Abstract
Hepatocellular carcinoma (HCC) represents a significant global health challenge due to its molecular heterogeneity and the immunosuppressive nature of its tumor microenvironment (TME). Tumor-associated macrophages (TAMs), particularly the M2-polarized subset, are crucial in HCC progression, contributing to immune evasion, angiogenesis, and metastasis. However, the mechanisms driving TAM reprogramming and their metabolic interactions within the TME remain poorly understood. In this study, we applied single-cell RNA sequencing (scRNA-seq) and spatial transcriptomics to identify a novel metabolic-immune axis in HCC. Our findings revealed increased oxidative phosphorylation (OXPHOS) in tumor cells, which correlated with enhanced ribosomal activity in M2 macrophages, suggesting a metabolic coupling between the two. Tumor cells, under elevated OXPHOS, secrete factors that recruit macrophages and promote M2 polarization, thereby reinforcing immune suppression. Further analysis of The Cancer Genome Atlas (TCGA) dataset led to the development of a prognostic model based on OXPHOS-related genes, showing strong predictive power across multiple cohorts. These results suggest a novel OXPHOS-ribosome-M2 polarization axis and offer potential therapeutic targets for immunometabolic therapies in HCC.
Similar content being viewed by others
Introduction
Hepatocellular carcinoma (HCC) remains a major global health issue, characterized by its molecular heterogeneity and the immunosuppressive nature of its tumor microenvironment (TME)1. Despite advances in understanding tumor biology, significant challenges remain in effectively targeting HCC, mainly due to the complexity of the TME and its influence on tumor progression. Tumor-associated macrophages (TAMs), particularly those polarized to the M2 phenotype, play a central role in immune suppression, angiogenesis, and metastasis in HCC2. However, the mechanisms by which TAMs interact with tumor cells through metabolic pathways to maintain an immunosuppressive microenvironment are still poorly understood. This gap in knowledge presents a major hurdle in the development of more effective therapies for HCC.
One of the key difficulties in addressing these issues is the lack of a comprehensive understanding of the metabolic-immune interactions within the TME. Evidence increasingly points to metabolic and translational pathways as key regulators of TAM polarization3,4. Among these, ribosome biogenesis and associated signaling cascades have emerged as potential drivers of macrophage adaptation toward an M2-like phenotype5,6. Ribosome-related pathways not only regulate protein synthesis but also orchestrate metabolic and immunological processes that fine-tune macrophage function in the TME7. Although studies have implicated these pathways in macrophage biology, their role in TAM-mediated immune suppression and interaction with tumor cells in HCC remains poorly accessed. Understanding how tumor-derived signals influence TAM ribosome pathways to sustain their immunosuppressive function could provide valuable insights into the feedback mechanisms that reinforce immunometabolic vulnerabilities in HCC progression.
To address this gap, we conducted an integrative analysis using single-cell transcriptomics, spatial metabolomics to investigate the TAM-tumor cell interplay in HCC (As shown in Fig. 1). Our findings reveal a previously uncharacterized metabolic-immune feedback loop: TAMs promote tumor cell OXPHOS activation, which subsequently induces the release of substances, acting on TAMs to activate ribosome-associated signaling pathways, thereby driving their polarization toward the M2-like phenotype. This establishes an OXPHOS-ribosome-M2 polarization axis that reinforces the immunosuppressive and pro-tumor functions of TAMs, ultimately sustaining a tumor-promoting microenvironment and facilitating disease progression.
In addition, we leveraged the TCGA-HCC dataset to evaluate the clinical significance of OXPHOS-related genes. Using these genes, we developed a robust prognostic model capable of stratifying HCC patients based on survival outcomes. The model highlights the clinical relevance of the OXPHOS pathway in HCC progression and provides a foundation for personalized risk assessment. Importantly, the prognostic signature derived from this analysis underscores the therapeutic potential of targeting the metabolic-immune axis in HCC.
By addressing the knowledge gap in the immune-metabolic interactions between TAMs and tumor cells, our study offers a new understanding of the metabolic drivers of immune evasion in HCC and provides innovative targets for therapeutic intervention. These findings have significant implications for improving immunometabolic therapy in HCC and advancing the development of precision medicine approaches for patients suffering from this aggressive cancer.
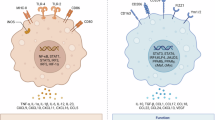
Overview of the study design and proposed mechanism. (A) Workflow diagram illustrating the key steps of the study. The analysis begins with single-cell RNA sequencing (scRNA-seq) and spatial transcriptomics data to identify the main tumor microenvironment (TME) cells and their distribution in HCC. In vitro experiments using a cell coculture system are conducted for further validation, including Western Blot (WB), qRT-PCR, and ELISA assays. A model construction section uses TCGA-LIHC transcriptome and clinical data to evaluate the expression of identified oxidative phosphorylation (OXPHOS) genes. Consensus clustering of the data helps stratify patients into distinct clusters based on their molecular profiles, with subsequent analyses to assess clinicopathological and prognostic significance, immune infiltration, and the development of a prognostic signature using GO and KEGG pathways. (B) Schematic representation showing the hypothesized mechanism. M2 macrophages within the TME of HCC drive an increase in OXPHOS in tumor cells. The elevated OXPHOS in tumor cells leads to the release of signal factors, which is recognized by its receptors on M2 macrophages. This signaling cascade activates the lysosomal pathway in M2 macrophages, enhancing their polarization and recruitment. As a result, M2 macrophages further promote tumor progression through the activation of the ribosome pathway, creating a positive feedback loop that supports tumor proliferation.
Results
Single-cell transcriptomics of macrophages in HCC
To delineate the tumor microenvironment (TME) landscape of hepatocellular carcinoma (HCC), we performed single-cell RNA sequencing (scRNA-seq) on tumor (n = 7) and non-tumor tissues (n = 4) spanning early to advanced disease stages (as shown in Supplementary Table 1). After filtering ineligible cells and yielded core cells for further analysis (Fig. S1A-C), UMAP clustering revealed 16 transcriptionally distinct resolved these clusters into major cellular components, including liver and tumor cells (Liver_tumorcells), macrophages, proliferative macrophages (prol.macro), Monocytes, NK cells and T cells (NK/T cells), B cells, fibroblasts, dendritic cells (DCs), and endothelial cells (ECs) (Fig. 2A, B), which allowed the classification of the clusters into distinct cell types. These annotated cell types were visualized in a UMAP plot (Fig. 2C). Figure 2D displays the relative proportions of the identified cell types across the tumor samples and different tumor stage. Notably, the proportion of macrophages expanded significantly in more advanced stages of HCC, as shown in the bar chart (Fig. 2E). This trend suggests a potential role for macrophages in driving tumor progression, prompting further investigation into their functional states.
