Abstract
In addition to identifying alterations suitable for molecularly targeted therapy (TT), comprehensive genomic profiling (CGP) may convey prognostic/predictive information in advanced non-small-cell lung cancer (aNSCLC). We investigated the impact of tumor suppressor gene alterations (TSGAs) on aNSCLC patients’ outcome in a real-world setting. The IMMINENT clinical-genomic database, which provides data on molecular and clinico-pathological features, treatment history, and patients’ outcomes of aNSCLC patients who underwent CGP using tissue- or blood-based next-generation sequencing (NGS) between May 2019 and November 2022 was interrogated to assess overall (OS) and progression-free survival (PFS). The Cox proportional hazard model was used to assess the prognostic and predictive role of genomic features. Two hundred and one patients were included in the analysis. Genomic alterations in STK11, KEAP1, and MYC were associated with poorer OS, while ALK alterations were associated with better OS; after adjustment, STK11 and KEAP1 alterations retained their independent negative prognostic impact. Moreover, KRAS:STK11 and KRAS:KEAP1 co-mutations cooperated to negatively influence OS. In terms of predictive performance, CDKN2A/B alterations predicted worse PFS on chemotherapy (CT) and TP53 alterations predicted worse PFS on TT, while both CDKN2A/B and TP53, as well as KRAS, alterations predicted better PFS on immunotherapy (IO). Extended CGP, encompassing TSGAs, provides valuable prognostic and predictive insights for aNSCLC, thereby adding to the mere identification of potential targets for TT. Our findings also underline the complexity of the lung cancer molecular landscape and suggest the need for studying a personalized therapeutic approach in patients with TSGAs.
Similar content being viewed by others
Introduction
Lung cancer ranks as the third cancer worldwide in terms of incidence and is the leading cause of cancer-related mortality 1. Most lung cancer cases are histologically identified as non-small-cell lung cancer (NSCLC), with around half of the patients presenting with metastatic disease at diagnosis1.
Molecular profiling in lung cancer has now become a stable reality in the daily clinical practice of treating patients with advanced NSCLC (aNSCLC). The introduction and integration of next-generation sequencing (NGS) into clinical practice have revolutionized the approach to molecular profiling. NGS technology enables the detection of a broad spectrum of genetic alterations (GAs) in a single test approach, which significantly enhances the precision of diagnosis and treatment planning. This method not only provides comprehensive results covering all major druggable alterations but also ensures timely outcomes while preserving the limited biopsy material. This is particularly crucial for patients with advanced disease, where tissue availability can be a limiting factor. Additionally, non-invasive testing, well known as a liquid biopsy, that utilizes, for example, blood as the biological source of either cell-free DNA (cfDNA) or circulating tumor DNA (ctDNA) for genomic profiling, is becoming more and more employed as a sequential strategy when tissue analysis is inappropriate or not feasible, or as a complementary approach to tissue-based profiling, with a liquid first approach not so far away on the horizon2.
International guidelines now recommend that all patients with advanced NSCLC undergo molecular profiling with an NGS panel that includes at least the following genes: EGFR, ALK, ROS1, HER2, KRAS, NTRK, BRAF, RET, and MET3. These genes have well-established roles in the pathogenesis of lung cancer and are targets for specific therapies that can be administered as first- or subsequent lines of treatment3,4.
In addition to these well-characterized GAs, there are several other genes whose roles in lung cancer are less well-defined, such as tumor suppressor gene alterations (TSGAs). Among TSGAs, alterations in STK11 and KEAP1 genes, promoting an immunosuppressive tumor microenvironment, have been associated with a lack of responsiveness to PD-(L)1 inhibitors5.
Our real-world study aims to shed light on the potential prognostic and predictive value of TSGAs analyzed with a comprehensive genomic profiling (CGP) approach. Understanding the impact of these alterations could provide new insights into the biology of lung cancer and help further shape the therapeutical pathway to be explored in prospective studies. Moreover, we aim to determine whether an extended profiling approach, which includes both well-known and less-characterized GAs, could lead to better clinical outcomes through the uncovering of additional therapeutic options to enable a more tailored treatment strategy for each patient.
