Abstract
The gastrointestinal tract (GIT) of cattle plays a vital role in nutrient absorption, immune function, and microbial homeostasis. While the importance of the GIT microbiome and epithelial barrier integrity has been increasingly recognized, the typical composition of microbial communities and the expression of tight junction proteins (TJPs) in feedlot cattle remains poorly characterized. We investigated microbial community structure and TJP expression at three GIT sites: the rumen (RU), small intestine (SI), and large intestine (LI) in 21 finish-fed feedlot steers sourced from 21 commercial feedyards in the Texas Panhandle. Samples of luminal contents and GIT tissue were collected from each region, as well as feces and liver abscess material. Microbial communities were characterized using 16S rRNA gene sequencing. TJP gene expression was quantified by RT-qPCR using synthetic standards, and protein expression was evaluated by immunohistochemistry (IHC) with both computer-generated and pathologist-generated scoring. Microbial community structures varied primarily by GIT region rather than by individual animals raised at different locations. Nine bacterial families were identified as core microbiome members, with Lachnospiraceae being the most abundant across the GIT. TJP gene expression varied considerably by site, with RU having significantly lower Claudin 1, Claudin 2, and E-Cadherin expression than the SI and LI. IHC results paralleled qPCR findings, with region-specific patterns of protein localization and intensity. Computerized and pathologist-generated H-scores showed moderate agreement but differed notably between epithelial and lamina propria regions. This study provides a comprehensive baseline of microbial and host factors associated with gut health in a uniquely diverse population of feedlot cattle. The identification of regional microbial communities and distinct TJP expression patterns offers foundational insights into gastrointestinal physiology and barrier function. This work establishes baseline data to support future investigations into the relationships among microbial ecology, epithelial barrier function, and cattle health and productivity.
Similar content being viewed by others
Introduction
The gastrointestinal tract (GIT) of cattle is a complex organ system with varied functions at different sites, all composed of structural cells, immune cells, and the associated microbiome. Functions related to digestion are well-described1, but recent research has begun to highlight the systemic influence on immune function associated with the GIT2,3. Correspondingly, the concepts of “gut health” have become more common in both human and animal health. However, little is known about the features that characterize a typical or “healthy” GIT in feedlot cattle. In the simplest terms, GIT health is directly related to the ability to digest and absorb nutrients while also providing barrier function, or the ability to allow some things (i.e., nutrients) to cross the epithelium while preventing others (i.e., pathogens) from crossing4. The maintenance of the GIT barrier is a complex biological function highly affected by tissue integrity, exemplified by tight junction proteins (TJPs), and the gastrointestinal microbiome5.
Tight junctions, or gap junctions, are the microscopic spaces between intestinal epithelial cells that regulate the passage of molecules across the epithelium6,7. There are several types of proteins contributing to this barrier from various protein families, including claudin, occludin, zona occludin, and cadherin8,9. However, the functions of individual proteins are complex; some promote connectivity between cells, while others promote leakiness10. Several TJPs are complementary and provide redundancy in their function9. Additionally, because TJPs are host-derived, they are influenced by luminal contents (e.g., pH, short-chain fatty acids—SCFA) and the microbiome.
The microbiome of cattle has long been recognized as influencing health. While the rumen and fecal microbiomes have been well characterized, the microbiota found in other gastrointestinal (GIT) sites remain less understood11. Furthermore, many studies have concentrated on young calves or lactating dairy cows12,13,14, but few have investigated the microbiomes of finish-fed beef animals produced in typical North American production systems. Annually, the United States feeds approximately 25 million cattle15, with approximately 6 million fed in the Southern Plains16. Despite the limited populations mentioned above, differences in the microbiomes found in cattle have been associated with dietary factors, production regions, and individual animals12,17. There is a growing impetus to use feed supplements in cattle to modify the microbiome for potential health benefits18, but there is a need to characterize core microbiome constituents and functions to provide a logical context for evaluating the utility of these products17.
Therefore, the objective of this study was to characterize the gastrointestinal microbiota and host tight junction protein (TJP) expression in clinically healthy, high-performing feedlot steers harvested in the Texas Panhandle. Specifically, we aimed to describe the composition and core members of microbial communities at distinct GIT sites (rumen, jejunum, and colon), and to quantify both protein and gene expression of select TJPs within corresponding tissues.
Materials and methods
Study overview
Feedlot steers finish-fed in 21 commercial feedlots in the Texas Panhandle were sampled at the time of harvest to characterize the microbiome and TJP characteristics at 3 GIT locations (rumen, jejunum, and colon). Additionally, fecal samples and, for cattle affected with liver abscesses, purulent material was collected. Microbial community compositions were characterized using 16S rRNA gene sequencing. Additionally, RNA extracted from epithelial surface scrapings was analyzed using qPCR to assess the expression of various TJP genes. Finally, tissue samples were evaluated for TJP expression using immunohistochemistry (IHC).
Study population
Samples for this study were obtained from steers that were finish-fed at 21 different large commercial feedlots (1 steer per feedlot) and harvested on the same day at the West Texas A&M University Caviness Meat Science & Innovation Center (USDA Est. #7124) as part of the Fed Beef Challenge competition sponsored by the Texas Cattle Feeders Association. Ownership of cattle was maintained by the feedlots from live animal production at their facilities through the harvest of saleable beef products. For this contest, managers submit one animal from each participating feedlot into a competition that ranks animals based on the total value of all harvested beef products. This provided a unique study population that was broadly representative of economically valuable cattle targeted in commercial beef production in the U.S. Detailed histories regarding the study subjects were not available, but animals were judged visually to likely have broad genetic diversity, and producers reported that all cattle were fed diets that were standard for animals raised on their respective premises. Given our knowledge of feedlot cattle production in the Texas Panhandle and their appearance, it is likely that these cattle were crossbred animals, although no genetic testing was conducted. The feedlot dietary rations at participating feedlots were formulated by nutritionists, and steam-flaked corn was the primary energy source. All animals were visually assessed by trained observers, including an inspector from the USDA Food Safety Inspection Service (FSIS), as being clinically healthy at the time of slaughter. Euthanasia and harvest processes were conducted under the supervision of the USDA–FSIS inspector using processes approved by that agency.
Sample collection
Cattle were rendered insensible by personnel employed by the abattoir using a captive bolt gun, and harvest procedures followed USDA-approved, industry-standard methods that are typically used in commercial abattoirs in the production of beef products intended for human consumption. Researchers were only involved in sampling after animals had been euthanized. Prior to hide removal, fecal samples were collected directly from the rectum using a clean plastic sleeve and glove for each animal. Samples were placed in sterile bags and stored on ice until further processing. Standardized carcass processing methods were applied uniformly across animals. Following hide removal and application of hot water and organic acid sanitizing of carcasses, approximately 20–30 min post-stunning, thoracic and abdominal viscera were removed. The entire gastrointestinal tract (GIT) was then transferred to a stainless-steel table in a room physically separated from the harvest floor.
