Abstract
Tumor cells express programmed cell death ligand 1 (PD-L1), which recognizes the immune checkpoint molecule programmed cell death 1 (PD-1) on T cells, suppressing the antitumor immune response. Inhibiting the PD-1:PD-L1 interaction has the potential to reactivate the immune response against tumors. Recent advancements in cancer therapy have demonstrated remarkable promise of immunotherapy, which exploits immune checkpoint inhibition by small molecules or monoclonal antibodies. This strategy has shown impressive clinical success in treating a wide range of cancer subtypes, albeit with certain limitations. This study aims to design a novel multi-epitope vaccine against PD-L1 by using an immunoinformatics approach. For attaining enhanced efficacy and minimize side effects, the vaccine was constructed using antigenic, non-allergenic, and non-toxic epitopes (5 CTL, 3 HTL, and 2 B-cell epitopes) predicted from the IgV domain of PD-L1. The vaccine design includes a large ribosomal subunit protein bL12 adjuvant, a 6xHis tag for purification, and appropriate linkers to connect the epitopes. The modelled 3D structure of the vaccine construct was docked with TLR4 immune receptor, demonstrating strong antigenic properties and stable binding, as validated by molecular dynamics simulations. Immune simulation studies suggest that the vaccine construct could potentially elicit significant immune regulators such as B cells, T-cells, and memory cells. Thus, the findings indicate that the vaccine may effectively suppress the PD-1:PD-L1 axis by targeting PD-L1, restoring the anticancer immune response. However, its efficacy needs to be validated in both in vitro and in vivo settings.
Similar content being viewed by others
Introduction
Despite the constant efforts and remarkable success in combating cancer, the incidence of cancer cases and related deaths continue to rise. One of the emerging approaches with immense potential for cancer management is cancer immunotherapy which activates the host immune system, allowing it to counter cancer cells. The immune system’s precision, dynamic nature, and memory enable it to target cancer cells while sparing healthy ones. The immune memory helps it to re-evaluate and eliminate the disease upon cancer recurrence. Immunotherapy consists of three main components: checkpoint inhibition, adoptive cell therapy, and vaccines. The checkpoint inhibition relies on the blockade of immune checkpoint molecules like cytotoxic T lymphocyte-associated antigen 4 (CTLA-4), the programmed cell death 1 (PD-1) receptor, and its ligand, programmed cell death ligand 1 (PD-L1). The PD-1 and PD-L1 are transmembrane proteins with immunoglobulin (Ig)-like extracellular domains that enable interaction and signal transduction to intracellular regions. PD-1, also known as CD279, is a type I transmembrane receptor found on the surface of T cells, B cells, monocytes, natural killer cells, and dendritic cells. It has two ligands that are found naturally: PD-L1 (B7-H1, CD274) and PD-L2 (B7-DC, CD273)1. PD-L1 has been reported to be overexpressed in a number of cancer types such as melanoma, lymphoma, glioblastoma, as well as carcinoma of ovary, colon, lung squamous cells, and breast2. The PD-1 and PD-L1 interaction sends a negative signal to the cytotoxic T-lymphocyte, inhibiting antitumor immunity. As a result, blocking the PD-1: PD-Ll interaction reactivates cytotoxic CD8 + T cells, reinstating antitumor immunity.
The development of immune checkpoint inhibitors has ushered in a new phase of immunotherapy for cancer. Monoclonal antibodies (mAbs) designed to target PD-1 and PD-L1 are critical in rescuing T-lymphocytes from exhaustion and reinvigorating the immune response against cancerous cells. Several anti-PD-1 and anti-PD-L1 mAbs have exhibited remarkable clinical outcomes and anticancer activity in patients with various cancers. The US FDA has approved three anti-PD-1 antibodies, namely nivolumab, pembrolizumab, and cemiplimab as well as three anti-PD-L1 antibodies, atezolizumab, durvalumab, and avelumab, for the treatment of various cancer types3. Moreover, several additional checkpoint inhibitors are currently being tested in clinical trials. While these mAb therapies as monotherapy have demonstrated notable clinical efficacy, challenges such as limited response rates, toxicity issues, resistance, steep costs, long half-life, and sophisticated therapeutic regimens remain important impediments4. Addressing these limitations will necessitate the development of more effective immune checkpoint inhibitors or novel combinational approaches. Another paradigm of the cancer immunotherapy includes development of vaccines.
Cancer vaccines promote antigen-specific immune responses by presenting tumor antigens to the patient’s immune system. The challenges for vaccination against cancer include limited penetrability, immune response waning, and development of resistance. Multi-target vaccines designed to target immunogenicity-optimized epitopes may be able to tackle some of these problems5. Understanding immune evasion mechanisms, designing effective formulations, and combining immunotherapy approaches can pave the way for future cancer vaccine development. Multi-epitope vaccines effectively activate both humoral and cellular immune responses by targeting T and B cell epitopes simultaneously, offering advantages such as high specificity, superior safety, ease of production and storage, and long-lasting efficacy6. Additionally, the incorporation of adjuvants in multi-epitope vaccines is expected to elicit long-lasting immunological responses and achieve high immunogenicity7. In this study, we assessed the T-cell and B-cell epitopes from human PD-L1 to design a multi-epitope cancer vaccine. This vaccine would potentially elicit both humoral and cell-mediated immunity, which will generate polyclonal antibodies targeting the PD-1: PD-L1 signaling axis, restoring cytotoxic T-cell functionality.
Materials and methods
A robust multi-peptide cancer vaccine against PD-L1 was designed using computational approach combining multiple bioinformatics tools and techniques as illustrated in Fig. 1.
Methodology outline for the designing of a multi-epitope vaccine candidate against PD-L1. The process includes antigen selection, epitope prediction, molecular docking, and MD simulation, followed by in silico simulation to verify the predicted efficacy of the vaccine candidate. The figure was prepared using Microsoft PowerPoint 2013.
Protein sequence retrieval and domain analysis
To predict potential epitopes in the protein and design a multi-epitope cancer vaccine, the PD-L1 protein sequence with accession number Q9NZQ7 was obtained from the Uniprot database (https://www.uniprot.org/) on September 14, 2023. Additionally, domain analysis was conducted using the Protter8 server (https://wlab.ethz.ch/protter/start/) with the same Uniprot accession number on February 12, 2024.
Cytotoxic T lymphocyte (CTL) epitopes prediction
A consensus list of high-binding and promiscuous cytotoxic T lymphocyte (CTL) epitopes was compiled by using the following webtools: NetCTL version 1.2 (https://services.healthtech.dtu.dk/services/NetCTL-1.2/), PickPocket version 1.1 (https://services.healthtech.dtu.dk/services/PickPocket-1.1/), and NetMHCpan − 4.1 (https://services.healthtech.dtu.dk/services/NetMHCpan-4.1/)9. All these web servers were accessed on September 14, 2023. NetCTL uses artificial neural networks to predict binding to MHC class I and proteasomal cleavage, and it employs a weight matrix to estimate TAP transport efficiency10,11. PickPocket, on the other hand, relies on position-specific weight matrices for its predictions12. NetCTLpan employs artificial neural networks for the epitope predictions. All the parameters were utilized in their default settings. The consensus list was created by selecting the top 10 affinity-sorted epitopes from PickPocket and comparing them with the high-scoring epitopes predicted by NetCTLpan and NetMHCpan to find common epitopes. This improved prediction diversity and accuracy. This analysis utilized a default set of 12 representative HLA supertypes and nonameric peptide epitopes. The 12 shared HLA supertypes in both algorithms were HLA-A*01:01, HLA-A*02:01, HLA-A*03:01, HLA-A*24:02, HLA-A*26:01, HLA-B*07:02, HLA-B*08:01, HLA-B*27:05, HLA-B*39:01, HLA-B*40:01, HLA-B*58:01, and HLA-B*15:01.
Helper T lymphocytes (HTL) epitopes prediction
The helper T lymphocytes (HTL) epitopes were predicted by using the NetMHCIIpan 4.0 (https://services.healthtech.dtu.dk/services/NetMHCIIpan-4.0/)9 server accessed on September 15, 2023, focusing on 15 amino acid sequences. Strong binding peptides were identified with a threshold of 2% of %Rank, while weak binding peptides were filtered at 10% of %Rank. Thirteen common HLA Class II alleles, including HLA-DRB1-0101, HLA-DRB1-0301, HLA-DRB1-0401, HLA-DRB1-0701, HLA-DRB1-0801, HLA-DRB1-0901, HLA-DRB1-1001, HLA-DRB1-1101, HLA-DRB1-1201, HLA-DRB1-1301, HLA-DRB1-1401, HLA-DRB1-1501, and HLA-DRB1-1601 were analysed to assess binding affinities and identify potential HTL epitopes.
