Abstract
Hepatitis B virus-related acute-on-chronic liver failure (ACHBLF) is a severe condition associated with short-term mortality without liver transplantation. Substantial evidence indicates that necroptosis and immune infiltration play critical roles in ACHBLF development. Therefore, the identification of necroptosis-related biomarkers may be beneficial for prognostic evaluations and may shed light on potential therapeutic targets for ACHBLF. In this study, we used integrated bioinformatics analysis and machine learning algorithms to investigate the correlation between necroptosis and immune infiltration using peripheral blood mononuclear cells from ACHBLF patients. First, after GSE168048 and GSE248217 were obtained from the Gene Expression Omnibus (GEO) database, differentially expressed genes (DEGs) between ACHBLF patients and normal controls were identified. With the help of the weighted gene coexpression network, 211 necroptosis-related DEGs were identified by intersecting the DEGs with necroptosis-related genes (NRGs). The functional characterization of these NRG-related DEGs was subsequently performed using GO, KEGG, and gene set enrichment analysis. Hub genes were identified through the integration of LASSO regression, support vector machines recursive feature elimination, and random forest. Immune-related functions were explored by analyzing the correlations between hub gene expression and immune cells infiltration. Furthermore, mRNA-miRNA-lncRNA interaction network, RNA-binding proteins (RBPs) and transcription factors were predicted using the miRDB, starBase and hTFtarget databases. Finally, target drugs were predicted using a connectivity map. A total of 7461 DEGs were identified between the ACHBLF and normal groups; 3123 genes were upregulated and 4338 genes were downregulated. A total of 211 NRG-related DEGs and 5 hub genes (FCRL3, CDC14A, KLHL22, RALY, and MAP4K1) were obtained. The hub genes were enriched in the T cell receptor signaling pathway, ubiquitin-mediated proteolysis, and the MAPK signaling pathway and were correlated with immune cell infiltration. A lncRNA XIST-miR-424-5p-CDC14A regulatory network was constructed, and 20RBPs and 2 transcription factors (GATA1 and CEBPB) were identified. In addition, 10 candidate drugs were predicted to target MAP4K1. The 5 hub genes could serve as biomarkers for predicting the prognosis of ACHBLF patients and provide clues for new potential therapeutic targets.
Similar content being viewed by others
Introduction
First defined in 2008 by the Asian Pacific Association for the Study of the Liver (APASL), acute-on-chronic liver failure (ACLF) is an acute hepatic insult in patients with previously diagnosed or undiagnosed chronic liver disease1. The main cause of ACLF in Asian is hepatitis B virus (HBV) reactivation, which leads to the specific subtype of ACLF known as hepatitis B virus-related acute-on-chronic liver failure (ACHBLF)2. The characteristic feature of ACHBLF is the high incidence of short-term mortality, which is 50–90%2. Although liver transplantation is the only effective therapy with a clear survival benefit in patients with severe ACHBLF, it cannot be widely applied due to diverse social, economic and cultural factors and a lack of strong legislation and liver donors3. Therefore, there is an urgent need to identify novel pathogenic mechanisms and therapeutic targets for ACHBLF.
Although many studies have focused on the pathophysiology of ACHBLF, the exact pathophysiology remains poorly understood. Recently, necroptosis in liver diseases, including chemical/drug-induced liver injury4, immune-mediated liver injury5, nonalcoholic steatohepatitis6, liver cirrhosis and hepatocellular carcinoma7, has garnered wide interest in the academic field. Necroptosis plays a potentially pathogenic role in ACLF in animal models and inhibiting necroptosis-necroinflammation provides hepatoprotection in ACLF8,9. Moreover, necroptosis plays a critical role in ACHBLF and is associated with poor clinical outcomes10. To identify new and effective biomarkers for the development of therapeutic strategies for ACHBLF, understanding the role of necroptosis during ACHBLF progression is essential.
ACHBLF is characterized by an immunological imbalance, leading to intense pro-inflammation and anti-inflammation, as well as multiple organ failures accompanied by the infiltration of monocytes, neutrophils, dendritic cells, lymphocytes and natural killer cells11. M2-like macrophages have been shown to play a hepatoprotective role by inhibiting necroptosis-related inflammation in an animal model of ACLF8. Exploring the precise role of immune cells infiltration in ACHBLF will provide novel perspectives on the mechanisms involved in the diagnosis and therapeutic strategy for ACHBLF.
Generally, necroptosis and immune infiltration are key triggers that promote ACHBLF progression. Analyzing the relationships between necroptosis and immune infiltration may provide novel insights into the pathogenesis and treatment of ACHBLF. We utilized integrated bioinformatics analysis and machine learning algorithms to explore the unique genes associated with necroptosis and disease-driven changes in immune infiltration in ACHBLF.
Results
Identification of DEGs between the ACHBLF and normal groups
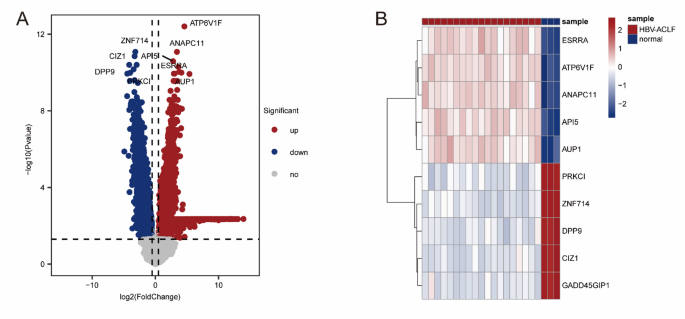
A total of 7461 DEGs were found between the ACHBLF and normal groups when |log2Fold Change|> 0.5 and P Value < 0.05 were used as cutoff values; 3123 genes were upregulated and 4338 genes were downregulated (Fig. 1 A, Table S2). The top 5 most significantly upregulated DEGs were ESRRA, ATP6V1F, ANAPC11, API5 and AUP1, and the top 5 most significantly downregulated DEGs were PRKCI, ZNF714, DPP9, CIZ1 and GADD45GIP1(Fig. 2B).
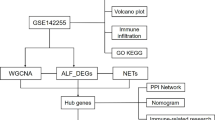
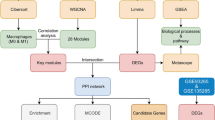
The flowchart of this study.
The DEGs from ACHBLF and normal groups from the GEO database. (A) The volcano of DEGs between ACHBLF and normal groups. (B) Heatmap clustering of the 5 top upregulated and downregulated DEGs.