The ventral sac of the rumen was opened, and 5 mL of the liquid phase of the luminal contents were collected in a sterile conical tube. Next, a 20 cm2 section of the rumen wall was removed using sterile forceps and a sterile disposable scalpel (approximately 20 cm from the cranial pillar). This was cut into 1 × 1 cm sections: the most cranial section was aseptically placed in a sterile 5 mL conical tube and flash frozen in an ethanol bath on dry ice, moving caudally another section was placed in a sterile 5 mL conical tube and stored on ice until further processed, and finally the two most caudal sections were pinned to Styrofoam and submerged in 10 × volume of 10% neutral buffered formalin. The same process was used to collect luminal and tissue samples from the anti-mesenteric jejunum, approximately 100 cm from the duodenum-abomasal junction. Finally, luminal and tissue samples were collected from the colon approximately 30 cm distal to the cecum as described above. Abscesses were identified in the livers of 3 animals (10 total abscesses: 2 animals had 2 abscesses each, and 1 animal had 6 abscesses); these livers were placed on ice in sterile bags at the time of evisceration until further processed.
Within 3 h of collection, samples were transported to the adjacent Texas A&M University VERO building. Sterile disposable spatulas were used to transfer approximately 4 ml of feces to sterile 5 mL conical tubes. Samples of feces, luminal contents, and tissue that were flash-frozen and those collected on ice were frozen at − 80 °C. Livers were sprayed with 90% ethanol and flamed to sterilize the external surfaces. Abscesses were incised using a sterile disposable scalpel, purulent material was collected using a sterile disposable spatula and placed in sterile cryotubes (2 mL Nalgene Cryogenic Tubes; Thermo Fisher Scientific, Waltham, MA), then frozen at − 80 °C. Tissue collected in formalin was fixed for 7 d, and then paraffin-embedded and processed for histology and immunohistochemistry at the Texas A&M University Gastrointestinal Laboratory.
DNA and RNA isolation
Epithelial surface scrapings were obtained by placing the tissue on a sterile surface and gently scraping the epithelial layer using a sterile disposable spatula. Approximately 100 mg feces, 100 mg epithelial surface scrapings, or 100 μL of luminal contents were used as input for the DNA extraction process. Samples were lysed prior to extraction using bead beating (Mini BeadBeater 24, BioSpec) with 5 × 1 min pulses (3,450 rpm) and 5 min rests in between each pulse. DNA was then extracted from these epithelial surface samples and the luminal contents using a commercial kit (QIAamp Power Soil Pro DNA, Qiagen, Hilden, Germany) with an automated nucleic acid extraction system (QIAcube Connect, Qiagen), following the manufacturer’s instructions. Recovered DNA was quantified (ng/µL) using fluorometry (Qubit Flex, Thermo Fisher Scientific). Extraction blanks were included when processing every batch of samples (n = 7) and included in downstream library preparation and sequencing.
Total RNA was extracted from flash-frozen tissue samples for investigation of TJP gene expression. Briefly, frozen tissue was placed on a sterile surface and approximately 20–30 mg of full-thickness tissue (i.e., serosa to epithelial surface) was excised using a sterile scalpel, placed in a sterile tube in an ice bath, then quickly moved into RNA isolation using a commercially available kit (RNeasy Mini, Qiagen) following the manufacturer’s instructions. Spectrophotometry (Nanodrop Eight, Thermo Fisher) and gel electrophoresis (TapeStation 4200, Agilent, Santa Clara, CA) were used to assess the quantity (ng/µL) and quality (RNA integrity number—RIN) of the RNA obtained from extractions.
16S rRNA amplicon library preparation and sequencing
The V3-V4 region of the 16S rRNA gene was amplified using the 341F/785R primer pair as previously described19. Steps for DNA amplification were conducted at 98 °C for 3 min, 98 °C for 30 s for 18 cycles, 72 °C for 1 min, and then 72 °C for 5 min. Amplification libraries were prepared according to Illumina’s protocol20 and pooled for sequencing in equimolar proportions. Then the pooled library was sequenced using an Illumina NovaSeq 6000 instrument using 2 × 250 base pair (bp) paired-end chemistry at the North Texas Genome Center with a targeted average sequencing depth of 1,100,000 paired-end reads per sample. Extraction blanks and no-template PCR controls (nuclease-free water) were included as negative controls and did not yield any detectable product. Thus, they were removed from downstream statistical analysis.
Bioinformatics
Demultiplexed sequencing reads were imported into QIIME2 version 2023.221. Amplicon sequencing variants (ASVs) were generated using DADA222. DADA2 also filters for read quality, removes chimeric sequences, and merges overlapping paired-end reads. Forward reads were trimmed at 17 bp, reverse reads at 21 bp, and all reads were truncated at 230 bp. Taxonomy was assigned using a Naïve Bayes classifier trained on SILVA 138.1 SSU NR 9923, and sequences were trimmed to only include the V3-V4 region delimited by the 341F/785R primer pair. Reads mapping to chloroplasts and mitochondria were removed from the ASV table and representative sequences. A mid-point rooted phylogenetic tree was built using ‘qiime alignment mafft’, ‘qiime alignment mask’, and ‘qiime phylogeny fasttree’ under default settings. The ASV table, representative sequences, and mid-point rooted tree were then imported to phyloseq24 using the ‘import_biome’ function. Finally, using the ‘import_qiime_sample_data’ function, metadata were imported and merged with the ASV table, representative sequences, and tree into a phyloseq object.
Richness and Faith’s observed phylogenic distance were quantified using the ‘estimate_richness’ and ‘estimate pd’ functions of the phyloseq and btools packages, respectively. Additionally, ASV counts were normalized using total sum scaling based on the median number of reads. Beta diversity was analyzed using generalized UniFrac distances25,26 and these distances were used for ordination plotting using non-metric multidimensional scaling (NMDS). Differences in community structure were tested by permutational multivariate analysis of variance (PERMANOVA) using the ‘vegan’27 and ‘pairwiseAdonis’ packages28. Additionally, a permutational analysis of dispersion (PERMDISP) was conducted for all PERMANOVA analyses to ensure differences were not generated from unequal dispersions of variance between groups. Using the ‘hclust’ function, hierarchical clustering was performed using Ward’s agglomeration method29 on the generalized UniFrac distances and plotted with the ggdendro package to create dendrograms. Relative abundances (RA) were calculated and plotted using phyloseq. Core members of the microbiome were characterized at the family level using the microbiome package in R, and a minimum detection threshold was set at 0.1% RA with the minimum prevalence set at 90%.
