Abstract
A novel colorimetric genosensor was developed for the detection of feline panleukopenia virus (FPV) DNA, based on the peroxidase-like activity of ZnFe2O4 (ZFO) nanoparticles. The synthesized ZFO nanoparticles catalytically oxidized 3,3′,5,5′-tetramethylbenzidine (TMB), resulting in the generation of a characteristic blue color. Upon the addition of 5% sulfuric acid stop solution, this intermediate was converted into a yellow diimine product, whose absorbance was measured at 450 nm using a microplate reader. ZFO nanoparticles were subsequently coated onto concave glass slides. 5′-amino-modified oligonucleotide probes were immobilized on APTES-modified glass surfaces to serve as capture probes. In the presence of the target sequence and subsequent hybridization with the immobilized probes, the spatial interaction between ZFO nanoparticles and the TMB substrate was inhibited, leading to a reduced intensity of the blue color. The limit of detection was determined to be 300 ng/µL for the amplified gene target in PCR reactions and 103 virus particles per gram of feces in clinical fecal samples. The results demonstrated that the proposed genosensor is simple, sensitive, and exhibits high selectivity, high reproducibility of fabrication, and stability. This platform may offer a practical tool for the screening of suspected feline panleukopenia cases in resource-limited settings. Moreover, with appropriate oligonucleotide probes, the genosensor can be adapted for the detection of a broad range of microorganisms.
Similar content being viewed by others
Introduction
Feline Panleukopenia (FP), caused by the Feline Panleukopenia Virus (FPV), is a highly contagious and often fatal disease in cats. The virus is primarily transmitted via the fecal-oral route and initially replicates in the oropharyngeal mucosa before disseminating systemically. Infected animals commonly exhibit symptoms such as diarrhea, leukopenia, anemia, thrombocytopenia, and cerebellar ataxia1,2. Studies have shown that FPV remains endemic in many regions. A high shedding prevalence during FP outbreaks is common and was reported to reach up to 95.5% with case-fatality rates reaching up to 90% in young, unvaccinated kittens3. Outbreaks can lead to significant mortality, especially in high-density environments such as animal shelters, rescue centers, and catteries. Due to its resilience in the environment and fecal-oral transmission, FPV can spread rapidly and cause severe economic burdens through quarantine measures, disinfection protocols, and animal loss. Although FPV has no known zoonotic risk, its impact on animal health, shelter management, and veterinary resource allocation makes it a critical target for improved diagnostics1,2,3. Without timely intervention, FPV can result in high mortality rates. Due to its rapid transmission and high fatality rate, accurate and early diagnosis is critical for outbreak control, effective treatment, and improved clinical outcomes.
Several diagnostic tools are available for FPV detection, including virus isolation, hemagglutination assays (HA), enzyme-linked immunosorbent assays (ELISA), immunofluorescence techniques (IF), and nucleic acid-based detection assays such as polymerase chain reaction (PCR)4,5,6. While these techniques offer high sensitivity and specificity, they often require sophisticated instruments, cold-chain storage, trained personnel, or prolonged processing times. In resource-limited or field settings, these requirements become major barriers to timely diagnosis. Furthermore, false negatives can occur in antigen-based tests, especially during early infection stages. Thus, there is a growing demand for alternative detection platforms that are rapid, affordable, easy to use, and suitable for point-of-care applications7,8,9,10,11.
The detection and quantification of nucleic acids from pathogenic microorganisms in clinical specimens are crucial for diagnostic microbiology and biotechnology12. Oligonucleotide hybridization serves as the foundational technique for nucleic acid detection in various diagnostic methods, including genosensors13,14. The development of miniaturized genosensors represents a strategic approach for genomic analyses, providing advantages such as rapidity, simplicity, cost-effectiveness, and high sensitivity. These sensors have broad applicability in both field and laboratory diagnostics15,16,17. Genosensors are designed using a variety of oligonucleotide probe binding mechanisms, including adsorption, covalent binding, and entrapment18,19, along with different transducers, such as semiconductors, metals, and carbon. Nanomaterials, which can be synthesized in various shapes and sizes using solvothermal and hydrothermal methods20,21,22, have demonstrated significant catalytic, optical, and electrical properties, offering considerable benefits in biosensor design16,18,23,24.
Colorimetric biosensing has recently gained attention due to its convenience, enabling visual detection by the naked eye without the need for specialized instruments, and its simplicity compared to other biosensing methods25. The main challenge in colorimetric measurements lies in converting detection signals into visible color outputs. Chromogenic substances, coupled with enzymes, provide a powerful combination for colorimetric assays26,27. The development of materials with enzyme-like properties has generated significant interest, addressing the limitations of traditional enzymes, such as high cost, storage difficulties, instability, and environmental sensitivity28,29. Metal oxide nanomaterials, including ZnFe₂O₄ (ZFO), CeO₂, Co₃O₄, and noble metal nanoparticles like AuNPs, Au/Pd NPs, and Ag/Pt NPs, exhibit peroxidase-like activity similar to that of horseradish peroxidase (HRP) on TMB substrates30,31,32,33,34,35,36. These nanomaterials are widely used in biosensing applications for nucleic acid analysis due to their large surface areas and controlled catalytic activities37. The 3,3′,5,5′-tetramethylbenzidine (TMB) substrate, commonly employed in enzyme-linked immunosorbent assays (ELISA) and immunodetection assays, is highly sensitive, non-mutagenic38,39, and commercially available in ready-to-use solutions. The peroxidase-like activity of Fe₂O₃, Fe₃O₄, and mixed iron oxide nanoparticles has been studied based on the catalytic oxidation of TMB in the presence of H₂O₂, demonstrating intrinsic peroxidase-like activity and colorimetric output40,41,42,43. Among these, ZFO has garnered significant interest due to its favorable stability and cost-effectiveness37.
Although the peroxidase-like properties of ZFO have been reported previously, their application in viral diagnostics particularly for veterinary pathogens such as feline FPV has not been explored. To our knowledge, this is the first study to utilize ZFO in a colorimetric assay platform for the sensitive and cost-effective detection of FPV. There are some reports on the use of genosensors for detecting parvoviruses in humans and dogs44,45, and no genosensor has been designed for FPV diagnosis to date. In this study, we report the development of a novel colorimetric DNA genosensor for the detection of FPV. The platform utilizes ZFO nanoparticles as peroxidase mimics and exploits hybridization-induced suppression of their catalytic activity toward TMB. This hybridization-mediated signal modulation enables sensitive and selective detection of FPV genomic sequences. To our knowledge, this is the first genosensor specifically designed for FPV detection, offering a portable and cost-effective diagnostic tool with potential applications in both clinical and field settings.
