Abstract
This study focused on evaluating the antioxidant properties of Octhochloa compressa, a Cholistani medicinal plant and to explore the interaction of its active phytochemical constituents with the biological macromolecule NADPH oxidase (PDB ID: 2CDU). Six different non-polar to polar solvent extracts were evaluated for their antioxidant activity using DPPH and FRAP assays. The key metabolites were then subjected to molecular docking against NADPH oxidase, density functional theory (DFT) and molecular dynamics (MD) simulations to validate the ligand-protein stability and conformational dynamics. The methanol extract revealed significantly highest scavenging potential (IC50 = 110 ± 2.34 µg/mL) in DPPH assay and displayed the greatest amount of total phenolic (97.34 ± 3.34 mg GAE/g) and flavonoid (98.72 ± 2.46 µg CE/g) contents. Phytochemical profiling of the methanol extract identified 43 major bioactive compounds by using HPLC and GC-MS. Silico molecular docking of selected metabolites (lignan, oxindole, apigenin, anthocyanin, acridone) with NADPH oxidase revealed stable interactions, with binding energies ranging from − 6.060 to -6.654 kcal/mol. DFT analysis revealed dipole moments of 1.36–7.52 D and HOMO–LUMO gaps of 3.15–11.74 eV with anthocyanin and apigenin, predicted as the most reactive ligands. MD simulations further confirmed these findings, as the representative 2CDU–ligand complex rapidly equilibrated and maintained the stability across 1 ns and 10 ns trajectories (RMSD ~ 0.2 nm, Rg 2.38–2.46 nm), with persistent non-covalent interactions (− 300 to − 400 kJ/mol) and well-defined free energy minima, validating favorable and thermo-dynamically stable binding. Collectively, these results suggest that O. compressa is a rich source of anti-oxidants phyto-metabolites, capable of interacting with NADPH oxidase, emphasizing its role in the management of disorders related to oxidative stress.
Similar content being viewed by others
Introduction
The oxidative stress arises when the production of reactive oxygen species (ROS) exceeds the anti-oxidant defense mechanism, leading to damage of key biomolecules including proteins, lipids and nucleic acids. The damage caused by oxidative stress plays a key role in the initiation and progression of chronic disorders like diabetes, aging, neurodegenerative and cardiovascular diseases and multiple carcinomas1,2. Accumulating evidence supports the contribution of plant-sourced antioxidants in mitigating oxidative damage, largely due to the existence of bioactive compounds with strong radical-scavenging properties3,4,5. Among the enzymatic sources of ROS, NADPH oxidase (NOX) is a major macromolecular complex responsible for a regulated ROS production across various cell types. Initially, characterized in the phagocytes for its antimicrobial function, NOX isoforms are now also known to be widely expressed in non-phagocytic cells like fibroblasts, tumor cells and vascular tissue cells, where they contribute to redox signaling and pathophysiological oxidative stress6. Dysregulated NOX activity contributes to the development of multiple chronic disorders, making it a molecular target of interest in redox biology. The natural antioxidants particularly polyphenols and flavonoids have demonstrated the ability to modulate or inhibit NOX-derived ROS generation, thus protecting the cellular components from oxidative injury7,8,9.
The exogenous anti-oxidants obtained from medicinal plants, fruits and vegetables play a complementary role to the endogenous antioxidant defense system. These plant-derived compounds like polyphenols, phenolic acids, tannins, terpenoids, flavonoids and alkaloids are well documented for their potential in neutralizing the production of free radicals, chelate pro-oxidant metals and to regulate anti-oxidant enzymes10,11,12. The antioxidant property of phenolic phyto-metabolites is largely attributed to their hydroxylated aromatic structures that confer high reactivity towards ROS and potential biological efficacy13,14,15. Hence, to investigate these bioactive compounds, HPLC and GC-MS are routinely employed to explore and characterize the chemical profile of extracted plants16,17. These methods allow for detailed qualitative and quantitative assessments of the plant, enabling the identification of compounds with potential therapeutic applications.
Octhochloa compressa (Poaceae), locally recognized as chimbar or ghoradhob, or hillu, is a long-lived grass, native to arid and semi-dry areas such as desert of Cholistan of Pakistan, as well as in Africa, Arabia and India18,19,20. Traditionally, greatly valued as a high-nutrition fodder for livestock, also used in ethnomedicine for treating kidney pain and inflammatory disorders in both animals and humans21. Pharmacological studies have demonstrated its in vitro anti-inflammatory effects and in vivo efficacy in carrageenan-induced rat paw edema model22, mediated by its terpenoid, flavonoid and polyphenolic constituents. Moreover, recent investigations have highlighted in vitro, in silico and in vivo anti-diabetic potential19 along with promising in vitro anti-cancer potential in HepG2 cell line and high thrombolytic activity23, further supporting the therapeutic versatility of the plant. Despite its well-documented traditional use and emerging ethno-pharmacological relevance, the structural characterization and mechanistic anti-oxidant basis of O. compressa still remained unexplored and extremely required in the scientific literature. To fill this gap, this study systematically evaluated the antioxidant potential of 6 different extracts; n-hexane, n-butanol, dichloromethane (DCM), ethyl acetate, methanol and aqueous of O. compressa. The methanol extract that exhibited the strongest antioxidant activity13,24, was investigated for the first time by using advanced structural elucidation techniques like HPLC and GC-MS, to explore and characterize its bioactive constituents. Additionally, computational molecular docking, density functional theory (DFT) calculations and molecular dynamics (MD) simulations were employed to explore the interaction of its bioactive compounds with oxidative stress-related macromolecules, particularly NADPH oxidase, thus, offering novel mechanistic insights into the antioxidant properties of the plant. These integrative approaches mark for the first comprehensive and extensive chemical and molecular investigation of O. compressa, underscoring its promising role as a natural source of antioxidants with great therapeutic effectiveness.
Materials and methods
Collection and extractions of the plant
The plant was obtained from the desert of Cholistan at Bahawalpur (Punjab, Pakistan) and carefully identified by Flora of Pakistan (www.efloras.org/index.aspx) and Online World Grass Flora database (https://www.kew.org/data/grasses-db/www/gen00423.htm), by following the institutional, national and international guidelines. For additional authenticity, standard herbarium methods were followed25 and was further identified and confirmed by Dr. Ghulam Sarwar (Taxonomist, Department of Botany, The Islamia University of Bahawalpur). The plant material was then deposited at the institutional herbarium at the Department of Botany, The Islamia University of Bahawalpur (voucher number: Oc-491; Supplementary Fig. 1). The collection of the plant material and the usage in the research interventions in different experiments were followed according to the national and international legislations and the study protocol was reviewed and granted the ethical approval by Institutional Ethical Review board, The Islamia University of Bahawalpur, Pakistan (18/2020/PAEC, Dated: 15-09-2020).
The entire plant was first rinsed, air dried under the shade and finally pulverized into powder. Approximately, 250gm of the dried plant powder was separately soaked in six different solvents: n-hexane, n-butanol, dichloromethane, ethyl acetate, methanol and aqueous solution. The mixtures were soaked for 72 h with periodic shaking, then filtration was done and extracts were concentrated by rotary evaporator and percentage yield was determined, as described earlier13,26.
Preliminary phytochemical analysis
The phytochemical analysis of each extract of O. compressa was conducted using established methods, as outlined earlier3.
Estimation of total phenolics and flavonoids contents
Total phenolic contents (TPC) of all extracts of O. compressa were quantified by using the Folin-Ciocalteu assay. Briefly, 1mL of respective plant extract was added in 5mL of freshly prepared Folin–Ciocalteureagent (diluted 10 fold with distilled water) and incubated for 10 min. Subsequently, 4mL of sodium bicarbonate solution was mixed and left for 30 min, then the absorbance was measured at 765 nm using a UV/vis spectrophotometer and expressed as milligrams of gallic acid equivalents (GAE) per gram of dry extract. For the measurement of total flavonoid contents (TFC), 5mL of aluminium trichloride (2% in methanol) was added to 5mL of each of the plant extract and mixed thoroughly. After 10 min, the absorbance was recorded at 415 nm and presented as milligrams of catechin equivalents (CE) per gram of each extract27.
HPLC quantification
HPLC analysis quantified the key polyphenolic compounds in the MetOH extract of O. compressa by following the previous standardized procedure13. Briefly, each extract was prepared by dissolving in a mobile phase of milliQ water with acetic acid (A) and acetonitrile with acetic acid (B) (93:7, v: v) with an addition of 20% DMSO. The sample, adjusted to 1 mg/250µL, was vortexed, sonicated and then injected (20µL) into an HPLC system equipped with a Waters of 600 solvent pump and detector of 2996PDA The separation was performed by using a column of C18 at 30 ± 1 °C, with UV/Vis detection across 200–500 nm. The gradient elution used a degassed water-acetonitrile mobile phase with acetic acid (3%). Finally, each target compound’s concentration was determined by use of calibration curve28. For reproducibility, authentic phenolic and heterocyclic compounds standards were procured from Sigma-Aldrich® and Merck (Darmstadt, Germany) with following product identifiers: gallic acid (CAS 149-91-7; product no. 8.42649), protocatechuic acid (CAS 99-50-3; product no. PHL89766), vanillic acid (CAS 121-34-6; product no. 94770), caffeic acid (CAS 331-39-5; product no. C0625), syringic acid (CAS 530-57-4; product no. S6881), p-coumaric acid (CAS 501-98-4; product no. C9008), ferulic acid (CAS 537-98-4; product no. Y0001013), 3,5-dimethoxy-4-hydroxycinnamic acid (CAS 4385-56-2; product no. 631310), flavanols (CAS 6203-18-5; product no. D4506), (+)-catechin (CAS 225937-10-0; product no. C1251), (–)-epicatechin (CAS 490-46-0; product no. 03940590), pyridines (CAS 110-86-1; product no. 33553), pyrroloindoles (CAS 13754-86-4; product no. 357839), quinazolines (CAS 2274-42-2; product no. 184365), quinolines (CAS 91-22-5; product no. 94517), oxindoles (CAS 59-48-3; product no. O9808), manzamines (CAS 13089-11-7; product no. 510246), indolizidines (CAS 72741-87-8; product no. 574775-M), bisindoles (CAS 603-76-9; product no. 193984), indoles (CAS 120-72-9; product no. PHR3524), imidazoles (CAS 288-32-4; product no. I2399), ergots (CAS 511-08-0; product no. PHL80440), ephedras (CAS 1124-11-4; product no. 95162), carbolines (CAS 78538-74-6; product no. E006) and acridones (CAS 1124-11-4; product no. 95162).
