Abstract
Immunoglobulin A (IgA) nephropathy (IgAN) is a prevalent primary glomerulonephritis with progressive potential. Early identification of high-risk patients is critical; however, current clinical and pathological markers are limited. This study aimed to identify epigenetic biomarkers in circulating CD8⁺ T cells that discriminate IgAN patients with different disease severity. Seventeen patients with biopsy-proven IgAN were stratified into early- and late-stage groups based on kidney function. CD8⁺ T cells were isolated and analyzed using transposase-accessible chromatin sequencing (ATAC-Seq) to assess chromatin accessibility. Differentially accessible regions (DARs) were identified and selected biomarkers were additionally analyzed with ATAC-qPCR. In total, 279 DARs were identified, of which 122 were selected as stage-specific biomarkers. CD8⁺ T cells from the early-stage group exhibited higher chromatin accessibility, and t-SNE showed a clear separation between the stages. Deconvolution analysis revealed the enrichment of naïve CD8⁺ T cells in the early-stage group and terminally differentiated effector memory CD8⁺ T cells in the late-stage group. Motif analysis uncovered distinct regulatory signatures: ETS1, LEF1, and RUNX2 in the early-stage, EOMES, TBX21, and IRF4 in the late stage. Receiver operating characteristic (ROC) analysis showed strong discriminatory power of the top biomarkers, enhanced by a composite weighted score. ATAC-qPCR confirmed the chromatin accessibility patterns observed using ATAC-Seq. This study defined stage-specific chromatin landscapes in the circulating CD8⁺ T cells of patients with IgAN and identified non-invasive epigenetic biomarkers associated with different disease severity, which may be utilized in early risk stratification and personalized management.
Similar content being viewed by others
Introduction
IgA nephropathy (IgAN) is a highly prevalent primary glomerulonephritis, with a significant proportion of patients progressing to end-stage kidney disease (ESKD)1. However, accurate assessment of disease severity and progression risk remains a major clinical challenge. Current prognostic markers, including proteinuria and histopathological findings, are limited in capturing patients with early disease dynamics before irreversible loss of kidney function occurs2,3. This delay in detection limits the opportunity for early intervention. Therefore, disease-specific and non-invasive biomarkers that reflect underlying immunopathology and can discriminate early-stage patients with progressive potential are needed to guide timely and personalized treatment.
Increasing evidence suggests that epigenetic dysregulation—including DNA methylation, histone modifications, and non-coding RNA activity—plays a critical role in IgAN pathogenesis. Altered expression of C1GALT1, a key enzyme for galactose-deficient IgA1, has been linked to microRNA upregulation4, while histone methylation changes have been associated with aberrant IgA receptor signaling5. Recent advances in chromatin accessibility profiling allow for cell-type–specific insights into transcriptional regulation and immune cell state dynamics in various diseases6,7,8,9. Since chromatin accessibility governs transcription factor binding and gene expression, this approach offers a powerful tool to elucidate disease-specific regulatory mechanisms and identify biomarkers that reflect disease heterogeneity, even among patients at similar clinical stages.10
Previous studies have highlighted the role of T cells in the pathogenesis of IgAN11,12. In particular, increased activated CD8⁺ T cells showed correlation with renal function deterioration12, while increased tubulointerstitial CD8⁺ T cell infiltration was associated with disease progression13. Recent single-cell analyses have identified specific CD8⁺ T cell subpopulations with distinct transcriptional signatures in IgAN patients, suggesting heterogeneous functional roles14. These findings collectively suggest that CD8⁺ T cells may contribute to both glomerular and interstitial injury through cytotoxic and pro-inflammatory functions. However, the precise immunological regulatory pathways of CD8⁺ T cells on pathogenesis and progression of IgAN require further investigation.
Thus, in the study, we applied Assay for Transposase-Accessible Chromatin using sequencing (ATAC-Seq) to sorted peripheral CD8⁺ T cells from biopsy-proven IgAN patients, and aimed to identify stage-specific chromatin accessibility profiles and uncover novel epigenetic biomarkers associated with discrimination of early IgAN. Furthermore, we evaluated the potential of these epigenetic markers as candidate biomarkers and explored their translational applicability in personalized therapeutic approaches.
Methods
Ethical statement
This study was approved by the Institutional Review Board (IRB) of Seoul National University Hospital (SNUH) (IRB No. 1903-128-1020). The use of peripheral blood samples stored in the SNUH Human Biobank was authorized with written informed consent. The authors adhered to the ethical principles of the latest version of the Declaration of Helsinki.
Study design and participants
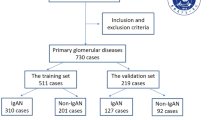
We screened adult patients (≥ 20 years old) with biopsy-confirmed IgAN who donated kidney tissue, blood, and urine samples to the SNUH human biobank between 2006 and 2019. Patients were stratified into early- and late-stage groups based on eGFR and urine protein-to-creatinine ratio (UPCR) value measured at the time of peripheral blood sampling and during follow up (median follow up: 3.16 years, interqurartile ranges 2.71–3.35). Early-stage disease was defined as having an eGFR ≥ 60 mL/min/1.73 m2 at the time of peripheral blood sampling, an average eGFR ≥ 30 mL/min/1.73 m2 (calculated from serial measurements during the follow-up period after peripheral blood sampling), or UPCR < 3 g/g. Those who did not meet the criteria were classified as late-stage group. In addition, those with a last follow up eGFR ≤15 mL/min/1.73 m2 were classified as late-stage irrespective of other criteria. Then, a total of 17 patients were enrolled and stratified into an early-stage group (n = 11) and a late-stage group (n = 6).
