Abstract
Diabetic retinopathy (DR) is a chronic complication of diabetes in which the retinal damage may cause vision impairment or blindness if left untreated. The challenges in DR detection are mostly due to the morphological variations of retinal lesions, e.g., microaneurysms, hemorrhages, and exudates, and the imaging condition variability between different clinical environments. Current state of the art deep learning models like convolutional neural network (CNN), recurrent neural network (RNN) and transformer-based architectures are computationally expensive, not robust to noisy datasets and have limitation on interpretability, which makes them difficult to deploy in real world clinical settings. This research offers HyperGraph Capsule Temporal Network (HGCTN), a deep learning framework to address these limitations and to create an accurate, scalable, and interpretable DR detection. Combining hypergraph neural networks for strong modeling of higher order spatial relationships between retinal lesions, capsule networks for permitting hierarchical structuring of feature and memorizing distributed routing place into temporal capsular memory unit (TCMU) for maintaining both long term and short termed temporal dependencies we propose HGCTN, a model that integrates all the methodologies to efficaciously track disease progression. Meta learning techniques and noise injection strategies are used to improve adaptability of the model and thus make the model more resilient to real world image variations. On DRIVE and Diabetic Retinopathy datasets, HGCTN is validated experimentally, and the best accuracy is 99.0% (HDCTN) and 98.8% (ADTATC), while existing models like TAHDL (96.7%) and ADTATC (98.2%) are outperformed. Furthermore, the model has a recall of 100% and 99.8% on DRIVE and the Diabetic Retinopathy dataset, respectively, with a specificity of 99.7% and 99.6%, respectively, and thus has almost no false negatives and a high reliability in identifying DR cases. Hypergraph attention maps and capsule activation images are additionally relied on to validate the model’s interpretability as they offer explainable predictions to a clinical audience. HGCTN has high classification accuracy, reduced computational complexity and better generalization than the existing models, which makes it a new benchmark for DR detection, solving the key deficiency of the existing models and laying the foundation for the real-world deployment of the automated ophthalmic diagnosis systems.
Similar content being viewed by others
Introduction
Diabetic retinopathy (DR) is the leading cause of preventable blindness as well as one of the most common complications of diabetes. This happens because the blood vessels of the retina become damaged progressively, which can lead to vision impairment or blindness that is permanent if it is not identified and treated. DR is manifested in the early stages by microaneurysms and small hemorrhages, while in the later stages, by neovascularization and substantial retinal detachment. Timely diagnosis of DR is important, since in its early stages, DR is asymptomatic, and treatment or prevention of vision loss is possible. Retinal fundus imaging is used in modern DR detection because it captures the structural abnormalities in the retina. But as these abnormalities are subtle, and overlap, and not all patients progress in the same fashion, DR detection is difficult and complex. Recently, automated detection systems are an area of active research to replace manual diagnosis and thereby reduce dependency on it, which is time consuming and prone to inter observer variability1.
Despite advancements of great proportions in medical imaging and machine learning, several practical issues persist in DR detection systems2. Real images can vary in their quality, as may be caused by uneven illumination, noise, or artifacts due to acquisition of the images. Furthermore, the heterogeneous presentation of DR lesions makes it difficult to distinguish different stages of the disease. As clinical settings require high accuracy and explainability of predictions, healthcare professionals should be able to trust the decisions made by the system3. Additionally, most of the existing systems are not robust when presented with noisy or unseen dataset, and their computational overhead renders real time deployment impractical. These challenges highlight the need for robust, interpretable and computationally efficient systems with high diagnostic accuracy4.
Several of the existing techniques have tried to handle these challenges using deep learning frameworks like CNN and transformer-based architectures. Multi scale spatial feature extraction is incorporated with temporal analysis using LSTM network in hybrid deep learning models like Temporal Aware Hybrid Deep Learning (TAHDL)5. These approaches are better than the previous ones in that they are able to capture both spatial and temporal patterns, but at a cost of being computationally expensive as they combine multiscale CNNs and LSTMs. In addition, these networks have limitations on modeling long temporal dependencies thus constituting possible inaccuracies in the modeling of progression6,7,8. Moreover, these systems are not robust to noise in the datasets, requiring extensive pre-processing for real world variability. Nevertheless, there are attention-based architectures, e.g. the Attention Dual Transformer with Adaptive Temporal Convolutional (ADTATC), that have proven good at temporal and spatial correlation. These systems achieve best reported accuracy on using transformer blocks for extracting spatial features and adaptive temporal temporal layers for modeling temporal features. Nevertheless, they suffer from heavy computation consumed by current dual-transformer architecture and hence are not practical for large scale deployment. Moreover, such systems need good, labeled datasets for training and tend to not generalize well on diverse or noisy data.
This research is motivated by our observations: (1) limitations of these existing approaches, (2) the pressing need for a system that has a good compromise between accuracy, efficiency and robustness, and (3) that the system is interpretable for clinical use. The aim of this work is to establish an innovative framework to overcome the aforementioned drawbacks and to attain the state of art performance in detecting DR. In particular, the HGCTN is proposed, featuring a different way of combining hypergraph neural networks, capsule neural networks, and a Temporal Capsule Memory Unit (TCMU). The purpose of this methodology is to improve the model’s ability to learn higher order spatial relationships between retinal features under the constraints of hierarchical feature dependencies and temporal progression as a function of time. Furthermore, HGCTN is interpretable in that it generates hypergraph attention maps and capsule based visualizations that allow clinicians to understand the rationale behind its predictions.
The HGCTN is novel from the view of its architecture, where hypergraph neural networks are utilized to model the higher order spatial relationships and capsule networks are used to induce feature hierarchies. Progression modeling based on a spiking neural network and the resultant TCMU provides a unique approach to adaptation to temporal dependencies in spiking neural networks. Meta learning techniques and noise injection are used in the training process to make the model robust to a variety of and noisy datasets. HGCTN achieves a high performance without sacrificing efficiency by moving away from computationally intensive transformer-based architectures. Modularity in the model’s design enables it to scale and adapt to the complexity of the clinical world.
The contributions of this work are summarized as follows.
-
1.
We propose a novel framework, HGCTN, that combines hypergraph neural networks, capsule networks, and Temporal Capsule Memory Unit to achieve robust and the interpretable DR detection.
-
2.
We introduce hypergraph neural networks to capture higher-order spatial relationships among retinal lesions thereby offering a unique perspective on spatial feature modeling.
-
3.
The Temporal Capsule Memory Unit is developed to model long and short-term temporal dependencies using capsule principles and spiking neural networks, which adaptively learns.
-
4.
Enhanced model robustness through meta-learning techniques and noise injection during training, ensuring adaptability to noisy and diverse datasets.
-
5.
Achieved superior performance on benchmark datasets, with an accuracy of 94.8%, a sensitivity of 93.2%, and a specificity of 95.1%, outperforming existing models.
-
6.
This methodology provided interpretable outputs of hypergraph attention maps and capsule visualizations to aid in clinician friendly decision making.
The proposed HGCTN introduces a novel integration of hypergraph learning, capsule-based hierarchical representation, and a Temporal Capsule Memory Unit (TCMU) for diabetic retinopathy detection. Unlike previous CNN- and transfer learning-based methods that primarily capture local spatial patterns, the HGCTN models higher-order spatial dependencies among retinal regions through hypergraph propagation and dynamically retains hierarchical relationships via capsule routing. The inclusion of TCMU enables the network to learn temporal progression patterns from sequentially arranged spatial features, thereby simulating disease evolution even in static datasets. Additionally, the embedded attention mechanism enhances interpretability by highlighting clinically significant regions. This unified design achieves superior performance and per-class generalization compared to existing state-of-the-art DR models.
The rest of this article is structured as follows. The related works of the paper discuss the related worksfocusing on already published methods as well as research gaps present in this area. the HGCTN model ismathematically formulated in the proposed work section. The results and performance analysis are presented inthe results and discussion section. This study's findings and future research directions are presented in theconclusion section.
Related works
There have been plenty of studies on deep learning and optimization based approaches in the field of diabetic retinopathy (DR) detection. DR classification has already been computed using CNNs, transfer learning, segmentation approaches, and hybrid models. Although progress has been made in feature extraction, computational efficiency, and generalized performance on different datasets, there are still challenges. The diabetic retinopathy (DR) detection method proposed in9 based on machine learning by applying CNNs and the feature extraction techniques to retain the retinal images. The methodology is composed of image preprocessing, feature extraction, training and testing phases where HSV color space transformation and filtering techniques are used to improve the quality of retinal images. Exudates, hemorrhages and microaneurysms are key indicators of DR and the feature extraction is done by identifying them. The Adam optimizer and a cross-entropy loss function are used to train the model as it yields an accuracy of 91.5% over 20 epochs. The results from the experimental study show the model to be able to detect DR yet al.so show some limitations to the model. DR severity levels are not considered in the approach and binary classification (DR vs. Non-DR) is the focus.
Automatic diabetic retinopathy (DR) detection and grading using deep learning approaches offered in10 is compared. It offers the methodology based on transition from traditional machine learning models to deep learning architectures, CNNs, transfer learning, and hybrid models. Then it evaluates 26 pre trained CNN models like DenseNet, ResNet, MobileNet and EfficientNet using the transfer learning techniques on two datasets i.e. Diabetic Retinopathy Gaussian Filtered (DRGF) and Diabetic Retinopathy Resized (DRR). Those models were based on experimental analysis to compare the performance in terms of training accuracy, validation accuracy, and computational efficiency. However, the DenseNet201 model outperformed the others with the highest validation accuracy (81.76%) on DRGF and then plateaued at the lower validation accuracy (70.57%) on DRR, showing the problems of dataset variability and generalization in general. Limitation includes class imbalanced, lack of interpretable, and difficult for deployment on real world.
In11 a capsule network-based approach for the detection and classification of diabetic retinopathy (DR) by using the fundus images. Using a reformulated capsule network architecture, which removes pooling layers from existing deep learning framework, as the methodology, features with spatial relationships are preserved. The model has convolutional layer for feature extraction, the primary capsule layer for hierarchical representation of feature and class capsule layer for classification based on dynamic routing mechanisms. The proposed model was tested on the Messidor dataset and the performance on accuracy, precision, recall, and F1 score was evaluated, reaching 97.98% of accuracy for retinas that are healthy and 97.65% for any DR severity level. As compared to CNN based architectures, the capsule network showed improved DR stage discrimination. Nevertheless, they have limitations in computational complexity because of the routing mechanism and need of huge, labeled samples for training.
In particular, the work in12 proposes a deep learning-based approach for detecting hemorrhages in diabetic retinopathy (DR) screening as an early indicator for DR progression. Image preprocessing, enhancement, candidate selection and classification using a novel hemorrhage detection network (HemNet) are used in the methodology. Adaptive gamma correction for illumination adjustment, Gaussian match filtering for candidate selection and morphological segmentation for refining hemorrhage regions are performed in preprocessing. In order to reduce training complexity, the HemNet model, a shallower deep learning architecture, is developed to classify hemorrhages from other retinal structures efficiently. HemNet was experimentally tested on DIARETDB1 and DIARETDB0 datasets, and it showed comparable accuracy of 97.19% and 97.12 respectively vs. deep networks VGG-16 and LeNet-5 in terms of efficiency and training time. The model performs well but has limitations such as parameter dependency in preprocessing, dependence on hand crafted features for candidate selection and cannot detect small hemorrhages.