GC-MS analysis
GC-MS analysis of MetOH extract of O. compressa was conducted using a Perkin Elmer Clarus 500 GC system equipped with an auto-sampler (AOC-20i) and a mass spectrometer at Textile Testing Lab, Quaid-e-Azam Industrial State, Lahore (Pakistan). The analysis was carried out at 70 eV in an electron impact mode, with 99.999% Helium gas serving as the carrier gas, with a constant 1 mL/min flow rate. A fused silica capillary column (Elite-1) with specific dimensions (30 × 0.25 mm ID×1µMdf), composed entirely of Dimethyl polydiloxane was employed. The injection volume was set at 0.5 µl with a 10:1 split ratio, while the temperature of injector and ion-source was kept at 250 °C and 280 °C, respectively. The temperature of oven was allowed to rise from 110 °C (held isothermally for 2 min) at a rate of 10 °C/min until reached at 280 °C, where it was held for 9 min. Mass spectra were recorded at 70 eV, with a 0.5 s scan interval, covering fragments from 40 to 450 Da. The entire GC run was lasted for 40–45 min. The relative percentage amounts of individual compounds were determined by comparison of their average peak areas to the total areas by employing Turbo Mass Ver 5.2.1 software for chromatogram and mass spectra handling17.
DPPH assay
The antioxidant potential of each extract of O. compressa was assessed by measuring their capacity to neutralize 1,1-diphenyl-2-picrylhydrazyl (DPPH) free radicals assay29. In brief, 20µL of the respective plant extract was added to 96-wells microplate containing DPPH, incubated for 30 min at 37 °C in darkness, OD517 was measured and the Radical Scavenging Activity (RSA) was calculated.
Ferric reducing antioxidant power (FRAP) assay
The antioxidant potential of each extract of O. compressa was assessed by FRAP assay. Briefly, in a microtiter plate, each plant extract (20µL) was mixed with 0.2 M phosphate buffer (90µL) and 1% w/v potassium ferricyanide (30µL) were added in the wells, followed by incubation for 20 min at 50 °C. Afterwards, 10% w/v trichloroacetic acid (30µL) and 0.1% w/v ferric chloride (30µL) were added and the plate was left for 10 min at room temperature. The data were recorded at OD700 by using ascorbic acid as the positive standard reference30.
In Silico study of NADPH oxidase
Retrieval of the target protein and associated ligands
In silico analysis of enzyme-inhibitor interactions was performed using the Schrodinger software suite (https://www.schrodinger.com/)31. Three-dimensional structure of the target protein, NADPH oxidase (PDB ID: 2CDU) was retrieved from the Protein Data Bank, as described (PDB, www.rcsb.org;32. For this study, the 2D molecular structures of the ligands were retrieved from PubChem (https://pubchem.ncbi.nlm.nih.gov) specifically of lignan (PubChem ID: 261166), oxindole (PubChem ID: 321710), acridone (PubChem ID: 2015), apigenin (PubChem ID: 5280443), anthocyanin (PubChem ID: 145858), quinazoline (PubChem ID: 9210), xanthone (PubChem ID: 7020), artimisine (PubChem ID: 68827), quinoline (PubChem ID: 7047), quinolizidine (PubChem ID: 119036), imidazole (PubChem ID: 795), stilbene (PubChem ID: 638088) and gallic Acid (PubChem ID: 370).
Preparation of NOX protein and ligands
For accurate docking simulations in the Maestro platform of Schrödinger, the target protein was prepared through a series of essential steps32. First, unwanted amino acid residues and any co-crystallized ligands were removed. The protein’s protonation states were then optimized using Epik, water molecule orientations were evaluated and hydrogen bonding networks were assigned with PropKa. Further structural refinement was achieved through energy minimization using the OPLS-2005 force field. The 2D structures of the selected compounds were imported and processed using the LigPrep module, where they were optimized geometrically at the relevant pH, taking into account the potential ionization and tautomeric forms. This streamlined preparation process laid the groundwork for precise and reliable docking simulations, providing a robust basis for subsequent analyses.
Molecular docking
To enable a precise docking, a receptor grid was generated, specifically tailored to the catalytic site of the NADPH oxidase protein. This grid construction was guided by pertinent literature, resulting in the creation of a grid box with dimensions measuring 14 × 15 × 19. The coordinates for the center of the grid were meticulously set as x = 18.9974, y = −5.774 and z = −1.808. Following the grid preparation, the ligands were flexibly docked onto this grid using Schrödinger’s Glide module that was operated in the standard precision (SP) mode. Each ligand was underwent 32 runs as part of the docking process. A detailed evaluation of the Glide outcomes, including qualitative and quantitative assessments was then performed, with a primary focus on examining the binding interactions. The Docking results were expressed as SP GScore in kcal/mol31,33.
Density functional theory (DFT) analysis
DFT computations were performed with Gaussian 09 to investigate the electronic properties and reactive behavior of the chosen ligands34. All ligands were first geometry-modified at the B3LYP/6-31G(d, p) theory level to achieve stable structures without any imaginary frequencies. Frontier molecular orbital analysis was then performed to identify the lowest unoccupied molecular orbital (LUMO), highest occupied molecular orbital (HOMO) and the corresponding energy gap (ΔE = E_LUMO − E_HOMO) that provides insights into the electronic reactivity and stability. Additionally, global reactivity parameters such as ionization potential (IP), electronegativity (χ), electron affinity (EA) and chemical hardness (η) were computed by following Koopmans’ theorem. These descriptors were then correlated with molecular docking outcomes to elucidate the electronic factors influencing the ligand binding to NADPH oxidase.
Molecular dynamics (MD) simulations
MD simulations was employed in GROMACS 2023.3 using the top-ranked NADPH oxidase–lignan complex (PDB ID: 2CDU), parameterized with CHARMM36 for protein and ACPYPE for ligand, solvated in a cubic TIP3P water box (1.0 nm margin) with Na⁺/Cl⁻ counter ions for neutralization. Energy minimization was carried out using the steepest descent method until the maximum force dropped below 1000 kJ/mol/nm. The system was equilibrated in two steps: first, 100 ps in the NVT ensemble at 300 K employing a velocity-rescaling thermostat, followed by100 ps in the NPT ensemble at 1 bar using the Parrinello–Rahman method for pressure equilibration, with positional restraints applied to the protein heavy atoms. An initial 1 ns MD production run was conducted, to evaluate the preliminary equilibration and stability of the complex. This was followed by a full 10 ns production run was achieved, under the conditions of periodic boundary, keeping the temperature at 300 K and at 1 bar of pressure. Bond constraints were enforced using the LINCS algorithm, while long-range electrostatic interactions were handled with the particle mesh Ewald (PME) procedure, applying a 1.0 nm cutoff for both interactions of van der Waals and Coulombic forces35.
Trajectory analyses were performed using GROMACS tools along with custom Python scripts. Structural stability was evaluated through root-mean-square deviation (RMSD), while the radius of gyration (Rg) was employed to assess the protein compactness. Residue-level flexibility was examined using root-mean-square fluctuation (RMSF) and hydrogen bond analysis quantified the interactions between the protein and ligand. Interaction energies including electrostatic and van der Waals components, were extracted from the simulations. Finally, free energy landscape (FEL) analysis along the first two principal components (PC1 and PC2) was performed to identify the most stable conformational basins of the 2CDU–lignan complex36.
Statistical analysis
The study employed one-way ANOVA followed by Dunnet’s t-test. The reported values represent the mean of three replicates with standard error of mean (SEM). The concentrations for 50% inhibition (IC50) of the extracts were determined using linear regression analysis37. The level of significance was set at p < 0.05.
Results
Phytochemical analysis and Estimation of total contents
The phytochemical analysis of O. compressa crude extracts revealed that alkaloids and flavonoids were abundantly detected in the Aqu, MetOH, EtAc and DCM extracts of O. compressa (Table 1), wherein, anthocyanin and betacyanin were exclusively found in the EtAc extract, while, most of the phytochemicals like phenols, steroids and steroids were present in all the extracts. Notably, cardiac glycosides, coumarins, phlobatannins and quinines were more abundant in Aqu, MetOH, EtAc and DCM extracts. Whereas, the saponins were predominantly identified in MetOH and EtAc extracts. Regarding the extraction yield of O. compressa, the highest percentage of extract yield was achieved in the Aqu extract (7.16%), followed by EtAc (6.10%) and MetOH (5.80%) extracts. Conversely, the lowest extract yield was observed in n-Hex (4.01%) extract (Table 2).
