Abstract
Triple-negative breast cancer (TNBC) is a highly aggressive malignancy lacking effective therapeutic strategies, resulting in a poor patient prognosis. In this study, we investigated the efficacy and underlying molecular mechanisms of the combination of FAK inhibitor Ifebemtinib and paclitaxel in TNBC. We found that FAK were significantly upregulated in breast cancer and correlated with poor prognosis in both breast cancer and TNBC patients. In vitro, the combination of Ifebemtinib and paclitaxel synergistically suppressed proliferation and colony-formation of TNBC cells. Through inactivating CDK1, they induced cell cycle arrest at G2/M phase. By down-regulating Bcl-2 and up-regulating Bax, the combination induced triggered apoptosis. By up-regulating E-cadherin, down-regulating N-cadherin and vimentin, they inhibited migration of TNBC cells. In vivo, the combination inhibited TNBC cell proliferation and spontaneous lung metastasis. Mechanistically, on one hand, the combination synergistically inhibited phosphorylation-mediated activation of FAK. On the other hand, they downregulated LSD1, thereby relieving LSD1-mediated transcriptional repression of PIK3IP1, which is a negative regulator of the PI3K/Akt pathway, leading to accumulation of PIK3IP1. By inhibiting FAK activation and upregulating PIK3IP1, Ifebemtinib and paclitaxel blocked the PI3K/Akt pathway, effectively suppressing TNBC proliferation and metastasis. Our findings suggest that FAK may serve as a potential therapeutic target for TNBC.
Similar content being viewed by others
Introduction
Breast cancer is a common malignant tumor. According to the GLOBOCAN2022 released by the International Agency for Research on Cancer (IARC), in 2022, the incidence and mortality of female breast cancer ranked second and fourth among malignant tumors, with both incidence and mortality ranking first among female malignancies1. TNBC refers to a type of breast cancer that is negative for estrogen receptors, progesterone receptors, and human epidermal growth factor receptor 2 (EGFR, HER-2), accounting for approximately 15% to 20% of all breast cancers. TNBC is one of the most challenging types of breast cancer to treat, characterized by early onset age, high heterogeneity, high malignancy degree, high risk of recurrence, and poor prognosis2, with a 3-year recurrence rate reaching 19%3. Due to the lack of effective treatment targets, chemotherapy is currently the main treatment for TNBC patients. Paclitaxel (PTX) is commonly used as a first-line chemotherapy drug for triple-negative breast cancer; however primary resistance and acquired resistance limit its clinical application and lead to disease progression. Therefore, exploring the mechanisms of primary resistance of paclitaxel in TNBC and developing new strategies to enhance sensitivity to PTX treatment has become crucial in improving paclitaxel therapy for TNBC.
Focal adhesion kinase (FAK) is a type of cytoplasmic non-receptor protein tyrosine kinase (PTKs). FAK can receive signals from integrins, growth factors, mechanical stimuli, etc., activate intracellular signaling pathways such as PI3K/Akt and Ras/MAPK, regulate cell growth and cytoskeleton rearrangement, and participate in important processes such as embryonic development, tumor occurrence and migration4,5. The FAK expression is frequently elevated in various cancers such as ovarian, lung squamous cell neoplasms, esophageal and uveal melanoma tumors, and it is associated with a poor patient prognosis6. In breast cancer, the activation of FAK has been confirmed to mediate brain and bone metastasis of breast cancer7,8.
Ifebemtinib (IN10018 or BI-853520) is one of the ATP-competitive FAK inhibitors9 that have shown anti-cancer effect in multiple preclinical tumor models and are being evaluated in combination regimens in early clinical studies. In preclinical, Ifebemtinib inhibits the proliferation of various human adenocarcinoma cell lines in vitro by inhibiting FAK phosphorylation activation10. In ovarian cancer, Ifebemtinib inhibits cell proliferation, migration, invasion through PI3K/AKT/mTOR signaling pathway11. Zhang. et al. reported that Ifebemtinib enhanced the immunogenic cell death of PEGylated liposome doxorubicin, further benefiting the anti-tumor effect of immune checkpoint blockade12. In pancreatic ductal adenocarcinoma, Ifebemtinib enhances antitumor response of radiation therapy through enhances CD8 + T cell infiltration13. In breast cancer, the FAK PROTECT degrader based on Ifebemtinib not only inhibits the proliferation, migration, and invasion of breast cancer cells but also can reverse their drug resistance14. Apart from being used alone, Ifebemtinib is also used in combination with other drugs to treat tumors. It is reported that Ifebemtinib combined with KRAS G12C inhibitors confer synergic anticancer effects in CDX and PDX tumor models through blocked the FAK-YAP signaling. In TNBC, combination of Ifebemtinib and crizotinib suppressed proliferation and promoted apoptosis15. Ifebemtinib improves the tumor uptake of antibody-drug conjugates to strengthen the anti-cancer responses in non-small cell lung as well as pancreatic cancer PDX models16. Due to excellent preclinical results, Ifebemtinib was therefore tested in phase I-III trials for the treatment of solid tumors. So far, more than 15 clinical trials have been conducted on Ifebemtinib17,18 including ovarian cancer, pancreatic cancer, cervical adenocarcinoma, gastric cancer and so on. Based some of these results, Ifebemtinib received fast track designation by the U.S. Food and Drug Administration (FDA) and breakthrough therapy designation by the China National Medical Products Administration (NMPA) for platinum-resistant ovarian cancer in 2021 and 2022, respectively.
Ifebemtinib is expected to overcome the tumor stroma fibrotic barrier, enhance local immune function, and thus improve the efficacy of targeted therapy, chemotherapy, and immunotherapy12,13,15,19. However, whether Ifebemtinib can enhance the effectiveness of paclitaxel in treating TNBC has not been reported.
In this study, we found that FAK is highly expressed in breast cancer and is associated with poor prognosis of breast cancer patients. In addition, we confirmed that the FAK inhibitor Ifebemtinib used in combination with paclitaxel can synergistically inhibit the proliferation and metastasis of TNBC cells both in vitro and in vivo, suggesting a novel therapeutic strategy for TNBC.
Results
The expression of FAK was associated with the poor prognosis of TNBC
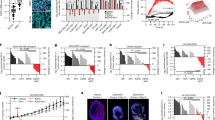
To investigate the clinical significance of FAK in TNBC, we firstly performed a comprehensive survey of mRNA levels in normal breast and breast cancer tissue. As shown in Fig. 1A, the level of FAK was higher in breast cancer tissue in comparison to normal breast tissue of breast invasive carcinoma (BRCA) patients. To further verify this result, we examined the expression of protein level of FAK in breast cancer tissues and adjacent tissues of 72 patients with breast cancer (Fig. 1B). The results in 1 C reveal that, the protein level of FAK in breast cancer tissues was significantly higher than that in adjacent tissues (P < 0.001). Further, the results in 1D reveal that, the protein level of FAK in TNBC tissues was significantly higher than that in adjacent tissues (P < 0.001). What’s more, the data form TIMER showed that high level of PTK2 is associated with poor prognosis both in BRCA (Fig. 1E) and BRCA-Basal (Fig. 1F) in which TNBC accounts for more than 85%.
