Abstract
This study aims to explore how CPS1 influences the progression of lung adenocarcinoma by affecting the ammonia-induced ROS/AMPK/P53/LKB1 signaling pathway. Bioinformatics analysis was conducted to identify differential gene expression in lung adenocarcinoma patients. A549 cells were infected with control (NC) or CPS1 knockdown (CPS1-KD) lentivirus. Cells were treated with or without AMPK agonists, AMPK inhibitors, P53 agonists, or P53 inhibitors, followed by Western blot analysis of CPS1, NOX2, NOX4, p-AMPK, p-P53, and LKB1 protein levels. The content of MDA and SOD was measured, and the expression of AMPK, caspase-3 and P53 in tumor cells was detected through immunofluorescence. Apoptosis-related protein expression and tumor cell apoptosis were assessed using Western blot and flow cytometry. Tumor cell proliferation was evaluated using CCK-8 assays and colony formation experiments. Tumor size was measured in xenograft models using nude mice. Bioinformatics analysis indicated that LKB1 positively regulates AMPK activity. CPS1 knockdown results in increased ammonia levels, with upregulated expression of NOX2, NOX4, p-AMPK, p-P53, and LKB1 in tumor cells. Elevated P53 levels, along with significant increases in Bax, Caspase-8,and Caspase-12 expression, were observed, promoting apoptosis and inhibiting tumor cell proliferation. AMPK and P53 act to inhibit lung adenocarcinoma progression. CPS1 promotes the progression of lung adenocarcinoma by suppressing ammonia-induced activation of the ROS/AMPK/P53/LKB1 signaling pathway.
Similar content being viewed by others
Introduction
Lung cancer remains the leading cause of cancer-related mortality globally, with an estimated 1.8 million deaths annually, accounting for 18% of all cancer fatalities1. As the most prevalent malignant tumor in China and worldwide, lung adenocarcinoma (lung adenocarcinoma) represents the predominant pathological subtype, comprising approximately 40% of lung cancer cases2. Lung adenocarcinoma is one of the most common pathological types of lung malignant tumours, and the exploration of key genes involved in the development of lung adenocarcinoma is of great significance for the discovery of targets for the prevention and treatment of lung cancer and clinical drug development.Previous studies have shown that CPS1, as a key rate-limiting enzyme in the urea cycle, promotes tumor proliferation in malignant tumors such as liver cancer and colorectal cancer through purine metabolism or urea production3,4,5. However, the mechanism of its role in lung adenocarcinoma has not been fully elucidated. Although some studies suggest that increased CPS1 expression is associated with poor prognosis in lung adenocarcinoma6,7, the specific molecular mechanism, especially how CPS1 affects tumor progression through metabolic reprogramming, still has significant knowledge gaps. Additionally, the synergistic role of the ROS/AMPK/P53/LKB1 signaling pathway in tumor metabolic stress has been reported8,9,10, but whether this pathway is regulated by urea cycle enzymes (such as CPS1) has not been systematically studied.
Carbamoylphosphate synthase 1 (CPS1) catalyzes carbamoyl phosphate synthesis from bicarbonate, ammonia, and ATP, requiring N-acetylglutamate for activation11. As the primary urea cycle regulator in hepatic metabolism12, CPS1 dysfunction contributes to hyperammonemia and liver pathologies including fibrosis and chronic HCV infection13. Intriguingly, CPS1 is frequently upregulated in cancers. For instance, high CPS1 levels predict unfavorable outcomes in rectal cancer patients receiving neoadjuvant radiotherapy3. Similarly, CPS1 expression levels were significantly higher in lung adenocarcinoma than in adjacent non-cancerous tissues6,7. Furthermore, CPS1 knockdown reduced cell proliferation and levels of interorganelle metabolites associated with pyrimidine metabolism4. Notably, P53 suppresses uremia production and ammonia clearance by downregulating CPS1 transcription10. Thus, loss of P53 function during carcinogenesis elevates CPS1 expression, which may contribute to poor prognosis14. AMPK has been observed to result in a reduction in CPS1 expression levels within hepatocellular carcinoma cells and xenografts. These observations indicate the potential involvement of CPS1 in the pathogenesis of malignant tumours and cellular transformation5.
P53 is able to bind to LKB1 to form a complex, Upon binding to a specific complex and subsequent activation, LKB1 initiates a process of self-phosphorylation13, which in turn activates substrates15. Among these substrates, AMP-activated protein kinase (AMPK), a complex enzyme consisting of the catalytic subunit AMPKα and the two regulatory subunits AMPKβ and AMPKγ, is of particular importance for maintaining cellular energy homeostasis16. The LKB1-AMPK signalling pathway, formed by LKB1 and AMPK, is primarily responsible for regulating energy metabolism, cell polarity establishment, cell cycle progression, and apoptosis in order to achieve its tumour suppressor effect. It is, however, important to note that the activation of this pathway is frequently impaired in many cancer patients.
This study has for the first time revealed the oncogenic mechanism of CPS1 through regulating the imbalance of ammonia metabolism, which activates the ROS/AMPK/P53/LKB1 signaling axis. Specifically, we found that knocking down CPS1 led to ammonia accumulation, triggering mitochondrial dysfunction and activation of NOX enzymes, thereby inducing excessive ROS production; ROS oxidatively modified the AMPKα subunit and changed the AMP/ATP ratio, activating AMPK, which then phosphorylated P53 and promoted its formation with the LKB1 complex, ultimately driving the expression of pro-apoptotic proteins (such as Bax, Caspase-8/12), inhibiting tumor proliferation. This mechanism not only fills the functional gap of urea cycle enzymes in the metabolic reprogramming of lung adenocarcinoma, but also innovatively integrates the cross-level regulatory network of metabolism (ammonia), oxidative stress (ROS), and apoptosis pathways (AMPK/P53/LKB1).