Cell sorting and sample preparation
Peripheral blood mononuclear cells (PBMCs) were isolated using a Ficoll–Histopaque gradient (GE Healthcare Life Sciences, Piscataway, NJ, USA). Isolated PBMCs were resuspended in a freezing medium (50% fetal bovine serum [FBS], 10% dimethyl sulfoxide [DMSO], and 40% RPMI-1640) (Invitrogen, Carlsbad, CA, USA) and stored in liquid nitrogen. For CD8+ T cell isolation, cryopreserved PBMCs were thawed, stained with fluorochrome-conjugated anti-human CD8 antibodies, and sorted using a FACSAria III cell sorter (BD Biosciences, San Jose, CA, USA).
ATAC library preparation and sequencing
ATAC libraries were prepared from 50,000 sorted viable CD8⁺ T cells per sample, as previously described15. Cells were washed with 1× cold Dulbecco’s phosphate-buffered saline (DPBS) at 4°C and pelleted by centrifugation at 500 × g for 5 min at 4°C. Pellets were resuspended in 50 μL of cold lysis buffer (10 mM Tris-HCl, pH 7.5, 10 mM NaCl, 3 mM MgCl2, 0.1% NP-40, 0.1% Tween-20, and 0.01% Digitonin) and incubated on ice for 3 min. Lysis was stopped with 1 mL of wash buffer (10 mM Tris-HCl, pH 7.5, 10 mM NaCl, 3 mM MgCl2, 0.1% Tween-20), followed by centrifugation at 500 × g for 10 min at 4°C to collect nuclei.
Isolated nuclei were incubated in transposase reaction mix (25 μL 2× TD buffer, 16.5 μL 1× cold DPBS, 2.5 μL Tn5 transposase [Illumina, San Diego, CA, USA], 0.5 μL 0.1% Tween-20, 0.5 μL 0.01% Digitonin, and 5 μL nuclease-free water), at 37°C for 45 min on a thermomixer at 1,000 rpm. Transposed DNA was purified using a Qiagen PCR cleanup kit. Library amplification was performed with indexing primers from the Nextera kit (Illumina). The optimal number of PCR cycles for each sample was determined via qPCR to avoid over-amplification. Final libraries were quantified using a Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA) and verified with the KAPA Library Quantification Kit (Roche Applied Science, Basel, Switzerland). Indexed libraries were pooled and sequenced on an Illumina NextSeq6000 platform using a paired-end 150 bp sequencing strategy.
ATAC-Seq data processing and biomarker selection
Paired-end reads from the sequenced ATAC-Seq libraries were aligned to the human reference genome (hg19) using Bowtie2 (v2.4.1) software with the default parameter set and the “-X 2000” option to accommodate the expected fragment size range16. Aligned reads were converted to BED format using the “bamToBed” function in BEDtools17. To correct for Tn5 transposase insertion bias, the 5′ ends of reads were adjusted by +4 bp for the positive strand and −5 bp for the negative strand, accounting for the 9 bp duplication introduced during tagmentation. Prior to downstream analysis, library quality and complexity were assessed according to the ENCODE ATAC-seq quality control guidelines, using parameters such as PBC1, PBC2, NRF, FRiP, and TSSE (Supplementary Table S1). Genome browser tracks were generated for each sample using MACS3 (v3.0.0a5) software with the “callpeak --bdg” option18. Sequentially, the resultant bedGraph files were converted to bigWig format using the “bedGraphToBigWig” function in KentUtils19. To enable inter-sample comparison, genome browser tracks were normalized using a scaling factor derived from stable control ATAC-peak regions identified across all samples. Initially, 20 control loci were used as described previously20, and subsequent validation confirmed that the normalization factor converged with expansion to >200 invariant loci. In total, 44,298 raw ATAC-peaks were detected across all CD8⁺ T-cell samples (q < 0.001). After removing sample-specific and low-confidence peaks, we defined a consensus set of 22,149 peaks that were present in ≥ 80% of donors and had an average ATAC-peak area above the median. These peaks were used for downstream analyses to ensure reproducibility. We then identified candidate ATAC-peaks that differentiated between the early and late groups by selecting regions with thresholds of fold change > 2.0 in either direction and P < 0.05 using a t-test. To maintain consistency with current genomic references, all genomic coordinates were converted from hg19 to hg38 using the UCSC liftOver tool. The complete list of liftover-converted differentially accessible regions (DARs) for the 279 ATAC-marker candidates is provided in Supplementary Table S2, and the statistical results from the negative-binomial generalized linear model (GLM) using edgeR21, incorporating age and sex as covariates (~ group + age + sex) are provided in Supplementary Table S3.
Enrichment score estimation of CD8+ T-cell subsets
To more accurately reflect disease-associated alterations in CD8+ T-cell subsets, we established a disease-specific reference framework using a single-cell RNA-sequencing (scRNA-seq) dataset generated from the same IgAN patient cohort (GSE285335; data under review). CD8⁺ T cells were clustered using Seurat (v5.2.1), and four major subsets were identified based on IL-7Rα expression and canonical differentiation markers, corresponding broadly to naïve-like+central memory and progressively differentiated effector-memory populations.
Signature genes from total four subsets were determined by differential expression analysis (one subset versus all others) using edgeR with thresholds of fold change > 1.5 and P < 0.05 21. Genes showing stable intra-cluster expression (coefficient of variation < 0.3) were retained, and overlapping genes between subsets were excluded. Promoter-associated ATAC-peaks in the bulk dataset were selected when these peaks overlapped to promoter regions (core-, proximal-, and distal promoters) of non-redundant signature genes.
