Abstract
Learning and memory, as fundamental components of human cognition, are heritable traits that are highly variable between individuals and within populations. Investigation into the genetic basis of cognition is a prominent area of research, with genetic associations being previously reported for a wide range of cognitive phenotypes. Here we utilise a genome-wide association study (GWAS) approach to evaluate the contribution of genetic variation to learning and memory phenotypes in a comprehensively phenotyped, well-characterised, healthy, and unrelated cohort of individuals (n = 613). Cognitive phenotypes were assessed using nine comprehensive test batteries consisting of twenty-one cognitive performance assessments including IQ, five measures for visual and verbal learning, and fifteen measures for semantic, working, episodic and prospective memory. Principal component analysis was utilised to amalgamate correlated test scores into additional new cognitive phenotypes. Our study identified genome wide significant associations for 13 loci across all phenotypes. A novel association was identified between the rs817826 SNP at 9q31.2 and verbal learning discrimination (p = 2.71 × 10− 9). GWAS of cognitive PCs identified three variants in the vicinity of thiamine (Vitamin B1) transporter gene SLC19A3 (most significant SNP rs12105620, p = 2.17 × 10− 9), a 3’ UTR variant in PPARD (rs9658167, p = 1.47 × 10− 8), and an intronic variant in RBFOX1 (rs17138790, p = 4.24 × 10− 8) associated with the cognitive PC related to visual and verbal learning. The cognitive PC relating to prospective and retrospective memory revealed a locus containing a synonymous variant in NXPE3 (rs2305990, p = 6.56 × 10− 9) and intronic variants in RD3 (rs17189035, p = 2.71 × 10− 8) and WLS/GNG12-AS1 (rs17130484, p = 4.13 × 10− 8). Pathway analysis identified olfactory, vitamin A, and cadherin pathways as being significantly overrepresented across multiple cognitive domains. The novel associations identified provide candidates for further investigation and necessitate replication in similarly characterised independent cohorts.
Similar content being viewed by others
Introduction
Memory is a cornerstone of cognitive function, acting as a sophisticated process that integrates sensory inputs and experiences to facilitate adaptive responses to information and stimuli. The fundamentals of memory are shaped by an intricate interplay of genetic and molecular factors, which over time have become more implicit as more is revealed about the foundation of human cognition. Cognition can be stratified into a number of domains – Sachdev et al. (2014) identify learning and memory, language, perceptual function, executive function, attention, and social cognition as six major neurocognitive domains1. Developing the understanding of these domains within a molecular genetic context may provide translatable insights into the treatment of disease-induced impairment, as well as personalised health insights for otherwise healthy individuals.
Genetic variability has been long understood to play a pivotal role in cognitive function, with large-scale studies using publicly available genetic datasets being undertaken. Cognition is a complex trait and is highly heritable, with studies indicating substantial genetic influences accounting for more than 80% of variability2. As genetic factors become more evident and datasets more comprehensive, genome-wide association studies (GWAS) have continued to implicate novel loci to explain genetic variability in cognitive differences3,4,5. A recent GWAS meta-analysis by Savage et al. comprising of 269,867 individuals has most recently identified 205 distinct genomic loci associated with a multidimensional set of cognitive performance tests6. Another large study of 300,486 individuals identified 709 genes and 148 independent loci associated with general cognitive function7. GWAS with larger sample sizes are required to continue to identify novel loci for investigation; however, smaller cohort studies with accurately reported phenotypes and greater uniformity in sample collection and processing methods can provide useful insight into the polygenic nature of cognition8,9,10.
Previous studies have evaluated the association of specific measures and batteries of tests with genetic variability, while neglecting to consider the relationships between tests and phenotypes. Principal component analysis (PCA) can be utilised to constellate sets of correlated variables into a set of new uncorrelated variables – principal components (PCs). These PCs retain the fundamental patterns of the grouped variables while reducing the dimensionality of the data, discarding the less relevant features, and revealing trends that may not be initially perceived11,12,13. This process has been utilised in numerous GWAS to highlight features that would have otherwise not been discernible5,7,14, and can provide significant insight into the identification of genetic variability within a cohort.
In this study, we investigated associations between genetic variants and cognitive measures in a healthy cohort, quantified using measures of intelligence, learning, and memory using a representative battery of standardised and validated tests. To uncover novel evidence implicating genetic variability in cognitive function, GWAS were performed for each individual test of a specific memory type, as well as for the principal components of cognitive measures. Genes and pathways related to known molecular processes and brain functions were implicated in this study, providing insights into the nature of cognition and informing future research. The deep phenotyping of cognition, using multiple standardised tests, and subsequent PCA of these measures supports PCA as a valid data reduction method in GWAS for correlated cognitive phenotypes.
Methods
Study population and cognitive phenotypes
The study was approved by both the Griffith University (MSC/01/09/HREC) and Queensland University of Technology (1300000486) Human Research and Ethics Committees and all experiments were performed in accordance with relevant guidelines and regulations. A cohort of 619 healthy participants were recruited from the Southeast Queensland region of Australia, principally the Brisbane and Gold Coast areas, through public advertisement. There were no age or education criteria, and participants with a history of psychiatric disorder or head injury were excluded to preserve the cohort’s capacity as a representative sample of cognitive and memory without additional confounding factors. All subjects provided written informed consent prior to collection and testing. Saliva samples were collected from all participants immediately post completion of all memory testing for genetic analysis using Oragene DNA Self Collection kits (DNA Genotek Inc., Ottawa, ON, Canada) with DNA extraction performed as per manufacturers protocol.
Memory, learning, and intelligence phenotypes were evaluated using eight cognitive battery tests as presented in Table 1 and as previously described15,16,17. All participants were assessed in a quiet, well-lit room by the same examiner, with a total of 21 measures recorded for all participants (Table 1). Principal component analysis (PCA) was applied to these 21 measures to transform the data into cognitive PCs using R (4.3.1) packages factoextra18 and FactoMineR19, with the first 3 PCs selected for further analysis after identifying the inflection point of the scree plot of the eigenvalues.
Genotyping and quality control
Background information was reviewed by the study team for participants to confirm that all met inclusion criteria as listed above, with six individuals being removed from the study for not satisfying the criteria. The remaining 613 samples were genotyped on Illumina Human OmniExpress-24 BeadChip Arrays (v1.0 and v1.1) (Illumina, Inc., San Diego, CA, USA). Only the overlapping SNPs in both arrays were included in the study (709,354 SNPs) to maintain a high genotyping rate. Four samples were excluded post genotyping due to the overall genotyping call rate being less than 95%.
Samples and SNPs were quality controlled and filtered using PLINK (v1.90b7)30, based on minor allele frequency > 0.01, Hardy-Weinberg equilibrium p-value > 0.001, genotyping call-rate > 0.95, and failure rate per-individual < 0.05. After quality control procedures, 598 individuals and 624,763 SNPs were retained. One individual recorded a missing WASI IQ measure – population characteristics were analysed with the full remaining cohort of 598 individuals, with the GWAS being performed with this individual excluded for a total of 597 individuals.
After quality control, population structure characteristics were analysed using PLINK (v1.90b7) and R packages factoextra, FactoMineR, and visualised using ggplot231. Population characterisation, performed using PCA, was used to elucidate the genetic substructure of the population and compared to self-reported ethnicity, the latter being utilised as a covariate in the association analysis. Correction for population substructure is essential in GWAS analysis to control for the structure of the cohort, and although self-reported ethnicity can assist in the identification and stratification of the population structure, it can be optimal to utilise the PCs of the allele frequencies as covariates to correct for heterogeneity in lieu of ethnicity11. The first 3 PCs were included as covariates after identifying the inflection point of the eigenvalues for the PCs of genetic substructure using the scree plot. Imputation was not considered for this dataset as the study cohort comprised of a range of ethnicities that are not included in the most widely used reference panels, reducing the probability of correct haplotype matches.
Genome-wide association analyses
Genomic inflation and association analyses were performed using PLINK (v1.90b7). An additive linear model to test association was fitted to each individual phenotype and to the PCs of the cognitive measures, with sex, age, IQ, and the first 3 PCs of the genetic substructure applied as covariates. For the WASI IQ and cognitive PC association analyses IQ was not included as a covariate as to not confound the analysis of these phenotypes. To account for multiple testing the genome wide significance threshold was set at the widely accepted p < 5 × 10− 8, with a suggestive threshold set at p < 1 × 10− 5.
