Abstract
The Foot-and-mouth disease virus (FMDV) is highly contagious, affecting cloven-hoofed animals and causing significant economic losses to the livestock industry. Its high genetic variability complicates vaccine development, necessitating frequent updates to traditional inactivated vaccines. Multi-epitope-based vaccines (MEBVs) offer a promising alternative by incorporating immunogenic epitopes to elicit robust humoral and cell-mediated immune responses, thereby providing broader protection against diverse FMDV strains. Using in silico approaches, we designed a novel FMDV-targeting MBEV specifically tailored to the BoLA alleles to ensure optimal epitope selection for cattle. The MEBV included non-toxic, antigenic, and non-allergenic epitopes combined with β-defensin 3 adjuvants, PADRE sequences, and linkers to enhance immunogenicity and stability. Molecular docking and dynamics simulations confirmed strong binding affinity and stable interactions with TLR3 and TLR7, and immune simulation analysis revealed notable increases in B-cell and T-cell populations, cytokines, and macrophages, with sustained levels of IFN-γ and IL-2. Codon optimization for Escherichia coli K12 enhanced expression, and in silico restriction cloning demonstrated its scalability for production. Our findings suggest that the MEBV design is a cost-effective and efficient approach to combat FMDV. While experimental validation is required, the in silico results also highlight its potential to trigger protective immune responses and address the limitations of traditional vaccines.
Similar content being viewed by others
Introduction
Foot-and-mouth disease (FMD) outbreaks have occurred in various parts of the world and significantly affected the livestock industry by infecting cloven-hoofed animals, such as cows, pigs, goats, and sheep1. These outbreaks had a major impact on global livestock production and the economy due to trade restrictions imposed on countries affected by this disease2. Despite the low mortality rate associated with FMD, countries wherein FMD infections are reported cannot participate in the international trade of animals and animal products due to the highly infectious nature of the disease3. Worldwide, FMD is estimated to cause economic losses of US$6.5–21 billion annually in endemic countries; costs in FMD-free countries exceed US$1.5 billion annually2,4. FMD is caused by the FMD virus (FMDV), a single-stranded positive RNA virus belonging to the Picornaviridae family5. FMDV has seven serotypes (O, A, C, Asia-1, SAT-1, SAT-2, and SAT-3), each of which has given rise to numerous variations and subtypes6. Among the seven serotypes, serotype O is prevalent in most regions with FMD outbreaks, including Asia, India, the Middle East, South America, and Africa7. Moreover, FMDV has an extremely high mutation rate because the virus-encoded RNA polymerase lacks a proofreading mechanism8,9. Hence, controlling and eradicating FMD has become challenging due to its numerous viral strains, highly contagious nature, and the high mutation rate of FMDV. After FMDV infection occurs, cattle and reared swine typically exhibit severe symptoms that appear 24–48 h later. At this stage, the serum of infected animals shows elevated viral counts, and the symptoms of FMD include fever, drooling with saliva, shivering, lameness, and the formation of blisters or vesicles on the nose, tongue, coronary bands, and teat epithelium10. These symptoms can vary depending on the age and species of the infected animal, as well as the strain and serotype of the virus.
Vaccination plays a crucial role in maintaining and controlling most FMD outbreaks worldwide. Different countries have adopted various vaccination approaches based on the epidemiological situation and prevailing strains. However, the efficacy of these vaccines is limited by their short-lived immunity and considerable antigenic variability11,12. As the development of the FMD vaccine began more than 50 years ago, it is inevitable for virologists and field veterinarians to have asserted inadequate vaccination success stories. This lack of success might be attributable to the persistent genetic evolution of the virus12. The effectiveness of FMDV vaccinations depends on identifying the prevalent strain, matching vaccine strains to field strains, and ensuring that the vaccine is effective in generating a long-lasting immune response13. Moreover, the particulars of regional vaccination campaigns, execution of regular vaccine matching studies, types of adjuvants employed, and volume of animals vaccinated during campaigns all affect the efficacy of vaccination campaigns4,11. Commercially available vaccines are typically prepared from the supernatants of FMDV-infected bovine hamster kidney 21 cell cultures. These supernatants are then partially purified, chemically inactivated, and combined with an adjuvant. However, the vaccine could potentially not be completely inactivated14. Thus, scientists have worked on developing novel vaccines using recombinant proteins, empty capsids, and genetically engineered subunits to address these issues15,16,17,18.
Advancements in bioinformatics and genomic science have shifted the traditional methods used for developing vaccines into modern vaccine design in an effort to improve and minimize risks. Recent developments in immunoinformatics have given rise to a new field dedicated to the design of multi-epitope-based vaccines (MEBV)19,20. The development of vaccines has been greatly accelerated in this discipline, allowing us to better understand host immune responses21. Hence, the current study aimed to create an MEBV by examining all structural proteins (VP1–4) from the FMDV. Upon entry into the host cell, the long open reading frame of the viral RNA primarily co-translates into four individual products (Lpro, P1, P2, and P3), of which P1 is further processed to form the four structural proteins, VP1, VP2, VP3, and VP4. VP1–3 collectively build the capsid surface of the virus, and VP4 forms the internal structure of the viral particle. Moreover, the viral capsid is composed of 60 capsomeres, each of which is formed by a single unit of VP1–422,23. By targeting these structural proteins, the effectiveness of the vaccine can be increased, and a wide range of FMDV strains can be targeted. Thus, we used a variety of immunoinformatics tools to screen all structural proteins to predict capable Sip epitopes (B-cell, cytotoxic T-lymphocyte [CTL], and helper T-lymphocyte [HTL] epitopes) and then constructed the final MEBV by joining these epitopes with specific linkers and an adjuvant. Different physicochemical and immunological characteristics were comprehensively assessed, and the three-dimensional structure of the MEBV modeled and refined. Molecular docking and molecular dynamics (MD) simulations were performed to better understand the binding interactions of the MEBV with two host immune Toll-like receptors (TLRs), TLR3 and TLR7. Further, the effectiveness of the designed MEBV in inducing an immune response in the host was assessed using immunoinformatics techniques.
Results
Retrieval and evaluation of structural proteins
In the MEBV design, targeting structural proteins, particularly capsid proteins, is crucial because of their immunogenicity, conservation, role in viral infection, and immune recognition. Owing to these features, structural proteins are essential for developing vaccines against viruses, such as FMDV, and ensuring effective and broad immunity induction. All structural proteins, including the VP1, VP2, VP3, and VP4 sequences, were retrieved from the UniProt database, and their antigenic propensities predicted using the Antigenic peptide tool. The average antigenic propensity scores for VP1, VP2, VP3, and VP4 were predicted as 1.0336, 1.0349, 1.0348, and 0.9720, respectively (Figure S1). Next, the antigenicity of all four structural proteins was evaluated using the VaxiJen v.2.0 and ANTIGENpro tools. Both tools predicted that the four structural proteins are antigenic (Table S1). Therefore, all four capsid proteins were considered for epitope prediction.