Methods
Bioinformatics analysis
The dataset GSE10072 related to lung adenocarcinoma was downloaded from the Gene Expression Omnibus (GEO) database. This dataset contains gene expression data of 49 healthy patients and 58 patients with lung adenocarcinoma, providing a valuable resource for studying the molecular mechanisms of lung adenocarcinoma. To ensure the accuracy and reliability of the data, the chip data were background-corrected, normalized, and averaged for multiple probes using the Robust Multi-array Average method, ensuring the consistency and comparability of the data. The processed data also underwent batch-effect correction using the Combat method to eliminate systematic deviations introduced during the experiment. Next, differential expression analysis was conducted using the limma package, with the screening criteria set as |log2 Fold Change| > 1 and p-value < 0.05, to identify genes with significant differences between lung adenocarcinoma patients and healthy control groups. To control for false positive rates, the p-values were corrected using the Benjamini-Hochberg method for multiple hypothesis testing, ensuring the statistical significance of the results. In terms of data visualization, the ggplot2 package was used to present the results of the differential analysis graphically, allowing researchers to intuitively understand the changes in gene expression. Subsequently, the differentially expressed genes identified were subjected to Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis through the ClusterProfiler package to deeply explore the functions of these genes in biological processes and the key pathways they are involved in. The GO analysis mainly covered Cellular Component (CC), Molecular Function (MF), and Biological Process (BP), while the KEGG pathway analysis aimed to reveal important biological pathways related to lung adenocarcinoma. Finally, the Spearman statistical method was used to analyze the correlation between differentially expressed genes to explore the potential relationships between gene variables and further reveal the molecular characteristics and biological mechanisms of lung adenocarcinoma.
Cell culture and reagents
Human lung adenocarcinoma A549 cells were obtained from Wuhan Procell Life Technology. AMPK agonist GSK621, p 53 inhibitor Pifithrin-α (PFTα) HBr, AMPK inhibitor GSK690693, p 53 agonist Kevetrin Hydrochloride, RIPA lysis buffer, MDA assay kit (Beyotime, China), and SOD assay kit (Beyotime, China) were used. Additional materials included HRP-conjugated IgG, a microplate reader, confocal laser microscope, flow cytometer, CCK-8 kit, optical microscope, and 5-week-old BALB/c nude mice were purchased from Henan Scripps Bio-technology Co.
Lentiviral infection
Post 2 × 105 cell plating and overnight culture, infection ensured 30–50% cell confluence. Virus usage was calculated based on MOI and titer, with 1µL polybrene per 1mL culture medium, adding virus to each well. Culture medium was replaced 12 h post-infection with complete medium. Post-infection, 72-hour observation under a fluorescence microscope showed an infection efficiency above 85%. After 3 weeks of puromycin selection, no cell death was observed. Cells were collected and stored for subsequent use.
Cell culture and grouping
A549 cells were cultured in low-glucose DMEM with 1% serum at 37℃ and 1% O2. Cells were divided into groups: NC (infected with control lentivirus), CPS1-KD (infected with CPS1-KD lentivirus), CPS1-KD + AMPK agonist (treated with GSK621), CPS1-KD + AMPK agonist + P53 inhibitor (treated with PFTα), CPS1-KD + AMPK inhibitor (treated with GSK690693), and CPS1-KD + AMPK inhibitor + P53 agonist (treated with Kevetrin Hydrochloride), all treatments dissolved in DMSO for 24 h.
Western blotting
Proteins CPS1, NOX2, NOX4, p-AMPK, p-P53, LKB1, Caspase-8 and Caspase-12 were analyzed using Western blot. The total protein of the corresponding cells was extracted using RIPA lysis buffer, and the protein concentration was determined using the BCA kit. The prepared protein samples were mixed with 4× loading buffer, heated in a boiling water bath for 5 min, and then subjected to electrophoresis separation after boiling. The separated proteins were transferred to a PVDF membrane, and then incubated at room temperature for 1 h for blocking. A primary antibody was added and incubated at 4 ℃ overnight. The membrane was washed 3 times with TBST for 10 min each time, and a corresponding secondary antibody was diluted according to the species source of the primary antibody at a ratio of 1 : 5000. The membrane was incubated at room temperature for 1.5 h, washed with TBST 3 times for 10 min each time, and then exposed and developed under the gel imaging instrument. The protein gray value was analyzed using Image J software, and the relative expression level of each protein was calculated based on the gray value of the target protein/gray value of the internal reference protein.
Flow cytometry
Cells were stained with Annexin V-FITC and PI post-infection and analyzed using flow cytometry to assess apoptosis. Before the experiment, cells were inoculated into 6-well plates according to the experimental requirements and the cell density was adjusted using a cell counter. The cells were intervened according to the experimental design. For adherent cells, trypsin without EDTA should be used for digestion before the experiment, and the trypsin digestion solution should be discarded promptly. Be careful to avoid false positives caused by prolonged digestion time. After trypsin digestion, the cells were washed twice with PBS solution, centrifuged at 800 g for 5 min in a low-temperature high-speed centrifuge, and the PBS solution was discarded. Then, 1–5 × 105 cells were collected. The collected cells were transferred to a flow cytometry tube, 500 µL of BindingBuffer was added to each flow cytometry tube, and the cells were gently mixed with a pipette to form a single-cell suspension. The suspension was gently vortexed with a vortex mixer to mix evenly. According to the experimental needs, 5 µL of AnnexinV-FITC or PI staining solution was added and mixed gently with the vortex mixer. The reaction was carried out under light protection for 5–10 min at room temperature. After the light protection ended, the flow cytometry tube should be vortexed again with the vortex mixer before loading onto the instrument. The double-staining reagents were detected using a flow cytometer. When detecting with the flow cytometer, the green fluorescence of AnnexinV-FITC selected the emission wavelength of 530 nm, and the red fluorescence of PI selected the emission wavelength of 630 nm.