Linear B-cell epitopes prediction
The ABCpred (https://webs.iiitd.edu.in/raghava/abcpred/index.html)13 web server was accessed on September 15, 2023, for predicting the linear B-cell epitopes of PD-L1. Default parameters were used for B-cell epitope prediction, with a length of 16 amino acid residues selected for prediction. The ten highest-ranking predicted epitopes were subsequently selected for further analysis. The ABCpred has been trained on B-cell epitopes sourced from the Bcipep database. It utilizes a recurrent neural network for classifying epitopes and non-epitopes, enhancing accuracy in the prediction process13.
Epitope screening
The best epitopes were selected based on their antigenicity, toxicity, and allergenicity due to the abundance of predicted epitopes. To predict the antigenicity of the epitopes, the VaxiJen v2.0 (https://www.ddg-pharmfac.net/vaxijen/VaxiJen/VaxiJen.html) server was employed14. This server is a versatile tool that can be used to calculate the antigenicity of a wide range of microorganisms, including bacteria, viruses, fungi as well as tumors, and parasites. Its prediction accuracy typically falls between 70% and 89%, making it a reliable choice for such analyses15. In this study, the tumor was the target, and an antigenicity threshold of 0.5 was set to identify the best epitopes for the vaccine. In addition to antigenicity, toxicity and allergenicity of the epitopes were also evaluated to ensure the safety of the vaccine. The ToxinPred (https://webs.iiitd.edu.in/raghava/toxinpred/multi_submit.php) server was used to predict the toxicity of the selected epitopes. In this study, the Swiss-Prot SVM-based method was employed to predict toxicity16. The AllerTOP v.2.0 (https://www.ddg-pharmfac.net/AllerTOP/) server was used to assess the allergenicity of the epitopes. Not all HTL epitopes trigger the production of cytokines, and even when they do, the cytokines released may vary among them. To further evaluate the selected epitopes, IL4pred (https://webs.iiitd.edu.in/raghava/il4pred/predict.php) and IFNepitope (https://webs.iiitd.edu.in/raghava/ifnepitope/predict.php) servers were used to predict their ability to induce the cytokines IL-4 and IFN-γ, respectively. For predicting IL-4 inducing HTL epitopes, a threshold of 0.2 was selected and an SVM-based model was utilized by the IL4pred server17,18. Similarly, for predicting IFN-γ inducing HTL epitopes, an SVM-based model was employed, but with an IFN-γ versus other cytokine models used by the IFNepitope server. All these web servers were accessed on September 17, 2023.
Worldwide human population coverage analysis
The population coverage of the selected epitopes was assessed by using the IEDB population coverage analysis (http://tools.iedb.org/population/)19 tool accessed on November 14, 2023. The assessment of the human population was conducted globally.
Multi-epitope vaccine construction
To enhance the immunogenicity of the selected epitopes, specific linkers such as EAAAK, GGGS, GPGPG, HEYGAEALERAG, AAY, and KK were used to connect different components in a rational manner. Additionally, to boost immune responses, adjuvant molecules were used. Four different adjuvants were tried, leading to the creation of four distinct constructs. These constructs were further analyzed for their physicochemical properties, antigenicity, allergenicity, and secondary structures. Additionally, the Pan DR epitope (PADRE – AKFVAAWTLKAAA) adjuvant was fused to serve as a stimulus for HTL. The process involved the sequential addition of CTL epitopes, followed by HTLs and B-cell epitopes. Finally, a 6xHis tag was added to the C-terminal portion for subsequent purification of the vaccine protein.
Physicochemical properties and solubility analysis of the vaccine
After designing the chimeric sequences, their physicochemical properties were evaluated using the ProtParam (https://web.expasy.org/protparam/) webserver20. Additionally, using the SoluProt 1.0 server (https://loschmidt.chemi.muni.cz/soluprot/), the solubility of chimaeras upon expression in bacteria was assessed21. The TargetTrack database served as the training set for the gradient-boosting machine algorithm that developed SoluProt21. Additionally, the solubility was also predicted using the SOLpro (https://scratch.proteomics.ics.uci.edu/) server22. According to estimates using tenfold cross validation, SolPro, an SVM-based method for predicting protein solubility from sequences, achieves a global accuracy exceeding 74% 22.
Evaluation of the antigenicity and allergenicity of the vaccine
In the development of vaccines, evaluating the antigenicity of the final vaccine construct is a crucial step. To predict the antigenic behaviour, two online servers were used, the VaxiJen v2.0 and ANTIGENpro (https://scratch.proteomics.ics.uci.edu/)23. Moreover, to ensure the safety of the vaccine, AllerTOP version 2.0 was employed to assess its potential allergenicity.
Prediction of the secondary structure
The percentage of secondary structure elements in the vaccine construct was determined by using the Prabi (https://npsa-prabi.ibcp.fr/cgi-bin/npsa_automat.pl?page=/NPSA/npsa_gor4.html) server 24.
Structure prediction and validation of the multi‑epitope vaccine
The 3D structure of the designed multi-epitope vaccine was predicted using the trRosetta server (https://yanglab.nankai.edu.cn/trRosetta25. The trRosetta is a web-based server that predicts protein structures using deep learning and Rosetta. A neural network predicts inter-residue geometries, which are used as restraints for energy minimization-based structure prediction with Rosetta25. The 3D model of the multi-epitope vaccine protein underwent a two-step refinement process. Initially, the ModRefiner server (https://seq2fun.dcmb.med.umich.edu//ModRefiner/)26 was used. Subsequently, refinement was carried out using the GalaxyRefine server (https://galaxy.seoklab.org/cgi-bin/submit.cgi?type=REFINE)27.
The protein structure was validated using two different servers: ProSA-web (https://prosa.services.came.sbg.ac.at/prosa.php) and SAVES v6.0 (https://saves.mbi.ucla.edu/)28. ProSA-web calculates the overall quality Z-score of a protein structure. If the Z-score is outside the typical range for native proteins, it suggests that there could be errors in the structure29. On the other hand, SAVES v6.0 uses the PROCHECK tool to evaluate the stereochemical quality of the protein structure by checking the geometry of individual residues and the overall structural geometry. This helps identify any anomalies or irregularities in the protein structure28.
Disulfide bond engineering in the multi-epitope vaccine
Disulfide engineering is a method to introduce disulfide bonds to protein structures. Such bonds stabilize the folded conformation of the protein by lowering its conformational entropy, thus increasing the free energy of the denatured state30. Residue pairs in the vaccine construct that may potentially mutate to cysteine and form interprotein disulfide bonds were identified using the Disulfide by Design 2.13 server available at (http://cptweb.cpt.wayne.edu/DbD2/.)31.
Discontinuous B cell epitope prediction
The Ellipro server (http://tools.iedb.org/ellipro/) was utilized to predict discontinuous B-cell epitopes in the designed vaccine construct. Ellipro uses a residue clustering algorithm and Thornton’s method to identify the epitopes, to which the PI (protrusion index) values were assigned15,32.
Interaction study of T cell epitopes with MHC molecules
Interactions of the selected CTL and HTL epitopes with MHC-I and MHC-II molecules respectively were evaluated by performing molecular docking analysis. Molecular docking is a computational method that determines the best orientation of a ligand in a complex with a receptor. For this purpose, the 3D structures of the selected MHC-I and MHC-II epitopes were modelled using the PEP-FOLD 4.0 online server(https://bioserv.rpbs.univ-paris-diderot.fr/services/PEP-FOLD4/)33. The best models for docking with their corresponding alleles: HLA-A*03:01 (PDB ID: 2XPG) for the MHC-I epitope and HLA-DRB1-1501 (PDB ID: 1BX2) for all MHC-II epitopes. Molecular docking was performed using the HPEPDOCK 2.0 (http://huanglab.phys.hust.edu.cn/hpepdock/ ) online server34, and the docking results of all epitopes with their corresponding MHC alleles were visualized using PyMOL 3.1.