Prediction of B-cell epitopes
B-cell epitopes are specific regions of antigens recognized by B-cell receptors or antibodies that then lead to the initiation of humoral immunity. Incorporating B-cell epitopes into the MEBV would enhance its ability to induce neutralizing antibodies that can block the entry of pathogens or neutralize their effects. When combined with T-cell epitopes, these vaccines could offer comprehensive protection against a wide range of pathogens, including FMDV. Computational tools have streamlined the identification and validation of these epitopes, ensuring that the designed vaccine is effective and specific. Incorporating B-cell epitopes into the MEBV is crucial for inducing a robust humoral response. Hence, all four FMDV protein sequences (VP1–4) were subjected to the ABCpred server to identify B-cell epitopes. Four epitopes from each protein with a 16-mer window were selected with the highest prediction scores (Table S2). To construct the MEBV, these 16-mer epitopes were selected as potential B-cell epitopes.
Prediction of cytotoxic and helper T-lymphocyte epitopes
Major histocompatibility class I (MHC-I)-specific CTL epitopes were predicted using the NetMHCpan-4.1 server. For each protein, 9-mer epitopes were selected based on their high binding affinity for different BoLA class-I alleles. Epitopes labeled as “strong interactions” with high prediction scores were considered for designing the MEBV. Similarly, owing to the pivotal role that HTLs play in vaccine design, MHC-II-specific HTL epitopes were predicted using the IEDB-MHCII server. Prediction was performed by selecting 15-mer epitopes and evaluating their binding interactions with various BoLA (Bovine Lymphocyte Antigen allele) class-II alleles, such as BoLA-DRB3*00101, BoLA-DRB3*00202, BoLA-DRB3*01101, BoLA-DRB3*01201, BoLA-DRB3*01501, and BoLA-DRB3*02703. Epitopes common in most of the BoLA alleles with the highest prediction scores were considered for vaccine construction. Detailed information on the selected CTL and HTL epitopes is provided in Table S3 and Table S4, respectively.
Moreover, the epitopes selected for MEBV design were mapped onto three-dimensional structures of the corresponding proteins that had been modeled using AlphaFold (Fig. 1). Constructing a vaccine subunit requires the selection of a nontoxic epitope. Hence, toxicity of the predicted epitopes was evaluated using the ToxinPred2 server (Table S5). The server results showed that all selected epitopes were predicted to be nontoxic; therefore, all were considered for MEBV construction.
Selected B-cell (light pink), cytotoxic T-lymphocyte (CTL; pale green), and helper T-lymphocyte (HTL; light blue) epitopes mapped onto their respective antigenic proteins, including VP1, VP2, VP3, and VP4, which were modeled using AlphaFold.
Designing the final multi-epitope-based vaccine subunit
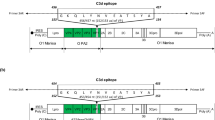
Different linkers were used to join the chosen epitopes that could trigger a cell-mediated immune response to construct the MEBV. The vaccine subunit was constructed by linking four B-cell, four CTL, and four HTL epitopes using KK, AAY, and GPGPG linkers. In addition, β-defensin 3 (an adjuvant) and a PADRE (Pan DR-binding epitope) peptide sequence were added to the N-terminal of the vaccine subunit through EAAAK linkers. Subsequently, a 6-histidine (6-His) tag was joined to the C-terminal of the vaccine with a GPGPG linker (Fig. 2A and B). The final MEBV subunit comprised 289 amino acid residues.
Illustration of the final multi-epitope-based vaccine (MEBV) construct. (A) Schematic representation of the arrangements of the adjuvant; PADRE sequence; B-cell, cytotoxic T-lymphocyte (CTL), and helper T-lymphocyte (HTL) epitopes; 6-His tag; and linkers used to construct the MEBV. (B) Complete sequence of the MEBV. (C) Three-dimensional structure of the final MEBV modeled using AlphaFold. The B-cell, CTL, and HTL epitopes are illustrated in light pink, pale green, and light blue, respectively. The adjuvant and PADRE sequences in the N-terminal and 6-His tag in the C-terminal are illustrated in light orange.
The antigenicity of the proposed vaccine was predicted using the VaxiJen v.2.0 and ANTIGENpro servers, and both tools predicted the proposed vaccine to be antigenic, with scores of 0.4572 and 0.933628, respectively. The AllerTop v.2.1, AllergenFP v.1.1, and AlgaPred 2.0 servers predicted the designed MEBV to be nonallergenic. The solubility of the vaccine subunit was assessed using the Protein-Sol server and found to be soluble, with a prediction score of 0.509 and isoelectric point (pI) of 10.3. Finally, the ToxinPred2 server was used to analyze the toxicity of the entire vaccine. With a prediction score of 0.27, the server predicted that the vaccine was nontoxic (Table S6). Physiochemical characterization using the Expasy ProtParam server revealed that the MEBV had a molecular weight of 31.513 kDa, with a pI score of 9.79. The total number of positive (+) and negative (−) residues were calculated as 10 and 31, respectively, with a total number of 4391 atoms and chemical formula “C1418H2171N399O387S16.” The server predicted that the aliphatic and instability indexes were 61.94 and 30.07, respectively, suggesting that the modeled vaccine was stable upon expression. Furthermore, the grand average hydropathicity (GRAVY) score of the protein sequence was calculated as − 0.433, indicating that the protein was predominantly hydrophilic and may have a higher affinity for aqueous environments, such as the cytoplasm.
Subsequently, the secondary structure of the final vaccine subunit was predicted using the GOR IV and PSIPRED servers. The servers predicted that the designed MEBV comprised 22.15%, 27.34%, and 50.25% α-helix, β-strand, and coil structures, respectively (Figure S2). Finally, the 3D structure of the MEBV was modeled using AlphaFold3 (Fig. 2C), and per-residue pLDDT score was used to assess the confidence in the modeled structure. Moreover, its stereochemical properties were validated using a Ramachandran plot, and the results state that 85.5% of the residues were plotted in the most favored region, 14.0% in the allowed region, and 0.4% in the disallowed region (Figure S3).
Binding interactions of the multi-epitope-based vaccine with Toll-like receptors
MEBVs are often docked with immunogenic receptors, such as TLRs, to ensure that they can effectively activate the immune system, which is important for initiating and shaping adaptive immunity. For FMDV vaccines, targeting specific TLRs, such as TLR3 and TLR7, is crucial due to the unique immune challenges imposed by the virus. TLR3 and TLR7 recognize dsRNA and ssRNA, respectively, which are the genetic material found in most viruses. Hence, a molecular docking study was performed using the HDOCK server to ensure that the epitopes could effectively interact with TLRs. Results revealed that the MEBV binds to TLR3 and TLR7 receptors with a binding affinity of − 348.34 and − 306.66 kcal/mol, respectively. The binding positions of the MEBV with TLRs are illustrated in Fig. 3A and B. Moreover, the residual interactions between the MEBV and TLRs were analyzed using the PDBsum tool and presented in Fig. 3C and D. According to the residual interaction profile, the MEBV formed 12 hydrogen bond (H-bond) interactions, 3 salt bridges, and 268 nonbonded contacts with TLR3. Similarly, it formed 9 H-bond interactions, 2 salt bridges, and 322 nonbonded contacts with TLR7. Although the number of H-bond interactions with TLR7 was lower than that with TLR3, the MEBV formed a more substantial number of nonbonded interactions with the TLR7 receptor (Figure S4). In contrast, the designed MEBV demonstrated strong binding interactions with immunogenic receptors.
