Abstract
The large yellow croaker (Larimichthys crocea) is a cornerstone of China’s mariculture. This study integrates transcriptomic, proteomic, and metabolomic analyses to determine hypoxia adaptation mechanisms in an anti-flowing F1 generation (FDTL) compared with non-selected (FDCL) counterparts under 24-h hypoxic stress (2.0 mg/L dissolved oxygen). FDTL exhibited higher survival (67% vs. 42%). Transcriptomic analysis identified 852 differentially expressed genes, with significant enriched pathways including hypoxia-inducible factor signaling, glycolysis/gluconeogenesis, and IL-17-mediated immunity. Metabolomic profiling revealed 463 differential metabolites, predominantly associated with glycerophospholipid metabolism, arachidonic acid metabolism, and VEGF signaling. Proteomic screening detected 388 differentially abundant proteins, uniquely enriched in the cytokine-cytokine receptor interaction pathway. Cross-omics integration uncovered 37 shared pathways, with VEGF, GnRH, and C-type lectin receptor signaling pathways being co-regulated at the transcriptomic-metabolomic level. Notably, Core glycolysis-related genes and hypoxia-inducible factor-associated genes were markedly downregulated. This study confirmed that the anti-flowing strain exhibits a lower oxygen threshold for metabolic reprogramming, enabling sustained aerobic metabolic homeostasis under reduced oxygen levels. The integration of immune regulation and angiogenesis establishes a multi-layered hypoxia resistance network, providing molecular targets for breeding stress-tolerant fish. These findings highlight the FDTL’s superior adaptability to high-density offshore aquaculture and validate the effectiveness of targeted breeding strategies.
Similar content being viewed by others
Introduction
The large yellow croaker (Larimichthys crocea), locally known as “huanghua” or “yellow fish,” is China’s most extensively farmed marine fish species, with an annual production of approximately 292,615 metric tons in 20241. Traditional coastal farming has long been the primary aquaculture model. However, this approach faces mounting challenges: (1) escalating land-based pollution and microplastic accumulation in nearshore waters2,3; (2) climate change-driven declines in the dissolved oxygen caused by rising sea temperatures4,5; and (3) ecological pressures from intensive farming practices, including antibiotic contamination and microbial proliferation6,7. To address these issues, offshore aquaculture has emerged as a strategic alternative. This approach leverages expansive open-ocean areas with nutrient-rich waters, high tidal exchange rates, and lower pollutant levels, enabling sustainable high-quality production when integrated with modern aquaculture equipment and advanced husbandry techniques8. However, offshore operations face formidable challenges, including strong waves, harsh marine conditions, and frequent typhoon impacts9. Furthermore, offshore aquaculture operates under high-density farming conditions, imposing stringent demands on the hypoxia tolerance of large yellow croaker, thus necessitating the development of specialized current-resistant strains optimized for these demanding environments10.
Dissolved oxygen (DO), referring to oxygen molecules dissolved in water, serves as a critical environmental determinant for fish survival11. Natural seawater typically maintains DO concentrations of 6.5–7.0 mg/L, with levels below 2.0 mg/L being classified as hypoxic12. Prolonged hypoxia in large yellow croaker triggers reactive oxygen species (ROS) accumulation, inducing oxidative stress and causing organ and cellular damage13,14. Through evolutionary adaptation, this species has developed multifaceted strategies to counter hypoxia. These include activating hypoxia-tolerant gene expression—such as stabilizing hypoxia-inducible factors (HIFs) that bind hypoxia response elements to regulate downstream targets including erythropoietin and vascular endothelial growth factor15—and rewiring energy metabolism via upregulation of hepatic glycolysis genes16. Morphological adaptations involve increasing the gill lamellar surface area and spacing to enhance oxygen uptake, while behavioral adjustments include accelerated ventilation rates or activation of accessory respiratory structures. Biochemical responses feature elevated antioxidant defenses, including superoxide dismutase (SOD), catalase (CAT), and glutathione (GSH) synthesis17,18. The effects of hypoxia have been extensively studied in aquaculture species such as large yellow croaker15,19,20, turbot (Scophthalmus maximus)21,22, and Atlantic salmon (Salmo salar)23,24,25,26,27, revealing both conserved and species-specific adaptation mechanisms (such as HIF pathway activation and gill remodeling). Such comparative insights advance our understanding of evolutionary adaptations to oxygen fluctuations, while informing targeted breeding programs for resilient aquaculture stocks.
Omics technologies, including transcriptomics, proteomics, and metabolomics, have allowed the determination of the intrinsic molecular mechanisms governing biological processes in large yellow croaker, offering critical insights into its adaptation to environmental stressors. Recent applications include transcriptome-metabolome analyses of hydrodynamic impacts28,29, integrated transcriptomic-proteomic investigations of the effects of acidification on larvae20, transcriptomic explorations of salinity adaptation30,31,32, and hypoxia-responsive gene profiling15,33. The liver, recognized as the metabolic hub and a stress-sensitive organ in aquatic species34, has served as a central research anchor for hypoxia adaptation studies across multiple species, including cobia (Rachycentron canadum)35, rainbow trout (Oncorhynchus mykiss)36, and large yellow croaker37,38. Although transcriptomics has identified hypoxia-responsive metabolic genes in the croaker liver16, single-omics approaches remain insufficient to comprehensively unravel the intricate regulatory networks governing hypoxia adaptation. Integrated multi-omics enables comprehensive profiling of genes, proteins, and metabolites, enhancing cross-validation and mechanistic clarity. For example, combined transcriptomic and metabolomic analyses revealed cold tolerance mechanisms in catfish through lipid metabolism adjustments39, proteome-metabolome integration identified key pathways governing feed efficiency in Chinook salmon (Oncorhynchus tshawytscha)40, and dual-omics studies revealed molecular adaptations in silver pomfret (Pampus argenteus) midgut physiology following jellyfish predation41. Despite these advances, no study has systematically applied transcriptomic, proteomic, and metabolomic integration to investigate hepatic hypoxia responses in current-resistant strains of L. crocea.
This study employed integrated transcriptomic, proteomic, and metabolomic analyses to compare the molecular responses to hypoxic stress between selectively bred current-resistant, anti-flowing F1 generation (FDTL), and conventional (FDCL) strains of large yellow croaker. By identifying distinct gene expression patterns, protein interactions, and metabolic shifts, we aimed to pinpoint critical regulators of current resistance and streamline the discovery of adaptive pathways. The results will establish a comprehensive biological database to evaluate stress tolerance in novel breeding lines, directly informing precision breeding strategies for offshore aquaculture systems exposed to dynamic marine conditions.
Results
The hypoxic tolerance test of anti-flowing F1 generation
As depicted in Fig. 1 survival curves, both FDTL and FDCL groups exhibited progressively declining survival rates with prolonged hypoxic exposure. Remarkably, the FDTL cohort maintained 67% survival after 24 h, whereas the FDCL group showed only 42% survival. These results demonstrate a significant association between large yellow croaker strain and survival under 24-h acute hypoxia (P < 0.001), with the current-tolerant strain (FDTL) displaying substantially higher viability than the non-selected strain (FDCL).
Survival curves under 24 h of hypoxic stress. FDTL, anti-flowing F1 generation; FDCL, non-selected control.
Transcriptome analysis
Illumina deep sequencing was carried out on the cDNA libraries constructed from the liver tissues of FDTL and FDCL. The number of original sequencing reads obtained and the number of reads obtained after quality control were counted. The details of the statistical analysis and comparison of the sequencing data are shown in Supplementary Table 1 and Supplementary Figs. 1–10.
Heatmap visualization demonstrated distinct gene expression differences between the groups, while high intra-group correlations indicated good stability of biological replicates within experimental and control groups, confirming appropriate sample grouping (Fig. 2A). Principal component analysis (PCA) analysis revealed tight clustering of intra-group samples with clear separation between the two strains across biological replicates. No outliers or aberrant sample distribution patterns were observed in the PCA plot (Fig. 2C).
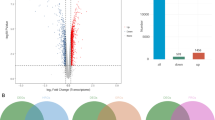
Transcriptome sample correlation heat map (A), differential volcano map (B), PCA analysis map (C), KEGG enrichment analysis. Left: Sankey diagram; Right: Bubble map (D). FDTL, anti-flowing F1 generation; FDCL, non-selected control; PCA, principal component analysis; KEGG, Kyoto Encyclopedia of Genes and Genomes.
A total of 852 significant DEGs were identified, comprising 534 upregulated and 318 downregulated genes. The volcano plot (Fig. 2B) visually depicts the distribution and magnitude of intergroup differential expression, highlighting both the quantity of gene variations and the directional patterns of regulation.
