Abstract
To examine associations between Kirsten rat sarcoma viral oncogene (KRAS) mutations and histopathological features in colorectal cancer and to evaluate their prognostic implications. A total of 256 patients with colorectal adenocarcinoma treated between January 2018 and December 2022 were retrospectively included. Rat sarcoma (RAS) gene mutations, covering 21 sites in KRAS and NRAS, were detected using the ADx-ARMS kit. Due to the low mutation rate of NRAS (3.9%) and its lack of significant correlation with pathological features (P > 0.05), further analysis focused on KRAS mutations. Tumor differentiation, mucinous components, and tumor budding were assessed by two pathologists in a double-blind manner. The relationship between KRAS mutation status and morphological features was analyzed using logistic regression, and the prognostic value was evaluated using the Cox proportional hazards model. The KRAS mutation rate was 52.7% (135/256), with the main subtypes being G12D (30.37%), G13D (20.00%), G12V (17.78%), and G12C (7.41%). Logistic regression showed that KRAS mutation independently predicted right-sided colon tumors (OR = 1.85, P = 0.017), high tumor budding (OR = 2.46, P = 0.002), and mucinous components (OR = 2.39, P = 0.012). It was also positively associated with PIK3CA co-mutation (OR = 2.15, P = 0.042). Overall survival (OS) was longer in patients with KRAS mutations than in those with wild-type KRAS (P = 0.013), was longer in KRAS-mutated than wild-type cases (P = 0.013), whereas progression-free survival (PFS) showed no significant difference. No survival differences were detected among KRAS subtypes. Multivariate Cox analysis showed that TNM stage IV (HR = 4.80), poor differentiation (HR = 2.00), and absence of primary tumor resection (HR = 3.20) were independent adverse prognostic factors. KRAS mutations are closely associated with aggressive pathological features in colorectal cancer, particularly tumor budding and mucinous differentiation. Histopathological evaluation may aid risk stratification alongside KRAS status. Prognostic assessment in clinical settings should take both TNM staging and KRAS status into account.
Similar content being viewed by others
Introduction
Colorectal cancer is one of the leading causes of cancer-related deaths worldwide. Its development involves the accumulation of multiple gene abnormalities and interactions with the tumor microenvironment1,2. With the advancement of precision medicine, molecular classification has become a key strategy in guiding personalized treatment for colorectal cancer. Among them, the mutation status of the RAS gene family (KRAS, NRAS, HRAS) directly influences the effectiveness of anti-EGFR targeted therapy and patient prognosis3,4. KRAS is the most critical driver gene in the RAS family, accounting for approximately 85–90% of RAS mutations in colorectal cancer5,6. Mutations in exons 2, 3, and 4 can lead to continuous activation of the MAPK signaling pathway, promoting tumor growth and spread7,8. In contrast, NRAS (8–12%) and HRAS (< 2%) mutations are less frequent and of limited clinical significance. Current guidelines recommend KRAS mutation status as a key predictor of resistance to anti-EGFR therapy8,9. In recent years, several studies have suggested that KRAS mutations may be related to tumor location and aggressive features10,11. For example, KRAS mutations are more common in right-sided colon tumors12, and specific subtypes like G12D and G13D may influence tumor differentiation and aggressiveness by affecting mucin production or epithelial-mesenchymal transition12. However, most studies have focused on molecular mechanisms, and the systematic link between KRAS mutations and tumor morphological features (such as tumor budding, mucinous components, and stromal response) remains unclear. As a cornerstone of clinical diagnosis, pathological morphology is vital for understanding tumor biology when integrated with molecular features. However, there is still a lack of direct frameworks connecting KRAS mutations to microscopic structural phenotypes. Therefore, this study aims to clarify the relationship between KRAS mutations and aggressive pathological features by integrating molecular testing and multi-parameter morphological evaluation in 256 colorectal adenocarcinoma cases, providing evidence to support a combined molecular-morphological diagnostic model.