Methods
Study design
This observational retrospective multicenter study used the IMMINENT clinico-genomic database6, which includes anonymized patient-level data of aNSCLC patients who underwent CGP between May 2019 and November 2022. Clinical data comprised clinico-pathological features, treatment history, and patients’ outcomes, while genomic data were mostly derived from NGS tests from Foundation Medicine, Inc. (FMI) performed on tissue and blood samples as part of routine care. Tumor tissue specimens were analyzed using FoundationOne®CDx (F1CDx)7. Blood samples were analyzed by FoundationOne®Liquid (F1L) assay and its successor FoundationOne®LiquidCDx (F1LCDx), launched in August 20208,9. The primary endpoints were overall survival (OS) and progression-free survival (PFS), which were defined as the time (in months) since the initiation of first-line (L1) therapy to death from any cause (OS), disease progression (PFS), or last follow-up (within February 2024). All the data were collected and stored by the University Hospital Trust in Verona in accordance with the Declaration of Helsinki. Patients provided written informed consent for secondary data use.
Patients
The study population included adult patients who were histologically diagnosed with aNSCLC, had successful tissue- or blood-based CGP testing, and received L1 systemic therapy. L1 treatment data were aggregated into four broad treatment categories: chemotherapy (CT), targeted therapy (TT), immunotherapy (IO), and chemo-immunotherapy (CT-IO). Exclusion criteria were lack of medical records or information regarding treatments and their outcomes, and unwillingness or inability to sign the written informed consent form.
Assay methods
FMI’s NGS tests have been described previously6. Briefly, F1CDx uses targeted high throughput hybridization-based capture technology to detect GAs (substitutions, insertion and deletion alterations, copy number alterations, and select gene rearrangements) in 324 genes, as well as genomic signatures including tumor mutational burden (TMB) and microsatellite instability (MSI), using DNA isolated from formalin-fixed, paraffin-embedded (FFPE) tumor tissue specimens10. Similarly, F1L and F1LCDx utilize circulating cfDNA extracted from plasma to detect and report all classes of GAs in 70 and 324 genes, respectively11. F1LCDx also provides blood TMB (bTMB) values, MSI, and circulating tumor DNA tumor fraction.
When available, standard testing results from tissue specimens collected for histological diagnosis were included in the analyses. Specifically, EGFR and KRAS mutations were tested using reverse trascriptase polymerase chain reaction (RT-PCR), while fluorescence in situ hybridization (FISH) was used to detect both ALK and ROS1 rearrangements.
In this study, we only analyzed GAs with a known or likely pathogenic functional status. Patients were labelled as mutant (Mut) for a gene if they carried at least one qualifying GA in that gene according to both their FMI and standard molecular test results. If no GAs were detected in either test, they were classified as wild-type (WT). TMB high (TMB-H) was designated as ≥ 10 mut/Mb for both tissue and liqud samples.
Programmed death-ligand 1 (PD-L1) expression was assessed using immunohistochemical (ICH) analysis, with PD-L1 positive defined as a tumor proportion score (TPS) ≥ 50%.
Statistical analysis
Descriptive statistics were performed using counts and percentages for categorical variables, while means, standard deviations (SD), medians, and interquartile range (IQR) were used to describe continuous variables. Frequencies were compared using the Chi-square test (or Fisher’s exact tests in cases where cell counts were low). Kaplan–Meier estimates were employed to analyze time-to-event data, and survival rates were compared among groups using the log-rank test. Hazard ratios (HRs) and their 95% confidence intervals (CIs) were estimated via univariate and multivariate Cox proportional hazards (PH) models, adjusting for age (< 65 vs. ≥ 65 years), sex, histology, smoking status, Eastern Cooperative Oncology Group performance status (ECOG PS), L1 treatment type, and the test timing. The PH assumption was tested using Schoenfeld residuals; covariates were stratified where necessary to prevent assumption violations. Missingness for confounders was treated as an additional category. Prognostic analyses focused on OS only, whereas predictive analyses considered both OS and PFS. Sensitivity analyses were conducted only in patients who underwent molecular testing before starting the L1 treatment. p-values were not adjusted for multiple testing due to the exploratory nature of the analyses. Statistical significance was set at p < 0.05. Statistical analyses were performed using R, version 4.3.2.
Prognostic effect analysis
A series of univariate and multivariate Cox PH regression models were performed to evaluate the prognostic role of each biomarker (altered genes, TMB, and PD-L1) as well as co-occurring genomic alterations. For co-mutations, a binary variable was defined to discern Mut/Mut patients from those WT/WT, WT/Mut, and Mut/WT. The independent effect of each gene on OS was evaluated by including a gene–gene interaction term in the multivariate model. Analyses were restricted to biomarkers with at least 10 patients in the exposure group and co-occurring genes present in at least 5 patients.
Predictive effect analysis
The predictive role of genomic biomarkers was assessed separately for each L1 treatment type and each genetic variable by including a biomarker-treatment interaction term in the multivariate Cox PH models. We restricted these analyses to genomic features with at least 20 patients in the exposure group and biomarker-drug interactions with at least 5 patients per stratum.