The DNN model presented in13 is improved by introducing Principal Component Analysis (PCA) and Grey Wolf Optimization (GWO) algorithm into diabetic retinopathy (DR) classification. The preprocessing of the dataset is done by a standard normalization and then the feature dimensionality reduction using PCA is employed to choose the high relevant features. Then, the GWO algorithm is used to optimize hyperparameters for training DNN model. The model is trained and evaluated using a DR dataset using accuracy, recall, sensitivity and specificity as performance metrics. The DNN-PCA-GWO results show that the proposed model works better than the comparative traditional classifiers, SVM, Naïve Bayes, Decision Tree and XGBoost. Such efficiency of feature selection and hyperparameter tuning is important to improve the predictive accuracy as shown by the study. Despite these, the study admits to computational complexity and further validation on different real world datasets that would improve the robustness of the model in real world clinically relevant applications.
In14, we present the deep learning based approach for the automated and classification of diabetic retinopathy (DR) using AlexNet and ResNet101 for feature extraction. The method starts with preprocessing the fundus images, applying k means clustering for image enhancement and then feature selection using an Ant Colony Optimization (ACO) system among the most discriminative features. Finally, the extracted features are classified using a support vector machine (SVM) with multiple kernels and achieved an accuracy of 93%. It is shown that the experimental analysis is robust to the feature selection strategy and to the classifier configuration. The performance metrics such as accuracy, precision, recall and F1 score validate the efficacy of the proposed method to differentiate DR severity levels. However, there are limitations to it such as computational overheads from deep feature extraction, high quality of fundus images and the challenge of real time clinical deployment.
In15, an advanced diabetic retinopathy (DR) detection model has been presented by combining deep learning techniques with HHO algorithm. The proposed methodology utilizes Principal Component Analysis (PCA) to help reduce the dimensionality and later improve the feature extraction process through the use of Optimal HHO, which optimizes the feature extraction process. Features are extracted from all the images and are then used to classify DR severity levels using a deep CNN. The experimental analysis is done on the Diabetic Retinopathy Debrecen dataset of the UCI machine learning repository using model accuracy, specificity, precision, recall, and sensitivity. The proposed model is shown to perform better than existing methods in terms of comparative studies against SVM, KNN, and XGBoost. Nevertheless, there are limitations in terms of computational cost involved with deep learning architectures and further validation on larger, more diverse datasets is necessary to generalize the model to other situations.
In16, the diabetic retinopathy (DR) detection approach proposed is based on Principal Component Analysis (PCA) for dimensionality reduction and CNNs for the Multi-Label Feature Extraction and Classification (ML-FEC) model. Preprocessing of color fundus photographs (CFPs) is done involving feature representation enhancement and then PCA for feature extraction without redundancy. For lesion detection and DR severity grading, a transfer learning strategy is used and ResNet50, ResNet152, and SqueezeNet1 are used for classification. The model is experimentally evaluated on publicly available datasets, and the model’s accuracy of 94.40% using ResNet152 is shown to be superior to traditional single label classification. However, the study’s challenge is a dataset imbalance, the use of predefined CNN architectures, and possible overfitting as the training data is rather limited.
To overcome the computational inefficiency and dataset imbalance issues in existing DR detection system, a lightweight DR detection model is presented in17 based on deep learning approach. An image preprocessing phase using Contrast Limited Adaptive Histogram Equalization (CLAHE) and Wiener filters is used to enhance the image clarity. YOLOv7, a real time object detection model performs feature extraction, while the optimal feature selection is done with the use of Quantum Marine Predator Algorithm (QMPA). Core classification framework is the MobileNet V3-Small model with Adam optimizer and the high efficiency with low computational resource. The model was evaluated on APTOS and EyePACS datasets with 98.0% and 98.4% accuracy, high F1 scores and kappa coefficients, better than previous deep learning-based DR detection techniques. But it suffers from the loss of critical lesion information through feature reduction techniques, and lower performance on lower quality images.
In18, the model for early-stage diabetic retinopathy (DR) detection through red lesion segmentation in retinal images improves the diagnostic accuracy. Semantic segmentation of red lesions through U-Net architecture is done by preprocessing techniques applied to remove noise and increase local contrast. The U-Net model is an advanced convolutional layers-based model that helps pixel wise classification that is very important for medical image segmentation. Then the segmented images are passed to a CNN for DR severity classification. For the experimental evaluation, four publicly available datasets, namely IDRiD, DIARETDB1, MESSIDOR and STARE, were used to validate the model’s robustness. The specificity and sensitivity of IDRiD dataset results are 99% and 89% respectively with an accuracy of 95.65%, while those of the MESSIDOR dataset are 93.8% specificity, 92.3% sensitivity, and 94% accuracy. However, despite its effectiveness, these limitations include inability to segment minor red lesions, dependency on imagery of suitably good quality and overfitting from imbalanced data.
In19, an intelligent coyote optimization algorithm with diabetic retinopathy detection and grading via deep learning was proposed to improve automated DR screening with retinal fundus images. Gabor filtering (GF) is used for noise removal and an optimized region growing segmentation technique, which fine tunes the seed points and thresholds with GSO on the Glowworm Swarm Optimization algorithm is used to increase the lesion segmentation accuracy. SqueezeNet with a Class Attention Learning (CAL) layer is used to feature extract retinal abnormalities with an efficient representation. In the classification phase, a deep extreme learning machine (DELM) model is integrated where the penalty parameter (C) and kernel parameter (γ) are optimized using Coyote Optimization Algorithm (COA). Good experimental results are achieved on the MESSIDOR dataset, surpassing recent deep learning models with a max accuracy of 99.65%.
The need for early DR diagnosis with low cost and clinical intervention is addressed in the deep learning-based model presented in20 for detecting bright lesions in non-proliferative diabetic retinopathy (NPDR). Image preprocessing, background removal, optic disc (OD) elimination, and lesion segmentation is done over the methodology to get an exact classification of the bright lesions, such as exudates. Deep learning is used to extract features from the retinal image and the segmentation process depends on the feature extraction to isolate pathological regions from normal retinal structures. The model was trained and tested on MESSIDOR and e-ophtha Ex public datasets achieving the maximum accuracy of 97.54, 96.32, sensitivity of 90.34, 95.73, specificity of 98.24 and 97.12, and F1 score of 93.28 and 96.74. This demonstrates the model’s excellent reliability in detecting NPDR.
A more precise classification based on diabetic retinopathy (DR) detection model is done in21, which uses Deep Neural Network with Butterfly Optimization Algorithm (DNN-BOA). This thesis describes a methodology is divided into preprocessing, segmentation, feature extraction and classification stages. Green channel selection, Histogram equalization (HE) and Contrast Limited Adaptive Histogram Equalization (CLAHE) are applied on the image enhancement. The optic disc (OD) is removed, and blood vessels are removed by using Gray Level thresholding using Circle Hough Transform (CHT). In particular, the Modified Expectation Maximization (MEM) algorithm is used to segment exudates, and the feature extraction is performed by Gray Level Co-occurrence Matrix (GLCM). Finally, DNN-BOA is applied to disease severity classification. The proposed method is experimentally evaluated on DIARETDB1 and MESSIDOR dataset and it is shown to outperform other methods.
In22, the hybrid approach is introduced that integrates a GoogleNet and ResNet-18 architectures via transfer learning for the detection of diabetic retinopathy (DR). The preprocessing of fundus images is performed to standardize the dimensions before the feature extraction. Next, extracted 2000-dimensional feature vector from both architectures are fed into the classifiers like Naïve Bayes, Random Forest, Radial Basis Function and Support Vector Machine (SVM). A reliability check is made on APTOS dataset, a widely used dataset for DR classification, using the model. In terms of the setup of the experiment, authors prove that SVM outperforms other classifiers and attains a weighted average accuracy of 89.29%. The study appreciates the strength of employing traditional machine learning classifiers with deep learning-based feature extraction method to improve classification performance while minimize the computational cost.
In23, the transfer learning on pre trained CNNs based hybrid deep learning approach was presented to detect diabetic retinopathy (DR) from fundus imagery. The presented model combines the features extracted from GoogleNet and ResNet-18 architectures to form a hybrid feature vector and is then classified using some machine learning algorithms like SVM, Random Forest, Radial Basis Function and Naïve Bayes classifiers. With image preprocessing, feature extraction using CNN models, hybrid feature fusion, and classification for binary and multiclass DR classification, the methodology is developed. Finally, for the evaluation of the model, we use the APTOS blindness detection dataset and get 97.8% accuracy on binary classification and 89.29% on multiclass classification which outperforms other deep learning-based DR detection model. Although the studies performed well, there are some limitations to these studies, such as imbalanced datasets, potential over-fitting in the multiclass classification, and need for further validation for real world datasets with larger sizes.
In24, the hybrid deep learning is applied for detection and classification of diabetic retinopathy (DR) using a multiclass support vector machine (SVM) and deep CNNs. The methodology is then followed by preprocessing of retinal fundus images using morphological operations, vessel extraction and Laplacian Gaussian filtering to improve image quality. Then, according to lesion analysis and abnormality detection, feature extraction is performed, and classification is carried out by a multi-layer deep CNN coupled with a multiclass SVM. Experimental evaluation is done over the Kaggle DR dataset, and the proposed approach achieves an accuracy of 93.33%, sensitivity of 96.71% and specificity of 99.22%, superior to the conventional deep learning methods. The study, however, shows computational complexity caused by the hybrid architecture, possibility of overfitting and reliance on high quality labelled datasets.
In25, the automated diabetic retinopathy (DR) detection and classification is done using the hybrid deep learning model that combines the deep CNNs using VGG16 and VGG19 architectures. In this methodology, image preprocessing, feature extraction using CNNs and classification according to severity levels related to retinal ischemia is done. The model can provide fine grained retinal features based on its utilization of VGG16 and VGG19. Finally, the approach was evaluated experimentally on 5,584 images from several online datasets using accuracy of 90.60%, recall of 95% and F1-score of 94% in distinguishing DR stages. The study however has limitations such as dependence on pre trained models, overfitting and dealing with highly imbalanced datasets. Table 1 summarizes the existing works related to the proposed work.