Materials and methods
General information
This retrospective study collected electronic pathology records and pathological specimens from patients diagnosed and treated with colorectal adenocarcinoma at a tertiary hospital between January 2018 and December 2022. The research protocol was approved by the hospital’s medical ethics committee and complied with the Declaration of Helsinki and related ethical guidelines. As patient information had been anonymized, informed consent was waived by the ethics committee. Inclusion criteria: (1) postoperative pathological confirmation of colorectal adenocarcinoma; (2) complete clinicopathological data with ≥ 30% tumor cell content on hematoxylin and eosin (H&E); and (3) available tumor tissue. Exclusion criteria: (1) history of other malignant tumors; (2) received neoadjuvant chemoradiotherapy; (3) poor-quality specimens (tumor cell content < 30%). Sample size estimation: assuming a RAS mutation rate of ~ 50% (α = 0.05, β = 0.20, effect size = 0.30), the minimum sample size was 200; 256 cases were included to improve precision. (Fig. 1).
KRAS Study Workflow.
RAS gene testing methods
Formalin-fixed, paraffin-embedded (FFPE) tumor tissues were reviewed by pathologists. Areas with > 50% tumor cells were selected, and 5-µm sections were cut under sterile blades13. Genomic DNA was extracted using the QIAamp DNA FFPE Tissue Kit (Qiagen, Germany), and DNA concentration was measured with a Qubit 3.0 fluorometer, with a minimum of ≥ 20 ng/µL required. RAS mutation detection was conducted using the CFDA-approved ADx-ARMS kit (AmoyDx, China), targeting 21 clinically relevant mutations in KRAS exons 2 (codons 12/13), exon 3 (codons 59/61), exon 4 (codons 117/146), and NRAS exons 2 (codons 12/13) and exon 3 (codons 59/61). All procedures followed the manufacturer’s protocol. Each run included a positive control (SW480 cell line DNA with KRAS G12V mutation) and a no-template control to monitor for contamination. PCR data were obtained using an ABI 7500 real-time PCR system. A result was considered positive when Ct < 28 with a typical sigmoidal amplification curve. Since NRAS mutations were infrequent (3.9%) and showed no statistically significant association with pathological features or survival, the subsequent analysis focused on KRAS mutations and their subtypes (G12D, G13D, G12V, G12C) in relation to clinicopathological features. Microsatellite Instability (MSI) / Mismatch Repair (MMR) status was assessed using immunohistochemistry to detect MMR-related proteins (MLH1, MSH2, MSH6, PMS2). Antibodies were purchased from ZSGB-Bio, and staining was performed using the EnVision secondary antibody system. Loss of any protein expression was classified as dMMR (MSI-H); intact expression indicated pMMR (MSS). Two experienced pathologists independently evaluated each case. Discrepancies were resolved through discussion. MSI status was assessed in all 256 patients. The evaluation was based on immunohistochemistry for MMR proteins, complemented by molecular testing when required.
Pathological morphology assessment
All pathological slides were independently evaluated in a double-blind manner by two pathologists with over 10 years of experience in gastrointestinal tumors. A pre-test of 30 cases was used to unify diagnostic standards (Kappa coefficient = 0.85). Evaluation parameters included four main aspects: (1) Tumor differentiation: according to the WHO Classification of Digestive System Tumors (5th Edition), tumors were classified as well, moderate, or poor differentiation on H&E staining. (2) mucinous component: Tumors in which mucin pools account for ≥ 50% of the total tumor area are defined as mucinous adenocarcinoma, according to the WHO Classification of Tumours of the Digestive System, 5th edition. (3) Invasive features: ①Vascular invasion: elastica van Gieson (EVG) staining showed tumor cells penetrating vein walls (yellow arrows indicating tumor thrombi); ② Perineural invasion: epithelial-specific staining (ET) confirmed nerve sheath disruption (blue-green areas); ③ Tumor budding: ckpan immunohistochemistry identified budding cell clusters (red circle area). Following the International Tumor Budding Consensus Conference (ITBCC) criteria, a hotspot with the highest density of buds at the invasive front was selected. Tumor buds were counted within a 0.785 mm2 field. Grading was based on the counts: low grade (0–4), intermediate grade (5–9), and high grade (≥ 10). (4) Stromal reaction: lymphocyte density at the invasive front was evaluated using the Klintrup-Mäkinen scoring system. The criteria were: 0, no increase in inflammatory cells; 1, mild increase limited to the invasive margin; 2, moderate infiltration with focal tumor cell destruction; 3, marked infiltration with extensive destruction. For analysis, scores 0–1 were grouped as “weak” and 2–3 as “strong” stromal reaction. (5) Poorly Differentiated Clusters (PDC): defined as clusters of ≥ 5 cancer cells at the invasive front lacking glandular formation. The evaluation was performed on HE slides, a hotspot at the invasive front was selected, and clusters were counted under a ×20 objective (0.785 mm² field). Classification was as follows: G1 (< 5), G2 (5–9), and G3 (≥ 10). For statistical purposes, G2–3 were combined as “PDC positive,” while G1 was treated as the negative group for binary analysis. For 16 cases (6.3%) with diagnostic discrepancies, consensus was reached through multidisciplinary review with additional immunohistochemistry when needed. (Fig. 2).