The expression of FAK was associated with the poor prognosis of TNBC. A The mRNA levels of PTK2 that encoding FAK in breast cancer tissues and normal breast tissues from patients with breast invasive cancer (BRCA) according to the TIMER database. B Representative images of FAK protein expression in breast cancer patient tissues detected by immunohistochemical staining. C The protein levels of FAK in breast cancer tissues and adjacent tissues of 72 breast cancer patients. D The protein levels of FAK in TNBC tissues and adjacent tissues of 38 breast cancer patients. E TIMER data showed that the high level of PTK2 was associated with poor outcomes for BRCA. F TIMER data showed that the high level of PTK2 is associated with poor outcomes for BRCA-basal. *** P < 0.001 versus NC.
Ifebemtinib and Paclitaxel synergistically reduced the survival of TNBC cells
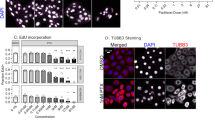
To develop novel sensitizer for paclitaxel in treatment of TNBC, CCK8 assay was used to determine the cytotoxicity of Ifebemtinib and paclitaxel alone or in combination on TNBC cells. The results in Fig. 2 revealed that each of Ifebemtinib or paclitaxel inhibited the cell viability of TNBC cells MDA-MB-231 and MDA-MB-436 in a concentration- and time- dependent manner (Fig. 2A and B). The IC50 of Ifebemtinib was (22.07 ± 2.03), (13.56 ± 2.87), (6.33 ± 1.31) µM on MDA-MB-231, and (21.47 ± 0.48), (17.53 ± 1.12), (11.81 ± 0.90) µM on MDA-MB-436 for 24 h, 48 h, 72 h, respectively. To test whether Ifebemtinib can enhance the anti-tumor effect of paclitaxel as a sensitizer, we tested the effects of Ifebemtinib, and paclitaxel used alone and in combination on cell viability of MDA-MB-231 and MDA-MB-436 for 24 h. According to the cytotoxicity of Ifebemtinib, 12 µM of Ifebemtinib were chosen in the following research. Notably, compared to each drug alone, lower cell viability was observed when Ifebemtinib or paclitaxel were used together in MDA-MB-231 (Fig. 2C) and MDA-MB-436 cells (Fig. 2D). To accurately clarify whether the combination of Ifebemtinib and paclitaxel acts synergistically, additively, or antagonistically, we calculated the combination index (CI) using CompuSyn (ComboSyn, Inc., Paramus, NJ, USA). As shown in Fig. 2E and F, at medium concentrations of paclitaxel, the CI was less than 1. Especially when the concentration of paclitaxel is around 3 µM, the CI value is the smallest, indicating that currently the synergistic effect of these two drugs is strongest. Thus, 4 µM of paclitaxel was used in the following experiments. Meanwhile, we observed the morphology of cells after drug treatment. As shown in the upper panel of Fig. 2G, MDA-MB-231 cells in the negative control (NC) group exhibit an epithelial cell-like growth morphology, mostly spindle-shaped, and can adhere firmly to the culture dish. After treating the cells with Ifebemtinib for 48 h, there was no significant change in cell status. Following treatment with paclitaxel for 48 h, most of the cells lost their spindle shape and became flat like pancakes or droplets with reduced attachment. In the combination group, cell morphology changed to larger flattened pancake shapes with a significant decrease in cell numbers. As can be seen in the lower panel of Fig. 2G, after treatment with Ifebemtinib, many vacuoles formed inside the cells. Treatment with paclitaxel caused the cells to become round and dark. When the two drugs were used together, the number of cells decreased significantly, and cell fragmentation occurred. These data suggested that Ifebemtinib and paclitaxel synergistically reduced the survival of TNBC cells.
Ifebemtinib and paclitaxel synergistically reduced the survival of TNBC cells. A, B In MDA-MB-231 (Fig. 2A) and MDA-MB-436 (Fig. 2B) TNBC cells, Ifebemtinib inhibited cell viability in a concentration- and time-dependent manner. C, D Ifebemtinib enhanced the proliferative inhibitory effects of PTX on MDA-MB-231 and MDA-MB-436 cells at 24 h. E, F The combination index (CI) values of Ifebemtinib and PTX in MDA-MB-231 and MDA-MB-436 cells. G The morphology of MDA-MB-231 and MDA-MB-436 cells after being treated with Ifebemtinib and PTX for 24 h. H Heat map of differentially expressed genes (DEGs) after treatment of Ifebemtinib (Ifeb), paclitaxel (PTX) or their combination (COM) of MDA-MB-231 cells. I Histograms of the number of DEGs between each two groups of negative control (NC), Ifebemtinib (Ifeb), paclitaxel (PTX) and the combination (COM). J GO analysis of DEGs between NC group and COM group. K KEGG analysis of DEGs between NC group and COM group.
Ifebemtinib and Paclitaxel synergistically delayed proliferation of TNBC cells in vitro
To clarify the mechanism of Ifebemtinib and paclitaxel synergistically inhibiting TNBC, we treated MDA-MB-231 cells with drugs and performed RNA sequencing. The heatmap of differentially expressed genes (DEGs) can be seen in Fig. 2H. Compared to the negative control (NC) group, over 2000 genes showed changes in mRNA levels after treatment with paclitaxel and drug combination of paclitaxel Ifebemtinib (COM), while only 75 genes changed after Ifebemtinib (Ifeb) treatment (Fig. 2I). Subsequently, we conducted GO analysis (Fig. 2J) and KEGG (Fig. 2K) analysis on the differentially expressed genes between NC vs. COM. The GO analysis showed that the enriched differentially expressed genes are involved in multiple pathways related to proliferation and mitosis (Red box in Fig. 2J). Based on this, we speculated that the combination of Ifebemtinib and paclitaxel may inhibit the proliferation of MDA-MB-231. Thus, we first detected cell proliferation using clone-formation assay. Both Ifebemtinib and paclitaxel alone can significantly inhibit the formation of cell clones in MDA-MB-231 (Fig. 3A and B) and MDA-MB 436 (Fig. 3C and D), and after using both drugs together, the cells can hardly form any clones (Fig. 3A-D). Next, cell mitosis was detected using CFDA-SE staining by flow cytometry. As shown in Fig. 3E, the percentages of cells in M3 gate in four groups was 0.17%-0.49% at 0 h, with no statistical difference between NC and treatment groups. After 24 h, the percentage of cells in M3 gate of NC, Ifebemtinib, paclitaxel and combination group increased to 77.35%, 74.77%, 33.24%, and 15.36% respectively (Fig. 3F), indicating that compared to NC, paclitaxel alone or drug combination significantly inhibited cell mitosis of MDA-MB-231. Meanwhile, this result was also validated in the MDA-MB-436 cells (Fig. 3G and H).
The combination of Ifebemtinib and paclitaxel inhibit the proliferation of TNBC cells. A, B The effect of Ifebemtinib and paclitaxel on colony-formation of MDA-MB-231 cells. D, D The effect of Ifebemtinib and paclitaxel on colony-formation of MDA-MB-436 cells. E, F The mitosis of MDA-MB-231 cells was detected by flow cytometry with CFDA-SE staining. F Compared to NC and Ifebemtinib, drug combination significantly inhibited cell mitosis of MDA-MB-231. G The mitosis of MDA-MB-436 cells was detected by flow cytometry with CFDA-SE staining. H Compared to NC, Ifebemtinib or paclitaxel alone, drug combination significantly inhibited cell mitosis of MDA-MB-436. ** P < 0.01 versus negative control (NC), *** P < 0.001 versus NC. # P < 0.05, ## P < 0.01, ### P < 0.001.