The corresponding promoter peaks were assembled into a subset-specific signature matrix after Z-score normalization to minimize the inter-sample variability. Subset enrichment within each bulk ATAC-seq sample was quantified using single-sample gene set enrichment analysis (ssGSEA) implemented in the GSVA R-package (v2.0.4). Enrichment scores derived from promoter accessibility and compared between early- and late-stage IgAN groups using Student’s t-test.
Calculation of weighted chromatin openness score
To quantify chromatin openness, a weighted composite score was calculated for each donor based on selected ATAC-based biomarkers. For each biomarker, the normalized peak area was binarized by assigning a value of 1 if it exceeded a predefined threshold, and 0 otherwise. Biomarkers were ranked by relative importance, and weights were assigned accordingly in descending order. The final weighted score for chromatin openness was obtained by multiplying each binary value by its corresponding weight and summing the products across all selected biomarkers.
Determination of cut-off for biomarkers and weighted score
To determine optimal cut-off values for predictive biomarkers based on normalized chromatin accessibility, Cutoff Finder tool22 was adopted. The cut-off value was established by comparing the early- and late-stage groups and selecting the point that minimized the Euclidean distance to the top-left corner of the ROC curve to achieve optimal sensitivity and specificity. The determination process for each DAR cut-off value consisted of two steps. First, the predictive performance of the cut-off value was evaluated by calculating the area under the ROC curve for each biomarker. Second, the discriminative ability of the cut-off value was assessed by calculating its sensitivity and specificity, ensuring robust separation between the early- and late-stage IgAN groups. These cut-off values were subsequently used for both individual biomarker analysis and the calculation of the weighted chromatin openness score.
Motif analysis and Identification of downstream genes
Transcription factor motifs within DARs were identified using the “findKnownMotifs.pl” function in the HOMER suite23, which assigns known motifs and calculates their statistical significance. Significant motifs were ranked based on ascending P-values, and top five significantly enriched motifs were selected independently for the early- and late-stage groups. Consensus motifs were then generated based on the frequency of actual sequence occurrences within assigned peaks. The visualization of consensus motifs was performed using the “seqLogo” R-package24, allowing graphical representation of base composition across motif positions.
To explore the downstream regulatory effects of the identified transcription factors, Ingenuity Pathway Analysis (IPA)25 was used. IPA enabled the prediction of transcriptional regulatory networks and the identification of functionally relevant target genes. To assess the expression profiles of these downstream genes, we used publicly available RNA-seq data from defined immune cell subsets in peripheral blood of healthy individuals (GSE227743). Only genes with a minimum expression level greater than 20 were included for visualization.
Quantitative assessment of chromatin openness using ATAC-qPCR
Chromatin accessibility at selected genomic loci was quantitatively assessed using real-time PCR with primers targeting regions proximal to the summits of previously identified ATAC- peaks. Primer pairs were designed using Primer-BLAST to generate amplicons of 100 – 150 base pairs. Primers predicted to produce off-target products under 500 base pairs or containing mismatches within the 3′ terminal three bases were excluded to ensure specificity. A comprehensive list of validated primer sequences for all target and control loci is provided in Supplementary Table S4.
ATAC-qPCR was performed using a 1:40 dilution of the original ATAC library as the template. Reactions were run in technical triplicates on a QuantStudio™ 6 Real-Time PCR System (Thermo Fisher Scientific, Waltham, MA, USA) using SYBR GreenER™ qPCR SuperMix (Thermo Fisher Scientific) and 200 nM each of forward and reverse primer. The thermal cycling protocol included UDG activation (50°C for 2 min), polymerase activation (95°C for 2 min), followed by 40 cycles of denaturation (95°C for 15 sec) and annealing/extension (60°C for 1 min). All primer sets underwent pre-evaluation using 1 ng of genomic DNA isolated from pooled PBMCs of IgAN patients (QIAamp® DNA Mini Kit, Qiagen) under the same cycling conditions. Only primer pairs exhibiting efficient and specific amplification, without primer-dimer formation (confirmed by melting curve analysis), were selected for subsequent analysis. To determine normalized chromatin openness, mean Ct values were transformed using the 2–Ct method. Relative chromatin openness was then calculated by normalizing these transformed values to the geometric mean of three control loci.
Statistical analyses
Data are presented as means ± standard deviation (SD) or means ± standard errors of the mean (SEM). Group comparisons were performed using Student’s t-test. Correlations were assessed using Pearson’s correlation coefficient. All statistical analyses were conducted using GraphPad Prism 8.02 (GraphPad Software, La Jolla, CA, USA) and R statistical software (v4.4.1; R Core Team 2021). Biomarker cut-offs were derived using Cutoff Finder algorithm22. ROC analysis was used to evaluate classifier performance. A two-tailed P < 0.05 was considered statistically significant. Data visualization was performed using GraphPad Prism 8.02 for volcano plot, dot plots, bar plots, pie charts, ROC curves, and waterfall plots. Genome browser tracks were visualized using the UCSC Genome Browser (https://genome.ucsc.edu/). Heatmaps were generated in R using the pheatmap package (v1.0.12). Regulatory network analyses were performed using Ingenuity Pathway Analysis (IPA, Qiagen).
Results
Clinical characteristics of IgAN patient cohort
We analyzed cryopreserved lymphocytes from the peripheral blood of 17 patients with biopsy-confirmed IgAN. The late-stage group showed a lower eGFR value at the time of kidney biopsy compared to the early-stage group (Supplementary Table S5).