Finally, non-transformed counts were used to conduct an analysis of compositions of microbiomes with bias correction (ANCOM-BC30) to test for differential abundance. Counts were pre-filtered to require at least an average of 10 counts per sample per family for inclusion in the ANCOM-BC model. The ANCOM-BC model included a fixed effect of body site (i.e. rumen, small intestine, large intestine, or feces) and no random effects. Liver abscess samples were excluded from the ANCOM-BC analysis. All comparisons were made to the fecal samples under the default settings of ANCOM-BC.
Immunohistochemistry for tight junction proteins
After 7 d of fixation in 10% neutral buffered formalin, tissue samples were processed and embedded in paraffin blocks by the Texas A&M University Gastrointestinal Laboratory (College Station, Texas, USA). Five-micrometer serial sections were obtained. One section was routinely stained with hematoxylin and eosin using an automated stainer (Gemini AS, Epredia, Kalamazoo, MI, USA). For immunohistochemistry, slides were processed using an automated platform (Leica Bond RXm, Leica Biosystems, Deer Park, IL, USA). Targets were Claudin 1 (Abcam, Cat. #ab15098), Claudin 2 (Invitrogen, Cat. #51-6100), Occludin (Invitrogen, Cat. #33-1500, Clone OC-3F10), CDH1 herin (BD Biosciences, Cat. #610181), and MAC-387 which is also known as calprotectin (Invitrogen, Cat. #MA1-80446). Complete methodology for each marker can be found in Table S1. Slides were scanned using a NanoZoomer S60 (Hamamatsu Photonics, Shizuoka, Japan).
Stained images were scored both objectively and subjectively. For objective scoring, images were uploaded into QuPath version 0.5.131 to generate pixelwise H scores. Pixelwise H scores are an objective measure of biomarker staining measured by computer systems at the pixel level. To generate the scores, areas of the epithelial surfaces were selected by a trained operator (JDY) with supervision by a board-certified pathologist (JPC) to exclude luminal contents and processing artifacts. Tissues were only considered if selected areas totaled 2.37 mm2. Pixelwise H-scores were calculated using modified scripts based on publicly available scripts implemented by Peter Bankhead. Subjective scores were generated by a panel of three board-certified pathologists (JPC, JE, KL) following recommendations by Meyerholz and Beck32. Pathologists were provided whole-slide digital images, which were visualized using NDP.view2 version 2.9.29 (Hamamatsu Photonics). They were instructed to consider entire images and then score the epithelium and the lamina propria (LP) of each tissue separately. In brief, scores were created based on the approximate percentage of cells with immunoreactivity (0–100%) and the estimated intensity of the immunoreactivity (mild (1), moderate (2), marked (3)). By multiplying the percent of positive staining cells by the percentage of staining that was mild, moderate, and marked, which were represented by the numerical values 1, 2, and 3, respectively, the H-score for the epithelium and lamina propria were determined by all three pathologists independently. Evaluators were blinded to the other pathologists’ scores and the computerized image analysis scores until after data were analyzed.
RT-qPCR for tight junction proteins
We used one-step RT-qPCR to quantify total RNA for 7 different genes related to GIT barrier function and health. Primers used are listed in Table S2. The genes of interest were Claudin 1 (CLDN1), Claudin 2 (CLDN2), Occludin (OCLDN), Tight Junction Protein 1 (TJP1), E-Cadherin (CDH1), and S100-A8 (A8) and S100-A9 (A9), which are subunits of the calprotectin protein. Primer sequences previously reported were used for 5 of the 7 targets33,34. Primer sequences for the 2 targets (CLDN2, and CDH1) that were not available for cattle in the published literature were designed using Primer-BLAST (National Center for Biotechnology and Information). Additionally, synthetic gene fragments (gBlocks, Integrated DNA Technologies, Coralville, IA—Table S3) were designed for each target and included in samples at set dilutions. Using the formula \(copies/\mu L=\left(C\right)\left(M\right)({10}^{-15}\frac{mol}{fmol})(6.022\times {10}^{23})\) where C is the concentration of the gene fragment (ng/µL) and M is the molecular weight of the gene fragment converted to moles and multiplied by Avogadro’s number35, a standard curve was developed for each target gene for the quantification of target expression on a per-copy basis in each sample.
Samples and standard curves were tested in triplicate using the reagent manufacturer’s protocol (iTaq™ Universal SYBR® Green One-Step Kit, Bio-Rad, https://www.bio-rad.com/webroot/web/pdf/lsr/literature/10032048.pdf). Briefly, the 20-µL final reactions consisted of 20 ng of RNA, 1X SYBR Green reaction mix and 1X reverse transcriptase, and 450 nM of forward and reverse primers. Thermal cycling was performed with a real-time qPCR system (QuantStudio™ 3, Applied Biosystems, Thermo Fisher Scientific). Cycling conditions included incubation at 50 °C for 10 min for reverse transcription, then 95 °C for 10 min, then 40 cycles of 95 °C for 15 s and 60 °C for 1 min. The process finished with the melt curve stage of 95 °C for 15 s, 60 °C for 1 min, and 0.15 °C ramp to 95 °C with a hold for 1 s.
Statistical analysis
Statistical analyses were conducted using R version 4.2.236. For multivariate comparisons (i.e., beta diversity), differences were tested using pairwise PERMANOVA with a Benjamini–Hochberg false discovery rate (FDR) correction for multiple comparisons and 9,999 permutations. Additionally, pairwise PERMDISPs were used with 9,999 permutations to test for differences in the variability of dispersions. All qPCR results were tested for normality with the Shapiro–Wilk test. Because the data were not normally distributed, a Kruskal–Wallis test was used to evaluate differences. When needed, a Wilcoxon pairwise rank-sum test with a Benjamini–Hochberg FDR correction was used to compare medians. Immunohistochemical data were evaluated for normality, and linear models were used when appropriate. Computer-generated data were analyzed using a linear model with sample type as the fixed effect and no random effects. Scores generated by the pathologists were also evaluated using a linear model that included sample type and evaluator as fixed effects. All linear models were calculated using the lme4 package of R37. Significant differences were determined at P ≤ 0.05. Bland–Altman analysis was used to investigate the agreement between the pathologist’s and the computer-generated H-scores, and plotted using the ggplot package. All graphics were generated in R, and legends were added using BioRender.com.
Results
16S rRNA sequencing
Samples with fewer than 250,000 reads per sample (n = 7) were excluded from downstream analysis (Table S4). Samples excluded from analysis were from a mixture of sample types (2 RU, 4 SI, and 1 LI). The number of reads identified in the remaining samples (n = 145) averaged 581,594 reads per sample (range = 270,390–886,923 Figure S1). There was no significant difference in sequencing depth among the animals (Kruskal Wallis Test P = 0.15). However, differences in sequencing depth were noted between sample sites (Kruskal Wallis Test P < 0.01), with fecal and liver abscess samples containing more reads than the RU, SI, and LI. More than 99% of all ASVs could be classified at the taxonomic ranks of phylum, class, order, and family, whereas only 63.7% of ASVs were classified at the genus level. There were differences in the richness and evenness (Kuskal Wallis Test P < 0.01; Figure S2). Generally, the liver abscess samples were less rich and less diverse than the gastrointestinal samples.