Binding interactions of the multi-epitope-based vaccine (MEBV) with (A) TLR3 and (B) TLR7. (C) and (D) represent the residual interaction profile of the MEBV with both receptors.
Structural stability assessment during molecular dynamics simulations
The dynamic behavior of the MEBV with TLRs was studied using 100-ns MD simulations. Structural stability of the TLR3-MEBV and TLR7-MEBV complexes was assessed through root-mean-square standard deviation (RMSD) and radius of gyration (Rg) calculations of the protein backbone atoms. The RMSD calculations suggested that after 70 ns, both complexes maintained their structural stability in the range of 0.57–0.70 nm. The RMSD plot showed that the TLR3-MEBV complex had lower RMSD values than the TLR7-MEBV complex did. Moreover, the probability distribution function (PDF) revealed values of 0.51 ± 0.07 nm and 0.69 ± 0.10 nm for the TLR3-MEBV and TLR7-MEBV complexes, respectively (Fig. 4A and B). Similarly, the TLR3-MEBV complex exhibited a lower Rg than the TLR7-MEBV complex did, with average values of 3.32 ± 0.05 and 3.59 ± 0.04 nm, respectively (Fig. 4C and D). Although the designed MEBV induced minimal structural alterations upon binding to TLR3, it maintained structural stability with both TLRs during the final 40 ns of the MD simulation. Root-mean-square fluctuation (RMSF) analysis indicated no significant changes in the TLR3 and TLR7 receptors upon MEBV-binding throughout the simulation. Residues 436–485 of TLR7 exhibited larger fluctuations because of its unique disordered loop region. However, the designed MEBV alone had a lower RMSF when binding to TLR3 than to TLR7, with average RMSF values of 0.39 and 0.45 nm, respectively (Figure S5).
Structural stability assessment through root-mean-square standard deviation (RMSD) and radius of gyration (Rg) calculations. (A) RMSD of the protein backbone. (B) Probability distribution function (PDF) of the RMSD. (C) Rg of the protein backbone. (D) PDF of the Rg. Green and blue colors represent the TLR3-MEBV and TLR7-MEBV complexes, respectively.
The number of H-bond interactions between the MEBV and TLRs was analyzed using all trajectory information, as illustrated in Fig. 5A and B. The designed MEBV formed an average of 23 H-bonds with TLR3 and 10 H-bonds with TLR7 during the MD simulation. The TLR3 receptor exhibited a stronger binding affinity for the MEBV, with a higher number of H-bond interactions than that with TLR7.
(A) Number of hydrogen bond (H-bond) interactions between the multi-epitope-based vaccine (MEBV) and Toll-like receptors (TLRs) during the molecular dynamics simulation. (B) Probability distribution function (PDF) of the H-bond interactions. Green and blue colors represent the number of H-bond interactions of the MEBV with TLR3 and TLR7, respectively.
Binding free energy and interaction energies between the multi-epitope-based vaccine and toll-like receptors
The binding affinity of the MEBV for TLR3 and TLR7 was studied using the final 50-ns trajectories from the MD simulation through MM-PBSA calculations, as illustrated in Fig. 6A. The designed MEBV exhibited a stronger binding affinity for TLR7 (− 2299.48 ± 189.025 kJ/mol) than for TLR3 (− 1983.33 ± 154.602 kJ/mol). Compared to the other free energy components, the polar solvation energy of the MEBV with TLR7 exhibited a stronger binding affinity, indicating a stronger binding energy associated with the electrostatic interactions between the MEBV-TLR7 complex and the solvent. Detailed information on each free energy term is provided in Table S7. Moreover, we calculated the total nonbonded interaction energy components between the MEBV and TLRs using all MD trajectory information (Fig. 6B). The total interaction energy was calculated as the sum of the short-range Coulombic and Lennard–Jones energies. The average total interaction energy of the MEBV with TLR3 and TLR7 was calculated as − 2312.15 and − 961.35 kJ/mol, respectively, which indicates that the MEBV formed a stronger nonbonded interaction with both TLRs, but particularly with TLR3.
(A) Free energy components calculated from MM-PBSA. VdW, Elec, Polar, SASA, and ΔG indicate the Van der Waals energy, electrostatics energy, polar solvation energy, solvent accessible surface area, and total binding free energy, respectively. (B) The total nonbonded interaction energy was calculated as the summation of Coulomb and Lennard–Jones interaction terms.
Immune simulation
The immunogenic profile of the proposed vaccine was generated using the C-ImmSim server and depicted in Fig. 7. Immune simulation results indicated that secondary and tertiary responses were produced at notably higher rates than those of primary responses. Enhanced responses were observed in the levels of various antibodies, including IgM, IgM + IgG, IgG1 + IgG2, IgG1, and IgG2. The simulation also demonstrated the efficacy of the vaccine, as elevated populations of B-cells and T-cells were observed. Furthermore, an increase in Th1 concentration was noted following each administered dose. The B-cell, T-cell, dendritic cell (D-cell), macrophage, and epithelial cell populations per entity state are shown in Figure S6. Further examination showed strong activation of these cells, notably a swift and prolonged rise in natural killer cell counts, which reached its highest point on the 100th day (approximately 398 cells/mm3) following initial immunization, with subsequent variations of approximately 330–370 cells/mm3. Additionally, the observed increase in macrophage and D-cell populations suggests that antigen-presenting cells effectively processed and presented antigens to CD4 + and CD8 + cells. Furthermore, the elevated levels of various cytokines observed indicated that the designed vaccine elicited favorable immune responses.
Immune simulation responses of the proposed multi-epitope-based vaccine (MEBV) generated from the C-ImmSim server. (A) Immunoglobulin responses to antigen exposure. (B) B lymphocyte population. (C) Plasma B lymphocyte count sub-divided per isotype. (D) CD4 T-helper lymphocyte count showing the total and memory counts. (E) CD8 T-cytotoxic lymphocyte count showing the total and memory counts. (F) Total count of natural killer cells (NK cells).