Ifebemtinib and Paclitaxel synergistically induced cell cycle arrest at G2/M phase
Meanwhile, the KEGG analysis results show that differentially expressed genes are enriched in cell cycle-related pathways (Red box, Fig. 2K). We analyzed the cell cycle distribution of TNBC cells treated with Ifebemtinib and paclitaxel. Figure 4A and B indicated that Ifebemtinib caused cell cycle arrest in G0/G1 phase, while PTX induces significant arrest in G2/M phase. After the combination of the two drugs, MDA-MB-231 cells are mainly arrested in the G2/M phase. A similar phenomenon of cell cycle arrest was also observed in MDA-MB-436 cells (Fig. 4C and D). The Western blotting in Fig. 4I revealed that combination of Ifebemtinib and paclitaxel inhibited CDK1 by up-regulating p-CDK1 (Y15) and down-regulating p-CDK1 (T161). In addition, p27 and the ratio of LC3B-I/LC3B-II were also increased, deterring that the combination of the two drugs may not only induce G2/M phase cell cycle arrest but also potentially induce autophagy in TNBC.
Ifebemtinib and paclitaxel synergistically induced cell cycle arrest at G2/M phase and apoptosis of TNBC cells. A, B Effect of Ifebemtinib, paclitaxel and their combination on cell cycle distribution of MDA-MB-231 cells. C, D Effect of Ifebemtinib, paclitaxel and their combination on cell cycle distribution of MDA-MB-436 cells. E, F Effect of Ifebemtinib, paclitaxel and their combination on apoptosis of MDA-MB-231 cells. G, H Effect of Ifebemtinib, paclitaxel and their combination on apoptosis of MDA-MB-436 cells. I Western blotting detected the effect of Ifebemtinib combined with paclitaxel on the cell cycle- and autophagy-related proteins in MDA-MB-231 cells. J Western blotting detected the effect of Ifebemtinib combined with paclitaxel on the apoptosis-related proteins in MDA-MB-231 cells. * P < 0.05 versus NC, ** P < 0.01 versus NC, *** P < 0.001 versus NC. ## P < 0.01.
Ifebemtinib and Paclitaxel synergistically induced apoptosis of TNBC cells through the mitochondrial pathway
Chemotherapy drugs usually inhibit tumor progression by inducing apoptosis in cells. Thus, we detected apoptosis of MDA-MB-231 and MDA-MB-436 cells after treatment. As shown in Fig. 4E and F, the apoptosis rates of the Ifebemtinib group and paclitaxel group were 16.69% and 26.61%, respectively, while the apoptosis rate of the combination drug group was 42.34%. Compared to the control group with an apoptosis rate of 5.58%, there was a statistical difference between Ifebemtinib and combination treatment, with a significant increase in apoptosis rate in the combination drug group compared to the paclitaxel monotherapy group. Although the apoptosis rates of MDA-MB-436 cells were lower after treatment with the same concentration of drugs, the apoptosis rate increased when the two drugs are used in combination (Fig. 4G). Apoptosis is a sophisticated and complex cascade regulatory process in which there are two main pathways: the extrinsic pathway (death receptor pathway) and the intrinsic pathway (mitochondrial pathway). At the same time, these two pathways are interconnected and jointly involved in regulating apoptosis. To understand how paclitaxel and Ifebemtinib synergistically triggered apoptosis in TNBC cells, we first measured the key regulators of apoptosis. As in Fig. 4J, after drug treatment, the anti-apoptotic protein Bcl-2 did not show significant changes, but the pro-apoptotic protein Bax was significantly upregulated. Cytochrome C was released into the cytoplasm. The activated apoptosis markers including caspase 9, caspase 3 and PARP1 were significantly upregulated, suggesting that the combination of the two drugs may primarily induce apoptosis synergistically through mitochondrial pathways.
Ifebemtinib and Paclitaxel reduced migration of TNBC cells in vitro
Compared to other types of breast cancer, TNBC is highly invasive, prone to metastasis, which is closely related to poor prognosis of patients. Therefore, we tested the effects of Ifebemtinib and paclitaxel alone or in combination on cell migration capability. The results indicated that, compared with single-drug therapy, drug combination of Ifebemtinib and paclitaxel has a more significant inhibitory effect on the migration of MAD-MB-231 cells (Fig. 5A and B) as well as MAD-MB-436 cells (Fig. 5C and D). Additional Western blot analysis in Fig. 5E demonstrated that the combination of Ifebemtinib and paclitaxel synergistically increased the level of E-cadherin and decreased level of N-cadherin and Vimentin, suggesting that the combination of Ifebemtinib and paclitaxel reversed EMT of TNBC cells.
Ifebemtinib and paclitaxel synergistically reduced migration of TNBC cells in vitro. A, B Effect of Ifebemtinib, paclitaxel and their combination on migration of MDA-MB-231 cells. C, D Effect of Ifebemtinib, paclitaxel and their combination on migration of MDA-MB-436 cells. E Effect of Ifebemtinib, paclitaxel and their combination on EMT-related proteins in MDA-MB-231 cells. * P < 0.05 versus the NC, ** P < 0.01 versus NC, *** P < 0.001 versus NC. # P < 0.05, ## P < 0.01.
Ifebemtinib and Paclitaxel suppressed the growth of TNBC cells in vivo
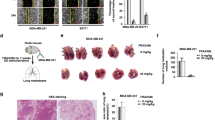
To assess the synergistic anticancer effect of Ifebemtinib and paclitaxel in vivo, we established MDA-MB-231xenograft models. At the end of the experiment, tumors of each group are shown in Fig. 6A. Compared with the vehicle control group, the tumor volumes (Fig. 6B) and weight (Fig. 6C) in Ifebemtinib and paclitaxel monotherapy groups showed a decrease. However, in the combination therapy group, both the average volume and weight of tumors were significantly lower than those in the vehicle control group, indicating that drug combination synergistically inhibited proliferation of MDA-MB-231 cells in vivo. The bodyweight of mice is an important indicator for preliminary evaluation of the toxicity of the treatments. At the end of therapy, we observed that there was no significant difference in bodyweight among the four groups of mice (Fig. 6D), suggesting that, compared to the mono-therapy, the combined treatment did not increase toxicity. The HE staining in Figure E indicated that in the vehicle group of mice, tumor cells grow densely in the tumor tissue. In the paclitaxel treatment group, the local tumor cell density decreased. In the Ifebemtinib treatment group, a small number of cavities were observed. After the two drugs were used in combination, the cell density in the tumor tissue significantly decreased and the number of cavities increased, indicating that combined therapy caused cell necrosis or apoptosis. The results of immunohistochemistry showed that over 85% of cells in the tumor tissue express the proliferation markers Ki67, indicative of tumor proliferation, in the vehicle group. In the single drug treatment group, about 50% of tumor cells in the tumor tissue express Ki67, while in the combination therapy group, only 20% of tumor cells express Ki67 (Fig. 6E), suggesting that combination therapy is more effective than single drug treatment. TUNEL staining was conducted to detect the apoptosis. We found that compared to the vehicle control group, there was a slightly increased number of green-positive signals in the tumor tissues of Ifebemtinib and paclitaxel monotherapy groups. Moreover, regardless of comparison with the vehicle control group or the monotherapy group, there was a significant increase in green-positive signals in the tumor tissues of the combination therapy group (Fig. 6F). This result suggests that the combined treatment of Ifebemtinib and paclitaxel synergistically induced apoptosis in vivo. These data suggest that, compared to monotherapy, the combined therapy effectively inhibited tumor growth, suppressed proliferation and promoted apoptosis.