Overall, microbial community structures did differ significantly among animals (PERMANOVA P < 0.01; R2 = 0.22, Fig. 1), but the PERMDISP statistic for this comparison was significant (PERMDISP P < 0.01), suggesting that unequal variances may have inflated the Type I error rate. Microbial community composition was significantly different among the various GIT locations based on generalized Unifrac distances (PERMANOVA P < 0.01; R2 = 0.44), but the PERMDISP statistic for this comparison was significant (PERMDISP P < 0.01), suggesting that unequal variances may have inflated the Type I error rate. Regardless, ordination and clustering evaluated using dendrograms visually demonstrated that the microbial communities of LA samples were distinctly different from all other sample types (Figure S3). For other samples, microbial communities found on GIT epithelial surfaces generally exhibited greater inter-sample variability than those in luminal content samples, with the most noticeable differences observed in SI epithelial surface samples. The RU epithelial surface and luminal samples clustered closely together, as did both LI sample types, fecal samples, and SI luminal samples; however, there was a clear separation between these two groups (Fig. 1, Figure S3). In contrast, microbial community composition in SI epithelial surface samples overlapped with all other sample types except for LA samples.
Non-metric multidimensional scaling (NMDS) ordination plot of microbial community composition in gastrointestinal samples from feedlot steers based on generalized UniFrac distances of 16S rRNA amplicon sequence variants (ASVs). Both luminal and epithelial surface samples are represented from 4 different gastrointestinal tract (GIT) sites: rumen (RU), small intestine (SI), large intestine (LI), feces (F), and from liver abscesses (LA). Ellipses represent the 95% confidence intervals for the group mean values. (A) Ordination of samples colored by individual animal ID (n = 21), showing that microbial communities do not cluster separately by animal. (B) Ordination of samples colored by GIT region and sample type. Clustering by GIT region was statistically significant (PERMANOVA, P < 0.001), with GIT locations showing site-specific microbial community structures and LA samples forming a distinct cluster. RU LC = rumen luminal fluid, RU ES = rumen epithelial surface, SI LC = small intestine luminal fluid, SI ES = small intestine epithelial surface, LI LC = large intestine luminal fluid, LI EC = large intestine epithelial surface.
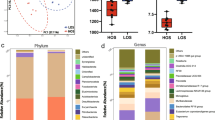
Analyses to identify core microbial communities were conducted for all sample sites and across the entire GIT; however, liver abscess (LA) samples were removed from all core analyses because of clear differences in microbial taxa (Figure S3). Across all GIT samples, 9 families were identified as core community members throughout the entire GIT (Coprostanoligenes group, Anaerovoracaceae, Atopobiaceae, Christensenellaceae, Erysipelotrichaceae, Lachnospiraceae, Methanobacteriaceae, Oscillospiraceae, and Ruminococcaceae; Fig. 2). Of these families, Lachnospiraceae was found in the greatest average relative abundance. However, more families were identified as core community members when samples were analyzed as subsets by sampling location. In the RU samples (both epithelial surface and luminal samples), 17 families were identified as core (Coprostanoligenes group, Acidaminococcaceae, Anaerovoraceae, Atopobiaceae, Uncultured Bacteroidales, Christensenellaceae, Erysipelatoclostridiaceae, Erysipelotrichaceae, F082, Lachnospiraceae, Methanobacteriaceae, Muribaculaceae, Oscillospiraceae, Prevotellaceae, Rikenellaceae, Ruminococcaceae, Selenomonadaceae, and Succinivibrionaceae; Fig. 3). In the SI samples, 10 families were identified as core (Anaerovoraceae, Atopobiaceae, Bifidobacteriaceae, Clostridiaceae, Eggerthellaceae, Erysipelotrichaceae, Lachnospiraceae, Methanobacteriaceae, Peptostreptococcaceae, and Ruminococcaceae). The LI had 15 core family taxa (Coprostanoligenes group, Anaerovoraceae, Atopobiaceae, Bifidobacteriaceae, Christensenellaceae, Clostridiaceae, Eggerthellaceae, Erysipelotrichaceae, Lachnospiraceae, Methanobacteriaceae, Oscillospiraceae, Peptostreptococcaceae, Prevotellaceae, and Ruminococcaceae). Finally, the feces had 20 families considered core (Coprostanoligenes group, Anaerovoraceae, Atopobiaceae, Bifidobacteriaceae, Christensenellaceae, Clostridiaceae, Eggerthellaceae, Erysipelotrichaceae, Lachnospiraceae, Methanobacteriaceae, Oscillospiraceae, Peptostreptococcaceae, Prevotellaceae, Rikenellaceae, and Ruminococcaceae). Within a sampling location, inter-animal variability was notable for all samples. Similarly, variation between luminal and epithelial surface samples was observed. However, when samples were analyzed together within the same sampling location (i.e., RU, SI, or LI), the RA for members of the core community tended to be similar. Finally, the overlap of core family composition in adjacent sample sites highlights a biologically logical flow of resident microbes from the proximal to distal ends of the GIT. However, differences in core community composition among sites underscore how each portion of the GIT has a unique microbial community structure.
The core bacterial families identified across all GIT regions sampled in feedlot cattle. Dots illustrate the average relative abundance (and 95% confidence interval) for the core microbiota identified in luminal and intestinal surface samples from the rumen (RU), small intestine (SI), and large intestine (LI), as well as feces, by sample type (luminal content = LC, epithelial surface = ES). Core taxa were defined as bacterial families present in ≥ 90% of samples with ≥ 0.1% mean relative abundance. Bubble size indicates the mean relative abundance for each sample type, and vertical lines represent the 95% confidence intervals. Liver abscess samples were excluded from this analysis. The consistent presence of Lachnospiraceae, Ruminococcaceae, Christensenellaceae, and other taxa associated with SCFA metabolism suggests microbial functional conservation across regions of the GIT in high-performing feedlot steers.
The core bacterial families identified within each of the GIT regions sampled in feedlot cattle. Each dot represents the relative abundance for a single sample for the core microbiota identified in luminal and intestinal surface samples by sampling location (rumen = RU, small intestine = SI, large intestine = LI, and feces), and by sample type (luminal content = LC, epithelial surface = ES). Core taxa were defined as bacterial families present in ≥ 90% of samples with ≥ 0.1% mean relative abundance. A total of 17 core families were identified in the rumen, 10 in the small intestine, 15 in the large intestine, and 20 in feces. Similar core community features were identified in luminal and epithelial surface samples from within tissues, and several families were shared across multiple locations, highlighting a gradient of microbial continuity throughout the GIT. Within-location comparisons revealed similar core community composition between luminal and mucosal samples, whereas across-location comparisons demonstrated both distinct core taxa by site, and prominent sharing of key features among locations that appear consistent with proximal-to-distal gradients in GIT function and structure.
