Abstract
Streptomycetes are biotechnologically valuable bacteria with complex cell division that produce extracellular vesicles (EVs), typically nanometre-sized but can reach 2.5 μm in diameter. Streptomyces also produce dividing wall-deficient L-forms (0.5–7 μm diameter) and, under hyperosmotic stress, non-dividing wall-deficient S-cells (3–4 μm diameter). The boundaries between EVs, L-forms and S-cells are not always clear, as large DNA-containing EVs can resemble small L-forms and S-cells in size. Both EVs and wall-deficient cells offer competitive advantages, such as inter-bacterial signalling, antibiotic transport, resistance and phage defence. However, their formation mechanisms remain poorly understood. We identified sco1758 (engA GTPase), sco0954 (methionine N-acetyltransferase), sco4439 (D-Ala-D-Ala carboxypeptidase), and sco4440 (GOLPH3-like) as important for wall-deficient cell formation in Streptomyces coelicolor under hyperosmotic sucrose conditions. Mutations in sco4439 and sco4440 increased tetra–tetra(Gly) and tetra(Gly)–penta(Gly) (4–3) peptidoglycan (PG) dimers, while sco1758 affected only the former. Complementation reversed these changes. sco0954 overexpression enhanced PG-associated methionine acetylation and oxidation. Our findings suggest that PG dimerisation and methionine modification may contribute to the formation of wall-deficient cells under hyperosmotic sucrose stress. Further research is required to elucidate how SCO1758, SCO0954 and SCO4439/40 modulate PG architecture and to evaluate their potential to promote EV production for biotechnological applications.
Similar content being viewed by others
Introduction
Streptomycetes are biotechnologically important bacteria that produce nearly two-thirds of clinically used bioactive secondary metabolites, including antibiotics, antitumour agents, and immunosuppressants1,2. Their complex developmental cycle involves programmed cell death, hyphal differentiation, and sporulation3,4. Cell division is unusually intricate, as vegetative and sporulation septa are differentially regulated: proteins such as FtsI, FtsL, FtsW, SsgA/B, CrgA, DynA/B, and SepH are required for sporulation but not vegetative division5,6,7,8,9,10,11,12, while SepX is specific to vegetative septation13. The tubulin-like GTPase FtsZ functions in both, recruiting the divisome14,15,16. Streptomyces is the only known bacterium in which ftsZ mutations do not abolish viability, although development is altered14.
Streptomyces cell biology is further complicated by the presence of non-replicative extracellular vesicles (EVs)17,18,19,20, which may carry proteins and metabolites, including antibiotics. While some lack DNA21, others may contain an entire chromosome, suggesting a role in genome evolution19. Vesicles have been linked to programmed cell death11,22 and division12. In addition, Streptomyces produce L-forms, wall-deficient cells capable of dividing independently of the FtsZ machinery23,24,25. Recently, non-dividing wall-deficient cells induced by hyperosmotic stress, termed S cells, have been identified in actinomycetes such as Kitasatospora viridifaciens, Streptomyces griseus and Streptomyces venezuelae26. These cells are induced by hyperosmotic stress in media supplemented with 0.64 M sucrose, 0.6 M NaCl or 1 M sorbitol26.
EVs, S-cells, and L-forms may confer adaptive benefits. L-forms of pathogenic bacteria are associated with persistent infections, resistance to cell wall-targeting antibiotics27,28,29, and phage protection30,31. L-forms in K. viridifaciens can take up extracellular DNA via endocytosis-like processes contributing to horizontal gene transfer32. EVs may contribute to defence, for example by transporting β-lactamases in Staphylococcus aureus or acting as phage decoys33. Nonetheless, the biological roles of EVs, S-cells, and L-forms in Streptomyces remain largely unclear.
The size of Streptomyces L-forms (0.5–7 μm in diameter)24,25 and actinomycete S-cells (3–4 μm in diameter)26 is significantly larger than that reported for most Streptomyces EVs (0.08–0.4 μm in diameter)17,18,19,20, but more comparable to EVs observed in S. venezuelae (2.5 μm in diameter)21. Their distinct size, support their classification as separate entities, although the boundaries between them are not always well defined, as the largest DNA-containing EVs can be similar in size with the smallest L-forms and S-cells. According to Welsh et al.34, EVs are lipid bilayer-bound particles released from cells. As L-forms divide, they fall outside this definition, while S-cells can be considered large EVs formed under hyperosmotic stress26.
EV formation in Gram-negative bacteria is relatively well understood, involving the outer membrane and up to 150 genes18. In contrast, Gram-positive EV biogenesis remains elusive. Heat-inactivated gram-positive bacteria do not produce EVs, indicating a dependence on metabolic activity35, and some regulatory genes have been implicated in modulating EV formation in these organisms. The sigma factor sigB and covRS influence EV synthesis in Listeria monocytogenes and Streptococcus pyogenes, respectively35. In Streptomyces venezuelae and Streptomyces lividans, EVs form at hyphal tips, likely due to mechanical stress from weakened apical walls20,21. In S. coelicolor, a contractile tail-like system (sco4256–4258) regulates vesicle formation during cell death22.
L-form division has been reported to depend on membrane fluidity and excess intracellular membranes36, as shown in E. coli37, L. monocytogenes38, and B. subtilis39, enabling passive, biophysical division40. Similar behaviour is seen in synthetic giant lipid vesicles41. Despite their passive nature, regulatory mechanisms affecting cell wall synthesis (e.g., peptidoglycan release) and intracellular membrane production are likely involved. Indeed, mutations affecting L-form formation have been found, as the murE gene repression in B. subtilis23 and mraY and ftsQ mutations in E. coli37.
S-cells are, by definition, induced by hyperosmotic stress in some actinomycetes26. While L-forms are not restricted to such conditions, they can also be triggered by hyperosmotic stress in actinomycetes26. The molecular regulation of S-cell and L-form formation remains unclear, but evidence for genetic control is growing. For example, filP mutation in K. viridifaciens prevents S-cell formation42. The stomatin-like protein StlP, involved in polar growth, was recently linked to the emergence of wall-deficient cells under hyperosmotic stress in S. coelicolor43, a strain previously not known to form S-cells or L-forms24,25,26, but which can produce EVs17,18,44.
In this study, we further explore the genetic modulation of S-cell and L-form formation in Streptomyces coelicolor. We identified sco1758 (EngA GTPase), sco0954 (methionine N-acetyltransferase), sco4439 (D-Ala-D-Ala carboxypeptidase), and sco4440 (GOLPH3-like), whose mutations alter peptidoglycan (PG) muropeptide composition and affect the redox and acetylation states of PG-bound methionine. These changes correlate with the emergence of wall-deficient cells under hyperosmotic sucrose stress (S-cells in sco0954 and sco4439/40, and both S-cells and L-forms in sco1758).
Results
The sco1760::Tn5, and sco4439/40::Tn5062 knockout mutants, as well as S. coelicolor overexpressing sco0954, exhibit hyphal bulging and the formation of large distinct round cells under conditions of hyperosmotic sucrose stress
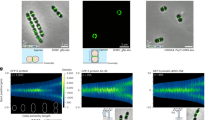
The sco1760::Tn5 mutant was generated in this study using the transposon-based random mutagenesis method of Xu et al.45. The phenotypes of 1,000 random mutants were analysed macroscopically (colony morphology, and production of undecylprodigiosin [red] and actinorhodin [blue]), and microscopically, using vital, membrane and cell-wall stains). The SCO1760::Tn5 mutant exhibited pronounced hyphal bulging and the formation of large extracellular round cells that contain DNA (arrows in Fig. 1b), most of which were viable (SYTO9 green staining; see methods).
Formation of wall-deficient cells and extracellular vesicles (EVs) under sucrose-induced hyperosmotic stress in the sco1760::Tn5, sco4439/40::Tn5062, and sco0954 knockout mutants, as well as in S. coelicolor overexpressing sco0954, grown on GYM medium supplemented with 0.64 M sucrose. Images correspond to confocal laser-scanning fluorescence microscopy of hyphae stained with SYTO9 and PI (DNA stains). Histograms represent EVs sizes (areas): grey bars correspond to the S. coelicolor wild-type strain (negative control, not producing EVs), used to establish the threshold area for EVs identification; blue bars represent EVs with an area above the 3.6 μm² threshold in mutants grown on GYM medium supplemented with 0.64 M sucrose. (a–d) Cultures on GYM medium supplemented with 0.64 M sucrose. (e–h) Cultures on GYM medium supplemented with 0.64 M sucrose. Representative images from at least three biological replicates are shown. Arrows indicate EVs and sucrose-driven hyperosmotic-stress-induced cells. Scale bars: 8 μm.