Materials and methods
Preparation and characterization of ZFO nanostructure
ZFO nanoparticles were synthesized following the solvothermal method described by Kurian et al.46. Stoichiometric amounts of metal salts, Zn(NO₃)₂·6 H₂O (analytical grade) and Fe(NO₃)₃·9 H₂O (analytical grade), were dissolved in a solution containing 100 mL ethylene glycol (C₂H₄(OH)₂) and 1.6 mL polyethylene glycol (H(OCH₂CH₂)nOH). After complete dissolution of the salts, ammonium acetate (CH₃CO₂NH₄) was added to the solution and stirred thoroughly. The resulting mixture was transferred to a Teflon-lined autoclave and heated at 180 °C for 8 h. The product was then washed several times with distilled water, acetone, and ethanol, followed by drying at 70 °C for 12 h. The synthesized ZFO nanoparticles were characterized using X-ray diffraction (XRD; Bruker D8 diffractometer), scanning electron microscopy (SEM), and VSM analyses.
Preparation of ZFO np’s coated glass slides
Single concave microscope glass slides (Eisco™; Thermo Fisher Scientific) were obtained for the study. The concave regions of the slides were initially modified using 1500 grit silicon carbide abrasive paper to enhance the binding of nanostructures to the glass surface. The slides were then oxidized by exposure to a Piranha solution (2:1 ratio of 18 M sulfuric acid to 30% hydrogen peroxide) for 2 h. After oxidation, the slides were thoroughly rinsed with distilled water to remove any residual Piranha solution before silanization. Then, the ZFO nanoparticles were applied to the modified slides to fill uniformly the abraded surface regions.
Prior to the coating procedure, the ZFO NPs were amine-functionalized using 3-aminopropyltriethoxysilane (APTES, 99%, Aldrich). Briefly, 0.1 g of synthesized ZFO NPs were treated with 100 µL of APTES in 10 mL of ethanol (99.9%, Merck). The mixture was stirred at room temperature overnight. The amine-modified nanoparticles were separated by centrifugation (10,000 rpm for 8 min at 4 °C). The supernatant was discarded, and the precipitate was resuspended in ethanol. This washing process was repeated four times to ensure thorough purification. The final product was dried overnight in an oven at 30 °C47.
APTES-functionalized ZFO suspensions at concentrations of 0.1%, 0.2%, 0.3%, and 0.4% were prepared in ethanol and homogenized by sonication (100 W). Subsequently, 10 µL of each suspension was transferred into the concave region of four modified glass slides. The slides were heated at 220 °C for 1 h to fix the APTES-functionalized ZFO NPs onto the slides48.
Before immobilizing oligonucleotide probes, the ability of the APTES-functionalized ZFO NPs to induce a colorimetric reaction with TMB was assessed. For this, 30 µL of TMB One Solution (Promega Corporation, Madison, WI) was added to each concave well. The slides were incubated at room temperature for 4 h. The oxidized TMB was then collected using a sampler, transferred into microtiter plates, and quantified by an ELISA reader (BioTek, ELX 800, USA) at wavelengths of 450 nm (after addition of 30 µL ELISA stop solution; sulfuric acid 5%) and 630 nm to evaluate the peroxidase-like activity of the ZFO NPs, based on their catalytic oxidation of TMB in the presence of hydrogen peroxide.
Immobilization of FPV specific oligonucleotides on the APTES functionalized glass slides
5′-Amino-modified oligonucleotides (Table 1), complementary to the VP1 structural protein of FPV49, were diluted in 0.1 M sodium bicarbonate buffer (NaHCO₃, pH 8.5) for immobilization on silane-functionalized substrates. Varying concentrations of the oligonucleotide solutions (10–102 pmol) were manually spotted (10 µL per well) onto the APTES-functionalized glass slide surfaces and incubated overnight at 37 °C in a humidity chamber to ensure proper binding.
Following immobilization, the substrates were washed sequentially with 0.1% Triton X-100, hydrochloric acid (pH 4), 0.1 M potassium chloride, and deionized water, as previously described50,51. To evaluate the effect of oligonucleotide binding on the catalytic activity of ZFO nanoparticles, 30 µL of TMB substrate was added to each well. The slides were incubated at room temperature for 4 h, and the resulting colorimetric changes were quantified using an ELISA reader, as described in the previous section.
Evaluation of FPV Geno sensing experiments
The attenuated Nobivac Tricat Trio vaccine (FPV strain MW-1; ≥10⁴.³ CCID₅₀) was utilized as the source of FPV in this study. Viral genomic DNA was extracted from the MW-1 strain using the DynaBio™ Virus Nucleic Acid Extraction Kit (Takapouzist, Iran), following the manufacturer’s protocol. The concentration and purity of the extracted nucleic acids were assessed via NanoDrop® spectrophotometry.
Subsequently, a 618 bp fragment corresponding to the VP1 gene of FPV was amplified using conventional PCR (polymerase chain reaction), as described by Parthiban et al.49. The specific primers used are presented in Table 1. PCR was performed under the following conditions: initial denaturation at 94 °C for 30 s, followed by 35 cycles of denaturation at 94 °C for 30 s, annealing at 55 °C for 45 s, and extension at 72 °C for 45 s. A final elongation step was carried out at 72 °C for 10 min. Amplified products were electrophoresed on 1.3% agarose gel and visualized under UV illumination (see Fig. 1). The quantity and purity of the PCR products were re-evaluated using NanoDrop®.
Amplification of the VP1-specific sequence from the Feline Panleukopenia Virus (FPV) MW1 strain by conventional PCR. Lane L: 100 bp DNA ladder (mid-range ladder); Lanes 1 and 2: 618 bp PCR product corresponding to the VP1 gene of the FPV vaccinal strain; Lane 3: negative control (distilled water as template) for the PCR reaction. The image is cropped from an original size gel electrophoresis image presented in supplementary files section.
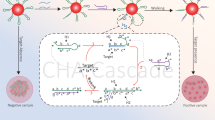
To assess the performance of the designed genosensor, 30 µL of the amplified VP1 fragment (at concentrations ranging from 100 to 1000 ng/µL, diluted in TBE buffer) was applied to the sensor and incubated overnight at 90 °C in a hybridization chamber. During the 90 °C overnight incubation, slides were placed in a humidified sealed chamber and covered with parafilm to minimize evaporation and maintain constant volume. After hybridization, the slides were washed using deionized water (AdvaHyb AH 100 wash solutions, Munich, Germany) pre-heated to 42 °C, according to the manufacturer’s instructions52. This was followed by the addition 30 µL of a commercial TMB substrate containing H₂O₂ for nanozyme-catalyzed color development. The slides were incubated for 4 h at room temperature. Following incubation, 50 µL of 5% H2SO4 was added to each well to terminate the reaction and convert the oxidized TMB from blue to yellow, as per standard ELISA protocols. Subsequently, the oxidized TMB solution was collected from each well, transferred to microtitre plates, and the absorbance was measured using an ELISA reader at 450 nm. A schematic overview of the genosensor reaction steps is presented in Fig. 2.
Schematic representation of the fabrication process and mechanism of the blue color production by the developed genosensor.