Results
Patient characteristics
Two hundred and forty-six patients with aNSCLC underwent tissue- or blood-based CGP within the previously reported IMMINENT study 6. Of these, 201 (81.7%) patients receiving L1 treatment met eligibility criteria and were included the present prognostic analysis. The baseline characteristics of the study population are presented in Table 1.
Median age at the time of systemic therapy initiation was 64 years (IQR: 56–70) and over a half of the patients were male (53.2%) and had a history of smoking (65.7%); ECOG performance status was 0 or 1 in 94.5% of patients. Adenocarcinoma (LUAD) was the most frequent histological subtype (81.1%). Nearly 74% of CGP tests were conducted before the start of L1 treatment and 66.7% were performed using the 324-gene CGP assays. The most common front-line therapy was CT alone (48.3%), followed by TT (22.9%), IO monotherapy (14.9%), and CT-IO (13.9%). Among TT, osimertinib (26.1%), gefitinib (23.9%), and alectinib (17.4%) were the most frequently used drugs, while pembrolizumab-based therapy was the most prescribed IO/CT-IO regimens (~ 96%) (Supplementary Table A1).
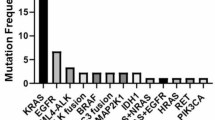
The genomic profile of aNSCLC patients is reported in Supplementary Table A2. Among patients with evaluable TMB (59.7%, n = 120), 24.2% (n = 29) were TMB-H. PD-L1 biomarker testing was conducted for 175 patients (87.1%), 53 (30.3%) having a TPS < 1% (negative), 86 (49.1%) having a TPS of 1–49%, and 36 (20.6%) having a TPS ≥ 50% (Supplementary Table A5).
At a median follow-up time of 15 months (IQR: 5–28), 123 patients had died, and 157 experienced disease progression or death, with PFS data available for 172 (85.6%) patients. Median OS from the start of L1 for the entire population was 23 months (95% CI: 19–27) and was significantly different according to the L1 treatment received: 30 months with TT, 23 months with CT, 23 months with IO, and 14 months with CT-IO. Median PFS to L1 for the entire population was 5.9 months (95% CI: 5.3–8.6), ranging from 8.7 months with TT, 7.5 months with IO, 5.8 months with CT-IO, and 5.3 months with CT (p = 0.038; Supplementary Fig. A1). Approximately 40% of patients received only L1, while approximately 60% received second or later (20%) lines of treatment.
Prognostic effect of biomarkers
Survival by mutation status is reported in Fig. 1. At univariate analysis, STK11, KEAP1, and MYC alterations were associated with worse OS (STK11: median OS 5.3 vs. 26 months, p < 0.001; KEAP1: median OS 3.6 vs. 23 months, p = 0.001; MYC: median OS 3.2 vs. 23 months, p = 0.002), whereas alterations in ALK appeared to be associated with longer survival (median OS: 78 vs. 23 months; p = 0.046). After adjusting the analysis for age, sex, histology, smoking status, ECOG PS and L1 treatment (Fig. 2), STK11 was significantly associated with reduced OS (aHR 2.89, p < 0.001). KEAP1 also reached the statistical significance (aHR 2.09, p = 0.043). Both TMB-H and PD-L1 + had no prognostic value. We also assessed the impact of KRAS G12C mutations on OS. Patients harboring KRAS G12C mutations have similar survival rates to WT patients (Fig. 1).
Kaplan–Meier curves of mutated genes. *For the genes not baited by F1L, the analyses were conducted in patients profiled by F1CDx and F1LCDx (N = 134). ^The analysis was conducted in the TMB-evaluable population (N = 120). †The analysis was conducted in the PD-L1-evaluable population (N = 175). p-values were from log-rank test. Mut, mutant; OS, overall survival; WT, wild-type.
Prognostic associations of genetic biomarkers and OS—univariate and multivariate analyses. ‡KRAS G12C vs. KRAS WT. *For the genes not baited by F1L, the analyses were conducted in the subgroup of patients profiled by F1CDx and F1LCDx (N = 134). ^The analysis was conducted in the TMB-evaluable population (N = 120). †The analysis was conducted in the PD-L1-evaluable population (N = 175). ¥Frequency of the Mut group. Violation of PH assumption for TP53, STK11, RB1, and MYC in univariate analyses, and TP53, MTAP, RB1, and MYC in multivariate analyses. aHR, adjusted HR; cHR, crude HR; CI,confidence interval; HR, hazard ratio; Mut, mutant; TMB-H, tumor mutational burden high; WT, wild-type.