KEGG analysis of liver DEGs under low oxygen conditions revealed significant enrichment in key biological pathways (Fig. 2D). The Sankey diagram depicts relationships between enriched pathways and associated DEGs, while the bubble plot quantifies their enrichment magnitude. The top enriched pathways included: The HIF-1 signaling pathway, the nuclear factor-kappa B (NF-κB) signaling pathway, Rheumatoid arthritis, Bile secretion, Central carbon metabolism in cancer, Glycolysis/Gluconeogenesis, African trypanosomiasis, the tumor necrosis factor (TNF) signaling pathway, Fructose and mannose metabolism, the Glucagon signaling pathway, the interleukin 17 (IL-17) signaling pathway, Starch and sucrose metabolism, Lipid and atherosclerosis, Epstein-Barr virus infection, Bladder cancer, Transcriptional misregulation in cancer, Amoebiasis, Pathways in cancer, Thyroid hormone signaling pathway, and Porphyrin metabolism.
Metabolome analysis
The heatmap demonstrated distinct metabolic differences between groups, while showing strong intra-group correlations (Fig. 3A), indicating effective group separation, with no deviant samples in either the experimental or control groups. PCA revealed tight intra-group clustering and clear inter-group separation of samples (Fig. 3B–C). No outliers were observed, further confirming the robustness of the grouping strategy. Further analysis of hepatic data from the FDTL and FDCL groups demonstrated complete separation in sample distribution (Fig. 3D–E), indicating distinct metabolic profiles between hypoxia-exposed anti-flowing and non-selected strains. PLS-DA model validation revealed an intercept > 0.05, with declining R2 and Q2 values as permutation retention decreased. The upward regression trend confirmed successful permutation testing, validating a robust model free from overfitting and suitable for predictive analysis (Fig. 3F–G). DEM screening identified 463 compounds, including 157 upregulated and 306 downregulated metabolites, visualized in a volcano plot (Fig. 3H).
Metabolome sample correlation heat map (A); PCA analysis charts (B, C); PLS-DA scatter diagram (D, E) and validation diagram corresponding to PLS-DA (F, G); Differential volcano map (H); Multidimensional enrichment cycle diagram of KEGG pathway enrichment analysis (I); KEGG enrichment bubble diagram (J). Metabolite negative data (B, D, F) and positive data (C, E, G). FDTL, anti-flowing F1 generation; FDCL, non-selected control; PCA, principal component analysis; KEGG, Kyoto Encyclopedia of Genes and Genomes; PLS-DA, partial least squares discriminant analysis.
KEGG pathway enrichment analysis was performed on the DEMs. The results mainly focused on glycerophospholipid metabolism, arachidonic acid metabolism, linoleic acid metabolism, steroid hormone biosynthesis, neuroactive ligand-receptor interaction, sphingolipid metabolism, the gonadotropin-releasing hormone (GnRH) signaling pathway, the vascular endothelial growth factor (VEGF) signaling pathway, the C-type lectin receptor signaling pathway, vascular smooth muscle contraction, galactose metabolism, autophagy-other, ether lipid metabolism, the erbB signaling pathway, starch and sucrose metabolism, the adipocytokine signaling pathway, glycosylphosphatidylinositol (GPI)-anchor biosynthesis, cellular senescence, the mitogen activated protein kinase (MAPK) signaling pathway, and biosynthesis of unsaturated fatty acids (P < 0.01) (Fig. 3J).
Among them, the levels of DEMs in pathways such as glycerophospholipid metabolism, arachidonic acid metabolism, linoleic acid metabolism, steroid hormone biosynthesis, and neuroactive ligand-receptor interaction showed an overall downward trend. In contrast, the levels of DEMs in the sphingolipid metabolism pathway showed an overall upward trend (Fig. 3I).
Proteomic analysis
The heatmap demonstrated distinct proteomic differences between the groups, while showing strong intra-group correlations (Fig. 4A), indicating good biological reproducibility within both the experimental and control groups, thereby validating the grouping strategy for subsequent analyses. PCA revealed tight clustering of biological replicates within each group and clear separation between the two strains (Fig. 4B). No outliers were observed, confirming significant differences in protein expression between anti-flowing and regular strains.
Proteome sample correlation heat map (A); PCA analysis charts (B); Differential volcano map (C); KEGG enrichment analysis. Left: Sankey diagram; Right: Bubble map (D). FDTL, anti-flowing F1 generation; FDCL, non-selected control; PCA, principal component analysis; KEGG, Kyoto Encyclopedia of Genes and Genomes.
We identified 388 significant DAPs in the FDTL_vs_FDCL comparison. Specifically, 177 DAPs were upregulated while 211 DAPs were downregulated. The volcano plot illustrates this differential expression pattern (Fig. 4C). Pathway analysis identified four significantly enriched pathways (P < 0.05): cytokine-cytokine receptor interaction, phenylalanine metabolism, glycosphingolipid biosynthesis—globo and isoglobo series, and calcium signaling pathway. Detailed enrichment results are presented in Fig. 4D, where the bubble plot (right panel) quantifies pathway enrichment levels, while the Sankey diagram (left panel) visualizes associated proteins. Notably, the calcium signaling pathway featured cardiac phospholamban, receptor tyrosine-protein kinase erbB-4 isoform X1, two-pore calcium channel protein 1 isoform X2, sarcoplasmic/endoplasmic reticulum calcium ATPase 1 isoform X1, and 1-phosphatidylinositol 4,5-bisphosphate phosphodiesterase gamma-1.The glycosphingolipid biosynthesis pathway included N-acetyllactosaminide alpha-1,3-galactosyltransferase-like isoform X1 and alpha-N-acetylgalactosaminidase isoform X1. Phenylalanine metabolism encompassed macrophage migration inhibitory factor, aromatic L-amino acid decarboxylase, and L-amino acid oxidase. The cytokine-cytokine receptor interaction pathway contained interleukin-1β, C–C motif chemokine 20-like protein, transforming growth factor beta receptor type 2, tumor necrosis factor receptor superfamily member 10B isoform X3, and interleukin-1 receptor type 2 isoform X1.
Multi-omics analysis
An integrated analysis was performed on the 328, 61, and 103 KEGG pathways enriched in the transcriptomic, metabolomic, and proteomic datasets, respectively. Venn diagram analysis revealed 37 pathways common to all three omics datasets. Specifically, 4 pathways were shared between the proteome and metabolome, 15 between the transcriptome and metabolome, and 55 between the proteome and transcriptome. Regarding unique pathways, the proteome had 7, the metabolome had 5, and the transcriptome had 221. The cumulative total of distinct pathways reached 344, indicating a highly complex metabolic network regulated by these three omics under hypoxic stress. Notably, pathways within the overlapping regions likely represent core regulatory nodes (Fig. 5A).Joint analysis of KEGG pathways co-enriched by transcriptomic, proteomic, and metabolomic profiling of large yellow croaker (Larimichthys crocea) liver tissue after 24 h of hypoxic stress revealed adaptive mechanisms in the anti-flowing strain. These mechanisms encompassed energy metabolism regulation, coordinated hypoxic response, neuroendocrine modulation, and immune defense activation. KEGG pathway enrichment analysis indicated distinct regulatory patterns across the different omics levels (Fig. 5B–C). At the transcriptome level, significant enrichment (P < 0.05) was observed for pathways including the Vascular Endothelial Growth Factor (VEGF) signaling pathway, Gonadotropin-Releasing Hormone (GnRH) signaling pathway, C-type lectin receptor signaling pathway, galactose metabolism, glycolysis/gluconeogenesis, starch and sucrose metabolism, adipocytokine signaling pathway, Mitogen-Activated Protein Kinase (MAPK) signaling pathway, cellular senescence, adrenergic signaling in cardiomyocytes, biosynthesis of neomycin, kanamycin, and gentamicin, and drug metabolism—other enzymes. Among these, the biosynthesis of neomycin, kanamycin, and gentamicin, glycolysis/gluconeogenesis, VEGF signaling pathway, GnRH signaling pathway, C-type lectin receptor signaling pathway, and cellular senescence pathways showed highly significant enrichment (P < 0.01).
Multi-omics Venn diagram (A); KEGG enrichment histogram (B); Triomics interaction network (C); KEGG enrichment bubble plot (D); Contains the TOP 10 pathway maps of differential proteins/metabolites/genes (E). KEGG, Kyoto Encyclopedia of Genes and Genomes.
Metabolomic analysis indicated significant enrichment (P < 0.05) in multiple pathways, including vascular smooth muscle contraction, galactose metabolism, arachidonic acid metabolism, sphingolipid metabolism, glycerophospholipid metabolism, neuroactive ligand-receptor interaction, VEGF signaling pathway, GnRH signaling pathway, and C-type lectin receptor signaling pathway. Notably, sphingolipid metabolism, neuroactive ligand-receptor interaction, glycerophospholipid metabolism, VEGF signaling pathway, GnRH signaling pathway, C-type lectin receptor signaling pathway, and arachidonic acid metabolism exhibited highly significant enrichment (P < 0.01), suggesting these are key pathways influencing differences in hypoxia tolerance. The calcium signaling pathway was significantly enriched (P < 0.05) only in the proteomic data, suggesting its involvement in the molecular regulatory network related to hypoxia tolerance potentially through post-translational modifications or dynamic protein expression differences.