The phenolic contents in O. compressa extracts ranged from 58.89 ± 5.67 to 97.34 ± 3.34 GAE mg/g of each plant extract. The highest TPC were observed in the MetOH (97.34 ± 3.34 GAE mg/g), DCM (95.23 ± 1.98GAE mg/g), EtAc (89.32 ± 4.56 GAE mg/g) and Aqu (73.43 ± 2.32 GAE mg/g) extracts. However, in TFC, the MetOH extract exhibited the highest value (98.72 ± 2.46 µg/CE) of catechin equivalents per gram of plant, followed by EtAc (95.27 ± 1.96 µg), DCM (91.44 ± 1.62 µg) and Aqu (65.45 ± 1.84 µg) extracts. The lowest TFC were recorded in the n-Hex extract (18.39 ± 2.27 µg/CE) (Table 2).
HPLC analysis
Phenolics and flavonols were quantified in the extracts by comparing their retention time and spectral properties with those of 25 standard analytes (Fig. 1). The MetOH extract of O. compressa revealed the presence of 14 compounds including stilbene (9.26 µg/mg), flavonoid/anthocyanin (9.15 µg/mg), imidazole (8.55 µg/mg), lignan (7.27 µg/mg), quinazoline (7.15 µg/mg), oxindole (6.06 µg/mg), quinolizidine (5.01 µg/mg), xanthone (4.99 µg/mg), quinoline (4.49 µg/mg), apigenin (4.29 µg/mg), tannins (3.86 µg/mg), artemisinin (2.72 µg/mg), acridone (2.47 µg/mg) and gallic acid (2.07 µg/mg) (Table 3; Figs. 2 and 3).
HPLC chromatograph of 25 standard compounds.
HPLC chromatograph of methanol extract of O. compressa. The results revealed the presence of 14 compounds including gallic acid (2.07 µg/mg), stilbene (9.26 µg/mg), flavonoid/anthocyanin (9.15 µg/mg), imidazole (8.55 µg/mg), lignan (7.27 µg/mg), quinazoline (7.15 µg/mg), quinoline (4.49 µg/mg), oxindole (6.06 µg/mg), quinolizidine (5.01 µg/mg), xanthone (4.99 µg/mg), apigenin (4.29 µg/mg), tannins (3.86 µg/mg), artemisinin (2.72 µg/mg) and acridone (2.47 µg/mg).
The structures of 14 compounds identified in the MetOH extract of O. compressaby HPLC.
Characterization by GC-MS
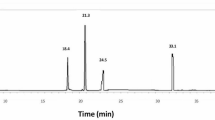
The GC-MS characterization of MetOH extract of O. compressa identified 29 compounds with retention time ranging from 5.095 to 33.866 min (Table 4; Figs. 4 and 5), including methyl palmitate (14.87 µg/mg), palmitic acid (14.16 µg/mg), methyl stearate (11.77 µg/mg), bis(2-ethylhexyl) phthalate (8.17 µg/mg), phenol,2,2’-methylenebis[6-(1,1-dimethylethyl)−4-methyl-(7.48 µg/mg), 2-methoxy-4-vinylphenol (4.52 µg/mg), stearic acid (4.14 µg/mg), dioctyl terephthalate (3.85 µg/mg), 9,12,15-octadecatrienoic acid methyl ester (3.12 µg/mg), 1,3,5-triphenylcyclohexane (3.04 µg/mg), methacrylic acid (2.42 µg/mg), dimethylamine (2.36 µg/mg), 2-palmitoylglycerol (1.87 µg/mg), isoeugenol (2.30 µg/mg), 3-piperidinemethanol, 1-methyl-(1.67 µg/mg), alloxan (1.52 µg/mg), syringol (1.50 µg/mg), pentasiloxane (1.37 µg/mg), cyclohexasiloxane (1.35 µg/mg), glycine (1.12 µg/mg), 9,12-octadecadienoic acid methyl ester (1.06 µg/mg), o-toluidine (1.03 µg/mg), gentisic acid (1.03 µg/mg), 2,4-di-tert-butylphenol (0.91 µg/mg), piperazine (0.90 µg/mg), cyclohexene 2-butyl-1,3,3-trimethyl-(0.70 µg/mg), octanamide, n,n dimethyl-(0.68 µg/mg), 3-amino isoxazole (0.63 µg/mg) and butylated hydroxytoluene (0.48 µg/mg).
GC-MS chromatograph of MetOH extract of O. compressa. GCMS identified 29 compounds with a retention time ranging from 5.095 to 33.866 min, including methyl palmitate (14.87 µg/mg), palmitic acid (14.16 µg/mg), methyl stearate (11.77 µg/mg), bis(2-ethylhexyl) phthalate (8.17 µg/mg), phenol, 2,2’-methylenebis [6-(1,1-dimethylethyl)−4-methyl- (7.48 µg/mg), 2-methoxy-4-vinylphenol (4.52 µg/mg), stearic acid (4.14 µg/mg), dioctyl terephthalate (3.85 µg/mg), 9, 12, 15-octadecatrienoic acid methyl ester (3.12 µg/mg), 1,3,5-triphenylcyclohexane (3.04 µg/mg), methacrylic acid (2.42 µg/mg), dimethylamine (2.36 µg/mg), 2-palmitoylglycerol (1.87 µg/mg), isoeugenol (2.30 µg/mg), 3-piperidinemethanol, 1-methyl-(1.67 µg/mg), alloxan (1.52 µg/mg), syringol (1.50 µg/mg), pentasiloxane (1.37 µg/mg), cyclohexasiloxane (1.35 µg/mg), glycine (1.12 µg/mg), 9,12-octadecadienoic acid methyl ester (1.06 µg/mg), o-toluidine (1.03 µg/mg), gentisic acid (1.03 µg/mg), 2, 4-di-tert-butylphenol (0.91 µg/mg), piperazine (0.90 µg/mg), cyclohexene 2-butyl-1 3 3-trimethyl (0.70 µg/mg), octanamide, n,n dimethyl- (0.68 µg/mg), 3-amino isoxazole (0.63 µg/mg), and butylated hydroxytoluene (0.48 µg/mg).
The structures of 29 compounds identified in the MetOH extract of O. compressaby GC-MS.
DPPH assay
All extracts of O. compressa exhibited varying degrees of antioxidant potential in a dose-dependent manner (Supplementary Fig. 2 A), with IC50 values ranging from 110 ± 2.34 µg/mL to 937 ± 1.21 µg/mL. Notably, the MetOH extract of O. compressa demonstrated strongest radical scavenging potential with IC50 value of 110 ± 2.34 µg/mL, followed by EtAc and DCM extracts of O. compressa with IC50 values of 155 ± 2.29 µg/mL and 198 ± 2.89 µg/mL, respectively. Conversely, the Aqu and n-Hex extracts showed the highest IC50 values with lowest percentage of RSA. In summary, the radical scavenging activity of O. compressa extracts followed the trend: MetOH > EtAc > DCM > n-But > n-Hex > Aqu (Table 5, Supplementary Fig. 3).
FRAP assay
In the FRAP assay, the MetOH and DCM extracts exhibited remarkably strong reducing power (Supplementary Fig. 2B), with lowest IC50 values < 42 µg/mL, followed by EtAc extract, achieved by donating the electrons to ferric cyanide. While, n-But and Aqu extracts showed comparatively moderate reducing power and exhibited considerable higher activity at increased concentrations, with high IC50 values (Fig. 6).
The relative percentage of FRAP assay (A) and IC50 (B) of O. compressa (Oc) compared to ascorbic acid (positive control). A: methanol (MetOH) and dichloromethane (DCM) extracts have shown strongest FRAP (%), B: least IC50 was observed by extract MetOH and DCM extracts. P.C: Positive control, Aqu: aqueous extract, n-But: n-butanol extract, EtAc: ethyl acetate extract, n-Hex: n-Hexane extract extract.
In Silico molecular Docking
According to the Ramachandran plot, 96.5% of residues in NADPH oxidase were positioned in the favorable regions, supporting the correctness of φ and ψ dihedral angles of the protein model (Fig. 7). Amongst, all compounds of MetOH extract, lignan, oxindole, acridone, apigenin and anthocyanin formed energetically favorable NOX-ligand complex with docking scores of −6.949, −6.683, −6.521, −6.298 and − 6.079 kcal/mol, respectively (Table 6). A comparative structure-activity relationships (SAR) offering critical insights, wherein lignan exhibited stable interactions of hydrogen bonding with specific residue Ala B:300 within the active site of the target enzyme (2CDU) (Fig. 8A). Similarly, oxindole showed stable hydrogen bonding interactions involving the NH group with Cys133 (Fig. 8B). However, acridone displayed hydrogen bond interaction of NH group with residue Thr112 (Fig. 8C). Additionally, apigenin demonstrated substantial hydrophobic interactions and stable interactions of hydrogen bonding between oxygen and hydroxyl group with Asp282 and Thr112 (Fig. 8D). Anthocyanin exhibited π–π interactions with the aromatic side chain of Phe245 (Fig. 8E). The existence of nitro and dihydroxy groups was identified as a key determinant for stable binding interactions within the active site of NADPH oxidase.
Structural validation and binding pocket analysis of NOX (2CDU). A: Ramachandran plot showing 96.5% of residues in favored regions, confirming proper φ and ψ dihedral geometry of the protein model, B: predicted binding pocket of 2CDU with docked ligand poses, C: corresponding molecular interaction view.