Within the mutated KRAS group, patients with KRAS G12C mutations tended to have a better prognosis compared to the other KRAS mutation variants (median OS: 42 vs. 14 months) (Fig. 2).
Sensitivity analysis results for the 148 patients who received molecular testing at baseline were similar to those obtained in the primary analysis (Supplementary Table A3), with MYC and KEAP1 reaching a statistical significance (p = 0.003 and 0.004, respectively).
The effect of co-mutations on OS is reported in Supplementary Table A4. Among the 20 pairs of co-occurring genes with absolute frequency ≥ 5, four co-mutations were associated with decreased OS (Fig. 3). Patients with alterations in both KRAS and STK11 had significantly worse OS than WT or single-mutant patients (aHR: 3.94, p < 0.001). Similar associations were found for the STK11:TP53 (aHR 2.55, p = 0.008), STK11:KEAP1 (aHR 2.70, p = 0.022), and KRAS:KEAP1 (aHR 2.35, p = 0.042). We then tested the independent effect of genes in each retained pair on survival. Patients with only KRAS or STK11 alterations appeared to have longer and shorter OS as compared to WT patients, respectively, while double-mutant patients had the worst outcomes (p for interaction = 0.027) (Supplementary Fig. A2). Notably, within the context of these co-occurring alterations, neither STK11 nor KEAP1 mutations retained independent statistical significance in terms of OS, suggesting that their prognostic impact is context-dependent and driven by interaction with other concurrent mutations.
Kaplan–Meier curve of OS for the combinations of (A) KRAS and STK11, (B) STK11 and TP53, (C) STK11 and KEAP1*, and (D) KRAS and KEAP1*. *As KEAP1 is not baited by F1L, the analyses were conducted in patients profiled by F1CDx and F1LCDx (N = 134). Mut, mutant; OS, overall survival; WT, wild-type.
Predictive effect of biomarkers
Supplementary Table A5 displays the frequencies of the biomarkers (with n > 20) in each treatment group. Most patients with KRAS and STK11 alterations were initially treated with CT-IO or IO (KRAS: 42.9% CT-IO and 33.3% IO vs. 22.7% CT and 6.5% TT, p = 0.004; STK11: 21.6% CT and 35.7% CT-IO vs. 3.3% IO and 2.2% TT, p < 0.001), whereas those with EGFR alterations most commonly received TT (63.0% TT vs. 9.3% CT, p < 0.001). Clinical outcomes for each treatment category according to biomarker status are shown in Fig. 4 (for PFS) and Supplementary Fig. A3 (for OS), while survival curves by mutation status and treatment arm, in case of significant associations, are shown in Supplementary Fig. A4.
Predictive associations of genetic biomarkers and PFS—multivariate analyses. *The analysis was conducted in the TMB-evaluable population (N = 102). †Frequency in each treatment group. Deviations from PH assumption of exposures were not assessed. aHR, adjusted HR; CI, confidence interval; CT, chemotherapy, CT-IO, chemo-immunotherapy; HR, hazard ratio; int-aHR, interaction-aHR; IO, immunotherapy; Mut, mutant; PFS, progression-free survival; TMB-H, tumor mutational burden high; TT, targeted therapy; WT, wild-type.
Within the subgroup of patients treated with TT, no significant difference in PFS or OS was observed between EGFR-mutant and EGFR WT patients. This result could be related, firstly, to presence of other actionanle molecular alterations (e.g. ALK traslocation) and the consequent TT in the EGFR WT group. Additionally, in our cohort, molecular profiling was often performed in later treatment lines, possibly associated to the relatively short PFS observed in EGFR-mutant. Patients with TP53 alterations showed a significant PFS benefit with IO vs. others (aHR 0.3, p = 0.008), while those without TP53 alterations had a reduced PFS (aHR: 3.57, p = 0.003; interaction p < 0.001) (Supplementary Fig. A4D). Similarly, KRAS and CDKN2A/B predicted better PFS on IO (interaction p = 0.037 and 0.005, respectively) (Supplementary Figu. A4B-C). Conversely, TP53 alterations were significantly associated with worse PFS with TT vs. other treatments (interaction p = 0.027) (Supplementary Fig. A4F). For patients with CDKN2A/B alterations, CT was associated with worse PFS than other therapies (interaction p = 0.007) (Supplementary Fig. A4A). Only TP53 confirmed its predictive value for OS (interaction p = 0.009 for TT and 0.001 for IO) (Supplementary Fig. A4E/G). Sensitivity analyses conducted on patients in whom CGP was performed before the start of L1 (n = 148) yielded results consistent with those obtained in the whole population (Supplementary Table A6). The association of genomic biomarkers and clinical outcomes was assessed further in each treatment group. Results are presented in Supplementary Figs. A5 and A6.