Multi-omics integration analysis (Fig. 5D, highlighted pathways) revealed that galactose metabolism, VEGF signaling pathway, GnRH signaling pathway, and C-type lectin receptor signaling pathway were co-significantly enriched (P < 0.05) in both the transcriptome and metabolome. This suggests that these pathways may represent key molecular mechanisms regulating the observed phenotypic differences in hypoxia tolerance. Furthermore, the VEGF signaling pathway, GnRH signaling pathway, and C-type lectin receptor signaling pathway reached highly significant levels (P < 0.01) in this integrated analysis (Fig. 5D).
Statistical analysis and visualization were performed on the top 10 Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways involving the highest numbers of differentially abundant proteins (DAPs), genes (DEGs), and metabolites (DEMs) identified in this study (Fig. 5E). In the transcriptome, prominent pathways were Pathways in cancer, PI3K-Akt signaling, and HIF-1 signaling. The cancer-related pathways reflect suppression of hypoxia-induced cell death via pro-survival networks in the hypoxia-tolerant strain, while HIF-1 activation drives glycolysis-related gene expression for anaerobic energy metabolism and induces angiogenesis/erythropoiesis to enhance oxygen transport. Key metabolomic pathways included glycerophospholipid metabolism, arachidonic acid metabolism, and glycine/serine/threonine metabolism. These collectively modulate membrane fluidity, generate signaling molecules, and provide precursors for antioxidant synthesis, facilitating adaptation to hypoxia-induced membrane potential shifts and oxidative stress. Significantly activated pathways included endocytosis in the proteome, which regulates internalization of membrane receptors to enhance nutrient uptake and signaling pathway activation, thereby supporting energy metabolism and hypoxia stress responses. Pathways highlighted in yellow (Fig. 5E)—cellular senescence, glycerophospholipid metabolism, arachidonic acid metabolism, neuroactive ligand-receptor interaction, and C-type lectin receptor signaling—exhibited significant co-enrichment (P < 0.05) in both transcriptomic and metabolomic datasets. This indicates their role not only as core biological processes engaging multiple molecular players but also reveals targeted, non-stochastic regulation characterized by both magnitude and statistical confidence, thereby pinpointing high-priority candidates for validating key hypoxia-tolerance mechanisms.
qRT-PCR verification
qRT-PCR analysis revealed significant downregulation of five genes (activating transcription factor 3, atf3; 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 3, pfkfb3; Krüppel-like factor 9, klf9; v-maf musculoaponeurotic fibrosarcoma oncogene homolog F, maff; and sonic hedgehog signaling molecule, shha) in the FDTL group compared to FDCL controls, while five other genes (ATPase phospholipid transporting 8A2, atp8a2; toll-like receptor 3, tlr3; signal peptide, CUB and EGF-like domain-containing protein 2, scube2; DNA damage-inducible transcript 4-like, ddit4l; and glycine amidinotransferase, gatm) exhibited significant upregulation. The expression trends of these selected genes were broadly consistent with RNA sequencing data, confirming high accuracy of the transcriptome analysis. Furthermore, correlation analysis demonstrated a highly significant linear relationship between qRT-PCR and RNA-Seq results (coefficient of determination R2 = 0.9709, P < 0.01), indicating robust reliability of the transcriptomic dataset (Fig. 6) .
Comparison of relative expression levels measured by RNA-seq and qRT-PCR. The actb gene was used as the internal control. Red bars represent qRT-PCR data and blue bars represent RNA-seq data. The vertical bar represents the mean ± S.D. (n = 3).
Discussion
Our 24-h hypoxia challenge experiment demonstrated significantly higher survival rates in the anti-flowing strain (FDTL) compared with that of conventional stocks (FDCL) (P < 0.01). Integrated analysis of transcriptomic, proteomic, and metabolomic data systematically uncovered distinct molecular responses between the strains, particularly in energy management adjustments during oxygen scarcity, activation of stress signaling pathways, rewiring of nerve-hormone communication networks, and maintenance of the immune balance. This study not only provides systematic analysis of hypoxia tolerance traits in the FDTL strain of large yellow croaker, but also demonstrates the strain’s dual stress-resistant advantages combining both current adaptation and hypoxia endurance.
Carbohydrate metabolism serves as a hallmark mechanism for hypoxia adaptation in aquatic species42.Under hypoxic stress, animals activate anaerobic glycolysis through HIF-1α-mediated metabolic reprogramming. While this adaptive strategy transiently sustains ATP production, its energy efficiency is significantly lower than that of oxidative phosphorylation43, and excessive lactate accumulation leads to pathological effects15,44. In this study, the glycolysis/gluconeogenesis pathway was significantly enriched in the FDTL group (P < 0.01), yet core glycolysis-related genes ldh (encoding lactate dehydrogenase), pk (ending pyruvate kinase ), and gapdh (encoding glyceraldehyde-3-phosphate dehydrogenase)) and hypoxia-inducible factor-associated genes were markedly downregulated. This indicated that FDTL exhibits reduced transcriptional activation of anaerobic glycolysis-related genes compared with FDCL under hypoxic conditions and suggested an optimized oxidative phosphorylation efficiency that maintains aerobic metabolic homeostasis at lower oxygen thresholds. This physiological adaptation, characterized by a reduced oxygen threshold for metabolic switching, likely underpins FDTL’s enhanced hypoxia tolerance. A parallel phenomenon was observed in Pacific oysters (Crassostrea gigas), where delayed anaerobic glycolysis activation correlated with reduced hypoxia sensitivity and higher survival rates45, aligning with FDTL’s adaptive profile. Furthermore, the upregulation of aldehyde dehydrogenase and galactokinase-related genes suggested a metabolic shift toward alternative ATP-generating pathways. By redirecting pyruvate to acetyl-CoA production, bypassing lactate generation, FDTL minimizes intracellular acidification, fuels the tricarboxylic acid (TCA) cycle, and mitigates lactic acidosis. Such metabolic rewiring aligns with cross-species adaptations: juvenile turbots leverage galactose metabolism to sustain their growth under hypoxia46 while tilapia (Oreochromis niloticus) shifts toward lipid metabolism by suppressing glycolysis/insulin signaling and enhancing unsaturated fatty acid synthesis42. Similarly, golden pompano (Trachinotus blochii) initially activates anaerobic glycolysis but transitions to lipid-dominated energy production as hypoxia persists47, a pattern mirrored in rainbow trout (Oncorhynchus mykiss), where lipid metabolism increases under severe hypoxia48. These comparative insights underscore a conserved evolutionary strategy, prioritizing metabolic flexibility to balance energy demands and cellular homeostasis during oxygen deprivation.
Beyond energy production, lipids play critical roles in maintaining membrane integrity and antioxidant defense during fish hypoxia adaptation49,50. Triploid crucian carp exposed to saline-alkaline stress exhibited similar sphingolipid-driven apoptotic regulation through lysophospholipid upregulation51, while symbiotic sphingolipids enhanced inflammatory responses in rainbow trout52. This regulatory pattern finds parallels in our findings, where coordinated lipid rewiring balanced structural stability and oxidative protection under hypoxia. The significant enrichment of sphingolipid metabolism (P < 0.01) and arachidonic acid (ARA) metabolism (P < 0.01) in our metabolomic data aligns with their dual functions in membrane fluidity adjustment and inflammatory regulation53. Notably, reduced levels of ceramide derivatives, Cer (D20:1/18:1 (12Z)-2OH (9,10) (FC = 0.0009), Cer (d18:0/Txb2), and Cer (T18:0/Pgf1α) suggest suppressed apoptotic signaling, mirroring observations in hepatic ischemia–reperfusion models where ceramide catabolism alleviates cell death54. Conversely, upregulated ARA derivatives, N-arachidonylethanolamine, N-arachidonoyl glutamic acid, and ethyl arachidonate, likely modulate the inflammatory balance via cyclooxygenase pathways, as demonstrated in Apostichopus japonicus during fungal infection55. Dietary ARA supplementation studies in crab (Scylla paramamosain), flounder (Paralichthys olivaceus), and rainbow trout (Oncorhynchus mykiss) further confirmed its role in enhancing innate immunity and antioxidant capacity56,57,58.