In silico cloning
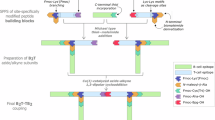
Maximizing vaccine expression is crucial in experimental research, where codon optimization of the amino acid sequence depends on the host expression system. Hence, the proposed vaccine was subjected to codon optimization for cloning into Escherichia coli K12 using an in silico approach (Table S9). The resulting cDNA had a codon adaptation index (CAI) and GC percentage of 1.0 and 50.73%, respectively, suggesting a high likelihood of expression in the vector. The optimized sequence was inverted to guarantee complementation in the vector translation direction. BamHI and XhoI restriction sites were added at the 5′- and 3′-ends, respectively. The sequence was then incorporated into the pET28a(+) vector using SnapGene software. The resulting recombinant plasmid, measuring 6043 bp in length, was designed for successful expression in the E. coli system (Fig. 8).
Proposed multi-epitope-based vaccine (MEBV) cloned into the pET28a(+) vector using XhoI and BamHI restriction sites, as illustrated using SnapGene. The MEBV and vector regions are represented in red and black, respectively.
Discussion
The FMDV has emerged as a highly contagious virus that affects cloven-hoofed animals, such as cattle, pigs, sheep, and goats. This virus belongs to the Picornaviridae family and causes significant economic losses to the livestock industry due to trade restrictions and decreased productivity2. FMD is particularly challenging to control because the FMDV has a high degree of genetic variability and can evolve quickly, making it challenging to develop effective vaccines. The seven viral serotypes, A, O, C, Asia-1, and SAT-1–3, make vaccine development even more challenging. Traditional vaccines, such as inactivated whole-virus vaccines, have been used to control and manage FMD. However, these vaccines require frequent updates owing to viral mutations and can lead to the production of non-neutralizing antibodies, which are not always effective in preventing infections11,12. Moreover, inactivated vaccines have limitations in terms of cost and safety. Therefore, there is an urgent need to develop a more effective and versatile vaccine approach. In this context, MEBVs represent an attractive alternative. MEBVs stimulate humoral and cell-mediated immune responses by incorporating several immunogenic epitopes from various viral regions. This may offer broader protection against various FMDV strains and lower the chances of viral escape due to mutations. Developing an MEBV against FMDV using in silico approaches offers several advantages, including the ability to predict and select the most immunogenic epitopes based on bioinformatics tools, thus reducing the time and costs associated with experimental vaccine development.
Advancements in structural bioinformatics and immunoinformatics have accelerated the development of accurate, safe, and cost-effective vaccines against various pathogens. In silico methods enable the prediction and selection of optimal epitopes by considering various factors, such as immunogenicity, antigenicity, and conservation across viral strains. Many MEBVs have been developed against the Ebola virus, Zika virus, SARS-CoV-2, Nipah virus, and HIV using an immunoinformatics approach24,25,26,27,28. Owing to the promising outcomes of these studies, researchers have opted to employ this approach to address livestock diseases. Cao et al., 2017 designed an MEBV that showed promise in preventing FMDV serotype A infection in pigs, demonstrating the potential of this MEBV immunoinformatics approach for FMDV infection in cattle29. Moreover, Bhutta et al., 2024 designed a novel MEBV by targeting the structural proteins of FMDV in bovines, where they used HLA alleles to screen for CTL and HTL epitopes without explicitly addressing BoLA alleles for the designed vaccines30. In contrast, our in silico analysis designed an MEBV against FMDV by considering the BoLA alleles to screen for epitopes, which yielded substantial findings.
The efficacy of an MEBV depends on the selected epitopes that can elicit both humoral and cell-mediated immune responses. B-cell epitopes are required for the activation of antibody-mediated immunity, whereas T-cell epitopes are required to activate CTLs, which are responsible for the clearance of infected cells31. Our in silico analysis successfully predicted several epitopes capable of binding to MHC-I and MHC-II molecules, suggesting their potential to induce T-helper and cytotoxic responses. Moreover, the selected epitopes fulfilled all the characteristics required for a potentially nontoxic, antigenic, nonallergenic, and soluble vaccine. In addition, the stability and immunogenicity of the MEBV were boosted by including an appropriate adjuvant (β-defensin 3) and linkers (KK, AAY, and GPGPG). The inclusion of appropriate adjuvants that enhance the immune response to epitopes is vital for ensuring vaccine effectiveness. Our results indicate that when coupled with the β-defensin 3 adjuvant, PADRE sequences, and a 6-His tag, the selected vaccine construct led to a stronger and more durable immune response. The 6-His tag added at the C-terminal of the vaccine plays a crucial role in the purification and stabilization of recombinant antigens. Next, the 3D structure of the proposed MEBV was modeled using the most advanced modeling tool, AlphaFold3. Previous studies have shown that several TLRs on the surface of immune cells trigger innate immune responses32. Therefore, molecular docking and MD simulations were used to validate the binding affinity and stability of the designed vaccine with TLR3 and TLR7. The results revealed a strong binding affinity and stable interaction with both TLRs, suggesting that the designed MEBV could potentially activate TLRs and induce strong immunological responses in the host body.
Immune simulation analysis revealed that during the simulation, B-cell and T-cell levels increased with each injection and maintained their population levels, suggesting that the vaccine successfully elicited immune responses. IFN-γ and IL-2 levels surged after the first injection and remained high after successive antigen exposure. This pattern indicates the notable presence of T-helper cells, contributing to efficient Ig synthesis and a powerful humoral response. Furthermore, a greater concentration of D-cells, cytokines, and macrophages increases the host defense against pathogens. Simulation of the immune response induced by our designed vaccine demonstrated that it could produce an adequate immune response after exposure. Thus, the key to designing a new-generation FMDV vaccine is to identify and screen these immunodominant peptides. Next, codon optimization was performed in the E. coli K12 strain to achieve better expression. Previous studies have recommended E. coli as the most preferred model for the bulk production of recombinant proteins26,27,30. The GC content and CAI values measured revealed a greater expression in the E. coli. Finally, in silico restriction cloning using the pET28(+) vector suggested that the designed vaccine could be easily purified and manufactured on a large scale. Overall, our results suggest that the proposed MEBV could effectively trigger an immune response and provide protection against the FMDV.
Although the constructed MEBV and immune simulation results are promising, they should only be regarded as a preliminary stage of vaccine development. The biological complexity of in vivo systems, including variations in antigen processing and presentation, and the possibility of unexpected cross-reactivity or reduced epitope immunogenicity, is not adequately considered by these approaches. Hence, to validate these computational findings, future research should focus on the experimental evaluation of the MEBV construct. This may include expression and purification in E.coli, in vitro assays such as ELISA, lymphocyte proliferation, and cytokine profiling, as well as in vivo immunization in suitable animal models. Although these experimental investigations are beyond the scope of the present study, such studies are essential to validate the predicted immunogenicity, safety, and protective potential of the designed vaccine and advance it toward practical application in FMDV control.