Replication cohorts
To validate the findings, results were compared with two replication cohorts: one consisting of a subset of the QIMR Twin cohort32 and the second being the Genetics of Brain Structure and Function (GOBS) study cohort33,34. In the QIMR Twin cohort, intelligence, semantic memory, and working memory scores was assessed in a cohort of Australian dizygotic and monozygotic twins of European ancestry using the same tests with the current study cohort; WASI IQ, WAIS INFO, and WAIS LNST respectively. Intelligence and semantic memory results were obtained from 2598 individuals and working memory results from for 956 individuals. Genotyping was performed using a combination of Illumina 317 K, Illumina 370 K, and Illumina 610 K microarray chips and GWAS was performed using METAL for 7,681,669 markers. In the GOBS study, 1709 participants of Mexican and American ancestry were genotyped using Illumina Human1Mv1, Human1M-Duov3, HumanHap550v3, and HumanExon510Sv1 BeadChips as previously described35. Each participant completing a battery of tests consisting of standardised measures36, with phenotypes equivalent to those assessed in this study compared for suggestively significant variants. Only data related to the associations with the cognitive measures, not the cognitive PC’s, was available for this cohort. A significance level of P < 0.05 was used for the replication analysis given the small sample size of the replication cohorts.
Pathway analysis
Pathway analysis was performed on the linear association outputs by collating all 624,763 variants into SNP-sets using PLINK. Loci were mapped to corresponding genes by genomic location using the hg19 reference genome. A 50 kb window around each gene was defined, an approach previously described to include SNPs within intergenic regulatory regions neighbouring genes, without sacrificing substantial specificity37,38,39. This set-based method was selected in lieu of tools that use a reference genome and calculate linkage disequilibrium (LD) against a reference dataset as the study population is un-imputed and of mixed ethnicity. Set-based methods allow for the consideration of multiple SNPs within a gene that alone may not be highly significant but could have greater combined genetic effects40. The set-based p-values for each gene were calculated using PLINK adaptive permutation, pruning SNPs in high LD, and the outputs subsequently analysed using the R package ActivePathways41 to produce overrepresented pathways. Gene Ontology (GO)42, Panther43, and Reactome44 databases were queried, and pathways were filtered to those within the range of 10 to 200 genes to minimise bias related to size and increase specificity in the pathway analysis; this range being well documented in previous studies39,45.
Results
Study population characteristics
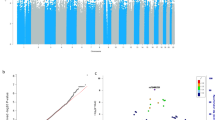
Summary statistics were generated for the 598 participants with cohort demographics presented in Table 2. The majority of participants were female (71.4%) with a median age of 20 (Mean age 23, SD 8.0) and an age range of 16–65. One individual recording a missing WASI_IQ measure was excluded from downstream analysis. Genomic inflation was calculated as 1.02 overall for the 21 measures and 1.00 for the 3 cognitive PCs in the multivariate models (QQ plots provided in Supplementary Fig. 1). The population substructure (Supplementary Fig. 2) demonstrated that self-reported ethnicity did not distinctly cluster and indicated unreported mixed ethnicity for some individuals, supporting the use of genetic PCs in the stratification of the population structure.
Principal component analysis of cognitive phenotypes
Twenty-one measures were transformed using PCA, with underlying variation illustrated in Fig. 1. The first 3 cognitive PCs explained 52% of the phenotype variance (Figs. 1 A and B). Cognitive PC1 explained 20% of the variance, with positive loading evident for HVLT and SVLT test batteries (0.33 to 0.74), representing visual and verbal learning (Figs. 1 A and C). Cognitive PC2 explained 17% of variance, with high positive loadings (0.74 to 0.94) observed for CAPM and PRMQ test batteries, representing prospective and retrospective memory (Figs. 1A and D), while PC3 explained 15%, with negative loading for SVLT (−0.63 to −0.72) and positive loading for HVLT (0.30 to 0.57) test scores (Figs. 1A and E).
PCA of cognitive measures. A: Summary of the first 3 PCs for the 21 cognitive measures. B: Scree plot of eigenvalues depicting variance of explained by PCs of cognitive measures, inflection point denoted by red dashed line. C: Loading plot of PC1 and PC2 of 21 cognitive measures to identify which measures have the largest effects of each component. D: Loading plot of PC2 and PC3. E: Loading plot of PC1 and PC3.
Genome-wide association results
Linear regression analysis was completed for 624,763 autosomal SNPs in a cohort of 597 healthy individuals adjusted for sex, age, IQ, and the first 3 PCs of the genetic substructure. For the WASI IQ and cognitive PC association analyses IQ was not included as a covariate. The multi-covariate additive model was fitted for each memory phenotype and the PCs of the phenotypes. In total, 13 SNPs passed the genome wide significance threshold of p < 5 × 10− 8 (Table 3), and 289 SNPs passed the suggestive threshold (p < 1 × 10− 5) across all 24 phenotypes (Supplementary Table 1). Manhattan plots of the association results for all phenotypes that show genome wide significant SNPs are presented in Figs. 2 and 3. One SNP reached the genome-wide significance threshold for the cognitive measures, rs817826, significantly associated with the HOP_DISCRIM measure for verbal learning (p = 2.71 × 10− 9), with the remaining 12 SNPs being significant for cognitive PC1 and cognitive PC2. No SNPs were significant at p < 5 × 10− 8 for PC3. Manhattan plots for the association results with no genome-wide significant SNPs are presented in Supplementary Figs. 3 and 4.
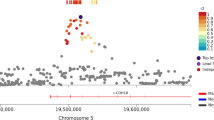
Manhattan plot for the HOP DISCRIM measure, with significant SNP rs817826 marked (p = 2.71 × 10− 9). The genome wide significance threshold (p < 5 × 10− 8) is indicated by the red line. The blue line indicates the suggestive threshold (p < 1 × 10− 5).
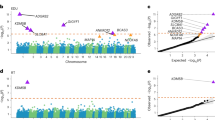
Manhattan plots of the GWAS results of the PCA of the cognitive measures, with loci reaching the genome wide significance threshold (p < 5 × 10− 8) indicated by the red line. The blue line indicates the suggestive threshold (p < 1 × 10− 5). Significant SNPS are denoted in Table 3. A: Plot of PC1 extracted from the cognitive measures. B: Plot of PC2 extracted from the cognitive measures.
Replication analysis of genome-wide association results in two independent cohorts
Replication analysis of genome-wide and suggestively significant (p < 1 × 10− 5) associations was undertaken in two independent cohorts with similar cognitive measures: the QIMR Twin cohort consisting of Australian twins of European ancestry, and the GOBS cohort consisting of Mexican-American pedigrees. Summary statistics were assessed for same or similar measures where available. Correlation between suggestively significant (p < 1 × 10− 5) SNPs in the GRC GOM cohort and nominal significant SNPSs (p < 0.05) the QIMR Twin cohort identified one locus, rs9948718, located downstream from MAPK4, as being associated with working memory in the QIMR Twin cohort and GRC GOM cohort (Table 4). Nominal associations were also identified between cognitive PCs and WASI_IQ, WAIS_LNST, and WAIS_INFO results. In the GOBS cohort, proximate SNPs rs1032704 and rs4931672 at 12p11.21 were nominally significant for the Penn Conditional Exclusion Test (PCET) Face Memory Test and reached the suggestively significant threshold in the GRC GOM cohort for the SVLT LI measure for visual learning (Table 5). Variant rs6035340, an intronic variant in SLC24A3, was nominally significant for the California Verbal Learning Test (CVLT) Delay Test and reached the suggestively significant threshold in the GRC GOM cohort for prospective memory measured by the MIST DELAY test. Replication data for the genome-wide significant SNPs is included in Supplementary Table 2. Of note, two SNPs, rs17189035 and rs17130484, located within the introns of the RD3 and WLS/GNG12-AS1 genes respectively, reached the genome wide significance threshold (p < 5 × 10− 8) in the GRC GOM cohort for cognitive PC2 and p < 0.05 in the QIMR Twin cohort.