Molecular docking of the designed vaccine with TLR4 receptor
The activation of a robust immune response depends on the interaction between an antigenic molecule and a specific immune receptor. The engagement of Toll-like receptors (TLRs) by vaccine epitopes is a critical step in initiating the immune response, with TLR4 being one of the key receptors implicated in recognizing pathogens and vaccine components. To identify the binding pockets or cavities in the (TLR4) receptor, the CASTp (http://sts.bioe.uic.edu/castp/) server was employed35. CASTp excels in identifying and measuring surface-accessible binding pockets, providing information about both accessible binding pockets and inner inaccessible cavities for protein molecules35. The interaction between the vaccine construct and TLR4 was evaluated by performing molecular docking. The docking analysis was conducted using ClusPro 2.0 server, where the refined model of the vaccine construct was submitted as the ligand and TLR4 (PDB ID: 3FXI) was submitted as the receptor. The PDBsum (https://www.ebi.ac.uk/thornton-srv/databases/pdbsum/) server was used to visualise the bonds formed between the residues of the vaccine construct and TLR4 in the docked complex36.
Molecular dynamics simulation of the vaccine‑receptor complex
In this study, we conducted molecular dynamic (MD) simulations to investigate the structural dynamics and specific interactions between the designed multiepitope vaccine and TLR4 by using Gromacs 22.4v37. The complex is composed of Chain A, and C of TLR4, and vaccine subunit chain V and their topology was generated by using AMBER99SB-ILDN force field. Then the complex was imported in SPC water box and added 195,640 solvents. The system was neutralised by adding 377 Na+ and 370 Cl − atoms to maintain the physiological pH with a concentration of 0.15 M. The MD simulations analyse several critical parameters to reveal the stability and specific binding interactions, providing insights into the potential efficacy of the vaccine design. In this study, MD simulations have been performed to elucidate the structural stability and binding efficacy of the vaccine-TLR4 complex. The long-range electrostatic interactions and hydrogen bond distances were handled using PME and LINCS algorithms respectively. The vaccine-TLR4 complex, comprising chains A and C of the receptor and chain V of the vaccine construct were equilibrated for 2 ns. The MD simulations were done for 175 ns with a time interval of 0.2 fs39 in which the root-mean-square deviation (RMSD) was monitored to assess structural stability. The RMSF was employed to determine the fluctuations of each amino acid present in the complex40. The root mean square distribution of the TLR4 and Vaccine construct was conducted by the cluster analysis. First, we have optimised the cutoff values from 0.4 − 0.2 and more clusters were obtained at 0.2 cutoff following the gromos method. Also, the hydrogen bonds occurred between the TLR4, and vaccine construct was generated by using the hbond module followed by the previous established protocols40.
Moreover, the structural compactness and dynamics of the TLR4 complexed with the designed multiepitope vaccine was further enriched by examining the radius of gyration (RoG) throughout the MD simulations. The RoG is a critical parameter that quantifies the molecule’s compactness, providing insights into the structural integrity and conformational changes over time. Therefore, such analysis was employed by various studies to determine the behaviour of the complex in the aqueous medium41.
Principal component and free energy landscape analysis
The module GROMACS gmx sham yielded meta-stable conformations, and two-dimensional free energy landscape images were generated using the gmx xpm2ps module. The Gibbs energy landscape is a very fundamental indicator of the thermodynamic characteristics associated with the simulated complex42. Analysis was done using tools available within GROMACS, beginning with the diagonalisation of the covariance matrix through modules of gmx covar and gmx anaeig, representing the energy of particular components42.
In Eq. (1), K represents the Boltzmann constant, while PC1 and PC2 denote the first two principal reaction coordinates. The fluctuations in enthalpy (ΔH), standard free energy (ΔG), and entropy (ΔS) were computed using the provided formula.
In Eq. (2), the symbols ΔH, TΔS, and ΔG represent enthalpy, temperature (in Kelvin), system entropy, and Gibbs free energy, respectively.
Molecular mechanics poisson boltzmann surface area (MM-PBSA) calculation
We have employed the MM-PBSA method to calculate the average binding free energy of vaccine construct with TLR4 using Gromacs 5.2v software. The TLR4 is composed of two chains A & C initially making two groups, (1) Vaccine construct (2) TLR4 Chain A&C. Following the bootstrap method was employed to calculate the average binding free energy by using “MmPbSaStat.py” code available on Kumari GitHub 2014. The required files were obtained from MD simulations (175 ns) for free energy calculations. The ΔGbind was calculated according to the following equation (i).
ΔGbinding = ΔGcomplex− (ΔGprotein + ΔGligand) (i).
In this particular approach, the computation of entropy via normal mode analysis is frequently omitted because its inclusion does not improve agreement with experimental data.
Immune simulation for vaccine efficacy
To understand the immunogenicity and immune response of the multi-epitope vaccine, in silico immune simulations were performed using the C-ImmSim server (https://kraken.iac.rm.cnr.it/C-IMMSIM/index.php), accessed on November 13, 202343, . To initiate the immune response generated by the designed chimeric antigen against PD-L1, this study adopted an immunization schedule similar to that was followed for the schistosomiasis vaccines rSh28GST and rSm14, which are currently under clinical trials44,45. The immunization schedule involves three doses administered at 4-week intervals ensuring optimal vaccine efficacy. Injections containing 1,000 vaccine proteins each were given at time-steps 1, 84, and 168, spaced four weeks apart, totalling 1,050 simulation steps. Each time-step is equivalent to 8 h in real-life, and time-step 1 corresponds to the injection at time = 0. The default simulation parameters were used for all other simulation aspects.
Codon optimization and in silico cloning of the final vaccine construct
The Java Codon Adaptation Tool (http://www.jcat.de/Start.jsp) was utilized to perform back translation and codon optimization for the final vaccine construct. The protein sequence of the vaccine was provides as input to JCat, and the host organism chosen for expressing the vaccine construct was E. coli (K12 strain). Within this server, two parameters were calculated: the codon adaptive index (CAI) and the GC content. These parameters play a crucial role in assessing protein expression levels. Following the addition of BamHI and HindIII sites to its 5’ and 3’ ends, the nucleic acid sequence was restriction digested and cloned into the pET-28a (+) vector using Snapgene software. The pET28a plasmid was chosen for cloning our vaccine construct due to its strong T7 promoter, common restriction sites, controlled expression with a lac operator, self-encoding lac repressor, and medium copy number, allowing high-level expression without overloading cells46.
Results
Retrieval of PD-L1 protein sequence and preliminary analysis
The PD-L1 protein sequence (Q9NZQ7) was retrieved from the Uniprot database. This protein, which is 290 amino acids long, belongs to the B7 family of type I transmembrane protein receptors47. The protein consists of two extracellular domains (IgV and IgC), a transmembrane domain, and a cytoplasmic domain47. The Ig-V domain spans from amino acid 19 to 127, while the IgC domain spans from amino acids 133 to 225, joined by a short stalk region covering amino acid residues 128–132. Another short stalk region connects the IgC domain to the transmembrane domain48. The IgV domain of PD-L1 serves as the sole interaction domain for PD-1 binding49. Therefore, in this study, we selected the segment of the protein spanning from residues 19 to 225 that encompasses the complete IgV domain as well as a brief stretch of amino acids at the C terminal of the IgV continuing to a few residues of IgC domain. The domains of PD-L1 are visualized using Protter, an open-source visualisation tool, as depicted in Fig. 2.
Primary structure of PD-L1 showing various functional domains. The visualisation of domain positions in the PD-L1 protein sequence in a transmembrane environment was done by using Protter server (https://wlab.ethz.ch/protter/start/).
Prediction of CTLs in PD-L1
The CTL epitopes are crucial for inducing cellular immunity mediated by CD8+ CTLs restricted by MHC class I molecules. Therefore, these epitopes are promising candidates for designing subunit vaccines targeting a range of diseases50. The CTLs targeting PD-L1 can identify and eliminate cancerous lymphoma cells and normal immune cells that express PD-L1, potentially boosting the effector stage of the immune response51.