The HIF, VEGF, and calcium signaling pathways are critical for hypoxia adaptation in fish. In yellow croaker (Larimichthys polyactis), acute hypoxia activates the hypoxia-inducible factor HIF-1α 13,19, while a study in rainbow trout (Oncorhynchus mykiss) identified HIF-1α as a central regulatory hub59. Similarly, HIF-1 signaling is activated in hypoxia-altered metabolism of golden pompano (Trachinotus blochii) and hybrid grouper49,60. In this study, transcriptomic data showed extreme enrichment of HIF and VEGF pathways (P < 0.01). Notably, HIF-1α was significantly downregulated in the FDTL strain, a phenotype aligning with its metabolic strategy to sustain aerobic homeostasis at lower oxygen thresholds, thereby reducing reliance on HIF-mediated anaerobic metabolism. Concurrently, VEGF signaling enrichment under acute hypoxia has been reported in largemouth bass (Micropterus salmoides) and blunt snout bream (Megalobrama amblycephala)61,62, with its activation also being observed in hypoxia-tolerant hybrids of Pelteobagrus fulvidraco (♀) × Leiocassis longirostris (♂)63. Intermittent hypoxia studies in largemouth bass further revealed calcium signaling reprogramming64, mirroring the proteomic enrichment of calcium signaling (P < 0.01) observed in this study. Functional analysis linked this pathway to VEGF-induced vasodilation, exemplified by the upregulation of the store-operated calcium entry regulator STIMATE. Under 24-h acute hypoxia, molecular responses in large yellow croaker exhibited distinct temporal dynamics: the transcriptomic and metabolomic layers showed multi-pathway activation, while proteomic changes were confined to calcium signaling. This hierarchical response aligns with acute stress regulatory kinetics65, in which transcriptional and metabolic adjustments precede proteomic shifts, a pattern also observed in silver pomfret (Pampus argenteus)41. The exclusive proteomic enrichment of calcium signaling suggests its role as a hypoxia-responsive core node, operating via rapid post-translational modifications rather than transcriptional regulation. While parallels exist with vascular regulation in hypoxic rainbow trout59, the FDTL strain uniquely enhances calcium recycling through upregulated sarco/endoplasmic reticulum calcium ATPase (SERCA) pumps, ATPase sarcoplasmic/endoplasmic reticulum Ca2⁺ transporting 3, and SERCA1 1.
While GnRH signaling primarily regulates reproduction in mammals through hypothalamic-pituitary–gonadal (HPG) axis control66, its role in fish is functionally more complex. Teleost studies reveal that GnRH multiplicity, dynamic neuronal interactions, and coordination with other neuroendocrine factors collectively modulate energy allocation alongside reproduction67,68. For instance, fish prioritize energy redistribution toward hypoxia adaptation by suppressing gonadal development, and diverting lipid stores from β-oxidation-driven ATP production69,70,71 to stress-response systems, including ion transport, acid–base balance, and respiratory adjustments72. The extreme enrichment of GnRH signaling in both the transcriptomic (P < 0.01) and metabolomic (P < 0.01) data aligns with this adaptive strategy. Downregulation of pyroglutamyl-3-methylhistidyl-prolinamide, a GnRH-associated metabolite, reflects HPG axis suppression. Transcriptional repression of gonadal regulators, olliculin interacting protein 1 and proheparin-binding EGF-like growth factor, further confirms inhibited reproductive hormone (e.g., luteinizing hormone (LH)/follicle stimulating hormone (FSH)) secretion. Such energy reallocation prioritizes survival over reproduction, exemplifying hepato-brain axis coordination in hypoxia adaptation. Parallel adaptations occur in Nile tilapia (Oreochromis niloticus), where hypoxia reduces the gonadosomatic index and increases the number of atretic follicles73,74, and in yellow catfish (Pelteobagrus fulvidraco), which exhibit gonadal dysfunction under chronic hypoxia75. These findings underscore a conserved trade-off: sacrificing reproductive investment to fuel hypoxia-responsive metabolic rewiring.
Low-oxygen stress triggers oxidative damage and inflammatory responses in fish, prompting immune cells to release mediators, such as chemokines, to clear hypoxic debris and pathogens76,77. This process involves complement system activation for pathogen recognition78 and protease-driven tissue repair79. Herein, the C-type lectin receptor (CLR) pathway showed extreme enrichment (P < 0.01), marked by upregulated nuclear factor of activated T-cells (NFAT), which enhances immune cell proliferation and antibody production.
Intriguingly, NFAT simultaneously suppressed cyclooxygenase-2 (COX2), key prostaglandin synthase, resulting in reduced prostaglandin metabolites. This dual regulation prevents inflammatory overactivation, because excessive prostaglandins exacerbate tissue damage during hypoxia80. Concurrently, MAPK and PI3K pathways were activated, driving immune cell metabolism via AP-1 transcription factors and PIP3-AKT signaling81,82,83.
ROS-induced NF-κB activation (P < 0.01) upregulated interleukin-8, immunoglobulin heavy chains, and LCK tyrosine kinase, priming rapid B/T-cell responses84. This pro-inflammatory mechanism synergized with CLR’s anti-inflammatory NFAT-COX2 axis to maintain immune homeostasis, comprising a “push–pull” strategy akin to mammalian ischemia–reperfusion recovery85,86, which is newly identified in hypoxia-tolerant fish. The FDTL strain’s coordinated CLR/NF-κB crosstalk demonstrates evolutionary innovation: leveraging NFAT to fine-tune prostaglandin synthesis while amplifying MAPK/PI3K-driven immunity. This balances pathogen clearance with inflammation control, a critical adaptation to survive prolonged hypoxia without excessive tissue damage.
Materials and methods
Animals and feeding protocols
The anti-flowing strain of large yellow croaker (Larimichthys crocea) was developed through genomic selection breeding. In January 2022, we performed deep resequencing of target loci in 800 reference population individuals and 400 candidates using targeted sequencing genotyping (GBTS) technology. Genotyping data underwent genome-wide association study (GWAS) analysis and genomic selection (GS) model construction. Population structure was assessed via principal component analysis (PCA) in both cohorts, revealing a preliminary heritability estimate of 0.51 for current tolerance. Based on marker SNP (mSNP) predictions from GWAS, genomic estimated breeding values (GEBV) were calculated, with lower values indicating stronger current tolerance87. The top 60 breeders (10% selection intensity) were selected (male:female ratio = 1:3). Hormonal induction with a single hormone, luteinizing hormone-releasing hormone analog (LRH-A3), was administered twice (January 27 and 28) followed by natural fertilization. Fertilized eggs were collected on January 29–30 using the static stratification method, yielding 4.1 kg of high-quality eggs. Post-hatch progeny constituted the anti-flowing F1 population. Since March 2022, this cohort has been cultured at the offshore integrated experimental platform of the East China Sea Fisheries Research Institute’s Fuding Base (Fujian Province, China). The non-selected control strain originated from fertilized eggs produced during the same period by a cohort of 200 “Fufa No.1” broodstock (natural mating; male:female ratio = 1:3), reared under identical conditions.
In May 2024, 300 specimens (24 months old) were collected, comprising 150 flow-resistant F1 individuals (FDTL strain) and 150 non-selected individuals (FDCL strain) from the same experimental platform. Their body weight was 670.48 ± 39.52 g, and their body length was 32.4 ± 2.25 cm. They were temporarily cultured in two cement ponds at the Fuding Experimental Base. During the temporary culture period, the environmental conditions (DO: 7.5 ± 0.5 mg/L,temperature: 23 ± 0.5 °C, salinity: 26 ± 1 ‰, pH: 8.1 ± 0.2, total ammonia nitrogen 0.18 ± 0.05 mg/L, 12 h natural light and 12 h dark) were kept consistent with those during mariculture in offshore cages. The fish were fed with compound feed (Fujian Tianma Science and Technology Group Co., Ltd., Ningde, China) twice a day until they were fully satiated. The water in the ponds was changed once a day, with half of the water volume being replaced each time. The fish were fasted for 24 h before the start of the experiment.
This study was approved by the Institutional Review Board (IRB) of the Laboratory Animal Ethics Committee of East China Sea Fisheries Research Institute (LAECECSFRI-2024-0424-1). Experimental procedures complied with the ARRIVE Guidelines 2.0 and the Chinese national standard GB/T 35892-2018 on laboratory animal welfare.
Hypoxia experiments and sample collection
Prior to formal trials, a pilot study was conducted to determine critical stifling points following established protocols (Chinese National Standard GB/T 18654.8-2008), utilizing 50 fish per strain. The anti-flowing F1 strain exhibited a critical stifling point of 1.69 ± 0.08 mg/L, while the non-selected strain measured 2.01 ± 0.05 mg/L. Based on these results, 2.0 mg/L dissolved oxygen (DO), corresponding to the non-selected strain’s critical stifling point, was selected as the hypoxic stress level.
The experiment comprised two groups: the experimental group FDTL (anti-flowing F1 generation, N = 100) and control group FDCL (non-selected group, N = 100). Each group was distributed across four independent circular 3000-L fiberglass-reinforced plastic tanks with 25 fish per tank, each constituting a biological replicate. Both groups were exposed to acute hypoxic stress at 2.0 mg/L DO for 24 h according to established protocols88. Throughout the experiment, all environmental parameters were rigorously maintained at identical levels between groups: temperature 23 ± 0.5 °C, salinity 26 ± 1 ‰, pH 8.1 ± 0.2, total ammonia nitrogen 0.18 ± 0.05 mg/L, 12 h natural light and 12 h dark. Water quality parameters were continuously monitored using a multiparameter water quality analyzer (Sigma AR8606, Suzhou, China), with the sole experimental variable being strain difference.