Conclusion
To eradicate, manage, and control FMDV infections in cattle, new control strategies must be implemented, such as the development of novel and effective vaccine candidates without side effects. In the current study, we designed an MEBV against FMDV infection in cattle using numerous structural and immunoinformatics approaches. By targeting all four structural proteins of the FMDV, we carefully selected antigenic and nonallergenic epitopes to construct a potentially immunogenic and safe vaccine candidate. Computational analyses suggested that the candidate vaccine possessed favorable immunological and structural properties that may elicit an effective immune response against the FMDV. However, although the in silico results are promising, they represent only the preliminary steps in vaccine development. Further in vitro and in vivo studies are essential to validate the efficacy and safety of this vaccine candidate, which could pave the way for its potential application in the control and management of FMDV infections in cattle.
Methods
Foot-and-mouth disease virus sequence retrieval
As serotype O is prevalent in most regions with FMD outbreaks, the 3D structure of the FMDV O serotype was retrieved from the protein databank (PDB ID: 7ENP). The FASTA sequence of the FMDV polyprotein was retrieved from the UniProt database (ID: P03305, retrieved on 18th March 2024), which consists of 2332 amino acids and a mass of 258,927 Da. Furthermore, the FASTA sequences of all FMDV capsid proteins (VP1–4) were assessed using PDB. The structural proteins, including VP1 (residues 725–935), VP2 (residues 287–504), VP3 (residues 505–724), and VP4 (residues 202–286), were found to be composed of 211, 218, 219, and 85 amino acids, respectively (Table S8). Subsequently, the antigenicity of all four structural proteins was examined using VaxiJen 2.0 (https://www.ddg-pharmfac.net/vaxijen/VaxiJen/VaxiJen.html) and ANTIGENpro (https://scratch.proteomics.ics.uci.edu/) at thresholds of 0.4 and 0.6, respectively33,34.
B-cell prediction
B-cell epitopes are crucial for designing effective vaccines because they directly elicit antibody production, enable memory responses, and can be tailored to neutralize pathogens. They also allow for the development of synthetic, peptide-based, and broadly protective vaccines that are safe and long-lasting. Proper identification and targeting of B-cell epitopes improve vaccine efficacy while minimizing potential adverse effects. Hence, the ABCpred server (https://webs.iiitd.edu.in/raghava/abcpred/, accessed on 25th July 2024) was used to predict B-cell epitopes, with a window length of 16 residues and threshold value of 0.8. The server uses machine-learning methods to predict epitopes that are immunogenic, unique, and continuous35.
MHC-I specific CTL epitope prediction
The first step in triggering an immune response against a disease is the interaction of antigens with CTLs via Major Histocompatibility class I (MHC-I)36. The current study accessed the NetMHCpan 4.1 server (https://services.healthtech.dtu.dk/services/NetMHCpan-4.1/, accessed on 22nd July 2024) to predict CTL epitopes. Based on the population coverage and significance of BoLA class-I supertypes, epitopes were predicted using the BoLA-6:01301 (HD6) and BoLA-1:02301 (D18.4) alleles37,38,39,40. The epitope length was set to 9-mers (nine residues), and epitopes with the highest prediction score and % rank > 0.5 considered good binders41. This server predicts the binding affinity of peptides for a wide range of MHC-I alleles using artificial neural networks. It covers nearly all MHC-I alleles in humans and many nonhuman alleles. Forecasting the peptide-MHC binding affinity and integrating this information with the population coverage and immunogenicity analysis would enable the development of novel, effective, and targeted vaccines against a range of pathogens.
MHC-II specific HTL epitope prediction
The interaction between HTLs and MHC-II molecules is important for eliciting an effective immune response, particularly for vaccines aimed at promoting long-term immunity. Incorporating HTL epitopes into vaccine design ensures the activation of helper T cells, thereby enhancing overall immunogenicity42. Hence, the IEDB MHC-II server (http://tools.iedb.org/mhcii/, accessed on 22nd July 2024) was accessed to evaluate the HTL epitopes of each structural protein using the default parameters. The best epitopes were sorted on the basis of their position, binding affinity, and occurrence in different BoLA class-II alleles (BoLA-DRB3*00101, BoLA-DRB3*00202, BoLA-DRB3*01101, BoLA-DRB3*01201, BoLA-DRB3*01501, and BoLA-DRB3*02703). This server uses a combination of machine-learning models and experimental repositories to predict peptide-MHC binding43,44.
Epitope toxicity prediction
The development of a subunit vaccine requires the selection of a nontoxic epitope. Therefore, the toxicity of each selected B-cell, CTL, and HTL epitope was evaluated using the ToxinPred2 server (https://webs.iiitd.edu.in/raghava/toxinpred2/)45. The threshold value was set to 0.6, and a prediction score > 0.6 considered toxic. After passing nontoxic filters, the epitopes were used to create the final MEBV.
Construction and physiochemical characterization of the multi-epitope-based vaccine
All epitopes selected using the above methodology (B-cell, CTL, and HTL epitopes from each capsid protein) were combined with different linkers, an adjuvant, PADRE sequence, and 6-His tag at the C-terminal to construct the final MEBV subunit. The KK, AAY, and GPGPG linkers were used to connect the selected epitopes. These linkers are crucial because they allow amino acid residues to exhibit maximum flexibility and enable proper epitope separation within the host. The adjuvant, β-defensin 3 (UniProt: P46161) of Bos taurus, which has a sequence length of 57 amino acids, was retrieved from UniProt and added to the N-terminal of the vaccine subunit46,47,48,49. A PADRE peptide sequence was added to the adjuvant via an EAAAK linker. The PADRE sequence plays a crucial role in computational MEBV design due to its immunogenic properties and ability to enhance vaccine efficacy32,50,51. Next, all selected B-cell, CTL, and HTL epitopes for all structural proteins were added by connecting them with the KK, AAY, and GPGPG linkers. Lastly, a 6-His (HHHHHH) tag was added to the C-terminal of the vaccine subunit, which would play a significant role in purifying and stabilizing the recombinant antigens. Moreover, its ability to improve antigen preparation and delivery platforms makes it indispensable for subunit vaccine development52,53,54.
Furthermore, different physiochemical properties of the final vaccine subunit, such as molecular weight, theoretical pI, aliphatic index, half-life, and GRAVY, were evaluated using the Expasy ProtParam server (https://web.expasy.org/protparam/)55. The antigenicity of the final vaccine construct was assessed using the VaxiJen v.2.0 and ANTIGENpro webservers33,34. The allergenicity of the final vaccine construct was examined using the AllerTop v.2.0 (https://www.ddg-pharmfac.net/allertop_test/) and AllergenFP v.1.0 (https://ddg-pharmfac.net/AllergenFP/) servers56.
Secondary structure information, such as the presence of α-helixes, β-sheets, and coils in the constructed vaccine, was examined using the PSIPRED v.4.0 (http://bioinf.cs.ucl.ac.uk/psipred/) and GOR IV (https://npsa-pbil.ibcp.fr/cgi-bin/npsa_automat.pl?%20page=/NPSA/npsa_gor4.html) web servers57,58. Next, the tertiary structure of the vaccine subunit was predicted using AlphaFold359. The GalaxyRefine2 server (https://galaxy.seoklab.org/cgi-bin/submit.cgi?%20type=REFINE2) was used to refine the vaccine structure generated from AlphaFold360,61. Finally, the stereochemical properties of the modeled structure were validated using the SAVES v.6.1 server (https://saves.mbi.ucla.edu/)62,63,64.