Histopathological features and invasion patterns of colorectal cancer. (a) Gland formation rate > 95%, good cell polarity, mild nuclear atypia, consistent with WHO Grade 1 differentiation; (b) approximately 70% gland formation, local stromal infiltration, increased mitotic figures (5–10/HPF), consistent with WHO Grade 2 differentiation; (c) gland formation < 50%, tumor cells arranged in solid sheets, increased nuclear-to-cytoplasmic ratio, pathological mitosis observed, consistent with WHO Grade 3 differentiation; (d) tumor cells penetrating vein wall with broken elastic fibers; (e) tumor cells encasing nerve bundles with disrupted perineurium; (f) EnVision staining for cytokeratin, discrete tumor cell clusters (≥ 5 cells/cluster) observed at the invasive front within red circle, meeting BD2 budding grade.
Statistical methods
Data were analyzed using R version 4.2.1 (R Foundation for Statistical Computing) and SPSS 26.0 (IBM Corp.). Continuous variables were tested for normality using the Shapiro-Wilk test. Between-group comparisons were performed using the independent t-test (for normally distributed data) or Mann-Whitney U test (for non-normally distributed data). Categorical variables were compared using the chi-square test or Fisher’s exact test (when expected frequencies were < 5). To control for confounding factors, variables with P < 0.1 in univariate analysis were included in a multivariate logistic regression model to calculate adjusted odds ratios (ORs) and 95% confidence intervals (CIs). Survival analysis was conducted using Kaplan-Meier curves with log-rank tests for group comparison. Prognostic factors were analyzed using univariate and multivariate Cox proportional hazards models to calculate hazard ratios (HRs) and 95% CIs. Variables with P < 0.1 in univariate analysis and known prognostic indicators such as TNM stage and tumor differentiation were included in the multivariate model for forced adjustment. To address multiple comparisons, associations between clinicopathological features and KRAS mutations were corrected using the Benjamini-Hochberg method for false discovery rate, with significance was set at two-sided P < 0.05. For survival analysis involving predefined clinicopathological variables, the traditional α = 0.05 significance level was retained. Data visualization was performed using the ggplot2 package. Key associations were illustrated using forest plots, and survival curves were annotated with risk numbers and median survival time.
Ethical statement
This retrospective study was conducted in accordance with the national regulations of China and the Declaration of Helsinki. The requirement for written informed consent was formally waived by the Ethics Committee of Jilin Provincial Cancer Hospital due to the retrospective nature of the study and anonymized data processing. The experimental protocol was reviewed and approved by the Ethics Committee of Jilin Provincial Cancer Hospital prior to data collection. All patient data were de-identified to ensure confidentiality.