Cell proliferation assay
Cells (2 × 103/well) were seeded in 96-well plates, treated with CCK-8 solution, and cultured. Absorbance at 450 nm was measured post-culture to assess proliferation.
Colony formation assay
Cells (400/well) were seeded in 6-well plates and cultured for 2 weeks, followed by PBS washing, 4% paraformaldehyde fixation, and crystal violet staining. Colonies with > 50 cells were counted.
MDA and SOD measurement
Detect the content of oxidative stress markers in the nude mice. Use the liquid sample SOD detection kit (WST8 method) and the liquid sample MDA detection kit (TBA method) to measure the levels of SOD and MDA in the serum of nude mice. Use the tissue SOD detection kit (WST-8 method) and the tissue sample MDA detection kit (TBA method) to measure the levels of SOD and MDA in the tissues of nude mice. All operations were carried out in accordance with the instructions provided by the reagent suppliers.MDA and SOD levels in A549 cells were assessed using commercial assay kits according to the manufacturer’s instructions, measuring absorbance at 450 nm.
Immunofluorescence
Cell coverslips in 24-well plates were seeded with 1 × 106 A549 cells. After fixation with 4% paraformaldehyde, blocking with 2% BSA, and peroxide treatment, coverslips were incubated with AMPK, Caspase-3 and P53 primary antibodies. Subsequent fluorescent secondary antibody staining was performed, with DAPI counterstaining, and observations made using a confocal laser microscope.
Xenograft tumor formation in nude mice
A549 cells in logarithmic growth phase were taken as 1 ml of A549 cell suspension (cell density of 1 × 105/mL) and inoculated in 6-well plates, and the A549 cells were divided into: the NC group (group a), with no additional treatment; the CPS1-KD group (group b), which was infected with the CPS1-KD lentivirus to the A549 cells; the CSP1-KD + AMPK-OE group (group c), which was infected with AMPK-OE lentivirus based on the same treatment as the CPS1- KD group (group b) was infected with AMPK-OE lentivirus on the basis of the same treatment; CSP1-KD + AMPK-OE + P53-KD group (group d) was additionally infected with P53-KD lentivirus on the basis of the same treatment in the CSP1-KD + AMPK-OE group (group c); and CSP1-KD + AMPK-KD group (group e) was infected with P53-KD lentivirus on the basis of the same treatment in the CPS1-KD group (group b) was infected with AMPK-KD lentivirus on the basis of the same treatment; CSP1-KD + AMPK-KD + P53-OE group (group f), was infected with P53-OE lentivirus on the basis of the same treatment in CSP1-KD + AMPK-KD group (group e).
A549 cells were injected subcutaneously into 5-week-old BALB/c nude mice in groups as described above (4 × 106 in 125 µL PBS), 6 mice per group.The nude mice were purchased from Henan SCBS Biotechnology Co., Ltd. They were male, weighing 18 ± 1 g, and a total of 36 mice were used. The nude mice were raised in a clean environment with a 12/12 hour day-night cycle and were provided with sufficient food and water.All mice were executed 4 weeks after inoculation with cancer cells in a 1% O2 environment with low sugar feeding. CO2 gas was administered at an equilibrium rate of 30–70% of cage volume per minute before the nude mice were placed in an ungassed euthanasia cage box. Each nude mouse was continuously monitored for changes in respiratory rate and eye colour. When all nude mice were determined not to be breathing and had lost eye colour, CO2 was continued for at least 1 min before the nude mice were removed. After nude mice were confirmed dead, tumours were collected and stored for subsequent volume and weight analysis.This study adhered to the 3R principles and the arrive guidelines.
Statistical analysis
SPSS 25.0 software was used and count data were expressed asS\(\overline {{\text{x}}} \pm {\text{s}}\); t-test was used for comparison between two groups.The LSD method was used when the variance was uniform. When the variance was not uniform, the Games-Howell method was used. p < 0.05 was regarded as statistically significant.
Results
Bioinformatics analysis.
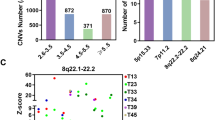
The lung adenocarcinoma dataset was examined for clustering patterns among sample groups through a PCA graph (Fig. 1A). Through in-depth analysis using the limma package, a total of 549 significantly differentially expressed genes were identified, including 185 up-regulated genes and 364 down-regulated genes. Among them, CPS1 was a significantly up-regulated gene (Fig. 1B). The expression changes of these genes provided important clues for subsequent biological research. Subsequently, we conducted GO (Gene Ontology) and KEGG (Kyoto Encyclopedia of Genes and Genomes) analyses using the ClusterProfiler tool. The results showed that these differentially expressed genes were significantly enriched in multiple biological pathways, indicating that they may play a key role in biological functions. The GO analysis (Fig. 1C) revealed that DEGs were significantly enriched in collagen-containing extracellular matrix, cell-cell junction, endoplasmic reticulum lumen, and apical part of cells; the DEGs were significantly enriched in extracellular matrix structural constituent, glycosaminoglycan binding, enzyme inhibitor activity, and growth factor binding in molecular functions; the DEGs were significantly enriched in response to reactive oxygen species, response to oxidative stress, negative regulation of phosphate metabolic process, and inositol phosphate-mediated signaling in biological processes, suggesting that they may play important roles in the response of tumor cells to oxidative stress. The KEGG pathway analysis (Fig. 1D/E) revealed that DEGs were significantly enriched in Complement and coagulation cascades, Protein digestion and absorption, P53 signaling pathway, ECM-receptor interaction, and Cell adhesion molecules, further emphasizing their multiple roles in tumor biology. Additionally, through Spearman correlation analysis, we found a positive correlation between reactive oxygen species (ROS) and AMPK (PRKAA1) (Fig. 1F), indicating that ROS may affect cell metabolism through the AMPK pathway. Similarly, a positive correlation was also observed between ROS and P53 (TP53) (Fig. 1G), suggesting the potential role of ROS in cellular stress and apoptosis. The positive correlation between AMPK (PRKAA1) and P53 (TP53) (Fig. 1H) further emphasized the interaction between these two in the cellular response mechanism. Finally, the positive correlation between LKB1 (STK11) and AMPK (PRKAA1) (Fig. 1I) may reveal the important role of LKB1 in regulating AMPK activity. What is particularly noteworthy is that CPS1 has a significant negative correlation with the core regulatory network of oxidative stress: it shows a significant negative correlation with reactive oxygen species (ROS, Fig. 1J), the DNA damage response hub P53 (TP53, Fig. 1K), the energy metabolism sensor AMPK (PRKAA1, Fig. 1L), and its upstream kinase LKB1 (STK11, Fig. 1M). These findings provide a new perspective for us to understand the response mechanism of cells under stress conditions.