Identification of differential chromatin landscapes and stage-specific biomarkers
We hypothesized that variations in chromatin accessibility in CD8⁺ T cells would reflect differences in disease severity. To explore this, we conducted ATAC-Seq to compare chromatin accessibility between the groups. CD8+ T cells were isolated from PBMCs using FACS, and ATAC libraries were prepared via Tn5 transposase-mediated tagmentation. Sequencing reads were mapped to the human reference genome (hg19) and ATAC-peak area values were normalized using control peaks for robust inter-sample comparison20 (Figure 1a).
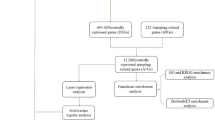
Workflow for ATAC-Seq library preparation, analysis, and biomarker selection in circulating CD8+ T cells from IgAN patients. a Schematic of the ATAC-Seq workflow. Circulating CD8+ T cells were isolated from peripheral blood mononuclear cells (PBMCs) of IgAN patients. Genomic DNA from open chromatin regions was fragmented via tagmentation using Tn5 transposase. These constructed ATAC libraries were sequenced and aligned to human reference genome (hg19). Chromatin accessibility for each sample was normalized using a factor derived from 20 control peaks. Normalized enrichment values within ATAC-peaks were converted to area values. DARs were identified by comparing early- and late-stage groups using fold change and Student’s t-test. Schematic created with BioRender.com. b Biomarker selection process. A total of 279 DARs with fold change > 2 and P < 0.05 between the early-stage (blue) and late-stage (red) group were identified (left, middle). Of these, 122 DARs with peak widths < 500 bp were selected as candidate biomarkers (right, top), including 104 early-stage group and 18 late-stage group enriched peak. Biomarkers were ranked based on relative distance and mean area values in the non-enriched group. A heatmap shows normalized area values as Z-scores ranging from ‒2 (navy) to +2 (yellow) (right, bottom).
From these data, 279 differentially accessible regions (DARs) were identified based on a fold-change > 2 and P < 0.05 from 330 accessible regions satisfying fold-change > 2 (Figure 1b, left and middle and Supplementary Table S2). Among them, 122 peaks < 500 base pair (bp) in width—representing mono- and di-nucleosome-sized ATAC-peaks—were further classified as potential stage-specific biomarkers. These were ranked based on inter-group differences in normalized peak areas (Figure 1b, right). Notably, the top-ranked biomarkers were predominantly enriched in early-stage group, whereas late-stage-specific peaks became apparent at lower ranks (after rank 31), suggesting stage-dependent regulatory signatures in CD8+ T cells (Figure 1b, right).
Chromatin landscape analysis reveals stage-specific regulatory signatures
The chromatin profiles were characterized in more detail. A total of 234 DARs were enriched in early-stage group, while 45 peaks were enriched in late-stage group, indicating higher overall chromatin accessibility in early-stage group (Figure 2a). Additionally, the generalized linear model confirmed the robustness of the identified DARs (Supplementary Table S3). Clustering using t-distributed stochastic neighbor embedding (t-SNE) based on normalized ATAC-peak area values revealed clear segregation of early- and late-stage groups (Figure 2b). Genomic annotation of DARs showed that 81.7% were located within gene bodies or promoter regions, underscoring potential impacts on transcriptional regulation (Figure 2c). The findings suggest a more accessible epigenetic landscape in early-stage IgAN and highlight the potential of chromatin accessibility signatures to reflect early IgAN, offering insight into underlying immune mechanisms before overt renal function decline.
Characterization of DARs in early and late-stage groups. a Volcano plot of 234 peaks enriched in the early-stage group (blue) and 45 peaks in the late-stage group (red) with fold change > 2 and P < 0.05. b t-SNE based on 279 DARs for 17 patients (early: blue, late: red). Each dot represents an individual patient. c Genomic annotation of the 279 DARs. Peaks were categorized into promoters, gene bodies, ncRNA, and intergenic regions. Promoters were further divided into distal (–30,000 to –10,000 bp from TSS), proximal (–10,000 to –500 bp), and core (–500 to +500 bp). Gene bodies included 5’-UTR, introns, exons, 3’-UTR, and TTS (+1,000 bp downstream from 3’-UTR). ATAC-peaks mapped to non-coding RNA regions were annotated as ncRNA. The other ATAC-peaks were grouped as intergenic regions.
Transcription factor motif analysis and CD8+ T-cell subset deconvolution
To investigate whether the observed DARs in CD8+ T cells reflected shifts in cellular composition and were regulated by distinct transcription factors governing differentiation, we re-estimated CD8+ T-cell subset composition using a disease-specific reference framework. This framework was derived from a single-cell RNA-sequencing dataset generated from the same IgAN patient cohort (GSE285335; data under review).
CD8+ T cells were clustered based on IL-7Rα expression and canonical differentiation markers, corresponding broadly to naïve-like + central memory (CM) and progressively differentiated effector-memory populations. Subset-specific signature genes were identified by differential expression analysis (one subset versus all others) and mapped to promoter-overlapping ATAC-peaks in the bulk dataset to construct a subset-specific signature matrix. Single-sample gene set enrichment analysis (ssGSEA) was then performed using promoter accessibility values to quantify subset enrichment in each sample (Figure 3a)26.