Results
Clinical and pathological characteristics of patients
Among 256 patients, 57.8% were male and 56.2% were aged ≥ 60 years. The tumors were mainly located in the colon (70.3%), and the mutation rate of the RAS gene was 52.7%. A low tumor budding rate was found in 39.8% of patients, while 60.2% exhibited tumor necrosis. Among the inflammatory cell types, the lymphocyte-dominant type was most common (46.9%), with moderate to strong infiltration intensity seen in 55.1% of cases. Mucinous components were present in 22.3% of tumors, and 56.6% showed an infiltrative growth pattern. PDC were present in 34.8% of patients. In terms of tumor invasion pattern, 32.8% showed a pushing border, 28.9% were intermediate, and 38.3% had an infiltrative pattern. Patients in TNM stage III–IV accounted for 59.8%, and distant metastasis occurred in 24.2%. The rates of neural invasion and vascular invasion were 31.2% and 37.5%, respectively. Poorly differentiated tumors made up 22.27%, while moderately differentiated ones accounted for 70.7%. The majority of patients were microsatellite stable (92.19%), with deficient mismatch repair (dMMR) found in only 7.81%. The mutation rates of KRAS, TP53, APC, and PIK3CA were 54.3%, 55.47%, 48.44%, and 16.8%, respectively. (Table 1).
KRAS gene mutation status
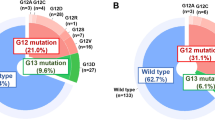
KRAS mutations were detected in 135 of 256 patients (52.7%). Twelve subtypes were identified across codons 12, 13, and 61. The most frequent mutation was G12D (30.37%), followed by G13D (20.00%), G12V (17.78%), and G12C (7.41%). Together, these four subtypes accounted for nearly 80% of all KRAS mutations and are considered classic hotspots. Other common variants included G12C (7.41%) and A146T (5.19%). Several rare mutations such as Q61H (2.22%), Q61L (1.48%), and A146V (0.74%) were also detected, indicating the diversity and heterogeneity of KRAS mutations. Approximately 94% were located at hotspot codons G12/13, indicating these remain the predominant sites of KRAS alterations. (Table 2).
Association of KRAS mutations with clinicopathological and morphological features
Compared with the KRAS wild-type group, the mutation group had a significantly higher proportion of right-sided colon tumors (53.2% vs. 35.6%), aggressive pathological features (high tumor budding: 70.7% vs. 32.5%; presence of mucinous component: 24.4% vs. 19.8%; moderate to high infiltration intensity: 34.7% vs. 17.5%) (P < 0.05). Moreover, co-mutations with PIK3CA were more frequent in the mutation group (24.5% vs. 12.5%). On the other hand, the wild-type group showed a higher proportion of left-sided colon tumors (71.8% vs. 88.1%), rectal tumors (36.2% vs. 52.4%), and TP53 co-mutations (47.1% vs. 58.4%). (Fig. 3)
Association between KRAS mutations and clinicopathological and morphological features. * indicates statistical significance at P < 0.05.
To determine the independent predictive value of the above clinical and molecular characteristics for KRAS mutation, logistic regression analysis was conducted. Univariate analysis revealed that right-sided colon tumors, presence of mucinous component, moderate to strong infiltration, high tumor budding, and PIK3CA co-mutations were all significantly associated with KRAS mutations (P < 0.05 for all). Multivariate analysis confirmed that right-sided colon tumor (OR = 1.85, P = 0.017), high tumor budding (OR = 2.46, P = 0.002), mucinous component (OR = 2.39, P = 0.012), and PIK3CA co-mutation (OR = 2.15, P = 0.042) were independent predictors of KRAS mutation, whereas TP53 co-mutation was protective (OR = 0.59, P = 0.047) (Table 3).
Correlation between KRAS mutation subtypes and clinicopathological and morphological features
Hotspot mutation subtypes from Table 2 were selected for analysis to compare their distribution across clinical, pathological, and molecular characteristics. The results showed that the G12V mutation was significantly associated with a higher proportion of high tumor budding compared with the non-G12V group (23/24, 95.83% vs. 65/111, 58.5%, P = 0.012), and it was also more frequent in tumors with mucinous components (P = 0.001) and moderate lymphocyte infiltration (P = 0.005). The G12C mutation was also more commonly observed in moderate to strong lymphocyte infiltration (P = 0.002) and highly differentiated tumors (P = 0.031) than in the non-G12C group. Other characteristics showed no significant difference between groups (P > 0.05). These findings indicate that G12V and G12C mutations have distinct distributions in certain pathological and molecular features. (Fig. 4)
Analysis of the relationship between G12V and G12C mutations and clinicopathological features. * indicates statistical significance at P < 0.05.