Screening and functional enrichment of NRGs-related DEGs
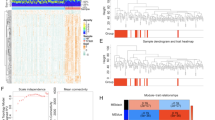
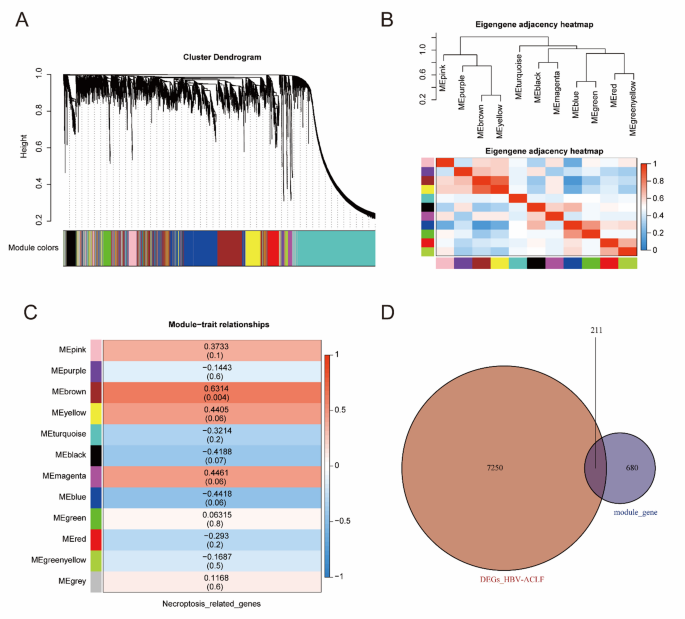
To screen NRG-related DEGs, WGCNA was performed, and 11 co-expression modules were acquired. Genes that were not assigned to any specific module were categorized into grey modules and excluded from further analysis (Fig. 3A). Module eigengenes (MEs) were used to explore the relationships and correlations of each module, and a dendrogram and heatmap were used to visualize the eigengene network (Fig. 3B). To elucidate the physiological significance of genes within modules, we correlated 12 MEs with NRGs and identified the most significant associations. On the basis of the module-trait relationships heatmap (Fig. 3C, Table S3), genes enriched in the brown module (r = 0.6314, P < 0.05) presented the most significant positive correlation with NRGs. Therefore, the brown module, which collectively including 891 genes, was employed for subsequent analyses (Table S4). NRG-related DEGs were identified by taking the intersection of DEGs and NRG-related module genes (Fig. 3D, Table S5).
Screening of the NRGs-related DEGs in ACHBLF. (A) WGCNA modules with different colors under the clustering tree. (B) Correlations and relationships among modules. (C) Heat map of module-trait relationships. (D) The Venn plot identified 211 NRGs-related DEGs among 680 module genes and 7250 DEGs of ACHBLF.
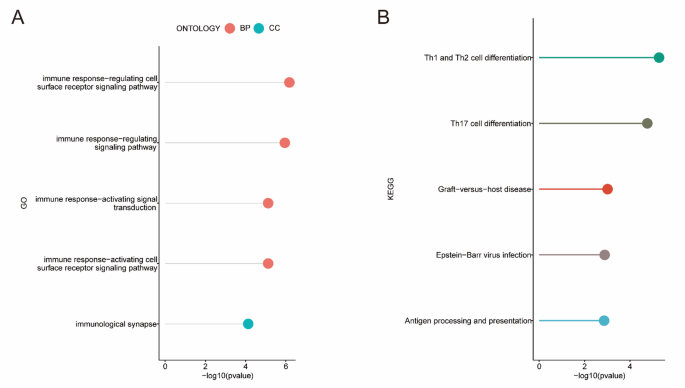
GO functional enrichment analyses revealed possible GO terms, including the CC terms “immunological synapse”, and BP terms “immune response-activating cell surface receptor signaling pathway”, “immune response-activating signal transduction”, “immune response-regulating signaling pathway”, and “immune response-regulating cell surface receptor signaling pathway” (Fig. 4B, Table S6). KEGG12,13,14 analysis revealed that 211 NRG-related DEGs were strongly associated with Th1 and Th2 cell differentiation, Th17 cell differentiation, graft-versus-host disease, Epstein-Barr virus infection and antigen processing and presentation (Fig. 4B, Table S7).
Analysis of GO and KEGG of NRGs-related DEGs. (A) GO enrichment analysis of NRGs-related DEGs. (B) KEGG pathway enrichment analysis of NRGs-related DEGs.
Screening by machine learning and functional enrichment of the hub genes
Three machine learning algorithms were used to screen hub genes. LASSO regression model identified 19 ACHBLF-related signature NRG-related DEGs (Fig. 5A and B), the RF algorithm was used to choose the top 30 genes as signature genes according to the feature weights (MDA and MDG) (Fig. 5C, D and E), and SVM-RFE identified 199 ACHBLF-related signature genes (Fig. 5F and G). A Venn diagram was used to identify the genes common to the three machine learning methods, and 5 NRG-signature genes (FCRL3, CDC14A, KLHL22, RALY, and MAP4K1) were identified as hub genes for subsequent analyses (Fig. 5H, Table S8).
Machine learning identified the signature NRGs-related DEGs of ACHBLF. (A, B) LASSO regression model screening of the signature NRGs-related DEGs of ACHBLF. (C) Random forest error rate versus the number of classification trees. (D) RF methods screen the signature genes according to MDA. (E) RF methods screen the signature genes according to MDG. (F, G) SVM-RFE selected the signature NRGs-related DEGs of ACHBLF. (H) Venn plot among three machine learning methods resulted in five hub genes of ACHBLF.