Deconvolution of CD8⁺ T-cell subsets and transcription factor motif analysis. a Schematic overview of the integrative analysis framework. Single-cell RNA-sequencing (scRNA-seq) data from the same IgAN patient cohort (GSE285335; data under review) were used to define disease-specific CD8+ T-cell subsets based on IL-7Rα expression and canonical differentiation markers. Subset signature genes were identified by differential expression analysis and mapped to promoter regions overlapping ATAC-peaks in the bulk dataset. Promoter accessibility values were converted to Z-score, and single-sample gene set enrichment analysis (ssGSEA) was performed to quantify subset-specific enrichment across individual ATAC-seq samples. Schematic created with BioRender.com. b Heatmap showing the scaled enrichment scores of four CD8+ T-cell subsets (Naïve-like+CM, IL-7Rαhigh EM, IL-7Rαmed EM, IL-7Rαlow EM) across early-stage and late-stage IgAN samples.c Comparison of ssGSEA enrichment scores for the Naïve+CM (top) and IL-7Rαlow EM (bottom) subsets between early-stage (blue) and late-stage (red) groups. Each point represents an individual patient; bars indicate mean ± SEM. Statistical significance was determined using two-tailed Student’s t-test. d Top 5 enriched Homer transcription factor motifs in the early-stage (left) and late-stage (right) groups, with consensus sequences derived from motif-matched genomic region of DARs.
Early-stage groups showed enrichment of naïve-like+CM CD8⁺ T cells, while IL-7Rαlow EM CD8⁺ T cells were more prevalent in late-stage groups (Figures 3b and c). Given that IL-7Rα expression on CD8+ T cells is known to decrease with aging27, epigenetic changes associated with advanced status of IgAN may be particularly prominent within terminally differentiated subset in the late-stages of the disease. In contrast, the enrichment of naïve-like+CM CD8⁺ T cells in early-stage IgAN suggests a more immunologically active and developmentally plastic state, potentially reflecting ongoing immune activation and early pathogenic processes.
We then conducted transcription factor (TF) motif analysis on stage-enriched DARs. Both groups showed enrichment for RUNX1, a key regulator of T cell homeostasis28. In the early-stage group, motifs for ETS1, LEF1, FLI1, and RUNX2 were enriched—TFs associated with T cell survival, proliferation, and memory formation (Figure 3d, left). Downstream targets of these TFs included cytotoxic effectors (GZMB and PRF1), apoptotic regulators (MYC), and differentiation markers (IL7R and TBX21), indicating active immune regulation in early-stage patients. These transcriptomic analyses collectively suggest that the early-stage group exhibits a predominance of cytotoxic CD8+ T cells, characterized by enhanced apoptotic potential and differentiation capabilities (Supplementary Figure S1a).
In contrast, late-stage DARs were enriched for motifs of EOMES, TBX21, IRF4, and GABPA (Figure 3d, right). These TFs are associated with terminal differentiation and T cell exhaustion in chronic immune activation29,30,31,32. Downstream targets included PRF1, PDCD1, and IL2RB, suggesting effector and inhibitory signatures in late-stage CD8⁺ T cells. Together, these results highlight distinct transcriptional regulatory networks between disease stages, with the late stage characterized by terminal differentiation and activation-induced exhaustion (Supplementary Figure S1b).
Diagnostic performance of chromatin accessibility biomarkers
To evaluate the diagnostic potential of stage-specific DARs, receiver operating characteristic (ROC) curve analysis was conducted to determine optimal cut-off values based on normalized area distributions. Optimal thresholds were selected by minimizing the Euclidean distance from the ideal point (100% sensitivity, 0% specificity), ensuring balanced discrimination between early- and late-stage groups (Figure 4a).
Evaluation of biomarker performance distinguishing early- and late-stage groups. a Schematic of biomarker assessment process. Area values representing chromatin openness were compared between groups. Cut-off values were determined using Euclidean distance in ROC analysis. Waterfall plots visualize area values after cut-off adjustment; filled and empty circles indicate patients above or below the cut-off, respectively. b, d Representative genome browser tracks of the top 10 biomarkers enriched in early-stage (b) and late-stage groups (d). c, e Classification of patients above (black circle) and below (open circle) the cut-off for each biomarker in the early-stage (c) and late-stage (e) groups. Sensitivity and specificity are color-coded from 50% (blue) to 100% (red).
The top 10 biomarkers from each group were selected. In early-stage group, seven biomarkers were located in gene bodies, one in a promoter, and two in intergenic regions (Figure 4b), achieving an average sensitivity of 98.3% (range: 83.3 – 100.0%) and specificity of 93.6% (range: 81.8 – 100.0%) (Figure 4c, Supplementary Figure S2). In late-stage group, five biomarkers each were located in gene bodies and intergenic regions (Figure 4d), with average sensitivity of 85.0% (range: 66.7 – 100.0%) and specificity of 89.1% (range: 72.7 – 100.0%) (Figure 4e, Supplementary Figure S3). Collectively, these results underscore the robustness and discriminative potential of stage-specific chromatin accessibility biomarkers in circulating CD8⁺ T cells for distinguishing IgAN patients according to the progressed status of the disease.
Composite biomarker scoring enhancing discriminatory power
To enhance stratification, we generated a composite weighted score by summing scores of individual biomarkers exceeding their respective cut-off values, as previously described 20 (Figures 4a and 5a). The combination of the top 10 biomarkers from each group revealed statistically significant differences between the early- and late-stage (P < 0.001 for both composite scores) groups. This composite score effectively distinguished early- and late-stage patients with AUROCs of 1.000 and 0.970, respectively. Sensitivity and specificity reached 100% and 100% in the early-stage group, and 100% and 90.9% in the late-stage group, respectively (Figures 5b and c). These results emphasize the value of integrating multiple chromatin biomarkers into a single composite score for sensitively distinguishing stage-specific disease severity based on CD8⁺ T-cell chromatin profiles.