Further comparison was made between G12D (n = 41) and G13D (n = 94) mutations, as well as between G13D (n = 27) and non-G13D (n = 108). Compared with the non-G12D group, patients with G12D mutations showed a significant age distribution difference (< 60 years: 18/41, 44.0% vs. 13/94, 13.8%; ≥60 years: 23/41, 56.0% vs. 81/94, 86.2%, P < 0.05). Regarding MSI/MMR status, G13D mutations also differed from non-G13D mutations (MSS/pMMR: 24/27, 88.9% vs. 82/108, 75.9%; MSI-H/dMMR: 3/27, 11.1% vs. 26/108, 24.1%, P < 0.05). No other clinicopathological features showed significant differences between these groups (P > 0.05). (Fig. 5)
Association of G12D and G13D mutations with clinicopathological and morphological features. * indicates statistical significance at P < 0.05.
Survival analysis of KRAS-Mutated patients
To assess the impact of KRAS mutation status and subtypes on the prognosis of colorectal cancer patients, Kaplan-Meier survival analysis was performed. As shown in Fig. 6a, patients with KRAS mutations had a slightly shorter PFS than wild-type patients, but the difference was not statistically significant (P > 0.05). However, in terms of OS, the KRAS mutation group showed significantly longer survival (Fig. 6b), suggesting a potential long-term survival advantage. Further analysis by KRAS mutation subtypes (G12C, G12D, G12V, G13D) indicated no significant differences in PFS (P = 0.618, Fig. 6c) or OS (P = 0.013, Fig. 6d), suggesting that KRAS subtypes did not significantly affect survival outcomes in this cohort. (Fig. 6).
Kaplan-Meier survival analysis for KRAS mutations and subtypes.
To identify the independent risk factors affecting OS and PFS, the study conducted Cox proportional hazards regression analysis on clinical and molecular characteristics. Univariate analysis showed that TNM stage II–III (OS: HR = 2.50, P = 0.002; PFS: HR = 2.20, P = 0.003) and stage IV (OS: HR = 6.80, P < 0.001; PFS: HR = 5.72, P < 0.001) significantly increased the risk of death and disease progression. Poor tumor differentiation (OS: HR = 2.40, P < 0.001; PFS: HR = 2.10, P = 0.001) and no resection of the primary tumor (OS: HR = 4.20, P < 0.001; PFS: HR = 3.50, P < 0.001) were also strongly linked to poor prognosis. PIK3CA mutation showed borderline statistical significance in OS (HR = 1.45, P = 0.049), while MSI-H/dMMR status acted as a protective factor in both OS and PFS (OS: HR = 0.40, P = 0.008; PFS: HR = 0.45, P = 0.009). Multivariate analysis further confirmed that TNM stage IV (OS: HR = 4.80, P < 0.01; PFS: HR = 4.15, P < 0.001), poor tumor differentiation (OS: HR = 2.00, P = 0.007; PFS: HR = 1.75, P = 0.018), and absence of primary tumor resection (OS: HR = 3.20, P < 0.001; PFS: HR = 2.80, P < 0.001) remained independent adverse prognostic factors. Although MSI-H/dMMR showed significance in the univariate analysis, it did not reach statistical significance in the multivariate model (OS: P = 0.098; PFS: P = 0.108). Mutations in TP53, APC, and PIK3CA did not show independent prognostic effects after adjustment. (Table 4).