Comparative binding poses (left side) and molecular interaction views (right side) of NOX enzyme (2CDU) with selected MetOH extract ligands. A: Lignan–Ala300, B: Oxindole–Cys133, C: Acridone–Thr112, D: Apigenin–Asp282/Thr112, E: Anthocyanin–Phe245. Stable hydrogen bonding and π–π interactions contributed to favorable binding affinities (− 6.949 to − 6.079 kcal/mol). Nitro and dihydroxy groups were identified as key determinants of ligand stability within the active site.
DFT calculations
DFT analysis was performed on the top-scoring ligands identified from molecular docking to characterize their electronic structures and reactivity profiles (Table 7; Fig. 9). Dipole moment values varied widely, from 1.36 D in anthocyanin to 7.52 D in apigenin, reflecting differences in molecular polarity that may affect the solubility and hydrogen-bonding capacity. Frontier molecular orbital analysis revealed the energy gaps (ΔE_gap) ranging from 3.15 eV (anthocyanin) to 11.74 eV (oxindole), indicating that anthocyanin is the most chemically reactive, whereas oxindole is the most electronically stable. Ionization potential (IP) and electron affinity (EA) further supported these observations: lignan showed moderate donor–acceptor capacity (IP 5.15 eV; EA 1.32 eV), while oxindole exhibited the highest IP (8.34 eV) and a negative EA (− 3.39 eV), suggesting a strong resistance to electron removal and poor electron-accepting ability. Global reactivity descriptors highlighted the clear distinctions in ligand behavior. Hardness (η) was highest for oxindole (5.87 eV), confirming its low reactivity, whereas anthocyanin displayed the lowest value (1.57 eV), indicating high softness and enhanced electron cloud polarizability. Electrophilicity indices further identified that acridone (3.26 eV) and apigenin (2.97 eV) are strong electrophiles having a high propensity for interaction with nucleophilic protein residues. Conversely, oxindole had the lowest electrophilicity (0.52 eV), consistent with its electronic inertness.
DFT analysis of five high scoring docked compounds. The electronic structure and reactivity parameters of lignan, oxindole, acridone, apigenin and anthocyanin were evaluated through frontier molecular orbital (FMO) analysis. Variations in HOMO–LUMO energy gaps, dipole moments and global reactivity descriptors revealed distinct chemical behaviors—anthocyanin showed the highest reactivity (lowest ΔE_gap), while oxindole exhibited maximum stability (highest ΔE_gap). Acridone and apigenin demonstrated strong electrophilic characteristics, suggesting a high potential for interaction with nucleophilic sites of the NOX enzyme.
MD simulations
The molecular dynamics simulations of 2CDU–lignan complex demonstrated consistent stability and favorable binding interactions over both 1 ns and 10 ns trajectories. During 1 ns simulation, the protein backbone RMSD rose rapidly within the first 0.1–0.2 ns to ~ 0.20–0.23 nm, after which it stabilized without any upward drift, indicating an equilibration of the system. Over the longer 10 ns trajectory, the RMSD continued to remain around ~ 0.2 nm, confirming the structural stability and absence of large-scale conformational deviations (Fig. 10). Flexibility analysis through RMSF revealed that most of the residues exhibited low fluctuations (~ 0.05–0.2 nm), consistent with a stable folded protein core, while flexible peaks reaching ~ 0.4–0.45 nm were observed in loop and terminal regions (Fig. 11A-B). The radius of gyration fluctuated narrowly between 2.38 and 2.46 nm in both short and extended runs, with balanced RgX, RgY and RgZ values, showing that the protein retained the compactness without unfolding (Fig. 11C-D).
RMSD profiles of NOX (2CDU) and its complex with lignan during molecular dynamics simulations. RMSD of the unbound NOX protein at 1 ns (A) and 10 ns (B). RMSD of the 2CDU–lignan complex at 1 ns (C) and 10 ns (D). The complex exhibited minimal structural deviation (~ 0.2 nm) throughout the simulation, indicating stable binding and overall conformational equilibrium.
RMSF profile of NOX (2CDU) protein. RMSF analysis revealed low residue fluctuations (~ 0.05–0.2 nm), indicating a stable protein core, with higher flexibility (~ 0.4–0.45 nm) confined to loop and terminal regions, confirming structural stability across both simulations at 1 ns (A) and 10 ns (B). Radius of gyration profile of NOX (2CDU) protein at 1 ns (C) and 10 ns (D). The Rg values fluctuated slightly between 2.38–2.46 nm throughout both simulations, with consistent RgX, RgY and RgZ components. These results indicate that the protein maintained structural compactness and stability without any signs of unfolding during the simulation.
This pattern was maintained across both 1 ns and 10 ns, indicating localized flexibility without destabilizing the overall structure. Hydrogen bonding remained dynamic but persistent, with 2–4 stable protein–ligand hydrogen bonds and occasional peaks of 5–6, supported by 6–10 donor–acceptor pairs within 0.35 nm. Interaction energies confirmed a favorable binding, with Lennard-Jones short-range energies between − 250 and − 320 kJ/mol, Coulombic short-range contributions from − 70 to − 110 kJ/mol and a total interaction energy of − 300 to − 400 kJ/mol throughout both simulations (Fig. 12A-B). Interaction energy decomposition confirmed a favorable binding at both 1 ns and 10 ns. Lennard-Jones short-range energies remained between − 250 and − 320 kJ/mol, while Coulombic short-range contributions fluctuated between − 70 and − 110 kJ/mol. The total interaction energy consistently ranged from − 300 to − 400 kJ/mol, highlighting the dominant role of van der Waals forces, with electrostatic interactions providing additional stabilization (Fig. 12C-D).
Hydrogen bond analysis of NOX (2CDU) protein at 1 ns (A) and 10 ns (B). Dynamic yet stable hydrogen bonding profile showing 2–4 consistent protein–ligand hydrogen bonds (peaking at 5–6) and 6–10 donor–acceptor interactions within 0.35 nm. Interaction energy profiles of NOX (2CDU)-lignan complex at 1 ns (C) and 10 ns (D). Lennard-Jones (− 250 to − 320 kJ/mol) and Coulombic (− 70 to − 110 kJ/mol) short-range energies yielded total interaction energy of − 300 to − 400 kJ/mol, confirming stable and favorable binding, primarily driven by van der Waals forces with supportive electrostatic interactions.
The free energy landscape (FEL), derived from principal component analysis, revealed multiple well-defined low-energy basins across the 10 ns trajectory, representing stable thermodynamic states of the protein–ligand complex. The persistence of these minima confirmed a robust binding mode and supported the reliability of docking predictions. Overall, the system setup, where the 2CDU protein was solvated in an explicit water box (TIP3P model) and neutralized with Na⁺/Cl⁻ ions, ensured a physiologically relevant simulation environment, while the FEL analysis confirmed the structural and energetic stability across both 1 ns and 10 ns simulations (Fig. 13).
System setup for MD simulation of 2CDU–lignan complex (A) and free energy landscape (FEL) representation derived from the simulation (B). MD simulation showing the 2CDU protein solvated in a TIP3P water box and neutralized with Na⁺/Cl⁻ ions. FEL derived from principal component analysis displaying multiple low-energy basins across 10 ns trajectory, indicating stable thermodynamic states and confirming a robust and energetically favorable protein–ligand binding mode.
Discussion
Medicinal plants serve as an abundant reservoir of bioactive phytochemicals that determine their great therapeutic potential26,38,39,40. The choice of solvent in extraction critically influences both the yield and profile of these compounds1. As likely in case of O. compressa, varying extraction yields were obtained from eleven South African medicinal plants, with the highest yield obtained in the aqueous extract, followed by DCM and MetOH extracts11. In the current study, the alkaloids and flavonoids like tannins, terpenes and saponins were maximally identified in the MetOH extract of the plant, followed by Aqu, EtAc and DCM extracts. These specific bioactive compounds are well-known due to their role in anti-oxidant, analgesic, anti-inflammatory and hyperglycemic effects, thus highlighting the medicinal significance of the plants in treating the infections, hyperglycemia, fever, pain and inflammation41,42,43,44. Specifically, terpenoids, flavonoids and phenolics contribute significantly to the antioxidant properties45, while anthocyanin, betacyanin, cardiac glycosides, coumarins, quinines and saponins further underscores the potential medicinal value of the plant8. Consistently with our observations, the methanolic extract displayed the highest TPC and TFC, in agreement with reports on Echium humile28, Beilschmiedia roxburghiana stem barks3 and Arabis carduchorum46. These polyphenols are associated with anti-oxidant, anti-inflammatory, antimicrobial, antipyretic and anti-diabetic activities, reinforcing their pharmaceutical significance27. Oxidative stress induced by free radicals lead to cellular and tissue damage, promoting inflammation, fever, liver damage, carcinogenesis, hematological and histological alterations15,47,48 Endogenous anti-oxidant enzymes like catalase, glutathione peroxidase and superoxide dismutase counteract these inflammatory reactions2,49,50, while plant-derived anti-oxidants further enhance this defense by scavenging the reactive oxygen species51.