To further evaluate the toxicity of the drug, we tested the blood routine and blood biochemistry of mice. The results of blood routine in Table 1 showed that, compared with the control group, the combination of drugs led to a significant decrease in white blood cells and neutrophils in mice. The results of blood biochemical tests are shown in Fig. 6G-L, and compared with the vehicle control group, there were no significant changes in total protein, albumin (ALB), direct bilirubin, and other indicators in peripheral blood of mice for both monotherapy and combination therapy groups, except for a significant increase in aspartate transaminase (AST). While, in the four groups, the AST levels remained within the normal physiological range, indicating no hepatic toxicity induced by the treatment. At the same time, there was no statistically significant difference in creatinine (Cre, Fig. 6M) and urea (Fig. 6N) levels in the peripheral blood of mice, suggesting that the combined use of drugs did not have a significant impact on renal function of mice. In addition, western blot revealed that the combination of Ifebemtinib and paclitaxel activated caspase 3, 7 and PARP1 through down-regulating Bcl-2 as well as p-FAK (Y397) and up-regulating Bax. The above data indicated that the combination of Ifebemtinib and paclitaxel synergistically inhibited TNBC cell proliferation in vivo without increasing toxicity in vivo.
Ifebemtinib and paclitaxel suppressed the growth and lung metastasis of TNBC in vivo. A The image of subcutaneous tumors of MDA-MB-231 cells at the end of therapy. B Tumor volume of MDA-MB-231 tumors during therapy. C Tumor weight of MDA-MB-231 tumors during the therapy. D Bodyweight mice during the therapy. E HE and IHC analysis of Ki67 and Vimentin in the subcutaneous tumor tissues. The scale bar is 30 μm. F TUNEL analysis was conducted to detect apoptosis in the subcutaneous tumor tissue. The scale bar is 100 μm. G-L Blood biochemistry analysis for evaluating liver function. Blood biochemistry analysis for evaluating liver function of mice. M, N Biochemical analysis of blood to evaluate kidney function of mice. O Western blotting detected the effect of Ifebemtinib combined with paclitaxel on the apoptosis-related proteins in subcutaneous implant tumor of MDA-MB-231 cells. * P < 0.05 versus NC, ** P < 0.01 versus NC, *** P < 0.001 versus NC.
Ifebemtinib and Paclitaxel synergistically repressed spontaneous metastasis of breast cancer to the lung
To further validate the inhibitory effects of Ifebemtinib combined paclitaxel on metastasis of breast cancer, we established orthotopic mouse lung metastasis model20. By counting the number of lung metastatic lesions in mice (Fig. 7A and B), we found that there was no significant difference in the number of lung metastatic lesions between the mono-drug treatment group and the vehicle group suggesting that Ifebemtinib and paclitaxel had a mild inhibitory effect on lung metastasis. The combination therapy led to significant decrease of lung metastatic lesions (Figure A, B and C). These data suggested that Ifebemtinib plus paclitaxel synergistically reduced spontaneous metastasis of breast cancer from the primary area to the lungs.
Ifebemtinib and paclitaxel blocked orthotopic lung metastasis of mouse BC cells in vivo. A, B Effect of Ifebemtinib, paclitaxel and their combination on orthotopic lung metastasis of 4T1 cells in vivo. C HE detection of mice lungs. The black square marks the metastatic lesions in the lung. The scale bar is 250 μm. * P < 0.05 versus NC.
Combination of ifebemtinib and Paclitaxel collaborated to block PI3K/Akt pathway
To further explore the synergistic anti-tumor mechanism of Ifebemtinib and paclitaxel, we conducted further analysis on the RNA sequencing results. As shown in the Venn diagram (Fig. 8A), there are a total of 26 genes in the intersection of differentially expressed genes (DEGs) from the three groups of NC (negative) vs. Ifebemtinib, NC VS PTX (paclitaxel), and NC vs. COM (combination). By performing series test of cluster, 9 clusters were produced (Fig. 8B). We first focused on the cluster 4, in which single drug use led to upregulation of genes, while in the combination group, these genes were further upregulated. Among these 26 genes, 17 genes were in the cluster 4 (Fig. 8C). We found that the negative regulator of the PI3K pathway PI3KIP1 (Fig. 8C, red box) was up regulated. PIK3IP1 negatively regulates PI3K activity by directly binding to the p110 catalytic subunit of PI3K. Through western blot, we found that both Ifebemtinib and PTX, either used alone or in combination, led to upregulation of PI3KIP1 protein expression in MDA-MB-231 (Fig. 8H). Additionally, the phosphorylation level of Akt, a downstream target of PI3K, was also inhibited, indicating the combination of Ifebemtinib and paclitaxel blocked PI3K/Akt pathway through upregulating of PI3KIP1. It has been reported that the transcription of PIK3IP1 is regulated by the histone lysine-specific demethylase (LSD1)21. The TIMER database shows that in breast cancer, the expression level of KDM1A is positively correlated with expression level of PIK3CA (encoding p110 catalytic subunit of PI3K, Fig. 8D) and PIK3CB (encoding p85 regulatory catalytic subunit of PI3K, Fig. 8E), suggesting that high expression of LSD1 may be a reason for the abnormal activation of PI3K/Akt in breast cancer. What’s more, KDM1A was found in cluster 2 in Fig. 8B in which single drug use led to downregulation of genes, while after combination drug use, these genes were further downregulated (Fig. 8F). Thus, we detected protein level of LSD1. After the two drugs were used in combination, the expression level of LSD1 protein decreased by more than 90% (Fig. 8H). The above results indicate that the combination of Ifebemtinib and PTX may downregulate LSD1, relieve LSD1’s transcriptional repression on PIK3IP1, thereby upregulating PIK3IP1 and inhibiting activation of the PI3K pathway. At the same time, we also detected another gene DDIT4 in Fig. 8C. As in Fig. 8H, whether used alone or in combination with both drugs, DDIT4 was upregulated. It has been reported that the upregulation of DDIT4 in TNBC promotes autophagy by inhibiting mTOR22. According to these data, we speculate that the combination of Ifebemtinib and PTX mainly promotes TNBC cell autophagy by up-regulating DDIT4. Finally, we analyzed the transcription factors of DEGs and found that only one gene (Fig. 8G), ELF3, was identified as an intersection among the three groups of DEGs. As in Fig. 8H, ELF3 was significantly upregulated in Ifebemtinib and PTX-treated cells. The co-treatment of both drugs further reduced the expression level of ELF3 protein. It is reported that the lung cancer cell migration rate can be enhanced by knockdown of ELF3, suggesting that it is a key regulator of epithelial-mesenchymal transition23. Based on this, we speculate that the combination of Ifebemtinib and PTX may promote E-cadherin and inhibit N-cadherin expression by suppressing ELF3 regulation, thus inhibiting EMT and migration of TNBC cells.