Gene-set based pathways analysis
To assess if individual loci were related to specific genes and pathways, gene-set pathways analysis was performed. Loci were mapped to genes, assessed for over-representation against cognitive phenotypes, and the resulting gene-sets analysed for enrichment against pathways. The pathway analysis results identified the most significantly associated pathways (p < 0.1) for the gene-sets for each phenotype. Figure 4 A summarises the cumulative number of significant pathways for each phenotype, with Figs. 4B and C illustrating the most significant pathways for each cognitive domain and for cognitive PCs 1–3. An average of 18 significant (Holm–Bonferroni corrected p < 0.1) pathways from the three databases were identified for each cognitive test (range: 2 to 37), and an average of 12 per cognitive PC (range: 6 to 24). The most significant pathways were identified to be related to cellular responses to ions, namely copper (GO:0071280, p = 3.93 × 10− 14), cadmium (GO:0071276, p = 3.07 × 10− 13), and zinc (GO:0071294, p = 1.80 × 10− 12) in the CAPM TOT test phenotype representing prospective memory. As for other test batteries, cadherin signalling (PANTHER PATHWAY: P00012, p = 2.63 × 10− 10) and sensory perception of taste (GO:0050913, p = 2.90 × 10− 5), and odorant binding (GO:0005549, p = 1.59 × 10− 3) pathways were identified to be overrepresented in HVLT outcomes, glucuronidation (GO:0052695, p = 1.02 × 10− 13), uronic acid (GO:0006063, p = 2.59 × 10− 13), retinoic acid (GO:0001972, p = 1.74 × 10− 10), and cadherin signalling (PANTHER PATHWAY: P00012, p = 7.17 × 10− 9) pathways identified in PRMQ test phenotypes, and growth hormone receptor binding (GO:0005131, p = 1.62 × 10− 9) identified in SVLT outcomes. Additional pathway results are presented in Supplementary Fig. 5 and Supplementary Table 3.
Results for Gene Ontology (GO), Panther, and Reactome pathway analysis across all phenotypes. All reported p-values are adjusted by Holm–Bonferroni correction. A: Cumulative count of significant (p < 0.1) terms for all phenotypes. B: Top significant terms for each cognitive domain C: Top significant terms for each cognitive PC extracted from the cognitive measures.
Discussion
The evolving landscape of genetic factors in cognitive research is marked by novel findings in both large-scale population-based studies and in smaller cohort studies, both providing insight into the complex phenotypes of memory and cognition. In this current study, we have performed a GWAS in a cohort of healthy individuals, who were extensively phenotyped for a range of cognitive traits, to discover genetic variants involved in intelligence, learning, and memory. As well as individual test scores, PCA was also utilised to extract underlying latent patterns in the data, allowing for more comprehensive, muti-phenotype analysis of the dataset. Results revealed that 13 SNPs passed the genome wide significance threshold of p < 5 × 10− 8 and 289 SNPs passed the suggestive threshold of p < 1 × 10− 5 across all 24 phenotypes.
A genome-wide significant association was identified between HVLT recall score, measuring verbal memory, and variant rs817826 in the GRC GOM cohort. Polymorphism rs817826, located between the RAD23B and KLF4 genes, has been identified as being associated with prostate cancer susceptibility48,49 though a mechanism is yet to be realized. KLF4 encodes a zinc finger protein that has been identified to be critical in neural stem cell differentiation50 and the regulation of axon growth51. KLF4 has been implicated in the regulation of neuroinflammation52,53, and specifically neuroinflammation and oxidative stress in Alzheimer’s disease (AD)54, both processes affecting memory performance. Consequently, the role of rs817826 in neurological function warrants further investigation.
Interrogation of the substructure of the cognitive measures was accomplished using PCA. Data reduction via PCA is widely used in association studies and allows for comprehensive analysis of correlated phenotypes12,13, though can only be applied when all phenotypes are measured for all samples55. In this study, an increase in the number of significant loci was gained when using the multi-phenotype PCA based approach, and, interestingly, none of these loci passed the genome wide significance threshold in any individual cognitive measure. One of the most notable findings from this analysis was the identification of multiple variants in the region of Thiamine Transporter 2 gene SLC19A3 (rs12105620, rs7578840, rs7579047); rs12105620 in the 3’ UTR being associated with the first cognitive PC that features high loading of visual and verbal learning scores. SLC19A3 encodes a transmembrane thiamine (vitamin B1) transporter that is ubiquitously expressed, and primarily located in the basement membrane and perivascular cells in cerebral blood vessels56. Pathogenic variants in this gene have been implicated in basal ganglia disease and encephalopathy57,58,59, and disrupted thiamine transport linked to AD pathogenesis60,61. Thiamine deficiency impairs learning ability associated with a related impaired hippocampal neurogenesis in mice models62, and plays a major role in alcohol dependent neurocognitive disorders63. Low levels of thiamine metabolites have also been associated with decreased Mini Mental State Examination (MMSE) Recall score64. These findings support the association between thiamine levels and metabolism and cognition, and further research into the role of SLC19A3 in neuropathology and cognition is warranted.
In addition to these results, associations were observed between the first cognitive PC and variants in the 3’ UTR of PPARD (rs9658167). PPARD encodes for a peroxisome proliferator-activated receptor (PPARδ), from a class of proteins which play a role in numerous metabolic functions65, including neurogenesis66, and demonstrates widespread localisation in the brain and central nervous system, particularly in the hippocampus and hypothalamus67. PPARD has been implicated in schizophrenia68 and major depressive disorder (MDD)69, with studies in mice identifying a role for PPARδ in the proliferation and differentiation of neuronal stem cells in the hippocampus70,71 and knockout studies demonstrating loss of neuronal and synaptic structure leading to memory dysfunction72. A recent cohort study by Insel et al. identified variant rs71567499, approximately 20 kb downstream from rs9658167 and exon 8 of the PPARD gene, as being associated with lower Preclinical Alzheimer Cognitive Composite scores in an asymptomatic pre-clinical AD cohort73. The effect size of the rs71567499 variant reported in Insel et al. is similar to the effect size observed for rs9658167 in the GRC GOM cohort (β= −2.16 and β= −2.99, respectively) supporting the findings of both studies and the potential role of PPARD in memory and cognition.
Also associated with cognitive PC1 is a variant in an intron of RBFOX1 (rs17138790). RBFOX1 encodes for an RNA binding protein and splicing factor that regulates the expression of numerous genes involved in neuronal and synaptic development74,75. Meta-analyses of 232,964 cases and 494,162 controls by the Cross-Disorder Group of the Psychiatric Genomics Consortium recently identified intronic variant rs7193263 in RBFOX1, (400 kb downstream from rs17138790), as a pleiotropic locus being associated with 7 neuropsychiatric disorders, including autism spectrum disorder (ASD), schizophrenia, bipolar disorder, MDD, and attention-deficit hyperactivity disorder (ADHD)76. Lahti et al. (2022) identified a suggestive association between intronic variant rs10852681 in RBFOX1 and verbal short-term memory in a sample of 44,874 individuals3. Variants in RBFOX1 have also been associated with amyloidosis and reduced cognitive performance in AD77. A recent study by O’Leary et al. (2022) demonstrated an increase in neural reactivity to emotional stimuli, captured using functional MRI, as well as reduced prefrontal activation during cognitive control, in participants carrying variant rs650074478, located 200 kb downstream from rs17138790 identified in this current study. Another recent study identified five significant SNPs in the RBFOX1 gene in close proximity and in high LD with lead variant rs75885813, as being significantly associated with working memory impairment in a juvenile ADHD cohort79. Visual memory was assessed in the study using the Rey–Osterrieth Complex Figure (ROCF)80, which involves copying a complex geometric shape, at a 20-minute delay. This component is similar to the SVLT DRI, also requiring the recognition of visual compositions following a 20-minute delay; consequently, although utilising different cognitive tests, this reinforces the findings of this current study and further implicates variants in the RBFOX1 locus as having a role in visual learning and memory.
