double-stranded library was purified using magnetic beads to eliminate residual reaction components. Subsequently, the library was amplified to add the full adapter sequences necessary for cluster generation. The finalized library was further purified to remove PCR contaminants. High-throughput sequencing was performed as single-end 75 sequencing using NextSeq 550 (Illumina, Inc., USA).
Data analysis
QuantSeq 3’ mRNA-Seq reads were aligned using Bowtie2 (Langmead and Salzberg, 2012). Bowtie2 indices were generated from either the genome assembly sequence or representative transcript sequences, facilitating alignment to both genome and transcriptome references. The resulting alignment files were used to assemble transcripts, estimate their abundances, and identify differential gene expression. Differentially expressed genes were determined by calculating unique and multiple alignment counts and analyzing coverage with Bedtools (Quinlan, 2010). Read count (RC) data were normalized using the TMM + CPM method within the EdgeR package in R (R Development Core Team, 2020) via Bioconductor (Gentleman et al., 2004). Gene classification was performed using searches through DAVID (http://david.abcc.ncifcrf.gov/) and Medline databases (http://www.ncbi.nlm.nih.gov/). Data mining and graphic visualization were performed using ExDEGA (Ebiogen Inc., Korea).
Ascorbic acid uptake assay
The ascorbic acid uptake assay was evaluated using the EZ-Ascorbic Acid Assay Kit (DG-ASC100, Seoul, Korea) according to the reported procedure. In this assay, the concentration of ascorbic acid was measured using a coupled enzyme reaction that produces a colorimetric product at 593 nm.
Cell viability assay
Cells were seeded at 1 × 104 cells per well on the 96-well plates, and cell viability was assayed using the CellTiter 96 AQueous One Solution Cell Proliferation Assay system (Promega) after incubating the cells with 10 μl of the CellTiter 96 AQueous One Solution reagent at 37 °C for 3 h. The absorbance was measured at 490 nm using a SpectraMax 190 microplate reader (Molecular Devices, Sunnyvale, CA, USA).
Statistical analysis
All experiments were independently repeated at least three times, and the data presented as the mean ± S.D. Statistical evaluation was assessed by the one-way analysis of variance (ANOVA). In each case, p < 0.05 was considered statistically significant.
Data availability
All data supporting the findings of this study are available within the paper and its Supplementary Information. The RNA-Seq sequencing data has been submitted to the NCBI Sequence Read Archive (SRA) (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1208065) and under accession number BioProject ID PRJNA1208065.
References
Harbeck, N. et al. Breast cancer. Nat. Rev. Dis. Primers 5, 66. https://doi.org/10.1038/s41572-019-0111-2 (2019).
Nolan, E., Lindeman, G. J. & Visvader, J. E. Deciphering breast cancer: From biology to the clinic. Cell 186, 1708–1728. https://doi.org/10.1016/j.cell.2023.01.040 (2023).
Liu, Y. et al. a key target for cancer therapy: A review. Onco. Targets Ther. 13, 2183–2191. https://doi.org/10.2147/OTT.S236532 (2020).
Tecalco-Cruz, A. C. & Ramirez-Jarquin, J. O. Mechanisms that increase stability of estrogen receptor alpha in breast cancer. Clin. Breast Cancer 17, 1–10. https://doi.org/10.1016/j.clbc.2016.07.015 (2017).
Kulkoyluoglu, E. & Madak-Erdogan, Z. Nuclear and extranuclear-initiated estrogen receptor signaling crosstalk and endocrine resistance in breast cancer. Steroids 114, 41–47. https://doi.org/10.1016/j.steroids.2016.06.007 (2016).
Farcas, A. M., Nagarajan, S., Cosulich, S. & Carroll, J. S. Genome-wide estrogen receptor activity in breast cancer. Endocrinology https://doi.org/10.1210/endocr/bqaa224 (2021).
Chen, P., Li, B. & Ou-Yang, L. Role of estrogen receptors in health and disease. Front. Endocrinol. (Lausanne) 13, 839005. https://doi.org/10.3389/fendo.2022.839005 (2022).
Thomas, C. & Gustafsson, J. A. Estrogen receptor mutations and functional consequences for breast cancer. Trends Endocrinol. Metab. 26, 467–476. https://doi.org/10.1016/j.tem.2015.06.007 (2015).