The sco4439/40::Tn5062 mutant, constructed previously46, lacks SCO4439, a D-Ala-D-Ala carboxypeptidase; this increases peptidoglycan cross-linking and causes notable spore swelling during germination46. This mutant also exhibited hyphal bulging and the formation of large cells in medium amended with 0.64 M sucrose (Fig. 1c).
sco0954 was identified in this study via RNA-seq comparing an ftsZ mutant14 with the S. coelicolor wild type. Given its marked upregulation in the mutant (Supplementary Table S1), we hypothesised that it may be involved in the FtsZ-independent division of L-forms. We overexpressed sco0954 in the wild-type strain to test this, observing a phenotype similar to the sco1760::Tn5 and sco4439/40::Tn5062 mutants, S-cell formation in hyperosmotic sucrose conditions (Fig. 1d).
No differences in viability were observed among the mutants, however, some of the large cells formed as well as the bulging hyphae, were dead (as indicated by red PI staining in Fig. 1; see additional images in Supplementary Fig. S1), perhaps reflecting an instability of these structures. Hyphal bulging and distinct large cells were not observed on 0.3 M sucrose (Fig. 1, right panels), a concentration typically used in laboratory media to protect wall-deficient cells from lysis (see examples of media designed to preserve protoplast integrity in Kieser et al.47.
Overexpression of sco0954 and mutations in sco1760 and sco4439/40 induce wall-deficient S-cells under hyperosmotic sucrose stress in Streptomyces coelicolor.
Overexpression of sco0954 and mutations in sco1760 and sco4439/40 induce wall-deficient S-cells under hyperosmotic sucrose stress in Streptomyces coelicolor.
We next examined whether the cells induced under hyperosmotic sucrose conditions observed here, like S- and L-cells, lack a cell wall. To do this, we used Alexa Fluor 488-WGA (green, cell wall) and FM5-95 (red, membrane) staining (Fig. 2). No large distinct round cells were detected in the S. coelicolor wild type (Fig. 2a). In contrast, all mutants grown on GYM medium with 0.64 M sucrose produced large distinct round cells with membranes (red) without detectable walls (green) (arrowheads in Fig. 2). Consequently, these cells fit the definition of EVs34. More precisely they are S-cells, i.e. wall-deficient cells induced under hyperosmotic stress (0.64 M sucrose) that are unable to divide. We also observed cells with thin walls (asterisks), thick walls (arrows), or localised wall patches (squares), which may represent remnants of original walls or sites of peptidoglycan regeneration. Some vesicles displayed excess internal FM5-95 staining.
Cells formed under sucrose-induced hyperosmotic stress in the sco1760::Tn5 and sco4439/40::Tn5062 mutants, as well as in the wild-type strain overexpressing sco0954, lack a cell wall. (a–g) Confocal laser-scanning microscopy images of cultures grown on GYM medium supplemented with 0.64 M sucrose and stained with FM5-95 (red membrane stain) and Alexa Fluor 488–WGA (green cell wall stain). (h) Membrane-to-cell-wall abundance quantified as the ratio between pixels stained with WGA and FM5-95. Asterisks indicate statistically significant differences compared to the wild-type strain: p < 0.05, *p < 0.01, **p < 0.001. Arrowheads indicate cells without detectable cell walls; asterisks denote cells with very thin cell walls; arrows indicate cells with thick peptidoglycan (PG) walls; and squares indicate cells with cell wall patches. Representative images from at least three biological replicates are shown. Scale bars represent 8 μm.
We quantified the membrane-to-wall ratio from confocal images using Fiji48, calculating the FM5-95 to WGA pixel ratio across at least three biological replicates (see Supplementary Fig. S1 for representative images). In wild-type cultures grown on 0.64 M sucrose, this ratio was slightly below 1, reflecting more peptidoglycan staining, as expected, as the cell wall is thicker than the membrane (Fig. 2a). In contrast, the sco0954-overexpressing strain and the sco4439/40::Tn5062 mutant showed significantly higher ratios (> 1), indicating membrane enrichment (Fig. 2h). The sco1760::Tn5 mutant did not differ significantly from wild type, likely due to its lower abundance of stress-induced vesicles (see next paragraph, Fig. 3b), with most stained pixels corresponding to hyphae with thick walls rather than wall-deficient cells.
Area and abundance of extracellular vesicles (EVs) formed under sucrose-induced hyperosmotic stress in the sco1760::Tn5, sco4439/40::Tn5062, and sco0954 knockout mutants, as well as in S. coelicolor overexpressing sco0954, grown on GYM medium supplemented with 0.64 M sucrose. (a) Osmotic-stress-induced cell/EVs area. (b) Osmotic-stress-induced EVs abundance. Box and whisker elements: centreline, median; box limits, upper and lower quartiles; whiskers, 1.5x interquartile range.
The sco1760::Tn5, and sco4439/40::Tn5062 knockout mutants, as well as S. coelicolor overexpressing sco0954, produce S-cells ranging from 4 to 14 µm2 (corresponding to diameters of 2.2 to 4.2 μm)
Extracellular S-cell sizes were quantified from confocal images using Fiji48. S. coelicolor wild-type cultures, which do not form EVs, defined a 3.6 μm² area cut-off (grey bars in Fig. 1 histograms), with smaller areas corresponding to minor hyphal thickenings (Fiji quantifies any round structure, including small hyphal thickenings occasionally observed in wild-type hyphae, see images in Fig. S1). All mutants produced EVs exceeding this threshold on 0.64 M sucrose but not on 0.3 M (blue bars, Fig. 1), indicating that these structures correspond to S-cells. Most vesicles ranged from 4 to 14 μm² (diameters of 2.2 to 4.2 μm) (Fig. 3a), consistent with reported sizes of Streptomyces L-forms (0.5–7 μm diameter; 0.2–38.4 μm² area)24,25, and K. viridifaciens S-cells (5.14 μm diameter; 20.73 ± 11.53 μm² area) and L-forms (3 μm diameter; 7.06 ± 5.87 μm² area)26. These sizes were larger than those from Streptomyces EVs (0.02 to 2.5 μm in diameter, 0.005–4.9 μm²)17,21, although they overlapped in the 4 to 4.9 μm² range.
Extracellular vesicle abundance was estimated from confocal images by calculating the percentage of SYTO9/propidium iodide-stained hyphal area occupied by vesicles (Fig. 3b; see Methods). Vesicle abundance in sco4439/40::Tn5062 and sco0954-overexpressing strains ranged from 2 to 14%, whereas sco1760::Tn5 showed lower levels (0–8%) (Fig. 3b), which as stated in the previous paragraph, corresponds with a lower membrane-to-wall ratio in this mutant (Fig. 2h).
Extracellular vesicle formation induced by hyperosmotic sucrose stress in sco1760::Tn5 and sco4439/40::Tn5062 mutants is complemented by sco1758 and sco4439/40, respectively
The Tn5 transposon in the sco1760::Tn5 mutant is inserted at chromosomal position 1,881,109, near the end of the sco1760 ORF (triangle, Fig. 4a), potentially affecting expression of downstream sco1759–58 (Fig. 4a). To pinpoint the gene(s) responsible for the phenotype, we performed complementation with various sco1760–58 combinations. Two promoters in this region, at positions 1,881,810 and 1,880,358 (P1 and P2, Fig. 4a), were previously identified by Jeong et al.49. Three complementation plasmids were constructed: one with sco1760 alone; one with sco1760 and sco1759; and one with sco1760, sco1759, and sco1758. All included upstream sequences covering the P1 promoter and were cloned into the integrative ΦC31 vector pRASK50.
Formation of extracellular vesicles (EVs) under sucrose-induced hyperosmotic stress in the sco1760::Tn5, sco4439/40::Tn5062, and sco0954 knockout mutants is complemented by sco1758 and sco4439/40. Images and histograms are as described in Fig. 1. (a) Genetic region disrupted by the Tn5 transposon; the arrowhead indicates the insertion site. P1 and P2 denote the two promoters identified by Jeong et al.49. (b-d) Images and histograms of the mutants (e) SCO1758 gene expression in 48-hour GYM solid cultures. (f-h) Images and histograms of the complemented mutant strains. (i, j) Images and histograms of the S.coelicolor wild-type strain harbouring the empty pRASK and pMS82 plasmids. All cultures were grown on GYM medium supplemented with 0.64 M sucrose. Asterisks indicate statistically significant differences compared to the wild-type strain: p < 0.05, p < 0.01, **p < 0.001. Arrows indicate EVs and sucrose-driven hyperosmotic stress-induced cells. Scale bars: 8 μm.
Neither sco1760 alone nor sco1760 with sco1759 restored the wild-type phenotype (absence of EVs under hyperosmotic sucrose stress) (Fig. 4b, c; blue bars; see Supplementary Fig. S1 for more images). Only the construct containing all three genes restored the wild-type phenotype (Fig. 4d), identifying sco1758 as being involved in EV formation under hyperosmotic sucrose stress. However, sco1760 and/or sco1759 may also contribute to the sco1760::Tn5 mutant phenotype.