Bioinformatics analysis of differentially expressed genes in lung adenocarcinoma (A): Sample difference PCA plot; (B): Sample difference volcano plot; (C): GO enrichment analysis histogram; (D): KEGG enrichment analysis circle plot; (E): KEGG enrichment analysis chord plot; (F): ROS and AMPK (PRKAA1) correlation scatterplot; (G): ROS and P53 (TP53) correlation scatterplot; H: AMPK (PRKAA1) and P53 (TP53) correlation scatterplot; (I): LKB1 (STK11) and AMPK (PRKAA1) correlation scatterplot.
Knockdown of CPS1 promotes oxidative stress in lung adenocarcinoma cells, thereby facilitating the expression of p-AMPK, P53 and LKB1
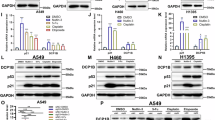
Tumour cells are strongly resistant to oxidative stress, in order to investigate the effect of CPS1 on oxidative stress in tumour cells, Western blot and immunofluorescence staining was performed on six groups of cells, and the expression of oxidative stress-related proteins NOX2 and NOX4 was significantly higher in the CPS1-KD group with knocked-down CPS1 compared with the NC group, and the expression of p-AMPK, p-P53 and LKB1 was also significantly higher. p-AMPK, p-P53, and LKB1 were also significantly increased. When AMPK agonist stimulation was given on the basis of CPS1 knockdown, the expressions of NOX2, NOX4, p-P53 and LKB1 were significantly increased; when P53 inhibitor stimulation was given again, the expressions of NOX2, NOX4, p-P53 and LKB1 were significantly decreased, but there was no significant change in the expressions of NOX2 and NOX4 when compared with that of the CPS1-KD group.CPS1 knockdown promotes oxidative stress, ROS promotes p-P53, and LKB1. oxidative stress, and ROS promoted the expression of p-AMPK, p-P53 and LKB1. To further investigate the relationship among ROS, p-AMPK, p-P53 and LKB1, AMPK inhibitor stimulation was given on the basis of CPS1 knockdown, and the expression of NOX2, NOX4, p-P53 and LKB1 was significantly reduced compared to that of the CPS1-KD group; and then P53 agonist stimulation was given, and the expression of NOX2, NOX4, p-P53 and LKB1 was significantly elevated, but there was no significant change in NOX2 and NOX4 expression compared with the CPS1-KD group. Tumor cells have a strong tolerance to oxidative stress. To study the effect of CPS1 on oxidative stress in tumor cells, we conducted experiments using Western blot, ELISA, and immunofluorescence staining. The Western blot results showed that compared with the NC group, the expression of CPS1 in the CPS1-KD group was significantly decreased; while there was no significant difference in the expression of CPS1 in the CPS1-KD group, the CPS1-KD + GSK621 group, the CPS1-KD + GSK621 + PFIα group, the CPS1-KD + GSK690693 group, and the CPS1-KD + GSK690693 + Kevetrin Hydrochloride group. Compared with the NC group, the expression of NOX2, NOX4, p-AMPK, p-P53, and LKB1 in the CPS1-KD group increased; compared with the CPS1-KD group, the expression of NOX2, NOX4, p-AMPK, p-P53, and LKB1 in the CPS1-KD + GSK621 group increased significantly; compared with the CPS1-KD + GSK621 group, the expression of NOX2 and NOX4 in the CPS1-KD + GSK621 + PFIα group was significantly lower, and there was no significant difference in p-P53 and LKB1 expression, while p-AMPK was not significantly different; compared with the CPS1-KD group, the expression of NOX2, NOX4, p-AMPK, p-P53, and LKB1 in the CPS1-KD + GSK690693 group was lower; compared with the CPS1-KD + GSK690693 group, the expression of NOX2, NOX4, p-P53, and LKB1 in the CPS1-KD + GSK690693 + Kevetrin Hydrochloride group increased, and there was no significant difference in p-AMPK; and compared with the CPS1-KD group, the expression of NOX2, NOX4, p-P53, and LKB1 in the CPS1-KD + GSK690693 group was lower; compared with the CPS1-KD + GSK690693 group, the expression of NOX2, NOX4, p-P53, and LKB1 in the CPS1-KD + GSK690693 + Kevetrin Hydrochloride group increased, and there was no significant difference in NOX2 and NOX4 compared with the CPS1-KD group. (Fig. 2A). The ELISA results showed that compared with the NC group, the MDA and SOD in the CPS1-KD group increased significantly; compared with the CPS1-KD group, the MDA and SOD in the CPS1-KD + GSK621 group increased; compared with the CPS1-KD + GSK621 group, the MDA and SOD in the CPS1-KD + GSK621 + PFIα group decreased, and there was no significant difference compared with the CPS1-KD group; compared with the CPS1-KD group, the MDA and SOD in the CPS1-KD + GSK690693 group decreased; compared with the CPS1-KD + GSK690693 group, the MDA and SOD in the CPS1-KD + GSK690693 + Kevetrin Hydrochloride group increased, and there was no significant difference compared with the CPS1-KD group (Fig. 2B). The immunofluorescence staining results showed that compared with the NC group, the relative fluorescence intensity of p-AMPK and p-P53 in the CPS1-KD group was significantly enhanced; compared with the CPS1-KD group, the relative fluorescence intensity of p-AMPK and p-P53 in the CPS1-KD + GSK621 group was enhanced; compared with the CPS1-KD + GSK621 group, the relative fluorescence intensity of p-AMPK and p-P53 in the CPS1-KD + GSK621 + PFIα group was weakened, and there was no significant difference compared with the CPS1-KD group; Compared with the CPS1-KD group, the relative fluorescence intensities of p-AMPK and p-P53 in the CPS1-KD + GSK690693 group were weakened; compared with the CPS1-KD + GSK690693 group, the relative fluorescence intensities of p-AMPK and p-P53 in the CPS1-KD + GSK690693 + Kevetrin Hydrochloride group were enhanced, and there was no significant difference compared with the CPS1-KD group (Fig. 2C). These results indicate that CPS1 knockdown promotes oxidative stress in lung adenocarcinoma cells, thereby promoting the expression of p-AMPK, P53 and LKB1.