Clinical specimen’s investigation
The potential and applicability of the developed genosensor for detecting FPV in clinical specimens were evaluated using fecal samples from a healthy cat (see Fig. 3). Briefly, 10-fold serial dilutions (ranging from 10⁰ to 10⁵) of the FPV vaccinal strain (MW-1) were prepared and individually mixed with 1 gram of feces collected from a clinically healthy, FPV-negative cat. Genomic DNA was then extracted from each spiked sample using a commercial genomic DNA extraction kit, following the manufacturer’s instructions.
Distribution of optical density values obtained following the addition of defined FPV titers to feline fecal samples for the evaluation of the developed genosensor. Black lines indicate median values; white lines represent individual data points; and shaded polygons illustrate the estimated data density. Polygons positioned above the horizontal dotted line correspond to samples diagnosed as positive by the genosensor, while those below the line represent samples not detected as positive.
The extracted DNA was subsequently analyzed using both the designed genosensor and a standard real-time PCR assay to determine and compare the limit of detection (LOD) of each method. Notably, this analysis was conducted without prior target sequence amplification to evaluate direct detection capability. As a negative control, DNA extracted from non-infected fecal samples of the same healthy cat was included to assess the specificity of the genosensor.
Stability, reproducibility, and selectivity
We conducted a stability test on the genosensor by storing it at 4 Cº in a dark case and evaluating its current response every week up to 7 weeks. For the assessment of the reproducibility of the sensor we carried out identical approach in three experiments on the feces specimens containing 1000 FPV particles per gram. To assess the selectivity of the biosensor in detecting FPV genome, we extracted DNA from feces of a healthy cat, feces containing 1000 FPV particles per gram of feces, and cat feces containing 1000 canine parvovirus particle (CPV) per gram (Nobivac DHPPi; Live CPV vaccine, strain 154 ≥ 107.0 TCID50) for interference experiments.
All experimental data were statistically analyzed using SPSS software (version 16.0). Differences between groups were evaluated, and p-values < 0.05 were considered statistically significant. Box plots were generated using BoxPlotR, a web-based tool for box plot visualization (available at: http://shiny.chemgrid.org/boxplotr/).
Results and discussion
The XRD pattern (Fig. 4a), referenced in 96-901-2442 JCPDS, validates the successful synthesis of zinc ferrite nanoparticles. The pattern reveals four peaks at 2θ values of 30, 35, 56, and 62˚, which correspond to the (022), (131), (151), and (044) planes. The crystallite size of ZFO nanoparticles was determined to be 20 nm using the Debye-Scherrer equation (Eq. 1), which is lower than values reported in other studies.
(a) The XRD pattern of ZnFe2O4 nanoparticles. (b) SEM images at various magnifications and (c) EDX results of ZnFe2O4 nanoparticles. (d) VSM analysis of ZnFe2O4 nanoparticles.
Figure 4b presents SEM images that reveal ZFO is made up of spherical particles roughly 30 nm in diameter, smaller than the sizes noted in other research47. The EDX results (Fig. 4c) indicate that the synthetic sample is impurity-free.
The results of the VSM analysis on the magnetic properties of zinc ferrite nanoparticles are presented in Fig. 4d. The lack of hysteresis loops verifies their superparamagnetic nature. Zinc ferrite’s saturation magnetization was measured at 55 emu/g, which exceeds values found in similar studies.
The peroxidase-like activity of the synthesized ZFO was evaluated through its catalytic oxidation of the chromogenic substrate TMB (Fig. 5). In the absence of ZFO, the TMB solution remained colorless; however, upon addition of ZFO, a distinct color change to the blue spectrum was observed (Fig. 6), indicating catalytic oxidation of TMB and confirming the intrinsic peroxidase-mimetic activity of ZFO nanoparticles, in agreement with previous findings42. This experiment was designed to directly assess the intrinsic catalytic (nanozyme-like) properties of ZFO, by monitoring its ability to oxidize TMB in the presence of H₂O₂, a hallmark of peroxidase-mimetic activity. The observed yellow color and absorbance at 450 nm confirmed the formation of the diimine oxidation product post acid addition. UV-Vis spectrophotometric analysis (Nanodrop) at 450 nm and 630 nm demonstrated a concentration-dependent increase in absorbance with rising ZFO levels (0.1–0.4%), with significant enhancement detected from 0.2% ZFO onward (p < 0.05). Based on these results, 0.2% and 0.3% ZFO concentrations were selected for subsequent experiments involving oligonucleotide probe immobilization. Furthermore, significant differences (p < 0.05) were observed between the two wavelengths tested, with measurements at 450 nm yielding higher absorbance values and greater sensitivity. Therefore, 450 nm was chosen for all subsequent colorimetric analyses.
Experimental workflow for evaluating the oxidizing effect of ZFO nanoparticle (NP)-coated slides on TMB substrate. Step 1: Glass slides were coated with different concentrations of ZFO NPs (0.4–0%). Step 2: The coated slides were incubated with TMB substrate for 4 h. Step 3: Then, TMB solution from each slide was collected, transferred to ELISA plates, and mixed with stop solution. Step 4: Optical density (OD) values were measured at 450 nm using an ELISA reader, and results were expressed as the mean of five independent replicates.
UV-Vis spectra of oxidized TMB resulting from the oxidation reaction in the presence of varying concentrations (0–0.4%) of ZnFe2O4 (ZFO), recorded at wavelengths of 450 and 630 nm. Statistical comparisons indicate the significance of differences in TMB oxidation across ZFO concentrations. Center lines represent medians; box limits correspond to the 25th and 75th percentiles as calculated using R software; whiskers denote minimum and maximum values; crosses indicate sample means; bars represent 95% confidence intervals of the means; and box widths are proportional to the square root of the sample size.
To achieve stable covalent immobilization of nucleic acid probes onto glass substrates, the surface was first modified using APTES via a silanization reaction. Under humid conditions, the ethoxy groups of APTES hydrolyze to form silanol groups, which subsequently condense with hydroxyl groups on the glass surface, forming covalent siloxane (Si–O–Si) bonds and generating a monolayer with exposed primary amine groups. This reaction is typically spontaneous and does not require an external initiator or catalyst, although mild heating or extended incubation may be used to improve the uniformity and stability of the coating. To enable covalent binding of terminally amino-modified oligonucleotides, the APTES-functionalized surface is then reacted with glutaraldehyde, a bifunctional aldehyde linker. Glutaraldehyde reacts with surface amines to introduce pendant aldehyde groups, which subsequently form Schiff base (imine) linkages with the primary amine groups on the nucleic acid probes. These imines can be stabilized by reductive amination, typically using sodium cyanoborohydride (NaBH₃CN) as a reducing agent, yielding stable secondary amine bonds. This process does not require enzymatic catalysis or radical initiators, as the reactions are chemically driven and proceed under mild aqueous conditions53. The described coupling strategy is accurate and complete for applications requiring stable nucleic acid immobilization on glass surfaces, such as genosensors and microarrays, and ensures high binding efficiency and signal reproducibility.