Polyphenols play a key role in radical scavenging due to their redox-active hydroxyl groups, a principle exploited in the DPPH activity, where DPPH radical reduction causes a colorimetric shift from purple to yellow46. In this study, the MetOH extract exhibited the strongest scavenging potential (IC50 of 110 µg/mL), followed by EtAc (155 µg/mL) and DCM (198 µg/mL) extracts, reflecting the abundance of hydroxyl-rich-polyphenols and flavonoids52. Similarly, antioxidant and anti-inflammatory potential has also been reported for Severinia buxifolia53, Justicia secunda54 and Anthurium plowmanii leaves55, supporting the relevance of polyphenolic compounds in mediating their bioactivities. One of our recent studies on another Cholistani desert plant (Neurada procumbens) also demonstrated a highest anti-oxidant potential of the plant due to its high phenolic contents13,24. Moreover, the methanolic extract of O. compressa exhibited the strongest FRAP among all extracts, consistent with its abundant TPC and TFC29,56,57 Similarly, Cypria and Cynara scolymus L. extracts rich in polyphenols and flavonoids were found to exhibit superior reducing potential58. The EtAc extract of Cynara scolymus L. also exhibited the highest reducing activity on in vitro and in vivo evaluations, due to high TPC and TFC7. These results also align with prior reports on Sideritis cypria, where extracts rich in polyphenols and flavonoids exhibited superior reducing potential59. Similarly, antioxidant evaluations of Borassus flabellifer L. extract, highlighted that phenolic acids and flavonoids, including quercetin, coumarin and gallic acid elucidate robust radical-scavenging and reducing activities12.
Due to strongest anti-oxidant potential of O. compressa, the methanol extract was selected for detailed characterization and structural elucidation of phenolic compounds by HPLC and GC-MS analysis. HPLC profiling of the plant performed against 25 standard analytes, enabled the identification of 14 compounds including stilbene, flavonoid/anthocyanin, lignan, quinazoline along with various concentrations of oxindole, quinolizidine, xanthone, quinoline, apigenin, tannins, artemisinin, acridone and gallic acid. In the same context, an extensive HPLC analysis of Cordia diffusa leaves revealed 8 phenolic compounds with potent DPPH radical scavenging activity60 and Trifolium repens L. leaves, wherein, 29 compounds were quantified in methanol extract, highlighting their potential as a natural anti-oxidant and anti-cholinesterase therapeutics16. Additionally, GC-MS profiling of MetOH extract of O. compressa identified 29 bioactive volatile and semi-volatile compounds with a retention time ranging from 5.095 to 33.866 min, several of which are strongly associated with anti-oxidant activity. Fatty acid esters like hexadecanoic acid methyl ester, diterpenoids such as phytol possess hydroxyl or unsaturated moieties are capable of donating electrons or stabilizing peroxyl radicals, thereby mitigating the oxidative stress. Such mechanisms have been experimentally validated in the previous reports linking these metabolites with significant DPPH and ABTS radical scavenging activities61. Consistent with this, GC–MS analyses of other medicinal plants, such as methanolic extract of Gardenia jasminoides revealed diverse phytochemical profiles with 54 bioactive compounds, demonstrating the anti-oxidant capacity of the plant17. While, MetOH and Aqu extracts of Moringa oleifera leaves and Catharanthus roseus identified 40 and 65 compounds respectively in GC-MS, with strong and significant anti-oxidant and anti-bacterial effects against three human pathogens (E. coli, B. subtilis, S. aureus)14.
The phenolic derivatives and sterols further reinforce the anti-oxidant response through hydrogen atom donation, metal chelation and modulation of endogenous defense systems, in line with prior phytochemical studies62. Alongside these metabolites, the peaks corresponding to siloxane derivatives like pentasiloxane and cyclohexasiloxane were also detected, although such signals are frequently attributed to column bleed or PDMS contamination63, however, the occurrence of silicon compounds in plants remains a subject of active debate64. Silicon is known to accumulate as phytoliths, soluble silica or organosilicon complexes, contributing to stress tolerance ability of plant65. Importantly, recent evidence has shown that GC-MS can successfully detect silicon-containing constituents in coffee beans, supporting the interpretation that the silicon-related peaks identified in O. compressa represent authentic metabolites rather than analytical artefacts64,66. Although confirmation with techniques such as Inductively Coupled Plasma–Mass Spectrometry (ICP–MS) or Scanning Electron Microscopy–Energy Dispersive X-ray Spectroscopy (SEM–EDS) is still required67, these findings highlight the growing recognition that silicon chemistry constitutes a genuine and underexplored aspect of plant metabolomes.
Importantly, in the computational study, molecular docking were employed to determine the interactions between the identified phenolic compounds with NADPH oxidase (NOX; PDB ID: 2CDU), a key enzyme complex involved in ROS generation and oxidative stress-mediated cellular injury. Among the tested phytochemicals, lignan, oxindole, acridone, apigenin and anthocyanin showed strong binding affinities (docking scores: −6.654 to − 6.060 kcal/mol), indicating their potential to inhibit the activity of NADPH oxidase. These computational predictions align with our recent in vitro and in vivo studies demonstrating significant anti-inflammatory activity of O. compressa19, as well as previous reports highlighting NOX-targeted antioxidant and cytoprotective activities of related phytochemicals in Piper cubeba extracts in the cardiovascular disease68. Similarly, acridine and apigenin also exhibited significant potential for free radical suppression through strong interactions with NADPH oxidase69,70. Notably, acridone and apigenin have been reported to mitigate the oxidative stress and cytotoxicity in models of aluminum phosphide–induced cardiomyocyte injury, highlighting their cardio-protective and anti-oxidant properties71. Moreover, anthocyanin demonstrated superior NOX inhibition, surpassing ascorbic acid and exhibiting DPPH radical-scavenging activity greater than that of vitamin E, consistent with its potent antioxidant capacity72,73. Additionally, lignan and oxindole effectively attenuate the oxidative stress across multiple models, contributing to reduced risks of atherosclerosis, diabetes, cancer and cardiovascular disease74.
The DFT calculations provided a mechanistic rationale for the docking outcomes by linking the ligand electronic properties to their predicted binding behavior. The markedly low HOMO–LUMO gaps of anthocyanin (3.15 eV) and apigenin (4.53 eV) reflect high chemical softness and facile charge transfer, characteristics that enable rapid electronic redistribution upon interaction with protein residues75. This property is particularly important in antioxidant ligands, as smaller energy gaps are associated with enhanced radical-scavenging ability and stabilization of electron-rich environment76,77. The presence of extended π-conjugation in anthocyanin and polyphenolic substitutions in apigenin not only reduces their ΔE < sub > gap</sub > but also increases the electrophilicity, thereby facilitating π–π stacking, hydrogen bonding and charge-transfer interactions within the catalytic pocket78,79. These descriptors rationalize their strong docking affinities and consistent stabilization during MD simulations, underscoring their suitability as reactive inhibitors of NADPH oxidase. In contrast, oxindole displayed a wide HOMO–LUMO gap (11.74 eV), negative electron affinity and the highest hardness (5.87 eV), electronic features that confer the stability and diminish reactivity80. The indole framework is electronically rigid, limiting delocalization and making the scaffold resistant to the charge perturbations. Although oxindole was structurally compatible with the binding site, its low electronic flexibility reduces the likelihood of dynamic charge-transfer interactions with active-site residues, explaining its relatively weaker docking interactions compared to other polyphenolic ligands81. Lignan and acridone, with intermediate HOMO–LUMO gaps, represent a balance between stability and reactivity, thus, these descriptors suggest moderate softness, allowing selective charge redistribution without compromising the structural integrity. This aligns with the previous reports about acridones engaging in π–π stacking and electrophilic interactions with nucleophilic residues, which is consistent with their observed docking stabilization and MD-derived binding energetics82,83. Moreover, the intermediate profile of lignans similarly supports their role as versatile stabilizers, able to adaptively engage in van der Waals and electrostatic interactions84.
MD simulations of the NOX–lignan complex provided the detailed insights into the structural stability and dynamic features of the docked ligand in the near-physiological state in GROMACS. The RMSD analysis (Fig. 10) revealed that the protein backbone equilibrated rapidly within 0.2 ns and maintained the plateau at ~ 0.2 nm throughout 10 ns, indicating that the docking-derived pose represents a near-optimal binding orientation with minimal structural rearrangements. This rapid stabilization is consistent with previous MD studies of FAD/NAD(P)H oxido-reductases, where ligands occupying pre-organized hydrophobic pockets maintain low RMSD values, due to constrained mobility85,86,87.
The radius of gyration (Rg) remained tightly distributed between 2.38 and 2.46 nm (Fig. 11), indicating that the overall compactness of 2CDU protein was preserved upon lignan binding; this stability suggests that the ligand complements the shape and hydrophobic character of the binding pocket without inducing global unfolding. In consistent with our observations, the previous MD studies narrate that tight-binding polyphenolic or aromatic ligands maintained protein structural integrity while optimizing the interactions within oxido-reductase active sites88,89. Residue-level flexibility analysis via RMSF (Fig. 11) revealed that fluctuations were largely confined to loop and terminal regions (~ 0.4–0.45 nm), while the catalytic core remained highly rigid (~ 0.05–0.2 nm), a mechanistically significant pattern that preserves the geometry of active site for stable catalysis while allowing loop mobility to facilitate ligand access and product release, as likely found in the prior MD studies on oxido-reductases and NADPH-dependent enzymes90. Hydrogen bond analysis (Fig. 12) showed 2–4 persistent interactions with 5–6 transient peaks, reflecting a dynamic yet stable network, mediated by water and solvent effects; a behavior characteristic of polyphenolic ligands that maintain the binding affinity while allowing conformational adaptability. This is consistent with previous MD studies on flavonoid–oxido-reductase complexes where reversible H-bonds facilitated both stability and flexibility86. Energetic decomposition (Fig. 12) revealed that van der Waals interactions (− 250 to − 320 kJ/mol) primarily stabilized the complex, complemented by electrostatic contributions (− 70 to − 110 kJ/mol), resulting in strongly negative total energies (− 300 to − 400 kJ/mol) and reflecting the hydrophobic nature of the NOX binding pocket, as consistent with prior findings that aromatic and hydrophobic ligands engage oxido-reductases mainly through dispersion forces86. Free energy landscape (FEL) analysis (Fig. 13) showed multiple deep minima along PC1 and PC2 over 10 ns, suggesting that the complex is stable in terms of kinetics and energetics and thus, reinforcing the potential biological significance of lignan binding15,36. Hence, the combined phytochemical, biochemical and multi-layered in silico analyses confirmed that O. compressa, particularly its methanolic extract, possesses potent anti-oxidant potential mediated in part by inhibition of NADPH oxidase and thus, supporting its promise as a source of plant-derived compounds for therapeutic applications in oxidative stress-related disorders.