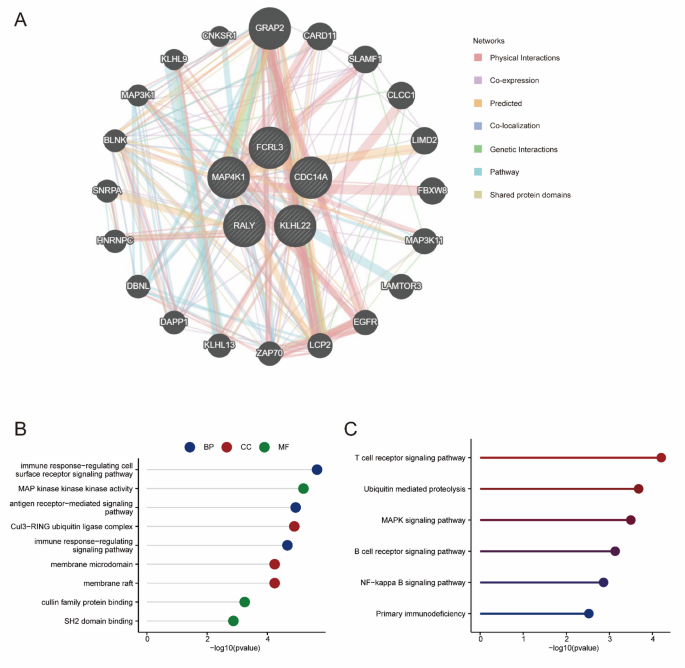
To investigate the biological roles of the hub genes in ACHBLF, protein-protein interaction (PPI) networks were constructed using GeneMANIA and analyzed by Cytoscape (Fig. 6A). GO and KEGG analyses were conducted to elucidate the potential signal pathways associated with the 5 hub genes and 20 hub gene-related genes. As indicated by the GO analysis, the genes were enriched in BPs, including the immune response-regulating cell surface receptor signaling pathway, the antigen receptor-mediated signaling pathway and the immune response-regulating signaling pathway. In terms of cellular components, these genes were predominantly enriched in the Cul3-RING ubiquitin ligase complex, membrane microdomains and membrane rafts. The genes were also enriched in molecular functions (MF), including MAP kinase kinase kinase activity, cullin family protein binding, and SH2 domain binding (Fig. 6B, Table S9). KEGG analysis revealed that these genes were strongly activated in the T cell receptor signaling pathway, ubiquitin-mediated proteolysis, MAPK signaling pathway, the B cell receptor signaling pathway, the NF-kappa B signaling pathway, and primary immunodeficiency (Fig. 6C, Table S10).
Functional enrichment of hub genes of ACHBLF. (A) PPI network analysis of hub genes. (B) GO enrichment analysis. (C) KEGG enrichment analysis.
Validation the expression of the hub genes with the GEO databases
Because no data have been reported on FCRL3, CDC14A, KLHL22, RALY, or MAP4K1 in ACHBLF, we compared the expression of FCRL3, CDC14A, KLHL22, RALY, and MAP4K1 between PBMCs from ACHBLF patients and normal controls. As shown in Fig. 7A, the expression of MAP4K1 was significantly greater in ACHBLF patients than in normal controls; in contrast, the expression levels of FCRL3, CDC14A, KLHL22, and RALY were significantly lower in ACHBLF patients than in normal controls. Moreover, Spearman correlation analysis revealed that CDC14A was significantly positively associated with FCRL3 and KLHL22, and RALY was significantly negatively correlated with MAP4K1 (Fig. 7B).
The expression profile of hub genes between ACHBLF and normal controls. (A) Comparisons of hub genes between ACHBLF and normal controls by boxplot. (B) The correlation analysis of hub genes in ACHBLF by the heatmap.
In the analysis of the molecular pathways of the hub genes in ACHBLF, ssGSEA revealed that CDC14A, the expression of which was upregulated in ACHBLF, was enriched in primary immunodeficiency, autoimmune thyroid disease, graft versus host disease, antigen processing presentation and ribosome (Fig. 8A); FCRL3, the expression of which was upregulated in ACHBLF, was enriched in the intestinal immune network for IgA production, antigen processing and presentation, graft versus host disease, autoimmune thyroid disease and allograft rejection (Fig. 8B); KLHL22, the expression of which was upregulated in ACHBLF, was enriched in graft versus host disease, ribosome, allograft rejection, autoimmune thyroid disease and asthma (Fig. 8C); MAP4K1, the expression of which was upregulated in ACHBLF, was enriched in graft versus host disease, primary immunodeficiency, antigen processing presentation, allograft rejection and viral myocarditis (Fig. 8D); and RALY, the expression of which was upregulated in ACHBLF, was enriched in lysosome, epithelial cell signaling in helicobacter pylori infection, hematopoietic cell lineage, viral myocarditis and cell adhesion molecules (Fig. 8E).
ssGSEA analysis of these hub genes. (A) ssGSEA of CDC14A. (B) ssGSEA of FCRL3. (C) ssGSEA of KLHL22. (D) ssGSEA of MAP4K1. (E) ssGSEA of and RALY.
Constructing a regulatory network of ceRNA, RBP and TFs
To explore the regulatory mechanisms of the hub genes in ACHBLF, an mRNA-miRNA-lncRNA interaction network was constructed, and the results revealed that the signature gene targeted by miR-424-5p is CDC14A and that the targeted lncRNA is XIST (Fig. 9A). Next, the starBase online database was used to search the mRNA/RBP pairs of the signature NRG-related DEGs, and the results revealed that 4 hub genes (CDC14A, MAP4K1, KLHL22 and RALY) had relevant pairing information. An RBP-mRNA regulatory network was generated using the starBase online database, which identified 24 nodes, 20RBPs, 4 mRNAs and 41 edges (Fig. 9B, Table S11). TFs are important regulators of genes expression. According to the hTFtarget database, interaction between 3 hub genes (CDC14A, MAP4K1, and RALY) and 2 TFs (GATA1 and CEBPB) were predicted (Fig. 9C, Table S12).
Constructing regulatory network of ceRNA, RBP and TFs. (A) An interaction network of mRNA-miRNA-lncRNA. Red color represented signature gene, yellow represented microRNA, and blue represented lncRNA. (B) RBP-mRNA regulatory network. Red color represented signature gene and blue represented RBP. (C) Predicted potential TFs of signature NRGs-related DEGs. Red color represented TFs and blue represented signature gene.
Evaluation of immune infiltration and correlation analysis of the hub genes
To compare immune infiltration between ACHBLF patients and normal controls, ssGSEA was conducted with 28 immune infiltration cell markers from the TISIDB database (Table S13). As shown in in Fig. 10A, the proportion of 28 immune cell subsets differed from one individual to another individual with ACHBLF (Fig. 10A), implying that immune cell may have crucial impacts on the pathophysiology of ACHBLF. Thus, immune cell infiltration was compared between ACHBLF patients and normal controls (Table S14). Compared with normal controls, ACHBLF patients had higher enrichment scores for immune infiltration of type 17 T helper cell, activated dendritic cell, plasmacytoid dendritic cell and monocytes (Fig. 10B). Additionally, correlations between the hub genes and immune cell infiltration were evaluated. As a consequence, the expression level of FCRL3 was negatively associated with plasmacytoid dendritic cells (R=−0.723, P < 0.001), monocytes (R=−0.774, P < 0.001) and activated dendritic cells (R=−0.814, P < 0.001); but positively associated with natural killer cells (R = 0.621, P = 0.002), activated CD8 T cells (R = 0.603, P = 0.003) and type 2 T helper cells (R = 0.648, P = 0.001). CDC14A expression was negatively correlated with activated dendritic cells (R=−0.674, P < 0.001) and plasmacytoid dendritic cells (R=−0.621, P = 0.002). KLHL22 expression was negatively correlated with only activated dendritic cells (R=−0.673, P < 0.001) (See Fig. 11).