Yasar, P., Ayaz, G., User, S. D., Gupur, G. & Muyan, M. Molecular mechanism of estrogen-estrogen receptor signaling. Reprod. Med. Biol. 16, 4–20. https://doi.org/10.1002/rmb2.12006 (2017).
Bjornstrom, L. & Sjoberg, M. Mechanisms of estrogen receptor signaling: Convergence of genomic and nongenomic actions on target genes. Mol. Endocrinol. 19, 833–842. https://doi.org/10.1210/me.2004-0486 (2005).
Petrossian, K. et al. ERalpha-mediated cell cycle progression is an important requisite for CDK4/6 inhibitor response in HR+ breast cancer. Oncotarget 9, 27736–27751. https://doi.org/10.18632/oncotarget.25552 (2018).
Chimento, A. et al. Estrogen receptors-mediated apoptosis in hormone-dependent cancers. Int. J. Mol. Sci. https://doi.org/10.3390/ijms23031242 (2022).
Patel, H. K. & Bihani, T. Selective estrogen receptor modulators (SERMs) and selective estrogen receptor degraders (SERDs) in cancer treatment. Pharmacol. Ther. 186, 1–24. https://doi.org/10.1016/j.pharmthera.2017.12.012 (2018).
Fuentes, N. & Silveyra, P. Estrogen receptor signaling mechanisms. Adv. Protein Chem. Struct. Biol. 116, 135–170. https://doi.org/10.1016/bs.apcsb.2019.01.001 (2019).
Pegram, M., Jackisch, C. & Johnston, S. R. D. Estrogen/HER2 receptor crosstalk in breast cancer: Combination therapies to improve outcomes for patients with hormone receptor-positive/HER2-positive breast cancer. NPJ Breast Cancer 9, 45. https://doi.org/10.1038/s41523-023-00533-2 (2023).
Dalal, H. et al. Clinical associations of ESR2 (estrogen receptor beta) expression across thousands of primary breast tumors. Sci. Rep. 12, 4696. https://doi.org/10.1038/s41598-022-08210-3 (2022).
Brufsky, A. M. & Dickler, M. N. Estrogen receptor-positive breast cancer: Exploiting signaling pathways implicated in endocrine resistance. Oncologist 23, 528–539. https://doi.org/10.1634/theoncologist.2017-0423 (2018).
Hanker, A. B., Sudhan, D. R. & Arteaga, C. L. Overcoming endocrine resistance in breast cancer. Cancer Cell 37, 496–513. https://doi.org/10.1016/j.ccell.2020.03.009 (2020).
Osborne, C. K. & Schiff, R. Mechanisms of endocrine resistance in breast cancer. Annu. Rev. Med. 62, 233–247. https://doi.org/10.1146/annurev-med-070909-182917 (2011).
Cottu, P. et al. Acquired resistance to endocrine treatments is associated with tumor-specific molecular changes in patient-derived luminal breast cancer xenografts. Clin. Cancer Res. 20, 4314–4325. https://doi.org/10.1158/1078-0432.CCR-13-3230 (2014).
Yan, S. et al. Targeting the crosstalk between estrogen receptors and membrane growth factor receptors in breast cancer treatment: Advances and opportunities. Biomed. Pharmacother. 175, 116615. https://doi.org/10.1016/j.biopha.2024.116615 (2024).
Altwegg, K. A. & Vadlamudi, R. K. Role of estrogen receptor coregulators in endocrine resistant breast cancer. Explor. Target Antitumor. Ther. 2, 385–400. https://doi.org/10.37349/etat.2021.00052 (2021).
Park, S. S. et al. Mutation SVCT2 promotes cell proliferation, invasion and migration in colorectal cancer. J. Cancer 12, 5385–5393. https://doi.org/10.7150/jca.57463 (2021).
Wohlrab, C., Phillips, E. & Dachs, G. U. Vitamin C transporters in cancer: Current understanding and gaps in knowledge. Front. Oncol. 7, 74. https://doi.org/10.3389/fonc.2017.00074 (2017).
Linowiecka, K., Foksinski, M. & Brozyna, A. A. Vitamin C transporters and their implications in carcinogenesis. Nutrients https://doi.org/10.3390/nu12123869 (2020).
Burzle, M. et al. The sodium-dependent ascorbic acid transporter family SLC23. Mol. Aspects Med. 34, 436–454. https://doi.org/10.1016/j.mam.2012.12.002 (2013).