Transcript analysis revealed a sevenfold decrease in sco1758 expression in the sco1760::Tn5 mutant versus wild type, while expression was restored in the strain complemented with sco1760-1758 (Fig. 4e), which also rescued the phenotype (Fig. 4d), supporting sco1758’s role in EVs formation.
In the sco4439/40::Tn5062 mutant, EVs formation under hyperosmotic sucrose stress was suppressed only when both sco4439 and sco4440 were co-expressed (Fig. 4f–h). Control plasmids pMS8251 and pRASK50 had no effect (Fig. 4i, j).
The sco1760::Tn5 mutant forms hyperosmotic sucrose stress-induced, division-capable cells (L-forms) when grown on GYM medium supplemented with 0.64 M sucrose
We next examined the dynamics of osmotic stress-induced vesicles using time-lapse microscopy. Observations began at 48 h on GYM medium with 0.64 M sucrose, when vesicles were already present in all mutants. Samples were stained with SYTO-9 (green; DNA stain) (Fig. 5; Movies S1–S3).
Dynamics of extracellular vesicles (EVs) induced by sucrose-driven hyperosmotic stress in GYM cultures supplemented with 0.64 M sucrose. (a) sco1760::Tn5 mutant. (b) Wild-type strain overexpressing sco0954. (c) sco4439/40::Tn5062 mutant (upper images in phase-contrast mode). Time-lapse imaging was initiated in 48-hour cultures, once EVs induced by sucrose-driven hyperosmotic stress had formed. Cultures were stained with SYTO-9 (green, DNA stain). Arrows indicate S-cells and L-forms. Time points (in minutes) are indicated; time 0 marks the start of time-lapse imaging (i.e. a 48-hour culture). Scale bars represent 8 μm.
In the sco1760::Tn5 mutant, we observed cells forming as lateral evaginations from hyphae, producing distinct round vesicles clearly separated from the original hyphae (Fig. 5a, upper panels; Movie S1). We also observed chromosomal division (indicated by arrows in Fig. 5a, lower panels; Movie S1) and the separation of two independent chromosomes, which appear to be two independent cells at the 240-minute time point (Fig. 5a, lower panels). These events resemble the L-forms described by Ramijan et al.26 in Kitasatospora viridifaciens, which arise under osmotic stress and are capable of division.
L-forms were not seen in the sco4439/40::Tn5062 mutant or in the S. coelicolor strain overexpressing sco0954 (Fig. 5b–c). Both strains produced stable vesicles resembling the S-cells defined by Ramijan et al.26 in 48-hour GYM cultures supplemented with 0.64 M sucrose, corresponding to the aerial hyphae stage preceding sporulation (Fig. 5b–c; Movies S2–S4). Some EVs in the sco4439/40::Tn5062 mutant lysed (see the large cell indicated by an arrow in Fig. 5c). Based on these data, we cannot currently determine the dynamics of EV formation in these two mutants.
The SCO1758 EngA GTPase, SCO0954 methionine N-acetyltransferase, SCO4439 D-Ala-D-Ala carboxypeptidase, and SCO4440 GOLPH3-like protein are highly conserved among Streptomyces species
In order to try to understand the mechanism by which mutations in sco1758, sco4439/40, and the overexpression of sco0954 lead to the formation of wall-deficient cells under hyperosmotic sucrose stress conditions, we studied the homology and predicted functions of these proteins across six model Streptomyces strains (S. lividans, S. avermitilis, S. venezuelae, S. scabies, S. griseus, and S. clavuligerus).
SCO1758 carries a putative EngA GTPase domain (CDD: TIGR03594), sharing 98% similarity across six model Streptomyces species (S. lividans, S. avermitilis, S. venezuelae, S. scabies, S. griseus, S. clavuligerus). EngA GTPases, part of a conserved GTPase superfamily, act as molecular switches regulating key processes such as chromosome segregation, cell division, and ribosome stability in E. coli52,53,54. Their role in wall-deficient cell formation is plausible; however, to the best of our knowledge, no study to date has described an effect of EngA GTPases on the formation of wall-deficient cells. The prototype EngA GTPases from C. jejuni, B. subtilis, H. influenzae, M. genitalium, and M. leprae are characterised by two tandem GTP-binding domains, each with G-1, G-3, and G-4 motifs55,56, which are also present in SCO1758 (Fig. 6a). The G-1 motif, a flexible loop found in many nucleotide-binding proteins, interacts with phosphate groups (Prosite: PS00017).
Homology of the SCO1758, SCO0954, SCO4439, and SCO4440 proteins. (a) Alignment of SCO1758 with the prototype bacterial EngA GTPases55, showing two GTP-binding domains, the G-1, G-3, and G-4 motifs, and the P-loop characteristic of EngA GTPases. (b) Conserved domain and protein conservation of SCO0954. (c) Conserved domains and protein conservation of SCO4439 and SCO4440. Conserved domain database accession numbers are provided, along with the average similarity of the protein orthologues across six model Streptomyces species: S. lividans, S. avermitilis, S. venezuelae, S. scabies, S. griseus, and S. clavuligerus.
sco0954 encodes a 160-residue protein with 75% average similarity in Streptomyces and contains a GCN5-related acetyltransferase (GNAT) MnaT domain (CDD: COG1247) (Fig. 6b). This domain defines the MddA subfamily of GNATs, first characterised in Salmonella enterica57. GNATs acetylate diverse molecules—amino acids, polyamines, antibiotics—modulating key pathways58. The S. enterica MddA specifically acetylates oxidative methionine derivatives, blocking their uptake and toxicity57.
SCO4439 contains a D-Ala-D-Ala carboxypeptidase domain (70% similarity) and a putative transcriptional regulator domain (44%), separated by a transmembrane region46. SCO4440 is highly conserved (93% similarity) and harbours a GOLPH3 (GPP34) domain (CDD: pfam05719) (Fig. 1c). Eukaryotic GOLPH3 proteins are conserved elements of the trans-Golgi network, directing protein trafficking via vesicle sorting59. GOLPH3-like proteins have not been previously described in prokaryotes. Human GOLPH3 (NP_071413.1) shares > 64% identity with animal homologues, and 39.73% with S. cerevisiae59, whereas SCO4440 shows only 27% identity with the human protein. Notably, sco4439–sco4440 synteny is conserved in all six model Streptomyces species (Fig. 1c). A search of the NCBI gene database (15 February 2023) identified 218 bacterial species encoding GOLPH3-domain proteins, 129 of which were actinomycetes (Supplementary Table S2).
Methionine acetylation and oxidation are significantly increased in peptidoglycan-anchored peptides of Streptomyces coelicolor overexpressing sco0954
As indicated in the previous paragraph, SCO0954 contains an MnaT domain (Fig. 6b), characteristic of the MddA family of methionine sulfoximine/L-methionine sulfone acetyltransferases. In Salmonella enterica, the MddA prototype acetylates extracellular oxidised methionine at its amino terminus, thereby preventing its uptake and avoiding cytosolic toxicity caused by oxidative damage57. We hypothesise that, analogous to S. enterica MddA, SCO0954 may acetylate extracellular methionine. While the primary peptidoglycan (PG) scaffold lacks methionine, sortases attach peptides to PG60, and the N-terminal methionine of these peptides could potentially be acetylated by SCO0954.
Quantifying acetylated methionine is challenging, as the most common method for amino acid quantification, o-phthalaldehyde derivatisation, is ineffective, since it targets the amino group, which is blocked by the acetyl moiety61. Moreover, acid hydrolysis, which is the most used method to release free amino acids, removes the acetyl group from methionine and is therefore unsuitable (Supplementary Fig. S2). To address these issues, we developed a method involving pronase digestion to release amino acids from PG samples, followed by hydrophilic interaction liquid chromatography (HILIC) to quantify acetylated methionine without derivatisation (see Methods). We measured levels of methionine (M) and N-acetylmethionine (Ac-M), as well as their oxidised forms: M sulfone, M sulfoxide, Ac-M sulfone, and Ac-M sulfoxide.
The S. coelicolor strain overexpressing sco0954 exhibited a marked increase in Ac-M and Ac-M sulfoxide levels, while Ac-M sulfone was only modestly elevated, relative to the wild-type strain carrying the empty pRASK-PermE* plasmid (Fig. 7a). Non-acetylated M sulfone was slightly elevated in the sco0954-overexpressing strain, whereas non-acetylated M sulfoxide levels remained comparable to the control strain (Fig. 7b). Overall, acetylated oxidised methionine species were more abundant than their non-acetylated counterparts, especially in the strain overexpressing sco0954 (compare Fig. 7a and b).