Additional variants were identified to be associated with cognitive PC2, featuring high loading of prospective and retrospective memory scores. One of these was a synonymous variant in NXPE3 (rs2305990), a gene encoding for a neurexophilin glycoprotein that is ubiquitously expressed in the brain81 and is involved in axonal adhesion82. NXPE3 is yet to be comprehensively investigated, though it has been implicated in adenocarcinoma of the lung83. An intronic variant in the RD3 (rs17189035) gene was also associated with cognitive PC2, and the encoded protein Retinal Degeneration Protein 3 has recently been reported to be localised to the retina84. Retinal Degeneration Protein 3 is involved in photoreceptor function85,86 and mutations in RD3 can lead to Leber congenital amaurosis, a disease that causes retinal degeneration and blindness87,88. It could be hypothesized that the association of RD3 variants with the cognitive phenotype identified in this study may be a result of its endogenous role in photoreceptor cells and retinal function, and subsequent impact on cognitive assessment, however further investigation is needed. Finally, an intronic variant rs17130484 was identified in the WLS/GNG12-AS1 locus, encoding the Wntless Wnt Ligand Secretion Mediator protein, an integral membrane transporter protein that mediates Wnt secretion89. Variants in WLS have been implicated in structural birth defects90, and Wnt signalling has been associated with hippocampal synaptic plasticity91 and memory92 in mouse models. Dysfunction of Wnt signalling has been implicated in neurological disorders including AD93,94; however, WLS is yet to be associated with cognitive function and its role in prospective and retrospective memory should be further explored.
Replication of suggestive associations in the GRC GOM cohort was performed in two cohorts, one a subset of the QIMR Twin cohort32 and the second the GOBS study cohort33,34. Variant rs9948718 was identified to nominally correlate with working memory in the GRC GOM and QIMR Twin cohorts and is located downstream of MAPK4. MAPK4 is yet to be linked to memory or cognition, although has been reported as being overexpressed in gliomas95. Variants rs1032704 and rs4931672, upstream from PKP2, were identified to be suggestively significantly associated with visual learning in the GRC GOM cohort, and nominally associated with visual memory in the GOBS cohort. These two variants are also yet to be associated with memory or cognition, however, upregulation of PKP2 has been associated with the progression of gliomas96, suggesting potential congruities between variants rs9948718, rs1032704, and rs4931672 that may warrant further investigation. Variant rs6035340, an intronic variant in SLC24A3, was suggestively significant in the GRC GOM cohort for MIST DELAY and replicated in the GOBS cohort for CVLT Delay47, associating this variant with visual learning and memory. SLC24A3 encodes a potassium-dependent sodium/calcium exchanger, highly expressed in the brain, and has been associated with abnormal motor learning, but not cognitive function, in mice97, and as a risk loci for migraine via GWAS98, implicating this gene in neurological phenotypes.
While specific variants identified as being significantly associated with cognitive phenotypes in our analysis have not been previously reported in similar studies, the genome-wide significant variants identified in this study map to several genes reported in previous cognitive GWAS. Variant rs10889966 is located upstream of NEGR1, which has been previously associated with cognitive performance in a previous GWAS99. NEGR1 has also been reported to be associated with MDD100,101 and AD102,103, with its role in neuronal development, cognition, and learning widely studied in murine models104,105,106. Similarly, variant rs17130484, significantly associated with cognitive PC2, is located in WLS GNG12-AS1 which has been associated with cognitive processing speed107, while rs17189035, located in RD3 and also significantly associated with cognitive PC2 was been previously associated with intelligence108. Differences in the cognitive phenotypes analysed between our study and similar studies, as well as the diversity of ancestry in our cohort, may both contribute to the novelty of our findings; however, these associations suggest convergent evidence at the gene level for our findings when compared to similar studies, despite variant-level novelty.
Pathway analysis using SNP sets identified the enrichment of odorant binding pathways in the WASI IQ, HVLT Delay and Retention, PRMQ RM, and SVLT RII subsets, implicating olfactory factors in the intelligence, verbal learning and memory, and retrospective memory domains. Enrichment of taste receptor pathways was also identified in HVLT Delay measures, specifically sensory perception of bitter taste. The association between odour and memory has been extensively explored, as reviewed by Saive et al. (2014)109 and Tong et al. (2014)110, with patterns of olfactory and verbally cued memory and associated brain activation identified in the evocation of autobiographical memory111. In addition, olfactory factors have been identified as being potential biomarkers for cognitive impairment with bearing on AD112,113, consolidating the associations between olfactory processes and memory. The mechanisms and relationship between taste and memory has been well documented114, with the association between taste perception and working memory also being reported115, in line with the findings in this study.
Enrichment of cadherin pathways were also identified in significant SNP sets in the HVLT discrimination and PRMQ RM measures representing verbal learning and retrospective memory, respectively. Cadherins are well understood to be critical in neuronal and synaptic architecture116,117, and have been functionally implicated in working memory and learning using mouse models118,119. Cadherins have been associated with working memory performance in an ADHD cohort120, and with neurocognitive disorders including ASD121,122, as well as being implicated in Aβ production and amyloidosis123,124. Further investigation into the role of the cadherin family of proteins in memory and cognition is justified.
Flavonoid and uronic acid metabolic processes were over enriched in the PRMQ PM, HVLT Learning, CAPM IADL, and WAIS LNST subsets, implicating these pathways in prospective memory, verbal learning, and working memory. Enrichment of these pathways in these sets are a result of several suggestively significant variants within a complex locus of uridine 5’-diphospho-glucuronosyltransferase (UGT) genes. UGTs are not ubiquitously expressed in the human brain; although, UGT1A4 has been reported to be expressed in endothelial and neuronal cells125. Glucuronate has also been associated with schizophrenia126, while uronic acid pathways have been implicated in AD and amyloidosis127. In a similar manner, enrichment of mineral ion metabolic processes in the CAPM Total test gene-sets are a result of a nominally significant 3’ UTR variant in Metallothionein 1 M gene MT1M, located in a multigenic region and in the vicinity of several other metallothionein genes. Enrichment of pathways in this manner is a limitation of the set-based method, and could be considered a trade-off when compared to other methods, such as SNP ratio tests, which cannot identify significant pathways that may contain multiple smaller effect loci and are unfit for unimputed or lower coverage data128,129.
Enrichment of the retinol metabolic process pathway in cognitive PC1 was also reported, this pathway being distinct from those associated with UGT genes. A-vitamins, namely retinol and its metabolite retinoic acid, are well established as being integral in normal cognitive function and neuroplasticity130,131. Contemporary studies have implicated vitamin A in ASD132, AD133,134,135, and cognitive function in later life136,137. These findings, notably the enrichment of retinol metabolic pathways for cognitive PC1 with high loading of visual and verbal learning scores, reinforces the proposition of a genetic link between cognition and vitamin metabolism that should be further explored.
Limitations
The small sample size is a limitation for this study when compared to large scale GWAS. Imputation was not considered due to the multi-demographic nature of the cohort and may have improved coverage. Specific significant variants identified in our current study were not replicated in similar cognitive GWAS. This is likely due to ethnicity differences in our cohorts that are not widely studied, or reported in similar studies, as well as differences in the cognitive phenotypes assessed in this study compared to similar studies. Our interrogation of the genetic substructure of our cohort indicates significant diversity of ancestry which supports this. The pathway analysis method is also sensitive to the overrepresentation of variants in multi-gene regions, and imputation may have allowed for more refined pathway analysis; however, as discussed above, it would have disputable impacts on the sensitivity of the analysis. For some of the findings in this study, there is currently insufficient evidence in the literature to explain the role of the genes identified. For the replication study, a significance level of P < 0.05 was used for analysis to avoid overly conservative replication criteria given the small sample size of the replication cohort. Further assessment in similar cohorts and via functional studies would be required to elucidate the mechanisms of the variants and genes implicated in this study.
Conclusion
Results identified 13 genome-wide significant SNPs and 289 SNPs passing the threshold of suggestive significance across all 24 phenotypes. We found a significant novel association between the rs817826 SNP located between the RAD23B and KLF4 genes and verbal learning discrimination. PCA identified three variants in the vicinity of thiamine transporter gene SLC19A3, including one within the 3’ UTR, which were associated with the cognitive PC explained by HVLT and SVLT test batteries that quantify of visual and verbal learning, respectively. This cognitive PC was also associated with a 3’ UTR variant in the PPARD gene and an intronic variant in RBFOX1. The cognitive PC with loadings dominated by CAPM and PRMQ test batteries, measuring prospective and retrospective memory, was identified to be associated with a synonymous variant in NXPE3, as well as intronic variants in RD3 and WLS/GNG12-AS1. Pathway analysis identified olfactory, vitamin A, and cadherin pathways as being notably overrepresented, exhibiting significant associations across multiple cognitive domains. These findings provide novel insights into the association between genetics and cognition in healthy individuals and provides a basis for future research.