Conclusions
This study presents the first comprehensive and detailed structural elucidation and functional evaluation of Octhochloa compressa, highlighting its potential as a natural antioxidant source. Among all its extracts, the methanole extract demonstrated the significantly highest TPC and TFC, along with superior DPPH radical scavenging and FRAP activities. Structural characterization using HPLC and GC-MS enabled the quantification of 14 phenolic and 29 volatile/semi-volatile metabolites in the plant, respectively. Furthermore, in computational study, molecular docking exhibited strong binding interactions of different phytometabolites with NADPH oxidase along with DFT calculations rationalized these interactions, while MD simulations further validated these findings. Collectively, all these experimental and computational results provide a mechanistic insight into the antioxidant activity of O. compressa and strongly support its future development as a safe phyto therapeutic agent for oxidative stress-related disorders.
Data availability
All the data of the current study are available in the manuscript.
References
Al-Omar, M. S. et al. Anti-microbial, anti-oxidant, and α-amylase inhibitory activity of traditionally-used medicinal herbs: A comparative analyses of pharmacology, and phytoconstituents of regional halophytic plants’ diaspora. Molecules 25 (22), 5457. https://doi.org/10.3390/molecules25225457 (2020).
Hussain, R. et al. Temporal and dosage impact of magnesium oxide nanoparticles on grass carp: unveiling oxidative stress, DNA damage and antioxidant suppression. Toxicol. Mech. Meth. 35 (1), 19–31. https://doi.org/10.1080/15376516.2024.2382801 (2025).
Khanal, L. N., Sharma, K. R., Pokharel, Y. R. & Kalauni, S. K. Phytochemical analysis and in vitro antioxidant and antibacterial activity of different solvent extracts of beilschmiedia Roxburghiananees stem barks. Sci. World J. 2022 (6717012). https://doi.org/10.1155/2022/6717012 (2022).
Karabekir, S. C. et al. Protective effect of Astaxanthin on histopathologic changes induced by bisphenol a in the liver of rats. Pak Vet. J. 44 (2), 244–251. https://doi.org/10.29261/pakvetj/2024.178 (2024).
Saeed, S. H. et al. Interactive effects of toxic metals on the total phenolic and flavonoid in Hydrocotyle umbellata L. Asian J. Agric. Biol. 2023122 (2024). (2024). (2) https://doi.org/10.35495/ajab.2023.122
Vermot, A., Petit-Härtlein, I., Smith, S. M. E. & Fieschi, F. NADPH oxidases (NOX): an overview from Discovery, molecular mechanisms to physiology and pathology. Antioxidants 10 (6), 890. https://doi.org/10.3390/antiox10060890 (2021).
Mejri, F. et al. Phytochemical analysis and in vitro and in vivo evaluation of biological activities of artichoke (Cynara scolymus L.) floral stems: towards the valorization of food by-products. Food Chem. 333, 127506. https://doi.org/10.1016/j.foodchem.2020.127506 (2020).
Oladeji, O. S., Adelowo, F. E., Ayodele, D. T. & Odelade, K. A. Phytochemistry and Pharmacological activities of cymbopogon citratus: A review. Sci. Afr. 6, e00137. https://doi.org/10.1016/j.sciaf.2019.e00137 (2019).
Stagos, D. Antioxidant activity of polyphenolic plant extracts. Antioxidants 9 (1), 19. https://doi.org/10.3390/antiox9010019 (2019).
Rashid, S. et al. Phytomedicine efficacy and prospects in poultry; a new insight to old anthelmintic resistance. Cont. Vet. J. 4 (1), 62–75. https://doi.org/10.71081/cvj/2024.009 (2024).
Ondua, M., Njoya, E. M., Abdalla, M. A. & McGaw, L. J. Anti-inflammatory and antioxidant properties of leaf extracts of eleven South African medicinal plants used traditionally to treat inflammation. J. Ethnopharmacol. 234, 27–35. https://doi.org/10.1016/j.jep.2018.12.030 (2019).
Tunit, P., Thammarat, P., Okonogi, S. & Chittasupho, C. Hydrogel containing Borassus flabellifer L. male flower extract for antioxidant, antimicrobial, and anti-inflammatory activity. Gels 8 (2), 126. https://doi.org/10.3390/gels8020126 (2022).
Aslam, J. et al. A multidirectional phytochemical profiling, antimicrobial, antioxidant and toxicity studies of Neurada procumbens L.: A desert medicinal plant. J. King Saud Uni. Sci. 35 102862 (2023a). (2023). https://doi.org/10.1016/j.jksus.2023.102862
Syeda, A. M. & Riazunnisa, K. Data on GC-MS analysis, in vitro anti-oxidant and anti-microbial activity of the catharanthus roseus and Moringa Oleifera leaf extracts. Data brief. 29, 105258. https://doi.org/10.1016/j.dib.2020.105258 (2020).
Yu, M., Gouvinhas, I., Rocha, J. & Barros, A. I. Phytochemical and antioxidant analysis of medicinal and food plants towards bioactive food and pharmaceutical resources. Sci Rep. 11 (1), 10041. https://doi.org/10.1038/s41598-021-89437-4 (2021).
Ahmad, S., Zeb, A., Ayaz, M. & Murkovic, M. Characterization of phenolic compounds using UPLC–HRMS and HPLC–DAD and anti-cholinesterase and anti-oxidant activities of trifolium repens L. leaves. Eur. Food Res. Technol. 246, 485–496. https://doi.org/10.1007/s00217-019-03416-8 (2020).
Saravanakumar, K. et al. Metabolite profiling of methanolic extract of gardenia jaminoides by LC-MS/MS and GC-MS and its anti-diabetic, and anti-oxidant activities. Pharmaceut 14 (2), 102. https://doi.org/10.3390/ph14020102 (2021).
Ahmad, F. et al. Taxonomic application of foliar anatomy in grasses of tribe Eragrostideae (poaceae) from salt range of Pakistan. Pak J. Bot. 43 (5), 2277–2284 (2011).
Aslam, J. et al. Exploring the antidiabetic potential of octhochloa compressa: a comprehensive study on chemical profiling, in vitro, in vivo and in Silico Pharmacological properties. Front. Pharmacol. 16, 1541482. https://doi.org/10.3389/fphar.2025.1541482 (2025).
Chaudhari, S. K. et al. Foliar epidermal anatomy of grasses from thal desert, district Khushab, Pakistan. Int. J. Biosci. 4 (8), 62–70. https://doi.org/10.12692/ijb/4.8.62-70 (2014).
Chaudhari, S. K. et al. Ethnobotanical evaluation of grasses from thal Desert, Pakistan. Arch. Sci. 66, 248–255 (2013).
Aslam, J. et al. In vitro and in vivo anti-inflammatory potential of octhochloa compressa extracts in Carrageenan induced rats. Pak-Euro J. Med. Life Sci. 4 (4), 265–274. https://doi.org/10.31580/pjmls.v4i4.1717 (2021).
Atif, M. et al. Exploring in-vitro antioxidant, cytotoxicity, hemolytic, thrombolytic and anticancer potentials of ochthochloa compressa (Forssk.) Hilu. PLoS One. 20 (9), e0332194. https://doi.org/10.1371/journal.pone.0332194 (2025).
Aslam, J. et al. Antioxidant and anti-inflammatory potentials of aerial and floral parts of Neurada procumbens extracts: In vitro and in vivo studies. J. King Saud Uni. Sci. 35 102822 (2023b). (2023). https://doi.org/10.1016/j.jksus.2023.102822
Wariss, H. et al. Floristic composition of the plants of the Cholistan Desert, Pakistan. Am. J. Plant. Sci. 4 (12A), 58–65. https://doi.org/10.4236/ajps.2013.412A1009 (2013).
Rehman, T. U. et al. In vitro anthelmentic efficiency of citrullus colocynthis (L.) Schrad on Haemonchus contortus. Vet. Arhiv. 91 (3), 309–318. https://doi.org/10.24099/vet.arhiv.1024 (2021).
Nguyen, N. V. T. et al. Effect of extraction solvent on total phenol, flavonoid content, and antioxidant activity of avicennia officinalis. Res. Appl. Chem. 12, 2678–2690. https://doi.org/10.33263/BRIAC122.26782690 (2021).