To further explore macrophage heterogeneity and polarization, macrophages were isolated for downstream analyses. A t-SNE plot of the macrophage population revealed distinct transcriptional states, indicative of functional heterogeneity (Fig. 2F). We focused on the polarization states of macrophages, categorizing them into classically activated M1 macrophages and alternatively activated M2 macrophages, based on the expression of subtype-specific marker genes (Fig. 2G). Temporal dynamics of macrophage polarization were assessed across different stages of HCC. The relative abundance of M2 macrophages increased progressively with tumor stage, while M1 macrophages diminished (Fig. 2H). This trend was corroborated by analyzing the M2/M1 ratio, which showed a marked rise in advanced stages of HCC (Fig. 2I, J). These findings emphasize the dominance of immunosuppressive M2 polarization in shaping the TME during HCC progression.
To explore macrophage polarization dynamics, we performed pseudotime trajectory analysis after correcting for batch effects to ensure robust stage-specific comparisons (Fig. S1D, E). This analysis revealed a gradual transition of macrophages along HCC progression (Fig. 2H). The develop tract of macrophages in different stage was shown in Fig. 2K. Notably, M2-specific marker genes displayed an increasing trend from normal tissue to stage B (Fig. 2L), highlighting the critical role of M2 macrophages in promoting tumor advancement.
Differential expression analysis between M2 and M1 macrophages identified ribosomal pathways as significantly enriched in M2 macrophages, indicating enhanced translational activity (Fig. 2M).
These results provide insights into the metabolic reprogramming underlying M2 polarization and its pro-tumorigenic functions.
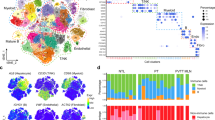
Single-cell RNA sequencing (scRNA-seq) analysis of the tumor microenvironment (TME) in HCC. (A) t-SNE plot showing the clustering of cells based on single-cell RNA sequencing data. Each color represents a distinct cell type identified in the TME, with the corresponding cell type labeled on the right. (B) Dot plot depicting the expression levels and the percentage of cells expressing specific markers for various immune and tumor cell types, including macrophages, monocytes, fibroblasts, and others. (C) UMAP plot visualizing the distribution of different cell types in the TME, with macrophages specifically highlighted. (D) Bar plots showing the ratio of different cell types across patients (left), distribution across tumor stages (middle), and the number of cells identified in the TME for each sample (right). (E) Bar graph depicting the proportions of different cell types in normal, BCLC-A, and BCLC-B HCC samples. (F) t-SNE plot showing the distribution of M1-like and M2-like macrophages in the TME based on gene expression profiles. (G) Dot plot comparing the average expression of M1-like and M2-like macrophage polarization markers across different cell populations. (H) Bar chart illustrating the polarization ratio of macrophages (M1 vs. M2) in normal, BCLC-A, and BCLC-B HCC samples. (I) Comparison of the cell ratios of M2 and M1 macrophages across different patients (left), by tumor versus normal tissue (middle), and the number of cells identified (right). (J) Line graph showing the increasing macrophage marker expression from normal to BCLC stages, with a trend indicating the shift towards M2 polarization as HCC progresses. (K) UMAP plot showing the clustering of macrophage subtypes (M1 and M2) and their relationship to the progression of the tumor stages (BCLC). (L) Pseudotime analysis showing the trajectory of macrophage polarization from normal tissue to advanced stages of HCC, with expression profiles of key macrophage markers such as APC1, CSF1, and others. (M) KEGG pathway enrichment analysis of differential genes between M2 and M1 macrophages, highlighting pathways such as oxidative phosphorylation (OXPHOS) and immune response pathways associated with tumor progression.
Spatial mapping reveals localization of M2 macrophages in the HCC microenvironment and functional validation of tumor-macrophage interactions
To explore the spatial organization of macrophages in HCC, we integrated scRNA-seq-derived cell type annotations with spatial transcriptomics (ST) data. This integration enabled the precise localization of macrophages in different polarization state and other immune cell subsets within HCC tissues (Fig. 3A-G, H-N, Fig. S2A, B). Notably, M2 macrophages were predominantly enriched in tumor-adjacent regions, co-localized with tumor cells, suggesting a strong spatial interaction between tumor cells and M2 macrophages. Given the spatial proximity observed, we hypothesized that tumor cells play a role in reprogramming macrophages toward the M2 phenotype.
For validating the observed spatial co-localization between tumor cells and M2 macrophages, we performed functional co-culture experiments using a transwell model (Fig. 3O). Quantitative RT-PCR (qRT-PCR) demonstrated that macrophages co-cultured with tumor cells exhibited significantly elevated expression of M2-specific marker genes (Fig. 3P). Consistently, Western blot analysis confirmed an increase in protein levels of CD206 and ARG-1 in co-cultured macrophages compared to controls (Fig. 3Q). Furthermore, ELISA analysis of the culture medium revealed enhanced secretion of cytokines TGF-β, and decreased TNF-α, IL-1β, and IL-6 under co-culture conditions (Fig. 3R). These results strongly suggest that tumor cells actively promote macrophage polarization toward an M2 phenotype, reinforcing their pro-tumorigenic roles in the HCC microenvironment.
Integration of scRNA-seq and spatial transcriptomics reveals the spatial distribution of macrophages in HCC and functional validation of M2 polarization. (A-G) Spatial transcriptomics (ST) of HCC tissue sections showing the overall transcriptional landscape. (H-N) Single-cell RNA sequencing (scRNA-seq)-derived cell type annotations mapped onto the ST data, identifying major cell types. (O) Transwell co-culture model illustrating the interaction between tumor cells (upper chamber) and macrophages (lower chamber). (P) Quantitative RT-PCR analysis comparing the expression of M2-specific marker genes in macrophages co-cultured with tumor cells versus macrophages cultured alone. (Q) Western blot analysis showing the protein levels of CD206, iNOS, ARG-1 in macrophages after co-culture with tumor cells compared to controls and quantifications (n = 3). (R) ELISA analysis of the culture medium for the secretion level of TNF-α, IL-1β, TGF-β, and IL-6 (n = 6).
Tumor cells with higher OXPHOS activity promote M2 polarization through chemerin signaling
Spatial transcriptomics analysis allowed for the classification of tumor cells into M2-proximal and M2-distal groups based on their spatial distance to M2 macrophages (Fig. 4A, Fig. S3A-F). Gene Ontology (GO) enrichment analysis and KEGG pathway analysis revealed that tumor cells closely interacting with M2 macrophages (M2-proximal) exhibited significantly enriched oxidative phosphorylation (OXPHOS) activity compared to M2-distal tumor cells (Fig. 4B, C).
Cell-cell communication analysis using CellChat identified more extensive interactions between tumor cells and M2 macrophages compared to M1 macrophages (Fig. 4D, E, Fig. S4A-C). Specifically, tumor cells and M2 macrophages were found to communicate through the chemerin signaling pathway, which mediates M2 macrophage polarization and contributes to shaping an immunosuppressive tumor microenvironment (Fig. 4F, G, Fig. S4D). This enriched chemerin signaling points to a mechanism by which tumor cells metabolically and functionally reorganize their microenvironment to promote an immunosuppressive and tumor-supportive phenotype.
Further spatial analysis demonstrated that M2 macrophages interact more strongly with tumor cells in tumor-associated regions compared to non-tumor regions, as evidenced by increased signaling intensity (Fig. 4H, I). Spatial enrichment analysis also confirmed a higher accumulation of M2 macrophages in the ribosome pathway of tumor cells within tumor regions relative to non-tumor regions (Fig. 4J, K), which was corroborating with the previous results.