Evaluate the immune cell infiltration between ACHBLF and normal controls. (A) Immune cell infiltration landscape analysis of ACHBLF. (B) Different enrichment scores of immune infiltration cells between ACHBLF and normal controls. ****p < 0.0001, ***p < 0.001, **p < 0.01, *p < 0.05.
The correlations between hub genes and the infiltration level of the immune cells. (A) Correlation between FCRL3 and plasmacytoid dendritic cell. (B) Correlation between CDC14A and activated dendritic cell. (C) Correlation between FCRL3 and Monocyte. (D) Correlation between FCRL3 and activated dendritic cell. (E) Correlation between KLHL22 and activated dendritic cell. (F) Correlation between CDC14A and plasmacytoid dendritic cell. (G) Correlation between FCRL3 and natural killer cell. (H) Correlation between FCRL3 and activated CD8 T cell. (I) Correlation between FCRL3 and type 2 T helper cell.
Association of the hub genes with cancer hallmark pathways and correlation analyses
Comparisons of the 50 cancer hallmark pathways were performed between ACHBLF patients and normal controls. DNA repair, MTORC1 signaling, the P53 pathway, PI3K-AKT-MTOR signaling, TGF-β signaling and Wnt/β-catenin signaling were significantly activated in ACHBLF patients compared with normal controls. However, Hedgehog signaling was suppressed in ACHBLF patients compared with normal controls (Fig. 12, Table S15). We also analyzed the correlations between the 5 signature NRG-related DEGs and 50 cancer hallmark pathways. The expression of CDC14A was significantly negatively correlated with coagulation, adipogenesis, the xenobiotic metabolism pathway, etc. The expression levels of FCRL3 were negatively associated with the reactive oxygen species pathway, glycolysis, peroxisome, etc. KLHL22 expression was negatively associated with many pathways, such as spermatogenesis, myogenesis, and the late estrogen response. MAP4K1 expression was significantly associated with adipogenesis, angiogenesis and pancreas β cells. In contrast, RALY expression was positively correlated with the reactive oxygen species pathway, glycolysis, fatty acid metabolism, etc. (Fig. 13)
Comparisons of the 50 cancer hallmark pathways. ****p < 0.0001, ***p < 0.001, **p < 0.01, *p < 0.05.
Correlations analysis of 5 hub genes with 50 cancer hallmark pathways. ****p < 0.0001, ***p < 0.001, **p < 0.01, *p < 0.05.
Identification of small-molecule drugs
MAP4K1 and highly associated genes were investigated to predict candidate drugs for ACHBLF therapy from the CMAP database (Table S16). The top 10 target drugs were tiaprofenic-acid, guanabenz, imatinib, etacrynic-acid, brivanib, vincamine, amiloride, lypressin, pizotifen and leflunomide (Table 1). However, no drugs have been approved by the National Medical Products Administration.
Discussion
Liver failure is the terminal stage of all liver diseases and is associated with high mortality worldwide due to the need for liver transplantation. Understanding the underlying mechanism of liver failure can significantly improve the effectiveness of liver failure management. Many studies have indicated that necroptosis and immune infiltration have a poor influence on the prognosis of ACHBLF. These findings provide a novel therapeutic approach for ACHBLF from the perspective of necroptosis suppression and immune regulation. Using integrated bioinformatics analysis and machine learning algorithms, our study identified 5 hub genes (FCRL3, CDC14A, KLHL22, RALY, and MAP4K1) associated with necroptosis and immune infiltration, and enrichment analysis demonstrated that these genes were enriched mainly in the T cell receptor signaling pathway, ubiquitin-mediated proteolysis, the MAPK signaling pathway, the B cell receptor signaling pathway, the NF-kappa B signaling pathway, and primary immunodeficiency. A hub gene-miRNA regulatory network, a hub gene-RBP network and a hub gene-TF network revealed that miR-424-5p, the lncRNA XIST, 20RBPs, and 2 TFs (GATA1 and CEBPB) was promising biomarkers for ACHBLF. Type 17 T helper cells, activated dendritic cells, plasmacytoid dendritic cells and monocytes was increased in ACHBLF patients compared with normal controls and were associated with the expression of hub genes. Candidate drugs, including imatinib and brivanib, which target these hub genes to treat patients with ACHBLF, were identified.