Prediction of chromatin openness using composite biomarker. a Schematic of weighted openness score calculation. For each biomarker, normalized values above or below the cut-off were scored as 1 or 0, respectively. Weighted sums were calculated across markers. b, c Openness scores for early-stage (blue) and late-stage (red) groups are shown as dot plots (left), with mean ± SEM for each group (early: n = 11, late: n = 6). Statistical significance was determined by Student’s t-test. ROC curves (middle) and waterfall plots (right) visualize predictive performance and individual openness scores.
Exploratory ATAC-qPCR assessment of ATAC-Seq derived biomarkers
Given the practical challenges of implementing ATAC-Seq in clinical settings—including cost, time, and technical complexity—we developed ATAC-qPCR as a streamlined and feasible alternative for quantifying chromatin accessibility at selected biomarker loci. This assay targets amplicons centered at the summits of ATAC-Seq peaks, which are regions enriched in accessible chromatin, using primers designed to span these open chromatin regions (Figure 6a). Chromatin accessibility was quantified using the ΔCt method, with normalization to the geometric mean of three reference loci.
Practical quantification of chromatin openness by ATAC-qPCR a Schematic of the ATAC-qPCR workflow. Amplicons centered on ATAC-peak summits (gold) were designed for qPCR quantification of chromatin openness (sky blue). Forward and reverse primers span accessible regions. Normalized chromatin openness was quantified using the 2–Ct method, normalized to the geometric mean of three control loci. Schematic created with BioRender.com. b Scatter plot showing correlation (Pearson’s correlation coefficient R2) between ATAC-peak area values and normalized chromatin openness for early (blue) and late (red) biomarkers. Linear regression (green) with 95% confidence interval (dashed lines) visualizes the correlation strength.
The ATAC-qPCR results showed a significant correlation with ATAC-Seq peak intensities, as demonstrated by the relationship between ΔCt values and ATAC-Seq peak areas for both early- and late-stage biomarker groups (R2 = 0.246 – 0.389) (Figure 6b). Notably, when assessed for clinical utility in discriminating disease stages, several early-stage biomarkers (n = 5) demonstrated good predictive accuracy (AUROC = 0.772 – 0.893), while late-stage biomarkers (n = 4) also exhibited high performance (AUROC = 0.848 – 0.924) (Supplementary Figures S4 and S5)33. These findings highlight the potential of ATAC-qPCR as a clinically translatable method for targeted assessment of chromatin accessibility at specific loci, offering a practical tool for evaluating the disease status of IgAN.
Discussion
In this study, we comprehensively profiled the chromatin accessibility landscapes of circulating CD8⁺ T cells in IgAN and identified distinct epigenetic signatures associated with different disease severity using ATAC-Seq. Through comparative analysis of early- and late-stage, we demonstrated 279 DARs, which were further characterized by distinct patterns of chromatin openness, immune cell subset composition, and transcription factor motif enrichment. These stage-specific chromatin accessibility patterns revealed chromatin accessibility signatures associated with disease severity and offered their utility as discriminatory tools for disease status stratification.
Circulating CD8⁺ T cells play diverse pathogenic roles across autoimmune diseases through cytotoxic activity and aberrant differentiation,34 as seen in Graves’ disease35, aplastic anemia, and systemic lupus erythematosus36,37. This functional heterogeneity is likely governed by disease-specific transcriptional and epigenetic regulation. In IgAN, our chromatin accessibility profiling of circulating CD8⁺ T cells revealed a phenotypic shift from naïve subsets in early-stage disease to terminally differentiated EMRA cells in late-stage disease. This phenotypic evolution is consistent with previous studies demonstrating increased activated CD8⁺ T-cell populations correlating with renal function deterioration12 and enhanced tubulointerstitial CD8⁺ T cell infiltration associated with disease progression13. The epigenetic profiles of circulating CD8⁺ T cells provide mechanistic insight into their transition from active effectors in early disease to dysfunctional or exhausted states in advanced IgAN. These dynamic changes support their potential as biomarkers for capturing immune state transitions and stratifying disease severity.
The early-stage group exhibited a greater number of DARs, predominantly located in promoters and gene bodies, suggesting heightened transcriptional activity and functional regulatory potential. Deconvolution analysis revealed a transition in subpopulations, from naïve cells in early-stage to EMRA cells in late-stage group, indicative of immune quiescence or exhaustion. These cellular shifts were accompanied by distinct transcription factor motif enrichments: early-stage group showed elevated ETS1 and LEF1 activity, regulators of memory formation, apoptosis, and proliferation38, while late-stage group were enriched for EOMES and TBX21 motifs, associated with terminal differentiation and chronic activation. Correspondingly, ETS1 and LEF1 targeted genes involved in cytotoxic function and differentiation (e.g., MYC, GZMB, and PRDM1), while ETS1, RUNX1, and FLI1 co-regulated IL7R and ADGRG1, markers of naïve or cytotoxic potential. In contrast, EOMES, TBX21, IRF4, and GABPA coordinated expression of effector, exhaustion, and metabolic genes (e.g., PDCD1, IL2RB, GATA3, and TCF7), underscoring transcriptional reprogramming of CD8+ T cells along the course of progression to advanced disease.
Importantly, ETS1 binding motifs were predominantly enriched in open chromatin regions of CD8⁺ T cells of early-stage group, suggesting a potential regulatory role in T-cell activation and differentiation during the early phase of IgAN. ETS1 has previously been identified as a key transcriptional regulator of IL-7Rα expression, which is critical for T-cell homeostasis39. ETS1 has also emerged as a significant susceptibility locus in GWAS of IgAN, raising the possibility that ETS1 may contribute to disease pathogenesis through immune-mediated mechanisms beyond its known role in mesangial cells. While the precise cellular targets remain to be clarified, our findings support a potential role for ETS1 in shaping the chromatin landscape of peripheral CD8⁺ T cells. Further studies are warranted to evaluate ETS1 as a biomarker of disease activity and to explore its relevance in modulating CD8+ T cell-mediated kidney injury, as a therapeutic target.