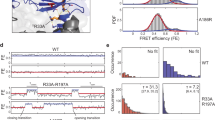
Acetylated/oxidised methionine in PG-associated peptides, and the effect of dithiothreitol on the inhibition of EVs formation, in the S. coelicolor strain overexpressing sco0954. (a) Levels of acetylated PG-associated methionine in the S. coelicolor strain overexpressing sco0954, compared to the S. coelicolor wild-type strain carrying the empty pRASK-PermE* plasmid. (b) Levels of non-acetylated, oxidised PG-associated methionine. (c–e) Absence of S-cells induced by sucrose-driven hyperosmotic stress in GYM medium supplemented with 0.64 M sucrose and 5 mM DTT, in the S. coelicolor strain overexpressing sco0954, and in the wild-type control strain with and without pRASK-PermE*. (f) Presence of hyperosmotic-stress induced cells in the sco4439/40::Tn5062 mutant in GYM medium supplemented with 0.64 M sucrose and 5 mM DTT. (g) Presence of hyperosmotic-stress induced cells in the sco1760::Tn5 mutant under the same conditions. Images and histograms are as in Fig. 1. Asterisks indicate statistically significant differences (p < 0.05, *p < 0.01, **p < 0.001). Error bars indicate SD.
These findings are consistent with SCO0954 possessing methionine acetyltransferase activity, potentially targeting oxidised methionine—particularly M sulfoxide, whose acetylated derivative showed the greatest increase. Nonetheless, further studies are required to fully elucidate the mechanism by which SCO0954 enhances levels of Ac-M and Ac-M sulfoxide in PG. As discussed below, these alterations in PG-associated peptides, may contribute to induction of wall-deficient cell formation.
Dithiothreitol inhibits wall-deficient cell formation in S. coelicolor overexpressing sco0954
Having found increased oxidised methionine (Ac-Met sulfoxide and Met sulfone) in S. coelicolor overexpressing sco0954 (Fig. 7a, b), we tested whether a strong reducing agent (5 mM dithiothreitol, DTT) affects wall-deficient cell formation. DTT had no effect in the wild-type strain (with or without pRASK-PermE*) grown on GYM with 0.64 M sucrose, as wall-deficient cells were still not produced (Fig. 7c, d; see Supplementary Fig. S1 for representative images). However, DTT blocked S-cell formation in the strain overexpressing sco0954 (Fig. 7e). This supports a role for SCO0954 in modulating PG oxidation, as reported for the Salmonella enterica MddA protein57, and suggests that redox changes in PG driven by sco0954 overexpression influence wall-deficient cell formation. Notably, DTT does not suppress the formation of wall-deficient cells in the sco4439/40::Tn5062 and sco1760::Tn5 mutants (Fig. 7f, g).
Methionine oxidation and acetylation are post-translational modifications analogous to phosphorylation62,63, capable of altering the conformation of PG-associated proteins and, in turn, PG structure.
The sco1760::Tn5 and sco4439/40::Tn5062 mutants, as well as the S. coelicolor wild-type strain overexpressing sco0954, exhibit altered peptidoglycan muropeptide composition
Peptidoglycan subunits, or muropeptides, differ between bacterial species64,65, and such variation affects PG architecture and cell shape66. We hypothesised that muropeptide composition might be altered in sco1760::Tn5 and sco4439/40::Tn5062 mutants, influencing PG properties and contributing to osmotic-stress-induced cell formation.
Hugonnet et al.67 previously characterised Streptomyces coelicolor PG muropeptides via LC-MS/MS (summarised in Fig. 8a). Here, we compared PG profiles of the wild-type (non-producing stress-induced cells) and mutants (producing stress-induced cells) (Figs. 8b–d), focusing on spores, as they derive from sporulating hyphae that are those involved in stress-induced cell production. While spores are easily isolated, sporulating hyphae are confined to the upper region of aerial hyphae and cannot be separated from non-sporulating forms68,69. We employed high-resolution mass spectrometry (ESI-Q-TOF Impact II HR-MS spectrometer) to identify muropeptides based on their exact monoisotopic masses, with a mass accuracy of ± 0.005 Da. Signal intensities were normalised to the tetra(Gly) monomer, the most abundant muropeptide in our PG analyses (see Methods for details).
Muropeptide composition of peptidoglycan in the sco4439/40::Tn5062 and sco1760::Tn5 mutants, and in the corresponding complemented strains. (a) S. coelicolor, PG monomer and dimer (4−3 and 3–3 PG cross-linkages) muropeptide structures, with calculated monoisotopic masses. (b) Heatmap illustrating muropeptide abundance in the sco4439/40::Tn5062 and sco1760::Tn5 mutants, compared to the S. coelicolor wild-type strain with or without the empty pRASK plasmid. (c) Heatmap showing muropeptide abundance in the sco1760::Tn5 mutant and its complemented strain, compared to the mutant harbouring pRASK. (d) Heatmap showing muropeptide abundance in the sco4439/40::Tn5062 mutant and its complemented strains, compared to the mutant harbouring pRASK. The relative abundances of the muropeptides shown in the heatmaps (rows) exhibit statistically significant differences in at least one of the analysed strains compared to the corresponding reference strain (reference strains are highlighted in red). Asterisks indicate significant differences (p < 0.05, p < 0.01, *p < 0.001).
Figure 8b shows altered muropeptide profiles in the sco4439/40::Tn5062 and sco1760::Tn5 mutants versus wild type (with or without pRASK). Both mutants exhibited increased tetra-tetra(Gly) (4–3) dimer (muropeptide 11); sco4439/40::Tn5062 also showed elevated muropeptide 14. This aligns with SCO4439’s D-Ala-D-Ala carboxypeptidase activity46, whose absence leads to pentapeptide accumulation and subsequent (4–3) dimer formation. Figure 8c indicates that complementation with sco1758–1760 restored muropeptide 11 to wild-type levels in sco1760::Tn5, matching phenotypic rescue (Fig. 4d). Figure 8d shows that sco4440 complementation reduced muropeptides 11 and 14 to wild-type levels in sco4439/40::Tn5062, whereas sco4439 alone or with sco4440 had minimal effect.
Overall, these findings suggest that SCO4439, SCO4440, and SCO1758 influence the PG muropeptide composition of S. coelicolor. As discussed below, these PG alterations appear to contribute to the induction of wall-deficient cell formation.
Discussion
As introduced above, non-replicative extracellular membrane vesicles and wall-deficient cells capable of division (L-forms) have been reported in many bacteria, including actinomycetes17,18,19,26,70, whereas wall-deficient cells unable to divide and induced by hyperosmotic stress (S-cells) have been specifically characterised in actinomycetes26. L-forms have been described as forming through a passive biophysical mechanism, dependent on an excess of intracellular membranes36,37,38. A similar passive mechanism for cell division has been proposed for the formation of EVs at the hyphal tips of Streptomyces venezuelae20,21. A biochemical modulation of EVs, S-cell, and L-form formation is compatible with this passive biophysical mechanism, as the intracellular membrane excess may be subject to genetic regulation. In this context, some genes (murE, mraY, ftsQ, filP, stlP) have already been reported to modulate L-form and S-cell formation in bacteria23,37,42,43. EV formation in gram-positive bacteria has been reported to be under genetic control (sigB, covRS)35. Our study advances this understanding by demonstrating that S-cell formation in Streptomyces coelicolor is triggered by the inactivation of sco1758, sco4439, and sco4440, as well as by the overexpression of sco0954. Moreover, the inactivation of sco1758 induces both L-form and S-cell formation (Fig. 1). We also found that wall-deficient cells are only present in 48-hour cultures, which correspond to the aerial hyphae stage preceding sporulation, suggesting a connection between hyphal differentiation and the formation of wall-deficient cells. Hyperosmotic stress (0.64 M sucrose) trigger wall-deficient cell formation, whereas lower osmotic stress (0.3 M sucrose, sufficient to prevent osmotic lysis of wall-deficient cells) does not induce their formation (Fig. 1). Consequently, wall-deficient cell formation appears to be actively triggered under the 0.64 M sucrose condition, rather than sucrose merely protecting wall-deficient cells, produced under all conditions, from osmotic lysis. As Streptomyces coelicolor lacks invertase, it is unable to metabolise sucrose71,72. The formation of wall-deficient cells at high sucrose concentrations in the mutants is therefore likely driven by osmotic stress, rather than by a metabolic effect of sucrose, as previously reported in other actinomycetes such as the wild-type strains of Kitasatospora viridifaciens, Streptomyces griseus and Streptomyces venezuelae26, as well as in the S. coelicolor strain lacking the stomatin-like protein StlP43. In these actinomycetes, S cells are induced not only by sucrose, but also by other osmolytes such as NaCl and sorbitol26,43. Further studies will be required to determine whether osmolytes other than sucrose can similarly trigger S-cell formation in the sco1760::Tn5 and sco4439/40::Tn5062 mutants, as well as in S. coelicolor overexpressing sco0954.