All tanks were covered with plastic film and aerated with 99% pure nitrogen to reduce DO from normoxia (~ 7.0 mg/L) to 2.0 mg/L within one hour. A custom DO controller maintained stability at 2.0 ± 0.1 mg/L16. Mortality was recorded every 6 h (death defined as loss of equilibrium, no opercular movement for 30 s, and no tail reflex). After 24-h exposure, 2 surviving fish per tank (8 per group) were anesthetized with 100 mg/L clove oil. Following biometric measurements using a Mettler Toledo XPR2002S/AC electronic balance (Shanghai, China) for body weight and Mitutoyo CD-15CP vernier calipers (Shanghai, China) for total length and standard length, specimens were dissected on ice. Liver tissue samples were immediately extracted, flash-frozen in liquid nitrogen, and stored at − 80 °C for subsequent analysis.
Transcriptome sequencing (RNA sequencing) and analysis
Liver samples from large yellow croaker were subjected to RNA sequencing and qPCR analysis, with three biological replicates per group. For each replicate, two randomly selected liver samples were pooled in equal amounts. Total RNA was isolated using TRIzol® Reagent (Invitrogen, CA), with concentration and purity assessed on a Nanodrop2000 instrument (Thermo Fisher Scientific, Waltham, MA). High-quality RNA (RIN ≥ 6.5, OD 260/280 = 1.8–2.2, OD 260/230 ≥ 2, 28S:18S ≥ 1) was used for library construction.
Sequencing libraries were prepared using the Illumina® Stranded mRNA Prep Kit following standard protocols. Fragmented products were PCR-amplified and sequenced on the DNBSEQ-T7 platform. Raw reads were processed with Fastp (v0.23.4; https://github.com/OpenGene/fastp) to remove adapter sequences, low-quality reads (Phred score < 20), reads with > 10% N bases, and sequences shorter than 50 bp after trimming. Cleaned reads were aligned to the large yellow croaker reference genome (NCBI: GCF_000972845.2) using TopHat2 (v2.1.1; http://tophat.cbcb.umd.edu/).
Gene expression was quantified with RSEM (v1.3.3; http://deweylab.biostat.wisc.edu/rsem/), normalized as TPM (transcripts per million) to account for gene length and sequencing depth89. Mean sequencing quality scores reached 94.08% (hypoxia group) and 94.04% (control group), confirming high-quality transcriptomes. A gene coverage analysis further validated data consistency. Principal component analysis (PCA) was conducted using scikit-learn in Python to visualize sample relationships.
Differentially expressed genes (DEGs) were identified with DESeq2 using thresholds of |log₂FC|≥ 1 (≥ twofold change) and adjusted p-value (P-adj) ≤ 0.05. DEGs from FDTL and FDCL groups underwent KEGG functional annotation (http://www.genome.jp/kegg/; release 2024.01). Enrichment analysis employed Fisher’s exact test90 with Benjamini–Hochberg FDR correction91. Pathways with FDR-adjusted P ≤ 0.05 were considered significantly enriched.
Metabolomic analysis
Metabolite extraction followed the protocol of Lin et al92. For each experimental group, six replicate liver samples from large yellow croaker were processed. Aliquots of 100 mg tissue per replicate were transferred to 2 mL microtubes containing grinding beads. Metabolites were extracted using a standard solution spiked with internal standards (methanol:water = 4:1, v/v) through sequential procedures: mechanical grinding, low-temperature ultrasonication, incubation at − 20 °C for 30 min, and centrifugation (13,000 g, 15 min). The resulting supernatants were collected for instrumental analysis.
A pooled quality control (QC) sample was generated by combining equal volumes of all sample extracts. This QC sample was injected every 5–10 analytical runs to monitor system stability.Chromatographic separation was performed on a SCIEX UPLC-TripleTOF 6600 system (SCIEX, Framingham, MA; supplied by Mibio Biotechnology, Shanghai) equipped with an HSS T3 column. Mass spectrometry detection was conducted in both positive and negative ionization modes to generate LC–MS raw data files. Raw peak features were extracted from LC–MS data using Progenesis QI (Waters Corporation, Milford, MA), with processing including baseline correction, peak alignment, spectral deconvolution, and peak integration. Internal standard peaks and known false-positive peaks (e.g., noise, column bleed, and derivatization reagent artifacts) were removed from the data matrix through deduplication and peak merging. MS and MS/MS spectra were matched against public metabolite databases (HMDB: http://www.hmdb.ca/, Metlin: https://metlin.scripps.edu/, KEGG, and Majorbio Cloud) for comprehensive metabolite annotation69.
Post-annotation data were uploaded to the Majorbio Cloud Platform (https://cloud.majorbio.com) for analysis. Metabolic features detected in at least 80% of samples within any experimental group were retained. Following filtration, samples with metabolite levels below the lower limit of quantification underwent minimum metabolite value imputation, with each metabolic feature subsequently normalized. To minimize technical variability from sample preparation and instrument instability, sum normalization was applied to mass spectrometry peak response intensities, yielding normalized data matrices. Quality control samples exhibiting > 30% relative standard deviation (RSD) were excluded, and the final data matrix for downstream analysis was obtained after log10 transformation.
Preprocessed data matrices underwent multivariate statistical analysis via Principal Component Analysis (PCA) and Orthogonal Partial Least Squares-Discriminant Analysis (OPLS-DA) using the ropls package (version 1.6.2) in R. This analysis was performed on the Majorbio Cloud Platform (www.majorbio.com). Differential metabolites (DEMs) were identified based on Variable Importance in Projection (VIP) scores from the OPLS-DA model and p-values from one-way ANOVA. Metabolites with VIP > 1.0 and p-value < 0.05 were defined as DEMs. The identified DEMs were mapped to the KEGG database (http://www.genome.jp/kegg/; release 2024.01) using the Python package SciPy (https://docs.scipy.org/doc/scipy/,version 1.12.0), followed by pathway enrichment and topology analysis69.
Proteome experiment procedure
Liver samples from large yellow croaker were processed with six biological replicates per group. For each replicate, 1 g of tissue was homogenized in a grinding tube using 8 M guanidine hydrochloride buffer supplemented with protease inhibitors (solution:inhibitor = 100:1). Following ultrasonication (2 min) and incubation (30 min), lysates were centrifuged at 12,000 × g (4 °C, 30 min) to collect supernatants.
Protein concentration was determined by bicinchoninic acid (BCA) assay against a bovine serum albumin (BSA) standard curve (0–2 mg/mL). After dilution, samples were incubated with BCA reagent (37 °C, 30 min), and absorbance measured at 562 nm using a Cypher microplate reader.Aliquots containing 100 μg protein were mixed with 100 mM triethylammonium bicarbonate (TEAB) buffer. Reduction was performed with 10 mM tris(2-carboxyethyl)phosphine (TCEP; 37 °C, 60 min), followed by alkylation with 40 mM iodoacetamide (RT, dark, 40 min). Proteins were precipitated with six volumes of pre-chilled acetone (− 20 °C, 4 h) and centrifuged (10,000 × g, 20 min). Pellets were redissolved in 100 μL 100 mM TEAB and digested overnight with trypsin (1:50 enzyme:protein ratio, 37 °C). Peptides were dried under vacuum, reconstituted in 0.1% trifluoroacetic acid (TFA), desalted using HLB cartridges, and re-lyophilized. Final quantification used the Pierce™ Quantitative Colorimetric Peptide Assay Kit (Thermo Fisher Scientific).
Equal peptide amounts from all samples were pooled, concentrated, and separated into 20 fractions by high-pH reversed-phase UPLC (Thermo Vanquish F system; Waters ACQUITY UPLC BEH C18 column; mobile phase A: 2% acetonitrile pH 10 adjusted with NH₄OH, B: 80% acetonitrile pH 10; flow rate: 200 μL/min; 47-min gradient). After concentration, fractions were analyzed by data-dependent acquisition (DDA) on a Q-Exactive HF-X mass spectrometer coupled to an EASY-nLC 1200 system (C18 column; A: 0.1% formic acid in 2% acetonitrile, B: 0.1% formic acid in 80% acetonitrile; 300 nL/min; 120-min gradient). Full MS scans (m/z 350–1300, resolution 70,000) were followed by HCD fragmentation of the top 20 precursors (MS/MS resolution 17,500). Raw data were processed in Proteome Discoverer (v2.5) using a Larimichthys crocea custom database.
Individual tryptic peptides were aliquoted equally, desalted, and quantified as described prior to data-independent acquisition (DIA) analysis. Peptides dissolved in MS loading buffer were separated on a Vanquish Neo UPLC system (Thermo Fisher Scientific) and analyzed on an Orbitrap Astral mass spectrometer (Thermo Fisher Scientific). DIA raw files were imported into Spectronaut™ 19 (Biognosys AG, Schlieren, Switzerland) and searched against the NCBI database (http://www.ncbi.nlm.nih.gov/). All data were uploaded to the Majorbio Cloud Platform (cloud.majorbio.com) for subsequent analysis.
Statistical analyses were performed using R v4.0.3. Principal component analysis (PCA) was conducted on preprocessed data matrices using Python’s scikit-learn package. Differential abundance proteins (DAPs) were identified by Student’s t-test with thresholds of P < 0.05 and |log₂FC|> 1. KEGG pathway analysis (http://www.genome.jp/kegg/; release 2024.01) was performed for DAP-associated metabolic pathways. Protein–protein interaction networks were constructed using STRING v12.0 (https://string-db.org/) with default parameters.