The practical applicability of our findings was demonstrated by the successful exploration of selected DARs using ATAC-qPCR, a cost-effective and clinically feasible method. Several biomarkers showed strong discriminatory performance, and a composite scoring approach further enhanced classification accuracy. These results highlight the translational potential of integrating chromatin accessibility biomarkers into clinical risk assessment for IgAN. Moreover, our study underscores the broader utility of peripheral immune cell epigenetic profiling as a non-invasive tool for identifying patients at higher risk of progression to CKD. By distinguishing early-stage patients with progressive potential from those with stable disease, this approach may enable earlier, more targeted interventions to prevent irreversible kidney damage. Furthermore, the consistency of DAR identification using edgeR’s implementation of the generalized linear modeling framework provides additional support for the robustness and reproducibility of our chromatin accessibility findings.
Several limitations should be considered. First, peripheral blood samples were collected at varying time points after kidney biopsy, which may introduce temporal variability in chromatin accessibility profiles and limit the ability to directly correlate pathological findings with epigenetic signatures. Second, the study cohort was composed of Korean patients, potentially limiting the generalizability of the findings to other ethnic populations. Third, although our study provides comprehensive chromatin accessibility data, it lacks matched transcriptomic information that could further validate the functional consequences of the observed epigenetic alterations. Integrating multi-omic data, including RNA sequencing and single-cell ATAC-seq, in future studies will be essential to delineate the transcriptional programs downstream of key regulatory loci and to confirm subset-specific chromatin dynamics at higher resolution. Finally, our analysis was restricted to circulating CD8⁺ T cells and therefore may not fully capture the immune cell interactions and heterogeneity within the renal microenvironment. Despite these limitations, our study provides valuable insights into systemic immune alterations associated with different disease severity in IgAN.
In conclusion, this study unveils distinct chromatin accessibility landscapes in circulating CD8+ T cells that are associated with the different stages of IgAN and elucidated the potential involvement of CD8⁺ T cells in the pathogenesis of IgAN. These findings highlight the potential of chromatin accessibility biomarkers for non-invasive disease staging and personalized management. Future investigations exploring the functional implications of these epigenetic alterations and their potential as therapeutic targets may pave the way for personalized treatment strategies in IgAN.
Data availability
The ATAC-Seq data and normalized genome browser tracks generated in this study have been deposited in the Gene Expression Omnibus (GEO) database, hosted by the National Center for Biotechnology Information (NCBI), under accession number GSE285347. The dataset is currently private, but can be accessed for peer review using the following secure token: mxutcgaadrstvap.
Change history
14 February 2026
The original online version of this Article was revised: In the original version of this Article, the Institutional Review Board (IRB) of Seoul National University Hospital (SNUH) approval number was incorrectly stated as “IRB No. 1903-128-1021”. The correct IRB number is 1903-128-1020. The original article has been corrected.
References
Rodrigues, J. C., Haas, M. & Reich, H. N. IgA nephropathy. Clin. J. Am. Soc. Nephrol. 12(4), 677–86 (2017).
Tomino, Y. Predictors of prognosis in IgA nephropathy. Kaohsiung J. Med Sci. 28(10), 517–20 (2012).
Joo, Y. S. et al. External validation of the international prediction tool in Korean patients with immunoglobulin a nephropathy. Kidney Res. Clin. Pract. 41(5), 556–66 (2022).
Yao, X. et al. MicroRNAs in IgA nephropathy. Ren Fail. 43(1), 1298–310 (2021).
Zhang, Y., Yang, H., Jiang, M. & Nie, X. Exploring the pathogenesis and treatment of IgA nephropathy based on epigenetics. Epigenomics. 15(19), 1017–26 (2023).
Shema, E., Bernstein, B. E. & Buenrostro, J. D. Single-cell and single-molecule epigenomics to uncover genome regulation at unprecedented resolution. Nat. Genet. 51(1), 19–25 (2019).
Spitz, F. & Furlong, E. E. Transcription factors: from enhancer binding to developmental control. Nat. Rev. Genet. 13(9), 613–26 (2012).
Ahmed, N. et al. Multi-omic analysis of longitudinal acute myeloid leukemia patient samples reveals potential prognostic markers linked to disease progression. Front. Genet. 15, 1442539 (2024).
Araki, Y., Fann, M., Wersto, R. & Weng, N. P. Histone acetylation facilitates rapid and robust memory CD8 T cell response through differential expression of effector molecules (eomesodermin and its targets: perforin and granzyme B). J. Immunol. 180(12), 8102–8 (2008).
Ma, S. et al. Chromatin potential identified by shared single-cell profiling of RNA and chromatin. Cell. 183(4), 1103–16 (2020).
Xie, S. et al. T cell responses in immune-mediated IgA nephropathy. J. Leukoc. Biol. 116(3), 523–35 (2024).
Ruszkowski, J. et al. T cells in IgA nephropathy: role in pathogenesis, clinical significance and potential therapeutic target. Clin. Exp. Nephrol. 23(3), 291–303 (2019).
Pei, G. et al. Renal interstitial infiltration and tertiary lymphoid organ neogenesis in IgA nephropathy. Clin. J. Am. Soc. Nephrol. 9(2), 255–64 (2014).