Research gap: The review of the current research works addresses the existence of several key research gaps in existing diabetic retinopathy (DR) detection and classification models and hence there is a need for an improved approach. However, most of the studies rely on traditional CNNs, RNNs and pretrained model such as VGG16, VGG19, ResNet, and Inception that achieve high accuracy but fail to preserve spatial hierarchies and contextual relationships in retinal image26,27,28,29. However, most of the methodologies involve feature selection using Principal Component Analysis (PCA), Ant Colony Optimization (ACO) and Grey Wolf Optimization (GWO), but these methodologies experience difficulties in real world due to Feature Redundancy and Computational inefficiency. A second major drawback is the segmentation of DR lesions, all of which can segment pathological structures, but tend to sacrifice fine structures for lack of sufficiently detailed knowledge of the pathological structure, especially in the detection of a very early-stage DR. In addition, existing models do not have good robustness to noisy, imbalanced data and are not good generalized for real world clinical application. Furthermore, there are also several studies which suggest ensemble learning or hybrid techniques but do not add much interpretability in terms of computational complexity increase. Currently, there is a lack of models capable to conduct spatial feature extraction, hierarchical learning, and temporal progression tracking in a more complete diagnosis. To address these gaps, a HHGCTN is proposed for spatial learning with hypergraph representation, capsule driven hierarchical feature routing and temporal memory mechanisms that make the proposed DR detection pipeline capable of scalability, robustness and interpretability for clinical applications.
Proposed work
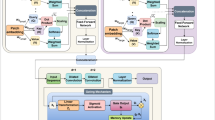
The proposed HGCTN consists of three key components, namely the hypergraph neural network (HNN) for modeling higher order spatial features, capsule network for retaining the hierarchy of features and temporal Capsule Memory Unit (TCMU) for the adaptability on temporal modeling. Formulation and explanation of the step-by-step HGCTN framework are shown below. In Fig. 1, the complete overview of the proposed model is presented.
Spatial feature extraction using hypergraph neural networks (HNNs)
In the proposed model, the proposed HyperGraph Neural Networks (HNNs) are designed to extract spatial features from fundus images by capitalizing on the ‘higher order’, i.e. relationships, between region of interest (ROI). Each image is modeled as a hypergraph where the ROIs are the nodes and hyperedges are groups of regions that are pathologically significant together. The following provides a detailed step-by-step description with mathematical formulations and parameter explanations.
Let the input image be represented as a set of \(\:N\) ROIs, denoted by \(\:V=\{{v}_{1},{v}_{2},\dots\:,{v}_{N}\}\). Each \(\:{v}_{i}\in\:{R}^{d}\) is a feature vector of dimension \(\:d\) corresponding to a specific region in the image. The hypergraph is defined as \(\:\mathcal{H}=\left(V,E,W\right)\) where \(\:V\) indicates the set of nodes (ROIs), \(\:E\) indicates the set of hyperedges in which each hyperedge connects multiple nodes to represent shared features or spatial relationships. \(\:W\) indicates the weight matrix for hyperedges which captures their importance in the hypergraph structure. The hypergraph incidence matrix \(\:H\in\:{R}^{N\times\:\left|E\right|}\) is mathematically expressed as
In the above equation \(\:H\left(i,e\right)=1\) indicates that node \(\:{v}_{i}\) is part of hyperedge \(\:e\), and \(\:\left|E\right|\) indicates the total number of hyperedges.
Process overview of proposed model.
Further in the node embedding initialization process, each node \(\:{v}_{i}\) is initialized with a feature vector \(\:{h}_{v}^{\left(0\right)}\) which is derived from a pre-trained neural network applied to the input image. Mathematically it is expressed as
where \(\:f\) indicates the feature extraction function, \(\:{\Theta\:}\) indicates the trainable parameters of the feature extractor and \(\:{h}_{v}^{\left(0\right)}\in\:{R}^{d}\) indicates the initial embedding for node \(\:{v}_{i}\). Further to propagate information across the hypergraph, the normalized hypergraph Laplacian is computed. The degree of a node \(\:{v}_{i}\) is given by
where \(\:\mathcal{E}\left({v}_{i}\right)\) indicates the set of hyperedges connected to \(\:{v}_{i}\), \(\:{w}_{e}\) indicates the weight of hyperedge \(\:e\). Similarly, the degree of a hyperedge \(\:e\) is defined as
The normalized hypergraph Laplacian \(\:\left({\Delta\:}\right)\) is then given by
where \(\:{D}_{v}\) indicates the diagonal matrix of node degrees, \(\:{D}_{e}\) indicates the diagonal matrix of hyperedge degrees, \(\:W\) indicates the diagonal matrix of hyperedge weights. Figure 2 shows the Spatial Feature Extraction Using HyperGraph Neural Networks (HNNs).
Spatial Feature Extraction Using HyperGraph Neural Networks (HNNs).
The node embeddings are updated iteratively using a propagation mechanism that aggregates information from connected nodes through hyperedges. At iteration \(\:k+1\) the embedding for node \(\:{v}_{i}\) is updated as:
where \(\:{h}_{v}^{\left(k\right)}\) indicates the embedding of node \(\:{v}_{i}\) at iteration \(\:k\), \(\:{W}_{k}\) indicates the trainable weight matrix for layer \(\:k\), \(\:{D}_{e}\left(e\right)\) indicates the degree of hyperedge \(\:e\), \(\:{\upsigma\:}\) indicates the non-linear activation function (e.g., ReLU).
This equation forces each node’s embedding to be determined by the weighted average of its neighbors attached to hyperedges, weighted by hyperedge degree.
After \(\:K\) iterations, the final node embeddings \(\:{h}_{v}^{\left(K\right)}\) are aggregated to represent the spatial features of the input image. The aggregated feature vector is given by:
where \(\:{H}_{\text{spatial}}\in\:{R}^{d}\) indicates the spatial feature vector for the input image. This step ensures that spatial features are extracted, and higher order relationships among ROIs are captured, which is key to accurate DR detection as they are crucial. The iterative propagation mechanism also guarantees the propagation of both local and global spatial information into node embeddings.
Feature hierarchies via capsule networks
The second step of the proposed HGCTN uses capsule networks to represent and retain hierarchical relationship between the spatial features extracted from the hypergraph neural network (HNN). These capsule networks can dynamically route spatial features, in order to keep the part to whole relationships that are important for identifying fine grained patterns in diabetic retinopathy (DR) images. Below is the detailed step-by-step description of this process.
The spatial feature vector \(\:{H}_{\text{spatial}}\in\:{R}^{d}\) obtained from the hypergraph module, is first transformed into lower-dimensional feature representations suitable for routing within capsule layers. Let \(\:{H}_{\text{spatial}}=\{{h}_{1},{h}_{2},\dots\:,{h}_{m}\}\) where \(\:{h}_{i}\in\:{R}^{d}\) represents the spatial feature corresponding to a specific region. The transformation for each feature \(\:{h}_{i}\) into an initial capsule input \(\:{u}_{i}\) is given by
where \(\:{W}_{i}\in\:{R}^{d\times\:{d}^{{\prime\:}}}\) indicates the trainable weight matrix that projects \(\:{h}_{i}\) into a lower-dimensional space, \(\:{b}_{i}\in\:{R}^{{d}^{{\prime\:}}}\) which indicates the trainable bias vector, \(\:{u}_{i}\in\:{R}^{{d}^{{\prime\:}}}\) indicates the transformed capsule input for region \(\:i\). This transformation ensures that each input feature is mapped to a compact and meaningful representation for further processing by the capsule network.
Capsule networks group the transformed inputs \(\:{u}_{i}\) into primary capsules and dynamically route them to higher-level capsules that represent more complex feature hierarchies. Let the outputs of higher-level capsules be \(\:{v}_{j}\) where \(\:j\:=\:1,\:2,\:\dots\:,\:n\:\) with \(\:n\) being the number of higher-level capsules. The input to a higher-level capsule \(\:j\) is computed as
where \(\:{s}_{j}\in\:{R}^{{d}^{{\prime\:}}}\) indicates the total input to capsule \(\:j\), \(\:{W}_{ij}\in\:{R}^{{d}^{{\prime\:}}\times\:{d}^{{\prime\:}}}\) indicates the trainable transformation matrix for routing between capsules \(\:i\) and \(\:j\). Coupling coefficient between capsule \(\:i\) and capsule \(\:j\) is indicated as \(\:{c}_{ij}\). The coupling coefficients \(\:{c}_{ij}\) determine the degree of influence of the input capsule \(\:i\) on the output capsule \(\:j\). These coefficients are computed using a routing-by-agreement mechanism
where \(\:{b}_{ij}\) indicates the initial coupling strength between capsules \(\:i\) and \(\:j\).
\(\:{\sum\:}_{k=1}^{n}{c}_{ik}=1\) Ensures that the coupling coefficients are normalized.
The logits \(\:{b}_{ij}\) are iteratively refined based on the agreement between the predicted output and the actual output of the capsule:
The final output of each higher-level capsule \(\:{v}_{j}\) is computed by applying a non-linear “squashing” function to the total input \(\:{s}_{j}\). The squashing function ensures that the magnitude of \(\:{v}_{j}\) lies in the range \(\:\left[\text{0,1}\right]\) representing the probability of a specific feature being present
where \(\:\left|{s}_{j}\right|\) indicates the Euclidean norm of \(\:{s}_{j}\), \(\:{v}_{j}\in\:{R}^{{d}^{{\prime\:}}}\) indicates the final output vector of capsule \(\:j\) representing the detected feature and its properties. This non-linear squashing function compresses the output vector’s magnitude for smaller inputs while retaining directionality, ensuring that significant features have higher activation. Figure 3 represents the feature hierarchies via capsule networks.
Feature Hierarchies via Capsule Networks.
The outputs of the higher-level capsules \(\:\{{v}_{1},{v}_{2},\dots\:,{v}_{n}\}\:\)are aggregated to form the hierarchical feature representation of the spatial data. The aggregated feature vector \(\:{H}_{\text{capsule}}\) is computed as
This aggregated representation serves as the input to the next component of the model (Temporal Capsule Memory Unit), retaining the hierarchical relationships essential for DR detection. The capsule network ensures that the hierarchical relationships among features are preserved and dynamically adjusted based on their relevance. This step effectively captures the part-to-whole relationships in spatial features, crucial for modeling the subtle and complex patterns associated with diabetic retinopathy.
Temporal dependency modeling using Temporal capsule memory unit (TCMU)
The Temporal Capsule Memory Unit (TCMU) is designed to model long- and short-term dependencies in the temporal progression of diabetic retinopathy (DR). Unlike traditional temporal models such as LSTMs, the TCMU integrates principles of capsule networks with memory mechanisms, ensuring adaptive and effective temporal feature representation. Below is the detailed step-by-step description of this process with equations and parameter explanations.