Among KRAS-mutated patients, the clinical features and the relationships between various KRAS subtypes and PFS and OS were further explored. Univariate analysis showed that TNM stage II–III (OS: HR = 2.30, P = 0.012; PFS: HR = 2.00, P = 0.024) and stage IV (OS: HR = 4.00, P < 0.001; PFS: HR = 3.50, P < 0.001) significantly shortened OS and PFS. Not undergoing resection of the primary tumor was also a notable poor prognostic factor (OS: HR = 3.20, P < 0.001; PFS: HR = 2.80, P < 0.001). In addition, patients with poorly differentiated tumors had significantly worse survival (OS: HR = 2.00, P = 0.014; PFS: HR = 1.80, P = 0.019). In multivariate analysis, patients with stage II–III and stage IV disease had significantly poorer OS and PFS (all P < 0.05). Poor tumor differentiation also showed a negative prognostic effect in multivariate regression (OS: HR = 1.85, P = 0.034; PFS: HR = 1.65, P = 0.049). Furthermore, patients who did not receive primary tumor resection still had a distinct survival disadvantage (OS: HR = 2.90, P = 0.001; PFS: HR = 2.60, P = 0.001). In contrast, KRAS subtype mutations (including G12C, G12D, G12V, G13D) and other related genes (TP53, APC, PIK3CA) did not show statistically significant differences in OS or PFS. Although the MSI-H/dMMR status appeared to be a potential protective factor in this cohort, it did not reach statistical significance in multivariate analysis (OS: P = 0.210; PFS: P = 0.230). (Table 5).
Discussion
The RAS gene family is one of the most important driver mutations in colorectal cancer, mainly including three subtypes: KRAS, NRAS, and HRAS. Among them, KRAS mutations are the most common, accounting for approximately 85–90% of RAS mutations in colorectal cancer, which is much higher than NRAS (8–12%) and HRAS (less than 2%)14. According to the NCCN guidelines and numerous clinical studies, KRAS mutations not only affect the biological behavior of tumors but also directly determine the effectiveness of targeted therapy (e.g., EGFR monoclonal antibodies)15. Therefore, research focusing on the morphological significance and subtype differences of KRAS mutations has both representativeness and clinical value.
As NRAS mutations occurred in only 3.9% of this study cohort and HRAS mutations are extremely rare in colorectal cancer, this study did not include NRAS or HRAS in the systematic analysis but instead focused on KRAS and its association with pathological structure and prognosis. Based on this, the study analyzed 256 colorectal cancer patients with confirmed RAS status, combining tissue sections and survival data to systematically examine the relationship between KRAS mutations and tumor morphological characteristics, further exploring the morphological variations and clinical relevance across different subtypes.
First, in terms of clinicopathological features, the study found that KRAS mutations were significantly enriched in right-sided colon tumors (53.2% vs. 35.6%, P < 0.05), suggesting an association with anatomical location. This result is consistent with the study by Isermann T et al., based on TCGA data (KRAS mutation rate of 58% in right-sided colon vs. 42% in left-sided colon)16. Previous studies have indicated that the right colon originates from the midgut during embryonic development, and its microenvironment (such as bile acid exposure and intestinal microbiota differences) may more easily trigger activation of the RAS signaling pathway17. Further, the KRAS mutation group showed significantly higher rates of tumor budding (70.7% vs. 32.5%) and mucinous components (87.5% vs. 71.4%), consistent with the mechanism proposed by Negrao M V et al.18, suggesting the KRAS-ERK pathway may be involved in the epithelial–mesenchymal transition process. Especially tumor budding, as a poor prognostic marker, showed a strong correlation with KRAS mutations in this study (OR = 2.46, P = 0.002).
Second, regarding KRAS mutation subtypes, the study found that G12D (30.37%), G13D (20.00%), and G12V (17.78%) were the most common, consistent with distributions reported in East Asian populations19, but different from Western cohorts where G12V was more common (e.g., 25–30% in North American patients20, suggesting that ethnicity or environmental factors may influence mutation patterns. Further analysis showed that G12V was associated with well-differentiated tumors (P = 0.012) and lymphocyte infiltration (P = 0.005), while G12C was often observed in cases with moderate to severe inflammatory infiltration (P = 0.002). The morphological and immunophenotypic evidence of KRAS G12C in colorectal cancer remains limited. Clinical studies have reached a more consistent conclusion: the incidence of G12C is relatively low (about 3–4%). It usually shows primary resistance to EGFR monotherapy. However, KRAS G12C inhibitors combined with EGFR blockade have demonstrated meaningful efficacy in patients who received prior treatments21. This combination has already obtained accelerated approval from the FDA. Therefore, the observations related to G12C in this study should be interpreted with caution and require confirmation in prospective research.