These differences are further highlighted by the differentially abundant taxa identified with ANCOM-BC (Figure S4). Clear spatial differentiation of microbial communities was observed across the cattle gastrointestinal tract. The rumen was dominated by fermentative and lactic acid–associated families, including Veillonellaceae and Succinivibrionaceae, consistent with its predominant role in carbohydrate fermentation. The small intestine showed enrichment of Lactobacillaceae and Streptococcaceae, taxa commonly associated with mucosal surfaces and nutrient assimilation. In contrast, the colon was characterized by enrichment of Ruminococcaceae and Lachnospiraceae, families known for secondary fermentation and short-chain fatty acid production. Fecal samples displayed higher abundance of Bacteroidaceae and Prevotellaceae, suggesting that while feces capture aspects of large-intestinal fermentation, they underrepresent taxa from more proximal compartments. These findings highlight the functional compartmentalization of the bovine GIT microbiome and caution against over-reliance on fecal samples as proxies for upstream microbial communities.
Immunohistochemistry for tight junction proteins
Immunohistochemical staining intensity and distribution varied greatly depending on the protein of interest. Generally, CLDN1 exhibited moderate epithelial staining in the rumen, small intestine, and large intestine, with less staining in the lamina propria. Epithelial cells in all three tissues showed positive staining diffusely in the cytoplasm of the enterocytes, with stronger cytoplasmic staining observed in goblet cells of the SI and LI. The cytoplasm of cells scattered throughout the lamina propria demonstrated strong immunoreactivity in all three tissues (Fig. 4). For CLDN2, diffuse, mild cytoplasmic staining was noted in the epithelial component of the small and large intestine; however, it was absent in the ruminal epithelium. The lamina propria was largely devoid of positivity, except for strong, diffuse, cytoplasmic staining found in scattered cells across all three tissues. Typically, OCLDN exhibited moderate, diffuse membranous, and mild cytoplasmic staining in the epithelium of all three tissue types, with slightly increased prevalence in the small and large intestines. The lamina propria in these tissues was largely devoid of immunoreactivity. For MAC-387, weak to strong cytoplasmic immunoreactivity was observed in the ruminal epithelium but was largely absent in the epithelium of the small or large intestine and was rarely present within the lamina propria of all three tissues. Finally, CDH1 exhibited strong, cytoplasmic, and membranous immunoreactivity in the epithelium of all three tissues and weak, cytoplasmic, and membranous reactivity in the lamina propria of all three tissues.
Representative images of immunohistochemical staining for protein expression in GIT tissues of feedlot steers. (A) Claudin 1 (CLDN1) in the colon: moderate, diffuse cytoplasmic and membranous staining is evident in the epithelium, with weaker staining in the lamina propria. (B) Claudin 2 (CLDN2) in the rumen: no detectable immunopositivity in the epithelium or lamina propria; observed staining is attributed to nonspecific uptake by debris. (C) Occludin (OCLDN) in the jejunum: moderate, diffuse cytoplasmic immunoreactivity in the epithelium with minimal staining in the lamina propria. (D) Calprotectin (MAC-387) in the rumen: weak to strong nuclear and cytoplasmic immunoreactivity observed in the epithelium. (E) E-cadherin (CDH1) in the jejunum: strong cytoplasmic and membranous immunoreactivity within the epithelium, and weak staining in the lamina propria. Scale bars: A–D = 100 µm; E = 250 µm.
There were no differences in average pixelwise H-scores generated by computerized image analysis for CLDN2 among the different tissue sites (F-Test P = 0.97; Fig. 5). However, the average H-scores for the RU were significantly greater (F-Test P < 0.01) for CLDN1, OCLDN, and MAC-387 than was found in the SI or LI. In contrast, image analysis of CDH1 staining found higher average scores in the SI and LI than in the RU (F-Test P < 0.01; RU sample mean = 174, 95% CI: 168–185; SI sample mean = 197, 95% CI: 187–206; LI sample mean = 199, 95% CI: 185–212; Table S5). However, given that the computerized image analysis generated a single pixelwise H-score for the entire tissue section, including the epithelium and the LP and all targets except MAC-387 were epithelium-associated targets, image analysis scores may have been skewed by an absence of staining in the LP.
Pixelwise H-scores for expression of TJPs and an inflammatory marker in GIT tissues of feedlot steers based on computerized image analysis. Dot and Boxplots show H-scores derived from image analysis of immunohistochemical staining of tissue sections for Claudin 1, Claudin 2, Occludin, E-Cadherin, and MAC-387 (Calprotectin) in the rumen (RU ES), jejunum (SI ES), and colon (LI ES). H-scores were generated using pixel-level quantification algorithms from QuPath software. Within each plot, boxes with different superscripts (a, b, c) indicate statistically significant differences among tissue types (P < 0.01).
In contrast, the average epithelium-specific OCLDN H-scores that were generated independently by a panel of three pathologists were not significantly different among the different tissue sites (F-Test P = 0.18; Fig. 6). However, the pathologist-generated H-scores were significantly greater in the SI (F-Test P ≤ 0.02) than the RU for CLDN1, CLDN2, and CDH1. The H-scores for these proteins were intermediate in LI and not significantly different from the RU or SI. MAC-387 H-scores for epithelial tissues were greatest on average in the RU exhibited (F-Test P < 0.01) and staining was rare in the SI and LI (RU average pathologists’ H-score = 157, 95% CI: 134–180; SI and LI average H-scores = 0, 95% CI = 0–0). However, the pathologists’ H-scores for MAC-387 staining in the LP were significantly greater in the SI compared to the RU (F-Test P < 0.01; Fig. 7). Similarly, the pathologists’ LP-specific scores were significantly greater for CLDN1, CDH1, and OCLDN in the SI and LI than the RU (F-Test P < 0.01), and CLDN2 staining was stronger in the SI than both the RU and LI (F-Test P < 0.01, RU sample mean = 4, 95% CI: 2–7; SI sample mean = 18, 95% CI: 10–26; LI sample mean = 10, 95% CI: 4–17). There were no significant differences among the pathologists’ H-scores when comparing cattle with liver abscesses to those with edible livers (F-Test P ≥ 0.06), but this could have been affected by the low liver abscess prevalence (2 of 21 subjects).
Epithelial H-scores for expression of TJPs and an inflammatory marker in GIT tissues of feedlot steers based on blinded, independent evaluations by three pathologists. Dot and Boxplots show H-scores of immunohistochemical staining of epithelium in tissue sections for Claudin 1, Claudin 2, Occludin, E-Cadherin, and MAC-387 (Calprotectin) in the rumen (RU ES), jejunum (SI ES), and colon (LI ES). Within each plot, boxes with different superscripts (a, b, c) indicate statistically significant differences among tissue types (P < 0.02).