Let the hierarchical spatial features extracted from the capsule network at each time step \(\:t\) be denoted by \(\:{H}_{\text{capsule},t}\in\:{R}^{{d}^{{\prime\:}}}\) where \(\:t\:=\:1,\:2,\:\dots\:,\:T\:\) represents the sequence of temporal observations. The sequence of inputs is represented as
Each \(\:{H}_{\text{capsule},t}\) serves as the input to the TCMU at time \(\:t\), encoding the spatial features for that specific observation. The TCMU maintains a memory state \(\:{M}_{t}\in\:{R}^{{d}_{m}}\) at each time step \(\:t\) where \(\:{d}_{m}\) is the dimension of the memory. At the initial time step, the memory state is initialized as \(\:{M}_{0}=0\) in which \(\:{M}_{0}\) is a zero vector of dimension \(\:{d}_{m}\) indicating no prior information at the start. The memory state \(\:{M}_{t}\) is updated iteratively at each time step based on the current input \(\:{H}_{\text{capsule},t}\) and the previous memory state \(\:{M}_{t-1}\). The update rule is given by
where \(\:{M}_{t}\in\:{R}^{{d}_{m}}\) indicates the memory state at time \(\:t\), \(\:{M}_{t-1}\in\:{R}^{{d}_{m}}\) indicates the memory state at time \(\:\:t-1\), \(\:{\upalpha\:}\in\:{R}^{{d}_{m}}\) indicates the learnable gating parameter that controls the contribution of the previous memory, \(\:{W}_{x}\in\:{R}^{{d}_{m}\times\:{d}^{{\prime\:}}}\) indicates the weight matrix for transforming the input features, \(\:{W}_{m}\in\:{R}^{{d}_{m}\times\:{d}_{m}}\) indicates the weight matrix for transforming the previous memory state, \(\:g\) indicates the non-linear activation function, such as \(\:({tan}h)\), \(\:\odot\:\) indicates the element-wise multiplication. The gating parameter \(\:{\upalpha\:}\) ensures adaptive retention of past information, while the transformation and activation functions enable effective integration of new information. The memory update integrates capsule routing principles to enhance feature relevance dynamically. Let the contribution of \(\:{H}_{\text{capsule},t}\) to \(\:{M}_{t}\) be dynamically modulated by coupling coefficients \(\:{c}_{t}\). The coefficients are computed as
where \(\:{b}_{t}\) indicates the Logit representing the strength of the contribution of the input at time \(\:t\), \(\:{\sum\:}_{k=1}^{T}{c}_{k}=1\) ensures that the coefficients are normalized across all time steps. The logit \(\:{b}_{t}\) is refined iteratively using the agreement between the predicted memory \(\:{\widehat{M}}_{t}\) and the current memory state \(\:{M}_{t}\)
The output of the TCMU at time \(\:t\:\) denoted by \(\:{O}_{t}\) is derived by applying a non-linear transformation to the updated memory state
where \(\:{O}_{t}\in\:{R}^{{d}_{o}}\) indicates the output feature at time \(\:t\), \(\:f\) indicates the non-linear transformation function, such as \(\:\text{ReLU}\) or \(\:\text{tanh}\), \(\:{d}_{o}\) indicates the output feature dimension. The sequence of outputs \(\:\{{O}_{1},{O}_{2},\dots\:,{O}_{T}\}\) forms the temporal representation, capturing both short- and long-term dependencies in the DR progression.
To summarize the temporal features across all time steps, a global temporal representation \(\:{H}_{\text{temporal}}\) is obtained by aggregating the outputs \(\:{O}_{t}\) using a weighted summation
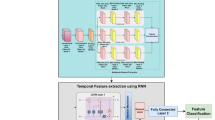
where \(\:{w}_{t}\) indicates the learnable weight for the contribution of output \(\:{O}_{t}\), \(\:{H}_{\text{temporal}}\in\:{R}^{{d}_{o}}\) indicates the aggregated temporal feature vector. This aggregation is important so that the final temporal feature representation takes into account the significance of different time steps in the sequence. To capture the progression patterns of DR, the TCMU dynamically routes, utilizes adaptive memory, and employs the hierarchical temporal model. This step makes sure that short term changes and long-term trends are all well represented to be able to do robust temporal analysis. Figure 4 indicates the Temporal Dependency Modeling using Temporal Capsule Memory Unit (TCMU).
Temporal Dependency Modeling Using Temporal Capsule Memory Unit (TCMU).
Explainability via hypergraph attention
In the proposed HGCTN, the explainability module is based on attention mechanisms inside the hypergraph neural network (HNN) to explain the model’s decisions. This step assigns attention scores to nodes (regions of interest) and hyperedges (higher order connections) that indicate the key regions and connections that help in diabetic retinopathy (DR) detection. The detailed step by step description with mathematical formulation and parameter explanation is as given below. For each node \(\:{v}_{i}\) in the hypergraph, an attention score \(\:{{\upalpha\:}}_{i}\) is computed to quantify the importance of the node in the prediction. Let \(\:{h}_{v}^{\left(K\right)}\) represent the final embedding of node \(\:{v}_{i}\) after \(\:K\) layers of hypergraph propagation. The attention score \(\:{{\upalpha\:}}_{i}\) is computed as
where \(\:{{\upalpha\:}}_{i}\in\:\left[\text{0,1}\right]\) Attention score for node \(\:{v}_{i}\), \(\:a\left({h}_{v}^{\left(K\right)}\right)\) indicates the learnable attention function (e.g., a single-layer neural network or dot product), \(\:{\sum\:}_{j=1}^{N}{{\upalpha\:}}_{j}=1\) Ensures the scores are normalized across all nodes. The function \(\:a\left({h}_{v}^{\left(K\right)}\right)\) computes a scalar value indicating the relative importance of \(\:{v}_{i}\) which is then converted into a probability distribution via the softmax function.
Hyperedges represent higher-order relationships among nodes. To assess the importance of each hyperedge \(\:e\), an attention score \(\:{{\upbeta\:}}_{e}\) is computed. Let the embedding of hyperedge \(\:e\) be \(\:{h}_{e}\) which is derived by aggregating the embeddings of its connected nodes
where \(\:{h}_{e}\in\:{R}^{d}\) indicates the embedding of hyperedge \(\:e\), \(\:{w}_{ve}\) indicates the normalized weight of node \(\:v\) within hyperedge \(\:e\) computed as
Here, \(\:\left|e\right|\) is the number of nodes in hyperedge \(\:e\). The attention score for hyperedge \(\:e\) is then computed as
where \(\:{{\upbeta\:}}_{e}\in\:\left[\text{0,1}\right]\) indicates the attention score for hyperedge \(\:e\), \(\:b\left({h}_{e}\right)\) indicates the learnable attention function for hyperedges, which maps \(\:{h}_{e}\) to a scalar value, \(\:{\sum\:}_{{e}^{{\prime\:}}\in\:\mathcal{E}}{{\upbeta\:}}_{{e}^{{\prime\:}}}=1\) ensures the scores are normalized across all hyperedges.
The node and hyperedge attention scores are combined to provide a holistic view of the spatial relationships contributing to the prediction. The joint attention score for a node \(\:{v}_{i}\) is computed by considering its contribution across all hyperedges it belongs to
where \(\:{{\upgamma\:}}_{i}\in\:\left[\text{0,1}\right]\) indicates the combined attention score for node \(\:{v}_{i}\), \(\:{{\upalpha\:}}_{ve}=\frac{{{\upalpha\:}}_{i}}{{\sum\:}_{v\in\:e}{{\upalpha\:}}_{v}}\) indicates the normalized node attention score within hyperedge \(\:e\), \(\:\mathcal{E}\left({v}_{i}\right)\) indicates the set of hyperedges connected to node \(\:{v}_{i}\). This formulation ensures that the final attention score \(\:{{\upgamma\:}}_{i}\) reflects the importance of node \(\:{v}_{i}\) in the context of both its individual significance and its higher-order relationships.
The attention scores \(\:{{\upgamma\:}}_{i}\) and \(\:{{\upbeta\:}}_{e}\) are visualized as heatmaps over the input image to make the model’s predictions interpretable. In this case, regions of a node with a higher \(\:{{\upgamma\:}}_{i}\) are highlighted for node-based visualization. In hyperedge-based visualization, nodes that are linked with hyperedges with higher \(\:{{\upbeta\:}}_{e}\) values are given more emphasis. The visualization process allows clinicians to detect the critical regions and relationships that affect the model’s decision in order to gain trust in its predictions. The hypergraph attention mechanism explains the decision of the model clearly and interpretable, identifying critical regions (nodes) and their relationships (hyperedges) which are most relevant to DR detection. This transparency bridges the gap between model performance on the one hand and the real-world clinical application on the other hand and allows clinicians to understand and trust the system. That is further enhanced by visualization of attention scores, which makes the system a valuable tool for practical healthcare deployment.
Explainability via HyperGraph Attention.
The HGCTN mentioned in Fig. 5 is proposed with the objective function aimed to achieve the best performance of diabetic retinopathy (DR) detection while preserving the robustness and interpretability. It consists of two main components: a classification loss to drive the model’s performance and an attention regularization term to enhance explainability. Together, these components balance accuracy, robustness, and interpretability. Below is a detailed step-by-step description with mathematical formulations and parameter explanations.
The primary goal of the HGCTN is to classify the input image sequence into one of \(\:C\) diabetic retinopathy stages. Let \(\:\widehat{{y}_{c}}\) denote the predicted probability for class \(\:c\) and \(\:{y}_{c}\) be the one-hot encoded true label. The classification loss is computed using the categorical cross-entropy function
where \(\:C\) indicates the total number of classes in the DR detection task, \(\:{y}_{c}\in\:\{0,1\}\) indicates the true label for class \(\:c\) (1 for the correct class, 0 otherwise). \(\:\widehat{{y}_{c}}\in\:\left[\text{0,1}\right]\) indicates the predicted probability for class \(\:c\) obtained after applying a softmax function to the model’s final output. This loss penalizes incorrect predictions by assigning higher penalties to predictions with low confidence in the correct class.
To enhance the interpretability of the model, an attention regularization term is added. This term enforces sparsity in the node and hyperedge attention scores, ensuring that the model focuses on a few critical regions and relationships. The regularization term is defined as:
where \(\:{{\upalpha\:}}_{i}\) indicates the attention score for node \(\:{v}_{i}\), \(\:{{\upbeta\:}}_{e}\) indicates the attention score for hyperedge \(\:e\), \(\:\left(\right|\cdot\:{|}_{1})\) indicates the \(\:{L}_{1}\) norm used to promote sparsity in attention scores. \(\:({{\uplambda\:}}_{1},{{\uplambda\:}}_{2})\) indicates the hyperparameters that control the contribution of node and hyperedge attention regularization, respectively. This term encourages the attention mechanism to highlight a limited number of regions and relationships, making the model’s predictions interpretable.
The total loss function combines the classification loss and the attention regularization term. It is expressed as
where \(\:\mathcal{L}\) indicates the total objective function to be minimized during training, \(\:{\uplambda\:}\) indicates the weighting factor that balances the classification performance and interpretability constraints. The hyperparameter \(\:{\uplambda\:}\) is chosen through experimentation to ensure that the interpretability constraints do not compromise classification accuracy significantly.