We identified 5 hub genes (FCRL3, CDC14A, KLHL22, RALY, and MAP4K1) associated with necroptosis and immune infiltration in patients with ACHBLF. The expression of MAP4K1 was significantly greater in ACHBLF samples than in normal controls. Mitogen-Activated Protein Kinase Kinase Kinase Kinase 1 (MAP4K1), also known as HPK1, is broadly expressed in multiple cell types, such as macrophages, neutrophils, dendritic cells and lymphocytes15. MAP4K1 has important functions in innate immunity and adaptive immune responses16. The dysregulation of MAP4K1 has been implicated in immune disorders (e.g., autoimmune diseases), hematologic cancers (e.g., acute myeloid leukemia)17, and several solid tumors (e.g., glioblastoma multiforme18. Previous studies have reported that MAP4K1 plays important roles in apoptosis, autophagy19, necrosis20, and ferroptosis21 in cancer and myocardial infarction. Necroptosis involves interactions with necrosis and apoptosis, processes that each can switch from activated to suppressed, and vice versa, in ACLF. Therefore, interference with the expression of MAP4K1 may be a potential therapeutic intervention for ACHBLF. FCRL3, a member of the immunoglobulin receptor superfamily22, is a multifaceted immune regulator with dual roles in activating and inhibitory signaling23. Genetic polymorphisms within FCRL3 are associated with either increased vulnerability or a protective effect regarding autoimmune diseases. For example, in systemic lupus erythematosus (SLE), the upregulation of FCRL3 expression in Tregs is correlated with the dysfunctional coexpression of TIGIT, thereby hindering effective immune regulation and indicating a link to T cell exhaustion24. FCRL3 could inhibit regulatory T cells function in malignancies and autoimmune diseases25. Regulatory T cells are redistributed from the periphery to the liver, and increased regulatory T cells predict a worse prognosis in ACHBLF patients26. These findings highlight FCRL3 as a critical immune checkpoint molecule with context-dependent roles in autoimmunity and inflammation. Therefore, FCRL3 is also a potential molecular target for the treatment of ACHBLF. CDC14A, a member of the dual specificity protein tyrosine phosphatase family, is involved in cell cycle regulation, adhesion and apoptosis27, particularly in the exit from mitosis and the initiation of DNA replication. The deletion of CDC14A in HCT116 cells results in slight impairment of DNA damage repair28. The overexpression of CDC14A delays mitotic entry by suppressing Cdc2 kinase activity, positioning it as a negative regulator of the cell cycle29. Necroptosis is a form of programmed cell death marked by cell swelling, plasma membrane rupture, and DNA damage and repair30; therefore, investing whether the modulation of CDC14A can alleviate necroptosis to improve the survival of ACHBLF patients is interesting research direction.KLHL22 not only regulates mitosis, chromosome alignment31, and DNA double-strand break repair28, but also prevent extreme T cell inhibition32. KLHL22 regulates PD-1 homeostasis by facilitating the degradation of underglycosylated PD-1 prior to its translocation to the cell membrane, thereby preventing the excessive suppression of T cells and fostering antitumor immune responses32. A deficiency in KLHL22 leads to the overaccumulation of PD-1, which hampers T cell effector functions and accelerates tumor progression32. Importantly, KLHL22 levels are significantly decreased in tumor-infiltrating T cells from patients with colorectal cancer32. Those findings are consistent with our results indicating that KLHL22 is a signature gene associated with necroptosis and immune infiltration. RALY, a multifunctional RBP, contributes to several cellular processes, including cell proliferation33, autophagy34, mitochondrial metabolism, and tumorigenesis35. RALY promotes colorectal cancer metastasis through exosome-mediated M2 macrophage activation36. Increased RALY expression enhances metastasis, poor outcomes and stemness in hepatocellular carcinoma37. Moreover, the downregulation of RALY expression suppresses cellular proliferation and counteracts aggressive biological behavior in HCC cells38. RALY modulates the expression of genes involved in immunity and inflammatory responses by influencing the splicing of regulatory factors and alternative splicing. These findings underscore the need for further exploration into the role of RALY in cancer immunology and its contribution to clinical resistance in the progression of cancer under specific therapeutic interventions39. Our study provided a new molecular feature of RALY in liver diseases. The identified functions of these hub genes are concordant with their substantial correlation with immune infiltration and GO and KEGG enrichment analyses. Therefore, these 5 hub genes may facilitate the pathophysiology of ACHBLF and inspire novel therapeutic strategies targeting ACHBLF.
Given that immune cells play critical roles in the pathogenesis of ACHBLF, we investigated immune infiltration in ACHBLF patients and normal controls. We found that type 17 T helper cells, activated dendritic cells, plasmacytoid dendritic cells and monocytes were more enriched in ACHBLF patients than in normal controls, findings that are consistent with the results of a previous study40. ACHBLF is characterized by enhanced proinflammatory responses, in addition to the increased infiltration of Th17 cells and dendritic cells41. Th17 cells can exacerbate hepatic inflammation and tissue damage42, whereas dendritic cells are crucial for antigen presentation and immune activation43. This study explored the links between key necroptosis-related genes and specific immune cell subsets using integrated bioinformatics and machine learning, offering new insights into ACHBLF immunopathogenesis. The correlations between the hub genes and specific immune cell subtypes were analyzed. FCRL3 was negatively associated with plasmacytoid dendritic cells, monocytes and activated dendritic cells but positively associated with natural killer cells, activated CD8 T cells and type 2 T helper cells. CDC14A expression was negatively correlated with activated dendritic cells and plasmacytoid dendritic cells. KLHL22 expression was negatively correlated with activated dendritic cells. Many studies have revealed that multiple immune cell types, including monocytes, dendritic cells, CD4 + T cells44, Kupffer cells, natural killer cells and cytotoxic T lymphocytes45, are present in ACHBLF and that the cytokines produced by such cells contribute to ACHBLF progression and mortality. On the basis of these findings, we speculate that immune cells infiltration might participate in the progression of necroptosis to aggregate ACHBLF.
To explore the potential molecular mechanisms of the 5 hub genes in ACHBLF, an mRNA-miRNA-lncRNA interaction network was constructed, and the results revealed that the lncRNA XIST-miR-424-5p-CDC14A axis might be involved in ACHBLF. The crosstalk network of the lncRNA XIST-miR-424-5p exists in liver cancer46, pituitary gland tumors and meningioma47. We also investigated RNA-binding proteins and transcription factors, but the results need further experimental validation.
In this study, we utilized the CMAP database to identify candidate drugs for ACHBLF therapy targeted MAP4K1 and its closely related genes, identifying several potential candidate small-molecule compounds. The ten most promising targeted drugs identified were tiaprofenic-acid, guanabenz, imatinib, etacrynic-acid, brivanib, vincamine, amiloride, lypressin, pizotifen and leflunomide. However, these predicted drugs have not yet been approved for clinical treatment of liver diseases from the National Medical Products Administration, and direct evidence for their regulation of necroptosis or immune infiltration remains insufficient. Additionally, there is a lack of direct evidence regarding how these candidate drugs regulate necroptosis or immune infiltration in ACHBLF. Consequently, our findings lay a theoretical foundation for future experimental studies in this area. The potential mechanisms by which these drugs regulate necroptosis or immune infiltration need to be further explored. Additionally, molecular docking studies to assess the binding affinity of these candidate drugs with key proteins have not been conducted. Therefore, the drug predictions in this study are primarily theoretical explorations, and the related mechanisms require further experimental validation. In future research, we will prioritize the selection of target drugs that have been approved for the treatment of liver diseases. By integrating techniques such as molecular docking, we will systematically evaluate their binding capabilities and mechanisms of action with key genes. This approach aims to provide a more robust theoretical basis and potential therapeutic options for the precise treatment of ACHBLF.