To evaluate the immobilization efficiency and functional effects of amino-modified oligonucleotide probes on the APTES-functionalized ZFO-coated glass slide substrates, complementary capture probes were applied at varying concentrations (0-100 pmol/µL) to concave wells coated with either 0.2% or 0.3% ZFO. Following probe immobilization, the catalytic activity of ZFO was assessed by adding TMB substrate to each well and incubating at room temperature for 4 h. The resulting chromogenic signals were measured using an ELISA reader to obtain optical density (OD) values (Fig. 7). Results demonstrated that glass slides coated with 0.3% ZFO generated higher OD values compared to those with 0.2% ZFO. In both ZFO concentrations, the highest OD values were observed on wells lacking immobilized probes (0 pmol), with significant reductions in OD detected upon application of 40 pmol and higher concentrations of oligonucleotide probes (p < 0.05). While both 0.2% and 0.3% ZFO surfaces exhibited statistically significant differences across the range of oligonucleotide concentrations tested, the 0.3% ZFO-coated slides displayed greater sensitivity to incremental probe concentrations, yielding more pronounced differences in OD values (Fig. 7). Notably, probe concentrations of 20 pmol did not significantly inhibit the catalytic oxidation of TMB, suggesting insufficient surface coverage or hybridization interference at this level. Therefore, the 0.3% APTES-ZFO-coated glass slides with 20 pmol immobilized oligonucleotides were selected for subsequent experiments to investigate the hybridization-dependent inhibition of ZFO-mediated TMB oxidation.
Effects of varying concentrations of oligonucleotide probes (0–100 picomol/µl) on the reduction of optical density (OD) values generated by TMB oxidation in the presence of 0.2 and 0.3% ZnFe2O4. Statistical significance levels are indicated for comparisons of the effects of different ZFO concentrations on TMB oxidation. Center lines represent medians; box limits correspond to the 25th and 75th percentiles as determined by R software; whiskers extend to the minimum and maximum values; crosses represent sample means; bars denote 95% confidence intervals of the means; and box widths are proportional to the square root of the sample size.
Hybridization of complementary DNA sequences was achieved using probe oligonucleotides covalently immobilized on the APTES-functionalized ZFO surfaces, as illustrated in Fig. 8. Under optimized conditions, the peroxidase-mimicking catalytic activity of the 0.3% ZFO-APTES-probe (20 pmol) system was evaluated via the oxidation of TMB in the presence of varying concentrations (0-1000 ng/µL) of the amplified VP1 fragment from the FPV genome (Fig. 9). Following overnight hybridization, the optical density (OD) corresponding to each target concentration was measured, with a maximum OD of 0.91 ± 0.06 observed in the absence of the target sequence.
Effects of hybridization with varying concentrations (0–1000 ng/µL) of the amplified VP1 target sequence, yielded by real-time PCR, on the reduction of optical density (OD) values generated by TMB oxidation in the presence of 0.3% ZnFe2O4 and 20 picomol/µL oligonucleotide probe. Statistical significance levels are presented for comparisons of the impact of different target fragment hybridization concentrations on TMB oxidation.
Detection workflow using designed colorimetric slides. Slides coated with 0.3% ZFO and oligo probes (20 pmol) were incubated overnight with varying concentrations of the target gene (1000− 100 ng/µL). After 4 h incubation with TMB substrate, the developed solution was transferred to ELISA plates with stop solution, and OD values were measured at 450 nm (mean of five replicates).
As depicted in Fig. 8, a significant decrease in OD was observed upon hybridization with VP1 fragment concentrations ≥ 300 ng/µL, while lower concentrations (≤ 200 ng/µL) produced minimal inhibition of catalytic activity. The OD values demonstrated a linear inverse correlation with increasing VP1 concentrations within the tested range (Fig. 10). A visible transition from dark blue to colorless could be discerned on the slides, allowing for qualitative detection by the naked eye. The regression equation for the colorimetric response was determined to be Y = 0X + 0.93, with a coefficient of determination (R²) of 0.9139. Based on these results, the LOD for the genosensor was established at 300 ng/µL.
Linear regression analysis of the relationship between FPV genome concentrations (VP1 amplified fragment) ranging from 0 to 1000 ng/µL and the optical density (OD) values obtained from TMB oxidation reactions, measured using an ELISA reader on the developed genosensor.
The presence of the FPV genome in fecal samples was successfully detected using both the real-time PCR and the developed colorimetric genosensor (Fig. 11). The real-time PCR method exhibited an LOD of 10 virus FPV/gr of feces, whereas the genosensor demonstrated an LOD of 1000 FPV/gr, indicating a markedly higher sensitivity of the real-time PCR in comparison to the colorimetric method. Importantly, no false-positive results were observed with either method, confirming the high specificity and reliability of the developed genosensor for FPV detection.
Comparison of the limit of detection (LOD) between real-time PCR and the designed genosensor using cat feces infected with 1–105 titers of FPV vaccinal particles. Detectable virus titers in fecal samples are shown in red, while undetectable virus titers are represented in blue for each assay.
As shown in Fig. 12A, after 7 weeks of storage, the sensor retained 93.2% of its initial current response, indicating good stability. Statistical analyses demonstrated that the sensor could detect 1000 LOD of FPV up to 6 weeks successfully, but the at the seventh week it could not detect such virus titer in a sample. Besides, Fig. 12B demonstrates the reproducibility of the biosensor. The difference in current response among three experiments which were fabricated using the identical approach, was found to be less than 1.6% when detecting the same titers (1000 LOD) of FPV particles in fecal specimens. Furthermore, Fig. 12C demonstrates the selectivity of the genosensor. Due to the genetic similarities in the DNA sequence of the VP1 gene between FPV and CPV, the sensor could detect both viruses in the fecal samples without any significant differences in OD values. But the comparison of FPV and CPV experiments with fecal samples from healthy cats revealed significant differences between OD values demonstrating the selectivity and specificity of the sensor for parvoviruses sharing genetic similarities in the targeted genes. These findings indicate that the genosensor exhibits excellent stability, reproducibility, and selectivity.
(A) Stability of the sensor at 4 °C in 7 weeks (W1–W7), (B) Reproducibility of the sensor in three experiments (1st, 2nd, and 3rd ), and (C) Selectivity test of the sensor detecting FPV versus canine parvovirus (CPV vaccine) and negative control containing unspecific DNA (NG). The vertical axis in all graphs demonestrates the OD values.
FPV is excreted in all body secretions during the active phase of infection, with particularly high concentrations found in gastrointestinal (GI) tract secretions2,54. Early detection of FPV in clinical specimens is crucial for identifying infected animals, implementing appropriate treatment, and preventing virus shedding in the environment. Various laboratory methods have been developed to detect the FPV genome or particles in clinical specimens, including PCR, virus isolation, immunochromatographic tests (IC), HA, ELISA, and IF tests4,9. Direct detection methods such as PCR, IC, HA, ELISA, and FA eliminate the need for virus propagation and allow for the direct identification of FPV in clinical samples. Nucleic acid-based methods, particularly PCR, are highly sensitive and specific compared to other techniques, and they offer the advantage of being rapid when the genome sequence of the circulating virus is known. However, despite these benefits, PCR techniques are associated with high costs, require trained personnel, and are susceptible to contamination that can lead to false results.