Ghanem, A. et al. Ascorbate kills breast cancer cells by rewiring metabolism via redox imbalance and energy crisis. Free Radic. Biol. Med. 163, 196–209. https://doi.org/10.1016/j.freeradbiomed.2020.12.012 (2021).
Hong, S. W. et al. SVCT-2 in breast cancer acts as an indicator for L-ascorbate treatment. Oncogene 32, 1508–1517. https://doi.org/10.1038/onc.2012.176 (2013).
Cho, S. et al. Hormetic dose response to (L)-ascorbic acid as an anti-cancer drug in colorectal cancer cell lines according to SVCT-2 expression. Sci. Rep. 8, 11372. https://doi.org/10.1038/s41598-018-29386-7 (2018).
Lv, H. et al. Vitamin C preferentially kills cancer stem cells in hepatocellular carcinoma via SVCT-2. NPJ Precis. Oncol. 2, 1. https://doi.org/10.1038/s41698-017-0044-8 (2018).
Wang, C. et al. SVCT-2 determines the sensitivity to ascorbate-induced cell death in cholangiocarcinoma cell lines and patient derived xenografts. Cancer Lett. 398, 1–11. https://doi.org/10.1016/j.canlet.2017.03.039 (2017).
Sasidharan Nair, V. & Huehn, J. Impact of vitamin C on the development, differentiation and functional properties of T cells. Eur. J. Microbiol. Immunol. (Bp) 14, 67–74. https://doi.org/10.1556/1886.2024.00017 (2024).
Zhitkovich, A. Nuclear and cytoplasmic functions of vitamin C. Chem. Res. Toxicol. 33, 2515–2526. https://doi.org/10.1021/acs.chemrestox.0c00348 (2020).
Mikkelsen, S. U., Gillberg, L., Lykkesfeldt, J. & Gronbaek, K. The role of vitamin C in epigenetic cancer therapy. Free Radic. Biol. Med. 170, 179–193. https://doi.org/10.1016/j.freeradbiomed.2021.03.017 (2021).
Mastrangelo, D., Pelosi, E., Castelli, G., Lo-Coco, F. & Testa, U. Mechanisms of anti-cancer effects of ascorbate: Cytotoxic activity and epigenetic modulation. Blood Cells Mol. Dis. 69, 57–64. https://doi.org/10.1016/j.bcmd.2017.09.005 (2018).
Bottger, F., Valles-Marti, A., Cahn, L. & Jimenez, C. R. High-dose intravenous vitamin C, a promising multi-targeting agent in the treatment of cancer. J. Exp. Clin. Cancer Res. 40, 343. https://doi.org/10.1186/s13046-021-02134-y (2021).
Salazar, K. et al. Role of vitamin C and SVCT2 in neurogenesis. Front. Neurosci. 17, 1155758. https://doi.org/10.3389/fnins.2023.1155758 (2023).
Yun, J. et al. Vitamin C selectively kills KRAS and BRAF mutant colorectal cancer cells by targeting GAPDH. Science 350, 1391–1396. https://doi.org/10.1126/science.aaa5004 (2015).
Rees, D. C., Johnson, E. & Lewinson, O. ABC transporters: the power to change. Nat. Rev. Mol. Cell Biol. 10, 218–227. https://doi.org/10.1038/nrm2646 (2009).
Alam, A. & Locher, K. P. Structure and mechanism of human ABC transporters. Annu. Rev. Biophys. 52, 275–300. https://doi.org/10.1146/annurev-biophys-111622-091232 (2023).
Xiao, H., Zheng, Y., Ma, L., Tian, L. & Sun, Q. Clinically-Relevant ABC transporter for anti-cancer drug resistance. Front. Pharmacol. 12, 648407. https://doi.org/10.3389/fphar.2021.648407 (2021).
Kim, E. H. et al. Role of p53 in transcriptional repression of SVCT2. Mol. Biol. Rep. 48, 1651–1658. https://doi.org/10.1007/s11033-021-06179-2 (2021).
Galban, S. & Duckett, C. S. XIAP as a ubiquitin ligase in cellular signaling. Cell Death Differ. 17, 54–60. https://doi.org/10.1038/cdd.2009.81 (2010).
Obexer, P. & Ausserlechner, M. J. X-linked inhibitor of apoptosis protein - a critical death resistance regulator and therapeutic target for personalized cancer therapy. Front. Oncol. 4, 197. https://doi.org/10.3389/fonc.2014.00197 (2014).