To validate the role of chemerin signaling and OXPHOS activity in tumor-macrophage interactions, a series of in vitro experiments were conducted. Analysis of TCGA-LIHC data showed that tumor tissues expressed the chemerin-encoding genes, ATP8A1, associating with mitochondrial function and OXPHOS marker, SDHB at a higher level compared to non-tumor tissues (Fig. 5A). As for the protein level, ATP8A1, SDHB, and Chemerin were up-regulated in the HCC tissues (Fig. 5B). In co-culture experiments, Huh7 cells exhibited elevated expression of ATP8A1, SDHB, and Chemerin when cultured with macrophages (Fig. 5C). Functional assays demonstrated that co-culture with macrophages enhanced tumor cell viability (Fig. 5D) and ATP production levels (Fig. 5E), indicating metabolic activation under these conditions.
On the other hand, immunofluorescence staining further revealed polarization of these macrophages toward an M2-like phenotype, characterized by decreased expression of iNOS and increased CD206 upon treatment with tumor cell-conditioned medium (Fig. 5F). To assess how tumor cell OXPHOS affects macrophage polarization, Huh7 cells were treated with oligomycin (0.5 µM and 2 µM) for 6 h to inhibit OXPHOS, then washed and cultured in fresh DMEM for 24 h. OXPHOS inhibition, especially at 2 µM, significantly reduced ATP5A1, SDHB, and Chemerin expression in Huh7 cells (Fig. 5G). Conditioned medium from these cells was then applied to PMA-differentiated THP-1 macrophages for 24 h. As oligomycin concentration increased, M2 marker expression in macrophages declined, indicating that stronger OXPHOS inhibition in tumor cells suppresses M2 polarization (Fig. 5H).
Additionally, macrophages co-cultured with tumor cells displayed increased expression of ribosome-related marker, UBTF, and the Chemerin receptor, CMKLR1, suggesting a functional response to tumor cell-secreted factors (Fig. 5I). α-NETA, CMKLR1 antagonist, was used to treated macrophages co-cultured with Huh7 cells, as shown in Fig. 5J, α-NETA increased the M1 marker while decreased the M2 marker.
Thus, we revealed that OXPHOS and chemerin expression when interacting with macrophages, while macrophages polarize toward an M2-like phenotype with increased CMKLR1 expression and lysosome activity. These findings highlight a synergistic metabolic and signaling crosstalk between tumor cells and M2 macrophages in shaping tumor progression.
Spatial and functional analysis of tumor-macrophage interactions and the role of chemerin signaling in HCC. (A) Spatial mapping of tumor cells categorized as M2-proximal or M2-distal based on their spatial distance to M2 macrophages, using spatial transcriptomics data. Tumor cells closely associated with M2 macrophages are highlighted as M2-proximal, while cells farther from M2 are labeled as M2-distal. (B) Gene Ontology (GO) enrichment analysis of differentially expressed genes between M2-proximal and M2-distal tumor cells. (C) KEGG pathway enrichment analysis of differentially expressed genes between M2-proximal and M2-distal tumor cells. (D) CellChat analysis of cellular communication between tumor cells and macrophages, demonstrating signaling interactions. (E) Quantitative analysis of signaling inputs and outputs for various cells in the HCC. (F, G) Visualization of the chemerin signaling pathway and its interactions within the tumor-macrophage network. (H) Interaction strength analysis between M2 macrophages and tumor cells in tumor regions compared to non-tumor regions. (I) Comparative interaction strength between tumor cells and M2 macrophages across tumor and non-tumor tissues. (J) Spatial GO enrichment analysis of M2 macrophages in tumor-associated regions versus non-tumor regions. (K) Enrichment KEGG analysis visualizing the spatial accumulation of M2 macrophages around tumor cells compared to non-tumor cells.
Validation of chemerin signaling and OXPHOS activity in tumor-macrophage interactions through in vitro experiments. (A) Gene expression analysis of chemerin-encoding genes in tumor and non-tumor tissues using TCGA-LIHC database. (B) Immunohistochemical (IHC) staining of ATP8A1, SDHB, and Chemerin in HCC tumor and adjacent normal tissues (scale bar: 100 μm), and IHC score quantification (n = 6). (C) Analysis of OXPHOS marker and Chemerin expression in tumor cells co-cultured with macrophages and quantifications (n = 3). (D) Assessment of tumor cell viability after co-culture with macrophages. (E) ATP production levels in tumor cells under co-culture conditions. (F) Immunofluorescence staining showing the expression level of iNOS, CD206 in macrophages cultured with tumor cell-conditioned medium to evaluate macrophage polarization (Scale bar: 50 μm). (G) Western blot analysis the expression level of ATP8A1, SDHB, and Chemerin in Huh7 cells treated with different concentration of oligomycin and quantification (n = 3). (H) Protein expression and quantification in macrophages treated with the conditioned medium collected after treating tumor cells with different concentrations of oligomycin (0.5 µM and 2 µM) and quantification (n = 3). (I) Expression of ribosome-related marker and the chemerin receptor, CMKLR1 in macrophages co-cultured with tumor cells and quantifications (n = 3). (J) The expression level of CD86 and CD206 analyzed by flow cytometry.
The signature of OXPHOS genes in HCC
To further analysis the influence OXPHOS genes in HCC, we obtained the OXPHOS related genes from KEGG (Supplementary Table 2), and among these 138 genes, 108 genes significantly up-regulated and 4 genes (SDHA, SDHB, SDHD, COX6A2) were significantly decreased in HCC tissues compared with the normal ones (Supplementary Table 3), and the heatmap of their expression levels displays in Fig. 6A. The co-expression network of the differentially expressed genes (DEGs) shows a positive correlation between the DEGs, which enriched in the pathway of OXPHOS (Fig. 6B-D).
(A) The heatmap shows the expression of differentially expressed OXPHOS genes. (B) The biological process enrichment analysis of the DEGs (|logfc|>1) using the STRING database. (C) PPI network offers the interaction among the DEGs by using the STRING database. (D) The co-expression network of the DEGs. DEGs, differentially expressed genes; PPI, protein-protein interaction.
Consensus clustering of OXPHOS genes identified two clusters of HCC patients with different pathological features and prognosis model construction
Due to limited knowledge on the role of OXPHOS states in HCC progression, we explored OXPHOS molecular subtypes. Based on the expression of known OXPHOS genes, the clearest clustering was observed when k = 2 (Fig. 7A-H), with the highest intra-group correlation and minimal inter-group interference (Fig. S5A-C). This led to the classification of HCC patients into two sub-clusters: Cluster1 and Cluster2. Kaplan-Meier (KM) curves showed a significant survival advantage for Cluster1 (P = 0.013, Fig. 7I), and most DEGs were highly expressed in Cluster2 with more patients in severe grades (P < 0.05) and advanced stages (P < 0.001) (Fig. 7J).