Here, the potential CTL epitopes of PD-L1 were predicted by using NetCTL-1.2, NetMHCpan-4.1, and PickPocket 1.1, employing the default 12 representative HLA supertypes. These supertypes are globally distributed, making them representative across populations52. Subsequently, the three prediction algorithms were employed to predict and compile a consensus list of top high binders. The consensus list was selected to enhance prediction accuracy by considering results from different algorithms. These predictions generated 73 CTL epitopes from NetCTL-1.2, 31 from NetMHCpan-4.1, and 120 from PickPocket-1.1 (Supplementary Table S1). The common epitopes from all three lists were considered for further analysis in designing the multi-epitope vaccine. We identified 23 common epitopes, from which 5 CTLs were selected for vaccine construction after using strict screening criteria.
Prediction of HTLs in PD-L1
The CD4 + HTLs play a crucial role in both humoral and cell-mediated immune responses53. Consequently, epitopes specific to HTL receptors are deemed crucial components of prophylactic and therapeutic vaccines. The HTLs play a pivotal role in initiating and sustaining long-term antitumor CTL responses54. In a recent study, Hirata-Nozaki et al., (2019)55 reported that HTLs specific to PD-L1 produce effector cytokines and augment cytotoxic activity against tumor cells expressing PD-L1. Notably, when PD-L1-specific HTLs were transferred into immunodeficient mice, there was a significant inhibition in the growth of PD-L1-positive human lung carcinoma55.
In this study, the potential HTL epitopes of PD-L1 were predicted by using NetMHCIIpan 4.0 as described in the Methods section. A total of 73 epitopes with high binding affinity were generated and are detailed in Supplementary Table S2. Out of these epitopes, finally three HTLs were selected for vaccine construction following certain screening criteria.
Prediction of linear B-cell epitopes in PD-L1
B-cell epitopes are regions on antigen surfaces that B-cell receptors (BCR) recognize, initiating immune responses. This process is fundamental to the adaptive immune system and is responsible for immunological memory and targeted responses to antigens in vertebrates56. Mapping B-cell epitopes is crucial for diagnostics and effective vaccine design57. Recently, Guo and colleagues showed that a PD-L1 B-cell epitope peptide vaccine produced robust immune responses and demonstrated significant antitumor immunity across multiple syngeneic mice models4.
Top ten B-cell epitopes with the highest scores are shown in Supplementary Table S3. The identified B cell epitopes were ranked based on their scores derived from a trained recurrent neural network. A higher score indicates a greater likelihood of the peptide being an epitope13. Considering all the selection criteria, finally two B cell epitopes were selected for vaccine construction.
Screening the predicted epitopes
The predicted epitopes were evaluated for antigenicity using the VaxiJen v2.0 server, for toxicity with ToxinPred v2.0, and for allergenicity with AllerTOP v2.0. Additionally, for HTL epitopes, the evaluation included their ability to induce IL-4 and IFN-γ (Supplementary Table S4). The IFN-γ and IL-4 play crucial roles in regulating the development and differentiation of immune cells, as well as the overall immune response of an organism. The cytokine IFN-γ promotes T helper type 1 (Th1) responses, while IL-4 stimulates T helper type 2 (Th2) responses58. IFN-γ enhances the antigen presentation ability of antigen-presenting cells and promotes the differentiation of CD4 + Th1 cells. On the other hand, IL-4 stimulates the proliferation of activated B cells59,60. Thus, the HTLs that stimulate IFN-γ and IL-4, which enhance immune response, are considered suitable vaccine candidates.
Only epitopes that demonstrated characteristics of being antigenic, non-toxic, and non-allergenic properties, specifically within or overlapping with the Ig-V domain, were chosen for the effective vaccine construction. Overlapping sequences were chosen due to the presence of amino acids within the Ig-V domain, which makes up the hotspot region for PD-1 binding, making it a potential target for PD-1: PD-L1 axis inhibition. Following all the screening criteria, the following ten epitopes were identified: five CTL epitopes (YRQRARLLK, KLQDAGVYR, ISYGGADYK, KRITVKVNA, ITVKVNAPY), three HTL epitopes (DLYVVEYGSNMTIEC, YGGADYKRITVKVNA, GGADYKRITVKVNAP), and two B-cell epitopes (HGEEDLKVQHSSYRQR, ALQITDVKLQDAGVYR). The attributes of the selected epitopes are presented in Table 1.
Worldwide human population coverage analysis
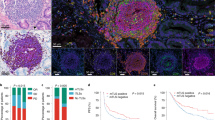
The IEDB population coverage calculation tool was used to estimate the anticipated Global response to a specific set of MHC-restricted epitopes. The finally selected CTL and HTL epitopes were subjected to population coverage analysis to assess their likelihood of binding to MHC molecules across the global population. The calculated world population coverage for MHC class I was found to be 25.83%, while for MHC class II, it was 33.41%. The cumulative population coverage for both MHC class I and II molecules was 50.61%. The Fig. 3 displays various region-wise data. It is to be noted that certain country-specific data were not available.
Population coverage of the selected HTL and CTL epitopes and their associated alleles. (A) Bar plots representing the population coverage for MHC-I and MHC-II. (B) A Global map illustrating the distribution of population coverage across different regions. Created using MapChart (https://www.mapchart.net/).
Multi-epitope vaccine construction
With the ten selected epitopes identified for the vaccine construction, four distinct constructs were designed, incorporating four different adjuvants. The following four adjuvants were considered: (1) Resuscitation-promoting factor RpfE, (2) Large ribosomal subunit protein bL12, (3) Beta-defensin 3, and (4) Cholera enterotoxin subunit B. The use of diverse adjuvants aimed to assess their compatibility with the designed vaccine and their ability to elicit strong and protective immune responses. These constructs were assessed for both antigenicity and allergenicity, and their secondary structures were predicted. To construct the multi-epitope vaccine, five CTL epitopes, three HTL epitopes, and two B-cell epitopes were combined using AAY, GPGPG, and KK linkers, respectively. An EAAAK linker was used to attach the adjuvant sequence to the N-terminal of the vaccine sequence. Additionally, a GGGS linker was added to connect PADRE with the first CTL epitope, and a HEYGAEALERAG linker was used for joining the last CTL epitope with the first HTL epitope. All adjuvants, when combined with epitopes, exhibited antigenicity and non-allergenicity. The various attributes of the four possible vaccine constructs designed with different adjuvants are shown in Table 2.
The graphical representation of the secondary structure configuration of the multi-epitope vaccine. The figure was obtained from the prabi server (https://npsa-prabi.ibcp.fr/cgi-bin/npsa_automat.pl?page=/NPSA/npsa_gor4.html).
The Large ribosomal subunit protein bL12 adjuvant construct showed 55.66% alpha helix, 16.67% extended strand, 27.67% random coil and an instability index of 22.11 (Table 2; Fig. 4). Considering the definition of random coil as protein regions lacking a specific secondary structure, it is noted that the adjuvant with Large ribosomal subunit protein bL12, which exhibited a lower percentage of random coil and lower instability index compared to the others (Table 2), was deemed to be the most suitable vaccine construct. The final vaccine construct with all the selected epitopes, linkers and adjuvants arranged in the construct is depicted in Fig. 5.
Multi-epitope vaccine construct and sequence. The construct includes an adjuvant, PADRE, CTL, HTL, and B-cell epitopes, connected by different linkers. The figure was prepared using Microsoft PowerPoint 2013.
Physicochemical properties and solubility analysis of the vaccine
Different physicochemical properties of the designed vaccine such as molecular weight, pI, half-life in eukaryotic as well as prokaryotic hosts, solubility, stability, GRAVY and aliphatic score were computed using bioinformatics tools as described in Methods section. The findings of the physicochemical analyses are presented in Table 3.
Evaluation of the antigenicity and allergenicity of the vaccine construct
The antigenicity of the vaccine construct was evaluated using the VaxiJen v2.0 and ANTIGENpro servers, which provided probabilities of 0.6256 and 0.834190, respectively. The AllerTOP v.2.0 server indicated that the vaccine was not allergenic. Overall, the vaccine construct was deemed a suitable candidate due to its strong antigenicity and non-allergenic nature.