Data availability
The GWAS summary results for all significant and suggestive SNPs are available in Supplementary Data. Full GWAS summary statistics are available in the NHGRI-EBI GWAS catalogue (https://ftp.ebi.ac.uk/pub/databases/gwas/summary_statistics/GCST90448001-GCST90449000/) under accessions GCST90448143-GCST90448166.
References
Sachdev, P. S. et al. Classifying neurocognitive disorders: the DSM-5 approach. Nat. Reviews Neurol. 10, 634–642. https://doi.org/10.1038/nrneurol.2014.181 (2014).
Hoekstra, R. A., Bartels, M. & Boomsma, D. I. Longitudinal genetic study of verbal and nonverbal IQ from early childhood to young adulthood. Learn. Individual Differences. 17, 97–114. https://doi.org/10.1016/j.lindif.2007.05.005 (2007).
Lahti, J. et al. Genome-wide meta-analyses reveal novel loci for verbal short-term memory and learning. Mol. Psychiatry. 27, 4419–4431. https://doi.org/10.1038/s41380-022-01710-8 (2022).
Davies, G. et al. Genome-wide association study of cognitive functions and educational attainment in UK biobank (N = 112 151). Mol. Psychiatry. 21, 758–767. https://doi.org/10.1038/mp.2016.45 (2016).
Davies, G. et al. Genetic contributions to variation in general cognitive function: a meta-analysis of genome-wide association studies in the CHARGE consortium (N = 53 949). Mol. Psychiatry. 20, 183–192. https://doi.org/10.1038/mp.2014.188 (2015).
Savage, J. E. et al. Genome-wide association meta-analysis in 269,867 individuals identifies new genetic and functional links to intelligence. Nat. Genet. 50, 912–919. https://doi.org/10.1038/s41588-018-0152-6 (2018).
Davies, G. et al. Study of 300,486 individuals identifies 148 independent genetic loci influencing general cognitive function. Nature Communications 9, (2098). (2018) https://doi.org/10.1038/s41467-018-04362-x
Papassotiropoulos, A. et al. A genome-wide survey of human short-term memory. Mol. Psychiatry. 16, 184–192. https://doi.org/10.1038/mp.2009.133 (2011).
Zenebe-Gete, S., Salowe, R. & O’Brien, J. M. Benefits of cohort studies in a Consortia-Dominated landscape. Front. Genet. 12, 801653. https://doi.org/10.3389/fgene.2021.801653 (2021).
Wijmenga, C. & Zhernakova, A. The importance of cohort studies in the post-GWAS era. Nat. Genet. 50, 322–328. https://doi.org/10.1038/s41588-018-0066-3 (2018).
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909. https://doi.org/10.1038/ng1847 (2006).
Schmid, P. R., Palmer, N. P., Kohane, I. S. & Berger, B. Making sense out of massive data by going beyond differential expression. Proc. Natl. Acad. Sci. U S A. 109, 5594–5599. https://doi.org/10.1073/pnas.1118792109 (2012).
Suo, C. et al. Analysis of multiple phenotypes in genome-wide genetic mapping studies. BMC Bioinform. 14, 151. https://doi.org/10.1186/1471-2105-14-151 (2013).
Tran, N. K. et al. Multi-phenotype genome-wide association studies of the Norfolk Island isolate implicate pleiotropic loci involved in chronic kidney disease. Sci. Rep. 11, 19425. https://doi.org/10.1038/s41598-021-98935-4 (2021).
Avgan, N. et al. Association Study of a Comprehensive Panel of Neuropeptide-Related Polymorphisms Suggest Potential Roles in Verbal Learning and Memory. Genes (Basel) 15. https://doi.org/10.3390/genes15010030 (2023).
Avgan, N. et al. BDNF Variants May Modulate Long-Term Visual Memory Performance in a Healthy Cohort. Int. J. Mol. Sci. 18, https://doi.org/10.3390/ijms18030655 (2017).
Avgan, N. et al. A CREB1 gene polymorphism (rs2253206) is associated with prospective memory in a healthy cohort. Front. Behav. Neurosci. 11, 86. https://doi.org/10.3389/fnbeh.2017.00086 (2017).
Kassambara, A. & Mundt, F. Factoextra: extract and visualize the results of multivariate data analyses. R package version 1.0.7. https://github.com/kassambara/factoextra (2020).
Lê, S., Josse, J., Husson, F. & FactoMineR An R package for multivariate analysis. J. Stat. Softw. 25, 1–18. https://doi.org/10.18637/jss.v025.i01 (2008).
Wechsler, D. Manual for the Wechsler Adult Intelligence Scale. San Antonio, TX: Psychological Corporation (1955).
Eadie, K. & Shum, D. Assessment of visual memory: a comparison of Chinese characters and geometric figures as stimulus materials. J. Clin. Exp. Neuropsychol. 17, 731–739. https://doi.org/10.1080/01688639508405163 (1995).
Shum, D. H., O’Gorman, J. G. & Eadie, K. Normative data for a new memory test: the shum visual learning test. Clin. Neuropsychol. 13, 121–135. https://doi.org/10.1076/clin.13.2.121.1967 (1999).
Brandt, J. The Hopkins verbal learning test: development of a new memory test with six equivalent forms. Clin. Neuropsychol. 5, 125–142. https://doi.org/10.1080/13854049108403297 (1991).
Dori, G. A. & Chelune, G. J. Education-stratified base-rate information on discrepancy scores within and between the Wechsler adult intelligence Scale–Third edition and the Wechsler memory Scale–Third edition. Psychol. Assess. 16, 146–154. https://doi.org/10.1037/1040-3590.16.2.146 (2004).
Wechsler, D. Wechsler Memory Scale, Third Edition (WMS-III), San Antonio, TX: Psychological Corporation (1997).
Crawford, J. R., Smith, G., Maylor, E. A., Della Sala, S. & Logie, R. H. The prospective and retrospective memory questionnaire (PRMQ): normative data and latent structure in a large non-clinical sample. Memory 11, 261–275. https://doi.org/10.1080/09658210244000027 (2003).
Hannon, R., Adams, P., Harrington, S., Fries-Dias, C. & Gipson, M. T. Effects of brain injury and age on prospective memory self-rating and performance. Rehabil Psychol. 40, 289 (1995).
Chau, L. T., Lee, J. B., Fleming, J., Roche, N. & Shum, D. Reliability and normative data for the comprehensive assessment of prospective memory (CAPM). Neuropsychol. Rehabil. 17, 707–722. https://doi.org/10.1080/09602010600923926 (2007).
Raskin, S. A. Memory for intentions screening test: psychometric properties and clinical evidence. Brain Impair. 10, 23–33. https://doi.org/10.1375/brim.10.1.23 (2009).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575. https://doi.org/10.1086/519795 (2007).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer-, 2016).
Wright, M. J. & Martin, N. G. Brisbane adolescent twin study: outline of study methods and research projects. Aust J. Psychol. 56, 65–78. https://doi.org/10.1080/00049530410001734865 (2004).
McKay, D. R. et al. Influence of age, sex and genetic factors on the human brain. Brain Imaging Behav. 8, 143–152. https://doi.org/10.1007/s11682-013-9277-5 (2014).
Kochunov, P. et al. Genetics of microstructure of cerebral white matter using diffusion tensor imaging. Neuroimage 53, 1109–1116. https://doi.org/10.1016/j.neuroimage.2010.01.078 (2010).
Hodgson, K. et al. Genome-wide significant loci for addiction and anxiety. Eur. Psychiatry. 36, 47–54. https://doi.org/10.1016/j.eurpsy.2016.03.004 (2016).
Knowles, E. E. et al. Genome-wide significant localization for working and Spatial memory: identifying genes for psychosis using models of cognition. Am. J. Med. Genet. B Neuropsychiatr Genet. 165b, 84–95. https://doi.org/10.1002/ajmg.b.32211 (2014).
De la Cruz, O., Wen, X., Ke, B., Song, M. & Nicolae, D. L. Gene, region and pathway level analyses in whole-genome studies. Genet. Epidemiol. 34, 222–231. https://doi.org/10.1002/gepi.20452 (2010).