In this context, the use of ZFO in colorimetric biosensors has garnered significant attention due to its low cost, stability, and ability to effectively conjugate with functionalized oligonucleotide probes. These probes, modified with functional groups, bind to target analytes with high affinity and specificity, serving as either identification or signal elements. ZFO nanocomposites, particularly when functionalized, have been extensively explored for diagnostic and therapeutic applications due to their ease of synthesis, chemical modification, and long shelf life37,55,56. Such biosensors, once fabricated, can be easily deployed in clinical or field settings without the need for specialized laboratory equipment or highly trained staff.
In this study, real-time PCR exhibited greater sensitivity and discriminative power for detecting FPV in fecal samples, confirming its advantages over the designed genosensor. Real-time PCR successfully detected FPV in fecal samples with a LOD of 10 FPV/gr, while the genosensor demonstrated an LOD of 1000 (Fig. 11). Although the LOD of 300 ng/µL may limit ultra-trace detection, it remains suitable for practical applications where target concentrations are relatively high, such as in symptomatic clinical samples or heavily contaminated materials. Despite its lower sensitivity, the genosensor offers significant advantages in terms of cost, ease of use, and accessibility. In contrast, real-time PCR requires expensive thermocyclers, specialized kits, and skilled personnel to analyze and interpret results, limiting its applicability, especially in resource-limited settings.
Table 2 provides a summary of previously reported biosensors designed for human and animals parvoviruses and compares them with the genosensor constructed in the present study. It could be seen that the introduction of VP1 protein for specific FPV DNA detection yields a sensitive platform, offering a broad detection range and low LOD. Therefore, the VP1-based colorimetric genosensor proves to be an effective and reliable platform for FPV genome detection.
A key challenge in genomic analysis from fecal samples is the quality and quantity of extracted DNA, which can be affected by the sample’s complex composition and the presence of inhibitors that interfere with DNA amplification or hybridization57. Variability in DNA yield and fragment size depending on the extraction method is a well-known issue that can influence downstream diagnostic assays.
This study also confirms the peroxidase-like activity of ZFO nanoparticles, demonstrated by their catalytic oxidation of TMB in the presence of hydrogen peroxide. The enzymatic mimicry of ZFO is influenced by several factors, including pH, nanoparticle concentration, and TMB concentration58. Consistent with previous reports, maximum absorbance of oxidized TMB occurred under acidic conditions (pH = 3.0), with activity declining sharply at higher pH levels likely due to rapid decomposition of H₂O₂ into water and oxygen, reducing its availability for TMB oxidation58,59. The catalytic activity increased with higher ZFO concentrations, with significant enhancement observed from 0.2% onwards; thus, this concentration was selected for further experiments. Additionally, a positive correlation was found between TMB concentration and peroxidase-like activity, although the fixed pH and concentration of commercial TMB substrates limited independent evaluation of these parameters.
While the present study demonstrates the successful application of ZFO as peroxidase mimics for colorimetric detection of FPV, it should be noted that a detailed kinetic evaluation, including determination of Michaelis–Menten parameters such as Km and Vmax, was not conducted. This is because our focus was primarily on developing a practical and sensitive genosensor suitable for point-of-care or laboratory diagnostics, using a commercially available substrate under fixed conditions. Nevertheless, future studies may explore the catalytic kinetics of ZFO nanozymes in greater depth to further elucidate their mechanistic properties and optimize their performance across a broader range of biosensing applications.
In optimizing the genosensor platform, glass substrates emerged as the most effective surface for nucleotide probe immobilization, offering superior stability and sensitivity. The performance of the glass-based genosensor is highly dependent on surface chemistry, with factors such as oxidation method, APTES functionalization time, probe binding duration, and hybridization time significantly influencing hybridization efficiency and overall sensitivity60,61. Treatment of the glass surface with Piranha solution for 2 h increased the density of silanol groups, thereby enhancing probe immobilization. Longer APTES reaction times resulted in a higher density of amine groups, facilitating improved probe attachment. However, excessive probe density caused by prolonged immobilization can create steric hindrance, which may reduce hybridization efficiency and consequently diminish the colorimetric response62.
The sensing mechanism of the developed genosensor is based on the peroxidase-mimetic catalytic activity of ZFO functionalized with APTES. These nanoparticles catalyze the oxidation of TMB in the presence of H₂O₂, yielding a blue-colored product with a characteristic absorbance peak at 630 nm. Upon immobilization of amino-modified single-stranded DNA (ssDNA) probes onto the ZFO surface, partial suppression of catalytic activity is observed. This is attributed to the electrostatic and steric effects of the negatively charged DNA molecules, which can interfere with substrate access to the nanoparticle surface. However, a more pronounced inhibition occurs after hybridization with complementary target DNA sequences (Fig. 2). The resulting dsDNA forms a more rigid and bulky structure, leading to greater surface coverage and enhanced blocking of active sites on the ZFO nanoparticles. Additionally, the local charge distribution and hydrophilicity are altered upon duplex formation, further suppressing electron transfer required for TMB oxidation. To validate this mechanism, control experiments were performed in which the TMB oxidation signal (absorbance at 450 nm) was compared across four conditions: (i) bare ZFO-coated surface; (ii) ZFO with immobilized ssDNA probe; (iii) ZFO with ssDNA probe hybridized with non-complementary DNA; and (iv) ZFO with ssDNA probe hybridized with complementary target DNA. Only the specific hybridization event in condition (iv) resulted in significant inhibition of the colorimetric signal, confirming that the catalytic suppression is induced by DNA hybridization rather than nonspecific adsorption. Therefore, the sensor operates on a hybridization-induced catalytic inhibition mechanism, wherein the presence of target DNA results in decreased TMB oxidation and reduced color intensity, which is proportional to target concentration. This catalytic inhibition is consistent with previous findings that DNA hybridization can suppress the peroxidase-mimicking activity of nanozymes through steric hindrance and surface blocking effects63,64,65.
This study also highlights the complex interplay between catalytic activity and surface functionalization in determining the performance of the ZFO-based genosensor. While several critical factors influencing sensor performance were identified, some variables such as the effects of TMB concentration and pH could not be fully explored due to experimental limitations. Future research should systematically investigate these parameters to further optimize the fabrication and functionality of ZFO-based genosensors. Despite the genosensor’s lower sensitivity compared to real-time PCR, its potential for rapid, cost-effective, and user-friendly diagnostics in field or clinical settings makes it a valuable alternative for detecting FPV, particularly in resource-limited environments. Although the current detection workflow involves steps similar to conventional ELISA techniques, our method simplifies key aspects by replacing enzyme-antibody conjugates with stable ZFO nanozymes and by conducting the assay on a concave glass slide using minimal reagent volumes. These features reduce costs and logistical complexity. Furthermore, the colorimetric output enables visual or smartphone-based detection, suggesting strong potential for field deployment with minimal infrastructure.