Molecular docking
Computationally designed vaccines are often docked with TLR molecules to predict their potential to activate the innate immune system in a host. TLRs are a class of pattern recognition receptors found on immune cells that recognize conserved molecular patterns of pathogens, making them crucial for initiating and shaping immune responses. Hence, the 3D AlphaFold structures of the bovine TLR3 (residues 27–699) and TLR7 (residues 26–830) ectodomains were retrieved from the UniProt database (UniProt IDs: Q5TJ59 and A1XC24, respectively). To understand the binding interactions occurring between these receptors (TLR3 and TLR7 ectodomains) and the modeled vaccine, a protein-protein docking study was performed using the HDOCK server65,66. Based on binding affinity, the model with the lowest docking score was considered for further analysis. The PDBsum program was used to analyze the protein-protein interactions between the vaccine and receptors67.
Molecular dynamics simulations
To understand the structural dynamics and binding interactions that occur between the vaccine and TLRs (TLR3 and TLR7) at the atomistic level, a 100-ns all-atom MD simulation was performed for both complexes using the GROMACS 2022.2 package68,69. As both TLRs work at a slightly acidic pH, we assessed the probable protonation states of the charged residues of TLR3 and TLR7 at pH 6.0 using the H + + server70. Subsequently, the systems were prepared using the AMBER-ff99SB-ILDN force field and explicit water-solvent models71. A triclinic periodic boundary condition at a distance of 1.6 nm was used with TIP3P water molecules. Appropriate Na+/Cl− counterions were added to neutralize the system. The steepest-descent algorithm was used to minimize the energy of the system with a 1000 kJ/mol/nm tolerance72. Both systems were equilibrated through NVT and NPT ensemble processes using the V-rescale and Parrinello–Rahman barostat coupling algorithm, respectively73,74. The parameters used for the MD simulation have been described in our previous communications23,75,76. The final MD production run was carried out for 100 ns, and 2-ps coordinates were saved throughout the trajectory. The last 50 ns of the trajectory information was used to calculate the binding free energy using the g_mmpbsa tool, as described in our previous studies77,78,79.
Codon optimization and in silico cloning
To maximize expression in E. coli, the vaccine subunit sequence was uploaded to the JCat server for reverse translation and codon optimization. The web server ensured the best expression, and the GC percentage and CAI were noted80,81. Following cloning of the vaccine into the pET28a(+) vector, restriction sites for XhoI and BamHI were appended to the N- and C-termini of the vaccine sequence, respectively. Finally, the SnapGene tool was used to clone the vaccine.
Immune response simulations
Immune response to the proposed vaccine was evaluated using the C-ImmSim server (https://kraken.iac.rm.cnr.it/C-IMMSIM). This program evaluates the antibody- and cell-mediated responses of a mammalian immune system to vaccine systems31,82. For most MEBVs currently in use, the recommended optimal interval between doses is typically four weeks. Consequently, the entire simulation spanned 1050 steps, encompassing three successive injections at time steps 1, 84, and 168 (with one time step being equivalent to 8 h). The simulation used default settings for all other parameters.
Data availability
All data generated or analyzed during this study are included in this published article and its Supplementary Information files.
References
Jamal, S. M. & Belsham, G. J. Foot-and-mouth disease: past, present and future. Vet. Res. 44, 116 (2013).
Knight-Jones, T. J. D. & Rushton, J. The economic impacts of foot and mouth disease – What are they, how big are they and where do they occur? Prev. Vet. Med. 112, 161–173 (2013).
Domingo, E., Baranowski, E., Escarmı́s, C. & Sobrino, F. Foot-and-mouth disease virus. Comp. Immunol. Microbiol. Infect. Dis. 25, 297–308 (2002).
Wubshet, A. K. et al. Foot and mouth disease vaccine efficacy in africa: a systematic review and meta-analysis. Front Vet. Sci 11, 1360256 (2024).
Order - Picornavirales. in. in Virus Taxonomy. 835–839 (eds King, A. M. Q., Adams, M. J., Carstens, E. B. & Lefkowitz, E. J.) (Elsevier, 2012). https://doi.org/10.1016/B978-0-12-384684-6.00070-7
Brito, B. P., Rodriguez, L. L., Hammond, J. M., Pinto, J. & Perez, A. M. Review of the global distribution of Foot-and-Mouth disease virus from 2007 to 2014. Transbound. Emerg. Dis. 64, 316–332 (2017).
Klein, J. Understanding the molecular epidemiology of foot-and-mouth-disease virus. Infect. Genet. Evol. 9, 153–161 (2009).
Domingo, E. et al. Emergence and selection of RNA virus variants: memory and extinction. Virus Res. 82, 39–44 (2001).
Haydon, D. T., Samuel, A. R. & Knowles, N. J. The generation and persistence of genetic variation in foot-and-mouth disease virus. Prev. Vet. Med. 51, 111–124 (2001).
Wong, C. L., Yong, C. Y., Ong, H. K., Ho, K. L. & Tan, W. S. Advances in the diagnosis of Foot-and-Mouth disease. Frontiers Veterinary Science 7, 477 (2020).
Maree, F. F. et al. Challenges and prospects for the control of foot-and-mouth disease: an African perspective. VMRR 5, 119–138 (2014).
Diaz-San Segundo, F., Medina, G. N., Stenfeldt, C. & Arzt, J. De Los santos, T. Foot-and-mouth disease vaccines. Vet. Microbiol. 206, 102–112 (2017).
Paton, D. J. et al. Selection of foot and mouth disease vaccine strains–a review. Rev. Sci. Tech. 24, 981–993 (2005).
Parida, S. Vaccination against foot-and-mouth disease virus: strategies and effectiveness. Expert Rev. Vaccines. 8, 347–365 (2009).
Guo, H. C. et al. Foot-and-mouth disease virus-like particles produced by a SUMO fusion protein system in Escherichia coli induce potent protective immune responses in Guinea pigs, swine and cattle. Vet. Res. 44, 48 (2013).
Porta, C. et al. Rational engineering of Recombinant picornavirus capsids to produce safe, protective vaccine antigen. PLoS Pathog. 9, e1003255 (2013).
Uddowla, S., Hollister, J., Pacheco, J. M., Rodriguez, L. L. & Rieder, E. A safe Foot-and-Mouth disease vaccine platform with two negative markers for differentiating infected from vaccinated animals. J. Virol. 86, 11675–11685 (2012).
Cao, Y. et al. Poly(I:C) combined with multi-epitope protein vaccine completely protects against virulent foot-and-mouth disease virus challenge in pigs. Antiviral Res. 97, 145–153 (2013).