Discussion
In our previous analysis, the genomic landscape of aNSCLC and its correlation with clinical characteristics were reported6. The present study further investigated the prognostic and predictive role of genomic biomarkers, focusing on TSGAs.
When analyzing the prognostic effects of individual genes, according to existing data12, we found that some TSGAs, such as STK11 and KEAP1, had a negative prognostic role. Alterations in STK11 and KEAP1 were strongly associated with poorer OS, in agreement with prior evidence indicating their role in promoting an immunosuppressive tumor microenvironment13. STK11-mutant or KEAP1-mutant tumors exhibit resistance to IO, particularly when co-mutated with KEAP1, which enhances tumor adaptability to oxidative stress and therapy resistance13. This suggests that patients harboring these mutations might benefit from alternative immunotherapeutic strategies, including combination regimens with CTLA-4 blockade, currently under investigation12,14.
Regarding the prognostic effect of co-mutations, the analysis identified four pairs among the most frequent co-occurring TSGAs significantly associated with reduced OS: KRAS:STK11, STK11:TP53, STK11:KEAP1, and KRAS:KEAP1 (Fig. 4). About KRAS:STK11, data from the literature are variable, showing similar15 or poorer OS16 concerning all NSCLC or other subsets. Papillon-Cavanagh and co-authors showed that the effects of KRAS and STK11 alterations are independent13. Regarding STK11:KEAP1, a search in a wide clinical-genomic database confirmed the shorter OS for this association15. The co-mutation of KEAP1 and STK11 has been associated with poor response also to PD-(L)1 inhibitors when administered as a single agent17 or in association with chemotherapy18. Together, they may account for half or more of the primary resistance to PD-(L)1 inhibitors in patients with NSCLC. In this regard, Skoulidis et al. demonstrated that in patients harboring this co-mutation, the addition of CTLA-4 blockade (tremelimumab) to chemotherapy combined with a PD-L1 inhibitor (durvalumab) could lead to a significant clinical benefit, thereby overcoming the intrinsic resistance conferred by this genetic pattern12. While STK11 and KEAP1 alterations were not independently associated with OS in the present analysis, both were linked to shorter OS in univariate and adjusted analyses upon co-occurrence; reliability of this finding, however, is limited by the fact that the F1L test cannot detect KEAP1 alterations, thus making the reduced sample size not sufficient to detect statistically significant differences in OS. Concerning KRAS:KEAP1, in a study analyzing 330 patients with aNSCLC harboring a KRAS mutation, the co-occurrence of KEAP1 or NFE2L2 alterations (both encoding for a protein that acts in the same pathway) was associated with reduced OS for the entire KRAS-mutant patient population19.
Interestingly, TP53 alterations were associated with longer PFS/OS in patients treated with IO, as compared with other treatments. Regarding its predictive role in the setting of IO treatment, a Korean study on 100 patients with aNSCLC treated with IO found that, among female patients, TP53 alterations were associated with longer PFS, as compared to TP53-wt, possibly due to higher PD-L1 expression20. Indeed, a wide analysis showed increased PD-L1 mRNA expression in the TP53 alteration subgroup, as compared to other subgroups; it was also demonstrated that TP53 alteration itself boosted PD-L1 expression20. Quite recently, another study demonstrated that patients with TP53 alterations showed improved PFS upon IO, again potentially due to an association with increased PD-L1 expression and tumor immunogenicity21. Our data also showed shorter PFS/OS in patients treated with TT, as compared to other treatments. In that respect, two recent meta-analyses, including patients with EGFR- or ALK-mutated aNSCLC treated with TT, showed that the co-occurrence of TP53 alteration was associated with significantly shorter PFS and OS 22,23.