Zheng, Y. et al. Single-cell transcriptomics reveal immune mechanisms of the onset and progression of IgA nephropathy. Cell Rep. 33(12), 108525 (2020).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods. 14(10), 959–62 (2017).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods. 9(4), 357–9 (2012).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics. 26(6), 841–2 (2010).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9(9), R137 (2008).
Consortium EP. An integrated encyclopedia of DNA elements in the human genome. Nature. 489(7414), 57–74 (2012).
Shin, H. M. et al. Chromatin accessibility of circulating CD8(+) T cells predicts treatment response to PD-1 blockade in patients with gastric cancer. Nat. Commun. 12(1), 975 (2021).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 26(1), 139–40 (2010).
Budczies, J. et al. Cutoff Finder: a comprehensive and straightforward Web application enabling rapid biomarker cutoff optimization. PLoS One. 7(12), e51862 (2012).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell. 38(4), 576–89 (2010).
Ivanek OBaR. seqLogo: Sequence logos for DNA sequence alignments. (2023).
Kramer, A., Green, J., Pollard, J. Jr. & Tugendreich, S. Causal analysis approaches in ingenuity pathway analysis. Bioinformatics. 30(4), 523–30 (2014).
Yi, M., Nissley, D. V., McCormick, F. & Stephens, R. M. ssGSEA score-based Ras dependency indexes derived from gene expression data reveal potential Ras addiction mechanisms with possible clinical implications. Sci. Rep. 10(1), 10258 (2020).
Wertheimer, A. M. et al. Aging and cytomegalovirus infection differentially and jointly affect distinct circulating T cell subsets in humans. J. Immunol. 192(5), 2143–55 (2014).
Mizutani, S. et al. Loss of RUNX1/AML1 arginine-methylation impairs peripheral T cell homeostasis. Br. J. Haematol. 170(6), 859–73 (2015).
Doering, T. A. et al. Network analysis reveals centrally connected genes and pathways involved in CD8+ T cell exhaustion versus memory. Immunity. 37(6), 1130–44 (2012).
Paley, M. A. et al. Progenitor and terminal subsets of CD8+ T cells cooperate to contain chronic viral infection. Science. 338(6111), 1220–5 (2012).
Man, K. et al. Transcription factor IRF4 promotes CD8(+) T cell exhaustion and limits the development of memory-like T cells during chronic infection. Immunity. 47(6), 1129–41 (2017).
Chandele, A. et al. Formation of IL-7Ralphahigh and IL-7Ralphalow CD8 T cells during infection is regulated by the opposing functions of GABPalpha and Gfi-1. J. Immunol. 180(8), 5309–19 (2008).
Simundic, A.-M. Diagnostic accuracy—part 1: basic concepts: sensitivity and specificity, ROC analysis STARD statement. Point Care. 11(1), 6–8 (2012).
Deng, Q. et al. The emerging epigenetic role of CD8+T cells in autoimmune diseases: a systematic review. Front. Immunol. 10, 856 (2019).
Uchimura, K. et al. The effects of CD40- and interleukin (IL-4)-activated CD23+ cells on the production of IL-10 by mononuclear cells in Graves’ disease: the role of CD8+ cells. Clin. Exp. Immunol. 128(2), 308–12 (2002).
Xing, L. et al. CD8+HLA-DR+ T cells are increased in patients with severe aplastic anemia. Mol. Med. Rep. 10(3), 1252–8 (2014).
Blanco, P. et al. Increase in activated CD8+ T lymphocytes expressing perforin and granzyme B correlates with disease activity in patients with systemic lupus erythematosus. Arthritis Rheum. 52(1), 201–11 (2005).
Jung, S. & Baek, J. H. The potential of T cell factor 1 in sustaining CD8(+) T lymphocyte-directed anti-tumor immunity. Cancers (Basel). 13(3), 515 (2021).
Grenningloh, R. et al. Ets-1 maintains IL-7 receptor expression in peripheral T cells. J. Immunol. 186(2), 969–76 (2011).
Acknowledgments
The biospecimens and data used for this study were provided by the Biobank of Seoul National University Hospital, a member of the Korea Biobank Network (project No. 2024ER050800). We thank Ms. Jiyoung Park (Core Research Facilities of Seoul National University College of Medicine) for sorting live CD8+ T cells. The schematic illustration was created with Biorender.com.
Funding
The study was supported in part by a National Research Foundation of Korea (NRF) grant funded by the Korean government (MSIT, Ministry of Science and ICT) (2021R1A2C2094586), Seoul National University Hospital Research Fund (0320213070), cooperative research fund from the Korean Society of Nephrology 2024 to D.K. Kim, and NRF grant RS-2021-NR056559 and RS-2023-NR076598 to H.-R. Kim.
Author information
Authors and Affiliations
Consortia
Contributions
H.-R.K., H.M.S. and D.K.K conceived and designed the project. G.K., S.L, S.L., S.S., S.P., J.H.K., S.C., Y.K., I.S.K., S.J.L., C.S., D.-S.L., D.K.K., H.-R.K., and H.M.S. analyzed the data. S.L, G.K., I.S.K, D.K.K., H.-R. K and H.M.S. wrote the manuscript with the help of all authors. D.K.K., H.-R. K and H.M.S. had full access to all data in the study and took responsibility for the integrity of the data, as well as for the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kim, G., Lee, S., Lee, S. et al. Chromatin accessibility of circulating CD8⁺ T cells differentiates disease severity in IgA nephropathy. Sci Rep 16, 1700 (2026). https://doi.org/10.1038/s41598-025-31213-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-31213-9