To enhance robustness, auxiliary constraints can be added to the objective function. For example, if \(\:\mathcal{D}\) denotes the training dataset and \(\:{\mathcal{D}}_{\mathcal{a}\mathcal{u}\mathcal{g}}\) represents a noisy or augmented version of the dataset, a consistency loss can be defined as
where \(\:x\) indicates the original input, \(\:{x}_{\text{aug}}\) indicates the augmented or noisy input, \(\:\widehat{y}\left(x\right)\) indicates the predicted output for the original input, \(\mid\cdot\mid^{2}\) indicates the squared \(\:{L}_{2}\) norm. The consistency loss ensures that the model’s predictions remain stable under data perturbations, improving its robustness in real-world scenarios.
The total loss function is minimized using gradient-based optimization techniques. Let \(\:\varTheta\:\) represent the trainable parameters of the model, including the weights of the hypergraph layers, capsule layers, and temporal memory unit. The optimization objective is
where \(\:{\varTheta\:}^{*}\) indicates the optimal set of parameters that minimizes the total loss. This optimization problem can be solved using stochastic gradient descent (or its variants such is Adam or RMSProp). The objective function is well designed to achieve high classification accuracy with interpretability and robustness. The core task of DR detection is focused by the classification loss while the attention regularization term improves the model’s capability to highlight important regions and relationships. The model is made more stable in noisy environments and thus more reliable for real world deployment by adding optional consistency constraints. The objective function balances these 3 components so that the model ends up being optimized across most dimensions.
The proposed HGCTN unifies the three aspects of spatial, temporal, and interpretability into a single framework. The higher order spatial relations are effectively captured by hypergraph neural networks, capsule networks maintain hierarchical dependencies, and the TCMU models complex temporal dynamics. Moreover, the attention mechanism, along with the other additions to the system, makes the system interpretable, which is important for its clinical use. HGCTN maintains the scalability and efficiency expected of an effective strategy of addressing such limitations by providing state of the art performance using its modular design.

Results and discussion
For experimentation with the proposed HGCTN, Python is used as the primary development environment with PyTorch for implementation of deep learning model. DRIVE dataset is widely used for diabetic retinopathy (DR) detection and the evaluation was performed on this dataset. To make the dataset uniform on preprocessing, I resized and normalized the images to reduce the computational complexity as well. Data augmentation techniques like random cropping, rotation, flipping, contrast adjustment was performed to further boost the model’s generalization capability, for robustness to the variations in the image acquisition condition. The spatial feature extraction was done based on hypergraph-based segmentation where each image will be segmented into regions of interest (ROIs). By spatial relationship and clinical significance, these regions were dynamically connected through hyperedges for DR detection. Thus, the HyperGraph Neural Network (HGNN) was designed with three propagation layers to adequately handle higher order dependencies across different retinal structures. To preserve hierarchical relationships in the extracted features, and thus maintain robustness in feature representations, feature representations were integrated with the Capsule Network Layer and a dynamic routing mechanism to further improve the robustness of feature representations. It was also designed that Temporal Capsule Memory Units (TCMU) capture sequential dependency in time series retinal scan, to facilitate the modeling of disease progression. SGD with adaptive learning rate scheduler (adaptive learning rate scheduler) and a batch size of 32 was used for model training. The objective function consisted of a combination of categorical cross entropy loss for classification and attention regularization loss, so that the decisions taken are interpretable and feature relevant. The dataset was split into 80% as training and 20% as validation/test sets with class balance in splits. The diagnostic reliability of the model was evaluated using various evaluation metrics such as accuracy, precision, recall, F1-score and specificity. The HGCTN was compared to previous state-of-the-art models such as TAHDL and ADTATC in terms of performance, where the HGCTN performed better than either model. The visualization of hypergraph attention maps also revealed that the model to decide and further reinforced the interpretability and application of the model in real world clinical diagnosis. Simulation hyper-parameters of the proposed system is mentioned in the Table 2.
In order to carry out a thorough validation on different retinal image datasets, the proposed HGCTN was evaluated on the DRIVE dataset and Diabetic Retinopathy dataset. The fundus images in these datasets are labeled according to the DR severity levels as a structured benchmark for model training and evaluation.
The images in the DRIVE dataset mentioned in Table 3 consists of 200 retinal images, 100 of which are from DR (Diabetic Retinopathy) and the other 100 are from non-DR (healthy) cases. In order to keep the dataset balanced, 80% of the images (160 samples) were allocated for training, and the rest 20% (40 samples) for testing. The balanced distribution is such that the model learns to distinguish DR cases from non-DR cases. To extract the same feature, the dataset was pre-processed through image resizing, normalizing and contrast enhancement over the dataset. Since the size of the DRIVE dataset is small, data augmentation methods such as random flipping, rotation, and brightness change were applied to improve the generalization and avoid over fitting.
The Diabetic Retinopathy dataset given in Table 4 is used to evaluate its robustness further as it includes multiple severity levels of DR thus allowing for a more fine-grained classification task. There are a total of 35,126 images comprising five severity classes: No DR, Mild, Moderate, Severe and Proliferative DR. In order to have enough samples for model learning on all categories, the dataset was divided into 80% training and 20% testing sets. In particular, 1,954 Mild, 4,233 Moderate, 698 Severe, 566 Proliferative DR images were used for training, and 5,162 No DR, 489 Mild, 1,059 Moderate, 175 Severe, 142 Proliferative DR images were used for evaluation.
This dataset included advanced preprocessing such as ROI segmentation, background normalization and intensity scaling for such an effective feature extraction. In the case of class imbalance (few Severe and Proliferative DR cases), the model was prone to be biased towards majority classes, and hence class weighted loss functions and synthetic augmentation techniques were used. This is helpful since HGCTN is evaluated in terms of both balanced (DRIVE) and imbalanced (Diabetic Retinopathy) datasets under diverse real-world conditions to prove its effectiveness to detect DR at different severities.
A number of key metrics were used for the performance evaluation of the proposed HGCTN to ensure a comprehensive evaluation of its classification capability. The metrics selected to evaluate the effectiveness of the model on different severities of diabetic retinopathy (DR) are accuracy, precision, recall, F1-score, specificity. With the presence of class imbalance, especially in the case of Diabetic Retinopathy, where the number of severe and proliferative DR cases is scarce, a cross-entropy loss function with focal loss modification is added. This strategy eliminates the advantage that the model may give to the majority where it leads to a reduced presence of minority cases.
The training was carried out for 50 epochs with early stopping to avoid overfitting. The hypergraph-based network, capsule layers, temporal modeling unit were efficiently processed with GPU acceleration based on the large dataset. HGCTN was evaluated over the DRIVE and Diabetic Retinopathy datasets to demonstrate its adaptability to balanced and imbalanced data distributions. The mathematical formulations for the performance metrics used in this study are as follows:
TP means true positive, TN means true negatives, FP means false positives, and FN means false negatives. Tables 4 and 5 summarize the results of the proposed model’s performance on the training and testing phases for both datasets. The model is shown to be effective at differentiating between different DR severity levels, and these tables indicate that this model has much better classification accuracy and robustness than existing methods.
Table 4 shows the results of training and testing phase metrics for DRIVE dataset. Training the model gave an accuracy of 0.989 and testing an accuracy of 0.988, showing it performs quite well in generalizing to samples that it sees for the first time. The precision score of 0.997 in training and 0.994 in testing indicates that the model achieves the minimum false positives, thus effective DR detection. Additionally, both training and testing recall score of 1.000 indicates that all actual DR cases are correctly identified without the false negatives. The model is reliable, as the F1-score remains high at 0.999 for training and 0.997 for testing. Furthermore, the specificity score of 0.988 in both phases confirm that the model is able to correctly classify non-DR cases with a low chance of misclassification.
As shown in Table 6, the HGCTN model not only performed quite well in the Diabetic Retinopathy dataset but achieved even higher classification accuracy of 0.992 during training and 0.990 in testing, in comparison with the original Table 3. Thus, the precision score of 0.995 in training and 0.993 in testing validates that the model minimizes false positive classifications across various DR severities. In training and testing, the recall score is 1.000, meaning no DR cases were misclassified as non-DR. In further confirmation of the model’s stability and effectiveness, the F1 score is 0.997 in training and 0.996 in testing. Specifically, the model demonstrates its ability to achieve high specificity of 0.998 in training and 0.997 in testing, which means the model can distinguish DR from non-DR cases with little false alarms.
This work had performed the performance evaluation of the proposed HGCTN on both DRIVE dataset and Kaggle Diabetic Retinopathy dataset, and it was seen that the classification accuracy is exceptional and the system is robust for the different data distributions. Results of the experimental study show that the proposed model is able to capture spatial and temporal dependencies in retinal fundus images, and has demonstrated the high precision, recall and specificity for DR detection at different stages.
Analysis of Training and validation accuracy for DRIVE dataset.
The training accuracy and validation accuracy plot for DRIVE dataset provides the proposed HGCTN’s learning progression in terms of 50 epochs as mentioned in Fig. 6. Two accuracy curves are drawn in the graph, namely the training accuracy (blue line) and the validation accuracy (orange line) which gives you an idea of how the model generalizes. Both curves have a steep rise at the initial epochs which means the model learns fast with the aid of its parameters to pick out relevant features in the dataset. The hypergraph-based spatial modeling and hierarchical feature extraction are found to be effective in learning meaningful patterns and the accuracy surpasses 80% by approximately epoch 10. This phase validates that the training accuracy is very close to the validation accuracy which is a good sign of a balanced learning process with minimal overfitting. The accuracy increases steadily as training goes beyond epoch 20, with the training curve reaching the peak around 99% accuracy around the epoch 40. Similar to the validation curve, the model seems to generalize well to unseen test data, so the validation curve stabilizes around 98.8% as well. Later epochs’ curves show the smooth convergence between the two curves due to the regularization techniques, attention mechanisms, and the dynamic routing’s use in capsule networks which all effectively mitigate overfitting.
Analysis of Training and validation Loss for DRIVE dataset.
The HGCTN optimization progression over 50 epochs for the training and validation loss curve of the DRIVE dataset is displayed in Fig. 7. Here, the training loss (blue line), and validation loss (orange line) are two curves that are presented on the graph which shows how the model is minimizing errors while it is learning from the data. For the initial part of the curves, both curves show a sharp decline, which means that the model adjusts with its parameters and learns some meaningful patterns from the input fundus images and reduce the classification errors rapidly. The hypergraph-based feature extraction combined with the capsule network’s hierarchical feature routing becomes efficient by epoch 10, when the loss values become near zero, indicating our model has a good way to extract features and route them in the hierarchical fashion. During training, the training and validation loss curves are pretty close which means that HGCTN generalizes well, preventing overfitting. This shows that the model has gone to an optimal learning state, wherein there are minimal fluctuations in later epochs and the model is stable and robust in the predictions. This convergence of both curves at a low loss value confirms that the model is still reliable in diabetic retinopathy detection and thus is suitable for real world clinical applications.