Certain limitations exist in our research. First, our study concentrated on bioinformatics analysis derived from public databases; this approach might not precisely replicate the actual circumstances. Second, the study is based solely on the transcriptomic data of PBMCs from ACHBLF patients for systematic bioinformatics analysis. Although PBMCs can reflect the immune status of the body to some extent, they cannot fully mimic the key pathological processes in the liver tissue microenvironment, such as necroptosis and immune cell infiltration. This may lead to discrepancies between the expression and regulatory mechanisms of some core molecules and the actual situation in liver tissue. Moreover, due to the significant difficulty in obtaining liver tissue samples in clinical practice, the study did not integrate liver tissue data for cross-validation. In the future research, we will combine multi-omics data from liver tissue samples and use tissue-level experimental methods to conduct in-depth functional validation of the key molecules and regulatory mechanisms identified in this study, thereby further enhancing the scientific validity and clinical translational value of the research conclusions. Third, the lncRNA XIST-miR-424-5p-CDC14A interaction network has yet to be validated in vitro and in vivo. Finally, the top 10 drugs that target the 5 hub genes in ACHBLF are merely database prediction results and have not been experimentally validated, although they have been used in clinical practice on the basis of prescribing information. Owing to time and resource constraints, our future research will focus on the following areas. We will detect the expression levels of signature genes in separate patient groups using qRT-PCR and Western blot. Next, we will further investigate their functions in cell apoptosis and necrosis using in vitro models such as PBMCs or hepatocyte lines through gene knockdown or overexpression. Flow cytometry will be used to evaluate how gene regulation affects different subsets of immune cells. Animal models will also be employed to elucidate the physiological functions of key genes in the pathogenesis of ACHBLF. These approaches will help clarify the role of signature genes in ACHBLF and lay a foundation for future clinical translation.
Conclusion
In conclusion, we identified 5 hub genes that target necroptosis and immune infiltration in ACHBLF through integrated bioinformatics analysis and machine learning algorithms. Analyses of the hub genes were performed to identify biological process and signaling pathway enrichment, correlations with immune cell subtypes, mRNA-miRNA-lncRNA interaction networks, RNA binding proteins, transcriptional regulators and candidate drugs. These findings suggest that the 5 hub genes critically contribute to ACHBLF pathogenesis and represent promising molecular targets for developing precision therapeutic strategies.
Methods
Data collection
Datasets GSE 168,048 and GSE248217 were obtained from the National Center for Biotechnology Information Gene Expression Omnibus (NCBI-GEO) (https://www.ncbi.nlm.nih.gov/geo/) using the GEOquery package in R (version 4.1.2). GSE 168,048 comprises expression profiles of PBMC from 16 patients with ACHBLF, and GSE248217 includes the same information from 3 ACHBLF patients and 3 normal individuals. After merging GSE 168,048 and GSE248217, 19 patients with ACHBLF and 3 normal individuals included, the SVA software package was used to remove bath effects, and principal component analysis (PCA) was used to assess the effectiveness of the correction. Subsequently, a normalized gene expression profile containing 19 patients with ACHBLF and 3 normal individuals was analyzed. The names and sources of the chosen necroptosis-related genes (NRGs) were presented in Table S1. The analytical flowchart of our study is presented in Fig. 1.
Validation of hub gene expression using GEO datasets
To verify the expression levels of the hub genes (FCRL3, CDC14A, KLHL22, RALY, and MAP4K1), we integrated two datasets, GSE168048 and GSE248217, and applied batch-effect correction to create a standardized expression matrix. We extracted expression data for these hub genes from all samples, including ACHBLF and normal control samples. To compare the expression levels of the hub genes between the ACHBLF group and the normal control group, the Wilcoxon rank-sum test was used. The results were visualized using box plots generated with the R package ggplot2 (version 3.3.6), with statistical significance defined as p < 0.05. Additionally, we analyzed the correlations among the expression levels of the hub genes using Spearman correlation coefficients, and these correlations were visualized with the R package corrplot (version 4.1.2).
Identification of differentially expressed genes
The R package Limma (version 3.50.0) was used to identify differentially expressed genes (DEGs) between the ACHBLF and normal groups. The criteria for screening were |log2Fold Change| > 0.5 and P Value < 0.05.
Functional enrichment analysis of NRGs -related DEGs in ACHBLF
Weighted gene coexpression network analysis (WGCNA) was performed to construct a coexpression network using the R package WGCNA (version 1.70–3.70). An empirical power value of 9 was selected to establish a weighted adjacency matrix. The relationships among modules were visualized through hierarchical clustering dendrograms of eigengene networks and corresponding eigengene heatmaps. Necroptosis-related DEGs (NRGs-related DEGs) were obtained from the intersection of DEGs and the component genes within the NRGs-correlated modules.
Gene Ontology (GO) enrichment analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses were performed using the R package clusterProfiler (version 4.2.2), with a significance threshold of P < 0.05.
Gene Set Enrichment Analysis (GESA) was also performed using the NRG-related DEGs. The R package Limma (version 3.50.0) was used to perform differential expression analysis of key genes between the high- and low-expression groups, yielding the fold change (FC) in gene expression. GSEA was conducted using the R package clusterProfiler (version 4.2.2) on an ordered list of all genes ranked by log2FC. The c2.cp.kegg.v7.5.1.symbols gene set, which was curated in the Molecular Signatures Database (MSigDB) (https://www.gsea-msigdb.org/gsea/msigdb), was utilized as the reference gene set. P < 0.05 were considered significant.
Machine learning
Support vector machines-recursive feature elimination (SVM-RFE) is a machine learning algorithm that identifies the most predictive features from different categories.
A least absolute shrinkage and selection operator (LASSO) regression model was constructed to compute and select linear models and identify valuable variables. The R package randomForest (RF) was used to perform random forest analysis. On the basis of the mean decrease accuracy (MDA) and mean decrease gini (MDG) of feature weights, the top 30 genes were selected as the signature genes screened by random forest. By integrating LASSO regression, SVM-RFE, and RF, the most significant feature genes were ultimately identified as NRG- signature genes in this study.
Key genes interaction network
The GeneMANIA website (https://genemania.org/) can be used to predict functional genomic relationships between functionally similar genes, including protein-protein interactions, protein-DNA interactions, pathways, physiological and biochemical reactions, coexpression, and colocalization. In the present study, the GeneMANIA website was used to construct an interaction network of the hub genes.
Constructing of a lncRNA-RBP-mRNA-transcription factors network
Since the role of competing endogenous RNA (ceRNA) in the pathogenesis of ACHBLF is not clear, starbase2.0 (https://starbase.sysu.edu.cn/starbase2/index.php) and miRDB (https://mirdb.org/index.html) were used to construct a ceRNA network to identify the upstream regulators of microRNAs and the lncRNAs associated with the common microRNAs of prognostic genes.