Third, regarding the prognostic value of KRAS mutations. Unlike most studies15, this study found better OS in patients with KRAS mutations than in those with wild-type KRAS (P = 0.013), while no significant differences were observed among subtypes. Possible explanations include: (1) a higher TP53 mutation rate in the wild-type group (58.4% vs. 47.1%), and TP53 loss may promote malignant progression through genomic instability22; (2) the KRAS-mutant group may harbor distinct co-mutation profiles or immunological microenvironmental characteristics, such as a higher frequency of PIK3CA co-mutations or more pronounced inflammatory responses, which could exert a protective effect on long-term survival. Further COX regression analysis showed that TNM stage (stage IV HR = 4.80) and lack of primary tumor resection (HR = 3.20) were independent prognostic factors, consistent with the conclusions of Siegel et al.23, suggesting that traditional pathological parameters remain central to prognosis assessment.
In summary, the above findings demonstrate the multifaceted role of KRAS mutations in clinicopathological characteristics, subtype distribution, and prognosis, and they also suggest potential applications of these morphological features in diagnostic practice. In this study, KRAS mutations were more frequently observed in right-sided tumors and were significantly associated with prominent tumor budding and abundant mucinous components. These features may serve as useful indicators in clinical practice, particularly in settings where molecular testing is limited or unavailable. For example, in the absence of genetic testing, the observation of marked tumor budding or pronounced mucinous differentiation may alert pathologists and clinicians to the possible activation of the RAS pathway. Such morphological–molecular correlations not only enhance the understanding of colorectal cancer biology but may also provide complementary guidance for individualized patient management.
Despite a large sample size and standardized morphological evaluation, several limitations remain: (1) the retrospective design may have introduced selection bias, such as the inclusion of only surgical cases; (2) NRAS and BRAF mutations were not analyzed, possibly affecting the completeness of molecular classification; (3) lack of treatment response data made it difficult to evaluate the impact of different mutations on EGFR-targeted therapy or chemotherapy. (4) because information on comorbidities, performance status, postoperative complications, and treatment distribution was unavailable, the observed association between KRAS mutations and improved OS may have been confounded by TP53 mutation imbalance, therapeutic variability, or other unmeasured factors; (5) the number of cases with rare KRAS variants such as G12F and A146V limited the statistical power of subtype-specific prognostic analyses. Future research should involve multicenter, prospective data, and incorporate spatial transcriptomics or single-cell sequencing to further reveal the dynamic interaction between KRAS subtypes and the tumor microenvironment, and explore the therapeutic potential of subtype-specific inhibitors (e.g., sotorasib) in colorectal cancer.
Data availability
The datasets generated and/or analyzed during the current study are not publicly available due to patient privacy and ethical restrictions but are available from the corresponding author on reasonable request.
Change history
14 January 2026
A Correction to this paper has been published: https://doi.org/10.1038/s41598-026-36079-z
References
Sun, D., Liu, Y., Li, P., Yang, P. & Yu, G. Clinicopathological features and prognosis of lung adenocarcinoma patients with K-RAS gene mutation. Appl. Immunohistochem. Mol. Morphol. 31 (6), 406–413 (2023).
Fujii, S. et al. Rapid screening using pathomorphologic interpretation to detect BRAF V600E mutation and microsatellite instability in colorectal cancer. Clin. Cancer Res. 28 (12), 2623–2632 (2022).
Ochman, B. et al. Association of SIGLEC9 expression with cytokine expression, tumor grading, KRAS, NRAS, BRAF, PIK3CA, AKT gene mutations, and MSI status in colorectal cancer. Curr. Issues Mol. Biol. 46 (12), 13617–13646 (2024).
Poter, P. et al. Association of PD-1 and PD-L1 protein expression with selected clinical and morphological parameters in colorectal cancers. Pol. J. Pathol. 75 (2), 105–114 (2024).
Aliyarov, Y. R. The role of KRAS and BRAF gene mutations as predictive factors in the treatment of locally advanced rectal cancer. Kazan Med. J. 106 (1), 98–104 (2025).