A limitation of the current study is the lack of recovery validation in biological matrices such as fecal samples. Although our proof-of-concept was established using purified nucleic acids, further testing in clinical or environmental samples is underway to assess matrix effects and enhance the robustness of the assay.
Conclusions
A ZFO nanostructure exhibiting peroxidase-like activity was synthesized and employed in the development of a colorimetric genosensor for detecting FPV. A specific oligonucleotide probe, designed from the FPV gene sequence, was immobilized onto APTES-functionalized glass slides to create a simple yet sensitive detection platform. Target viral DNA and various nucleic acid sequences were hybridized with the immobilized probes, and hybridization efficiency was assessed via colorimetric analysis. The genosensor demonstrated high sensitivity and specificity for FPV, with no cross-reactivity or false-positive signals observed in the presence of non-specific nucleic acids, including those extracted from feline fecal samples. Additionally, validation with clinical specimens confirmed the sensor’s applicability and effectiveness for FPV detection in real-world diagnostic settings.
Data availability
The datasets used and analysed during the current study available from the corresponding author on reasonable request.
References
Hartmann, K., Hofmann-Lehmann, R. & Sykes, J. E. Feline Leukemia Virus Infection. In Greene’s Infect. Dis. Dog Cat, 5 Ed. 382–413 https://doi.org/10.1016/B978-0-323-50934-3.00032-X (Elsevier, 2022).
Jacobson, L. S., Janke, K. J., Giacinti, J. & Weese, J. S. Diagnostic testing for feline panleukopenia in a shelter setting: a prospective, observational study. J. Feline Med. Surg. 23, 1192–1199. https://doi.org/10.1177/1098612X211005301 (2021).
Rehme, T. et al. Feline panleukopenia outbreaks and risk factors in cats in animal shelters. Viruses 14.6, 1248. (2022).
Truyen, U. et al. Feline panleukopenia ABCD guidelines on prevention and management. J. Feline Med. Surg. 11, 538–546. https://doi.org/10.1016/j.jfms.2009.05.002 (2009).
SOMA, T., OHTA, K., YAMASHITA, R. & SASAI, K. Anti-feline panleukopenia virus serum neutralizing antibody titer in domestic cats with the negative or low hemagglutination Inhibition antibody titer. J. Vet. Med. Sci. 81, 252–255. https://doi.org/10.1292/jvms.18-0472 (2019).
Sykes, J. E. Feline panleukopenia virus infection and other viral enteritides. In Canine Feline Infect. Dis., 187–194. https://doi.org/10.1016/B978-1-4377-0795-3.00019-3. (Elsevier Inc., 2013).
Fiscus, S. A., Mildbrand, M. M., Gordon, J. C., Teramoto, Y. A. & Winston, S. Rapid enzyme-linked immunosorbent assay for detecting antibodies to canine parvovirus. Am. J. Vet. Res. 46, 859–863 (1985).
Hofmann-Lehmann, R. et al. Prevalence of antibodies to feline parvovirus, calicivirus, herpesvirus, coronavirus, and immunodeficiency virus and of feline leukemia virus antigen and the interrelationship of these viral infections in free-ranging lions in East Africa. Clin. Diagn. Lab. Immunol. 3, 554–562. https://doi.org/10.1128/cdli.3.5.554-562.1996 (1996).
Schunck, B., Kraft, W. & Truyen, U. A simple touch-down polymerase chain reaction for the detection of canine parvovirus and feline panleukopenia virus in feces. J. Virol. Methods. 55, 427–433. https://doi.org/10.1016/0166-0934(95)00069-3 (1995).
Ryser-Degiorgis, M. P. et al. Epizootiologic investigations of selected infectious disease agents in free-ranging Eurasian Lynx from Sweden. J. Wildl. Dis. 41, 58–66. https://doi.org/10.7589/0090-3558-41.1.58 (2005).
Veijalainen, P. M. L., Neuvonen, E., Niskanen, A. & Juokslahti, T. Latex agglutination test for detecting feline panleukopenia virus, canine parvovirus, and parvoviruses of fur animals. J. Clin. Microbiol. 23, 556–559. https://doi.org/10.1128/jcm.23.3.556-559.1986 (1986).
Bouchez, T. et al. Molecular microbiology methods for environmental diagnosis. Environ. Chem. Lett. 14 423–441. https://doi.org/10.1007/s10311-016-0581-3. (2016).
Babaei, A. et al. Genosensors as an alternative diagnostic sensing approaches for specific detection of virus species: A review of common techniques and outcomes. TrAC - Trends Anal. Chem. 155, 116686. https://doi.org/10.1016/j.trac.2022.116686 (2022).
Paniel, N., Baudart, J., Hayat, A. & Barthelmebs, L. Aptasensor and genosensor methods for detection of microbes in real world samples. Methods 64, 229–240. https://doi.org/10.1016/j.ymeth.2013.07.001 (2013).
Nanofabrication and Biosystems: Integrating Materials Science, Engineering. Google Books, (n.d.). (accessed January 17, 2025); https://books.google.com/books?hl=en&lr=&id=sW2poBEATN8C&oi=fnd&pg=PR7&ots=J5Vtg2dzOF&sig=NifqJeQNABWaA5-9z3nCSaG3eC0#v=onepage&q&f=false
Rahi, A., Sattarahmady, N. & Heli, H. Zepto-molar electrochemical detection of Brucella genome based on gold nanoribbons covered by gold nanoblooms. Sci. Rep. 5, 18060. https://doi.org/10.1038/srep18060 (2015).
Biosensors and Modern Biospecific Analytical Techniques - Google Books. (n.d.). (accessed January 17, 2025); https://books.google.co.uk/books?hl=en&lr=&id=Xjw3B8pg0KYC&oi=fnd&pg=PR1&dq=%5D+L.+Gorton+(Ed.),+Biosensors+and+Modern+Biospecific+Analytical+Techniques,+Elsevier+Science,+2005.&ots=iGc_lqU7V-&sig=paazpj2iF0WbXf2Uf-Dvgj0M9RY&redir_esc=y#v=onepage&q=%5DL.Gorton(Ed.)%2CBiosensorsandModernBiospecificAnalyticalTechniques%2CElsevierScience%2C2005.&f=false
Nazari-Vanani, R., Heli, H. & Sattarahmady, N. An impedimetric genosensor for leishmania infantum based on electrodeposited cadmium sulfide nanosheets. Talanta 217 https://doi.org/10.1016/j.talanta.2020.121080 (2020).
De Silva, A. P., Moody, T. S. & Wright, G. D. Fluorescent PET (Photoinduced electron Transfer) sensors as potent analytical tools. Analyst 134, 2385–2393. https://doi.org/10.1039/b912527m (2009).