Our findings suggest that CDKN2A/B alterations may serve as predictive biomarkers of immunotherapy efficacy, at least in terms of PFS—an association that is not well established in the current literature. While previous studies have reported inconsistent results regarding the predictive role of CDKN2A/B alterations 24, our data could add a novel perspective to this debate. One possible explanation lies in the heterogeneity of these alterations (single nucleotide variants vs larger structural variations). Notably, structural alterations affecting the CDKN2A/B locus have been linked to co-deletion of type I interferon genes in lung and other thoracic malignancies, a mechanism that could influence immune responsiveness 25,26. This study has several limitations. First, at a difference with the current therapeutic landscape of advanced NSCLC, CT was the most common first-line therapy (48.3%) at the time when patients included in the present analysis were profiled. Second, at the time of patients’ enrollment, certain driver alterations (e.g. BRAF mutations, KRAS G12C mutations, etc.) did not yet have specific TT available in clinical practice. Third, the study cohort was limited to patients who received CGP, which may have introduced a selection bias, i.e., left truncation bias. In this case, patients with longer survival are more likely to undergo NGS testing and be included in the study, potentially leading to an OS overestimation when compared to the general population27. Due to the unavailability of the exact CGP testing date, proper adjustment for left truncation bias was not feasible28. Nevertheless, landmark analysis, which includes only patients profiled before treatment initiation, supported the primary conclusion. Furthermore, the analysis was constrained by the small sample sizes in some subgroups, which resulted in wide confidence intervals and reduced the precision of the survival estimates. Lastly, p-values were not adjusted for multiple testing because of the exploratory nature of the analyses, which should be considered when interpreting the results.
Nevertheless, results of the IMMINENT study further underscore the growing importance of CGP in correctly classifying NSCLC from a molecular prognostic and predictive standpoint. While conventional biomarker testing aims at detecting oncogenic drivers with therapeutic implications, an extended CGP approach incorporating TSGAs provides a more refined stratification of patients’ prognosis and treatment response29. This is well exemplified by the recently reported results of the EPROPA experience30; indeed, while only 5% of tested patients benefitted from CGP-identified clinical trial options, an additional 20% obtained relevant information of co-occurring alterations, 25% of EGFR-mutant patients obtained relevant information about potentially targetable genomic mechansims of acquired resistance to osimertinib, and 3% obtained relevant information about potential germline alterations potentially associated with hereditary cancer syndromes. Overall, large-scale real-world analyses confirmed that patients undergoing CGP-guided therapy may experience improved survival outcomes compared to those receiving standard-of-care treatment31. Additionally, timely CGP implementation has been associated with better utilization of targeted therapies, prolonged treatment duration, and reduced reliance on non-targeted chemotherapy12. This is particularly true for EGFR-mutant NSCLC, emphasizing the need for CGP even in oncogene-driven tumors32 and supporting the integration of TSGA assessment into routine CGP to identify patients unlikely to benefit from standard TT and facilitate a shift toward precision-based combinatorial strategies. Future studies to validate these findings and potentially inform clinical practice will require larger databases and prospective patient enrollment and assessment; for these reasons, our data will feed the ATLAS interactive database platform (https://biomarkersatlas.com/)33,34, which contains data coming from most Italian centers on gene mutations in lung cancer, enabling further analysis using artificial intelligence and comparisons with public datasets.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Zhou, J. et al. Global burden of lung cancer in 2022 and projections to 2050: Incidence and mortality estimates from GLOBOCAN. Cancer Epidemiol. 93, 102693 (2024).
Rolfo, C. et al. Liquid biopsy for advanced NSCLC: A consensus statement from the international association for the study of lung cancer. J. Thorac. Oncol. 16(10), 1647–1662 (2021).
Mosele, M. F. et al. Recommendations for the use of next-generation sequencing (NGS) for patients with advanced cancer in 2024: A report from the ESMO precision medicine working group. Ann. Oncol. 35(7), 588–606 (2024).
Hendriks, L. E. et al. Oncogene-addicted metastatic non-small-cell lung cancer: ESMO clinical practice guideline for diagnosis, treatment and follow-up. Ann. Oncol. 34(4), 339–357 (2023).
Skoulidis, F. et al. Co-occurring genomic alterations define major subsets of KRAS-mutant lung adenocarcinoma with distinct biology, immune profiles, and therapeutic vulnerabilities. Cancer Discov. 5(8), 860–877 (2015).
Sposito, M. et al. Tissue- and liquid-biopsy based NGS profiling in advanced non-small-cell lung cancer in a real-world setting: the IMMINENT study. Front Oncol. 14, 1436588 (2024).
Milbury, C. A. et al. Clinical and analytical validation of FoundationOne®CDx, a comprehensive genomic profiling assay for solid tumors. PLoS ONE 17(3), e0264138 (2022).
Woodhouse, R. et al. Clinical and analytical validation of FoundationOne Liquid CDx, a novel 324-Gene cfDNA-based comprehensive genomic profiling assay for cancers of solid tumor origin. PLoS ONE 15(9), e0237802 (2020).
Clark, T. A. et al. Analytical validation of a hybrid capture-based next-generation sequencing clinical assay for genomic profiling of cell-free circulating tumor DNA. J Mol Diagn. 20(5), 686–702 (2018).