Takeda, M. et al. The role of KRAS mutations in colorectal cancer: biological insights, clinical implications, and future therapeutic perspectives. Cancers 17 (3), 428 (2025).
Hashimoto, T. et al. The SCRUM-MONSTAR cancer-omics ecosystem: striving for a quantum leap in precision medicine. Cancer Discov. 14 (11), 2243–2261 (2024).
Naumov, S. S. et al. Clinicopathological features of colon cancer depending on the dMMR status of the tumor. Bull. Sib Med. 22 (4), 57–64 (2024).
Healy, F. M., Prior, I. A. & MacEwan, D. J. The importance of Ras in drug resistance in cancer. Br. J. Pharmacol. 179 (12), 2844–2867 (2022).
Escher, T. E. & Satchell, K. J. RAS degraders: the new frontier for RAS-driven cancers. Mol. Ther. 31 (7), 1904–1919 (2023).
Punekar, S. R., Velcheti, V., Neel, B. G. & Wong, K. K. The current state of the Art and future trends in RAS-targeted cancer therapies. Nat. Rev. Clin. Oncol. 19 (10), 637–655 (2022).
Siegel, R. L., Kratzer, T. B., Giaquinto, A. N., Sung, H. & Jemal, A. Cancer statistics. CA Cancer J. Clin. 75 (1), 10 (2025).
Kobayashi, Y. et al. Silent mutations reveal therapeutic vulnerability in RAS Q61 cancers. Nature 603 (7900), 335–342 (2022).
Ono, H. et al. Schimmelpenning-Feuerstein-Mims syndrome induced by HRAS Gly12Ser somatic mosaic mutation: case report and literature review. J. Dermatol. 50 (9), 1213–1215 (2023).
Yaeger, R. et al. Efficacy and safety of adagrasib plus cetuximab in patients with KRAS G12C-mutated metastatic colorectal cancer. Cancer Discov. 14 (6), 982–993 (2024).
Isermann, T., Sers, C., Der, C. J. & Papke, B. KRAS inhibitors: resistance drivers and combinatorial strategies. Trends Cancer. 11 (2), 91–116 (2025).
Johnson, C., Burkhart, D. L. & Haigis, K. M. Classification of KRAS-activating mutations and the implications for therapeutic intervention. Cancer Discov. 12 (4), 913–923 (2022).
Negrao, M. V. et al. Comutations and KRASG12C inhibitor efficacy in advanced NSCLC. Cancer Discov. 13 (7), 1556–1571 (2023).
Ge, Z. et al. Recent advances in targeted degradation in the RAS pathway. Future Med. Chem. 17 (6), 693–708 (2025).
Ilhan, N. et al. Regional and gender-based distribution of KRAS mutations in metastatic colorectal cancer patients in Turkey: an observational study. Med. (Kaunas). 61 (4), 694 (2025).
Fakih, M. G. et al. Sotorasib plus panitumumab in refractory colorectal cancer with mutated KRAS G12C. N Engl. J. Med. 389 (23), 2125–2139 (2023).
Krishnan, T., Roberts-Thomson, R., Broadbridge, V. & Price, T. Targeting mutated KRAS genes to treat solid tumours. Mol. Diagn. Ther. 26 (1), 39–49 (2022).
Sacher, A. et al. Single-agent divarasib (GDC-6036) in solid tumors with a KRAS G12C mutation. N Engl. J. Med. 389 (8), 710–721 (2023).
Author information
Authors and Affiliations
Contributions
L.G. and S.W. conceived and designed the study. Y.W., L.G., and C.Y. performed the experiments and analyzed the data. W.Q. and S.W. contributed to data interpretation. Y.W. and S.W. wrote the manuscript. All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: The original version of this Article contained an error in the order of the author names, which was incorrectly given as Yifei Wang, Lili Guo, Chunxue Yang, Wenchao Qi, Shuai Wang.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Guo, L., Wang, Y., Yang, C. et al. Correlation analysis between RAS gene mutations and pathological morphological features in colorectal cancer. Sci Rep 15, 43515 (2025). https://doi.org/10.1038/s41598-025-26317-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-26317-1