Multi-omics analysis
Transcriptomic, metabolomic, and proteomic data from experimental (FDTL) and control (FDCL) groups were analyzed to extract KEGG pathway annotations for significantly differential genes (DEGs), metabolites (DEMs), and proteins (DAPs) (screening criteria detailed in Sects. “Transcriptome sequencing (RNA sequencing) and analysis”, “Metabolomic analysis” and “Proteome experiment procedure”). Intersections among enriched pathways across the three omics datasets were computed using the VennDiagram R package (v1.7.3), visualizing shared pathway counts and omics-specific distributions.
For co-enriched pathways (intersection set), all annotated DEGs/DEMs and DAPs were extracted. Using sample-level quantitative data (normalized expression/abundance matrices), Pearson correlation coefficients (PCC) were calculated for all gene-metabolite and protein-metabolite pairs. Each PCC underwent t-testing with Benjamini-Hochberg (BH) correction, retaining significant correlations meeting |PCC|> 0.7 and FDR < 0.05. Molecular networks were constructed with molecules as nodes and significant correlations as edges (edge weight =|PCC|; positive/negative correlations colored red/green) using igraph (v1.6.0).
To determine functional weights of key pathways, co-enriched pathways were visualized first, followed by visualization of the union of all significantly enriched pathways. Based on differential analysis results, we quantified the number of significant differential molecules per KEGG pathway (DEGs for transcriptome, DEMs for metabolome, DAPs for proteome). Pathways were ranked by molecule count (descending order), with the top 10 pathways per omics layer visualized using ggplot2 (v3.0.0).
qRT-PCR verification Protocol
Liver samples from large yellow croaker underwent quantitative real-time PCR (qRT-PCR) analysis with three biological replicates per group. Each replicate consisted of two randomly selected liver samples pooled in equal amounts. Ten key genes representing distinct functional categories were selected for transcriptome validation: ddit4l (hypoxia sensing/signaling), pfkfb3 and gatm (energy metabolism reprogramming), maff (oxidative stress defense), atf3, klf9, and atp8a2 (cellular stress/survival regulation), shha and scube2 (tissue repair/remodeling), and tlr3 (immune-inflammatory modulation), with normalization to the reference gene β-actin and Rpl893.
Total RNA (1 μg) was reverse-transcribed to cDNA using Hifair™ III Reverse Transcriptase (Yeasen Biotechnology, Shanghai). qPCR reactions (10 μL total volume) comprised 5 μL Hieff® qPCR SYBR Green Master Mix (Yeasen Biotechnology), 0.2 μM forward/reverse primers, and 4.8 μL cDNA template. All gene-specific primer sequences (Supplementary Table 2) were validated via NCBI Primer-BLAST, the specific information of the genes is provided in Supplementary Table 3. Amplification was performed on an Applied Biosystems QuantStudio 5 system (Foster City, CA, USA) under standardized conditions: initial denaturation at 95 °C for 30 s, followed by 40 cycles of 95 °C for 10 s and 60 °C for 30 s. Gene expression levels were calculated using the 2−ΔΔCT method94.
Statistical analysis
Comprehensive omics data processing methodologies are detailed in corresponding sections. Survival rates between FDTL and FDCL groups under hypoxic stress were analyzed using Cox proportional hazards regression from multivariate survival analysis. Other parametric data underwent analysis via Student’s t-test, while multi-group parametric comparisons employed one-way ANOVA followed by Tukey’s post-hoc testing. For datasets violating homogeneity of variance assumptions, nonparametric rank-sum tests were applied. Normality assumptions were formally verified for all relevant analyses. All statistical procedures were conducted using SPSS Statistics (version 25.0, IBM Corp., Armonk, NY) or GraphPad Prism 9 (GraphPad Software, San Diego, CA), with results expressed as mean ± standard deviation. Statistical significance was defined at P < 0.05 throughout this investigation.
Data availability
The raw data of RNA-seq were available in NCBI database under the accession number PRJNA1285809.
References
Ministry of Agriculture and Rural Affairs of China. China Fishery Statistical Yearbook, 22 (China Agriculture Press, Beijing, 2025).
Xu, H., Gao, Q. & Yuan, B. Analysis and identification of pollution sources of comprehensive river water quality: Evidence from two river basins in China. Ecol. Ind. 135, 108561 (2022).
Pantusa, D. Assessment of coastal vulnerability to land-based sources of pollution and its application in Apulia, Italy. Sci. Total Environ. 886, 163754 (2023).
Ma, J. The effects of seawater thermodynamic parameters on the oxygen minimum zone (OMZ) in the tropical western Pacific ocean. Mar. Poll. Bull. 187, 114579 (2023).
Bertagnolli, A. D. Microbial Niches in Marine Oxygen Minimum Zones (2018).
Duan, N. Impacts of an intensive traditional mariculture model on offshore environments as evidenced by dissolved organic matter and bacterial communities. Water Res. 267, 122530 (2024).
Hain, E. et al. Geospatial and co-occurrence analysis of antibiotics, hormones, and UV filters in the Chesapeake Bay (USA) to confirm inputs from wastewater treatment plants, septic systems, and animal feeding operations. J. Hazard. Mater. 460, 132405 (2023).
Li, S. et al. Differences in physiological performance and gut microbiota between deep-sea and coastal aquaculture of Thachinotus ovatus: A metagenomic approach. (2023).
Zhao, Y.-P., Liu, H.-F., Bi, C.-W., Cui, Y. & Guan, C.-T. Numerical study on the flow field inside and around a semi-submersible aquaculture platform. Appl. Ocean Res. 115, 102824 (2021).
Yu, J., Yan, T. & Kong, H. Exploring the management mode for breeding new species suitable for deep sea mariculture in China: Best practices, challenges, and prospects. Ocean Coast. Manag. 261, 107508 (2025).
Torrans, L., Ott, B. & Bosworth, B. Impact of minimum daily dissolved oxygen concentration on production performance of hybrid female channel catfish × male blue catfish. N. Am. J. Aquac. 77, 485–490 (2015).
Li, Y., Li, J. & Wang, Q. The effects of dissolved oxygen concentration and stocking density on growth and non-specific immunity factors in Chinese shrimp, Fenneropenaeus chinensis. Aquaculture 256, 608–616 (2006).
Wang, L. et al. Understanding the mechanisms of hypoxia-induced tissue damage in fish: The role of GasderminEa/b in Larimichthys crocea. Water Biol. Secur. 4, 100322 (2025).
Wang, X. et al. Large yellow croaker peroxiredoxin IV protect cells against oxidative damage and apoptosis. Mol. Immunol. 127, 150–156 (2020).
Chen, H. et al. The molecular mechanism under the behavioral and physiological changes of large yellow croaker (Larimichthys crocea) in response to hypoxia. Aquaculture Rep. 38, 102309 (2024).
Ding, J. et al. Transcriptome and physiology analysis identify key metabolic changes in the liver of the large yellow croaker (Larimichthys crocea) in response to acute hypoxia. Ecotoxicol. Environ. Saf. 189, 109957 (2020).
Serradell, A. et al. Gill oxidative stress protection through the use of phytogenics and galactomannan oligosaccharides as functional additives in practical diets for european sea bass (Dicentrarchus labrax) juveniles. Animals 12, 3332 (2022).
Mu, Y. et al. Transcriptome analysis reveals molecular strategies in gills and heart of large yellow croaker (Larimichthys crocea) under hypoxia stress. Fish Shellfish Immunol. 104, 304–313 (2020).
Wang, Y. et al. Transcriptome analysis reveals molecular strategies in gills and heart of large yellow croaker (Larimichthys crocea) under hypoxia stress. Vet. Sci. 9, 632 (2022).
Liu, M. et al. Proteomic and transcriptomic analysis of large yellow croaker (Larimichthys crocea) during early development under hypoxia and acidification stress. Aquaculture 577, 739982 (2023).
Wu, Z., You, F., Wen, A., Ma, D. & Zhang, P. Physiological and morphological effects of severe hypoxia, hypoxia and hyperoxia in juvenile turbot (Scophthalmus maximus L.). Aquac. Res. 47, 219 (2016).
Jia, Y., Wang, J., Gao, Y. & Huang, B. Hypoxia tolerance, hematological, and biochemical response in juvenile turbot (Scophthalmus maximus L.). Aquaculture 535, 736380 (2021).
Gamperl, A. K. et al. The impacts of increasing temperature and moderate hypoxia on the production characteristics, cardiac morphology and haematology of Atlantic Salmon (Salmo salar). Aquaculture 519, 734874 (2020).
Barnes, R. K., King, H. & Carter, C. G. Hypoxia tolerance and oxygen regulation in Atlantic salmon, Salmo salar from a Tasmanian population. Aquaculture 318, 397–401 (2011).
Remen, M., Oppedal, F., Imsland, A. K., Olsen, R. E. & Torgersen, T. Hypoxia tolerance thresholds for post-smolt Atlantic salmon: Dependency of temperature and hypoxia acclimation. Aquaculture 416–417, 41–47 (2013).