We developed a risk signature using OXPHOS-related genes. A total of 1694 DEGs were selected via univariate Cox regression analysis (Supplementary Table 4), and a 10-gene signature was built through LASSO regression based on the optimal λ value (Fig. S5D, E). The method for calculating the risk score is outlined in Table 1.
To validate the signature, ROC curves were generated to assess predictive sensitivity and specificity, revealing an AUC of 0.816 for the risk score, higher than for age and gender. The AUCs for 1-year, 3-year, and 5-year survival were 0.725, 0.736, and 0.816, respectively (Fig. 7K, L). The signature outperformed age and stage in comprehensive prognosis prediction, as indicated by the C-index (Fig. S5F). KM curves (Fig. 7Q) showed worse prognosis for high-risk patients. Risk score and survival status curves (Fig. 7M) demonstrated a correlation between higher risk scores and increased mortality. PCA and t-SNE projections (Fig. 7N, O) confirmed the risk model’s effective clustering ability. A combined clinical prediction model (‘Clinical’) and the risk score model (‘Riskscore’) demonstrated a net benefit (NB) between 0.2 and 0.5 for risk score, suggesting the superior value of the gene signature over clinical factors in predicting outcomes (Fig. 7P). Univariate Cox regression identified the risk score as an independent prognostic factor, with multivariate analysis further confirming it after adjusting for confounders (Fig. 7R, S). A nomogram was created to aid in prognosis prediction (Fig. 7T), with high sensitivity and accuracy confirmed by calibration curves (Fig. 7U).
Consensus clustering of OXPHOS genes identified and prognosis model construction. (A-H) Consensus matrix plots for cluster numbers 2 through 9. (I) KM survival analysis comparing two clusters. (J) heatmap of DEG expressions of the two clusters and clinicopathological features between the two clusters. (K) ROC curves for risk score and clinicopathological features. (L) ROC curves for the prognostic signature at 1, 3, and 5 years. (M) Heatmap displaying OXPHOS-related gene expression in risk groups; scatterplot based on survival status; and risk curve by sample risk score. (N) PCA-based dot plot for low- and high-risk groups. (O) t-SNE plot. (P) DCA curves for different predicting models. (Q) KM curves for overall survival in high- and low-risk groups. (R) Univariate analysis. (S) Multivariate analysis. (T) Nomogram for prognosis prediction. (U) Calibration curves for predicting 1, 3, and 5-year OS. KM curve, Kaplan–Meier curve; OS, overall survival; DEG, differentially expressed genes; ROC curve, receiver operating characteristic curve; t-SNE, t-distributed stochastic neighbor embedding; PCA, principal component analysis; DCA, Decision Curve Analysis. **** P < 0.001.
Functional enrichment and evaluation of immune infiltration, TMB
The analysis of Tumor Mutational Burden (TMB), immune infiltration, and metabolic pathways is essential for a deeper understanding of HCC prognosis and treatment responses. TMB serves as a critical biomarker reflecting the mutational landscape of tumors, with higher TMB often correlating with a more aggressive disease and poorer survival outcomes, as shown in Fig. 8A. Combining TMB with the risk score (Fig. 8B) allows for more refined patient stratification into four groups, each exhibiting markedly different prognoses (P < 0.001), which can guide personalized treatment approaches.
Moreover, KEGG and GO analyses are key to uncovering the molecular mechanisms underlying these findings. KEGG analysis (Fig. 8C) revealed that the genes associated with the TMB and risk score are primarily linked to metabolic pathways, which are crucial for tumor growth and progression. GO analysis (Fig. 8D) further emphasized that these genes are primarily involved in metabolic functions, indicating that metabolic dysregulation is a significant feature of HCC.
Additionally, immune infiltration plays a central role in tumor progression and response to therapy. Figure 8E illustrates how risk scores are correlated with immune cell infiltration, particularly M2 macrophages, which are known to promote tumor progression and immune evasion. Higher immune function expression in the high-risk group (Fig. 8F) underscores the importance of immune responses in shaping the tumor microenvironment and influencing patient outcomes.
(A) The KM curves for the patients with high- and low-TMB. (B) The KM curves for patients with high TMB + high-risk score, high TMB + low-risk score, low TMB + high-risk score, and low TMB + low-risk score. (C) KEGG analysis of ten OXPHOS-related genes. (D) GO analysis of ten OXPHOS-related genes. (E) The heatmap of the correlation between risk score and tumor-infiltrating immune cells. (F) Differences in the immune functions between the two risk groups.
Discussion
The interplay between tumor metabolism and immune modulation is an emerging area of research, with growing evidence supporting the existence of mutual reprogramming between tumor cells and tumor-associated macrophages (TAMs)8. In this study, we uncover a mechanistic axis linking oxidative phosphorylation (OXPHOS) in hepatocellular carcinoma (HCC) tumor cells to M2 macrophage reprogramming via chemerin signaling and subsequent activation of ribosomal pathways.
Recent studies have highlighted the critical role of OXPHOS in sustaining tumor growth and immune evasion. Interestingly, while key components of the OXPHOS machinery, including SDHA, SDHB, SDHD, and COX6A2, are downregulated in HCC, OXPHOS activity appears to be maintained. This is consistent with findings from other studies showing that tumors can compensate for the loss of specific OXPHOS subunits by upregulating alternative metabolic pathways or other OXPHOS-related genes. Previous studies have demonstrated that alternative subunits of the NDUF family and ATP synthase are often upregulated in cancer cells, helping to preserve mitochondrial function despite the loss of key respiratory chain components9. Tumor cells with high OXPHOS activity often exhibit enhanced metabolic flexibility, enabling them to adapt to nutrient-deprived and hypoxic conditions in the tumor microenvironment (TME)10,11. Pharmacologically targeting OXPHOS, using inhibitors such as metformin, could potentially disrupt this metabolic flexibility, thereby weakening the tumor’s ability to adapt and promoting a less immunosuppressive TME by reducing immune cell suppression and enhancing anti-tumor immunity12. Previous study has been demonstrated that, in the invasive zone, macrophage recruitment and subsequent M2 polarization could be triggered, further enhancing HCC cell OXPHOS activity, establishing local immunosuppression, and potentially driving tumor progression13. On the other hand, OXPHOS upregulation has also been linked to drug resistance, tumor aggressiveness, and immunosuppressive TME formation, where highly metabolically active tumor cells modulate immune components to their advantage14. These findings provide a solid foundation for our observation that OXPHOS-active HCC tumor cells promote the recruitment and functional reprogramming of M2 TAMs.
The role of chemerin (encoded by RARRES2) in immune cell recruitment and polarization has been widely recognized in cancer research. Conventionally, chemerin is reported to promote macrophage polarization toward an M1-like phenotype with pro-inflammatory and anti-tumor functions, primarily through its interaction with the receptor CMKLR115,16. However, our findings diverge from this established perspective, demonstrating that OXPHOS-high HCC tumor cells secrete chemerin, which instead drives macrophage polarization toward the immunosuppressive M2 phenotype. This discrepancy underscores the context-dependent nature of chemerin signaling within the TME. Several explanations could account for this M2-polarizing effect in HCC. Post-translational modifications, such as protease-driven cleavage, are known to modulate chemerin activity and receptor specificity, possibly skewing its effects toward immune suppression in the TME17,18. Second, the unique immunosuppressive landscape of HCC, characterized by hypoxia, high interstitial pressure, and altered cytokine profiles, might reshape the functional response of macrophages to chemerin secretion19. These conditions could enhance the sensitivity of TAMs to M2-driving cues, thereby overriding chemerin’s conventional M1-polarizing effects.