Combination of Ifebemtinib and paclitaxel collaborated to block LSD1/PI3K/Akt pathway. A Venn diagram displaying the overlaps of DEGs among different groups of MDA-MB-231 cells. B Series Test of Cluster of DEGs between group of NC and COM. C Heat map of 26 genes of overlaps of DEGs among NC vs. Ifebemtinib, NC VS PTX and NC vs. COM. D, E According to the TIMER database, the mRNA level of KDM1A is positively correlated with expression level of PIK3CA and PIK3CB, respectively. F Heat map of KDM1A in different groups. G Venn diagram displaying the overlaps of transcription factors from DEGs among different groups. H Western blotting detected the effect of Ifebemtinib combined with paclitaxel on the LSD1/PI3K/Akt pathway.
Discussion
Breast cancer is the most common malignant tumor among women, with TNBC accounting for approximately 15% to 20%. Compared to other subtypes of breast cancer, TNBC is a highly heterogeneous disease characterized by poor differentiation, high invasion, and an earlier and more frequent occurrence of recurrence and metastasis. More than one-third of patients with TNBC will experience recurrence or distant metastasis. Recurrent or metastatic TNBC generally has a poor prognosis, with a five-year survival rate of less than 15%, significantly lower than the overall five-year survival rate for breast cancer patients (31%). Since TNBC does not express ER, PR, or HER2, commonly used targeted therapies are often ineffective. Therefore, in the latest NCCN guidelines, treatment plans for advanced TNBC still primarily include paclitaxel chemotherapy24. However, in clinical practice, we found that patient responses to paclitaxel are highly heterogeneous. Some patients show poor response to paclitaxel, exhibiting rapid disease progression shortly after first-line treatment, indicating the “primary resistance” of paclitaxel in TNBC. This makes it urgent to develop paclitaxel sensitizers and discover new strategies for treating TNBC.
As we know, mutations in the PI3K/AKT/mTOR pathway are particularly common in breast cancer, with approximately 70% of breast cancer patients exhibiting such mutations25,26. Inhibitors of this pathway are considered potential treatment strategies for breast cancer. For example, the selective PI3Kα inhibitor (such as Alpelisib) has been approved for use in combination with estrogen receptor antagonists for the treatment of advanced HR+ breast cancer27,28,29. However, inhibiting PI3K is insufficient to treat TNBC; combinations of PI3K inhibitors with chemotherapy and other treatments are being developed30. For example, Akt inhibitors combined with EZH2 inhibitors have been shown to effectively suppress TNBC31.
FAK is a non-receptor tyrosine kinase located in the cytoplasm, which is coupled with integrins and growth factor receptors to regulate cell adhesion, proliferation, migration, invasion, and metastasis. FAK is overexpressed and abnormally activated in various tumors, including TNBC. High expression of FAISL in tumor tissues of TNBC patients is associated with high expression of FAK and poor prognosis32. FAK signaling is an important regulatory factor for the colonization of breast cancer cells in the brain, while FAK inhibitors can effectively reduce the probability of brain metastasis by cancer cells8. Additionally, FAK inhibitors can block EZH2-induced bone metastasis in breast cancer7. The FAK inhibitor Ifebemtinib (also known as BI 853520 or Ifebemtinib) has shown effects on inhibiting tumor growth and metastasis across various cancers and has received Fast Track designation from the U.S. Food and Drug Administration (FDA). The combination treatment of FAK inhibitor Ifebemtinib with ROS1 inhibitor crizotinib for TNBC15. The combination therapy of KRAS G12C inhibitors and FAK inhibitors Ifebemtinib can achieve a synergistic anti-cancer effect19. Furthermore, Ifebemtinib can enhance immunotherapy efficacy by promoting immunogenic cell death through doxorubicin liposomes12. A PROTAC drug targeting FAK based on Ifebemtinib has also been confirmed to inhibit the proliferation and migration of triple-negative breast cancer cells in vitro14. However, there have been no reports on whether Ifebemtinib can sensitize paclitaxel in TNBC. In this study, we first reveal the high expression of FAK in breast cancer and reveal the synergistic effects and potential mechanism of the FAK inhibitor Ifebemtinib combined with paclitaxel in the treatment of TNBC. This finding provides a new theoretical basis and potential therapeutic strategies for the treatment of TNBC.
In vitro, we observed that the combined application of Ifebemtinib and paclitaxel significantly synergized to inhibit the survival and colony formation of TNBC cells, directly reflecting a significant suppression of tumor cell proliferation ability by combination therapy. The combination of Ifebemtinib and paclitaxel induced cell cycle arrest by inhibiting CDK1 activation. They promoted apoptosis and autophagy induction by activating mitochondrial apoptotic pathways, suggesting that the combination provides an important avenue for the clearance of tumor cells. At the same time, they synergistically reversed EMT by up-regulating ELF3 to inhibit the migration of TNBC cells, suggesting that the capacity for further growth and spread of tumor cells is severely restricted. These data demonstrate that the combined use of both agents has powerful efficacy in suppressing malignant biological behavior in TNBC cells. These data aligned with previous research reports suggesting that combination therapies can enhance inhibitory effects on tumor cells, further confirming the feasibility and effectiveness of combinatorial treatment strategies in cancer therapy.
In vivo data further validated the findings from in vitro experiments. The combination of Ifebemtinib and paclitaxel can synergistically inhibit the proliferation and lung metastasis of breast cancer cells. This result demonstrated a dual inhibitory effect on tumor growth and metastasis, which has important guiding significance for clinical treatment. Inhibiting tumor cell proliferation can directly control the size of the tumor, while inhibiting lung metastasis can reduce the mortality rate associated with tumors and extend patient survival. As the lungs are a common site for breast cancer metastasis, inhibiting this spread is crucial for improving patient prognosis. However, it is worth noting that the main adverse reaction of paclitaxel is a decrease in white blood cells and neutrophils. And in our mouse experiment, we observed that the combination of Ifebemtinib and paclitaxel further reduced the number of neutrophils and white blood cells in mice. However, this phenomenon may require further experimental validation.
Mechanically, we found that the combination of Ifebemtinib and paclitaxel synergistically inhibits the phosphorylation activation of FAK. As a key regulator of cancer, the phosphorylation activation of FAK plays an important role in survival, proliferation, and migration of tumor cells. Inhibiting the phosphorylation activation of FAK can block a series of downstream pro-cancer signaling pathways, thereby suppressing malignant behaviors in tumor cells. On the other hand, we discovered for the first time that the combination inhibits LSD1 to upregulate PIK3IP1, which synergistically inactivates the PI3K/Akt pathway to suppress cell survival. The PI3K/Akt pathway plays a central role in tumor cell survival and anti-apoptosis processes and is commonly activated in TNBC. By regulating this pathway, combined treatment effectively inhibited survival, cell cycle progress and promoted apoptosis of TNBC cells. This revelation provides crucial clues for our understanding of the synergistic effect of Ifebemtinib and paclitaxel in treating TNBC.
In summary, in this study, we clarified that Ifebemtinib sensitizes paclitaxel to inhibit proliferation and metastasis of TNBC cells mainly by blocking the PI3K/Akt pathway through the LSD1/PIK3IP1 axis, which provides new evidence to develop new strategies of FAK inhibitors and paclitaxel for therapy of TNBC.
Materials and methods
TIMER database
The differential expression of PTK2 that encoding FAK between tumor and adjacent normal tissues for PTK2 across all TCGA tumors and the survival differences of breast invasion carcinoma patients were analyzed according to the Tumor IMmune Estimation Resource (TIMER) database (https://cistrome.shinyapps.io/timer/).