SUMMARY OF SAFETY AND EFFECTIVENESS DATA (SSED). FoundationOne®CDx., (2024). https://www.accessdata.fda.gov/cdrh_docs/pdf17/P170019S054B.pdf February 1, 2024.
SUMMARY OF SAFETY AND EFFECTIVENESS DATA (SSED). FoundationOne®Liquid CDx., (2024). https://www.foundationmedicine.com/sites/default/files/media/documents/2025-04/v16_RAL-0035_P190032_S024_F1LCDx_Technical_Clean%20Label_10DEC2024.pdf February 1, 2024.
Skoulidis, F. et al. CTLA4 blockade abrogates KEAP1/STK11-related resistance to PD-(L)1 inhibitors. Nature 635(8038), 462–471 (2024).
Papillon-Cavanagh, S. et al. STK11 and KEAP1 mutations as prognostic biomarkers in an observational real-world lung adenocarcinoma cohort. ESMO Open. 5(2), e000706 (2020).
Skoulidis, F. et al. TRITON: Phase 3b study of tremelimumab (T)+ durvalumab (D) vs pembrolizumab (P), in combination with chemotherapy (CT), in non-squamous (NSQ) metastatic NSCLC (mNSCLC) with STK11 and/or KEAP1 and/or KRAS mutations. J. Clin. Oncol. 42(16), TPS8655–TPS9655 (2024).
Julian, C. et al. Overall survival in patients with advanced non-small cell lung cancer with KRAS G12C mutation with or without STK11 and/or KEAP1 mutations in a real-world setting. BMC Cancer 23(1), 352 (2023).
Shire, N. J. et al. STK11 (LKB1) mutations in metastatic NSCLC: Prognostic value in the real world. PLoS ONE 15(9), e0238358 (2020).
Ricciuti, B. et al. Diminished efficacy of programmed death-(Ligand)1 inhibition in STK11- and KEAP1-mutant lung adenocarcinoma is affected by KRAS mutation status. J. Thorac. Oncol. 17(3), 399–410 (2022).
Alessi, J. V. et al. Clinicopathologic and genomic factors impacting efficacy of first-line chemoimmunotherapy in advanced NSCLC. J Thorac. Oncol. 18(6), 731–743 (2023).
Arbour, K. C. et al. Effects of co-occurring genomic alterations on outcomes in patients with KRAS-mutant non-small cell lung cancer. Clin. Cancer Res. 24(2), 334–340 (2018).
Choi, S. et al. Association of TP53 mutation status and sex with clinical outcome in non-small cell lung cancer treated with immune checkpoint inhibitors: A retrospective cohort study. Cancer Res. Treat. 57(1), 70–82 (2025).
Feng, J. et al. Prognostic and predictive effects of TP53 co-mutation in patients with non-small cell lung cancer with rare treatable driver mutations. Lung Cancer 204, 108452 (2025).
Qin, K. et al. Prognostic value of TP53 concurrent mutations for EGFR- TKIs and ALK-TKIs based targeted therapy in advanced non-small cell lung cancer: A meta-analysis. BMC Cancer 20(1), 328 (2020).
Ferrara, M. G. et al. Meta-analysis of the prognostic impact of TP53 co-mutations in EGFR-mutant advanced non-small-cell lung cancer treated with tyrosine kinase inhibitors. Crit. Rev. Oncol. Hematol. 184, 103929 (2023).
Provencio-Pulla, M. et al. Identification of non-actionable mutations with prognostic and predictive value in patients with advanced or metastatic non-small cell lung cancer. Clin. Transl. Oncol. 26(6), 1384–1394 (2024).
Grard, M. et al. Homozygous co-deletion of type I interferons and CDKN2A genes in thoracic cancers: Potential consequences for therapy. Front Oncol. 11, 695770 (2021).
Peng, Y. et al. Co-occurrence of CDKN2A/B and IFN-I homozygous deletions correlates with an immunosuppressive phenotype and poor prognosis in lung adenocarcinoma. Mol. Oncol. 16(8), 1746–1760 (2022).
Franchi, M., Cortinovis, D. & Corrao, G. Treatment patterns, clinical outcomes and healthcare costs of advanced non-small cell lung cancer: A real-world evaluation in Italy. Cancers (basel). 13(15), 3809 (2021).
Backenroth, D. et al. Accounting for delayed entry in analyses of overall survival in clinico-genomic databases. Cancer Epidemiol. Biomarkers Prev. 31(6), 1195–1201 (2022).
Le, X. Partners in crime: Co-occurring genetic alterations in EGFR-mutant NSCLC. J. Thorac. Oncol. 19(2), 190–192 (2024).