Ribosomal biogenesis and translational activity are essential for macrophage functionality, particularly in the M2 subtype, given their dependence on protein synthesis to sustain pro-tumoral activities such as cytokine secretion, angiogenesis, and extracellular matrix remodeling20,21. Previous studies have shown that M2 macrophages exhibit heightened ribosomal activity compared to M1 macrophages, aligning with our finding of ribosomal pathway enrichment in the M2 subtype22. Furthermore, enhanced ribosomal activity in M2 macrophages has been linked to tumor-promoting roles in cancers23. This evidence supports the downstream effects of tumor signaling in activating ribosomal pathways and sustaining M2 macrophage polarization, as demonstrated in our study.
The robustness of the OXPHOS-ribosome-M2 polarization loop is further supported by published multi-omic studies that demonstrate metabolic crosstalk between tumor and immune cells at the spatial and functional levels. Metabolically active tumor cells are known to secrete immunomodulatory factors, influencing immune cell polarization and function24. In the case of M2 macrophages, studies have shown that tumor-derived metabolites (e.g., lactate, lipids) can skew macrophage polarization while simultaneously rewiring their metabolic activity25.
In HCC, tumor cells balance glycolysis and OXPHOS via metabolic flexibility to modulate M2 polarization. Spatial transcriptomics shows M2-proximal tumor cells have elevated OXPHOS (with upregulated OXPHOS-related genes) while retaining glycolytic activity, indicating coexistence of both pathways. OXPHOS specifically drives chemerin secretion to promote M2 polarization, as shown by in vitro experiments where OXPHOS inhibition reduces chemerin and M2 polarization. Glycolysis persists to support biosynthesis for proliferation. This is context-dependent: glycolysis may dominate in hypoxic, M2-distal regions (aiding immune suppression via lactate), while M2-proximal cells enhance OXPHOS for chemerin secretion while retaining glycolysis. This balance lets tumor cells adapt to the microenvironment and regulate M2 polarization via the OXPHOS-chemerin axis.
While our study provides compelling evidence for the OXPHOS-ribosome-M2 polarization axis, several limitations remain. First, chemerin’s specific mechanism in driving M2 polarization requires further investigation, particularly its interaction with CMKLR1 and potential downstream signaling pathways. Second, validation in larger, ethnically diverse patient cohorts is necessary to enhance generalizability. Third, independent validation of the OXPHOS-related risk model in external cohorts, such as those from ICGC, is needed to confirm its robustness and applicability. Fourth, in vivo models should be employed to better understand the temporal dynamics and causal relationships within this axis. Lastly, the interplay between this pathway and other metabolic or immune-regulatory mechanisms remains unexplored, highlighting the need for further studies to identify combinational strategies, such as targeting both OXPHOS and chemerin signaling, to disrupt M2-enriched TMEs.
In summary, our findings illuminate a novel metabolic-immune regulatory loop in HCC wherein OXPHOS-active tumor cells secrete chemerin to polarize macrophages into the M2 phenotype, enhancing their ribosomal activity and further sustaining tumor progression. This mechanism aligns with existing literature on TAM-driven metabolic plasticity and tumor immunity, adding mechanistic precision to current knowledge. Given the strong preclinical and clinical evidence supporting the relevance of this axis, the OXPHOS-ribosome-M2 polarization loop represents a promising therapeutic target in HCC and potentially other solid tumors.
Methods
Single-cell RNA sequencing data collection and analysis
Single-cell transcriptomic profiles and tumor stages of hepatocellular carcinoma (HCC) were acquired from publicly accessible repositories in the Gene Expression Omnibus (GEO) (GSE202642). Raw sequencing matrices were processed through the Seurat v5 pipeline26. Rigorous quality control excluded cellular barcodes exhibiting suboptimal sequencing metrics, including features < 100 or mitochondrial gene ratios > 25%. Transcriptomic normalization was performed via log-linear transformation, followed by identification of variably expressed genes for downstream dimensionality reduction. Principal component decomposition preceded systematic removal of heterotypic doublets using the DoubletFinder algorithm. Cross-sample batch correction was implemented through Harmony integration27.
Cellular identification
Based on Harmony-integrated data, we performed clustering analysis using the FindNeighbors and FindClusters functions (dims.use = 1:25), selecting an optimal resolution of 0.5 after evaluating multiple parameters. Cell distribution was visualized via Uniform Manifold Approximation and Projection (UMAP) for dimensionality reduction. Cellular identities were resolved through canonical lineage markers: NK cells and T cells (CD2, TRAC), B-lineage (CD79A, CD19), monocyte compartment (S100A8, S100A9), macrophages (CD14, CSF1R), proliferation macrophages (STMN1, TUBB), fibroblasts (ACTA2, PDPN), dendritic subsets (FCER1A, CLEC10A), vasculature (FLT1, EGFL7), hepatocytes and HCC cells (Liver_Tumorcells) (TF, FGB).
Macrophage polarization scoring
Cells were first subsetted to focus on macrophages by selecting the “Macrophages” cell type. To integrate the data and adjust for batch effects, the Harmony algorithm was applied, and dimensionality reduction was performed using the FindNeighbors function with 1 to 30 components. Outlier cells were manually selected by visual inspection using the DimPlot function with UMAP reduction. The CellSelector tool was used to interactively select and remove outlier cells, ensuring that only relevant macrophages were retained for further analysis. After outlier removal, the data were filtered, and the remaining macrophages were used for downstream analysis. To assess macrophage polarization, a set of genes characteristic of M1 (e.g., IL1B, TNF, CXCL10, STAT1) and M2 (e.g., IL10, TGFB1, CD163, ARG1) polarization states was used to calculate polarization scores. The AddModuleScore function in Seurat was employed to generate M1 and M2 scores, which were then normalized by scaling using the formula: M1_score_norm = (M1_score − mean(M1_score))/sd(M1_score); M2_score_norm = (M2_score − mean(M2_score))/sd(M2_score). The difference between M1 and M2 scores was calculated as: M1_M2_diff = M1_score_norm − M2_score_norm. Macrophages were classified into polarization states based on the M1_M2_diff score with the following thresholds: macrophages with M1_M2_diff > 0 were classified as M1 polarized, while those with M1_M2_diff < 0 were classified as M2 polarized. This classification was added as a new metadata column called polarization in the Seurat object, where each macrophage was labeled as either “M1” or “M2”.