Human specimens of breast cancer patients
A total of 72 cases of breast cancer tissue specimens and their paired adjacent normal breast tissue specimens were obtained from the Affiliated Cancer Hospital of Zhengzhou University & Henan Cancer Hospital. This study was conducted in accordance with the criteria of the 2013 Declaration of Helsinki. This study was approved by the Medical Ethics Committee of Henan Cancer Hospital (Approval No. 2022-KY-0001-002) and informed consent was obtained from all subjects or their legal guardians.
Human breast cancer cell lines and cell culture
Human TNBC cell lines MDA-MB-231 and MDA-MB-436 as well as mouse mammary cancer cell 4T1 were purchased from Procell (Wuhan, China) and were verified by STR analysis. The MDA-MB-231 cells were cultured in DMEM medium (Gibco™, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum (Gibco™, Thermo Fisher Scientific, Waltham, MA, USA) 100 units/mL penicillin, and 100 µg/mL streptomycin (Solarbio, Beijing, China). The MDA-MB-436 cell lines were cultured in DMEM medium with 15% FBS, 10 µg/ml Insulin (Procell, Wuhan, China), 100 units/mL penicillin, and 100 µg/mL streptomycin. Both MDA-MB-231 and MDA-MB-436 cells were cultured in a humidified incubator at 37 °C with 5% CO2.
Drugs and antibodies
Ifebemtinib was provided by InxMed (Nanjing, China). Paclitaxel Injection was purchased from Hainan Quanxing Pharmaceutical Co.Ltd (Haikou, China).
CCK-8 assay
Cell viability was assessed using a CCK-8 Kit (ZETA LIFE Inc, Menlo Park, CA, USA) according to the manufacturer’s protocol. Briefly, 2000–2500 TNBC cells were inoculated into 96-well plates for each well. The next day, the cells were treated with a series of concentrations of paclitaxel for 24–72 h. At the end of the drug treatment, 10 µL of CCK-8 solution was added to each well for another 2 h-incubation. Then the absorbance at 450 nm was detected. Graphpad Prism 10 software was used to plot cell survival curves, and SPSS 20.0 software was used to calculate the IC50 values of PXT and Ifebemtinib. Drug interactions were analyzed using the median effect principle (Talalay-Chou method). The combination index (CI) was calculated using the CompuSyn software, where a CI = 1 was indicative of an additive effect, a CI < 1 was indicative of a synergistic effect, and a CI > 1 was indicated of an antagonistic effect33.
Clone-Formation assay
Cell proliferation ability analyzed by clone-formation assay. Briefly, on the first day, MDA-MB-231 and MDA-MB-436 cells were seeded at a density of 500 cells per well in the six-well plate. On the second day, the cells were treated with paclitaxel, Ifebemtinib, and paclitaxel plus Ifebemtinib for 24 h respectively. After drug treatment, the cells were cultured for an additional 7 days. Subsequently, the cells were fixed with a 4% formaldehyde solution and stained using crystal violet. The number of cell clones was calculated using Image J software.
Carboxyfluorescein diacetate, succinimidyl ester (CFDS, SE) assay
The cell proliferation ability was evaluated by the CFDASE staining assay. MDA-MB-231 and MDA-MB-436 cells were seeded in a 6-well plate, cultured overnight, and then treated with drugs for 24 h. Subsequently, after removing the culture medium, washing twice with PBS, 2 ml of CDFA SE (5 µM, Yeasen Biotechnology Co., LTD, Shanghai, Chnia) staining solution was added to each well and incubated at 37℃ in dark for 30 min. After washing the cells with PBS, flow cytometry was used to detect the signal of carboxyfluorescein succinimidyl ester (CFSE), which is a hydrolysis product of CFDA SE.
Cell cycle distribution analysis
Cell cycle distribution was detected by flow cytometry (ACEA Biosciences, Agilent Technologies, Santa Clara, CA, USA) using the PI/RNase Staining Buffer (#550825, BD Pharmingen, San Diego, CA, USA). TNBC cells were treated with Ifebemtinib, PTX or their combination for 24 h and were fixed and permeabilized by ethyl alcohol (70%, v/v) at -20℃ for one week. After that, 0.5 mL PI/RNase staining buffer were added to each sample, incubating for 15 min at room temperature before analysis by flow cytometry.
Migration assay
Migration of TNBC cells was measured by wound healing assay. One ibidi Culture-Insert was put in each well of the 12-well plate, and 30,000–35,000 cells in 70 µL medium were seeded in each room of the culture-Insert. After the cell attachment, remove the Culture-Insert and add 1 ml medium containing 2% FBS and the drug Ifebemtinib, paclitaxel or the combination. 24 h later, the medium containing drugs was replaced with fresh medium containing 2% FBS. For MDA-MB-231 cells, microscopy images of the cell-free gap were captured at 0 h and 24 h and for MDA-MB-436 cells, the microscopy images were captured at 0 h and 48 h.
Apoptosis assays
Cell apoptosis was detected by flow cytometry using the FITC Annexin V Apoptosis Detection Kit I (#556547, BD Pharmingen, San Diego, CA, USA). According to the manufactory’s instruction, following different treatments, cells were collected after being digested by trypsin without EDTA and then stained with 100 µL FITC Annexin V and PI staining solution, followed by a 15-min incubation in dark on ice. Before detection, the sample was added another 400 µL 1 × binding buffer and placed on ice. The fluorescence signal of FITC Annexin V and PI was detected by flow cytometry within 30 min.
Western blotting
MDA-MB-231 and MDA-MB-436 cells were treated with Ifebemtinib, paclitaxel or their combination for 24 h. Then, the cells were rinsed by cold PBS for twice before lysing by RIPA lysis buffer on ice for 20 min. After centrifugation, the cleared cell lysates were collected. Bicinchoninic Acid (BCA) assay was used for protein quantitation. Then, 6× loading buffer was added to the samples before heated at 95 °C for 5min34. The protein samples were loaded onto a 10% SDS-polyacrylamide gel and subsequently transferred to PVDF membranes. To detect proteins of different molecular weights and to reduce antibody consumption, the membranes were cut to an appropriate size before antibody hybridization. For further analysis, the membranes were incubated with the primary antibodies: p-FAK overnight at 4 °C. The cell membranes were incubated with Goat anti-Mouse IgG (H + L) Secondary Antibody (#C31430100, Thermo Fisher Scientific, Waltham, MA, USA) or Goat anti-Rabbit IgG (H + L) Secondary Antibody (#C31460100, Thermo Fisher scientific) secondary antibodies conjugated with horseradish peroxidase (HRP) for 1 h at room temperature. Finally, enhanced chemiluminescence was used to detect the protein-antibody complex. β-Actin was used as the loading control. Relative protein levels were semi-quantified by Image J software35. The immunoreactive bands were detected using an enhanced chemiluminescence (Pierce ECL Western, Thermo Fisher scientific) system and exposed to medical X-ray film for visualization. The full panel of western gels for Figs. 4, 5, 6 and 8 are available as Supplementary Material 1.