The starBase online database (https://starbase.sysu.edu.cn/tutorialAPI.php#RBPTarget) was used to analyze the RNA-binding protein (RBP)-mRNA interactions. The significant RBPs-mRNAs was screened under the following conditions: clusterNum ≥ 10, clipExpNum ≥ 10 and P value < 0.05. An RBPs-mRNA network was subsequently constructed using Cytoscape software (version 3.9.1). Cytoscape is an open-source bioinformatics platform that allows users to visualize, analyze, and model molecular interaction networks and biological pathways; it provides a graphical representation of complex biological networks, such as protein-protein interactions and gene regulatory networks, with customizable node and edge attributes.
The necroptosis-related signature genes encoding transcription factors (TFs) were predicted using the hTFtarget database (http://bioinfo.life.hust.edu.cn/hTFtarget#!), which describes the TFs and direct target genes deprived from PBMCs in a human gene signature. Furthermore, an mRNA-TF regulatory network was constructed using Cytoscape software.
Immune infiltration analysis and correlation analysis of immune infiltration and NRG-signature genes
Data for 28 immune cell types were obtained from the TISIDB (http://cis.hku.hk/TISIDB/index.php). The correlations between the hub genes and the infiltration of 28 immune cell types were estimated by single-sample (ss) GSEA. The relative enrichment scores of each immune cell type were calculated on the basis of gene expression. The R package ggplot2 (version 3.3.6) was employed to display variations in immune infiltration between the ACHBLF and normal groups.
To explore the functions of the hub genes, 50 cancer hallmark gene sets were accessed from the Molecular Signature Database (MSigDB). ssGSEA was used to calculate the enrichment scores of the 50 cancer hallmark gene on the basis of the gene expression matrix from a standard transcriptomic dataset. The R package ggplot2 (version 3.3.6) was used to analyze the variations in the enrichment scores for the 50 cancer hallmark gene between the ACHBLF and normal groups. The correlations of enrichment scores between the NRG signature genes and 50 cancer hallmark gene were calculated via Spearman correlation analysis.
Identification of drug candidates
To identify the drug candidates for ACHBLF, Connectivity Map (https://www.broadinstitute.org/connectivity-map-cmap) analysis was conducted to analyze the upregulated expression of the hub genes (MAP4K1 and those genes highly correlated with MAP4K1). A negative enrichment score indicated that small-molecule drugs could reverse gene expression, thus demonstrating their potential therapeutic value.
Statistical analyses
All the statistical analyses were carried out with R studio software (version 4.1.2). Spearman’s correlation was used to calculate the relationships between two variables. The Wilcoxon test was used to calculate the difference between two paired groups; comparisons between three or more independent groups were conducted with the Kruskal-Wallis Test. All the statistical tests were two-sided, and a P value less than 0.05 was considered statistically significant.
Data availability
The GEO summary statistics used in this study were publicly accessed from GEO database (https://www.ncbi.nlm.nih.gov/geo/): GSE168048: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE168048;GSE248217: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE248217.
References
Sarin, S. K. et al. Acute-on-chronic liver failure: consensus recommendations of the Asian Pacific association for the study of the liver (APASL). Hep. Intl. 3, 269–282 (2008).
Sarin, S. K. et al. Acute-on-chronic liver failure: consensus recommendations of the Asian Pacific association for the study of the liver (APASL): an update. Hep. Intl. 13, 353–390 (2019).
Hibi, T., Wei Chieh, A. K., Chi-Yan Chan, A. & Bhangui, P. Current status of liver transplantation in Asia. Int. J. Surg. 82, 4–8 (2020).
Dara, L., Liu, Z-X. & Kaplowitz, N. Questions and controversies: the role of necroptosis in liver disease. Cell. Death Discovery. 2, 16089 (2016).
Suda, J. et al. Knockdown of RIPK1 markedly exacerbates murine immune-mediated liver injury through massive apoptosis of hepatocytes, independent of necroptosis and Inhibition of NF-κB. J. Immunol. 197, 3120–3129 (2016).
Afonso, M. B. et al. Necroptosis is a key pathogenic event in human and experimental murine models of non-alcoholic steatohepatitis. Clin. Sci. (Lond). 129, 721–739 (2015).
Li, X., Dong, G., Xiong, H. & Diao, H. A narrative review of the role of necroptosis in liver disease: a double-edged sword. Annals Translational Med. 9, 422–422 (2021).
Bai, L. et al. M2-like macrophages exert hepatoprotection in acute-on-chronic liver failure through inhibiting necroptosis-S100A9-necroinflammation axis. Cell Death Dis. 12, 93 (2021).
Kondo, T. et al. The role of RIPK1 mediated cell death in acute on chronic liver failure. Cell. Death Dis. 13, 5 (2021).
Chen, L. et al. Circulating Receptor-Interacting protein kinase 3 are increased in HBV patients with Acute-on-Chronic liver failure and are associated with clinical outcome. Front. Physiol. 11, 526 (2020).
Li, J. et al. PBMC transcriptomics identifies immune-metabolism disorder during the development of HBV-ACLF. Gut 71, 163–175 (2020).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53, D672–D677 (2025).
Kanehisa, M. Toward Understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951 (2019).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Kiefer, F., LA. T, M. A., BW. Z, A. J., L, N. & P, T. JR. W, NN. I: HPK1, a hematopoietic protein kinase activating the SAPK/JNK pathway. EMBO J. 15, 7013–7025 (1996).
He, T. S. et al. The kinase MAP4K1 inhibits cytosolic RNA-induced antiviral signaling by promoting proteasomal degradation of TBK1/IKKε. Microbiol. Spectr. 9, e0145821 (2021).
Ling, Q. et al. MAP4K1 functions as a tumor promotor and drug mediator for AML via modulation of DNA damage/repair system and MAPK pathway. eBioMedicine 69. (2021).
Sun, J-M. et al. Glioblastoma cellular MAP4K1 facilitates tumor growth and disrupts T effector cell infiltration. Life Sci. Alliance 6(12), e202301966 (2023).
Bai, Y. et al. PDIA6 modulates apoptosis and autophagy of non-small cell lung cancer cells via the MAP4K1/JNK signaling pathway. EBioMedicine 42, 311–325 (2019).
Lei, J. J. et al. Long noncoding RNA CDKN2B-AS1 interacts with transcription factor BCL11A to regulate progression of cerebral infarction through mediating MAP4K1 transcription. FASEB J. 33, 7037–7048 (2019).