Aouadi, K. et al. HPLC/MS phytochemical profiling with antioxidant activities of echium humileDesf. Extracts: ADMET prediction and computational study targeting human Peroxiredoxin 5 receptor. Agronomy 11 (11), 2165. https://doi.org/10.3390/agronomy11112165 (2021).
Elhakem, H. et al. DPPH, FRAP and TAEC assays with postharvest cabbage (Brassica oleracea) parameters during the packaging process. Pak J. Biol. Sci. 24 (2), 182–187. https://doi.org/10.3923/pjbs.2021.182.187 (2021).
Chaves, N., Santiago, A. & Alías, J. C. Quantification of the antioxidant activity of plant extracts: analysis of sensitivity and hierarchization based on the method used. Antioxidants 9 (1), 76. https://doi.org/10.3390/antiox9010076 (2020).
Beniaich, G. et al. GC-MS characterization, in vitro antioxidant, antimicrobial, and in Silico NADPH oxidase Inhibition studies of Anvillea radiata essential oils. Horticulturae 8 (10), 886. https://doi.org/10.3390/horticulturae8100886 (2022).
Yan, B. & Wu, Q. Evaluating the cardioprotective efficacy of diosmin against LPS-induced cardiac dysfunction: in Silico Docking and experimental investigations. Pak Vet. J. 44 (4), 1201–1208. https://doi.org/10.29261/pakvetj/2024.288 (2024).
Al Shammari, B. R. Antioxidant, antimicrobial and in Silico NADPH oxidase Inhibition of chemically-analyzed essential oils derived from Ballota deserti (Noë) jury. Molecules 27 (19), 6636. https://doi.org/10.3390/molecules27196636 (2022).
Deka, J. et al. Photocatalytic and antibacterial activities of alternanthera dentata mediated Bio-nanocomposite of CuO nanoparticles and DFT study on the selective capping behavior of phytocompounds. J. Elect. Mater. 53 (10), 6508–6523. https://doi.org/10.1007/s11664-024-11344-x (2024).
Vieira, I. et al. (ed P, H.) Visual dynamics: a WEB application for molecular dynamics simulation using GROMACS. BMC Bioinform. 24 1 107 https://doi.org/10.1186/s12859-023-05234-y (2023).
Rodríguez-Martínez, A. et al. Enhancing MD simulations: asgard’s automated analysis for GROMACS. J. Biomol. Struct. Dyn. 1–13. https://doi.org/10.1080/07391102.2024.2349527 (2024).
Nazir, A., Mustafa, R. & Iqbal, M. In vitro & in vivo phytochemical evaluation of bioactive components against hyperglycemic-induced oxidative stress in Streptozocin rat model: a histopathological investigation. Biointerface Res. Appl. Chem. 20, 7321–7341. https://doi.org/10.33263/BRIAC126.73217341 (2021).
Bilal, M. F. et al. In vitro evaluation of antimicrobial efficacy of Azadirachta indica (Neem) extract against Staphylococcus aureus isolates from subclinical mastitis in Cholistani cattle. Pak-Euro J. Med. Life Sci. 7 (Sp 2), S287-S298 (2024). https://doi.org/10.31580/pjmls.v7isp2.3179
Malik, S. et al. Characterization and identification of bioactive natural products in the ethanol extracts of acacia nilotica, melia azedarach, and euphorbia Hirta from Cholistan desert, Pakistan. Asian J. Agric. Biol. 2024 (2), 2023180. https://doi.org/10.35495/ajab.2023.180 (2024).
Al-Farraj, D. A. et al. Enhanced wound healing effects of herbal gel formulations in a rabbit model: a comparative study. Asian J. Agri. Biol. 2023190 (2024). (2024). (1) https://doi.org/10.35495/ajab.2023.190
Dai, W., Long, L., Wang, X., Li, S. & Xu, H. Phytochemicals targeting Toll-like receptors 4 (TLR4) in inflammatory bowel disease. Chin. Med. 17 (1), 53. https://doi.org/10.1186/s13020-022-00611-w (2022).
Lara, M. V. et al. Stone fruit as biofactories of phytochemicals with potential roles in human nutrition and health. Front. Plant. Sci. 11, 562252. https://doi.org/10.3389/fpls.2020.562252 (2020).
Khan, M. T. S. et al. Therapeutic effects of medicinal plants on immunology and growth (a review). Cont. Vet. J. 3 (2), 43–54. https://doi.org/10.71081/cvj/2023.019 (2023).
Štrbac, F. et al. In vitro anthelmintic potential of selected essential oils against Gastrointestinal nematodes of sheep. Pak Vet. J. 44 (4), 1053–1062. https://doi.org/10.29261/pakvetj/2024.295 (2024).
Agatonovic-Kustrin, S., Gegechkori, V., Kustrin, E. & Morton, D. W. The effect of lactic acid fermentation on extraction of phenolics and flavonoids from Sage leaves. Appl. Sci. 12 (19), 9959. https://doi.org/10.3390/app12199959 (2022).
Uba, A. I. et al. Antioxidant and enzyme inhibitory properties, and HPLC–MS/MS profiles of different extracts of arabis carduchorum boiss.: an endemic plant to Turkey. App Sci. 12 (13), 6561. https://doi.org/10.3390/app12136561 (2022).
Kalsoom, R. et al. Bifenthrin induced toxic effects on haematological, reproductive and histo-morphological profile in adult male quail (Coturnix japonica). Asian J. Agric. Biol. 2024 (1) (2023357). https://doi.org/10.35495/ajab.2023.357 (2024).
Yang, B. et al. Exposure to the herbicide Butachlor activates hepatic stress signals and disturbs lipid metabolism in mice. Chemosphere 283, 131226. https://doi.org/10.1016/j.chemosphere.2021.131226 (2021).
Afzal, G. et al. Effects of sub-lethal concentrations of Lindane on histo-morphometric and physio-biochemical parameters of Labeo Rohita. PLoS ONE. 19 (7), e0304387. https://doi.org/10.1371/journal.pone.0304387 (2024).
Mahmood, Y. et al. Multi-biomarker approach to assess oxidative stress and antioxidants profile in male albino rats exposed to ZnO nanoparticles. Asian J. Agric. Biol. 2024115 (2024). (2024). (4) https://doi.org/10.35495/ajab.2024.115
Rasheed, M. et al. Prophylactic effects of methylene blue, coconut and Olive oils supplements on hemato-biochemical and histo-pathological parameters against p-Phenylenediamine toxicity in male albino rats. Pak Vet. J. 44 (3), 840–846. https://doi.org/10.29261/pakvetj/2024.193 (2024).
Ghoora, M. D., Haldipur, A. C. & Srividya, N. Comparative evaluation of phytochemical content, antioxidant capacities and overall antioxidant potential of select culinary microgreens. J. Agri Food Res. 2, 100046. https://doi.org/10.1016/j.jafr.2020.100046 (2020).
Truong, D. H. et al. Evaluation of the use of different solvents for phytochemical constituents, antioxidants, and in vitro anti-inflammatory activities of Severinia buxifolia. J. Food Quality, 8178294 (2019). (2019). https://doi.org/10.1155/2019/8178294
Świątek, Ł. et al. Chemical characterization of different extracts of Justicia secunda Vahl and determination of their anti-oxidant, anti-enzymatic, anti-viral, and cytotoxic properties. Antioxidants 12 (2), 509. https://doi.org/10.3390/antiox12020509 (2023).
Ramadhania, N. R., Harun, F., Purnomo, A. S. & Fatmawati, S. Anti-oxidant and anti-bacterial activities of anthurium plowmanii leaves extracts. Mal J. Fund Appl. Sci. 15, 194–199. https://doi.org/10.11113/mjfas.v15n2.1040 (2019).
Catarino, M. D. et al. Applications of antioxidant secondary metabolites of sargassum spp. Mar. Drugs. 21 (3), 172. https://doi.org/10.3390/md21030172 (2023).
Park, J. S., Han, J. M., Surendhiran, D. & Chun, B. S. Physicochemical and biofunctional properties of sargassum thunbergii extracts obtained from subcritical water extraction and conventional solvent extraction. J. Supercritical Fluids. 182, 105535. https://doi.org/10.1016/j.supflu.2022.105535 (2022).
Porro, C. et al. Functional and therapeutic potential of Cynara scolymus in health benefits. Nutrients 16 (6), 872. https://doi.org/10.3390/nu16060872 (2024).
Lytra, K. et al. Bio-guided investigation of sideritiscypria methanol extract driven by in vitro antioxidant and cytotoxic assays. Chem. Biodiv. 18 (3). https://doi.org/10.1002/cbdv.202000966 (2021). e2000966.
Prabu, K., Rajasekaran, A., Bharathi, D. & Ramalakshmi, S. Anti-oxidant activity, phytochemical screening and HPLC profile of rare endemic cordia diffusa. J. King Saud Uni Sci. 31 (4), 724–727. https://doi.org/10.1016/j.jksus.2018.04.025 (2019).
Chen, N. et al. The relationship between polysaccharide structure and its antioxidant activity needs to be systematically elucidated. Int. J. Biol. Macromol. 270, 132391. https://doi.org/10.1016/j.ijbiomac.2024.132391 (2024).
Khan, A. et al. Fatty acid composition, phenolic compounds, phytosterols, and lipid oxidation of single-and double-fractionated Olein of safflower oil produced by low-temperature crystallization. ACS Omega. 9 (6), 6787–6796. https://doi.org/10.1021/acsomega.3c08099 (2024).