Lamina propria H-scores for expression of TJPs and an inflammatory marker in GIT tissues of feedlot steers based on blinded, independent evaluations by three pathologists. Dot and Boxplots show H-scores of immunohistochemical staining of lamina propria in tissue sections for Claudin 1, Claudin 2, Occludin, E-Cadherin, and MAC-387 (Calprotectin) in the rumen (RU ES), jejunum (SI ES), and colon (LI ES). Within each plot, boxes with different superscripts (a, b, c) indicate statistically significant differences among tissue types (P < 0.02).
The pathologists’ H-scores for CLDN1 staining in the epithelial tissues were generally higher than the computer-generated pixel-wise H-scores and lower for the LP, as demonstrated through Bland–Altman analysis (Fig. 8). This result is perhaps not surprising, given that the computer-generated image analysis provided scores for staining in both the epithelial and LP tissues. However, when the pathologists’ H-scores for CLDN1 staining were averaged for the epithelium and LP of the tissue section, the mean bias was very small relative to the computer-generated H-scores (mean bias = 0.511). However, the pathologists’ epithelium scores for CLDN2 had a relatively small mean bias when compared to the computerized analysis (mean bias = 7.85), and much lower than computer-generated scores for the LP with an unequal variance across the range of values. The averaged pathologists’ scores for CLDN2 demonstrated modest agreement with the computer-generated score (mean bias = -33.78). This same trend was noted for OCLDN and CDH1 wherein the pathologists’ epithelial scores were greater, and the LP scores were lesser when compared to the computerized analysis scores, and the pathologists’ averaged scores were similar to the computerized analysis score (OCLDN mean bias = 2.27, CDH1 mean bias = -46.63). Finally, there was a broader range of differences between the pathologists’ and the computer-generated H-scores, with a mean bias suggesting modest agreement with the pathologists’ averaged scores (mean bias = -38.32).
Bland–Altman plots evaluating agreement between scoring methods of immunohistochemical staining of TJPs and an inflammatory marker in GIT tissues of feedlot steers (Claudin 1 = CLDN1, Claudin 2 = CLDN2, Occludin = OCLDN, E-cadherin = E-CAD, MAC-387 = Calprotectin). Plots compare the mean H-scores for samples generated by a panel of 3 pathologists for epithelial tissues (Epi H), lamina propria (LP), or a combined average of scores of the epithelium and lamina propria (AVG) vs the computer-generated pixelwise H-score (automated QuPath analysis) for the epithelium and lamina propria combined. Solid blue lines indicate mean bias, and red dashed lines represent the 95% confidence interval limits.
The potential relationships between tight junction protein expression and microbial community composition was explored by categorizing computer-generated H-Scores as quartiles for each of the proteins evaluated by immunohistochemistry (CLDN1, CLDN2, OCLDN, CDH1, and MAC-387). Analyses of beta diversity stratified by protein and tissue site did not identify any significant differences in microbial communities for any proteins with different H-Score intensity categories at any tissue site (PERMANOVA P > 0.16).
RT-qPCR for tight junction proteins
There were no significant differences among most sample types regarding the quality of RNA recovered from flash-frozen tissue samples (Kruskal Wallis Test P = 0.27; mean RIN for RU samples = 4.27, 95% confidence interval (95% CI): 3.01–5.54; SI sample mean = 3.10, 95% CI: 2.74–3.45; LI sample mean = 2.66, 95% CI: 2.31–3.01). The low average RIN values for samples may suggest that RNA had been moderately degraded at the time of sampling, which could have affected qPCR results. This is further supported by the observation that RIN values were unobtainable from the electrophoresis analysis for many rumen samples (n = 11).
There were no differences detected in OCLDN expression across sample types (Kruskal Wallis Test P = 0.08; Figure S5). The SI had lower (Kruskal Wallis Test P = 0.03) TJP1 expression compared to the LI, while the RU was intermediate. However, the SI exhibited greater (Kruskal Wallis Test P ≤ 0.02) expression of CDH1 and CLDN1 than the RU, with the LI being intermediate. Both the SI and the LI displayed increased (Kruskal Wallis Test P < 0.01) expression of CLDN2 compared to the RU. In contrast, for TJP1, the LI demonstrated increased expression compared to the SI, with the RU being intermediate. Despite attempts using two different published primer sets34,38, A8 and A9 consistently failed to amplify in all sample types (data not shown). No differences were associated with the sample batch (plate) or regarding the occurrence of LA in the animal (Kruskal Wallis Test P ≥ 0.32). However, the number of LAs limited our ability to investigate this variable thoroughly.
Discussion
This study provides foundational insight into the microbial and host components of the GIT that could affect the health and productivity of finish-fed feedlot cattle, a population underrepresented in existing research despite its central importance to beef production in North America. We observed pronounced variations in both microbial communities and TJP expression across GIT sites (e.g., RU, SI, LI, feces), consistent with prior reports14,39. Notably, site-specific differences were much greater than variability attributable to individual animals. Inclusion of cattle produced at 21 different commercial feedlots provided a unique strength to this investigation, as these animals are broadly representative of economically valuable cattle targeted in commercial beef production in large feedlots in the U.S. The limited inter-animal variability in findings compared to differences in the GIT site was striking, given their complete separation in environments, management practices, and sources for dietary components. While dietary composition has been associated with differences in the GIT microbiome40,41,42,43, specifics of diet formulations were not available for this population though rations were mixed onsite at the feedlots (i.e., diets were not produced at a central milling location and distributed to multiple feedlots) and it is likely that there were similarities in major energy and protein sources, e.g., steam-flaked corn, distiller’s grains44, which may have contributed to the similarities of the microbial communities at the time of harvest. Still, inter-animal differences in the microbiomes were more pronounced in epithelial surface than luminal samples, particularly in the small intestine, suggesting host-specific factors may more strongly influence epithelial-associated microbial populations.
While differences in TJP expression by GIT site have been reported in calves8,45, this study provides novel data for mature feedlot cattle. Notably, both the small and large intestines exhibited significantly higher expression of several TJPs evaluated in this study using IHC (CLDN1, CLDN2, CDH1, and TJP1) compared to the rumen, supported by RT-qPCR. These proteins are essential for regulating epithelial permeability and maintaining tissue integrity, and their increased expression in the intestines likely reflects a greater need for selective absorption and immunological defense in these regions46,47,48. Interestingly, OCLDN was expressed uniformly across all GIT sites, suggesting a conserved functional role maintained regardless of location. The protein expression analyzed using computerized and pathologist-generated H-scores was generally similar at the tissue level, suggesting that either method may serve as a proxy in future studies. However, we found image analysis limited in its ability to readily analyze different tissue layers (e.g., epithelium vs. LP), which may pose an important limitation depending on the study objectives and protein target. While RT-qPCR results supported its potential use as a proxy for IHC values, caution must be exercised in interpreting the results of this study’s RT-qPCR due to lower RNA quality scores (RIN). Despite best efforts to quickly preserve RNA when tissues were sampled, RNA quality was likely impacted by the inherent time lapse present in standard commercial slaughter procedures from the time that cattle are rendered senseless until the removal of GIT organs.