Passiglia, F. et al. Actionable NSCLC mutation identification by comprehensive genomic profiling for clinical trial enrollment: The European program for the routine testing of patients with advanced lung cancer (EPROPA). J. Thorac. Oncol. 16, 614–624 (2024).
Stockhammer, P. et al. Co-occurring alterations in multiple tumor suppressor genes are associated with worse outcomes in patients with EGFR-mutant lung cancer. J. Thorac. Oncol. 19(2), 240–251 (2024).
Skoulidis, F. et al. P4.11D.01 TRITON: Tremelimumab + Durvalumab + Chemotherapy (CT) vs Pembrolizumab + CT in mNSCLC with STK11, KEAP1 and/or KRAS mutations. J. Thorac. Oncol. 19(10), S389 (2024).
Novello S, Troncone U, Malapelle U. ATLAS, Biomarkers atlas https://biomarkersatlas.com. September 13, 2024.
Malapelle, U. et al. The biomarkers ATLAS: An audit on 1100 non-small cell lung cancer from an Italian knowledge-based database. Lung Cancer 191, 107787 (2024).
Acknowledgements
We acknowledge the participating centers and the relevant healthcare personnel involved in this research: Policlinico Universitario A.Gemelli (Roma), Azienda Ospedaliera San Giovanni Addolorata (Roma), Policlinico Umberto I (Roma); Azienda Ospedaliera Ospedali Riuniti Villa Sofia Cervello (Palermo); Ospedale San Vincenzo Taormina (Taormina); Azienda Ospedaliero Universitaria Policlinico “G.Rodolico—San Marco” (Catania); Azienda Ospedaliera Universitaria Integrata Verona (Verona); Ospedale P. Pederzoli (Peschiera); Ospedale San Bortolo di Vicenza (Vicenza); Ospedale Santa Chiara di Trento (Trento); Agenzia Ospedale Università (Padova); Azienda Ospedaliero Universitaria San Giovanni di Dio Ruggi d’Aragona (Palermo). LB is supported by grant from Associazione Pietro Casagrande ONLUS. L.B. and M.M. are supported by Piano Nazionale di Ripresa e Resilienza (PNRR) projects HEAL Italia, INNOVA and by Research Projects of National Relevance (PRIN) TELUAD. SP is supported by the Associazione Italiana per la Ricerca sul Cancro (AIRC, Next Gen Clinician Scientist 2023 n° 30204). We also acknowledge AdRes Health Economics & Outcomes Research, that provided statistical analyses, and SEEd Medical Publishers, that provided publishing support and journal styling services, both funded by Roche SpA.
Funding
This work was supported by Roche S.p.A. The funder had no role in the data collection, study design, analysis, interpretation of data, and writing of the report and had no access to patient-level data. The corresponding author had full access to all the data in the study and had final responsibility for the decision to submit it for publication.
Author information
Authors and Affiliations
Contributions
M.S: Conceptualization, Data curation, Investigation, Methodology, Resources, Validation, Writing—review and editing. L.B: Conceptualization, Data curation, Investigation, Resources, Supervision, Validation, Writing—review and editing. R.N.: Conceptualization, Data curation. J.I.: Data curation. I.M.S.: Data curation. J.M.: Data curation. M.S.: Data curation. A.L.: Data curation. F.B.: Data curation. F.V.: Data curation. F.S.: Data curation. G.A.: Data curation. L.C.: Data curation. M.O.: Data curation. D.M.: Data curation. A.V.: Data curation. F.L.: Data curation. H.J.S.P.: Data curation. F.F.: Data curation. C.S.: Data curation. C.P.: Data curation, Formal analysis, Methodology, Software, Validation, Visualization, Writing—original draft preparation. L.P.: Supervision, Writing—review and editing. E.S.: Conceptualization, Funding acquisition, Project administration, Supervision, Writing—review and editing. S.C.: Conceptualization, Supervision, Writing—review and editing. U.M.: Supervision. F.P.: Writing—review and editing. E.B.: Data curation, Supervision. S.P.: Data curation, Supervision. M.M.: Conceptualization, Data curation, Funding acquisition, Investigation, Methodology, Resources, Supervision, Writing—review and editing. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The study was conducted in accordance with the Declaration of Helsinki, and approved by the Institutional Ethics Committee of clinical research of AOUI Verona (protocol code 59115 of 11/10/2021).
Informed consent
Informed consent was obtained from all subjects involved in the study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sposito, M., Belluomini, L., Nocini, R. et al. Prognostic and predictive implications of tumor suppressor gene alterations in non-small cell lung cancer. Sci Rep 15, 34807 (2025). https://doi.org/10.1038/s41598-025-18662-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-18662-y