The PG structure is altered in all the S. coelicolor mutants producing wall-deficient cells. Acetylated and oxidised methionine from PG-associated peptides is highly increased in the S. coelicolor strain overexpressing sco0954 (Fig. 7a), whereas PG muropeptide composition is altered in the sco4439/40::Tn5062 and sco1760::Tn5 mutants (Fig. 8). PG muropeptide composition is fundamental in defining PG 3D structure, PG resistance, and bacterial cell shape. In fact, the shape of purified PG is reminiscent of the bacterial cell73,74. Methionine oxidation and acetylation also influence PG structure, as both post-translational modifications significantly affect protein conformation and function, in a manner similar to that of other PTMs such as amino acid phosphorylation62,63.
We propose the model outlined in Fig. 9, describing the effect of SCO0954, SCO4439, SCO4440 and SCO1758 on PG structure and wall deficient cell formation under hyperosmotic sucrose conditions. First, sco4439/40::Tn5062 mutant lack D-Ala-D-Ala carboxypeptidase activity46, leading to the accumulation of pentapeptides, which serve as substrates for increased formation of 4–3 cross-linked dimers (Fig. 8b). Second, sco0954 overexpression increases acetylated oxidised methionine (Fig. 7a), consistent with SCO0954 acting as an N-acetyltransferase of oxidised methionine, similar to the prototype MddA N-acetyltransferase from Salmonella enterica57. Third, down-regulation of SCO1758 (EngA GTPase) in sco1760::Tn5 (Fig. 4e) increases 4–3 dimers (Fig. 8b). Analogously to other GTPases involved in PG biosynthesis as the FtsZ GTPase75 or the CpgA GTPase76, SCO1758 might interact with specific PG reshuffling enzymes, positioning them in the hyphae destined to form wall-deficient cells. Fourth, SCO4440 also influences 4–3 muropeptide dimers, as their levels decrease when sco4439/40::Tn5062 is complemented with sco4440 (Fig. 8d). SCO4440 contains a GOLPH3-like domain, a feature of eukaryotic trans-Golgi proteins (Fig. 6c). While not previously reported in prokaryotes, GOLPH3-like proteins are widespread in actinomycetes (Supplementary Table S2), hinting at a role in a putative secretory network similar to the eukaryotic Golgi59,77. Like other cytoskeleton-interacting proteins involved in S/L-form (the Kitasatospora viridifaciens FilP cytoskeletal protein)42 and in the formation of intracellular membrane vesicles (the S. coelicolor SCO4256-4258 contractile cytosolic bacteriophage tail-like injection systems)22, SCO4440 might link membrane proteins with cytoskeleton elements.
Model illustrating the roles of SCO4439, SCO4440, SCO0954, and SCO1758 in modulating peptidoglycan structure. The putative functions of SCO4439 (labelled as 1), SCO0954 (2), SCO1758 (3), and SCO4440 (4) in influencing PG muropeptide composition and the acetylation of oxidised methionine in PG-associated peptides are depicted. Solid arrows indicate direct modifications to the PG, while dashed arrows represent indirect effects on its structure. See Discussion for details.
In summary, SCO0954 (N-acetyltransferase), SCO4439 (carboxypeptidase), SCO4440 (GOLPH3-like), and SCO1758 (GTPase) contribute to wall-deficient cell formation under hyperosmotic sucrose stress (non-dividing S-cells) and SCO1758 is also involved in the formation of divisible L-forms. Alterations in muropeptide composition, particularly 4–3 dimers and PG-associated acetylated and oxidised methionine, modulate this process. S. coelicolor stress-induced wall-deficient cells (4–14 μm²; Fig. 3a) are up to four times larger than Streptomyces EVs (0.02–4.9 μm²)17,21, with both overlapping in the 4–4.9 μm² range, suggesting they are distinct structures that may nonetheless share mechanisms of formation and regulation. Additional research is needed to further elucidate how these proteins modulate PG architecture, determine whether homologous proteins induce wall-deficient cells in other bacteria, and to explore the potential of these genes to promote EV formation for biotechnological applications.
Methods
Bacterial strains and culture conditions
All Streptomyces and Escherichia coli strains used are listed in Supplementary Table S3. Spores were harvested from SFM solid plates47 after incubation at 30 °C for 7 days. Osmotic-stress-induced cell formation was assessed on GYM78 (5 g/L glucose, 4 g/L yeast extract, 5 g/L malt extract, 0.5 g/L MgSO₄·7 H₂O, 20 g/L agar; post-autoclaving, supplemented with sterile 0.5 g/L K₂HPO₄). Twenty-five millilitres of medium were dispensed into 90-mm, 3-vent Petri dishes. Plates were overlaid with cellophane, inoculated with 3 × 10⁷ fresh (non-frozen) spores, and incubated at 30 °C. Hyperosmotic stress to induce wall-deficient S-cell formation was created by the addition of 0.64 M sucrose, as described by Ramijan et al.26.
E. coli strains were grown in LB or 2×TY at 37 °C. Antibiotics for plasmid and mutant selection included: ampicillin (100 µg/mL), apramycin (100 µg/mL for E. coli, 25 µg/mL for S. coelicolor), hygromycin (100 µg/mL for E. coli, 200 µg/mL for S. coelicolor), kanamycin (50 µg/mL), thiostrepton (50 µg/mL), chloramphenicol (25 µg/mL), and nalidixic acid (25 µg/mL).
Random mutagenesis
The S. coelicolor sco1760 ORF, disrupted by mini-Tn5 (sco1760::Tn5), was identified from a random mutagenesis library generated in our laboratory using the method of Xu et al.45.
Directed mutagenesis
The sco4439 ORF was disrupted in a previous study46 using the Tn5062-based single-gene knockout library developed by Fernández-Martínez et al.79. As shown by Rioseras et al.46, the transposon insertion caused a deletion encompassing the promoter regions of sco4439 and sco4440, thereby abolishing expression of both genes.
Complementation of the sco4439/40::Tn5062, and the sco1760::Tn5 mutants
The sco4440 was previously cloned into pMS82 by Rioseras et al.46. In this study, sco4439 was excised from pTOPO [sco4439]46 using EcoRV and SpeI, and cloned into similarly digested pRASK50, yielding pRASK [sco4439].
sco4440 and the sco4440-41-42 fragment were excised from their respective pTOPO constructs46 using the EcoRV and SpeI and cloned into pRASK, generating pRASK [sco4440] and pRASK [sco4440-41-42], respectively.
To construct pRASK [sco4439-4440], a PCR product from wild-type genomic DNA was amplified using primers sco4439/4440-F and sco4439/4440-R, cloned into pCR™-Blunt II-TOPO (Thermo Fisher), sequenced, and subcloned into pRASK via SpeI and EcoRV.
sco1760, sco1760-1759, and sco1760-1758 (including its upstream P1 promoter region) were amplified using primer pairs sco1760F/sco1760R, sco1760F/SCO1759R, and sco1760F/sco1758R, respectively. Products were cloned into pCR™-Blunt II-TOPO (Thermo Fisher), sequenced, and subcloned into pRASK50.
The pRASK plasmids containing the genes used for complementation were introduced into the mutants via conjugation47. Integration of ɸC31 was confirmed using the primers sco3798intF and intR80.
Protein sequence analyses
Orthologous amino acid sequences of S. lividans, S. avermitilis, S. venezuelae, S. scabies, S. griseus, and S. clavuligerus for SCO0954, SCO4439, SCO4440, and SCO1758 were retrieved from StrepDB (http://strepdb.streptomyces.org.uk/). Conserved domains were identified using the Conserved Domain Database (CDD) (https://www.ncbi.nlm.nih.gov/Structure/cdd/cdd.shtml). Pairwise amino acid similarities were calculated with EMBOSS Needle (https://www.ebi.ac.uk/Tools/psa/emboss_needle/). Sequence alignments were generated using MUSCLE via the Phylemon platform (http://phylemon.bioinfo.cipf.es/)81.
DNA and RNA extraction
Genomic DNA was isolated using standard methods47. RNA was extracted with Direct-zol™ RNA columns (Zymo-Spin™), treated with TURBO™ DNase (Thermo Fisher) and assessed using a Nanodrop 2000 (Thermo Fisher) and a 2100 Bioanalyzer (Agilent).
FtsZ mutant transcriptomic analyses
Next-generation sequencing (NGS) was performed by Stab Vida (Caparica, Portugal) using two biological replicates of the S. coelicolor FtsZ mutant14 and the M145 wild-type strain. Ribosomal RNA was depleted with the Ribo-Zero Bacteria Kit (Illumina), and cDNA libraries were prepared using the TruSeq Stranded mRNA Library Preparation Kit (Illumina). Sequencing was conducted on the Illumina HiSeq 2500 platform with 100-bp paired-end reads. Bioinformatic analyses were performed on Linux using FastQC (quality control), Cutadapt (trimming), Bowtie2 (mapping to the S. coelicolor genome), and Cuffdiff (differential expression analysis)82.