Han, Z. et al. New insights into Vitamin C function: Vitamin C induces JAK2 activation through its receptor-like transporter SVCT2. Int. J. Biol. Macromol. 173, 379–398. https://doi.org/10.1016/j.ijbiomac.2021.01.120 (2021).
Jung, S. A. et al. L-Ascorbic acid can abrogate SVCT-2-dependent cetuximab resistance mediated by mutant KRAS in human colon cancer cells. Free Radic. Biol. Med. 95, 200–208. https://doi.org/10.1016/j.freeradbiomed.2016.03.009 (2016).
Guo, D. et al. The involvement of ascorbic acid in cancer treatment. Molecules https://doi.org/10.3390/molecules29102295 (2024).
Heaney, M. L. et al. Vitamin C antagonizes the cytotoxic effects of antineoplastic drugs. Cancer Res. 68, 8031–8038. https://doi.org/10.1158/0008-5472.CAN-08-1490 (2008).
Moore, J. M., Bell, E. L., Hughes, R. O. & Garfield, A. S. ABC transporters: Human disease and pharmacotherapeutic potential. Trends Mol. Med. 29, 152–172. https://doi.org/10.1016/j.molmed.2022.11.001 (2023).
Thomas, C. & Tampe, R. Structural and mechanistic principles of ABC transporters. Annu. Rev. Biochem. 89, 605–636. https://doi.org/10.1146/annurev-biochem-011520-105201 (2020).
Sajid, A., Rahman, H. & Ambudkar, S. V. Advances in the structure, mechanism and targeting of chemoresistance-linked ABC transporters. Nat. Rev. Cancer 23, 762–779. https://doi.org/10.1038/s41568-023-00612-3 (2023).
Nedeljkovic, M., Tanic, N., Prvanovic, M., Milovanovic, Z. & Tanic, N. Friend or foe: ABCG2, ABCC1 and ABCB1 expression in triple-negative breast cancer. Breast Cancer 28, 727–736. https://doi.org/10.1007/s12282-020-01210-z (2021).
Lu, J. F., Pokharel, D. & Bebawy, M. MRP1 and its role in anticancer drug resistance. Drug Metab. Rev. 47, 406–419. https://doi.org/10.3109/03602532.2015.1105253 (2015).
Hendig, D. et al. Characterization of the ATP-binding cassette transporter gene expression profile in Y79: A retinoblastoma cell line. Mol. Cell. Biochem. 328, 85–92. https://doi.org/10.1007/s11010-009-0077-6 (2009).
Sasaki, N. et al. Alpha-fetoprotein-producing pancreatic cancer cells possess cancer stem cell characteristics. Cancer Lett. 308, 152–161. https://doi.org/10.1016/j.canlet.2011.04.023 (2011).
Jang, S. Y. et al. Curcumin inhibits human cancer cell growth and migration through downregulation of SVCT2. Cell Biochem. Funct. 41, 696–703. https://doi.org/10.1002/cbf.3824 (2023).
Funding
This research was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (NRF-2022R1A2C1004297 and RS-2023–00264573). M.T.R is funded by the European Research Council (G.A. no 101053122).
Author information
Authors and Affiliations
Contributions
Conceptualization, S.Y.J, C.H.Y., and S.P.; methodology, S.Y.J., J.K., E.H., Y.J., E.J., and Y.N.; software, S.Y.J.; validation, S.Y.J., Y.N., Y.J., Y.J.Y., M.T.R, and C.H.Y.; visualization, J.K. and E.H.; formal analysis, S.Y.J. J.K., E.H., and Y.J.; data curation, J.K., and E.H.; investigation, S.Y.J., J.K., E.H., and Y.J.; resources, J.H.K, Y.N. and C.H.Y.; writing-original draft preparation, S.Y.J., E.J., and S.P.; writing-review and editing, C.H.Y., Y.J.Y., M.T.R., and S.P.; supervision, S.P.; project administration, Y.N., Y.J.Y., E.J., M.T.R., and S.P.; funding acquisition, S.P. All Authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Jang, S.Y., Hong, E., Jo, Y. et al. Estrogen receptor α regulates SVCT2 protein level in human breast cancer cells. Sci Rep 15, 27629 (2025). https://doi.org/10.1038/s41598-025-11758-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-11758-5