Kabakchiev, B. & Silverberg, M. S. Expression quantitative trait loci analysis identifies associations between genotype and gene expression in human intestine. Gastroenterology 144, 1488–1496. https://doi.org/10.1053/j.gastro.2013.03.001 (2013). 1496.e14811483.
Pei, G. et al. Investigation of multi-trait associations using pathway-based analysis of GWAS summary statistics. BMC Genom. 20, 79. https://doi.org/10.1186/s12864-018-5373-7 (2019).
Weng, L. et al. SNP-based pathway enrichment analysis for genome-wide association studies. BMC Bioinform. 12, 99. https://doi.org/10.1186/1471-2105-12-99 (2011).
Paczkowska, M. et al. Integrative pathway enrichment analysis of multivariate omics data. Nat. Commun. 11, 735. https://doi.org/10.1038/s41467-019-13983-9 (2020).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The gene ontology consortium. Nat. Genet. 25, 25–29. https://doi.org/10.1038/75556 (2000).
Mi, H. & Thomas, P. PANTHER pathway: an ontology-based pathway database coupled with data analysis tools. Methods Mol. Biol. 563, 123–140. https://doi.org/10.1007/978-1-60761-175-2_7 (2009).
Croft, D. et al. Reactome: a database of reactions, pathways and biological processes. Nucleic Acids Res. 39, D691–D697. https://doi.org/10.1093/nar/gkq1018 (2010).
O’Dushlaine, C. et al. Psychiatric genome-wide association study analyses implicate neuronal, immune and histone pathways. Nat. Neurosci. 18, 199–209. https://doi.org/10.1038/nn.3922 (2015).
Kurtz, M. M., Ragland, J. D., Moberg, P. J. & Gur, R. C. The Penn conditional exclusion test: a new measure of executive-function with alternate forms of repeat administration. Arch. Clin. Neuropsychol. 19, 191–201. https://doi.org/10.1016/s0887-6177(03)00003-9 (2004).
Delis, D. C., Kramer, J. H., Kaplan, E. & Ober, B. A. California Verbal Learning Test, Second Edition, San Antonio, TX: Pearson Assessments (2000).
Xu, J. et al. Genome-wide association study in Chinese men identifies two new prostate cancer risk loci at 9q31.2 and 19q13.4. Nat. Genet. 44, 1231–1235. https://doi.org/10.1038/ng.2424 (2012).
Lachance, J. et al. Genetic hitchhiking and population bottlenecks contribute to prostate cancer disparities in men of African descent. Cancer Res. 78, 2432–2443. https://doi.org/10.1158/0008-5472.Can-17-1550 (2018).
Qin, S. & Zhang, C. L. Role of Kruppel-like factor 4 in neurogenesis and radial neuronal migration in the developing cerebral cortex. Mol. Cell. Biol. 32, 4297–4305. https://doi.org/10.1128/mcb.00838-12 (2012).
Moore, D. L. et al. KLF family members regulate intrinsic axon regeneration ability. Science 326, 298–301. https://doi.org/10.1126/science.1175737 (2009).
Kaushik, D. K., Gupta, M., Das, S. & Basu, A. Krüppel-like factor 4, a novel transcription factor regulates microglial activation and subsequent neuroinflammation. J. Neuroinflammation. 7, 68. https://doi.org/10.1186/1742-2094-7-68 (2010).
53 Alder, J. K. et al. Kruppel-like factor 4 is essential for inflammatory monocyte differentiation in vivo. J. Immunol. 180, 5645–5652. https://doi.org/10.4049/jimmunol.180.8.5645 (2008).
Li, L., Zi, X., Hou, D. & Tu, Q. Krüppel-like factor 4 regulates amyloid-β (Aβ)-induced neuroinflammation in alzheimer’s disease. Neurosci. Lett. 643, 131–137. https://doi.org/10.1016/j.neulet.2017.02.017 (2017).
Zhang, W. et al. PCA-Based Multiple-Trait GWAS Analysis: A Powerful Model for Exploring Pleiotropy. Anim. (Basel) 8. https://doi.org/10.3390/ani8120239 (2018).
Kevelam, S. H. et al. Exome sequencing reveals mutated SLC19A3 in patients with an early-infantile, lethal encephalopathy. Brain 136, 1534–1543. https://doi.org/10.1093/brain/awt054 (2013).
Yamada, K. et al. A wide spectrum of clinical and brain MRI findings in patients with SLC19A3mutations. BMC Med. Genet. 11, 171. https://doi.org/10.1186/1471-2350-11-171 (2010).
Serrano, M. et al. Reversible generalized dystonia and encephalopathy from thiamine transporter 2 deficiency. Mov. Disord. 27, 1295–1298. https://doi.org/10.1002/mds.25008 (2012).
Zeng, W. Q. et al. Biotin-responsive basal ganglia disease maps to 2q36.3 and is due to mutations in SLC19A3. Am. J. Hum. Genet. 77, 16–26. https://doi.org/10.1086/431216 (2005).
Ramamoorthy, K. et al. Alzheimer’s disease is associated with disruption in Thiamin transport physiology: A potential role for neuroinflammation. Neurobiol. Dis. 171, 105799. https://doi.org/10.1016/j.nbd.2022.105799 (2022).
Gibson, G. E. et al. Reduced activities of Thiamine-Dependent enzymes in the brains and peripheral tissues of patients with alzheimer’s disease. Arch. Neurol. 45, 836–840. https://doi.org/10.1001/archneur.1988.00520320022009 (1988).
Zhao, N. et al. Impaired hippocampal neurogenesis is involved in cognitive dysfunction induced by thiamine deficiency at early pre-pathological lesion stage. Neurobiol. Dis. 29, 176–185. https://doi.org/10.1016/j.nbd.2007.08.014 (2008).
Butterworth, R. F., Kril, J. J. & Harper, C. G. Thiamine-dependent enzyme changes in the brains of alcoholics: relationship to the Wernicke-Korsakoff syndrome. Alcohol Clin. Exp. Res. 17, 1084–1088. https://doi.org/10.1111/j.1530-0277.1993.tb05668.x (1993).
Lu, J. et al. Correlation of thiamine metabolite levels with cognitive function in the non-demented elderly. Neurosci. Bull. 31, 676–684. https://doi.org/10.1007/s12264-015-1563-3 (2015).
Tyagi, S., Gupta, P., Saini, A. S., Kaushal, C. & Sharma, S. The peroxisome proliferator-activated receptor: A family of nuclear receptors role in various diseases. J. Adv. Pharm. Technol. Res. 2, 236–240. https://doi.org/10.4103/2231-4040.90879 (2011).
D’Angelo, M. et al. PPARs and Energy Metabolism Adaptation during Neurogenesis and Neuronal Maturation. Int. J. Mol. Sci. 19, https://doi.org/10.3390/ijms19071869 (2018).
Woods, J. W. et al. Localization of PPARδ in murine central nervous system: expression in oligodendrocytes and neurons. Brain Res. 975, 10–21. https://doi.org/10.1016/S0006-8993(03)02515-0 (2003). https://doi.org/.
Li, X., Liu, S., Kapoor, K. & Xu, Y. PPARD May Play a Protective Role against the Development of Schizophrenia. PPAR Res. 2020(3480412), https://doi.org/10.1155/2020/3480412 (2020).
Yang, T. et al. PPARD May Play a Protective Role for Major Depressive Disorder. PPAR Res. 2021(5518138), https://doi.org/10.1155/2021/5518138 (2021).
Ji, M. J. et al. Hippocampal PPARδ Overexpression or Activation Represses Stress-Induced Depressive Behaviors and Enhances Neurogenesis. Int. J. Neuropsychopharmacol. 19, https://doi.org/10.1093/ijnp/pyv083 (2015).
Chen, F. et al. Hippocampal genetic knockdown of PPARδ causes Depression-Like behaviors and neurogenesis suppression. Int. J. Neuropsychopharmacol. 22, 372–382. https://doi.org/10.1093/ijnp/pyz008 (2019).
Espinosa-Jiménez, T. et al. Peroxisomal Proliferator-Activated receptor β/δ deficiency induces cognitive alterations. Front. Pharmacol. 13, 902047. https://doi.org/10.3389/fphar.2022.902047 (2022).