Overexpression of sco0954
The sco0954 gene was amplified from S. coelicolor using primers sco0954F/R (Table S3), which included NdeI and SpeI. The amplicons were cloned into pCR™-Blunt II-TOPO and sequenced by Sanger using M13F/R. The ORFs were excised from pCR™-Blunt II-TOPO with NdeI and SpeI and subcloned into pNG483 using the same enzymes. PermE-sco0954 was excised from pNG4 with MfeI and BglII and cloned into pRASK50 (pRASK-PermE[sco0954]). The empty pRASK-PermE* plasmid was constructed by digesting pRASK-PermE*[sco0954] with NdeI and HpaI, blunting with S1 nuclease (Thermo Fisher™) and religating.
Real-time quantitative reverse-transcription PCR (qRT-PCR)
cDNA was synthesised from 1.5 µg of RNA from two biological replicates using the High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems, Waltham, MA, USA). Real-time PCR was performed on an FQD-96 A fluorescence detection system (BIOER, China) in triplicate reactions containing 2 µL cDNA, 10 µL PowerTrack™ SYBR Green Master Mix (Thermo Fisher), and 300 nM of gene-specific primers (Table S3) in a final volume of 20 µL. sco1758 was amplified with primers q1758F2/R2 (Table S3). Negative controls (RNA or water in place of cDNA) were used to assess DNA contamination and primer-dimer formation. Amplification conditions: 2 min at 50 °C, 10 min at 95 °C, then 40 cycles of 15 s at 95 °C and 1 min at 60 °C.
Absolute quantification was performed using standard curves generated from PCR-amplified sco1758 fragments84, all with correlation coefficients > 0.99. Transcript abundance was normalised by amplicon size and molecular weight, and expressed as transcript copies per microgram of cDNA. Three biological replicates were analysed per gene. Statistical significance was assessed by two-sided t-tests; p-values < 0.05, < 0.01, or < 0.001 were considered significant.
Cell morphology, viability, membrane, cell-wall staining, time-lapse analyses
Cell morphology and viability were assessed using PI and SYTO 9 stains from the LIVE/DEAD BacLight Bacterial Viability Kit (Invitrogen). Stains were prepared in 103 g/L sucrose to prevent osmotic lysis. Samples were imaged using a Leica TCS-SP8 confocal laser-scanning microscope (excitation: 488 nm and 568 nm; emission: 530 nm [green] and 640 nm [red])85.
Cell walls were stained with Alexa Fluor 488-conjugated WGA (Invitrogen), which binds N-acetylglucosamine and N-acetylmuramic acid. Streptomyces hyphae were scraped from cellophane with a spatula and processed as described15. Samples were imaged at excitation/emission wavelengths of 498 nm/520 nm.
Time-lapse imaging was performed using FM5-95 (3.95 µg/mL) and SYTO9 (0.5 µmol/L) in solid cultures. Cultures were grown on cellophane, pre-incubated at 30 °C for 48 h, and transferred to fresh medium containing the stains. The sheets were inverted onto coated µ-dishes (Ibidi GmbH) and incubated at 30 °C. Imaging was performed every 13 min over 17 h on a Leica TCS-SP8 confocal microscope (excitation: 488 nm and 522 nm; emission: 530 nm [SYTO9] and 782 nm [FM5-95]). The microscope chamber was equilibrated for 3 h prior to imaging. Unstained cultures served as autofluorescence controls. Images were processed using Fiji software48 to adjust histograms, add scales, and assemble time-lapse videos.
Quantification of osmotic-stress-induced EVs size and abundance
Cell size and abundance under osmotic stress were quantified from confocal images stained with PI and SYTO 9, or SYTO9 plus FM5-95. Vesicle size was measured using a Fiji macro48 with the Stardist plugin86, which extracted parameters including area, perimeter, circularity, Feret diameter, minimum Feret diameter, aspect ratio (AR), roundness, and solidity. A circularity threshold of 0.8 was applied (1.0 = perfect sphere). The vesicle area threshold was defined as 3.6 μm² based on control images of the wild-type strain (with or without empty plasmids). Over 100 vesicles exceeding this threshold were analysed from images taken from at least three biological replicates.
Vesicle abundance was calculated as the percentage of vesicle-stained area relative to total stained area per image. Vesicle area was quantified as above, and total stained area was measured by counting pixels above background and converting to µm². Background levels were estimated from hyphae-free regions. At least 100 vesicles were analysed from three biological replicates.
Statistical differences were assessed using a two-sided t-test, with p-values of < 0.05, < 0.01, or < 0.001 considered significant.
Quantification of membrane and cell wall abundance
The membrane-to-cell wall ratio was determined from FM5-95- and WGA-stained confocal images by quantifying pixels above background using Fiji48. Background levels for each stain were calculated per image from hyphae-free regions. For each strain, data from at least three biological replicates were combined, with a minimum of 1 × 10⁶ pixels above background analysed per replicate. The FM5-95/WGA pixel ratio reflects membrane abundance: a ratio of 1 indicates equal membrane and wall content; <1 indicates more wall than membrane; and > 1 indicates more membrane than wall. Statistical significance was assessed using a two-sided t-test, with p-values < 0.05, < 0.01, or < 0.001 considered significant.
Muropeptide abundance quantification
Peptidoglycan was purified from Streptomyces spores as described previously46,87, resuspended in 3 mL mQ and stored at − 80 °C. Streptomyces muropeptides67 were quantified by high-resolution LC-MS. PG (100 µL) was digested with 50 U/mL mutanolysin (A&A Biotechnology) and 2 mg/mL lysozyme in 25 mM phosphate buffer (pH 7) with 0.1 M MgCl₂. Lactoyl peptides were released using 32% ammonium hydroxide88, dried, and resuspended in 50 µL mQ water with 0.1% formic acid. Five µL were analysed by LC-HRESI-MS using a Dionex Ultimate 3000 UHPLC system and Bruker Impact II Q-TOF in positive mode (m/z 50–2500). Chromatographic separation used a Zorbax® Eclipse Plus C18 column (50 × 2.1 mm, 1.8 μm) at 0.25 mL/min with a formic acid/acetonitrile gradient.
Data were processed with DataAnalysis v4.3, calibrated with internal standards. Muropeptides were identified using a ± 0.005 Da mass and ± 0.1 min RT tolerance. [M + H]⁺ was dominant in monomers, [M + 2 H]²⁺ in dimers. Relative abundances (peak areas) were normalised to the most abundant Tetra (Gly) monomer. Statistical analyses were performed in MetaboAnalyst 6.089, with missing values imputed as 1/5 of the minimum, log₁₀-transformed, and Pareto scaled. Comparisons included wild-type and mutant strains with or without plasmids. Triplicate samples were analysed. Significance thresholds were p < 0.05, < 0.01, or < 0.001.
Quantification of N-acetylmethionine, N-acetylmethionine sulfone, and N-acetylmethionine sulfoxide
One hundred µL of purified PG were digested overnight at 37 °C with 50 U/mL mutanolysin (A&A Biotechnology) and 2 mg/mL lysozyme (GoldBio) in 25 mM phosphate buffer (pH 7) containing 0.1 M MgCl₂. The reaction was stopped by boiling for 3 min. After centrifugation (16,000 × g), the supernatant was digested with 0.2 mg/mL pronase (MedChem Express) at 37 °C for 20 h. Proteins were precipitated with 5% TCA on ice for 30 min, followed by centrifugation and vacuum-drying of the supernatant. The pellet was resuspended in 100 µL ultrapure water, vortexed, and diluted with 900 µL acetonitrile (ACN). An aliquot (25 µL) was injected into an Agilent 6410 LC-MS/MS system (ESI-QqQ) operated in MRM mode. Parameters included a 35 psi nebulizer, + 4000 V capillary voltage, and nitrogen gas at 12 L/min and 350 °C.
Chromatography was performed using a XBridge® Amide column (Waters) (4.6 × 150 mm, 3.5 μm; Waters) at 1 mL/min with a gradient of Solvent A (0.1% formic acid in water) and Solvent B (ACN): 85–80% B (0–5 min), 80–50% B (5–15 min), and re-equilibrated at 85% B for 4 min. Standards included N-acetyl-L-methionine, its sulfoxide and sulfone forms (AOZEAL and Sigma-Aldrich), as well as non-acetylated forms (Sigma-Aldrich). Instrument parameters are listed in Supplementary Table S4. Three biological replicates per sample were analysed; statistical significance was set at p < 0.05, < 0.01, or < 0.001.
Data availability
The authors declare that the data supporting the findings of this study are available within the article and its supplementary information files or from the corresponding authors on request. The RNAseq raw data are available on the SRA database (https://www.ncbi.nlm.nih.gov/sra) under the PRJNA1024652 accession number.
References
Berdy, J. Bioactive microbial metabolites. J. Antibiot. (Tokyo). 58, 1–26 (2005).
Hopwood, D. A. Streptomyces in Nature and Medicine: the Antibiotic Makers (Oxford University Press, 2007).
Flardh, K. & Buttner, M. J. Streptomyces morphogenetics: dissecting differentiation in a filamentous bacterium. Nat. Rev. Microbiol. 7, 36–49 (2009).