Visualization, trajectory and enrichment analysis
For all visualizations, ggplot2, pheatmap, and other custom plotting functions were employed to generate UMAP, t-SNE, and trajectory plots of macrophage polarization and trajectories. Statistical analysis, including density plots and violin plots, was performed to examine the differences in gene expression and polarization scores. All plots were annotated to highlight the changes in polarization states and the association with different tumor stages (Normal, BCLC-A, and BCLC-B). Trajectory analysis was performed using Monocle328, reducing the dataset to three components with reduce_dimension and clustering cells using cluster_cells. Trajectories were visualized in reduced space using plot_cells, colored by polarization state, to identify key genes and pathways involved in the transition from M1 to M2 polarization in HCC. Developmental genes influencing the trajectory were identified using graph_test with the principal_graph neighbor graph. Significant genes, such as CD163, CD206, ARG1, IL10, VEGF, and CSF1R, were visualized in pseudotime to examine expression changes across polarization stages. At the same time, KEGG pathway enrichment was performed on DEGs using enrichKEGG and visualized with dot plots. Results were saved to a file. Further enrichment was done with MSigDB KEGG gene sets using enricher, and the top pathways were also visualized in a dot plot.
Spatial transcriptomics
The spatial transcriptomics data was obtained from GEO (GSE238264). The analysis starts by loading 10X RNA-seq and spatial transcriptomic data, creating Seurat objects, and performing SCTransform normalization. The datasets are merged, and PCA, clustering, and UMAP are used for dimensionality reduction and visualization. For single-cell data, 5000 cells are randomly selected, and common features are identified for integration. SCT normalization is applied, and FindTransferAnchors and TransferData are used to transfer cell type annotations from the single-cell dataset to the spatial data, with predicted cell types stored in the metadata. For each cell type in the single-cell RNA-seq dataset, marker gene enrichment analysis was performed using the enricher function. Genes specific to each cell type were compared to predefined gene sets, and adjusted p-values were calculated for enrichment significance.
To investigate the spatial relationship between liver tumor cells (“Liver_Tumorcells”) and M2 macrophages (“M2”), the nearest-neighbor distances were calculated using their spatial coordinates. A threshold of 30 units was set to define whether tumor cells were spatially “near” or “not near” M2 cells. Based on this classification, liver tumor cells were grouped into two subsets (“near_M2” and “not_near_M2”) for downstream differential expression analysis. DEGs between these two groups were identified using a threshold of logFC > 0.5 and p-value < 0.05. Significant DEGs were further subjected to functional enrichment analysis, including Gene Ontology (GO) analysis, using the clusterProfiler package. Enriched functional categories with adjusted p-values (p.adjust) < 0.05 were considered statistically significant. Bar plots and bubble plots were generated to visualize the results.
Cell culture
Human HCC cell lines Huh-7 were the generous gifts from Dr. Zhou (Chongqing Medical University) and cultured in DMEM containing 10% fetal bovine serum (FBS), and 1% penicillin streptomycin. And Human monocytic leukemia THP-1 were purchased from Sunncell Biotech (Wuhan, China), and cultured in RPMI1640 plus 10% FBS and 1% penicillin streptomycin. Macrophage differentiation was achieved by treating THP-1 cells with 5 ng/ml PMA for a duration of 48 h. For co-culture system, Huh7 cells or macrophages were seeded in the lower compartment of the Boyden chamber, while other cells were placed in the upper compartment.
Western blot
Protein lysates were obtained by lysing cells and subjected to electrophoresis on sodium dodecyl sulfate-polyacrylamide gels (Bio-Rad). The separated proteins were transferred onto polyvinylidene fluoride (PVDF) membranes, which were then blocked with 5% skim milk. Membranes were incubated with primary antibodies in blocking buffer overnight at 4 °C, followed by incubation with horseradish-peroxidase-conjugated secondary antibodies (Proteintech) at room temperature for 1 h. Protein bands were visualized using enhanced chemiluminescence detection reagents, and signals were captured with an ECL blotting analysis system (Bio-OI, Guangzhou, China). ImageJ 2.0.0 (NIH) was used to quantify band intensity. Western blot experiments were repeated twice, producing consistent results. β-tubulin and β-actin served as the endogenous loading control. The following antibodies were used: anti-CD206 monoclonal antibody (1:500; Proteintech; 2A6A10), anti-ARG-1 polyclonal antibody (1:1000; Proteintech; 16001-1-AP), anti-iNOS polyclonal antibody (1:1000; Proteintech; 22226-1-AP), anti-β-tubulin monoclonal antibody (1:500; Beyotime; AF2835), anti-ATP8A1 polyclonal antibody (1:1000; HUABIO; ER1902-61), anti-Chemerin recombinant antibody (1:1000; Proteintech; 83550-4-RR), anti-β-actin monoclonal antibody (1:500; Beyotime; AF0003), anti-UBTF polyclonal antibody (1:1000; Proteintech; 20660-1-AP), anti-UBTF polyclonal antibody (1:1000; Proteintech; 20660-1-AP), and anti-CMKLR1 polyclonal antibody (1:1000; HUABIO; ER63003).
RNA extraction and real-time PCR analysis
Total RNA was extracted using AG RNAex Pro RNA reagent (Accurate Biology, AG21101), and RNA concentrations were measured by absorbance at 260/280 nm. Complementary DNA (cDNA) was synthesized using a cDNA synthesis kit (Accurate Biology, AG11705). Real-time PCR was performed on the ABI 7300 System (Applied Biosystems, CA, USA) using SYBR Green Pro Taq HS (Accurate Biology, AG11759) to measure mRNA expression. Primers for CD36, CD163, and β-actin are listed in Supplementary Table 5. Each 25 µl reaction contained Maxima SYBR Mix, primers (400 nM), and 100 ng cDNA. Cycling conditions included 50 °C for 2 min, 95 °C for 10 min, followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min. Gene expression was calculated using the 2−ΔΔCT method and normalized to β-actin levels.
Enzyme-linked immunosorbent assay (ELISA)
Enzyme-linked immunosorbent assay (ELISA) kits were used to quantify the levels of tumor necrosis factor TNF-α, IL-1β, IL-6, transforming growth factor (TGF)-β (eBioscience, San Diego, CA, USA). All procedures adhered to the manufacturer’s guidelines.
Immunohistochemical (IHC) analysis
Tumor, margin (1 cm from tumor edge), and para-tumor (≥ 2 cm from edge) tissues from six HCC patients were collected at The First Affiliated Hospital of Chongqing Medical University with informed consent and IRB approval, in accordance with the Declarations of Helsinki and Istanbul. Tissues were fixed in 4% paraformaldehyde, embedded in paraffin, and sectioned at 4 μm. Sections were deparaffinized, rehydrated, and underwent antigen retrieval in citrate buffer (pH 6.0) for 20 min. After blocking with 5% BSA for 30 min, primary antibodies were applied overnight at 4 °C, followed by HRP-conjugated secondary antibodies for 1 h at room temperature. DAB was used for detection, with hematoxylin counterstaining. The IHC score was determined by multiplying staining intensity (0–3) by the proportion of positive cells (0–4), where intensity ranged from colorless (0) to dark brown (3), and positivity from 0% (0) to 76–100% (4).