CPS1 knockdown induces oxidative stress through ammonia accumulation and activates the AMPK/P53/LKB1 pathway. (A): Western blot was used to detect the expressions of CPS1, NOX2, NOX4, p-AMPK, p-P53, and LKB1 in the NC group, the CPS1-KD group, the CPS1-KD + GSK621 group, the CPS1-KD + GSK621 + PFIα group, the CPS1-KD + GSK690693 group, and the CPS1-KD + GSK690693 + Kevetrin Hydrochloride group; (B): ELISA detected the intracellular levels of MDA and SOD; (C): Results from cell climbing assay are shown, with green fluorescence representing p-P53 and red fluorescence representing p-AMPK. Data are expressed as mean ± standard deviation. N = 3, P < 0.05 is considered statistically significant. nsP > 0.05; *P < 0.05; **P < 0.01.
Knocking down CPS1 promotes the expression of P53 and LKB1, thereby inhibiting the proliferation of lung adenocarcinoma cells and promoting their apoptosis
To study the effects of P53 and LKB1 on the development of lung adenocarcinoma, we conducted experiments using western blot, CCK-8, immunofluorescence staining, colony formation, and flow cytometry. The results of Western Blot showed that compared with the NC group, the expression of Bax, Caspase-8, and Caspase-12 in the CPS1-KD group was significantly increased. Compared with the CPS1-KD group, the expression of Bax, Caspase-8, and Caspase-12 in the CPS1-KD + GSK621 group was even higher. Compared with the CPS1-KD group and the CPS1-KD + GSK621 group, the expression of Bax, Caspase-8, and Caspase-12 in the CPS1-KD + GSK621 + PFIα group was significantly decreased. Compared with the CPS1-KD group, the expression of Bax, Caspase-8, and Caspase-12 in the CPS1-KD + GSK690693 group was lower. Compared with the CPS1-KD + GSK690693 group, the expression of Bax, Caspase-8, and Caspase-12 in the CPS1-KD + GSK690693 + Kevetrin Hydrochloride group was higher (Fig. 3A). The results of the CCK-8 kit assay showed that at 72 h, compared with the NC group, the OD value of the CPS1-KD group decreased; compared with the CPS1-KD group, the OD value of the CPS1-KD + GSK621 group was significantly decreased; compared with the CPS1-KD group and the CPS1-KD + GSK621 group, the OD value of the CPS1-KD + GSK621 + PFIα group was significantly increased; compared with the CPS1-KD group, the OD value of the CPS1-KD + GSK690693 group increased; compared with the CPS1-KD + GSK690693 group, the OD value of the CPS1-KD + GSK690693 + Kevetrin Hydrochloride group decreased (Fig. 3B). The results of the immunofluorescence staining experiment showed that compared with the NC group, the relative fluorescence intensity of Caspase-3 in the CPS1-KD group was significantly enhanced; compared with the CPS1-KD group, the relative fluorescence intensity of Caspase-3 in the CPS1-KD + GSK621 group was significantly enhanced; compared with the CPS1-KD group and the CPS1-KD + GSK621 group, the relative fluorescence intensity of Caspase-3 in the CPS1-KD + GSK621 + PFIα group was significantly decreased; compared with the CPS1-KD group, the relative fluorescence intensity of Caspase-3 in the CPS1-KD + GSK690693 group was significantly decreased; compared with the CPS1-KD + GSK690693 group, the relative fluorescence intensity of Caspase-3 in the CPS1-KD + GSK690693 + Kevetrin Hydrochloride group was significantly enhanced, with no significant difference compared to the CPS1-KD group (Fig. 3C). The results of the colony formation experiment showed that compared with the NC group, the number of colonies in the CPS1-KD group decreased; compared with the CPS1-KD group, the number of colonies in the CPS1-KD + GSK621 group decreased; compared with the CPS1-KD group and the CPS1-KD + GSK621 group, the number of colonies in the CPS1-KD + GSK621 + PFIα group significantly increased; Compared with the CPS1-KD group, the number of clones in the CPS1-KD + GSK690693 group increased; compared with the CPS1-KD + GSK690693 group, the number of clones in the CPS1-KD + GSK690693 + Kevetrin Hydrochloride group decreased (Fig. 3D). The results of flow cytometry showed that compared with the NC group, the apoptosis rate of cells in the CPS1-KD group was significantly increased; compared with the CPS1-KD group, the apoptosis rate of cells in the CPS1-KD + GSK621 group was significantly increased; compared with the CPS1-KD group and the CPS1-KD + GSK621 group, the apoptosis rate of cells in the CPS1-KD + GSK621 + PFIα group was significantly decreased; compared with the CPS1-KD group, the apoptosis rate of cells in the CPS1-KD + GSK690693 group was decreased; compared with the CPS1-KD + GSK690693 group, the apoptosis rate of cells in the CPS1-KD + GSK690693 + Kevetrin Hydrochloride group was significantly increased (Fig. 3E). These results indicate that CPS1 knockdown promotes the expression of P53 and LKB1, thereby inhibiting tumor cell proliferation and promoting their apoptosis.