Cell-derived xenograft (CDX) studies
Twenty Female BALB/c nude mice aged 4–6 weeks were purchased from Hangzhou Ziyuan Laboratory Animal Technology Co., Ltd. The mice were housed in an SPF-grade animal facility in individually ventilated cages, with free access to food and water. After one week of acclimatization, experiments were initiated. Generally, mice were subcutaneously inoculated with 7 × 106 MDA-MB-231 cells per animal. When the average volume of tumors reached 100 to 150 mm3, five animals were allocated to each group according to the tumor volume: vehicle control, Ifebemtinib, paclitaxel, and combination group. The vehicle control group received saline once a day. The mice in Ifebemtinib group were administrated 10 mg/kg/d of Ifebemtinib (Intragastric injection). The mice in paclitaxel group were administrated 5 mg/kg/3d of paclitaxel (Intraperitoneal injection). The mice in the combination group were administrated 10 mg/kg/d of Ifebemtinib and 5 mg/kg/3d of paclitaxel. The treatment was administered until the tumor volume in the mice reached 2000 mm³. At the end of the treatment, peripheral blood was collected from the mice by enucleation of the eyeball in mice, followed by euthanasia via cervical dislocation. After excision, each mouse tumor was bisected; one half was snap-frozen at − 80 °C, and the other half was fixed in 4% formaldehyde and embedded in paraffin within 48 h for further analysis.
Orthotopic mouse lung metastasis model
Twenty female BALB/c mice (aged seven-weeks) were purchased from (Hangzhou Ziyuan Laboratory Animal Technology Co., Ltd., Hangzhou, China). The mice were housed in an SPF-grade animal facility in individually ventilated cages, with free access to food and water. After one week of acclimatization, a total of 2 × 105 mouse breast cancer 4T1 cells were injected into the mammary pads of After three weeks post-injection, twenty mice were randomly divided into 4 groups: the vehicle, Ifebemtinib (10 mg/kg/d, intragastric injection), paclitaxel (5 mg/kg/3d, intraperitoneal injection), and combination. After 3 weeks, the mice were euthanized by cervical dislocation. The lungs of mice were collected, and metastatic nodules were counted. Significance was determined at P < 0.05 using Tukey’s multiple comparisons test.
RNA-seq
APTBI (Shanghai, China) performed the following RNA-seq analyses. Genes with adjusted P-value < 0.05 and absolute log2 fold changes > 1 were identified as differentially expressed. First, total RNA was isolated from MDA-MB-231 cells treated with Ifebemtinib, paclitaxel, or a combination using Trizol (Thermo Fisher scientific). The RNA was then purified and subjected to library preparation and sequencing on the Illumina Hiseq platform. Differential expression analysis of the resulting data from the three datasets was conducted using the DESeq2 R software package (version 1.20.0). Genes were considered significantly differentially expressed if their p-values were less than 0.05. Further analyses of these differentially expressed genes (DEGs) involved gene ontology (GO), and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses performed with the cluster Profiler R software package.
Hematoxylin and Eosin (HE) staining
The sections were deparaffinized, stained with hematoxylin for 5 min, rinsed, and differentiated using 1% hydrochloric acid in alcohol. They were then incubated in Eosin solution before being rinsed and paraffinized. Finally, the slides were mounted in neutral balsam.
Immunohistochemistry
Immunohistochemistry was performed on formalin-fixed and paraffin-embedded tissues following a previously described protocol34. The tissue sections were de-paraffinized, rehydrated, and subjected to antigen retrieval in boiling citric acid solution for 5 min followed by low-temperature microwave heating for 20 min. Subsequently, the sections were treated with H2O2 (3%), washed with PBS three times, and blocked with 10% sheep serum at room temperature for 30 min. The primary antibody was applied and incubated overnight at 4 °C. This was followed by incubation with a biotin-labeled secondary antibody for 30 min. DAB dye was then added dropwise to visualize color development under a microscope. Staining was stopped when the desired color intensity was achieved by rinsing in water, followed by counterstaining with hematoxylin. The expression of FAK was analyzed based on the following criteria: (1) score of immunostaining intensity: 0 (None), 1 (Weak), 2 (Moderate), 3 (Strong). (2) score of immunostaining positive rate: 0 (None), 1 (below 25%), 2 (25%-49%), 3 (50%-74%), 4 (above 75%). The expression score = (1) × (2). The difference was analyzed by the paired t-test.
TdT-mediated dUTP Nick end labeling (TUNEL) assay
In the tumor tissue, cell apoptosis is detected using the TUNEL assay. Paraffin sections of tumor tissues were deparaffinized using routine methods, and then baked at 60 °C for 60 min. After washing with PBS, the Proteinase K working solution (20 µg/ml) was added and incubated at 37 °C for 30 min followed by three washes with PBS. Subsequently, each section was treated with 50 µl of TUNEL working solution (containing TdT and FITC-labeled dUTP), incubated at 37 °C for 60 min, and washed three times with PBS. Finally, after adding 5 µl of antifade mounting medium (Beyotime, Shanghai, China), cover slips were used to mount the slides. Green fluorescence signals were observed under a microscope.
Blood biochemical assay and routine analysis
The safety of treatments is primarily assessed through biochemistry and routine analysis of blood. The complete blood of animals was collected in the tubes containing anticoagulant EDTA-2 K. Routine analysis of blood was performed using animal blood.
analyzer (HEMAVET 950FS, Drew Scientific, TX, USA). Following centrifugation for 1500 rpm, 10 min at room temperature, the serum was obtained. Blood biochemistry is tested using blood biochemistry equipment.
Statistical analysis
This study was analyzed using GraphPad Prism 8. ANOVA compared the four groups, while the Student’s t-test compared the two groups. Statistical significance levels were defined as follows: ns (not significant) for P > 0.05, * for P < 0.05, ** for P < 0.01, and *** for P < 0.001.
Data availability
The datasets analyzed during the current study are available in the TIMER2 database (https://cistrome.shinyapps.io/timer/). The data generated during and/or analyzed during the current study are available from the corresponding author.
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 74, 229–263 (2024).
Foulkes, W. D., Smith, I. E. & Reis-Filho, J. S. Triple-negative breast cancer. N Engl. J. Med. 363, 1938–1948 (2010).
Lin, N. U. et al. Clinicopathologic features, patterns of recurrence, and survival among women with triple-negative breast cancer in the National comprehensive cancer network. Cancer 118, 5463–5472 (2012).
Hu, H. H. et al. Roles and inhibitors of FAK in cancer: current advances and future directions. Front. Pharmacol. 15, 1274209 (2024).
Wu, X. et al. Leucine rich repeat containing 15 promotes triple-negative breast cancer proliferation and invasion via the ITGB1/FAK/PI3K signalling pathway. Sci. Rep. 15, 14535 (2025).
Dawson, J. C., Serrels, A., Stupack, D. G., Schlaepfer, D. D. & Frame, M. C. Targeting FAK in anticancer combination therapies. Nat. Rev. Cancer. 21, 313–324 (2021).
Lorusso, G. et al. Connexins orchestrate progression of breast cancer metastasis to the brain by promoting FAK activation. Sci. Transl Med. 14, eaax8933 (2022).
Zhang, L. et al. EZH2 engages TGFbeta signaling to promote breast cancer bone metastasis via integrin beta1-FAK activation. Nat. Commun. 13, 2543 (2022).
Pang, X. J. et al. Drug discovery targeting focal adhesion kinase (FAK) as a promising cancer therapy. Molecules 26, 4250 (2021).