Insel, P. S., Kumar, A., Hansson, O. & Mattsson-Carlgren, N. For the alzheimer’s disease Neuroimaging, I. Genetic moderation of the association of β-Amyloid with cognition and MRI brain structure in alzheimer disease. Neurology 101, e20–e29. https://doi.org/10.1212/WNL.0000000000207305 (2023).
Wamsley, B. et al. Rbfox1 mediates Cell-type-Specific splicing in cortical interneurons. Neuron 100, 846–859e847. https://doi.org/10.1016/j.neuron.2018.09.026 (2018).
Lee, J. A. et al. Cytoplasmic Rbfox1 regulates the expression of synaptic and Autism-Related genes. Neuron 89, 113–128. https://doi.org/10.1016/j.neuron.2015.11.025 (2016).
Genomic & Relationships Novel Loci, and pleiotropic mechanisms across eight psychiatric disorders. Cell 179, 1469–1482e1411. https://doi.org/10.1016/j.cell.2019.11.020 (2019).
Raghavan, N. S. et al. Association between common variants in RBFOX1, an RNA-Binding Protein, and brain amyloidosis in early and preclinical alzheimer disease. JAMA Neurol. 77, 1288–1298. https://doi.org/10.1001/jamaneurol.2020.1760 (2020).
O’Leary, A. et al. Behavioural and functional evidence revealing the role of RBFOX1 variation in multiple psychiatric disorders and traits. Mol. Psychiatry. 27, 4464–4473. https://doi.org/10.1038/s41380-022-01722-4 (2022).
Zhong, Y. et al. RBFOX1 and working memory: from genome to transcriptome revealed posttranscriptional mechanism separate from Attention-Deficit/Hyperactivity disorder. Biol. Psychiatry Global Open. Sci. 3, 1042–1052. https://doi.org/10.1016/j.bpsgos.2022.08.006 (2023).
Zhang, X. et al. Overview of the complex figure test and its clinical application in neuropsychiatric Disorders, including copying and recall. Front. Neurol. 12, 680474. https://doi.org/10.3389/fneur.2021.680474 (2021).
Fagerberg, L. et al. Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Mol. Cell. Proteom. 13, 397–406. https://doi.org/10.1074/mcp.M113.035600 (2014).
Missler, M. & Südhof, T. C. Neurexophilins form a conserved family of neuropeptide-like glycoproteins. J. Neurosci. 18, 3630–3638. https://doi.org/10.1523/jneurosci.18-10-03630.1998 (1998).
Song, T. et al. Integrative identification by Hi-C revealed distinct advanced structural variations in lung adenocarcinoma tissue. Phenomics 3, 390–407. https://doi.org/10.1007/s43657-023-00103-3 (2023).
Aravindan, S. et al. Retinal degeneration protein 3 (RD3) in normal human tissues: novel insights. Sci. Rep. 7, 13154. https://doi.org/10.1038/s41598-017-13337-9 (2017).
Peshenko, I. V. et al. Retinal degeneration 3 (RD3) protein, a retinal Guanylyl cyclase regulator, forms a monomeric and elongated four-helix bundle. J. Biol. Chem. 294, 2318–2328. https://doi.org/10.1074/jbc.RA118.006106 (2019).
Wimberg, H., Janssen-Bienhold, U. & Koch, K. W. Control of the nucleotide cycle in photoreceptor cell extracts by retinal degeneration protein 3. Front. Mol. Neurosci. 11, 52. https://doi.org/10.3389/fnmol.2018.00052 (2018).
Azadi, S., Molday, L. L. & Molday, R. S. RD3, the protein associated with leber congenital amaurosis type 12, is required for guanylate cyclase trafficking in photoreceptor cells. Proc. Natl. Acad. Sci. 107, 21158–21163. https://doi.org/10.1073/pnas.1010460107 (2010).
Zulliger, R., Naash, M. I., Rajala, R. V., Molday, R. S. & Azadi, S. Impaired association of retinal degeneration-3 with guanylate cyclase-1 and guanylate cyclase-activating protein-1 leads to leber congenital amaurosis-1. J. Biol. Chem. 290, 3488–3499. https://doi.org/10.1074/jbc.M114.616656 (2015).
Bänziger, C. et al. Wntless, a conserved membrane protein dedicated to the secretion of Wnt proteins from signaling cells. Cell 125, 509–522. https://doi.org/10.1016/j.cell.2006.02.049 (2006).
Chai, G. et al. A human pleiotropic multiorgan condition caused by deficient Wnt secretion. N Engl. J. Med. 385, 1292–1301. https://doi.org/10.1056/NEJMoa2033911 (2021).
Chen, J., Park, C. S. & Tang, S. J. Activity-dependent synaptic Wnt release regulates hippocampal long term potentiation. J. Biol. Chem. 281, 11910–11916. https://doi.org/10.1074/jbc.M511920200 (2006).
Maguschak, K. A. & Ressler, K. J. Wnt signaling in amygdala-dependent learning and memory. J. Neurosci. 31, 13057–13067. https://doi.org/10.1523/jneurosci.3248-11.2011 (2011).
Caricasole, A. et al. Induction of Dickkopf-1, a negative modulator of the Wnt pathway, is associated with neuronal degeneration in alzheimer’s brain. J. Neurosci. 24, 6021–6027. https://doi.org/10.1523/jneurosci.1381-04.2004 (2004).
Liu, C. C. et al. Deficiency in LRP6-mediated Wnt signaling contributes to synaptic abnormalities and amyloid pathology in alzheimer’s disease. Neuron 84, 63–77. https://doi.org/10.1016/j.neuron.2014.08.048 (2014).
Ren, J. et al. MAPK4 predicts poor prognosis and facilitates the proliferation and migration of glioma through the AKT/mTOR pathway. Cancer Med. 12, 11624–11640. https://doi.org/10.1002/cam4.5859 (2023).
Zhang, D. et al. Up-regulation of plakophilin-2 is correlated with the progression of glioma. Neuropathology 37, 207–216. https://doi.org/10.1111/neup.12363 (2017).
Tran, D. N., Jung, E. M., Yoo, Y. M., Lee, J. H., Jeung, E. B. & 97,, , & Potassium-dependent sodium/calcium exchanger 3 (Nckx3) depletion leads to abnormal motor function and social behavior in mice. J. Physiol. Pharmacol. 71 https://doi.org/10.26402/jpp.2020.4.08 (2020).
Hautakangas, H. et al. Genome-wide analysis of 102,084 migraine cases identifies 123 risk loci and subtype-specific risk alleles. Nat. Genet. 54, 152–160. https://doi.org/10.1038/s41588-021-00990-0 (2022).
Lee, J. J. et al. Gene discovery and polygenic prediction from a genome-wide association study of educational attainment in 1.1 million individuals. Nat. Genet. 50, 1112–1121. https://doi.org/10.1038/s41588-018-0147-3 (2018).
Hyde, C. L. et al. Identification of 15 genetic loci associated with risk of major depression in individuals of European descent. Nat. Genet. 48, 1031–1036. https://doi.org/10.1038/ng.3623 (2016).
Wray, N. R. et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat. Genet. 50, 668–681. https://doi.org/10.1038/s41588-018-0090-3 (2018).
Ni, H. et al. The GWAS risk genes for depression May be actively involved in alzheimer’s disease. J. Alzheimers Dis. 64, 1149–1161. https://doi.org/10.3233/jad-180276 (2018).
Raghavan, N. S., Vardarajan, B. & Mayeux, R. Genomic variation in educational attainment modifies alzheimer disease risk. Neurol. Genet. 5, e310. https://doi.org/10.1212/nxg.0000000000000310 (2019).
Noh, K. et al. Negr1 controls adult hippocampal neurogenesis and affective behaviors. Mol. Psychiatry. 24, 1189–1205. https://doi.org/10.1038/s41380-018-0347-3 (2019).
105 Singh, K. et al. Neural cell adhesion molecule Negr1 deficiency in mouse results in structural brain endophenotypes and behavioral deviations related to psychiatric disorders. Sci. Rep. 9, 5457. https://doi.org/10.1038/s41598-019-41991-8 (2019).
Singh, K. et al. Neuronal growth and behavioral alterations in mice deficient for the psychiatric Disease-Associated Negr1 gene. Front. Mol. Neurosci. 11, 30. https://doi.org/10.3389/fnmol.2018.00030 (2018).