Immunofluorescence staining
Immunofluorescence (IF) staining of cells were performed according to standard protocols. In brief, macrophages cultured in RPMI1640 plus 10% FBS and 1% penicillin streptomycin or Huh7 cell-conditioned medium on coverslips were washed with ice-cold PBS and fixed with methanol. Cells were permeabilized with PBS containing 0.2% Triton X-100. The cells expressing iNOS, CD206 were incubated with anti-CD206 mouse monoclonal antibody (1:200, Proteintech; 2A6A10), anti-iNOS polyclonal antibody (1:200; Proteintech; 22226-1-AP) for overnight followed by FITC-Affini Pure Goat Anti-Mouse IgG (1:200, Yeasen, 33207ES60) and Rhodamine (TRITC) AffiniPure Goat Anti-Rabbit IgG (1:200, Yeasen, 33109ES60) for 1 h. The coverslips were mounted in DAPI-containing mounting medium (Abcam, USA) and examined by a confocal laser-scanning microscope (CLSM, Leica TCS SP8, Mannheim, Germany).
Flow cytometry
Macrophages were suspended in PBS with 2% FBS, fixed, and stained with CD86, then washed, permeabilized, and stained with CD206 antibodies. CD86 and CD206 expression was analyzed using a BD flow cytometer, with data processed via FlowJo.
Consensus clustering analysis and prognostic signature construction
RNA-sequencing data (50 normal liver tissues and 369 HCC tissues) along with clinicopathological details for the TCGA-LIHC cohort were retrieved from The Cancer Genome Atlas (TCGA) (https://cancergenome.nih.gov/). OXPHOS-related genes were identified via KEGG, as detailed in Supplementary Table S1. Differentially expressed OXPHOS genes between normal and tumor tissues were analyzed using the “limma” R package with a threshold of P < 0.05. The “igraph” R package29 was used to construct the co-expression network of these DEGs, while the STRING database (https://string-db.org/) was employed to build and visualize the protein-protein interaction (PPI) network for OXPHOS-related DEGs. Consensus clustering of HCC patients was performed using the “ConsensusClusterPlus” R package30 based on RNA expression of OXPHOS DEGs, employing the K-means algorithm repeated 50 times for robust OXPHOS subtyping. The optimal number of clusters was determined using cumulative distribution function (CDF), delta area, and cluster-consensus (CLC). HCC patients were divided into two sub-clusters, and overall survival (OS) between these subtypes was compared via Kaplan–Meier analysis using “survival” and “survminer” R packages. Clinical and pathological differences between the two sub-clusters were assessed with the “limma,” “pheatmap,” and “ggpubr” R packages, while prognosis was analyzed using “survival” and “survminer.” DEGs between the sub-clusters (|log2FC| > 1, FDR < 0.001) were identified using “limma.” Functional enrichment of DEGs, including Gene Ontology (GO) and KEGG pathway analyses, was carried out using the “clusterProfiler” R package.To facilitate clinical application, univariate Cox regression analysis (R package “survival”) was used to evaluate correlations between DEG expression and survival. A prognostic signature was developed using LASSO regression (R package “glmnet”31) with the formula: risk score = Σ (βi × Expi) (β: coefficient, Exp: RNA expression level). Based on the risk score, HCC patients were categorized into low- and high-risk groups.
Assessing the predictive power of the prognostic model for HCC
To assess the predictive power of the prognostic model for HCC, receiver operating characteristic (ROC) curves were generated using the “survivalROC” R package to evaluate overall survival (OS) performance across various clinicopathological factors and the risk model. The concordance index (C-index), calculated using the “pec” R package, was employed to compare the model’s accuracy with clinicopathological characteristics. Kaplan-Meier (KM) survival curves illustrated OS differences between high- and low-risk subgroups. Principal component analysis (PCA) and t-distributed stochastic neighbor embedding (t-SNE) were performed using the “Rtsne” R package to visualize group distributions. Univariate and multivariate Cox regression analyses (“Survival” R package) evaluated the prognostic value of the risk score. A nomogram integrating the risk score and clinical variables was developed using the “rms” R package to predict 1-, 3-, and 5-year OS for HCC patients.
Analyses of relation between tumor mutation burden, and prediction of the effective response of postoperative immunotherapy
Seven algorithms (TIMER, XCELL, MCPcounter, QUANTISEQ, CIBERSORT, EPIC, and CIBERSORT-ABS) were applied to estimate the infiltration status of different types of immune cells between the two risk groups32. And ssGSEA was performed to explore the different infiltration degrees of immune checkpoints in the two risk groups using the R package “GSVA”33. The somatic mutation data (MuTect2 Variant Aggregation and Masking) was downloaded from the TCGA-LIHC dataset, and the frequency of gene mutation in specific patients was calculated by the R package “maftools”34. And tumor mutation burden (TMB) was defined as the total number of somatic mutations per million bases and analyzed by the “maftools” package, too. The KM analysis was conducted to reveal different OS based on the different TMB groups.
Statistical analysis
Statistical analysis was performed using R Studio (version 4.4.2). Continuous variables, such as gene expression levels, were analyzed using one-way analysis, while Pearson’s chi-square test was applied to categorical variables. Kaplan-Meier analysis was used to compare OS between groups. Univariate and multivariate Cox regression evaluated the prognostic value of the risk model. Spearman correlation assessed the relationships between TMB, risk scores, single-gene expression, and immune checkpoint expression. Two-way ANOVA (GraphPad Prism 10) was used for quantitative western blot analysis.
Data availability
Sequence data that support the findings of this study were obtained from GEO database (GSE202642, GSE238264) and TCGA-LIHC, and the code is available upon request from the correspondence author. KEGG analysis.
References
Llovet, J. M. et al. Hepatocellular carcinoma. Nat. Rev. Dis. Primers. 7, 6. https://doi.org/10.1038/s41572-020-00240-3 (2021).
Wu, T. & Dai, Y. Tumor microenvironment and therapeutic response. Cancer Lett. 387, 61–68. https://doi.org/10.1016/j.canlet.2016.01.043 (2017).
Kieler, M., Hofmann, M. & Schabbauer, G. More than just protein Building blocks: how amino acids and related metabolic pathways fuel macrophage polarization. Febs j. 288, 3694–3714. https://doi.org/10.1111/febs.15715 (2021).
Zhu, L., Zhao, Q., Yang, T., Ding, W. & Zhao, Y. Cellular metabolism and macrophage functional polarization. Int. Rev. Immunol. 34, 82–100. https://doi.org/10.3109/08830185.2014.969421 (2015).
Mayer, K. A. et al. The energy sensor AMPK orchestrates metabolic and translational adaptation in expanding T helper cells. Faseb j. 35, e21217. https://doi.org/10.1096/fj.202001763RR (2021).
Gubin, M. M. et al. High-Dimensional analysis delineates myeloid and lymphoid compartment remodeling during successful Immune-Checkpoint cancer therapy. Cell 175, 1014–1030e1019. https://doi.org/10.1016/j.cell.2018.09.030 (2018).
Metge, B. J. et al. Ribosomal RNA biosynthesis functionally programs Tumor-Associated macrophages to support breast cancer progression. Cancer Res. https://doi.org/10.1158/0008-5472.Can-24-0707 (2025).