Feng, S. H. & Li, G. H. Hydrothermal and solvothermal syntheses. In: Mod. Inorg. Synth. Chem. 2 Ed, 73–104. https://doi.org/10.1016/B978-0-444-63591-4.00004-5 (Elsevier Inc., 2017).
Lai, J., Niu, W., Luque, R. & Xu, G. Solvothermal synthesis of metal nanocrystals and their applications. Nano Today. 10, 240–267. https://doi.org/10.1016/j.nantod.2015.03.001 (2015).
Aliofkhazraei, M. Handbook of nanoparticles. https://doi.org/10.1007/978-3-319-15338-4 (2015).
Sattarahmady, N., Heli, H. & Dehdari Vais, R. A flower-like nickel oxide nanostructure: synthesis and application for choline sensing. Talanta 119, 207–213. https://doi.org/10.1016/j.talanta.2013.11.002 (2014).
Heli, H., Majdi, S., Jabbari, A., Sattarahmady, N. & Moosavi-Movahedi, A. A. Electrooxidation of dextromethorphan on a carbon nanotube-carbon microparticle-ionic liquid composite: applied to determination in pharmaceutical forms. J. Solid State Electrochem. 14, 1515–1523. https://doi.org/10.1007/s10008-009-0979-y (2010).
Liu, D. M., Xu, B. & Dong, C. Recent advances in colorimetric strategies for acetylcholinesterase assay and their applications. TrAC - Trends Anal. Chem. 142, 116320. https://doi.org/10.1016/j.trac.2021.116320 (2021).
Melikishvili, S., Piovarci, I. & Hianik, T. Advances in colorimetric assay based on Aunps modified by proteins and nucleic acid aptamers. Chemosensors 9, 281. https://doi.org/10.3390/chemosensors9100281 (2021).
Liang, X., Wang, X., Zhang, Y., Huang, B. & Han, L. Selective Inhibition toward dual Enzyme-like activities of iridium nanozymes for a specific colorimetric assay of malathion without enzymes. J. Agric. Food Chem. 70, 3898–3906. https://doi.org/10.1021/acs.jafc.1c06954 (2022).
Xie, J. et al. Analytical and environmental applications of nanoparticles as enzyme mimetics. TrAC - Trends Anal. Chem. 39, 114–129. https://doi.org/10.1016/j.trac.2012.03.021 (2012).
Zhang, X. Z., Zhou, Y., Zhang, W., Zhang, Y. & Gu, N. Polystyrene@Au@prussian blue nanocomposites with enzyme-like activity and their application in glucose detection. Colloids Surf. Physicochem Eng. Asp. 490, 291–299. https://doi.org/10.1016/j.colsurfa.2015.11.035 (2016).
Gao, L. et al. Intrinsic peroxidase-like activity of ferromagnetic nanoparticles. Nat. Nanotechnol. 2, 577–583. https://doi.org/10.1038/nnano.2007.260 (2007).
Wei, H. & Wang, E. Nanomaterials with enzyme-like characteristics (nanozymes): Next-generation artificial enzymes. Chem. Soc. Rev. 42, 6060–6093. https://doi.org/10.1039/c3cs35486e (2013).
Asati, A., Santra, S., Kaittanis, C., Nath, S. & Perez, J. M. Oxidase-Like activity of Polymer‐Coated cerium oxide nanoparticles, Angew. Chemie Int. Ed. 48, 2308–2312. https://doi.org/10.1002/anie.200805279 (2009).
Mu, J., Wang, Y., Zhao, M. & Zhang, L. Intrinsic peroxidase-like activity and catalase-like activity of Co3O4 nanoparticles. Chem. Commun. 48, 2540–2542. https://doi.org/10.1039/c2cc17013b (2012).
Chen, H., Li, Y., Zhang, F., Zhang, G. & Fan, X. Graphene supported Au-Pd bimetallic nanoparticles with core-shell structures and superior peroxidase-like activities. J. Mater. Chem. 21, 17658–17661. https://doi.org/10.1039/c1jm13356j (2011).
Zheng, X. et al. Catalytic gold nanoparticles for nanoplasmonic detection of DNA hybridization, Angew. Chemie - Int. Ed. 50, 11994–11998. https://doi.org/10.1002/anie.201105121 (2011).
Zheng, C. et al. One-pot synthesized DNA-templated ag/pt bimetallic nanoclusters as peroxidase mimics for colorimetric detection of thrombin. Chem. Commun. 50, 13103–13106. https://doi.org/10.1039/c4cc05339g (2014).
Wu, S., Duan, N., Qiu, Y., Li, J. & Wang, Z. Colorimetric aptasensor for the detection of Salmonella enterica serovar typhimurium using ZnFe2O4-reduced graphene oxide nanostructures as an effective peroxidase mimetics. Int. J. Food Microbiol. 261, 42–48. https://doi.org/10.1016/j.ijfoodmicro.2017.09.002 (2017).
Bruun, L., Koch, C., Pedersen, B., Jakobsen, M. H. & Aamand, J. A quantitative enzyme-linked immunoassay for the detection of 2,6- dichlorobenzamide (BAM), a degradation product of the herbicide dichlobenil. J. Immunol. Methods. 240, 133–142. https://doi.org/10.1016/S0022-1759(00)00190-3 (2000).
Bos, E. S., van der Doelen, A. A., van Rooy, N. & Schuurs, A. H. W. M. 3,31,5,51 -Tetramethylbenzidine as an Ames test negative chromogen for Horse-Radish peroxidase in Enzyme-Immunoassay. J. Immunoass. 2, 187–204. https://doi.org/10.1080/15321818108056977 (1981).
Chung, K. T. Mutagenicity studies of benzidine and its analogs: Structure-Activity relationships. Toxicol. Sci. 56, 351–356. https://doi.org/10.1093/toxsci/56.2.351 (2000).
Johnsen, A. R. & Jacobsen, O. S. A quick and sensitive method for the quantification of peroxidase activity of organic surface soil from forests. Soil. Biol. Biochem. 40, 814–821. https://doi.org/10.1016/j.soilbio.2007.10.017 (2008).
Song, L. et al. Graphene oxide-based Fe2O3 hybrid enzyme mimetic with enhanced peroxidase and catalase-like activities. Colloids Surf. Physicochem Eng. Asp. 506, 747–755. https://doi.org/10.1016/j.colsurfa.2016.07.037 (2016).
Mitra, K., Ghosh, A. B., Sarkar, A., Saha, N. & Dutta, A. K. Colorimetric Estimation of human glucose level using γ-Fe 2O3 nanoparticles: an easily recoverable effective mimic peroxidase. Biochem. Biophys. Res. Commun. 451, 30–35. https://doi.org/10.1016/j.bbrc.2014.07.028 (2014).
Khatri, R. et al. A novel amperometric genosensor for rapid detection of canine parvovirus in feces. J. Nanosci. Nanotechnol. 21, 3524–3530. https://doi.org/10.1166/jnn.2021.18998 (2021).