Donati, G., Dumontheil, I. & Meaburn, E. L. Genome-Wide association study of latent cognitive measures in adolescence: genetic overlap with intelligence and education. Mind Brain Educ. 13, 224–233. https://doi.org/10.1111/mbe.12198 (2019).
Hill, W. D. et al. A combined analysis of genetically correlated traits identifies 187 loci and a role for neurogenesis and myelination in intelligence. Mol. Psychiatry. 24, 169–181. https://doi.org/10.1038/s41380-017-0001-5 (2019).
Saive, A. L., Royet, J. P. & Plailly, J. A review on the neural bases of episodic odor memory: from laboratory-based to autobiographical approaches. Front. Behav. Neurosci. 8, 240. https://doi.org/10.3389/fnbeh.2014.00240 (2014).
Tong, M. T., Peace, S. T. & Cleland, T. A. Properties and mechanisms of olfactory learning and memory. Front. Behav. Neurosci. 8, 238. https://doi.org/10.3389/fnbeh.2014.00238 (2014).
Arshamian, A. et al. The functional neuroanatomy of odor evoked autobiographical memories cued by odors and words. Neuropsychologia 51, 123–131. https://doi.org/10.1016/j.neuropsychologia.2012.10.023 (2013). https://doi.org/https://doi.org/
Song, J. et al. A potential biomarker of cognitive impairment: the olfactory dysfunction and its genes expression. Ann. Clin. Transl. Neurol. 9, 1884–1897. https://doi.org/10.1002/acn3.51680 (2022).
Wilson, R. S. et al. Olfactory identification and incidence of mild cognitive impairment in older age. Arch. Gen. Psychiatry. 64, 802–808. https://doi.org/10.1001/archpsyc.64.7.802 (2007).
Bermúdez-Rattoni, F. Molecular mechanisms of taste-recognition memory. Nat. Rev. Neurosci. 5, 209–217. https://doi.org/10.1038/nrn1344 (2004).
Lim, S. X. L. et al. The capacity and organization of gustatory working memory. Sci. Rep. 12, 8056. https://doi.org/10.1038/s41598-022-12005-x (2022).
Fannon, A. M. & Colman, D. R. A model for central synaptic junctional complex formation based on the differential adhesive specificities of the cadherins. Neuron 17, 423–434. https://doi.org/10.1016/S0896-6273(00)80175-0 (1996).
Elste, A. M. & Benson, D. L. Structural basis for developmentally regulated changes in Cadherin function at synapses. J. Comp. Neurol. 495, 324–335. https://doi.org/10.1002/cne.20876 (2006).
Schrick, C. et al. N-cadherin regulates cytoskeletally associated IQGAP1/ERK signaling and memory formation. Neuron 55, 786–798. https://doi.org/10.1016/j.neuron.2007.07.034 (2007).
Asada-Utsugi, M. et al. Mice with cleavage-resistant N-cadherin exhibit synapse anomaly in the hippocampus and outperformance in Spatial learning tasks. Mol. Brain. 14, 23. https://doi.org/10.1186/s13041-021-00738-1 (2021).
Arias-Vásquez, A. et al. CDH13 is associated with working memory performance in attention deficit/hyperactivity disorder. Genes Brain Behav. 10, 844–851. https://doi.org/10.1111/j.1601-183X.2011.00724.x (2011). https://doi.org/.
Alistair, T. P.et al. Rare familial 16q21 microdeletions under a linkage peak implicate cadherin 8 (CDH8)in susceptibility to autism and learning disability. J. Med. Genet.48, 48 (2011). https://doi.org/10.1136/jmg.2010.079426.
Wang, K. et al. Common genetic variants on 5p14.1 associate with autism spectrum disorders. Nature 459, 528–533. https://doi.org/10.1038/nature07999 (2009).
Asada-Utsugi, M. et al. N-cadherin enhances APP dimerization at the extracellular domain and modulates Aβ production. J. Neurochem. 119, 354–363. https://doi.org/10.1111/j.1471-4159.2011.07364.x (2011).
Agiostratidou, G., Muros, R. M., Shioi, J., Marambaud, P. & Robakis, N. K. The cytoplasmic sequence of E-cadherin promotes non-amyloidogenic degradation of A beta precursors. J. Neurochem. 96, 1182–1188. https://doi.org/10.1111/j.1471-4159.2005.03616.x (2006).
Ghosh, C. et al. Expression and functional relevance of UGT1A4 in a cohort of human drug-resistant epileptic brains. Epilepsia 54, 1562–1570. https://doi.org/10.1111/epi.12318 (2013).
Xuan, J. et al. Metabolomic profiling to identify potential serum biomarkers for schizophrenia and Risperidone action. J. Proteome Res. 10, 5433–5443. https://doi.org/10.1021/pr2006796 (2011).
Kurano, M., Saito, Y. & Yatomi, Y. Comprehensive analysis of metabolites in postmortem brains of patients with alzheimer’s disease. J. Alzheimers Dis. 97, 1139–1159. https://doi.org/10.3233/jad-230942 (2024).
O’Dushlaine, C. et al. The SNP ratio test: pathway analysis of genome-wide association datasets. Bioinformatics 25, 2762–2763. https://doi.org/10.1093/bioinformatics/btp448 (2009).
Wang, K., Li, M. & Bucan, M. Pathway-based approaches for analysis of genomewide association studies. Am. J. Hum. Genet. 81, 1278–1283. https://doi.org/10.1086/522374 (2007).
Shearer, K. D., Stoney, P. N., Morgan, P. J. & McCaffery, P. J. A vitamin for the brain. Trends Neurosci. 35, 733–741. https://doi.org/10.1016/j.tins.2012.08.005 (2012).
McCaffery, P., Zhang, J. & Crandall, J. E. Retinoic acid signaling and function in the adult hippocampus. J. Neurobiol. 66, 780–791. https://doi.org/10.1002/neu.20237 (2006).
Liu, Z. et al. Research progress in vitamin A and autism spectrum disorder. Behav. Neurol. 2021 (5417497). https://doi.org/10.1155/2021/5417497 (2021).
Behl, T. et al. Therapeutic insights elaborating the potential of retinoids in alzheimer’s disease. Front. Pharmacol. 13, 976799. https://doi.org/10.3389/fphar.2022.976799 (2022).
Goodman, A. B. Retinoid receptors, transporters, and metabolizers as therapeutic targets in late onset alzheimer disease. J. Cell. Physiol. 209, 598–603. https://doi.org/10.1002/jcp.20784 (2006).
Chakrabarti, M. et al. Molecular signaling mechanisms of natural and synthetic retinoids for Inhibition of pathogenesis in alzheimer’s disease. J. Alzheimers Dis. 50, 335–352. https://doi.org/10.3233/JAD-150450 (2016).
Yuan, C. et al. Long-Term intake of dietary carotenoids is positively associated with Late-Life subjective cognitive function in a prospective study in US women. J. Nutr. 150, 1871–1879. https://doi.org/10.1093/jn/nxaa087 (2020).
Rutjes, A. W. et al. Vitamin and mineral supplementation for maintaining cognitive function in cognitively healthy people in mid and late life. Cochrane Database Syst. Rev. 12, Cd011906 (2018). https://doi.org/10.1002/14651858.CD011906.pub2
Acknowledgements
We would like to thank all participants who participated in this study. We thank Lauren K. Spriggens for collecting the GRC GOM cohort phenotype data. We thank Nick G. Martin, Margaret J. Wright, and Scott D. Gordon for their collaboration concerning the QIMR Twin Cohort replication study.
Author information
Authors and Affiliations
Contributions
L.R.G., R.A.L., H.G.S. and D.H.K.S. designed and supervised the overall study; D.H.K.S. designed the memory performance evaluations; H.G.S. and N.A. prepared the GRC GOM cohort samples for genotyping; L.N.H. and N.A. analysed and interpreted the data; E.E.M.K., J.B., and D.C.G. were responsible for the GOBS replication study; L.N.H. and N.A. wrote the draft paper; H.G.S., L.M.H, R.A.L., F.E.F., D.H.K.S., and L.R.G. edited the paper and approved the final draft.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hopkins, L.N., Avgan, N., Sutherland, H.G. et al. Genome-wide association study identifies novel variants in olfactory, vitamin A, vitamin B, and cadherin pathways associated with learning and memory. Sci Rep 16, 2911 (2026). https://doi.org/10.1038/s41598-025-32828-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-32828-8