Wood, A. T., Taylor, R. S., Quezada-Rodriguez, P. R. & Wynne, J. W. Hydrogen peroxide treatment of Atlantic salmon temporarily decreases oxygen consumption but has negligible effects on hypoxia tolerance and aerobic performance. Aquaculture 540, 736676 (2021).
Wood, A. T., Clark, T. D., Elliott, N. G., Frappell, P. B. & Andrewartha, S. J. The effects of constant and cyclical hypoxia on the survival, growth and metabolic physiology of incubating Atlantic salmon (Salmo salar). Aquaculture 527, 735449 (2020).
Lv, H. et al. Integration of transcriptomics and metabolomics reveals the effects of sea currents on overwintering of large yellow croaker Larimichthys crocea in cage culture. Aquaculture 578, 740054 (2024).
Chai, R. et al. Effects of constant water flow on endurance swimming and fatigue metabolism of large yellow croaker. JMSE 11, 270 (2023).
Lu, Z. et al. Liver transcriptome analysis reveals the molecular responses to low-salinity in large yellow croaker Larimichthys crocea. Aquaculture 517, 734827 (2020).
Yang, J., Liu, M., Zhou, T., Li, Q. & Lin, Z. Transcriptome and methylome dynamics in the gills of large yellow croaker (Larimichthys crocea) during low-salinity adaption. Front. Mar. Sci. 10, 1082655 (2023).
Zhang, Y. et al. Effects of salinity stress on methylation of the liver genome and complement gene in large yellow croaker (Larimichthys crocea). Fish Shellfish Immunol. 129, 207–220 (2022).
Mu, Y., Li, W., Wu, B., Chen, J. & Chen, X. Transcriptome analysis reveals new insights into immune response to hypoxia challenge of large yellow croaker (Larimichthys crocea). Fish Shellfish Immunol. 98, 738–747 (2020).
Li, C. et al. High lipid diet can keep liver health but cause muscle oxidative stress of triploid rainbow trout treated with acute hypoxic challenge. Aquac. Rep. 40, 102575 (2025).
Huang, J. et al. Effects of hypoxia-reoxygenation conditions on serum chemistry indicators and gill and liver tissues of cobia (Rachycentron canadum). Aquac. Rep. 20, 100692 (2021).
Aksakal, E. & Ekinci, D. Effects of hypoxia and hyperoxia on growth parameters and transcription levels of growth, immune system and stress related genes in rainbow trout. Comp. Biochem. Physiol. A Mol. Integr. Physiol. 262, 111060 (2021).
Yue, Y. et al. Whole-genome methylation sequencing of large yellow croaker (Larimichthys crocea) liver under hypoxia and acidification stress. Mar. Biotechnol. 25, 567–579 (2023).
Zeng, L., Ai, C.-X., Zhang, J.-S. & Li, W.-C. Pre-hypoxia exposure inhibited copper toxicity by improving energy metabolism, antioxidant defence and mitophagy in the liver of the large yellow croaker Larimichthys crocea. Sci. Total Environ. 708, 134961 (2020).
Liu, M. et al. Comparative transcriptomes and metabolomes reveal different tolerance mechanisms to cold stress in two different catfish species. Aquaculture 560, 738543 (2022).
Esmaeili, N. et al. Proteomic investigation of liver and white muscle in efficient and inefficient Chinook salmon (Oncorhynchus tshawytscha): Fatty acid metabolism and protein turnover drive feed efficiency. Aquaculture 542, 736855 (2021).
Wang, Y. et al. Proteomic and transcriptomic analysis of the effects of dietary jellyfish on the midgut of Pampus argenteus. Aquaculture 561, 738689 (2022).
Ma, J.-L. et al. Multi-omics analysis reveals the glycolipid metabolism response mechanism in the liver of genetically improved farmed Tilapia (GIFT, Oreochromis niloticus) under hypoxia stress. BMC Genomics 22, 105 (2021).
Hand, S. C. et al. Metabolic restructuring during energy-limited states: Insights from Artemia franciscana embryos and other animals. J. Insect Physiol. 57, 584–594 (2011).
Luo, S.-Y. et al. Scavenging reactive oxygen species is a potential strategy to protect Larimichthys crocea against environmental hypoxia by mitigating oxidative stress. Zool. Res. 42, 592–605 (2021).
Meng, J., Wang, T., Li, L. & Zhang, G. Inducible variation in anaerobic energy metabolism reflects hypoxia tolerance across the intertidal and subtidal distribution of the Pacific oyster ( Crassostrea gigas ). Mar. Environ. Res. 138, 135–143 (2018).
Li, X. et al. Long-term phosphate exposure effects on juvenile turbot (Scophthalmus maximus): Growth, metabolism, and adaptive responses. Mar. Pollut. Bull. 213, 117618 (2025).
Sun, J. L. et al. Golden pompano (Trachinotus blochii) adapts to acute hypoxic stress by altering the preferred mode of energy metabolism. Aquaculture 542, 736842 (2021).
Wu, S., Huang, J., Li, Y. & Zhao, L. Comparative transcriptomics combined with physiological and functional analysis reveals the regulatory mechanism of rainbow trout (Oncorhynchus mykiss) under acute hypoxia stress. Ecotoxicol. Environ. Saf. 278, 116347 (2024).
Wang, T. et al. Acute hypoxia and reoxygenation alters glucose and lipid metabolic patterns in Hulong hybrid grouper ( Epinephelus fuscoguttatus ♀ × E. lanceolatus ♂ ). Aquaculture 586, 740827 (2024).
An, X. et al. A new strategy: Dietary lipids protected the liver from hypoxia injury in multiple ways in grass carp (Ctenopharyngodon idella). Aquaculture 560, 738523 (2022).
Yuan, F. et al. Analysis of immunosuppression and antioxidant damage in diploid and triploid crucian carp (Carassius auratus) induced by saline-alkaline environmental stress: From metabolomic insight. Metabolites 14, 721 (2024).
Sanchez, M., Sepahi, A., Casadei, E. & Salinas, I. Symbiont-derived sphingolipids regulate inflammatory responses in rainbow trout (Oncorhynchus mykiss). Aquaculture 495, 932–939 (2018).
Sissener, N. H. et al. Dietary 18: 2n–6 affects EPA (20: 5n–3) and ARA (20: 4n–6) content in cell membranes and eicosanoid production in Atlantic salmon (Salmo salar L.). Aquaculture 522, 735098 (2020).
Jiang, Y., He, X., Simonaro, C. M., Yi, B. & Schuchman, E. H. Acid ceramidase protects against hepatic ischemia/reperfusion injury by modulating sphingolipid metabolism and reducing inflammation and oxidative stress. Front. Cell Dev. Biol. 9, 633657 (2021).
Qi, H. et al. Effects of dietary arachidonic acid (ARA) on immunity, growth and fatty acids of Apostichopus japonicus. Fish Shellfish Immunol. 127, 901–909 (2022).
Fang, F. et al. Alteration of growth performance, antioxidant capacity, tissue fatty acid profiles, and lipid metabolism of mud crab (Scylla paramamosain) juvenile in response to different dietary arachidonic acid levels. Aquac. Nutr. 2022, 1–20 (2022).
Medagoda, N. & Lee, K.-J. Effects of dietary arachidonic acid supplementation in high plant protein diets on growth, feed utilization, and immunity of olive flounder, Paralichthys olivaceus. Aquaculture 571, 739431 (2023).
Hong, J., Ortiz, J. G., Sealey, W. M. & Small, B. C. Effects of dietary arachidonic acid supplementation in low fishmeal and fish oil-free diets on growth performance, inflammatory response, gut histology, and non-specific immunity in sub-adult rainbow trout Oncorhynchus mykiss. Aquaculture 580, 740272 (2024).
Wu, S. Dynamic and systemic regulatory mechanisms in rainbow trout (Oncorhynchus mykiss) in response to acute hypoxia and reoxygenation stress. Aquaculture 572, 739540 (2023).
Jiang, T. et al. Hypoxia alters glucose and lipid metabolisms in golden pompano (Trachinotus blochii). Aquaculture 562, 738747 (2023).
Gong, D. et al. Comparative analysis of liver transcriptomes associated with hypoxia tolerance in the improve blunt snout bream (Megalobrama amblycephala). Aquaculture 592, 741190 (2024).
Sun, J. L. et al. MiRNA-mRNA integration analysis reveals the regulatory roles of miRNAs in the metabolism of largemouth bass (Micropterus salmoides) livers during acute hypoxic stress. Aquaculture 526, 735362 (2020).
Wang, H. et al. The hybrid Pelteobagrus fulvidraco (♀) × Leiocassis longirostris (♂) exhibits improved trait on hypoxia-tolerance. Aquaculture 562, 738859 (2023).
Zhao, L. et al. Intermittent hypoxia promotes largemouth bassheart adaptation through increasing angiogenesis, reprogramming calcium signaling, and upregulating mitochondrial turnover. Aquaculture 579, 740195 (2024).