These findings also highlight the potential for using TJP profiles as biomarkers of GIT function and health. Establishing baseline expression patterns in healthy cattle offers critical context for interpreting changes associated with disease, nutritional interventions, or management strategies. Moreover, the integration of host gene expression with microbiome data presents new opportunities to understand how microbial populations influence, or are influenced by, epithelial integrity—an interaction with implications for nutrient utilization, disease susceptibility, and feed efficiency33,49.
The microbiome analyses revealed distinct, region-specific community structures consistent with previous findings in cattle and other species14,39. Core bacterial families were identified for each GIT region, representing abundant, prevalent taxa that are well-suited to thrive within the GIT of feedlot cattle and likely fulfill important functions in a specific gastrointestinal ecosystem50, with Lachnospiraceae emerging as the most consistently abundant group. When core taxa were analyzed collectively across all sites, nine families met stringent prevalence and abundance criteria, suggesting they may play essential roles in maintaining GIT homeostasis. The consistent presence of Lachnospiraceae, Ruminococcaceae, Christensenellaceae, and other taxa associated with SCFA metabolism support this hypothesis regarding microbial functional conservation across regions of the GIT in high-performing feedlot steers, which may have been influenced by high-energy feedlot diets that were likely fed to these cattle41,51. SCFAs are known to contribute to both local and systemic host physiology and tissue health. Additional taxa were as being core within specific sample types, reflecting how each anatomical site supports a unique microbial niche50,52.
The insight into the spatial expression of TJPs provided by IHC added further granularity to our understanding of TJPs in tissues, which was generally supported by RT-qPCR findings. Overall, TJPs were more abundant in the small and large intestines compared to the rumen, with the exception of OCLDN, which showed consistent expression across tissues. The absence of variation in OCLDN expression may indicate that while other TJPs adapt to local conditions, OCLDN fulfills a foundational role in epithelial integrity across the GIT. Differences in staining between epithelial and lamina propria (LP) regions were common and highlight the importance of evaluating distinct tissue layers. While computerized image analysis allows for high-throughput scoring, pathologist-assigned H-scores revealed regional disparities not fully captured by automated approaches. Bland–Altman plots demonstrated moderate agreement between methods and suggested that hybrid approaches—leveraging both expert evaluation and computational scoring—may offer the most robust assessment of tissue-specific protein expression32. Nonetheless, scoring algorithms likely need to separately quantify epithelial and LP regions, especially as these regions may exhibit different functional roles in barrier integrity and immune function.
Data availability
All sequencing data are available through BioProject PRJNA1207998 at the NCBI’s Sequence Read Archive. The code and instructions for the bioinformatic and statistical analyses can be found at DOI: (https://doi.org/10.5281/zenodo.16954014). Images of immunohistochemically stained tissue sections that were analyzed in this study and RT-qPCR data can be found at DOI: (https://doi.org/10.18738/T8/SX1YFM).
Change history
08 January 2026
A Correction to this paper has been published: https://doi.org/10.1038/s41598-025-32675-7
References
The Ruminant Animal: Digestive Physiology and Nutrition (Waveland Press Inc., 1988).
Tulic, M. K., Piche, T. & Verhasselt, V. Lung–gut cross-talk: evidence, mechanisms and implications for the mucosal inflammatory diseases. Clin. Exp. Allergy 46, 519–528 (2016).
Marsland, B. J., Trompette, A. & Gollwitzer, E. S. The gut-lung axis in respiratory disease. Ann. Am. Thorac. Soc. 12, S150–S156 (2015).
Steele, M. A., Penner, G. B., Chaucheyras-Durand, F. & Guan, L. L. Development and physiology of the rumen and the lower gut: Targets for improving gut health. J. Dairy Sci. 99, 4955–4966 (2016).
Bischoff, S. C. ‘Gut health’: A new objective in medicine?. BMC Med. 9, 1–14 (2011).
Di Tommaso, N., Gasbarrini, A. & Ponziani, F. R. intestinal barrier in human health and disease. Int. J. Environ. Res. Public Health 18, 12836 (2021).
Chase, C. & Kaushik, R. S. Mucosal immune system of cattle: All immune responses begin here. Vet. Clin. N. Am. Food Anim. Pract. 35, 431–451 (2019).
Malmuthuge, N., Li, M., Goonewardene, L. A., Oba, M. & Guan, L. L. Effect of calf starter feeding on gut microbial diversity and expression of genes involved in host immune responses and tight junctions in dairy calves during weaning transition. J. Dairy Sci. 96, 3189–3200 (2013).
Zihni, C., Mills, C., Matter, K. & Balda, M. S. Tight junctions: from simple barriers to multifunctional molecular gates. Nat. Rev. Mol. Cell Biol. 17, 564–580 (2016).
Aschenbach, J. R. et al. Symposium review: The importance of the ruminal epithelial barrier for a healthy and productive cow. J. Dairy Sci. 102, 1866–1882 (2019).
Holman, D. B. & Gzyl, K. E. A meta-analysis of the bovine gastrointestinal tract microbiota. FEMS Microbiol. Ecol. 95, 72 (2019).
Lin, L., Lai, Z., Zhang, J., Zhu, W. & Mao, S. The gastrointestinal microbiome in dairy cattle is constrained by the deterministic driver of the region and the modified effect of diet. Microbiome 11, 10 (2023).
Mao, S., Zhang, M., Liu, J. & Zhu, W. Characterising the bacterial microbiota across the gastrointestinal tracts of dairy cattle: membership and potential function. Sci. Rep. 5, 1–14 (2015).
Plaizier, J. C. et al. The duration of increased grain feeding affects the microbiota throughout the digestive tract of yearling Holstein steers. Microorganisms 8, 1854 (2020).
USDA Livestock Poultry Grain and Market news. National daily beef and cattle summary (2024). https://www.ams.usda.gov/mnreports/lsddcbs.pdf
Economic Impact: Cattle Feeding Supports Stronger Communities. Texas Cattle Feeders Association Website (2024). https://www.tcfa.org/economic-impact.html
Kinross, J. M., Darzi, A. W. & Nicholson, J. K. Gut microbiome-host interactions in health and disease. Genome Med. 3, 14 (2011).