CPS1 knockdown promotes tumor cell apoptosis and inhibits proliferation by activating P53/LKB1. (A): Western blot was used to detect the expression of Bax, Caspase-8 and Caspase-12 in the NC group, the CPS1-KD group, the CPS1-KD + GSK621 group, the CPS1-KD + GSK621 + PFIα group, the CPS1-KD + GSK690693 group and the CPS1-KD + GSK690693 + Kevetrin Hydrochloride group; (B): Cell proliferation results detected by CCK8 assay; (C): Immunofluorescence staining experiment was used to detect the expression of caspase-3.; (D): Colony formation assay to detect cell proliferation; (E): Flow cytometry detected cell apoptosis. Data are expressed as mean ± standard deviation. N = 3, P < 0.05 is considered statistically significant. nsP > 0.05; *P < 0.05; **P < 0.01.
CPS1 knockout promotes the expression of AMPK and P53, thereby inhibiting tumor formation in nude mice
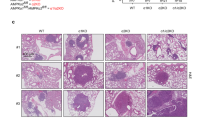
In order to observe more intuitively the effects of CPS1, AMPK and P53 on the development of lung adenocarcinoma, lung adenocarcinoma cells were inoculated on the back of nude mice, and the size of the transplanted tumours was observed. Compared with the NC group, the weight and volume of the transplanted tumors in the CPS1-KD group were significantly reduced; compared with the CPS1-KD group, the weight and volume of the transplanted tumors in the CPS1-KD + AMPK-OE group were significantly reduced; compared with the CPS1-KD + AMPK-OE group, the weight and volume of the transplanted tumors in the CPS1-KD + AMPK-OE + P53-KD group were significantly increased, and there was no significant difference between the CPS1-KD + AMPK-KD group and the CPS1-KD group; compared with the CPS1-KD + AMPK-KD group, the weight and volume of the transplanted tumors in the CPS1-KD + AMPK-KD + P53-OE group were significantly reduced, and there was no significant difference between the CPS1-KD + AMPK-KD + P53-OE group and the CPS1-KD group(Fig. 4A, B). It has been shown to initiate the process of apoptosis through the activation of P53. In summary, CPS1 knockdown initiates a series of intricate signalling cascades by affecting ammonia metabolism, which ultimately results in apoptosis and the growth inhibition of tumour cells by regulating reactive oxygen species and the expression of P53, AMPK, LKB1 and JNK. We propose that CPS1 promotes lung adenocarcinoma progression by inhibiting ammonia-induced activation of the ROS/AMPK/P53/LKB1 signalling pathway (Fig. 5). The original Western blot membranes for this study are provided in Supplementary File 1, and the raw data are provided in Supplementary File 2.
CPS1 knockdown inhibits the growth of transplanted tumors in nude mice through the AMPK/P53 pathway (A): The experimental tumor results of subcutaneous tumor transplantation in nude mice for the NC group, CPS1-KD group, CPS1-KD + AMPK-OE group, CPS1-KD + AMPK-OE + P53-KD group, CPS1-KD + AMPK-KD group and CPS1-KD + AMPK-KD + P53-OE group; (B): Statistical graphs of tumor volume and tumor weight for each group.N = 6, P < 0.05 is considered statistically significant. nsP > 0.05; *P < 0.05; **P < 0.01.
Schematic diagram illustrating the mechanism by which CPS1 promotes the progression of lung adenocarcinoma by inhibiting the ammonia-induced ROS/AMPK/P53/LKB1 signaling pathway. After CPS1 knockdown, the intracellular ammonia metabolism becomes unbalanced, resulting in a significant increase in ammonia levels. The accumulated ammonia induces oxidative stress and generates excessive reactive oxygen species (ROS). Excessive ROS are activated through oxidative modification and phosphorylate AMPK, increasing its phosphorylation level. Phosphorylated AMPK further activates P53 and promotes its formation with LKB1 into a complex, enhancing the activity of LKB1. The activated P53 and LKB1 complex upregulates the expression of pro-apoptotic proteins (such as Bax, Caspase-8, and Caspase-12), thereby inhibiting the proliferation of tumor cells and promoting their apoptosis.
Discussion
Lung cancer is a leading global malignancy, with over 2 million new cases annually. Adenocarcinoma constitutes ~ 45% of cases and contributes to ~ 18% of cancer deaths, posing a major public health burden17. Early symptoms of lung adenocarcinoma are insidious, and most of the patients are diagnosed at advanced stages with poor prognosis18,19. Given the differences in the molecular characteristics of different lung adenocarcinoma patients20, the identification of specific biomarkers is essential for personalised treatment and early diagnosis and treatment, which can help to improve the cure rate of patients.
AMPK is an energy sensor in the cell. The activation of AMPK is primarily mediated by phosphorylation of key proteins in response to metabolic stress, thereby biasing metabolism towards increased catabolism and decreased anabolism. It has been demonstrated that activation of AMPK is essential for alleviating the metabolic and energetic stresses associated with tumour progression. Furthermore, activation of AMPK has been observed to induce a number of cell death mechanisms. These include effects on cell cycle checkpoints, autophagy, mitochondrial autophagy, and apoptosis. AMPK has been demonstrated to promote autophagy and mitochondrial autophagy through the activation of UNC-51-like kinase 1 (ULK1) and death-associated protein 1 (DAP1), respectively9,10.