Tertiary structure modelling and validation of the multi‑epitope vaccine
The 3D structure of the vaccine protein was predicted using trRosetta, generating five different models. Among them, one model with a high template modelling (TM) score of approximately 0.461 was chosen for further analysis. The tertiary structure of the vaccine is depicted in Fig. 6A. The TM-score, ranging from 0 to 1, signifies the accuracy of the model’s topology, and a score surpassing 0.5 typically indicates correctly predicted topology61. While our TM-score is close to 0.5, suggesting a reasonably accurate model, a two-step refinement was initiated. The model was subjected to refinement using ModRefiner, and the downloaded refined model exhibited a significantly improved TM-score of 0.9823. Further refinement on the GalaxyRefine server resulted in five models. Based on the quality scores, “model 1” was chosen as the final 3D model for further characterization, exhibiting parameters of GDT-HA (0.9733), RMSD (0.362), and MolProbity (1.460).
The generated models underwent inspection using ProSA (Fig. 6B) to identify potential errors in the 3D structures. ProSA assigned a Z-score to each model; if the score deviated from the range typical for native proteins, it suggested potential structural errors. The Z-score represented the energy separation between the model and the average of misfolded structures in standard deviation units62. The generated model’s Z-score hovered around the borderline of that of native proteins, indicating a cautious assessment. Additionally, the 3D structure underwent validation through the online server SAVES v6.0. The ERRAT value for the refined models was determined to be 95.319 (Fig. 6C). A protein model with an ERRAT score exceeding 50 was deemed of excellent quality63. The Ramachandran plot revealed that 94.9% of residues resided in the core regions, 4.0% in additional allowed regions, 0.4% in generously allowed regions, and only 0.7% in disallowed regions (Fig. 6D), affirming the overall good quality of the predicted model64.
Evaluation and validation of the modelled 3D structure of the designed vaccine. (A) Displays the 3D structure of the vaccine construct modelled using trRosetta and visualized using PyMOL 3.1, (B) Shows the Z-score plot of the modelled vaccine structure obtained using ProSA server, (C) and (D) Show the ERRAT score plot and Ramachandran plot respectively both obtained using SAVES v6.0 server, (E) Disulfide engineering of the vaccine protein, visualized using PyMol 3.1, with residue pairs displayed in ball-and-stick representation and highlighted in magenta, (F) Visualization of discontinuous epitopes on the 3D structure of the vaccine construct, with the epitopes highlighted in orange using PyMol 3.1.
Disulfide bond engineering in the multi-epitope vaccine
The Disulfide by Design 2.13 server identified 32 potential residue pairs in the refined vaccine construct model capable of forming disulfide bonds (Supplementary Table S5). According to Craig and Dombkowski31, in their study on 1,505 native disulfide bonds across 331 non-homologous proteins, the χ3 angle typically peaks at -87° and + 97°, with around 90% of naturally occurring disulfide bonds exhibiting an energy value below 2.2 kcal/mol. Based on these criteria, specifically a χ3 angle between − 87° and + 97° and an energy threshold below 2.2 kcal/mol, we selected two residue pairs for disulfide bond formation: Gln 60-Ala 134 and Gly 237-Tyr 247( Fig. 6E).
Discontinuous B cell epitope prediction
The ElliPro server identified discontinuous B-cell epitopes within the vaccine’s tertiary structure (Fig. 6F). The predicted epitopes had scores ranging from 0.515 to 0.984 (Supplementary Table S6).
Interaction of the selected T cell epitopes with MHC molecules
Molecular docking was applied to determine if the MHC peptide binding groove could recognize the selected epitopes. Epitopes were docked with globally prevailing MHC Class I and II alleles, detected by population coverage studies, which indicated that HLA-A*03:01 and HLA-DRB1-1501 had the highest coverage. HPEPDOCK is an online server, and docking was performed on the same using all the epitopes; every epitope possessed different energy values with its respective HLA alleles. For MHC class I, YRQRARLLK exhibited a binding energy of -216.537 kcal/mol, while KLQDAGVYR showed -202.923 kcal/mol. Other epitopes included ISYGGADYK and KRITVKVNA, with binding energies of -218.253 kcal/mol and -210.788 kcal/mol, respectively. Additionally, ITVKVNAPY had a binding energy of -210.971 kcal/mol. For MHC class II, the binding energies were as follows: the epitope DLYVVEYGSNMTIEC had a binding energy of -233.335 kcal/mol, while YGGADYKRITVKVNA and GGADYKRITVKVNAP exhibited binding energies of -213.556 kcal/mol and -207.665 kcal/mol, respectively. The complete data is provided in Supplementary Table S7 while the graphical output is shown in Fig. 7.
Interaction of selected peptides with MHC alleles. (A-F) Cytotoxic T lymphocyte (CTL) epitopes of the vaccine protein interact with HLA-A*03:01. (G-J) Helper T lymphocyte (HTL) epitopes of the vaccine protein interact with HLA-DRB1-1501. Panels A and G show co-crystal peptide docking with respective alleles for a visual representation of how the vaccine epitopes bind with their corresponding MHC molecules, visualized using PyMol 3.1.
Interaction of the designed vaccine with TLR4
The identification of protein binding and hydrophobic interaction sites on the protein surface was conducted using the CASTp server. A binding pocket, located between residue 32 and 266, was identified as a potential binding site for TLR4. The molecular surface area of this pocket was 957.918 Å2, with a molecular surface volume of 1053.979 ų. Molecular docking between the final vaccine construct model and TLR4 was then conducted using the ClusPro server. The best-docked complex, which showed the highest binding affinity with a total free energy of -1324.4 kcal/mol, was chosen. The optimal docking configuration between the final construct model and the TLR4 complex is shown in Fig. 8. Visualization and analysis of the docked complex were performed using the PDBsum server.
The CASTp server identified specific amino acids within the binding pocket of TLR4, including Val32, Thr37, Tyr38, Gln39, Asn58, Leu59, Asp60, Val82, Asp84, Thr106, Ile108, Thr110, Lys130, Leu131, Val132, Val134, Glu154, Leu155, Asn156, Ala158, His159, His179, Leu180, Asp181, Ser183, Ser184, Ser207, Leu208, Asp209, Ser211, Leu212, Lys230, Thr232, Arg234, Phe263, and Glu266. Post PDBsum analysis, it was revealed that Gln115, Asn137, Gln163, Lys186, Asp238, Glu266, Gly267, Glu135, Ala139, His159, Asn160, Leu161, Ser184, Asn185, Leu212, Pro214, Arg234, Asn235, Phe263 amino acids of TLR4 interacted with the vaccine construct (Fig. 8). Specifically, Glu266, His159, Ser184, Leu212, Arg234, Phe263 were found to be situated in the binding pocket, indicating their involvement in binding with the vaccine construct. Conversely, amino acids from the vaccine construct, including Lys295, Tyr291, Ser289, Val286, His288, His279, Asp283, Leu284, Gly280, Arg294, Gln287, Lys285, Glu282, Glu281, were identified to interact with TLR4. The vaccine constructs form interactions with TLR4 through 9 hydrogen bonds, 3 salt bridges, and 154 non-bonded contacts. Additionally, the vaccine construct establishes bonds with MD-2 (also known as lymphocyte antigen 96) involving 8 hydrogen bonds, 2 salt bridges, and 245 non-bonded contacts. This binding of the vaccine construct with TLR4 provides optimal immune response initiation and guarantees vaccine effectiveness.
The interaction between the designed vaccine construct and the TLR4 receptor in the docked complex. PyMol 3.1 was used to generate surface-filled structures, and the PDBsum (https://www.ebi.ac.uk/thornton-srv/databases/pdbsum/Generate.html) was used to obtain 2D interaction figures.