Welch, C. B., Ryman, V. E., Pringle, T. D. & Lourenco, J. M. Utilizing the gastrointestinal microbiota to modulate cattle health through the microbiome-gut-organ axes. Microorganisms 10, 1391 (2022).
Klindworth, A. et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 41, e1–e1 (2013).
Illumina Inc. 16S Metagenomic sequencing library preparation. 15044223, Preprint at (2013).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Quast, C. et al. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2012).
McMurdie, P. J. & Holmes, S. phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217 (2013).
Chen, J. et al. Associating microbiome composition with environmental covariates using generalized UniFrac distances. Bioinformatics 28, 2106–2113 (2012).
Paulson, J. N., Stine, O. C., Bravo, H. C. & Pop, M. Differential abundance analysis for microbial marker-gene surveys. Nat. Methods 10, 1200–1202 (2013).
Oksanen, J. et al. vegan: Community ecology package (2022).
Martinez Arizu, P. pairwiseAdonis: Pairwise multilevel comparison using adonis (2020).
Murtagh, F. & Legendre, P. Ward’s hierarchical agglomerative clustering method: Which algorithms implement ward’s criterion?. J. Classif. 31, 274–295 (2014).
Lin, H. & Peddada, S. D. Analysis of compositions of microbiomes with bias correction. Nat. Commun. 11, 3514 (2020).
Bankhead, P. et al. QuPath: Open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Meyerholz, D. K. & Beck, A. P. Principles and approaches for reproducible scoring of tissue stains in research. Lab. Invest. 98, 844–855 (2018).
Pederzolli, R.-L.A. et al. Effect of ruminal acidosis and short-term low feed intake on indicators of gastroin-testinal barrier function in Holstein steers. J. Anim. Sci 96, 108–125 (2018).
Koy, M. et al. Recombinant bovine S100A8 and A9 enhance IL-1β secretion of interferon-gamma primed monocytes. Vet. Immunol. Immunopathol. 155, 162–170 (2013).
Tyra, H. Tips for working with gBlocks gene fragments (2017).
Team, R. C. R: A Language and Environment for Statistical Computing. Preprint at www.r-project.org (2020).
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67 (2015).
Zhao, F. et al. S100A9 a new marker for monocytic human myeloid-derived suppressor cells. Immunology 136, 176–183 (2012).
Malmuthuge, N., Griebel, P. J. & Guan, L. L. Taxonomic identification of commensal bacteria associated with the mucosa and digesta throughout the gastrointestinal tracts of preweaned calves. Appl. Environ. Microbiol. 80, 2021–2028 (2014).
Deusch, S., Tilocca, B., Camarinha-Silva, A. & Seifert, J. News in livestock research - use of Omics-technologies to study the microbiota in the gastrointestinal tract of farm animals. Comput. Struct. Biotechnol. J. 13, 55–63 (2014).
Mccann, J. C., Wickersham, T. A. & Loor, J. J. High-throughput methods redefine the rumen microbiome and its relationship with nutrition and metabolism. Bioinform. Biol. Insights 8, BBI.S15389 (2014).
Zened, A. et al. Microbial ecology of the rumen evaluated by 454 GS FLX pyrosequencing is affected by starch and oil supplementation of diets. FEMS Microbiol. Ecol. 83, 504–514 (2013).
Clemmons, B. A., Voy, B. H. & Myer, P. R. Altering the gut microbiome of cattle: considerations of host-microbiome interactions for persistent microbiome manipulation. Microb. Ecol. 77, 523–536 (2019).
Samuelson, K. L., Hubbert, M. E., Galyean, M. L. & Löest, C. A. Nutritional recommendations of feedlot consulting nutritionists: The 2015 New Mexico State and Texas Tech University survey1. J Anim. Sci. 94, 2648–2663 (2016).
Walker, M. P., Evock-Clover, C. M., Elsasser, T. H. & Connor, E. E. Short communication: Glucagon-like peptide-2 and coccidiosis alter tight junction gene expression in the gastrointestinal tract of dairy calves. J. Dairy Sci. 98, 3432–3437 (2015).
Kinashi, Y. & Hase, K. Partners in leaky gut syndrome: Intestinal dysbiosis and autoimmunity. Front. Immunol. 12, 1390 (2021).
Steed, E., Balda, M. S. & Matter, K. Dynamics and functions of tight junctions. Trends Cell Biol. 20, 142–149 (2010).
Horowitz, A., Chanez-Paredes, S. D., Haest, X. & Turner, J. R. Paracellular permeability and tight junction regulation in gut health and disease. Nat. Rev. Gastroenterol. Hepatol. 20, 417–432 (2023).
Zou, B. et al. Protective effects of niacin on rumen epithelial cell barrier integrity in heat-stressed beef cattle. Animals 14, 313 (2024).
Perlman, D. et al. Concepts and consequences of a core gut microbiota for animal growth and development. Annu. Rev. Anim. Biosci. 10, 177–201 (2022).
Meehan, C. J. & Beiko, R. G. A phylogenomic view of ecological specialization in the lachnospiraceae, a family of digestive tract-associated bacteria. Genome Biol. Evol. 6, 703–713 (2014).
Neu, A. T., Allen, E. E. & Roy, K. Defining and quantifying the core microbiome: Challenges and prospects. Proc. Natl. Acad. Sci. 118 (2021).
Acknowledgements
The authors thank the employees of the West Texas A&M University Research Feedlot, Caviness Meat Science & Innovation Center, and Dana Burk, Rachel Huff, and Nerissa Bechtol from the VERO Research Laboratory for their contributions to this project. This research was funded by Texas A&M University, and portions of this research were conducted using advanced computing resources provided by Texas A&M High Performance Research Computing Center.
Funding
This work was funded by Texas A&M University.
Author information
Authors and Affiliations
Contributions
PSM was the principal investigator overseeing all aspects of the research. PSM, LJP, and JDY conceived and planned the study with assistance from TEL and JTR. TEL oversaw the animal harvest processes. JDY and TEL conducted sample collection. CAW oversaw sample handling in the laboratory, sequencing and RT-qPCR procedures with assistance from PSM, LJP, and MES. JDY conducted laboratory and data analyses. JPC conducted all immunohistochemical analysis and developed protocols for the computerized and pathologists’ histological analysis of tissues. JDY conducted the computerized image analysis of. JPC, KLL, and JAE scored immunohistochemical staining in GIT tissues. JDY conducted bioinformatic and statistical analysis with assistance from LJP. JDY prepared initial manuscript drafts with assistance from PSM and LJP. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: The original version of this Article contained an error where Figure S4 was incorrectly duplicated as Figure 8.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Young, J.D., Pinnell, L.J., Wolfe, C.A. et al. Microbial communities and tight junction protein expression in the gastrointestinal tract of feedlot cattle. Sci Rep 15, 39055 (2025). https://doi.org/10.1038/s41598-025-25982-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-25982-6