The P53 gene, which functions as a tumour suppressor, has been the subject of extensive investigation in numerous studies. It has been demonstrated that mutations in this gene are present in more than 50% of cancer cases21. Furthermore, P53 plays a pivotal role in the cellular response to DNA damage. In response to cellular stress or DNA damage, P53 becomes stable and initiates gene transcription, thereby influencing the cell’s future fate. Given that P53 is a redox protein containing cysteine clusters, it is susceptible to attack by ROS22. In this study, Phosphorylation of AMPK activates P53 phosphorylation, phosphorylated P53 binds to LKB1 in the nucleus to form a complex that promotes LKB1 transcription, Bax and caspase family protein expression is promoted, apoptosis is facilitated, and tumour cell proliferation is inhibited. In this study, the AMPK agonist further increased the levels of p-P53 and apoptotic proteins, while the AMPK inhibitor reversed this effect, confirming that AMPK is an important upstream regulatory factor for P53.Liver kinase B1 (LKB1) is a member of the serine/threonine protein kinase family, which plays a pivotal role in oncogenesis by activating the downstream kinase AMPK23. LKB1 enhances the stability of P53 and the nuclear binding of phosphorylated P53, and jointly induces apoptosis by facilitating the activation of JNK. The bioinformatics analysis in this study revealed that LKB1 is positively correlated with AMPK and P53 expression, suggesting that LKB1 may exert tumor suppression by coordinating the activity of AMPK and P53-dependent apoptosis.CPS1 knockdown has been verified to increase intracellular ammonia levels and decrease urea levels in lung adenocarcinoma cells, and the elevated toxic ammonia allows for increased ROS levels and promotes the expression of AMPK and P53. AMPK promotes apoptosis in lung adenocarcinoma cells by activating P53 to form a complex with KLB1, while AMPK negatively regulates ROS levels, and KLB1 also promotes AMPK and regulates P53 by promoting JNK expression.
Under normal conditions, CPS1 is able to convert NH4 + to urea through the urea cycle, but when CPS1 is knocked down, the urea cycle is inhibited, and the intracellular ammonia content is increased and the urea content is decreased.Ammonia induces excessive production of ROS through multiple pathways. This study shows that knockdown of CPS1 significantly upregulates the expression of NOX2 and NOX4, which are the main sources of ROS. At the same time, the accumulation of ammonia may deplete antioxidant molecules such as glutathione (GSH), leading to a decrease in SOD activity (as shown in the experimental results, MDA increased and SOD decreased), thereby exacerbating oxidative stress.ROS accumulation activated multiple pathways and had different effects on cancer cell survival 8, ROS promote the expression of AMPK and P53. ROS may induce conformational changes by oxidizing the Cys130 and Cys174 residues of the AMPKα subunit, thereby promoting its phosphorylation (such as at the Thr172 site). Additionally, ATP depletion induced by ammonia may indirectly activate AMPK by increasing the AMP/ATP ratio. And the activation of AMPK can, in turn, inhibit the accumulation of ROS, forming a regulatory loop.
Recent systems biology studies have revealed a positive correlation between P53 gene expression and AMPK expression and oxidative stress. Additionally, KEGG pathway analysis indicated that differentially expressed genes (DEGs) in lung adenocarcinoma were significantly enriched in the P53 signaling pathway.This study further validated the function of the P53 gene through experimental verification, confirming that P53 is a core executor of metabolic stress: P53 serves as a key downstream effector of the ROS/AMPK pathway, mediating tumor cell apoptosis, while CPS1 can indirectly inhibit P53 activation by suppressing the ammonia-ROS-AMPK axis24. In addition, studies have shown that lung cancer stem cells (CSCs) exhibit embryonic stem cell-like characteristics and EMT phenotypes, and maintain CSC stemness through activation of the PI3K/AKT/NF-κB pathway.Although this study did not directly detect CSCs, there is a potential mechanism linking these findings, and the metabolic perspective expands this framework: CPS1, as a metabolic hub gene, promotes tumor survival by inhibiting the ammonia-induced ROS/AMPK/P53/LKB1 pathway, while the ROS burst caused by CPS1 inhibition simultaneously weakens tumor stem cell self-renewal25. This study not only investigates the anti-tumor functions of P53 and AMPK but also reveals the core triggering role of the metabolite ammonia: ammonia accumulation-induced ROS bursts may weaken CSC self-renewal by activating DNA damage responses.targeting CPS1 can synergistically eliminate CSCs. Additionally, incorporating CPS1 into the molecular subtyping system of lung adenocarcinoma (especially the metabolic subtype) may optimize combined treatment strategies targeting EMT and CSCs. In the future, we will validate the effects of CPS1 on stemness markers and EMT in CSC models to further elucidate the role of metabolic regulation in CSCs.
This indicates that CPS1 indirectly inhibits the production of ROS by maintaining ammonia homeostasis, thereby weakening the activation of the AMPK/P53/LKB1 pathway, providing favorable conditions for tumor cell survival. Knockdown of CPS1 can induce oxidative stress through ammonia accumulation, activate the AMPK/P53/LKB1 pathway, promote tumor cell apoptosis, and inhibit proliferation. However, this study has not yet clarified the differences in the dependence of different molecular subtypes of lung adenocarcinoma (such as EGFR mutant type) on this pathway, which will be our focus for subsequent research.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries [J]. CA Cancer J. Clin. 74 (3), 229–263 (2024).
Herbst, R. S., Morgensztern, D. & Boshoff, C. The biology and management of non-small cell lung cancer [J]. Nature 553 (7689), 446–454 (2018).