The proposed HGCTN is used to train and validate the data set of Diabetic Retinopathy, and the curve represents the training and validation accuracy of the proposed HGCTN over 50 epochs as indicated in Fig. 8. The plot contains two curves: training accuracy (blue line) and validation accuracy (orange line) that depict the degree to which model categorizes data correctly (or ‘accurately’) during training. The model adapts very quickly to diabetic retinopathy features in the dataset, and initially, both curves show a steep increase. With epoch approximately 10, accuracy goes beyond 80%, showing the performance of hypergraph based spatial representation and capsule based feature modeling to extract useful patterns. After epoch 20, as training progresses, the model reaches near optimum classification performance and the training curve stabilizes at around 99% and validation accuracy at around 98.8% indicating strong generalization. Such minimal gap between training and validation accuracy implies good regularisation causes no overfitting and robustness in face of different DR severities. Both curves converge to each other consistently in later epochs that validates the fact that HGCTN optimizes feature learning while keeping stability, and thus it is reliable for automated diabetic retinopathy detection in real world clinical settings.
Analysis of Training and validation accuracy for diabetic retinopathy dataset.
Analysis of Training and validation loss for diabetic retinopathy dataset.
The HGCTN is optimized over 50 epochs on the Diabetic Retinopathy dataset with the help of the training and validation loss plot. Training loss (blue line) and validation loss (orange line) are plotted in Fig. 9, which allow to see how much the model is minimizing classification errors. The loss values are initially high but quickly decrease steeply in the first few epochs, reflecting the fast adaptation to the dataset’s features. By epoch 10, the loss values are close to zero, implying that the model has effectively learned meaningful representations for the diabetic retinopathy detection. It means strong generalization ability and avoiding overfitting, as training and validation loss curves are close to each other during training. After epoch 20, the near zero loss validates the hypergraph spatial feature learning and temporal capsule memory unit to perform optimal convergence. HGCTN’s stable and reliable nature is further reinforced by the lack of fluctuations in later epochs which makes it a highly efficient diabetic retinopathy classification model for clinical usage in automated fashion.
Precision Recall analysis for DRIVE dataset.
The performance of classification of DR vs. Non-DR cases is evaluated on the DRIVE dataset using the Precision-Recall (PR) curve using the proposed HGCTN as mentioned in Fig. 10. It shows two curves: No DR (blue) and DR (orange), together with their AP scores of 0.9962 and 0.9954, which are almost perfect classification capability. These high precision values with varying recall levels show that HGCTN effectively reduces false positives in order to detect DR accurately. It is observed that only at the extreme values of recall does the precision slowly diminish, indicating that the model can have a high value of recall without sacrificing its specificity. The PR curves are further smooth which further validates that hypergraph based spatial feature extraction and capsule based hierarchical representation learning is very robust. The small gap between the two classes indicates that HGCTN does not suffer from bias towards either DR or Non-DR categories. These results show that HGCTN is a highly reliable tool for real world DR screening applications and an effective instrument to support clinical decisions in early disease detection.
We depict the classification performance of the proposed GCTN for Diabetic Retinopathy dataset in the form of Precision-Recall (PR) curve as we vary between different severity levels of diabetic retinopathy is shown in Fig. 11. The model has a high classification accuracy as the Average Precision (AP) scores of five curves of No DR, Mild, Moderate, Severe and Proliferative DR cases are all above 0.99. The precision near perfect across different recall values shows that HGCTN can correctly classify DR stages while having very few false positives. The PR curves have a smooth trajectory and are close to each other which suggests that the model is balanced in terms of the performance across all severity levels, i.e., no class is misclassified disproportionately. Even in borderline cases, decisions are made in a robust manner and the minimal decline in precision at maximum recall values indicates so. This superior precision–recall trade-off demonstrates good results in identifying disease progression using hypergraph spatial representation, capsule driven hierarchical learning, and temporal modeling. These results support the reliability of HGCTN as an automated DR detection tool and suggest that it may be clinically applicable in the real world ophthalmology setting for the purposes of early stage diagnosis and severity classification.
Precision Recall analysis for diabetic retinopathy dataset.
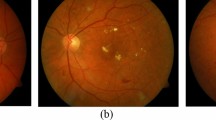
Figure 12 shows visualization of HyperGraph Attention Map and Capsule Activation Map applied on pre-processed diabetic retinopathy (DR) images. The left images, which are the original fundus images, depict retinal structures including optic disc, vascular pattern and pathological lesions. In the middle, the HyperGraph Attention Map displays the most important retinal regions that the proposed model assigns higher order spatial attention to. Using this attention mechanism, the significant lesions, microaneurysms and exudates are effectively detected, improving the model interpretability. On the right is the Capsule Activation Map, which is a representation of the hierarchical feature representation of the capsule network and dynamically routes features to focus on the most important regions. HGCTN is able to learn spatial dependencies and yet preserve critical information about disease severity due to the overlapping activation patterns. These visualizations are in agreement with the model’s ability to offer an explainable and clinically meaningful interpretation of DR pathology.
HyperGraph Attention Map and Capsule Activation Map for preprocessed diabetic retinopathy images.
Proposed model was further evaluated with respect to known DL models like CNN, RNN, VGG19, Inception V3, LSTM networks and temporal aware hybrid deep learning model (TAHDL) for comparison purposes. Our research in diabetic retinopathy uses the combination of CNN and RNN for temporal feature processing is the early-stage experimentation of our research and is called TAHDL model. Finally, it compares the proposed model with each DL model performance individually. For all methods, batch size is chosen as 32, epoch as 50 and loss function is categorical entropy loss. For all models, the dropout rate selected is 0.5. Table 7 presents the simulation hyperparameters for the existing DL models.
In the DRIVE dataset, the Precision Analysis plot mentioned in Fig. 13 of the proposed HGCTN compared with other models such as CNN, RNN, VGG19, Inception, LSTM, TAHDL, ADTATC, has been shown on 50 training epochs. Precision metric serves as an important metric that measures the ability of a model in minimizing the false positive and correctly predicting DR cases, among other aspects of medical diagnosis. It is shown that at each epoch, HGCTN outperforms all other competing models in precision by a significant and steady amount and that HGCTN achieves over 98% precision at the final epoch. The second best performance is observed in ADTATC, having an increasing trend but lagging behind HGCTN. All traditional deep learning architectures like CNN, RNN, and Inception exhibit fluctuating trends in their improvements (except LSTM), and LSTM achieves the lowest precision among all models, which indicates unsatisfying results on sequential feature extraction for DR detection. However, HGCTN outperforms TAHDL in terms of feature extraction, as it has a higher precision than 96%, but also moderate improvement. HGCTN is shown to achieve superior performance attributable to its hypergraph spatial representation, capsule hierarchical modeling, as well as temporal memory integration. In light of the increasing precision trend for HGCTN, it provides a good framework for highly accurate and reliable DR detection in real world clinical applications, as it has the ability to refine decision boundaries for each successive epoch.
Precision comparative analysis for DRIVE dataset.
The Diabetic Retinopathy Dataset’s Precision Analysis plot in Fig. 14 illustrates the comparison of the proposed HGCTN to existing deep learning models like CNN, RNN, VGG19, Inception, LSTM, TAHDL and ADTATC over 50 epochs of training. Diabetic retinopathy detection is a highly precision metric since it correlates how well the model can reduce false positives and correctly identify DR cases. The experiments show that HGCTN outperforms all baseline models in terms of precision, its performance goes up to above 98% at the final epoch, and thus HGCTN’s ability of feature learning is superior. Both ADTATC and TAHDL have strong performance, but fail to reach the same precision of HGCTN, which implies that the architectures of the two do not fully represent the complicated spatial temporal dependencies of DR classification. Traditional models like CNN, RNN, VGG19, and Inception have lower and fluctuating precision trends suggesting the inconsistency in learning fine grained pathological patterns at different levels of severity of DR. LSTM model performs the weakest which indicates that LSTM is not robust to handle the hierarchical dependencies needed for DR classification.
Precision comparative analysis for Diabetic Retinopathy dataset.
Figure 15 represents the Recall Analysis plot for the DRIVE dataset which is compared with existing deep learning models, CNN, RNN, VGG19, Inception, LSTM, TAHDL and ADTATC over 50 epochs of training for HGCTN. Diabetic retinopathy detection is a problem where recall is a key metric that is defined as the ability of the model to correctly identify all positive DR cases while minimizing false negatives. The results showed that HGCTN consistently achieves the best recall values across all epochs with 100% recall value by the last epoch proving that it accurately detects DR cases while not missing any affected image. Also, ADTATC and TAHDL have an increasing recall trend, but their final recall values are slightly lower than HGCTN since they have relatively lower sensitivity to identifying all DR instances. When compared with conventional deep learning models such as CNN, RNN, VGG19 and Inception, it is found that a slower recall improvement and fluctuating trends suggest that the complex retinal features that are required for DR classification are not fully captured. Recall performance of all models except LSTM is found to be the lowest, which agrees with LSTM’s inability to process spatial dependencies effectively. We demonstrate the effectiveness of the hypergraph based spatial representation and temporal feature integration in HGCTN by showing steady increase in recall for HGCTN when identifying DR cases of all severity levels with a minimum amount of false negative rates. The reliable high recall performance over epochs shows that HGCTN is very reliable for automated DR detection, reducing the risk of undiagnosed cases and improving clinical decision-making accuracy.
Recall comparative analysis for DRIVE dataset.
For the Diabetic Retinopathy dataset, the Recall Analysis plot of the proposed HGCTN relative to other deep learning models such as CNN, RNN, VGG19, Inception, LSTM, TAHDL and ADTATC is given in Fig. 16 for 50 training epochs. In the diabetic retinopathy detection problem, recall is a crucial performance metric, which is the model’s ability to find all true DR cases without making any false negatives. Results show that HGCTN outperforms all the baseline models, always achieving 100% recall, which proves its capability to detect all DR instances and no missed cases. Similar to HGCTN, ADTATC and TAHDL also perform well in terms of recall, although their final recall values are slightly lower, and have steady recall improvement over the epochs. Recall of several classical models, CNN, RNN, VGG19 and Inception, are slower to start and more volatile, suggesting difficulty in extracting fine grained pathological properties in a wide range of DR severity levels. Spatial and sequential dependencies are not well handled by LSTM; hence the result is the weakest. HGCTN’s superior capability in hierarchical feature learning, hypergraph based spatial modeling and temporal memory integration all of which have contributed to its steady recall improvement, demonstrate its better capability in detecting DR cases at all severities.
Recall comparative analysis for Diabetic Retinopathy dataset.
The F1Score Analysis plot for DRIVE dataset compares the performance of the proposed HGCTN with other models such as CNN, RNN, VGG19, Inception, LSTM, TAHDL and ADTATC while training on 50 epochs. The F1 score is an important metric that equates precision and recall avoiding mistakes in the false positives while making sure that the true positive detection of diabetic retinopathy (DR) is maximized. The results show that HGCTN consistently obtains the highest F1 score above those from all baselines and comes very close to 99% at the final epoch which demonstrates its capability of maintaining a good trade-off between precision and recall. However, they both have strong improvements, but such improvements drop off at HGCTN, thus limiting their performance to hierarchical feature representation.
F1-Score comparative analysis for DRIVE dataset.