Zhang, Y. et al. Genetics responses to hypoxia and reoxygenation stress in Larimichthys crocea revealed via transcriptome analysis and weighted gene co-expression network. Animals 11, 3021 (2021).
Oka, Y. Neuroendocrine regulation of reproduction by GnRH neurons: Multidisciplinary studies using a small fish brain model. Endocr J 70, 343–358 (2023).
Somoza, G. M., Mechaly, A. S. & Trudeau, V. L. Kisspeptin and GnRH interactions in the reproductive brain of teleosts. Gen. Comp. Endocrinol. 298, 113568 (2020).
Zohar, Y., Zmora, N., Trudeau, V. L., Muñoz-Cueto, J. A. & Golan, M. A half century of fish gonadotropin-releasing hormones: Breaking paradigms. J. Neuroendocrinol. 34, e13069 (2022).
Zhang, F. et al. Effects of soya lecithin on the growth, gonad development, transcription of genes related with fatty acid synthesis and inflammation between sexes of sea urchin Strongylocentrotus intermedius (A. Agassiz, 1864). Aquaculture 575, 739774 (2023).
Planas, J. V., Athos, J., Goetz, F. W. & Swanson, P. Regulation of ovarian steroidogenesis in vitro by follicle-stimulating hormone and luteinizing hormone during sexual maturation in salmonid fish1. Biol. Reprod. 62, 1262–1269 (2000).
Turkmen, S. et al. Parental nutritional programming and a reminder during juvenile stage affect growth, lipid metabolism and utilisation in later developmental stages of a marine teleost, the gilthead sea bream (Sparus aurata). Br J Nutr 118, 500–512 (2017).
Wang, M. et al. Dynamic brain responses and systemic regulatory mechanisms in yellow catfish (Pelteobagrus fulvidraco) exposed to acute hypoxia. Aquaculture 599, 742177 (2025).
Ali, M. J. et al. Modulation of chronic hypoxia on ovarian structure, oxidative stress, and apoptosis in female Nile Tilapia (Oreochromis niloticus). Aquaculture 590, 741081 (2024).
Konkal, P. & Ganesh, C. B. Exposure to hypoxia inhibits pituitary-testicular activity in the fish Oreochromis mossambicus. Aquaculture 515, 734552 (2020).
Zhao, C. et al. Multi-omics analysis revealed the dysfunction of ovary and testis induced by chronic hypoxia in Pelteobagrus fulvidraco. Aquaculture 584, 740668 (2024).
Gao, F. et al. Signalling regulation of reactive oxygen species in fish inflammation. Rev. Aquac. 16, 1266–1285 (2024).
Shi, X. et al. Role of HIF in fish inflammation. Fish Shellfish Immunol. 143, 109222 (2023).
Faught, E. & Schaaf, M. J. M. Molecular mechanisms of the stress-induced regulation of the inflammatory response in fish. Gen. Comp. Endocrinol. 345, 114387 (2024).
Gómez-Abellán, V. et al. Professional phagocytic granulocyte-derived PGD2 regulates the resolution of inflammation in fish. Dev. Comp. Immunol. 52, 182–191 (2015).
Gómez-Abellán, V. & Sepulcre, M. P. The role of prostaglandins in the regulation of fish immunity. Mol. Immunol. 69, 139–145 (2016).
Deng, W., Yang, T., Dong, R., Yan, Y. & Jiang, Q. Astaxanthin protects tilapia head kidney cells against polystyrene microplastics-induced inflammation through MAPK and NF-κB signaling pathways. Aquaculture 574, 739686 (2023).
Yu, J. et al. Genome-wide identification of MKK and MAPK gene families and their expression analysis under abiotic stress in largemouth bass (Micropterus salmoides). Aquaculture 561, 738688 (2022).
Zhou, Y., Wang, Y.-Y., Fu, H.-C. & Huang, H.-Z. MicroRNA expression and analysis of immune-related putative target genes in ISKNV-infected spleen of mandarin fish (Siniperca chuatsi). Aquaculture 547, 737450 (2022).
Xie, J. et al. Study on mechanism of synthetic astaxanthin and Haematococcus pluvialis improving the growth performance and antioxidant capacity under acute hypoxia stress of golden pompano (Trachinotus ovatus) and enhancing anti-inflammatory by activating Nrf2-ARE pathway to antagonize the NF-κB pathway. Aquaculture 518, 734657 (2020).
Banaei, S. Novel role of microRNAs in renal ischemia reperfusion injury. Ren. Fail. 37, 1073–1079 (2015).
Quesnelle, K. M., Bystrom, P. V. & Toledo-Pereyra, L. H. Molecular responses to ischemia and reperfusion in the liver. Arch Toxicol 89, 651–657 (2015).
Wang, J. et al. Development and evaluation of liquid SNP array for large yellow croaker (Larimichthys crocea). Aquaculture 563, 739021 (2023).
Ding, J. et al. Genomic selection for hypoxia tolerance in large yellow croaker. Aquaculture 579, 740212 (2024).
Wagner, G. P., Kin, K. & Lynch, V. J. Measurement of mRNA abundance using RNA-seq data: RPKM measure is inconsistent among samples. Theory Biosci. 131, 281–285 (2012).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: New perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, D353–D361 (2017).
Tang, Y. et al. Mechanistic insight into the roles of Pseudomonas plecoglossicida clpV gene in host-pathogen interactions with Larimichthys crocea by dual RNA-seq. Fish Shellfish Immunol. 93, 344–353 (2019).
Lin, X. et al. Metabolomics provides insights into the alleviating effect of dietary Caulerpa lentillifera on diquat-induced oxidative damage in zebrafish (Danio rerio) liver. Aquaculture 584, 740630 (2024).
Ding, J. et al. Genome-wide association and transcriptome analysis provide the SNPs and molecular insights into the hypoxia tolerance in large yellow croaker (Larimichthys crocea). Aquaculture 573, 739547 (2023).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Funding
This research was funded by the Biological Breeding-Major Projects (2023ZD0405501), Central Public-interest Scientific Institution Basal Research Fund, CAFS (NO. 2025TD03) and Spark Project of Fujian Provincial Science and Technology Plan (2025S0062).
Author information
Authors and Affiliations
Contributions
**Bingfei Li** : Writing—review &; editing, Writing—original draft. **Yabing Wang** : Writing—review &; editing, Methodology, Conceptualization. **Guangde Qiao** : Validation. **Qiaozhen Ke** : Validation. **Xiaoshan Wang** : Resources. **Shengyu Liu** : Resources. **Kunhuang Han** : Writing—review &; editing, Project administration, Conceptualization. **Shiming Peng** : Writing—review &; editing, Resources, Funding acquisition, Conceptualization.
Declaration of generative AI
No generative AI technologies were used in any aspect of this manuscript’s preparation. All content was solely created by the human authors, who take full responsibility for the scientific integrity, data accuracy, and intellectual contributions of this work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Glossary
Glossary
Catalase (CAT): Enzyme that decomposes hydrogen peroxide into water and oxygen.
Data-independent acquisition (DIA): Mass spectrometry method fragmenting all ions in predefined m/z ranges.
Differentially abundant proteins (DAPs): Proteins showing statistically significant abundance changes between experimental groups.
Differentially expressed genes (DEGs): Genes with statistically significant expression differences between conditions.
Differentially expressed metabolites (DEMs): Metabolites with statistically significant concentration variations between groups.
Dissolved oxygen (DO): Concentration of oxygen molecules (O₂) dissolved in water (mg/L).
False discovery rate (FDR): Statistical threshold controlling expected proportion of false positives in multiple testing.
Fold change (FC): Ratio of expression/abundance levels between experimental and control groups.
Glutathione (GSH): Tripeptide antioxidant that scavenges free radicals and maintains redox balance.
Hypoxia-inducible factors (HIFs): Transcription factors activating gene expression in response to low oxygen.
KEGG pathway: Kyoto Encyclopedia of Genes and Genomes database for biological pathway annotation.
Liquid chromatography-mass spectrometry (LC–MS): Analytical technique combining separation and detection of molecules.
Multi-omics integration: Combined analysis of transcriptomic, proteomic, and metabolomic datasets.
Orthogonal partial least squares-discriminant analysis (OPLS-DA): Multivariate method for identifying group-discriminating variables.
Principal component analysis (PCA): Unsupervised dimensionality reduction method visualizing sample clustering.
Protein–protein interaction (PPI): Functional/physical connections between proteins within biological networks.
qRT-PCR: Quantitative reverse transcription polymerase chain reaction for gene expression measurement.
Reactive oxygen species (ROS): Chemically reactive oxygen-containing molecules causing oxidative damage.
Superoxide dismutase (SOD): Enzyme catalyzing superoxide radicals into oxygen and hydrogen peroxide.
Variable importance in projection (VIP): Metric quantifying a variable’s contribution to group separation in OPLS-DA.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, B., Wang, Y., Qiao, G. et al. Multi-omics analysis reveals hypoxia adaptation mechanisms in an anti-flowing F1 generation of large yellow croaker (Larimichthys crocea). Sci Rep 16, 2100 (2026). https://doi.org/10.1038/s41598-025-31844-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-31844-y