Usmani, S. S., Kumar, R., Bhalla, S., Kumar, V. & Raghava, G. P. S. Chapter Seven - In Silico tools and databases for designing Peptide-Based vaccine and drugs. in Advances in Protein Chemistry and Structural Biology (ed Donev, R.) vol. 112 221–263 (Academic, (2018).
Mortazavi, B., Molaei, A. & Fard, N. A. Multi-epitope vaccines, from design to expression; an in Silico approach. Hum. Immunol. 85, 110804 (2024).
Ananya et al. Vaccine design and development: exploring the interface with computational biology and AI. International Reviews Immunology 43(6), 361–380. https://doi.org/10.1080/08830185.2024.2374546 (2024).
Rodríguez Pulido, M. & Sáiz, M. Molecular mechanisms of Foot-and-Mouth disease virus targeting the host antiviral response. Front. Cell. Infect. Microbiol. 7, 252 (2017).
Sahoo, S., Lee, H. K. & Shin, D. Structure-based virtual screening and molecular dynamics studies to explore potential natural inhibitors against 3 C protease of foot-and-mouth disease virus. Frontiers Veterinary Science 10, 1340126 (2024).
Ullah, M. A., Sarkar, B. & Islam, S. S. Exploiting the reverse vaccinology approach to design novel subunit vaccines against Ebola virus. Immunobiology 225, 151949 (2020).
Kumar Pandey, R., Ojha, R., Mishra, A. & Kumar Prajapati, V. Designing B- and T-cell multi-epitope based subunit vaccine using immunoinformatics approach to control Zika virus infection. J. Cell. Biochem. 119, 7631–7642 (2018).
Pandey, R. K., Ojha, R., Aathmanathan, V. S., Krishnan, M. & Prajapati, V. K. Immunoinformatics approaches to design a novel multi-epitope subunit vaccine against HIV infection. Vaccine 36, 2262–2272 (2018).
Naz, A. et al. Designing Multi-Epitope vaccines to combat emerging coronavirus disease 2019 (COVID-19) by employing Immuno-Informatics approach. Front Immunol 11, 1663 (2020).
Majee, P., Jain, N. & Kumar, A. Designing of a multi-epitope vaccine candidate against Nipah virus by in Silico approach: a putative prophylactic solution for the deadly virus. J. Biomol. Struct. Dynamics. 39, 1461–1480 (2021).
Cao, Y. et al. Rational design and efficacy of a multi-epitope Recombinant protein vaccine against foot-and-mouth disease virus serotype A in pigs. Antiviral Res. 140, 133–141 (2017).
Bhutta, M. S. et al. A novel immunoinformatics approach for developing a poly-epitope vaccine targeting foot and mouth disease virus, exploiting structural VP proteins. Journal Biomol. Struct. Dynamics 0, 1–17 (2024).
Pathak, R. K., Lim, B., Kim, D. Y. & Kim, J. M. Designing multi-epitope-based vaccine targeting surface Immunogenic protein of Streptococcus agalactiae using immunoinformatics to control mastitis in dairy cattle. BMC Vet. Res. 18, 337 (2022).
Zhu, X., Wang, X., Liu, T., Zhang, D. & Jin, T. Design of multi-epitope vaccine against Porcine rotavirus using computational biology and molecular dynamics simulation approaches. Virol. J. 21, 160 (2024).
Doytchinova, I. A. & Flower, D. R. VaxiJen: a server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC Bioinform. 8, 4 (2007).
Magnan, C. N. et al. High-throughput prediction of protein antigenicity using protein microarray data. Bioinformatics 26, 2936–2943 (2010).
Saha, S. & Raghava, G. P. S. Prediction of continuous B-cell epitopes in an antigen using recurrent neural network. Proteins Struct. Funct. Bioinform. 65, 40–48 (2006).
Hewitt, E. W. The MHC class I antigen presentation pathway: strategies for viral immune evasion. Immunology 110, 163–169 (2003).
Yılmaz Çolak, Ç. Computational design of a Multi-epitope vaccine against clostridium chauvoei: an immunoinformatics approach. Int. J. Pept. Res. Ther. 27, 2639–2649 (2021).
Gaafar, B. B. M., Ali, S. A., Abd-elrahman, K. A. & Almofti, Y. A. Immunoinformatics Approach for Multiepitope Vaccine Prediction from H, M, F, and N Proteins of Peste des Petits Ruminants Virus. Journal of Immunology Research 6124030 (2019). (2019).
Mollazadeh, S., Bakhshesh, M., Keyvanfar, H. & Nikbakht Brujeni, G. Identification of cytotoxic T lymphocyte (CTL) epitope and design of an Immunogenic multi-epitope of bovine ephemeral fever virus (BEFV) glycoprotein G for vaccine development. Res. Vet. Sci. 144, 18–26 (2022).
Kolla, H. B. et al. Immuno-informatics study identifies conserved T cell epitopes in non-structural proteins of bluetongue virus serotypes: formulation of a computationally optimized next-generation broad-spectrum multi-epitope vaccine. Front Immunol 15, 1424307 (2024).
Reynisson, B., Alvarez, B., Paul, S., Peters, B. & Nielsen, M. NetMHCpan-4.1 and NetMHCIIpan-4.0: improved predictions of MHC antigen presentation by concurrent motif Deconvolution and integration of MS MHC eluted ligand data. Nucleic Acids Res. 48, W449–W454 (2020).
Oli, A. N. et al. Immunoinformatics and vaccine development: an overview. ImmunoTargets Therapy. 9, 13–30 (2020).
Wang, P. et al. A systematic assessment of MHC class II peptide binding predictions and evaluation of a consensus approach. PLoS Comput. Biol. 4, e1000048 (2008).
Wang, P. et al. Peptide binding predictions for HLA DR, DP and DQ molecules. BMC Bioinform. 11, 568 (2010).
Sharma, N., Naorem, L. D., Jain, S. & Raghava, G. P. ToxinPred2: an improved method for predicting toxicity of proteins. Brief. Bioinform. 23, bbac174 (2022).
Jiang, Q. et al. Screening of Bovine Coronavirus Multiepitope Vaccine Candidates: An Immunoinformatics Approach. Transboundary and Emerging Diseases 5986893 (2024). (2024).
Samad, A. et al. Immune epitopes identification and designing of a multi-epitope vaccine against bovine leukemia virus: a molecular dynamics and immune simulation approaches. Cancer Immunol. Immunother. 71, 2535–2548 (2022).
Sinha, P. R. et al. In Silico development of a Multi-Epitope subunit vaccine against bluetongue virus in Ovis Aries using immunoinformatics. Pathogens 13, 944 (2024).
Mackenzie-Dyck, S., Kovacs-Nolan, J., Snider, M. & Babiuk, L. A. Drunen Littel-van Den hurk, S. Inclusion of the bovine neutrophil Beta-Defensin 3 with glycoprotein D of bovine herpesvirus 1 in a DNA vaccine modulates immune responses of mice and cattle. Clin. Vaccine Immunol. 21, 463–477 (2014). van.