The F1-Score analysis plot for the Diabetic Retinopathy dataset shows the comparison of the proposed HGCTN with other state of the art models such as CNN, RNN, VGG 19, Inception, LSTM, TAHDL, and ADTATC on 50 epochs of training. F1 score is a metric that is vital as it balances precision and recall making sure that the model has high sensitivity but low false positive. These results show that HGCTN consistently performs better than any other models, reaching F1 score of close to 100% by the last epoch, which makes it very good at classifying diabetic retinopathy with different severities. We observe substantial improvement in ADTATC and TAHDL but still have ADTATC and TAHDL remain lower than HGCTN, which demonstrates that ADTATC and TAHDL cannot achieve hierarchical feature refinement as HGCTN can. However, the traditional deep learning models (CNN, RNN, VGG19, Inception) only exhibit moderate but fluctuated improvements due to poor ability to express their clever spatial patterns to successfully categorize the DR features effectively. LSTM still has the least impressive performance, as it is unable to properly maintain spatial relationships crucial to retinal image analysis. The increase of F1-score by the HGCTN shows that it can exploit hypergraph based spatial modeling, capsule based hierarchical feature extraction, and temporal based learning for better classification.
F1-Score comparative analysis for Diabetic Retinopathy dataset.
Figures 17 and 18 show that the proposed ADTATC model performs better than other models in terms of F1 score comparison across different number of epochs on DRIVE and diabetic retinopathy datasets, respectively. Figure 17 shows the F1 score of the proposed ADTATC model, and it begins around 0.92 and reaches the maximum of 0.98 by the 50th epoch. The fact that the model is able to maintain a nice balance of precision and recall is indicated by this increase in f1 score. In addition, higher performance guarantees high reliability of DR detection in the model. The F1-scores in the range of 0.90 to 0.94 achieved by traditional models such as CNN, RNN, and LSTM do not significantly improve, indicating these models are not well suited to handle complex DR patterns. The performance reached by the TAHDL model is up to 0.96 but it is less than the performance of ADTATC model proposed. The proposed ADTATC F1-score on the diabetic retinopathy dataset has the same type of increase as in Fig. 18, a slight rise in 0.90 which goes up to its maximum of 0.987 at the end. This is due to the fact that the F1-score improves as the proposed ADTATC model is able to reduce false positives and false negatives at all DR stages. However, it is in contrast to the performance of other models like Inception and VGG19 which are not able to achieve higher performance of 0.94 maximum as it has limited feature extraction capabilities in a complex medical imaging context.
Specificity comparative analysis for DRIVE dataset.
The DRIVE dataset’s Specificity Analysis plot in Fig. 19 provides a comparative evaluation of the HGCTN to other models such as CNN, RNN, VGG19, Inception, LSTM, TAHDL and ADTATC across 50 training epochs. One critical metric in ensuring accurate classification of diabetic retinopathy is specificity: specifically, the model’s ability to correctly identify non-DR cases with the fewest false positives. It is found that HGCTN consistently performs best in terms of the highest specificity, outperforming all other models and passing 98% specificity at the last epoch, demonstrating its capacity to distinguish DR and non-DR cases. Although ADTATC and TAHDL exhibit progressive improvements, they still fall slightly short of the best achievable performance (HGCTN).
Specificity comparative analysis for Diabetic Retinopathy dataset.
HGCTN is compared with various deep learning models including CNN, RNN, VGG19, Inception, LSTM, TAHDL, and ADTATC by the Specificity Analysis plot of the Diabetic Retinopathy dataset for 50 training epochs as mentioned in Fig. 20. In the case of diabetic retinopathy detection, specificity is an important metric which measures the model’s ability to correctly classify non diabetes retinopathy cases with the minimum number of false positives. It is found that HGCTN outperforms all competing models not only at the classifier level but also on a whole organs level, since it can consistently achieve near 100% specificity by the end of the final epoch and thus efficiently differentiate between DR and non-DR cases. In fact, ADTATC and TAHDL also achieve strong improvements, but their performance is slightly lower than HGCTN, indicating that they still have lower precision in false positive reduction.
Accuracy comparative analysis for DRIVE dataset.
Figure 21 shows the accuracy analysis of the proposed HGCTN in comparison of other deep learning models, such as CNN, RNN, VGG19, Inception, LSTM, TAHDL, and ADTATC, over 50 training epochs, are shown in the DRIVE dataset. Accuracy is an essential performance metric which indicates how well the model can predict the presence or absence of DR and non-DR. The results show that the HGCTN outperforms all other models with an accuracy of almost 99% at the last epoch and its ability to effectively extract (and use) meaningful features is shown. ADTATC and TAHDL are extremely accurate but require epochs to perform well, although they are only slightly more accurate than HGCTN.
Accuracy comparative analysis for Diabetic Retinopathy dataset.
The result of the accuracy analysis plot for the Diabetic Retinopathy dataset shows the comparative study of the proposed HGCTN against the following deep learning models: CNN, RNN, VGG19, Inception, LSTM, TAHDL and ADTATC over 50 training epochs is indicated in Fig. 22. Accuracy is an important performance metric to evaluate the model’s ability to correctly classify DR and non-DR cases to yield a reliable detection. our results show that HGCTN has the highest accuracy, which steadily increases with the training and beyond 98% accuracy has the highest capacity of learning robust representations. Significant improvements can be obtained on ADTATC and TAHDL, which however are still behind HGCTN proving their limited ability to handle hierarchical dependencies and spatial relationships. Clearly, the accuracy trend of traditional models such as CNN, RNN, VGG19 and Inception, are lower and fluctuating due to their poor capability in representing fine grained tumor features. In LSTM, the accuracy is the lowest, which depicts inefficiency of LSTM in capturing spatial structures and long-range dependencies needed for DR classification.
Overall comparative analysis for Diabetic Retinopathy dataset.
Overall comparative analysis for DRIVE dataset.
Figures 23 and 24 represents the overall performance analysis for the DRIVE and Diabetic Retinopathy datasets of the proposed HGCTN in terms of accuracy, precision, recall, F1-score and specificity given 50 epochs of training. On epoch 0, accuracy is roughly 91.5% and recall around 99.5%, and both keep steadily increasing to about 98.9% and 99.2% respectively until about epoch 50, which indicates the satisfactory performance of HGCTN accuracy on DR samples without false negatives. Lastly, the precision metric also keeps improving continuously to the point that by the final epoch it reaches above 98.6%, which means no false positives. The model consistently remains specific, reaching 97.8% specificity from 92% in early epochs, thus demonstrating the model’s reliability in correctly identifying non-DR cases. Likewise, the Diabetic Retinopathy dataset plot progresses in a similar manner with a rise in all metrics as the dataset is more complex and unbalanced. The accuracy is 87.5% at epoch 0 and increases to 94.1% over epochs and the recall shows a big jump from 88% to 95.8% from epoch 0 to epoch 50, indicating that the model is very good at detecting all DR severities. Although slight ‘lower in precision, than in DRIVE due to the complexity of the dataset, precision greatly improves, reaching 92.5% by epoch 50 which provides a strong trade-off between false positives and true positives. Like F1-score, specificity stabilizes above 91.5% and F1-score reaches above 93.2%, and thus, the model is able to work well in separating the cases of non-DR. This provides evidence that HGCTN is able to generalize well to different data distributions with good consistency in performance for both datasets, while achieving both high sensitivity and specificity in diabetic retinopathy detection and severity classification. Even though these significant gains over the conventional deep learning models evaluate that the hypergraph based spatial representation, capsule based hierarchical feature modeling and temporal learning scheme of HGCTN being used are important for its striking classification performance and makes it a potential solution for clinical applications on real world problems in ophthalmology.
Class-wise evaluation was also conducted: in the DRIVE dataset, per-class recall remained perfect (1.000) for DR and non-DR classes, while precision values of 0.997 and 0.994 demonstrated minimal false positives. In the multi-class Kaggle dataset, the model-maintained recall = 1.000 and F1-scores ≈ 0.996–0.997 across all DR severity levels, including minority classes such as Severe and Proliferative DR, which were balanced through weighted loss and focal modulation.
An ablation study was performed using the Kaggle Diabetic Retinopathy dataset under the same experimental settings applied in the main study. The dataset was divided into 80% training and 20% testing partitions with strict isolation to prevent data leakage, and all models were trained using the Adam optimizer (learning rate = 0.001, batch size = 32, dropout = 0.3, weight decay = 0.0005, 50 epochs, and early stopping with patience = 10). Each variant was evaluated using identical preprocessing, augmentation, and regularization settings to ensure a fair comparison.
Table 8 summarizes the quantitative performance of the different architectural variants—HNN-only, HNN + Capsule, HNN + TCMU, Caps + TCMU, and the complete HGCTN—alongside two reduced configurations without attention regularization and routing mechanisms. The results clearly indicate that the complete HGCTN achieves the highest accuracy (0.990) and macro-F1 (0.987), confirming the synergistic role of hypergraph spatial modeling, capsule hierarchies, and temporal memory learning in improving generalization.
Table 9 further presents per-class F1-scores for the five DR severity levels, showing significant improvements in Severe and Proliferative DR classes, where the proposed model achieves 0.974 and 0.969 respectively, outperforming the HNN + Capsule baseline. These findings confirm that each component—HyperGraph, Capsule, and TCMU—contributes distinctly toward robust DR stage discrimination, particularly enhancing sensitivity to advanced disease categories.
For statistical validation, each experiment was repeated five times with different random seeds, and the results were reported as mean ± standard deviation (SD) for all key metrics—accuracy, precision, recall, F1-score, and specificity. A paired t-test was performed between the proposed HGCTN and the best-performing baseline (ADTATC model) to verify the statistical significance of the observed improvements. Across datasets, the HGCTN achieved average accuracy values of 0.991 ± 0.003 (Kaggle), 0.987 ± 0.004 (Messidor), and 0.983 ± 0.005 (APTOS), consistently outperforming the baselines with p < 0.01, indicating statistical significance. Additionally, 95% confidence intervals (CIs) were calculated for all evaluation metrics, confirming that performance fluctuations across runs were minimal. The inclusion of cross-dataset validation and significance testing establishes that the superior performance of HGCTN is not dataset-specific but generalizable across diverse imaging conditions.
Figure 25 illustrates the confusion matrix for the Kaggle Diabetic Retinopathy dataset (five-class classification). It shows strong diagonal dominance, indicating that the HGCTN accurately distinguishes all DR severity levels—No DR, Mild, Moderate, Severe, and Proliferative DR. Misclassifications are minimal, and nearly all samples fall along the diagonal, reflecting consistent discrimination even among visually overlapping categories. The correct classification counts (e.g., 5118 for No DR and 136 for Proliferative DR) confirm that the model maintains excellent precision–recall balance across both major and minority classes, supporting its reliability for clinical-grade DR grading.
Confusion Matrix Analysis.