Huang, J., Wu, Q., Geller, D. A. & Yan, Y. Macrophage metabolism, phenotype, function, and therapy in hepatocellular carcinoma (HCC). J. Transl Med. 21, 815. https://doi.org/10.1186/s12967-023-04716-0 (2023).
Galber, C., Acosta, M. J., Minervini, G. & Giorgio, V. The role of mitochondrial ATP synthase in cancer. Biol. Chem. 401, 1199–1214 (2020).
Zheng, J. Energy metabolism of cancer: Glycolysis versus oxidative phosphorylation (Review). Oncol. Lett. 4, 1151–1157. https://doi.org/10.3892/ol.2012.928 (2012).
Greene, J., Segaran, A. & Lord, S. Targeting OXPHOS and the electron transport chain in cancer; molecular and therapeutic implications. Semin Cancer Biol. 86, 851–859. https://doi.org/10.1016/j.semcancer.2022.02.002 (2022).
Huang, X. et al. Metformin reprograms Tryptophan metabolism to stimulate CD8 + T-cell function in colorectal cancer. Cancer Res. 83, 2358–2371. https://doi.org/10.1158/0008-5472.Can-22-3042 (2023).
Wu, L. et al. An invasive zone in human liver cancer identified by Stereo-seq promotes hepatocyte-tumor cell crosstalk, local immunosuppression and tumor progression. Cell. Res. 33, 585–603. https://doi.org/10.1038/s41422-023-00831-1 (2023).
Ashton, T. M., McKenna, W. G., Kunz-Schughart, L. A. & Higgins, G. S. Oxidative phosphorylation as an emerging target in cancer therapy. Clin. Cancer Res. 24, 2482–2490. https://doi.org/10.1158/1078-0432.Ccr-17-3070 (2018).
Goralski, K. B. et al. Chemerin, a novel adipokine that regulates adipogenesis and adipocyte metabolism. J. Biol. Chem. 282, 28175–28188. https://doi.org/10.1074/jbc.M700793200 (2007).
Ji, Z. S. et al. Chemerin promotes the pathogenesis of preeclampsia by activating CMKLR1/p-Akt/CEBPɑ axis and inducing M1 macrophage polarization. Cell. Biol. Toxicol. 38, 611–628. https://doi.org/10.1007/s10565-021-09636-7 (2022).
Zabel, B. A. et al. Chemoattractants, extracellular proteases, and the integrated host defense response. Exp. Hematol. 34, 1021–1032. https://doi.org/10.1016/j.exphem.2006.05.003 (2006).
Li, W., Li, F., Zhang, X., Lin, H. K. & Xu, C. Insights into the post-translational modification and its emerging role in shaping the tumor microenvironment. Signal. Transduct. Target. Ther. 6, 422. https://doi.org/10.1038/s41392-021-00825-8 (2021).
Blitz, S. E. et al. Tumor-Associated Macrophages/Microglia in glioblastoma oncolytic virotherapy: A Double-Edged sword. Int. J. Mol. Sci. 23 https://doi.org/10.3390/ijms23031808 (2022).
Zhang, L. et al. Ribosomal protein S3A (RPS3A), as a transcription regulator of colony-stimulating factor 1 (CSF1), promotes glioma progression through regulating the recruitment and autophagy-mediated M2 polarization of tumor-associated macrophages. Naunyn. Schmiedebergs Arch. Pharmacol. https://doi.org/10.1007/s00210-024-03601-x (2024).
Hu, A. et al. Therapeutic targeting of CNBP phase separation inhibits ribosome biogenesis and neuroblastoma progression via modulating SWI/SNF complex activity. Clin. Transl Med. 13, e1235. https://doi.org/10.1002/ctm2.1235 (2023).
Tabatabaei, N., Hou, S., Kim, K. W. & Tahmasebi, S. Signaling pathways that control mRNA translation initiation in macrophages. Cell. Signal. 73, 109700. https://doi.org/10.1016/j.cellsig.2020.109700 (2020).
Donati, G., Montanaro, L. & Derenzini, M. Ribosome biogenesis and control of cell proliferation: p53 is not alone. Cancer Res. 72, 1602–1607. https://doi.org/10.1158/0008-5472.Can-11-3992 (2012).
Kumar, S. et al. Metabolic reprogramming in tumor-Associated macrophages in the ovarian tumor microenvironment. Cancers 14, 5224 (2022).
Noe, J. T. et al. Lactate supports a metabolic-epigenetic link in macrophage polarization. Sci. Adv. 7, eabi8602. https://doi.org/10.1126/sciadv.abi8602 (2021).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304. https://doi.org/10.1038/s41587-023-01767-y (2024).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with harmony. Nat. Methods. 16, 1289–1296. https://doi.org/10.1038/s41592-019-0619-0 (2019).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502. https://doi.org/10.1038/s41586-019-0969-x (2019).
Mora, A. & Donaldson, I. M. iRefR: an R package to manipulate the iRefIndex consolidated protein interaction database. BMC Bioinform. 12, 455. https://doi.org/10.1186/1471-2105-12-455 (2011).
Wilkerson, M. D. & Hayes, D. N. ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics 26, 1572–1573. https://doi.org/10.1093/bioinformatics/btq170 (2010).
Engebretsen, S. & Bohlin, J. Statistical predictions with Glmnet. Clin. Epigenetics. 11, 123. https://doi.org/10.1186/s13148-019-0730-1 (2019).
Li, T. et al. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res. 48, W509–w514. https://doi.org/10.1093/nar/gkaa407 (2020).
Chen, Y. et al. A novel Immune-Related gene signature to identify the tumor microenvironment and prognose disease among patients with oral squamous cell carcinoma patients using ssgsea: A bioinformatics and biological validation study. Front. Immunol. 13, 922195. https://doi.org/10.3389/fimmu.2022.922195 (2022).
Mayakonda, A., Lin, D. C., Assenov, Y., Plass, C. & Koeffler, H. P. Maftools: efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 28, 1747–1756. https://doi.org/10.1101/gr.239244.118 (2018).
Funding
This study was sponsored by Natural Science Foundation of Chongqing, China (cstc2021jcyj-msxmX0292) and Program for Youth Innovation in Future Medicine from Chongqing Medical University.
Author information
Authors and Affiliations
Contributions
All authors contributed to the work presented in this paper. Conceptualization, Y.N. AND Q.L.; dataset resourcing, analysis, validation and visualization, Y.N., X.C., AND B.Z.; writing—original draft, Y.N. and W.; review and editing, Q.L.; supervision, C.D.; experiment achievement, Y.N. and Z.L. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Nong, Y., Chen, X., Zhang, B. et al. M2 macrophages and tumor cells engage in a metabolic feedback loop to drive HCC progression. Sci Rep 15, 41701 (2025). https://doi.org/10.1038/s41598-025-25673-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-25673-2
Keywords
This article is cited by
-
HMGA2 promotes hepatocellular carcinoma progression by regulating tumor-associated macrophage via Notch1/CCL2 signaling
Molecular and Cellular Biochemistry (2026)