Yagüe, P. et al. Pre-sporulation stages of streptomyces differentiation: State-of-the-art and future perspectives. FEMS Microbiol. Lett. 342, 79–88 (2013).
McCormick, J. R. Cell division is dispensable but not irrelevant in streptomyces. Curr. Opin. Microbiol. 12, 689–698 (2009).
Jakimowicz, D. & van Wezel, G. P. Cell division and DNA segregation in streptomyces: how to build a septum in the middle of nowhere? Mol. Microbiol. 85, 393–404 (2012).
Willemse, J. et al. Positive control of cell division: Ftsz is recruited by Ssgb during sporulation of streptomyces. Genes Dev. 25, 89–99 (2011).
Del Sol, R. et al. Influence of Crga on assembly of the cell division protein Ftsz during development of streptomyces coelicolor. J. Bacteriol. 188, 1540–1550 (2006).
Schlimpert, S. et al. Two dynamin-like proteins stabilize Ftsz rings during streptomyces sporulation. Proc. Natl. Acad. Sci. U S A. 114, E6176–E6183 (2017).
Ramos-Leon, F. et al., A conserved cell division protein directly regulates ftsz dynamics in filamentous and unicellular actinobacteria. Elife, 10: (2021).
Yagüe, P. et al. Subcompartmentalization by cross-membranes during early growth of streptomyces hyphae. Nat. Commun. 7, 12467 (2016).
Celler, K. et al. Cross-membranes orchestrate compartmentalization and morphogenesis in streptomyces. Nat. Commun. 7, ncomms11836 (2016).
Bush, M. J. et al. Hyphal compartmentalization and sporulation in streptomyces require the conserved cell division protein Sepx. Nat. Commun. 13, 71 (2022).
McCormick, J. R. et al. Growth and viability of streptomyces coelicolor mutant for the cell division gene Ftsz. Mol. Microbiol. 14, 243–254 (1994).
Schwedock, J. et al. Assembly of the cell division protein Ftsz into ladder-like structures in the aerial hyphae of streptomyces coelicolor. Mol. Microbiol. 25, 847–858 (1997).
den Blaauwen, T., Hamoen, L. W. & Levin, P. A. The divisome at 25: the road ahead. Curr. Opin. Microbiol. 36, 85–94 (2017).
Schrempf, H. et al. Extracellular streptomyces vesicles: amphorae for survival and defence. Microb. Biotechnol. 4, 286–299 (2011).
Faddetta, T. et al. ,. Streptomyces coelicolor vesicles: Many molecules to be delivered. Appl Environ Microbiol, 88: e0188121 (2022).
Faddetta, T. et al. Unravelling the DNA sequences carried by streptomyces coelicolor membrane vesicles. Sci. Rep. 12, 16651 (2022).
Schrempf, H. & Merling, P. Extracellular streptomyces lividans vesicles: composition, biogenesis and antimicrobial activity. Microb. Biotechnol. 8, 644–658 (2015).
Frojd, M. J. & Flardh, K. Extrusion of extracellular membrane vesicles from hyphal tips of streptomyces Venezuelae coupled to cell-wall stress. Microbiol. (Reading). 165, 1295–1305 (2019).
Casu, B. et al., Cytoplasmic contractile injection systems mediate cell death in streptomyces. Nat. Microbiol., (2023).
Leaver, M. et al. Life without a wall or division machine in Bacillus subtilis. Nature 457, 849–853 (2009).
Gumpert, J. Growth characteristics and ultrastructure of protoplast type l-forms from streptomycetes. Z. Für Allgemeine Mikrobiologie. 22, 617–627 (1982).
Innes, C. M. & Allan, E. J. Induction, growth and antibiotic production of streptomyces viridifaciens l-form bacteria. J. Appl. Microbiol. 90, 301–308 (2001).
Ramijan, K. et al. Stress-induced formation of cell wall-deficient cells in filamentous actinomycetes. Nat. Commun. 9, 5164 (2018).
Owens, W. Isolation of Staphylococcus aureus l forms from experimentally induced bovine mastitis. J. Clin. Microbiol. 25, 1956–1961 (1987).
Kita, E. et al. Conversion of Salmonella typhimurium to l-forms contributes to the maintenance of acquired immunity against murine typhoid. Immunology 86, 206 (1995).
Allan, E., Hoischen, C. & Gumpert, J. Bacterial l-forms. Adv. Appl. Microbiol. 68, 1–39 (2009).
Ongenae, V. et al. Reversible bacteriophage resistance by shedding the bacterial cell wall. Open. Biol. 12, 210379 (2022).
Wohlfarth, J. C. et al. L-form conversion in gram-positive bacteria enables escape from phage infection. Nat. Microbiol. 8, 387–399 (2023).
Kapteijn, R. et al. Endocytosis-like DNA uptake by cell wall-deficient bacteria. Nat. Commun. 13, 5524 (2022).
Depelteau, J. S., Brenzinger, S. & Briegel, A. Bacterial and archaeal cell structure. (2019).
Welsh, J. A. et al. Minimal information for studies of extracellular vesicles (misev2023): from basic to advanced approaches. J. Extracell. Vesicles. 13, e12404 (2024).
Briaud, P. & Carroll, R. K. Extracellular vesicle biogenesis and functions in gram-positive bacteria. Infect. Immun., 88: (2020).
Mercier, R., Dominguez-Cuevas, P. & Errington, J. Crucial role for membrane fluidity in proliferation of primitive cells. Cell. Rep. 1, 417–423 (2012).
Siddiqui, R. A. et al. The analysis of cell division and cell wall synthesis genes reveals mutationally inactivated Ftsq and Mray in a protoplast-type l-form of Escherichia coli. FEMS Microbiol. Lett. 258, 305–311 (2006).
Briers, Y. et al. Intracellular vesicles as reproduction elements in cell wall-deficient l-form bacteria. PLoS One. 7, e38514 (2012).
Mercier, R., Kawai, Y. & Errington, J. Excess membrane synthesis drives a primitive mode of cell proliferation. Cell 152, 997–1007 (2013).
Errington, J. L-form bacteria, cell walls and the origins of life. Open. Biol. 3, 120143 (2013).
Castro, J. M., Sugiyama, H. & Toyota, T. Budding and division of giant vesicles linked to phospholipid production. Sci. Rep. 9, 165 (2019).
Ultee, E. et al. Formation of wall-less cells in Kitasatospora viridifaciens requires cytoskeletal protein filp in oxygen-limiting conditions. Mol. Microbiol. 115, 1181–1190 (2021).
Zhong, X. et al. The stomatin-like protein stlp organizes membrane microdomains to govern Polar growth in filamentous actinobacteria under hyperosmotic stress. Nat. Commun. 16, 2669 (2025).
Meyer, K. J. & Nodwell, J. R. Streptomyces extracellular vesicles are a broad and permissive antimicrobial packaging and delivery system. J. Bacteriol. 206, e0032523 (2024).
Xu, Z. et al., Large-scale transposition mutagenesis of streptomyces coelicolor identifies hundreds of genes influencing antibiotic biosynthesis. Appl. Environ. Microbiol., 83: (2017).
Rioseras, B. et al. ,. Characterization of sco4439, a d-alanyl-d-alanine carboxypeptidase involved in spore cell wall maturation, resistance, and germination in streptomyces coelicolor Sci Rep, 6: 21659 (2016).
Kieser, T. Practical Streptomyces Genetics (John Innes Foundation, 2000).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods. 9, 676–682 (2012).
Jeong, Y. et al. The dynamic transcriptional and translational landscape of the model antibiotic producer streptomyces coelicolor a3(2). Nat. Commun. 7, 11605 (2016).
Fernández-García, G. et al. The deor-like pleiotropic regulator sco1897 controls specialised metabolism, sporulation, spore germination, and phosphorus accumulation in streptomyces coelicolor. Commun. Biology. 7, 1457 (2024).
Gregory, M. A., Till, R. & Smith, M. C. Integration site for streptomyces phage phibt1 and development of site-specific integrating vectors. J. Bacteriol. 185, 5320–5323 (2003).
Caldon, C. E., Yoong, P. & March, P. E. Evolution of a molecular switch: universal bacterial Gtpases regulate ribosome function. Mol. Microbiol. 41, 289–297 (2001).
Bharat, A. & Brown, E. D. Phenotypic investigations of the depletion of Enga in Escherichia coli are consistent with a role in ribosome biogenesis. FEMS Microbiol. Lett. 353, 26–32 (2014).
Robinson, V. L. et al. Domain arrangement of der, a switch protein containing two Gtpase domains. Structure 10, 1649–1658 (2002).
van Doorn, L. J. et al. Molecular discrimination between Campylobacter jejuni, Campylobacter coli, Campylobacter Lari and Campylobacter upsaliensis by polymerase chain reaction based on a novel putative Gtpase gene. Mol. Cell. Probes. 11, 177–185 (1997).