Hosseini, F. et al. Application of biosensors in diagnosis of human parvoviruses. J. Liaquat Univ. Med. Heal Sci. 22, 89–93. https://doi.org/10.22442/jlumhs.2023.01006 (2023).
Kurian, J. & Mathew, M. J. Structural, optical and magnetic studies of CuFe2O4, MgFe2O4 and ZnFe2O4 nanoparticles prepared by hydrothermal/solvothermal method. J. Magn. Magn. Mater. 451, 121–130. https://doi.org/10.1016/j.jmmm.2017.10.124 (2018).
Wang, Y. & Liu, B. Conjugated polymer as a signal amplifier for novel silica nanoparticle-based fluoroimmunoassay. Biosens. Bioelectron. 24, 3293–3298. https://doi.org/10.1016/j.bios.2009.04.020 (2009).
Yang, S. et al. Effect of reaction temperature on grafting of γ-aminopropyl triethoxysilane (APTES) onto kaolinite. Appl. Clay Sci. 62–63. https://doi.org/10.1016/j.clay.2012.04.006 (2012).
Parthiban, M., Aarthi, K. S., Balagangatharathilagar, M. & Kumanan, K. Evidence of feline panleukopenia infection in cats in india, virusdisease. 25 497–499. (2014). https://doi.org/10.1007/s13337-014-0231-y
Guo, Z., Guilfoyle, R. A., Thiel, A. J., Wang, R. & Smith, L. M. Direct fluorescence analysis of genetic polymorphisms by hybridization with oligonucleotide arrays on glass supports. Nucleic Acids Res. 22, 5456–5465. https://doi.org/10.1093/nar/22.24.5456 (1994).
Festag, G. et al. Optimization of gold nanoparticle-based DNA detection for microarrays. J. Fluoresc. 15, 161–170. https://doi.org/10.1007/s10895-005-2524-4 (2005).
Staji, H., Tonelli, A., Zahraei Salehi, T., Iorio, M. & Lopes, F. Genetic characterization of Salmonella typhimurium isolates from faeces of children with gastroenteritis hospitalized in Baqiatollah-Azam hospital, tehran, Iran. J. Med. Microbiol. Infect. Dis. 3 (1), 29–34 (2015).
Zammatteo, N. et al. Comparison between different strategies of covalent attachment of DNA to glass surfaces to build DNA microarrays. Anal. Biochem. 280 (1), 143–150 (2000).
Tuzio, H. Feline panleukopenia. Infect. Dis. Manag Anim. Shelter. Second Ed., 337–366. https://doi.org/10.1002/9781119294382.ch15 (2021).
Park, J. Y., Jeong, H. Y., Il Kim, M. & Park, T. J. Colorimetric detection system for Salmonella typhimurium based on peroxidase-like activity of magnetic nanoparticles with DNA aptamers. J. Nanomater. https://doi.org/10.1155/2015/527126 (2015).
Nimjee, S. M., Rusconi, C. P. & Sullenger, B. A. Aptamers: an emerging class of therapeutics. Annu. Rev. Med. 56, 555–583. https://doi.org/10.1146/annurev.med.56.062904.144915 (2005).
Alinezhad, P., Staji, H. & Sani, R. N. Comparison of three methods including temperature, H2O2/ascorbic acid/sonication, and nitrous acid treatments for overcoming the inhibitory effect of heparin on DNA amplification in realtime-PCR. Int. J. Biol. Macromol. 209, 1298–1306. https://doi.org/10.1016/j.ijbiomac.2022.04.113 (2022).
Bian, B. et al. Pt and ZnFe 2 O 4 nanoparticles immobilized on carbon for the detection of glutathione, ACS appl. Nano Mater. 4, 9479–9488. https://doi.org/10.1021/acsanm.1c01954 (2021).
Sahoo, R. et al. Hierarchical growth of ZnFe 2 O 4 for sensing applications. New. J. Chem. 40, 1861–1871. https://doi.org/10.1039/C5NJ02547H (2016).
Bruce, K. L. Development of oligonucleotide modified substrates for the selective detection of harmful algal bloom causative microalgae (2016).
Walsh, M. K., Wang, X. & Weimer, B. C. Optimizing the immobilization of single-stranded DNA onto glass beads. J. Biochem. Biophys. Methods. 47, 221–231. https://doi.org/10.1016/S0165-022X(00)00146-9 (2001).
Dugas, V., Depret, G., Chevalier, Y., Nesme, X. & Souteyrand, É. Immobilization of single-stranded DNA fragments to solid surfaces and their repeatable specific hybridization: covalent binding or adsorption? Sens. Actuators B Chem. 101, 112–121. https://doi.org/10.1016/j.snb.2004.02.041 (2004).
Gao, L. et al. Intrinsic peroxidase-like activity of ferromagnetic nanoparticles. Nat. Nanotechnol. 2 (9), 577–583 (2007).
Wang, L. et al. A novel colorimetric aptasensor for sensitive Tetracycline detection based on the peroxidase-like activity of Fe3O4@ Cu nanoparticles and sandwich oligonucleotide hybridization. Microchim. Acta. 189 (3), 86 (2022).
Chen, X. et al. Ultratrace antibiotic sensing using aptamer/graphene-based field-effect transistors. Biosens. Bioelectron. 126, 664–671 (2019).
Mao, S. et al. A DNA sensor based on CbAgo effector protein and on a dual electrochemical signal amplification strategy for B19 parvovirus detection. Bioelectrochemistry 162, 108860 (2025).
Yang, Z. et al. Detection of parvovirus B19 genomic fragments using an electrochemical biosensor based on argonaute-assisted silver metallization. Microchem. J. 208, 112293 (2025).
Kim, Y. K. et al. A novel assay for detecting canine parvovirus using a quartz crystal microbalance biosensor. J. Virol. Methods. 219, 23–27 (2015).
Yamakawa, A. C., Basso, C. R., de Albuquerque Pedrosa, V. & Júnior, J. P. Canine parvovirus 2 detection using a LSPR biosensing method with gold nanoparticles. Sens. Diagn. 2 (1), 122–131 (2023).
Acknowledgements
We would like to express our gratitude to Mrs. Behnaz Raeisian, Mrs. Mansoureh Kanani, and Mrs. Leila Bagheripour for their valuable assistance with the experimental procedures.
Author information
Authors and Affiliations
Contributions
H. S. Conceptualization, Methodology, Software, Data curation, Writing, Original draft preparation. Z. A., H. S., and Z. B. Visualization, Investigation. H.S. Supervision. H.S. and Z. B. Writing—Reviewing and Editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Abdoli, Z., Staji, H. & Bahrami, Z. A novel colorimetric DNA sensor for rapid detection of feline panleukopenia virus using ZnFe2O4 nanoparticles and TMB modulation. Sci Rep 15, 36031 (2025). https://doi.org/10.1038/s41598-025-20071-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-20071-0