Molecular Dynamic Simulation
The development of vaccines leveraging innate immune recognition mechanisms has been a focal point in immunological research. The RMSD analysis indicated that the system reached equilibrium at approximately 35 ns. The RMSD of the entire complex was analysed by superimposing it onto the respective structures, which helped to elucidate the compact behaviour of the vaccine subunit complexed with the TLR4 domain and the RMSD analysed after system reached to equilibrium. The RMSD value for the TLR4 ranged between 0.25 and 0.5 nm, black trajectory while the RMSD for the TLR4-vaccine complex spanned between 1 and 1.45 nm, red trajectory. The vaccine superimposed on the TLR4 complex showed an RMSD in increasing trend between 0.5 and 2.5 nm till 20 ns and decrease suddenly till 30 − 35 ns of simulation. Therafter the vaccine construct achieved stability throughout the simulation period and the RMSD ranged between 1.75 and 2.25 nm, green trajectory. The smaller deviations in the RMSD reflects the robust bindings of vaccine construct with the TLR4. The vaccine subunit alone exhibited an RMSD range of 1.5 to 1.75 nm, blue trajectory. Throughout the simulations, the RMSD values for the vaccine and TLR4 complex remained relatively stable, maintaining linearity (Fig. 9A). For a more in-depth analysis, cluster analysis was performed (Fig. 9B) with an RMSD-based cutoff value of 2 Å. For the whole TLR4-vaccine complex, the RMSD ranged from 0.112 to 1.659 nm, with an average RMSD of 0.611 nm. A total of 1758 structures were included in the matrix, and the energy of the matrix was 50.34. The analysis identified 339 clusters within the given cutoff (Fig. 9C). For the TLR4 complex alone, the RMSD ranged from 0.0939 to 0.753 nm, with an average of 0.288 nm. This matrix also included 1758 structures, and the energy was 29.43, with 30 clusters identified (Fig. 9D). For the vaccine subunit, the RMSD ranged from 0.113 to 1.961 nm, with an average of 0.729 nm. The energy for this matrix was 56.93, and 481 clusters were identified (Fig. 9E).
Molecular dynamics simulation of the vaccine construct and TLR4 with cluster analysis. (A) Graphical representation of the RMSD for the vaccine construct and TLR4 complex, highlighting the stability of the system over the course of the simulation, (B) MD analysis plots showing the RMSD matrix for cluster formation, displaying the number of clusters identified from this analysis, (C) RMSD matrix cluster plot for the TLR4-vaccine complex, (D) RMSD matrix cluster plot for TLR4 alone, (E) RMSD matrix cluster plot for the vaccine construct, illustrating the conformational variations and cluster distribution. Plots were generated using XMGrace 5.1.25.
An in-depth examination of the structural flexibility and local dynamics within the TLR4 complexed with the multiepitope vaccine was performed through root-mean-square fluctuation (RMSF) analysis. The RMSF provides insights into the flexibility of different regions within the protein complex, which is crucial for understanding the dynamic behaviour and functional implications of the vaccine-receptor interaction. For chains A and C (representing TLR4), the RMSF average values observed were within the range of 0.2 nm for TLR4 Chain A and for Chain C the RMSF analysed were ranged between 0.2 and 0.45, indicating general stability across the protein backbone with localized regions of flexibility (Fig. 10A &B). Notably, the analysis revealed that the higher fluctuations were predominantly localized at the terminal ends of each chain, a common characteristic in protein dynamics attributed to the inherent flexibility of terminal regions. This flexibility could play a significant role in facilitating the necessary conformational adjustments for optimal interaction between the vaccine construct and TLR4. In contrast, the 3D construct of the vaccine subunit exhibited a broader range of fluctuations with 5 different transitions ranged amino acids from 1 to 40, 60–70, 80–180, 180–240 and 250–320, with RMSF values spanning from 1 to 1.75, 0.4–1.25, 0.5–1.1, 0.25–0.75, and 0.25–0.5 nm respectively (Fig. 10C). The fluctuation within the vaccine subunit underscores its dynamic nature, potentially reflecting the presence of highly flexible regions that might be crucial for its interaction with the immune receptor and possibly its immunogenicity. The RMSF analysis sheds light on the dynamic landscape of the TLR4-vaccine complex, highlighting regions of both stability and flexibility within the complex. The higher fluctuations observed at the terminal ends of TLR4 and within certain regions of the vaccine construct suggest areas that may undergo significant conformational changes, essential for the biological function and interaction of the complex. Such flexible regions could be key in enabling the vaccine to effectively mimic natural immunogenic structures, thus enhancing receptor engagement and downstream signalling. Furthermore, the differential flexibility patterns observed between the TLR4 chains, and the vaccine subunit may reflect the distinct roles these components play within the complex. While the relative stability of TLR4 suggests a robust scaffold for interaction, the flexibility within the vaccine construct might facilitate the exposure of epitopes and binding sites, crucial for effective immune recognition and response activation. The RMSF analysis provides valuable insights into the structural dynamics and flexibility within the TLR4-vaccine complex, revealing critical aspects of its functional interaction.
Molecular dynamics (MD) analysis of vaccine construct interaction with TLR4. (A-C) RMSF analysis, indicating the flexibility of amino acid residues in the vaccine construct throughout the simulation, (D) RoG analysis, showing the structural compactness and stability over time, (E) Hydrogen bond analysis, displaying the number of hydrogen bonds formed between the vaccine construct and TLR4. Plots were generated using XMGrace 5.1.25.
During the 175 ns of MD simulations the linearity in RoG indicates a transition towards a more compact and stable structure of the complex, suggesting effective folding and interaction dynamics of the vaccine with TLR4. The linearity in the RoG during the initial phase of the simulation highlights the conformational adjustments the vaccine-TLR4 complex undergoes to achieve a stable and functionally viable configuration (Fig. 10D). A more compact structure, as suggested by the reduced RoG, could enhance the interaction efficacy between the vaccine epitopes and TLR4, potentially facilitating the receptor’s activation and subsequent signal transduction pathways critical for initiating an immune response. The stabilization of the RoG values after 15 ns corroborates the RMSD findings, further confirming that the system reaches equilibrium and maintains structural integrity throughout the remainder of the simulation period. This trend in RoG is indicative of the molecular interactions and folding processes that are critical for the vaccine’s functionality. The compaction of the complex might also influence the exposure and orientation of key epitopes and interaction sites, which are essential for the immunogenicity of the vaccine. The correlation between the compactness of the vaccine-TLR4 complex, as indicated by the RoG, and its stability and binding interactions, underscores the importance of structural dynamics in vaccine design and efficacy. The RoG analysis complements the RMSD findings, providing a comprehensive view of the vaccine-TLR4 complex’s structural dynamics during MD simulations. The observed linearity in RoG, culminating in a stable and compact structure during simulations, highlights the potential of the multiepitope vaccine to effectively interact with TLR4. Furthermore, hydrogen bond analysis shows that the interaction is stable with an interaction strength of 7–17 hydrogen bonds with TLR4 as depicted in Fig. 10E. These results support the idea that the vaccine construct exhibits a constant binding affinity throughout the entire simulation process, suggesting its suitability as a candidate for further experimental validation and development.
To handle the big data, we employed the post-MD simulations analysis such as principal component analysis (PCA) to capture the motions of the simulated TLR4-vaccine construct complex with an energy matrix. This was achieved by reducing the dimensionality of the trajectory and suggesting the stability of each frame of the TLR4 and Vaccine Construct. Individually, the motions of TLR4 Chain A and C, the vaccine construct, and the whole TLR4-vaccine construct complex for 175 ns were conducted to comprehensively explore the simulated complexes’ structural dynamics and clustering patterns (Fig. 11A-C). We further conducted a comprehensive analysis to assess the Gibbs energy landscape of TLR4 Chain A and C, the vaccine construct, and the whole TLR4-vaccine construct complex. This analysis involved calculating potential energy or overall energy profiles for individual complexes specified above, which provided insights into the interactions within their respective environments throughout the 175 ns simulation period. The free energy landscape (FEL) of TLR4-vaccine construct complex TLR4, and the vaccine construct, were 21.9, 20.3, and 24.9 kJ/mol respectively. The colors of the FEL histogram showed the conformations that occurred during simulations (Fig. 11D-F). These findings align with previous studies emphasizing the importance of receptor-ligand stability and specific residue interactions in vaccine efficacy65. This study demonstrates that the designed multiepitope vaccine exhibits stable binding and specific interactions with TLR4, suggesting its potential effectiveness in eliciting an immune response.
PCA and Gibbs free energy landscape analysis of TLR4 and vaccine construct. PCA was performed to assess the clustering of the TLR4-vaccine complex (A), TLR4 alone (B), and the vaccine alone (C). The Gibbs free energy landscape, color-coded to reflect the system’s energy states, was plotted using the first two principal components (PC1 and PC2), with lower energy systems indicated by deeper blue regions on the contour map. Panels D, E and F show the Gibbs free energy landscapes for the TLR4-vaccine complex, TLR4 alone, and the vaccine alone, respectively. Plots were generated using XMGrace 5.1.25.