Caldon, C. E. & March, P. E. Function of the universally conserved bacterial Gtpases. Curr. Opin. Microbiol. 6, 135–139 (2003).
Hentchel, K. L. & Escalante-Semerena, J. C. In Salmonella enterica, the gcn5-related acetyltransferase Mdda (formerly ynca) acetylates methionine sulfoximine and methionine sulfone, blocking their toxic effects. J. Bacteriol. 197, 314–325 (2015).
Burckhardt, R. M. & Escalante-Semerena, J. C. Small-molecule acetylation by gcn5-related n-acetyltransferases in bacteria. Microbiol. Mol. Biol. Rev., 84: (2020).
Sechi, S. et al. The multiple cellular functions of the oncoprotein golgi phosphoprotein 3. Oncotarget 6, 3493–3506 (2015).
Schneewind, O. & Missiakas, D. M. Protein secretion and surface display in gram-positive bacteria. Philos. Trans. R Soc. Lond. B Biol. Sci. 367, 1123–1139 (2012).
Roth, M. Fluorescence reaction for amino acids. Anal. Chem. 43, 880–882 (1971).
Boschi-Muller, S. et al. The enzymology and biochemistry of methionine sulfoxide reductases. Biochim. Biophys. Acta. 1703, 231–238 (2005).
Hollebeke, J., Van Damme, P. & Gevaert, K. N-terminal acetylation and other functions of nalpha-acetyltransferases. Biol. Chem. 393, 291–298 (2012).
Doyle, R. J., Chaloupka, J. & Vinter, V. Turnover of cell walls in microorganisms. Microbiol. Rev. 52, 554–567 (1988).
Horcajo, P., de Pedro, M. A. & Cava, F. Peptidoglycan plasticity in bacteria: Stress-induced peptidoglycan editing by noncanonical d-amino acids. Microb. Drug Resist. 18, 306–313 (2012).
Desmarais, S. M. et al. Peptidoglycan at its peaks: how chromatographic analyses can reveal bacterial cell wall structure and assembly. Mol. Microbiol. 89, 1–13 (2013).
Hugonnet, J. E. et al. Peptidoglycan cross-linking in glycopeptide-resistant actinomycetales. Antimicrob. Agents Chemother. 58, 1749–1756 (2014).
Chater, K. F. Regulation of sporulation in streptomyces coelicolor a3(2): A checkpoint multiplex? Curr. Opin. Microbiol. 4, 667–673 (2001).
Manteca, A., Fernández, M. & Sanchez, J. Mycelium development in streptomyces antibioticus atcc11891 occurs in an orderly pattern which determines multiphase growth curves. BMC Microbiol. 5, 51 (2005).
Lazenby, J. J., Li, E. S. & Whitchurch, C. B. Cell wall deficiency - an alternate bacterial lifestyle? Microbiol. (Reading), 168: (2022).
Parche, S. et al. Sugar uptake and utilisation in streptomyces coelicolor: A Pts view to the genome. Antonie Van Leeuwenhoek. 78, 243–251 (2000).
Santamaría, R. I. et al. Morphological and physiological changes in streptomyces lividans induced by different yeasts. Arch. Microbiol. 177, 259–266 (2002).
Ago, R. et al. Relationship between the rod complex and peptidoglycan structure in Escherichia coli. Microbiologyopen 12, e1385 (2023).
Rohs, P. D. A. & Bernhardt, T. G. Growth and division of the peptidoglycan matrix. Annu. Rev. Microbiol. 75, 315–336 (2021).
Barrows, J. M. & Goley, E. D. Ftsz dynamics in bacterial division: what, how, and why? Curr. Opin. Cell. Biol. 68, 163–172 (2021).
Absalon, C. et al. The Gtpase Cpga is implicated in the deposition of the peptidoglycan sacculus in Bacillus subtilis. J. Bacteriol. 190, 3786–3790 (2008).
Buschman, M. D., Xing, M. & Field, S. J. The golph3 pathway regulates golgi shape and function and is activated by DNA damage. Front. Neurosci. 9, 362 (2015).
Novella, I. S., Barbes, C. & Sanchez, J. Sporulation of streptomyces antibioticus Ethz 7451 in submerged culture. Can. J. Microbiol. 38, 769–773 (1992).
Fernández-Martinez, L. T. et al. A transposon insertion single-gene knockout library and new ordered cosmid library for the model organism streptomyces coelicolor a3(2). Antonie Van Leeuwenhoek. 99, 515–522 (2011).
Fernández-García, G. et al. The sco2102 protein harbouring a Dnaa Ii protein-interaction domain is essential for the sco2103 methylenetetrahydrofolate reductase positioning at streptomyces sporulating hyphae, enhancing DNA replication during sporulation. Int. J. Mol. Sci. 23, 4984 (2022).
Sanchez, R. et al. Phylemon 2.0: A suite of web-tools for molecular evolution, phylogenetics, phylogenomics and hypotheses testing. Nucleic Acids Res. 39, W470–474 (2011).
Trapnell, C. et al. Differential gene and transcript expression analysis of rna-seq experiments with tophat and cufflinks. Nat. Protoc. 7, 562–578 (2012).
Gonzalez-Quinonez, N. et al. New phibt1 site-specific integrative vectors with neutral phenotype in streptomyces. Appl. Microbiol. Biotechnol. 100, 2797–2808 (2016).
Rutledge, R. G. & Cote, C. Mathematics of quantitative kinetic Pcr and the application of standard curves. Nucleic Acids Res. 31, e93 (2003).
Manteca, A. et al. ,. Mycelium differentiation and antibiotic production in submerged cultures of streptomyces coelicolor Appl Environ Microbiol, 74: 3877–3886 (2008).
Schmidt, U. et al. Cell detection with star-convex polygons 265–273.
Atrih, A. et al. Structural analysis of Bacillus subtilis 168 endospore peptidoglycan and its role during differentiation. J. Bacteriol. 178, 6173–6183 (1996).
Arbeloa, A. et al. Synthesis of mosaic peptidoglycan cross-bridges by hybrid peptidoglycan assembly pathways in gram-positive bacteria. J. Biol. Chem. 279, 41546–41556 (2004).
Pang, Z. et al., Metaboanalystr 3.0: Toward an optimized workflow for global metabolomics. Metabolites, 10: (2020).
Acknowledgements
We thank the “Ministerio de Ciencia, Innovación Universidades/Agencia Estatal de Investigacion/Fondo Europeo de Desarrollo Regional” (PID2021-122911OB-I00, PID2024-156811OB-I00) and the “Consejería de Empleo, Industria y Turismo del Principado de Asturias” (SV-PA-21-AYUD/2021/51399 and IDE/2024/000742). Sergio Alonso was funded by a “Severo Ochoa” predoctoral grant (grant no. PA-20-PFBP19-006) from “Consejería de Ciencia, Innovación y Universidad del Principado de Asturias”. Paula Díez was funded by a “Ramón y Cajal” contract (RYC2022-035568-I)” from the Spanish Ministry of Science and Innovation. We thank Angel Martinez Nistal and Marta Alonso Guervos (Servicios Científico-Técnicos, Universidad de Oviedo) for their support with confocal microscopy and Pablo Manrique (Servicios Científico-Técnicos, Universidad de Oviedo) for his support with LC-HRMS.
Funding
This work was funded by the “Ministerio de Ciencia, Innovación Universidades/Agencia Estatal de Investigacion/Fondo Europeo de Desarrollo Regional” (PID2021-122911OB-I00, PID2024-156811OB-I00) and by the “Consejería de Empleo, Industria y Turismo del Principado de Asturias” (SV-PA-21-AYUD/2021/51399 and IDE/2024/000742). Sergio Alonso was funded by a “Severo Ochoa” predoctoral grant (PA-20-PFBP19-006) from “Consejería de Ciencia, Innovación y Universidad del Principado de Asturias”. Paula Díez was funded by a “Ramón y Cajal” contract (RYC2022-035568-I)” from the Spanish Ministry of Science and Innovation.
Author information
Authors and Affiliations
Contributions
S.A.F., G.F.G, M.T.F.D.C.G, E.H.H., D.G.G. and I.G.D.R. performed the main experiments. A.M., S.A.F., G.F.G., M.M.B., P.D. D.G.G., I.G.D.R., and F.L. designed experiments. A.M., M.M.B, P.D. and F.L. supervised the study. All authors wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary Material 7
Supplementary Material 8
Supplementary Material 9
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Alonso-Fernández, S., Gutiérrez-Del-Río, I., Lombó, F. et al. Peptidoglycan-reshuffling proteins SCO0954, SCO1758, SCO4439, and SCO4440 modulate the formation of wall-deficient cells in Streptomyces coelicolor under hyperosmotic sucrose stress. Sci Rep 15, 32112 (2025). https://doi.org/10.1038/s41598-025-15457-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-15457-z