Hirt, U. A. et al. Efficacy of the highly selective focal adhesion kinase inhibitor BI 853520 in adenocarcinoma xenograft models is linked to a mesenchymal tumor phenotype. Oncogenesis 7, 21 (2018).
Li, H., Gao, Y. & Ren, C. Focal adhesion kinase inhibitor BI 853520 inhibits cell proliferation, migration and EMT process through PI3K/AKT/mTOR signaling pathway in ovarian cancer. Discov Oncol. 12, 29 (2021).
Zhang, B. et al. Targeting of focal adhesion kinase enhances the Immunogenic cell death of pegylated liposome doxorubicin to optimize therapeutic responses of immune checkpoint Blockade. J. Exp. Clin. Cancer Res. 43, 51 (2024).
Osipov, A. et al. Inhibition of focal adhesion kinase enhances antitumor response of radiation therapy in pancreatic cancer through CD8 + T cells. Cancer Biol. Med. 18, 206–214 (2021).
Xu, M. S. et al. A novel FAK-degrading PROTAC molecule exhibited both anti-tumor activities and efficient MDR reversal effects. Acta Pharmacol. Sin. 45, 2174–2185 (2024).
Tan, X. et al. Simultaneous Inhibition of FAK and ROS1 synergistically repressed triple-negative breast cancer by upregulating p53 signalling. Biomark. Res. 12, 13 (2024).
Zhang, B. et al. Targeting FAK improves the tumor uptake of antibody-drug conjugates to strengthen the anti-cancer responses. iScience 28, 111536 (2025).
de Jonge, M. J. A. et al. Phase I study of BI 853520, an inhibitor of focal adhesion Kinase, in patients with advanced or metastatic nonhematologic malignancies. Target. Oncol. 14, 43–55 (2019).
Doi, T. et al. Phase I study of the focal adhesion kinase inhibitor BI 853520 in Japanese and Taiwanese patients with advanced or metastatic solid tumors. Target. Oncol. 14, 57–65 (2019).
Zhang, B. et al. Focal adhesion kinase (FAK) Inhibition synergizes with KRAS G12C inhibitors in treating cancer through the regulation of the FAK-YAP signaling. Adv. Sci. (Weinh). 8, e2100250 (2021).
Peyvandi, S. et al. Tumor-educated Gr1 + CD11b + cells drive breast cancer metastasis via OSM/IL-6/JAK-induced cancer cell plasticity. J Clin. Invest 134, e166847 (2024).
Lee, K., Kitagawa, M., Liao, P. J., Virshup, D. M. & Lee, S. H. A Ras-LSD1 axis activates PI3K signaling through PIK3IP1 suppression. Oncogenesis 9, 2 (2020).
Jiang, T. et al. Targeting LncRNA DDIT4-AS1 sensitizes triple negative breast cancer to chemotherapy via suppressing of autophagy. Adv. Sci. (Weinh). 10, e2207257 (2023).
Lin, J. et al. Switch of ELF3 and ATF4 transcriptional axis programs the amino acid insufficiency-linked epithelial-to-mesenchymal transition. Mol. Ther. 32, 1956–1969 (2024).
NCCN Clinical Practice. Guidelines in Oncology (NCCN Guidelines®), Breast Cancer, Version 2.2024, March 11, (2024).
Hinz, N. & Jucker, M. Distinct functions of AKT isoforms in breast cancer: a comprehensive review. Cell. Commun. Signal. 17, 154 (2019).
Lopez-Knowles, E. et al. PI3K pathway activation in breast cancer is associated with the basal-like phenotype and cancer-specific mortality. Int. J. Cancer. 126, 1121–1131 (2010).
Nik-Zainal, S. et al. Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature 534, 47–54 (2016).
Kim, M. et al. A protein interaction landscape of breast cancer. Science 374, eabf3066 (2021).
Stephens, P. J. et al. The landscape of cancer genes and mutational processes in breast cancer. Nature 486, 400–404 (2012).
Garrido-Castro, A. C. et al. Phase 2 study of buparlisib (BKM120), a pan-class I PI3K inhibitor, in patients with metastatic triple-negative breast cancer. Breast Cancer Res. 22, 120 (2020).
Schade, A. E. et al. AKT and EZH2 inhibitors kill TNBCs by hijacking mechanisms of Involution. Nature 635, 755–763 (2024).
Zhang, Y. et al. LncRNA FAISL inhibits Calpain 2-Mediated proteolysis of FAK to promote progression and metastasis of triple negative breast cancer. Adv. Sci. (Weinh). 11, e2407493 (2024).
Wang, S. Q. et al. Geridonin and Paclitaxel act synergistically to inhibit the proliferation of gastric cancer cells through ROS-mediated regulation of the PTEN/PI3K/Akt pathway. Oncotarget 7, 72990–73002 (2016).
Wang, S. Q. et al. Preclinical studies of the triazolo[1,5-a]pyrimidine derivative WS-716 as a highly potent, specific and orally active P-glycoprotein (P-gp) inhibitor. Acta Pharm. Sin B. 12, 3263–3280 (2022).
Wang, S. Q. et al. Steroidal dimer by001 inhibits proliferation and migration of esophageal cancer cells via multiple mechanisms. Cancer Chemother. Pharmacol. 83, 179–189 (2019).
Funding
This work was supported by the National Natural Science Foundation of China (No. 82573686 for X.C.), Henan Science and Technology Development Programs (No. 242102311123 for Q.M.), Henan Province Science and Technology Development Joint Fund Key Project (No. 235200810034 for X.C). Medical Science and Technique Foundation of Henan Province (No. SBGJ202301004 for X.C.), Natural Science Foundation of Henan Province (No. 232300421119 for X.C.), Henan Province Triple-100 Program: Overseas Fellowship for Medical Science and Technology Talents (No. HNMOT2024015 for S.W.), High-level Talents Internationalization Training Program of Henan Province (No.53 for S.W.), Young and Middle-aged Health Science and Technology Innovation Talent Training Program of Henan Province (No. YXKC2022048 for S.W.).
Author information
Authors and Affiliations
Contributions
Y.L. and X.C. conceptualized and designed the study. Y.L., Q.M., and S.Z. performed the study. Y.L. and W.Z. drafted the manuscript. L.L. and S.L. collected the data. Y. S., S.Z. and W.Z. analyzed the data. Y.L. and S.W. revised the manuscript. All authors reviewed the results and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Approval of animal experiments
The animal experiment was performed in accordance with guidelines of the Committee of Laboratory Animal Center of Zhengzhou University (Ethics approval license: ZZULAC2023072502). The results of animal experiments were reported according to the ARRIVE guidelines.
Approval for human experiments
A total of 72 cases of cancer tissue specimens and their paired adjacent normal breast tissue specimens from BC patients were obtained from the Affiliated Cancer Hospital of Zhengzhou University & Henan Cancer Hospital. This study was conducted in accordance with the criteria of the 2013 Declaration of Helsinki. This study was approved by the Medical Ethics Committee of Henan Cancer Hospital (Approval No. 2022-KY-0001-002) and informed consent was obtained from all subjects or their legal guardians.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, Y., Li, L., Mao, Q. et al. Ifebemtinib and paclitaxel synergistically inhibit the proliferation and metastasis of TNBC by blocking PI3K/Akt pathway through LSD1/PIK3IP1 axis. Sci Rep 15, 42672 (2025). https://doi.org/10.1038/s41598-025-26828-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-26828-x