Jiang, Y., Qiao, Y., He, D., Tian, A. & Li, Z. Adaptor protein HIP-55-mediated signalosome protects against ferroptosis in myocardial infarction. Cell. Death Differ. 30, 825–838 (2023).
Lin, J., Zhou, J. & Xu, Y. Potential drug targets for multiple sclerosis identified through Mendelian randomization analysis. Brain 146, 3364–3372 (2023).
Kraus, Z. J., Agarwal, S., Dement-Brown, J. & Tolnay, M. FCRL3 modulates the function of human thymic regulatory T cells. J. Immunol. 198, 8011–8011 (2017).
Bahabayi, A. et al. FCRL3 expression is upregulated and closely correlates with TIGIT expression in regulatory T cells of patients with systemic lupus erythematosus. Eur. J. Immunol. 54, e2350739 (2024).
Kim, S. et al. Dual-mode action of scalable, high-quality engineered stem cell-derived SIRPα-extracellular vesicles for treating acute liver failure. Nat. Commun. 16(1), 1903 (2025).
Shen, C. et al. Increased CD4 + CD25 + regulatory T cells correlate with poor short-term outcomes in hepatitis B virus-related acute-on-chronic liver failure patients. J. Microbiol. Immunol. Infect. 48, 137–146 (2015).
Jiang, P. et al. MicroRNA-146a-5p induces cell cycle arrest and enhances apoptosis in gastric cancer via targeting CDC14A. Frontiers Cell. Dev. Biol. 11, 1181628 (2023).
Lin, H. et al. Cdc14A and Cdc14B redundantly regulate DNA double-strand break repair. Mol. Cell. Biol. 35, 3657–3668 (2023).
Bembenek, J. & Yu, H. Regulation of the anaphase-promoting complex by the dual specificity phosphatase human Cdc14a. J. Biol. Chem. 276, 48237–48242 (2001).
Hoblos, H., Cawthorne, W., Samson, A. L. & Murphy, J. M. Protein shapeshifting in necroptotic cell death signaling. Trends Biochem. Sci. 50, 92–105 (2025).
Song, Y. et al. KLHL22 regulates the EMT and proliferation in colorectal cancer cells in part via the Wnt/β-catenin signaling pathway. Cancer Manage. Res. 12, 3981–3993 (2020).
Zhou, X. A. et al. KLHL22 maintains PD-1 homeostasis and prevents excessive T cell suppression. Proc. Natl. Acad. Sci. U S A. 117, 28239–28250 (2020).
Liu, S. et al. O-GlcNAcylated RALY contributes to hepatocellular carcinoma cells proliferation by regulating USP22 mRNA nuclear export. Int. J. Biol. Sci. 20, 3675–3690 (2024).
Qin, W. et al. Nuclear ribonucleoprotein RALY targets virus nucleocapsid protein and induces autophagy to restrict Porcine epidemic diarrhea virus replication. J. Biol. Chem. 298, 102190 (2022).
Hu, H. et al. The RNA binding protein RALY suppresses p53 activity and promotes lung tumorigenesis. Cell Rep. 42(4), 112288 (2023)
Zhou, J. et al. RNA binding protein RALY facilitates colorectal cancer metastasis via enhancing exosome biogenesis in m6A dependent manner. Int. J. Biol. Macromol. 273, 133112 (2024).
Wang, X. et al. RALYL increases hepatocellular carcinoma stemness by sustaining the mRNA stability of TGF-β2. Nat. Commun. 12, 1518 (2021).
Zhu, Z. et al. Overexpression of RALY promotes migration and predicts poor prognosis in hepatocellular carcinoma. Cancer Manag Res. 10, 5559–5572 (2018).
Liang, Z. et al. RALY regulate the proliferation and expression of immune/inflammatory response genes via alternative splicing of FOS. Genes Immun. 23, 246–254 (2022).
Wu, Z. et al. Comparative analysis of monocyte-derived dendritic cell phenotype and T cell stimulatory function in patients with acute-on-chronic liver failure with different clinical parameters. Front. Immunol. 14, 1290445 (2023).
Wang, L-Y. et al. Increased frequency of Circulating Th17 cells in acute-on-chronic hepatitis B liver failure. Dig. Dis. Sci. 57, 667–674 (2012).
Lafdil, F., Miller, A. M., Ki, S. H. & Gao, B. Th17 cells and their associated cytokines in liver diseases. Cell Mol. Immunol. 7, 250–254 (2010).
Hilligan, K. L. & Ronchese, F. Antigen presentation by dendritic cells and their instruction of CD4 + T helper cell responses. Cell Mol. Immunol. 17, 587–599 (2020).
Zhang, Y. et al. Downregulated VISTA enhances Th17 differentiation and aggravates inflammation in patients with acute-on-chronic liver failure. Hepatol. Int. 17, 1000–1015 (2023).
Zhang, T. et al. Disruption of the gut-liver axis in the pathogenesis of acute-on-chronic liver failure. Eur. J. Gastroenterol. Hepatol. 30, 130–135 (2018).
Ning, D. et al. The crosstalk network of XIST/miR-424-5p/OGT mediates RAF1 glycosylation and participates in the progression of liver cancer. Liver Int. 41, 1933–1944 (2021).
Ghafouri-Fard, S., Abak, A., Hussen, B., Taheri, M. & Sharifi, G. The emerging role of non-coding RNAs in pituitary gland tumors and meningioma. Cancers (Basel). 13, 5987 (2021).
Acknowledgements
We thank the participants who provided samples for the National Center for Biotechnology Information Gene Expression Omnibus (NCBI-GEO) datasets used in this study and the investigators who made the GEO data publicly available.
Funding
This project was supported by the Provincial Natural Science Foundation of Hunan (Grant Number 2025JJ70165, 2025JJ70166).
Author information
Authors and Affiliations
Contributions
Chuangjie Cao prepared Figs. 1, 2, 3, 4, 5, 6, 7, 8 and 9;Dan Luo prepared Figs. 10, 11, 12 and 13; Table 1;Xia Xie wrote the original draft. Chengyun Dou designed this trial and supervised and critically revised the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The GEO statistics used in this study are publicly available, and therefore, no specific ethics approval was needed.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cao, C., Luo, D., Xie, X. et al. Identification of key biomarkers associated with necroptosis and immune infiltration in hepatitis B virus-related acute-on-chronic liver failure. Sci Rep 15, 37440 (2025). https://doi.org/10.1038/s41598-025-21273-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-21273-2