Osuala, O. et al. Antimicrobial potential of aqueous and ethanol extracts of unripe carica Papaya L. Acad. Nut Dietet. 2 (1), 1–10. https://doi.org/10.20935/AcadNutr7471 (2025).
Panayides, J. L., Riley, D. L., Hasenmaile, F. & Van Otterlo, W. A. The role of silicon in drug discovery: a review. RSC Med. Chem. 15 (10), 3286–3344. https://doi.org/10.1039/D4MD00169A (2024).
Chibesa, M. C. et al. Silicon solubilisation from soil minerals and soil by root exudate compounds. Geoderma 459, 117375. https://doi.org/10.1016/j.geoderma.2025.117375 (2025).
Aytar, E. C. & Aydın, B. Investigation of chemical composition, antioxidant properties, and molecular docking in different roasting stages of coffee beans. Food Bioproc. Technol. 18 (2), 1464–1482 (2025). https://doi.org/10.1007/s11947-024-03539-1
Redondo-Velasco, M. et al. New analytical platform for the sensitive detection and quantification of silicon compounds in plastic pyrolysis oil. Microchem J. 114703 https://doi.org/10.1016/j.microc.2025.114703 (2025).
Jayaraj, P. et al. Sesamol: A powerful functional food ingredient from Sesame oil for cardioprotection. Food Funct. 11 (2), 1198–1210. https://doi.org/10.1039/c9fo01873e (2020).
Imran, M. et al. Apigenin as an anticancer agent. Phytother Res. 34 (8), 1812–1828. https://doi.org/10.1002/ptr.6647 (2020).
Waheed, A. et al. Insights into Pharmacological potential of apigenin through various pathways on a nanoplatform in multitude of diseases. Curr. Pharma Des. 29 (17), 1326–1340. https://doi.org/10.2174/1381612829666230529164321 (2023).
Hafez, A. A. et al. Calcitriol attenuates the cytotoxicity induced by aluminium phosphide via inhibiting mitochondrial dysfunction and oxidative stress in rat isolated cardiomyocytes. Pest Biochem. Physiol. 176, 104883. https://doi.org/10.1016/j.pestbp.2021.104883 (2021).
Polkaew, M. et al. Applicability of butterfly pea flower extract as an alternative natural dye in histopathological canine mast cell tumor diagnosis. Pak Vet. J. 44 (2), 266–273. https://doi.org/10.29261/pakvetj/2024.163 (2024).
Festa, J., Da Boit, M., Hussain, A. & Singh, H. Potential benefits of berry anthocyanins on vascular function. Mol. Nut Food Res. 65 (19), 2100170. https://doi.org/10.1002/mnfr.202100170 (2021).
VanáVo, Q. et al. The antioxidant activity of tetrahydrofuran lignans from anogeissusrivularis: theoretical insights into the radical scavenging activity and enzyme Inhibition. New. J. Chem. 47, 17314–17322. https://doi.org/10.1039/D3NJ03344A (2023).
Sarıkaya, E. K., Ekincioğlu, Y., Bahçeli, S. & Dereli, Ö. Analysis of the nonlinear optical properties, vibrational spectra, DFT method and photovoltaic performance of cyanidin-3-rutinoside chloride. Opt. Quant. Electron. 56 (8), 1383. https://doi.org/10.1007/s11082-024-07294-7 (2024).
Atalay, V., Atish, I., Shahin, K., Kashikchi, E. & Karahan, M. DFT study: ranking of antioxidant activity of various candidate molecules. UNEC J. Eng. Appl. Sci. 2, 33–40 (2022).
Boulmokh, Y., Belguidoum, K., Meddour, F. & Amira-Guebailia, H. Enhanced antioxidant properties of novel Curcumin derivatives: a comprehensive DFT computational study. Struc Chem. 35 (3), 825–839. https://doi.org/10.1007/s11224-023-02237-6 (2024).
Dudek, A., Spiegel, M., Strugała-Danak, P. & Gabrielska, J. Analytical and theoretical studies of antioxidant properties of chosen anthocyanins; a structure-dependent relationships. Inter J. Mol. Sci. 23 (10), 5432. https://doi.org/10.3390/ijms23105432 (2022).
Zheng, X., Du, Y., Chai, Y. & Zheng, Y. A DFT – Based Mechanism Analysis of the Cyclodextrin Inclusion on the Radical Scavenging Activity of Apigenin. Antioxidants, 12 (11), (2018). (2023) https://doi.org/10.3390/antiox12112018
Rauf, A. et al. In silico, SwissADME, and DFT studies of newly synthesized oxindole derivatives followed by antioxidant studies. J. Chem. 2023 (1) (5553913). https://doi.org/10.1155/2023/5553913 (2023).
Pirzada, A. S. et al. Physicochemical properties, Pharmacokinetic studies, DFT approach, and antioxidant activity of nitro and Chloro Indolinone derivatives. Front. Chem. 12, 1360719. https://doi.org/10.3389/fchem.2024.1360719 (2024).
Arunachalam, A., Sankar, M., Pandi, B., Paul, S. & Thilagar, S. Evaluation of Rapanone and Nectandrin B as novel inhibitors for targeting the metastatic regulator protein BACH1 using breast cancer cell line Mcf-7. J. Biomol. Struct. Dyn. 42 (20), 11185–11200. https://doi.org/10.1080/07391102.2023.2260880 (2024).
Sulthanudeen, S., Imran, P. M., Selvakumaran, M. & Kubaib, A. Novel acridone derivatives probed using DFT, including design, synthesis, characterization with anti-oxidant and anti-mitotic screening. Results Chem. 5, 100753. https://doi.org/10.1016/j.rechem.2022.100753 (2023).
Kumar, R. C. S., Jayaraman, A. & Venkatachalapathy, R. Computational investigation of lignans as potential target for non-alcoholic fatty liver disease: insights from network pharmacology, docking, DFT, and dynamics simulation analysis. Hum. Gene. 201457 https://doi.org/10.1016/j.humgen.2025.201457 (2025).
Kamiński, K. et al. Rossmann-toolbox: a deep learning-based protocol for the prediction and design of cofactor specificity in Rossmann fold proteins. Brief. Bioinform. 23 (1), bbab371. https://doi.org/10.1093/bib/bbab371 (2022).
Ouyang, X. et al. A multifunctional flavoprotein monooxygenase HspB for hydroxylation and CC cleavage of 6-hydroxy-3-succinoyl-pyridine. Applied Environ. Microbiol. 90 (3), e02255-02223 (2024). https://doi.org/10.1128/aem.02255-23. Toledo-Patiño, S., Pascarelli, S., Uechi, G.-i., Laurino, P. Insertions and deletions mediated functional divergence of Rossmann fold enzymes. Proceed. Nat. Acad. Sci. 119 (48), e2207965119 (2022). https://doi.org/10.1073/pnas.2207965119.
Kumar, A., Nimsarkar, P. & Singh, S. Probing the interactions responsible for the structural stability of trypanothione reductase through computer simulation and biophysical characterization. Prot. J. 41 (2), 230–244. https://doi.org/10.1007/s10930-022-10052-x (2022).
Sonkar, K. S., Achary, V. M. M., Sahoo, S., Reddy, M. K. & Arockiasamy, A. Biochemical and structural characterization of a robust and thermostable ascorbate recycling monodehydroascorbate reductase (MDHAR) from stress adapted Pearl millet. Biochem. Biophys. Res. Commun. 662, 135–141. https://doi.org/10.1016/j.bbrc.2023.04.034 (2023).
Hoda, A., Kolaneci, V., Lika, M. & Koleci, X. Computational insights into mercuric reductase from Pseudomonas fluorescens: a bioinformatic and molecular dynamics approach for mercury detoxification. Bioremed J. 1–17. https://doi.org/10.1080/10889868.2024.2428721 (2024).
Narang, P. K. et al. Genome-based identification and comparative analysis of enzymes for carotenoid biosynthesis in microalgae. World J. Microbiol. Biotechnol. 38 (1), 8. https://doi.org/10.1007/s11274-021-03188-y (2022).
Acknowledgements
The authors extend their appreciation to “Ongoing Research Funding Program” (ORF-2025-965), King Saud University, Riyadh, Saudi Arabia.
Author information
Authors and Affiliations
Contributions
J. Alam: Data curation, Methodology, Software, Writing – original draft, Writing – review & editing. T. Hussain: Investigation, Methodology, Writing – original draft. M.I. Shahzad: Conceptualization, Investigation, Supervision. H.M. Ali: Conceptualization, Writing – original draft, Writing – review & editing. M. Aslam: Investigation, Data curation, Writing – original draft. M. Shahid: Conceptualization, Investigation, Supervision. A. Jamal: Methodology, Data curation, Writing – review & editing. U. Farooq: Methodology, Investigation, Writing – review & editing. S. Asif: Formal Analysis, Writing – review & editing. K.S. Abass: Formal analysis, Validation, Writing – review & editing. D. Fouad: Formal analysis, Funding acquisition, Writing – original draft. F.S. Ataya: Validation, Funding acquisition, Writing – review & editing. K. Ahmadova: Investigation, Software, Writing – review & editing. Z. Chachar: Investigation, Formal Analysis, Writing – review & editing. All authors critically read and gave their consent for publication of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Consent to publish
All listed authors agreed to the publication of the present manuscript.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Aslam, J., Hussain, T., Shahzad, M.I. et al. Chemical finger-printing, antioxidant activity and in silico validation of phytometabolites of Octhochloa compressa. Sci Rep 15, 41339 (2025). https://doi.org/10.1038/s41598-025-25238-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-25238-3