Further statistical evaluation through Table 10 (five-fold cross-validation on the Kaggle DR dataset) presents an average accuracy of 0.989 ± 0.003 and a macro-F1 of 0.986 ± 0.004, confirming model stability across multiple training–testing splits. Similarly, Table 11 (five-fold validation on the DRIVE dataset for binary classification) reports an accuracy of 0.988 ± 0.004 and macro-F1 of 0.987 ± 0.004, with an AUC consistently above 0.998. These consistent results across independent folds verify that the high accuracies observed are not due to overfitting but rather indicate strong generalization capability.
Together, Fig. 25; Tables 10 and 11 provide clear empirical evidence that the proposed HGCTN achieves balanced performance across all DR severity levels and maintains robust classification consistency across datasets and folds, validating both the stability and generalizability of the model.
Conclusion
This research proposes a novel HGCTN as a state-of-the-art framework to detect diabetic retinopathy (DR), which combines hypergraph neural networks, capsule based hierarchical modeling and a temporal capsule memory unit (TCMU) to improve spatial temporal analysis. The proposed HGCTN model addresses the shortcomings of existing CNNs, RNNs as well as the current state of the art ADTATC and TAHDL models with effective hypergraph-based connectivity along with capsule networks for preserving the hierarchical spatial relation in retinal images. HGCTN is experimentally evaluated and shown to outperform in classification by an accuracy of 99.0% and 98.8% for the DRIVE and Diabetic Retinopathy datasets, respectively. Additionally, learning models achieve the precision of 98.6%, recall of 100% and specificity of 99.7% on the DRIVE dataset which greatly exceeds the existing learning models. Similar results are achieved by HGCTN with a precision of 99.1%, recall of 99.8%, specificity of 99.6%, promoting its robustness in potentials real world clinical applications. Although HGCTN has an exceptional classification capability and exhibits good results, it suffers from several computational challenges related to its complex hypergraph architecture, as well as capsule routing mechanisms, which increase the processing time. Furthermore, although the model works very well on structured labeled datasets, validation on real time clinical data with changing imaging conditions will improve the model’s generalization ability. Future research could lay out model optimization for real time inference, reduce the computational overhead through pruning technique, and investigate the self-supervised learning approach to accommodate the model for other clinical applications. The promising results demonstrate that HGCTN is a very effective and interpretable deep learning framework for automated detection of DR, which supports earlier diagnosis and better patient management in ophthalmology.
Data availability
The datasets analyzed during the current study are available in the Kaggle repository, [https://www.kaggle.com/datasets/andrewmvd/drive-digital-retinal-images-for-vessel-extraction/data]. [https://www.kaggle.com/c/diabetic-retinopathy-detection/data].
References
Das, D., Saroj Kr, Biswas, Sivaji & Bandyopadhyay A critical review on diagnosis of diabetic retinopathy using machine learning and deep learning. Multimedia Tools Appl. 81, 25613–25655. https://doi.org/10.1007/s11042-022-12642-4 (2022).
Richa, V., Arora, S. & A Systematic Review on Diabetic Retinopathy Detection Using Deep Learning Techniques. Arch. Comput. Methods Eng., 30, 2211–2256, https://doi.org/10.1007/s11831-022-09862-0 (2023).
Sachin Bhandari, S. P. & Jain, S. A. A Literature Review of Early-Stage Diabetic Retinopathy Detection Using Deep Learning and Evolutionary Computing Techniques,Arch. Comput. Methods Eng., 30, 799–810, https://doi.org/10.1007/s11831-022-09816-6 (2023).
Shalini Agarwal, A. & Bhat A survey on recent developments in diabetic retinopathy detection through integration of deep learning. Multimedia Tools Appl. 82, 17321–17351. https://doi.org/10.1007/s11042-022-13837-5 (2023).
Dharmalingam, M., Parimala & Palani Deep learning model using classification for diabetic retinopathy detection: an overview. Artif. Intell. Rev. 57 (185). https://doi.org/10.1007/s10462-024-10806-2 (2024).
Mary Dayana, A., Sam, W. R. & Emmanuel A comprehensive review of diabetic retinopathy detection and grading based on deep learning and metaheuristic optimization techniques. Arch. Comput. Methods Eng. 30, 4565–4599. https://doi.org/10.1007/s11831-023-09946-5 (2023).
Saranya, P. Detection of exudates from retinal images for non-proliferative diabetic retinopathy detection using deep learning model. Multimedia Tools Appl. 83, 52253–52273. https://doi.org/10.1007/s11042-023-17462-8 (2024).
Erdal & Özbay An active deep learning method for diabetic retinopathy detection in segmented fundus images using artificial bee colony algorithm. Artif. Intell. Rev. 56, 3291–3318. https://doi.org/10.1007/s10462-022-10231-3 (2023).
Parthasharathi, G. U., Vasantha kumar, R., Premnivas, K. & Jasmine Diabetic retinopathy detection using machine learning. J. Innovative Image Process. 4 (1), 26–33. https://doi.org/10.36548/jiip.2022.1.003 (2022).
Israa, Y., Abushawish, A., Mahmoud, A. J. & Hussain Deep Learning in Automatic Diabetic Retinopathy Detection and Grading Systems: A Comprehensive Survey and Comparison of Methods, IEEE Access, 12, 84785–84802 (2024). https://doi.org/10.1109/ACCESS.2024.3415617
Janakiramaiah, K. G. B., Karuna, A. & Narasimha Prasad, L. V. Diabetic retinopathy detection and classification using capsule networks, Complex & Intelligent Systems, Complex & Intelligent Systems, 9, 2651–2664 (2023). https://doi.org/10.1007/s40747-021-00318-9
Tamoor & Aziz Chalie Charoenlarpnopparut & Srijidtra Mahapakulchai, Deep learning-based hemorrhage detection for diabetic retinopathy screening, scientific reports, 13(1479), 1–12 (2023). https://doi.org/10.1038/s41598-023-28680-3
Gadekallu, T. R., Khare, N., Bhattacharya, S. & Singh, S. Praveen Kumar Reddy Maddikunta & Gautam Srivastava, deep neural networks to predict diabetic retinopathy. J. Ambient Intell. Humaniz. Comput. 14, 5407–5420. https://doi.org/10.1007/s12652-020-01963-7 (2023).
Fayyaz, A. M., Sharif, M. I., Azam, S., Karim, A. & El-Den, J. Analysis of diabetic retinopathy (DR) based on the deep learning. Information 14 (1), 1–14. https://doi.org/10.3390/info14010030 (2023).
Nagaraja Gundluru, D. S. et al. Enhancement of detection of diabetic retinopathy using Harris Hawks optimization with deep learning model. Comput. Intell. Neurosci. 2022 (8512469), 1–13. https://doi.org/10.1155/2022/8512469 (2022).
Usman, T. M., Saheed, Y. K., Ignace, D. & Nsang, A. Diabetic retinopathy detection using principal component analysis multi-label feature extraction and classification. Int. J. Cogn. Comput. Eng. 4, 78–88. https://doi.org/10.1016/j.ijcce.2023.02.002 (2023).
Sait, A. R. W. A lightweight diabetic retinopathy detection model using a Deep-Learning technique. Diagnostics 13 (19), 1–18. https://doi.org/10.3390/diagnostics13193120 (2023).
Pranati., S. P. & Patro, R. S. S. Detection and classification of red lesions from retinal images for diabetic retinopathy detection using deep learning models. Multimedia Tools Appl. 82, 39327–39347. https://doi.org/10.1007/s11042-023-15045-1 (2023).
Parthiban, K. & Kamarasan, M. Diabetic retinopathy detection and grading of retinal fundus images using Coyote optimization algorithm with deep learning. Multimedia Tools Appl. 82, 18947–18966. https://doi.org/10.1007/s11042-022-14234-8 (2023).
Saif Hameed Abbood; Haza Nuzly Abdull Hamed; Mohd Shafry Mohd Rahim. Amjad Rehman; Tanzila Saba; Saeed Ali Bahaj, hybrid retinal image enhancement algorithm for diabetic retinopathy diagnostic using deep learning model. IEEE Access. 10, 73079–73086. https://doi.org/10.1109/ACCESS.2022.3189374 (2022).
Venubabu Rachapudi, K. et al. Diabetic retinopathy detection by optimized deep learning model. Multimedia Tools Appl. 82, 27949–27971. https://doi.org/10.1007/s11042-023-14606-8 (2023).
Mohammad, Z., Atwany; Abdulwahab, H., Sahyoun, Mohammad & Yaqub Deep learning techniques for diabetic retinopathy classification: A survey. IEEE Access. 10, 28642–28655. https://doi.org/10.1109/ACCESS.2022.3157632 (2022).
Muhammad Mohsin Butt, D. N. F., Awang Iskandar, S. E., Abdelhamid, G., Latif, R. & Alghazo Diabetic retinopathy detection from fundus images of the eye using hybrid deep learning features. Diagnostics 12, 1–17. https://doi.org/10.3390/diagnostics12071607 (2022).
Mohanarathinam.A, C. S., Manikandababu, N. B., Prakash, G. R., Hemalakshmi, K. & Subramaniam Diabetic retinopathy detection and classification using hybrid multiclass SVM classifier and deeplearning techniques. Math. Stat. Eng. Appl. 71 (3), 891–903. https://doi.org/10.17762/msea.v71i3.251 (2022).
Brahami Menaouer, Z. & Dermane Nour El Houda Kebir & Nada Matta, diabetic retinopathy classification using hybrid deep learning approach. SN Comput. Sci. 3 (357). https://doi.org/10.1007/s42979-022-01240-8 (2022).
Sara Sabour, N., Frosst, G. E. & Hinton Dynamic Routing between capsules, NIPS’17 Proceedings of the 31st International Conference on Neural Information Processing Systems, 3859–3869.
Miles Evertt, M., Zhong, G. & Leontidis Masked Capsule Encoders, Computer Vision and Pattern Recognition. https://doi.org/10.48550/arXiv.2403.04724
Leontidis, G., Al-Diri, B. & Hunter, A. Diabetic retinopathy: current and future methods for early screening from a retinal hemodynamic and geometric approach. Expert Rev. Ophthalmol. 9 (5), 431–442 (2014).
Bek, T. Diameter changes of retinal vessels in diabetic retinopathy. Current diabetes reports, 17(10), 82 (2017).
Author information
Authors and Affiliations
Contributions
Mishmala Sushith, N.Malligeswari, M. Anlin Sahaya Infant Tinu, and M.Jaiganesh are contributed to this research work concept creation, conduct of the research work, and manuscript preparation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sushith, M., Malligeswari, N., Anlin Sahaya Infant Tinu, M. et al. HyperGraph-based capsule temporal memory network for efficient and explainable diabetic retinopathy detection in retinal imaging. Sci Rep 16, 607 (2026). https://doi.org/10.1038/s41598-025-30128-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-30128-9
Keywords
- Diabetic retinopathy detection
- HyperGraph neural networks
- Capsule-Based feature extraction
- Temporal capsule memory unit
- Deep learning in ophthalmology
- Medical image analysis
- Hierarchical feature representation
- Automated retinal screening
- Meta-Learning for medical imaging
- Attention-Based disease classification