The RMSD of the complexes to be used for analysis to find the overall superposition of the vaccine-TLR4 complex with time. The RMSD values were observed at different time points as: 1 ns = 14.49, 25 ns = 6.36, 50 ns = 6.51, 75 ns = 6.07, 100 ns = 5.15, 125 ns = 6.17, 150 ns = 3.42, and 175 ns = 0. Figure 12A and H describe these, while Fig. 12I shows the superposition and alignment of TLR4 complex along with vaccine subunit at various time intervals: 1 ns, 25 ns, 50 ns, 75 ns, 100 ns, 125 ns, 150 ns, and 175 ns. The RMSD of each complex in relation to other aligned complexes was calculated from the isolated structures at various time intervals.
The insights gained from the MD simulations provide valuable information for the further design and optimization of multiepitope vaccines targeting TLR4. Future studies should focus on experimental validation of the vaccine’s immunogenicity and protective efficacy in vivo. Future studies should aim to correlate these structural dynamics with immunogenic response outcomes, paving the way for the optimization of vaccine designs targeting innate immune receptors like TLR4.
RMSD analysis and superposition of the vaccine-TLR4 complex over time. (A-H) Shows different conformational changes over time, (I) shows the illustration of the superposing and alignment of the TLR4 complex with the vaccine subunit at different time intervals: 1 ns, 25 ns, 50 ns, 75 ns, 100 ns, 125 ns, 150 ns and 175 ns. The colour red is used to denote the complex at 25 ns, while other timepoints are coloured differently, as can be seen in this figure: cyan at 50 ns, orange at 75 ns, green at 100 ns, purple at 125 ns, silver at 150 ns, and pink at 175 ns. From the isolated structures at different time intervals, the RMSD of each complex was determined with respect to other aligned complexes. Images were generated using VMD 1.9.3 software.
Free energy binding of vaccine construct with TLR4
We have employed the MM-PBSA approach to calculate the binding energy of the vaccine construct to TLR4 receptor. The vaccine construct has shown efficient binding free energy calculated from 201 after every 1000 frames till 175 ns of simulation. This method concludes the present designed vaccine construct against TLR4 would be highly effective as the binding energy reaches to -896.74 ± 6.51 kcal/mol. The other components from where the free binding energy was calculated are illustrated in below Table 4. The non-covalent forces between atoms due to attraction at short distances know as Van der Waals (VDW) Interactions. Comparably strong van der Waals interactions can lead to significant negative contribution to binding energy, favouring binding was found significant in the present case. The electrostatic energy arise from charged amino acid residues interacting with each other or with oppositely charged amino acid. The strong electrostatic attractions can significantly lower the binding energy was observed in the TLR4 and vaccine construct complex.
Furthermore, we have calculated the contribution of each amino acids of TLR4-vaccine construct complex. The interaction between the vaccine construct (chain V) and TLR4 (chains A and C) in terms of hydrogen bonds and their contribution energies obtained from MM-PBSA calculations. MM-PBSA gives assistance to the binding free energy between biomolecular complexes, which is crucial for understanding how stable the vaccine construct might be in interaction with TLR4. The contribution energy of interactome, amino acids take part in hydrogen bonding between vaccine construct and TLR4 were assessed in detail. The contribution binding energy from the interactome was observed among amino acids from vaccine construct, TLR4 chain A and from chain C were illustrated in Table 5 respectively. These interactions collectively stabilize the binding between Chain V and its partner (TLR4). Lower (more negative) energy values suggest stronger, more favorable binding.
Immune simulation for vaccine efficacy
The immune simulation results from the C-ImmSim server match actual immunological responses. The initial response is marked by increased IgM levels, as illustrated in Fig. 13A. After the second and third vaccine doses, there is a significant rise in the levels of IgG1, IgG1 + IgG2, IgM, and IgG + IgM antibodies (Fig. 13A). The IgM + IgG reached its peak after the third dose at approximately 160,000, and the IL-2 level after the second dose is around 900,000, indicating a robust humoral immune response. Remarkably, the antigen population decreases after the second exposure, indicating enhanced clearance by the immune system. B-cell population, particularly B-memory cells (> 700 cells per mm3), experiences an increase peaking after the third dose (Fig. 13B). Similarly, cytotoxic T-cell and helper-T cell populations exhibit elevated responses with corresponding memory development lasting for several months (Fig. 13C-D). The levels of IFN-γ and IL-2 significantly increased following the first vaccine dose and remained at peak levels with subsequent antigen exposures (Fig. 13E). The Simpson index, D, indicative of diversity, predicts a diverse immune response for this vaccine candidate, likely attributable to the inclusion of different epitopes.
The in-silico simulation of the immune response to the designed vaccine construct, as predicted by the C-ImmSim (https://kraken.iac.rm.cnr.it/C-IMMSIM/index.php) server. (A) The production of IgG1 + IgG2 and IgM indicates the advancement of secondary and tertiary immune responses, (B) The B-cell population following three injections, (C) Shows elevated levels in the populations of active T-cytotoxic cells and (D) T-helper cells per state following the injections, respectively, (E) Production of cytokines and interleukins along with the Simpson Index as a measure of the immune response.
Codon optimization and in silico cloning of the final vaccine construct
For achieving maximal expression of the recombinant vaccine, the amino acid sequence was codon optimized based on the host expression system. The JCat tool was employed for the back translation and codon optimization of the multi-epitope vaccine for expression in E. coli (K12) host. The optimized nucleotide sequence (954 bases) of the vaccine exhibited a CAI of 1.0 and a GC content of 49.58%. In order to create sticky ends following restriction digestion, BamHI and HindIII sites were added to the 5’ and 3’ ends of the vaccine sequence, respectively. To clone the vaccine construct into the pET28a (+) vector, Snapgene software was used (Fig. 14).
In silico cloning of the vaccine construct was performed between the BamHI and HindIII restriction sites of the pET-28a (+) vector using SnapGene 7.0.2 software free trial (https://www.snapgene.com/free-trial/). The red segment represents the vaccine sequence, while the black segment denotes the backbone of the pET-28a (+) vector.
Discussion
Immune checkpoint blockade using antibodies is the most explored immunotherapeutic strategy, with several FDA-approved mAbs available in clinics. In the context of immune checkpoints, PD-L1 is a promising candidate for vaccine development owing to its low expression in healthy tissues and overexpression on cancer cells4. Moreover, there is a pressing requirement to devise safer, more precise and effective therapy for cancer treatment, which is being attempted through developing multi-epitope vaccines. The use of immunoinformatics approach to predict prospective epitopes and design novel multi-epitope vaccines with suitable properties can expedite this process. These vaccines provide unique benefits over their monovalent counterparts because they can more effectively stimulate humoral, cellular, and innate immune responses in concert66.
There have been several attempts to create prospective PD-L1 vaccines that block the interaction with PD-1 because targeting PD-L1 offers certain beneficial properties67. Several clinical studies have also been carried out; these can be found on ClinicalTrials.gov identifier NCT03381768 and NCT03042793. A novel chimeric B-cell peptide epitope vaccine against PD-L1, introduced by Guo et al. (2022)4, demonstrated high immunogenicity and antigenicity in several animal models. It produced polyclonal antibodies (IgG1, IgG2a, IgG2b, and IgG3) that inhibited tumor growth and induced apoptosis by blocking PD-1/PD-L1 interaction4.
The current study focused on using computational immunoinformatics techniques to design a multi-epitope PD-L1 vaccine. The extracellular domain of PD-L1 spanning amino acids 19 to 225, was the target antigen chosen for epitope prediction. The predicted epitopes were carefully evaluated for allergenicity, toxicity, and antigenicity. When creating multi-epitope vaccines, it is important to account for the properties of the epitopes, adjuvants and linkers, as to how they are arranged within the chimeric sequence. Epitopes were joined by linkers so that they could be combined with other chimera parts. Artificial protein fusion using linker peptides has been prompted by studies of naturally occurring multiple domain proteins68. Linkers provide benefits to multi-epitope vaccines, including decreased risk of junctional antigen formation and improved antigen presentation and processing69. However, linkers also influence other important parameters like structural rigidity and flexibility. We have utilised the following linkers: EAAAK, GGGS, AAY, HEYGAEALERAG, GPGPG, and KK. Large ribosomal subunit protein bL12, which is derived from Mycobacterium tuberculosis, has been shown in multiple studies to have an affinity for TLR415. Its Uniport accession number is G8FRW4. As a result, we added it as an adjuvant to increase the immunogenicity of the vaccine. In order to reduce potential interference from other protein regions in the adjuvant-receptor interaction, EAAAK was utilised to provide rigidity70. One of the helper T-cell epitopes, the PADRE sequence, is important for boosting CTL responses to various antigens71. However, GGGS contributed in terms of flexibility. The remaining linkers were employed primarily because of their capacity to trigger the HTL immune response (GPGPG) and serve as cleavage sites for the lysosomal (HEYGAEALERAG) and proteasomal (AAY, HEYGAEALERAG) systems72. To maintain their distinctive immunogenic activity, the B-cell epitopes were connected by a KK linker73. Ultimately, the C-terminal section received a 6xHis tag added for use in ensuing purification experiments.