Lee, H. H. et al. Biomarkers of favorable vs. Unfavorable responses in locally advanced rectal cancer patients receiving neoadjuvant concurrent chemoradiotherapy [J]. Cells, 11(10):1611. (2022).
Tang, Z. et al. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis [J]. Nucleic Acids Res., 47(W1): W556–w60. (2019).
Zhang, H. et al. Blockade of AMPK-Mediated cAMP-PKA-CREB/ATF1 signaling synergizes with aspirin to inhibit hepatocellular carcinoma [J]. Cancers (Basel), 13(7):1738. (2021).
Pham-danis, C. et al. Urea cycle sustains cellular energetics upon EGFR Inhibition in EGFR-Mutant NSCLC [J]. Mol. Cancer Res. 17 (6), 1351–1364 (2019).
Itzel, T. et al. Hepamine - A liver disease microarray database, visualization platform and Data-Mining resource [J]. Sci. Rep. 10 (1), 4760 (2020).
Wu, M. et al. Hepatic AMPK signaling dynamic activation in response to REDOX balance are Sentinel biomarkers of exercise and antioxidant intervention to improve blood glucose control [J]. Elife, 11:e79939. (2022).
Naik, P. P. et al. Secretory clusterin promotes oral cancer cell survival via inhibiting apoptosis by activation of autophagy in AMPK/mTOR/ULK1 dependent pathway [J]. Life Sci. 264, 118722 (2021).
Shen, W. et al. Trifluoperazine activates AMPK / mTOR / ULK1 signaling pathway to induce mitophagy in osteosarcoma cells [J]. Chem. Biol. Interact. 392, 110904 (2024).
Khoja, S. et al. A constitutive knockout of murine carbamoyl phosphate synthetase 1 results in death with marked hyperglutaminemia and hyperammonemia [J]. J. Inherit. Metab. Dis. 42 (6), 1044–1053 (2019).
Nitzahn, M. & Lipshutz, G. S. CPS1: looking at an ancient enzyme in a modern light [J]. Mol. Genet. Metab. 131 (3), 289–298 (2020).
El-sheikh, R. M. et al. Carbamoyl phosphate synthetase 1 (CPS1) as a prognostic marker in chronic hepatitis C infection [J]. Apmis 127 (2), 93–105 (2019).
Chandrashekar, D. S. et al. UALCAN: an update to the integrated cancer data analysis platform [J]. Neoplasia 25, 18–27 (2022).
Molina, E., Hong, L. & Chefetz, I. AMPKα-like proteins as LKB1 downstream targets in cell physiology and cancer [J]. J. Mol. Med. (Berl). 99 (5), 651–662 (2021).
Steinberg, G. R. & Hardie, D. G. New insights into activation and function of the AMPK [J]. Nat. Rev. Mol. Cell. Biol. 24 (4), 255–272 (2023).
Zhang, Y. et al. Global variations in lung cancer incidence by histological subtype in 2020: a population-based study [J]. Lancet Oncol. 24 (11), 1206–1218 (2023).
Liu, Z. et al. The screening of immune-related biomarkers for prognosis of lung adenocarcinoma [J]. Bioengineered 12 (1), 1273–1285 (2021).
Jia, Y. et al. Prognosis of early-stage lung adenocarcinoma in young patients [J]. Clin. Exp. Pharmacol. Physiol. 50 (10), 826–832 (2023).
Wei, Q. et al. Molecular subtypes of lung adenocarcinoma patients for prognosis and therapeutic response prediction with machine learning on 13 programmed cell death patterns [J]. J. Cancer Res. Clin. Oncol. 149 (13), 11351–11368 (2023).
Bacher, S. & Schmitz, M. L. Understanding and therapeutic targeting of the P53 network [J]. Cancers (Basel), 15(18):4461. (2023).
Shi, T. & Dansen, T. B. Reactive oxygen species induced P53 activation: DNA damage, redox signaling, or both?? [J]. Antioxid. Redox Signal. 33 (12), 839–859 (2020).
Shimada, T. et al. The distinct roles of LKB1 and AMPK in P53-Dependent apoptosis induced by cisplatin [J]. Int. J. Mol. Sci., 23(17):10064. (2022).
Roudi, R. et al. Identification of hub genes, modules and biological pathways associated with lung adenocarcinoma: A system biology approach [J]. Gene Rep. 27, 101638 (2022).
Roudi, R. et al. Evidence for embryonic stem-like signature and epithelial-mesenchymal transition features in the spheroid cells derived from lung adenocarcinoma [J]. Tumour Biol. 37 (9), 11843–11859 (2016).
Funding
2024 Government Funding for Excellence in Clinical Medicine Project, ZF2024175. Medical Science Research Proiect of Hebei, No.20250138.
Author information
Authors and Affiliations
Contributions
Yanchao Luan, Liru Liu and Chao Liang were responsible for designing the experiments and constructing the general framework of the study. The actual operation of the experiment was undertaken by Yanchao Luan, Liru Liu and Jiakun Liu. Data analysis was undertaken by Zhao Li.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
This study was reviewed by the Ethics Committee of Hebei Provincial Chest Hospital, and was in accordance with the principles of animal protection, animal welfare and ethics, and with the national regulations on experimental animal welfare and ethics,2022051. All methods in this experiment were performed in accordance with the relevant guidelines and regulations and conformed to THE RULES OF 3R and ARRIVE guidelines.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Luan, Y., Liu, L., Liu, J. et al. CPS1-promoted the progression of lung adenocarcinoma via suppressing ammonia induced the activation of ROS/AMPK/P53/LKB1 signaling pathway. Sci Rep 15, 29610 (2025). https://doi.org/10.1038/s41598-025-14443-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-14443-9
Keywords
This article is cited by
-
Prognostic and immunological roles of ammonia-induced cell death-related genes in non-small cell lung cancer
BMC Pulmonary Medicine (2026)