Uddin, M. B. et al. A candidate multi-epitope vaccine against lumpy skin disease. Transbound. Emerg. Dis. 69, 3548–3561 (2022).
Zhang, Z. et al. Identification of B-cell epitopes on structural proteins VP1 and VP2 of senecavirus A and development of a multi-epitope Recombinant protein vaccine. Virology 582, 48–56 (2023).
Mahmoud, N. A., Elshafei, A. M. & Almofti, Y. A. A novel strategy for developing vaccine candidate against Jaagsiekte sheep retrovirus from the envelope and gag proteins: an in-silico approach. BMC Vet. Res. 18, 343 (2022).
Bahadori, Z. et al. Design, development, and assessment of a novel multi-peptide vaccine targeting pspc, psaa, and PhtD proteins of Streptococcus pneumoniae. Int. J. Biol. Macromol. 258, 128924 (2024).
Hakami, M. A. An immunoinformatics and structural vaccinology approach to design a novel and potent multi-epitope base vaccine targeting Zika virus. BMC Chem. 18, 31 (2024).
Gasteiger, E. et al. ExPASy: the proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res. 31, 3784–3788 (2003).
Dimitrov, I., Bangov, I., Flower, D. R. & Doytchinova, I. AllerTOP v.2—a server for in Silico prediction of allergens. J. Mol. Model. 20, 2278 (2014).
McGuffin, L. J., Bryson, K. & Jones, D. T. The PSIPRED protein structure prediction server. Bioinformatics 16, 404–405 (2000).
Kouza, M., Faraggi, E., Kolinski, A. & Kloczkowski, A. The GOR method of protein secondary structure prediction and its application as a protein aggregation prediction tool. In Prediction of Protein Secondary Structure (eds Zhou, Y. et al.) 7–24 (Springer, 2017). https://doi.org/10.1007/978-1-4939-6406-2_2.
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with alphafold 3. Nature 630, 493–500 (2024).
Heo, L., Park, H. & Seok, C. GalaxyRefine: protein structure refinement driven by side-chain repacking. Nucleic Acids Res. 41, W384–W388 (2013).
Lee, G. R., Won, J., Heo, L. & Seok, C. GalaxyRefine2: simultaneous refinement of inaccurate local regions and overall protein structure. Nucleic Acids Res. 47, W451–W455 (2019).
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Colovos, C. & Yeates, T. O. Verification of protein structures: patterns of nonbonded atomic interactions. Protein Sci. 2, 1511–1519 (1993).
Pontius, J., Richelle, J. & Wodak, S. J. Deviations from standard atomic volumes as a quality measure for protein crystal structures. J. Mol. Biol. 264, 121–136 (1996).
Yan, Y., Zhang, D., Zhou, P., Li, B. & Huang, S. Y. HDOCK: a web server for protein–protein and protein–DNA/RNA Docking based on a hybrid strategy. Nucleic Acids Res. 45, W365–W373 (2017).
Yan, Y., Tao, H., He, J. & Huang, S. Y. The HDOCK server for integrated protein–protein Docking. Nat. Protoc. 15, 1829–1852 (2020).
Laskowski, R. A. et al. Structural summaries of PDB entries. Protein Sci. 27, 129–134 (2018).
Van Der Spoel, D. et al. GROMACS: fast, flexible, and free. J. Comput. Chem. 26, 1701–1718 (2005).
BEKKER, H. et al. GROMACS - A SIMULATIONS: 4th International Conference on Computational Physics (PC 92). PHYSICS COMPUTING ’92 252–256 (1993).
Gordon, J. C. et al. H++: a server for estimating p Ka s and adding missing hydrogens to macromolecules. Nucleic Acids Res. 33, W368–W371 (2005).
Lindorff-Larsen, K. et al. Improved side-chain torsion potentials for the amber ff99SB protein force field. Proteins 78, 1950–1958 (2010).
Meza, J. C. Steepest descent. WIRE Comput. Stat. 2, 719–722 (2010).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: A new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Sahoo, S., Gosu, V., Lee, H. K. & Shin, D. Deciphering the conformational changes induced by high-risk NsSNPs in β-lactoglobulin. Heliyon 10, e40040 (2024).
Sahoo, S. et al. Impact of NsSNPs in human AIM2 and IFI16 gene: a comprehensive in Silico analysis. J. Biomol. Struct. Dynamics. 42, 2603–2615 (2024).
Sahoo, S., Purohit, P., Samantaray, S. & Meher, B. R. Identification of Antiviral Phytocompounds as Potential Anti-Dengue Agents against DENV NS2B/NS3 Protease: An Integrated Molecular Modelling and Dynamics Approach. ChemistrySelect 9, e202400384 (2024).
Sahoo, S. et al. In vitro and in Silico studies to explore potent antidiabetic inhibitor against human pancreatic alpha-amylase from the methanolic extract of the green microalga chlorella vulgaris. J. Biomol. Struct. Dynamics. 42, 8089–8099 (2024).
Sahoo, S., Lee, H. K. & Shin, D. Elucidating the structural dynamics induced by active site mutations in 3 C protease of foot-and-mouth disease virus. PLOS ONE. 20, e0321079 (2025).
Carbone, A., Zinovyev, A. & Képès, F. Codon adaptation index as a measure of dominating codon bias. Bioinformatics 19, 2005–2015 (2003).
Grote, A. et al. JCat: a novel tool to adapt codon usage of a target gene to its potential expression host. Nucleic Acids Res. 33, W526–W531 (2005).
Rapin, N., Lund, O., Bernaschi, M. & Castiglione, F. Computational immunology Meets bioinformatics: the use of prediction tools for molecular binding in the simulation of the immune system. PLOS ONE. 5, e9862 (2010).
Acknowledgements
The authors thank the Jeonbuk National University for providing the necessary facilities for carrying out this study.
Funding
The authors declare that financial support was received for the research, authorship, and/or publication of this article. This study was supported by the Ministry of Education of the Republic of Korea and the National Research Foundation of Korea (NRF-2022R1A2C4002510); and the Science and Technology Project Opens the Future of the Region through the INNOPOLIS FOUNDATION grant funded by the Ministry of Science and ICT of Korea (2022-DD-UP-0333).
Author information
Authors and Affiliations
Contributions
Conceptualization: S.S. and D.S.; Formal analysis: S.S.; Investigation: S.S.; Supervision: D.S. and H.L.; Writing–original draft: S.S.; Writing–review and editing: S.S., H.L., and D.S.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sahoo, S., Lee, HK. & Shin, D. An integrated structural and immunoinformatic approach to design a multi-epitope based vaccine against the foot-and-mouth disease virus. Sci Rep 15, 35911 (2025). https://doi.org/10.1038/s41598-025-19826-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-19826-6