The final vaccination construct was found to be basic in nature and stable within the physiological pH range, as anticipated by physiochemical analysis. Also, the vaccine’s estimated aliphatic index score showed that it is more thermostable, and its negative GRAVY suggested that it is highly hydrophilic, meaning that it can interact with water molecules. Upon expression in bacteria, the construct had been found to be soluble. The predicted continuous and discontinuous B-cell epitopes of the vaccine construct showed some possible interactions with antibodies and flexibility. Further, disulfide engineering was performed to strengthen the vaccine construct’s stability, as disulfide bonds confer protein stability in biological environments6,74,75. A molecular docking study was performed for the prediction of interaction of the designed vaccine construct with TLR4, MHC-I, and MHC-II molecules, aiming to understand the immune response elicited by the final vaccine structure. In all three molecules, it has been seen that the vaccine had strong binding inside the receptor pockets. The immune receptor TLR4 is expressed 100 times more frequently in human cervical cancer HeLa cells than other TLRs, indicating a link between TLR4 and the advancement of cervical cancer15. Many studies have reported and emphasized the importance of TLR476,77,78,79. Consequently, TLR4 was used for the vaccine’s molecular docking study. Strong interactions between the vaccine and TLR4 were shown by the molecular docking results. The stability of the vaccine construct was also assessed using MD simulation on the vaccine-TLR4 docked complex. The TLR4 complexed with vaccine submit were simulated and meticulously investigated the stability and binding pattern during simulation. The vaccine subunit sticks with the TLR4 and forms a more compact complex. This was achieved by the isolating the Vaccine-TLR4 complexes at different time intervals of the simulations.
According to Ghaffari-Nazari et al. (2015)80, the inclusion of PADRE adjuvant in a vaccine formulation containing CpG-oligodeoxynucleotides (CpG-ODN) and multi-epitope protein enhanced the immune response against lobular carcinoma by causing the expansion of CD4 + and CD8 + subpopulations that produce IFN-γ. Immunosimulation results were similar to those of Saha et al., (2022) and Rahman et al., (2023)66,81. We found that the target vaccine construct effectively elicits a significant T-cell and B-cell-mediated response in our investigation. Effective activation of cell-mediated immunity was demonstrated by the population of CTL and active HTL increasing following the first dose and further amplifying following the second. After the first dosage, a similar pattern was seen for B-cells, which was followed by increased IgM and IgG levels, indicating an antibody-mediated response. After the initial vaccination dose, IFN-γ and IL-2 levels rise noticeably and remain at their peak even after several antigen exposures. The physicochemical properties anticipated for the multi-epitope candidate indicate that heterologous expression and antigen purification are feasible processes. To address this, we have cloned the vaccine candidate codons in silico using the widely used expression vector pET28a (+) and optimised them based on the E. coli strain K12.
Previously, several studies have explored different PD-L1-based vaccine constructs, but each has certain limitations. For example, Lin et al. (2019) designed a PD-L1-based cancer vaccine using the extracellular domain of PD-L1 (PD-L1E) linked to the C-terminal of the diphtheria toxin translocation domain (DTT)67. However, their approach did not identify specific PD-L1 T-cell epitopes, limiting the evaluation of the T-cell response in tumor control. Similarly, Jørgensen et al. (2020) developed a peptide vaccine (IO103) using a 19-amino-acid sequence from the signal peptide of PD-L182, but it focused solely on this short peptide region. Guo et al. (2022)4 identified a PD-L1 B-cell epitope peptide (PDL1-Vaxx), which elicited strong immune responses and antitumor immunity4, but their study was limited to B-cell epitopes only. In contrast, our vaccine construct takes a multi-epitope approach, incorporating both T-cell and B-cell epitopes against PD-L1. This strategy provides broader immune coverage and may enhance the overall immune response, addressing the limitations of the previous vaccines by targeting multiple pathways in tumor control.
Since this work is entirely in silico, our next step is to move towards experimental validation. The future course of experiments will lead us to clone and purify the vaccine construct, followed by conducting wet lab experiments to evaluate its efficacy. These experiments will include immunological assays to assess the immune response, as well as in vivo studies to determine the vaccine’s potential in tumor control. This will help validate the in silico predictions and further develop the vaccine for potential therapeutic applications.
A key limitation of this study is that the findings are based on computational predictions, which, while informative, may not fully capture the complexities of biological systems. The in silico analysis provides a strong foundation, but experimental validation is essential to confirm the predicted immune responses, structural stability, and overall effectiveness of the vaccine construct.
In this study, we harnessed a suite of bioinformatics tools to design a multi-epitope vaccine directed against PD-L1, showcasing encouraging outcomes. Nonetheless, it’s imperative to acknowledge that this study’s scope remains confined to computational predictions and modelling. Therefore, the principal limitation lies in the indispensable requirement for subsequent experimental validation, spanning in vitro assays, animal models (in vivo studies), and eventually clinical trials. These rigorous investigations are essential to corroborate the efficacy, immunogenicity, and safety profile of the proposed vaccine candidate before its potential translation into clinical practice can be realized.
Conclusion
The findings of this study suggest that the designed vaccine against PD-L1 has favorable structural, physiochemical, and immunological properties, making it a promising candidate for further evaluation under in vitro and in vivo set ups. Immune simulation results indicate that the designed vaccine has ability to trigger remarkable immune response. The recombinant vaccine should be produced using the designed construct and tested for its binding affinity with purified TLR4 in vitro, followed by serological assays to confirm the expected immune response.
Data availability
All data generated or analysed during this study are included in this published article and its supplementary information files.
References
Zak, K. M. et al. Structural Biology of the Immune checkpoint receptor PD-1 and its ligands PD-L1/PD-L2. Structure 25, 1163–1174 (2017).
Pandey, P. et al. Revolutionization in Cancer therapeutics via Targeting Major Immune checkpoints PD-1, PD-L1 and CTLA-4. Pharmaceuticals 15, 335 (2022).
Twomey, J. D. & Zhang, B. Cancer Immunotherapy Update: FDA-Approved checkpoint inhibitors and Companion Diagnostics. AAPS J. 23, 39 (2021).
Guo, L., Overholser, J., Darby, H., Ede, N. J. & Kaumaya, P. T. P. A newly discovered PD-L1 B-cell epitope peptide vaccine (PDL1-Vaxx) exhibits potent immune responses and effective anti-tumor immunity in multiple syngeneic mice models and (synergizes) in combination with a dual HER-2 B-cell vaccine (B-Vaxx). OncoImmunology 11, 2127691 (2022).
Lahiri, A. et al. Lung cancer immunotherapy: progress, pitfalls, and promises. Mol. Cancer. 22, 40 (2023).
Rani, N. A. et al. Development of multi epitope subunit vaccines against emerging carp viruses cyprinid herpesvirus 1 and 3 using immunoinformatics approach. Sci. Rep. 14, 11783 (2024).
Rouzbahani, A. K., Kheirandish, F. & Hosseini, S. Z. Design of a multi-epitope-based peptide vaccine against the S and N proteins of SARS-COV-2 using immunoinformatics approach. Egypt. J. Med. Hum. Genet. 23, 16 (2022).
Omasits, U., Ahrens, C. H., Müller, S. & Wollscheid, B. Protter: interactive protein feature visualization and integration with experimental proteomic data. Bioinformatics 30, 884–886 (2014).
Reynisson, B., Alvarez, B., Paul, S., Peters, B. & Nielsen, M. NetMHCpan-4.1 and NetMHCIIpan-4.0: improved predictions of MHC antigen presentation by concurrent motif deconvolution and integration of MS MHC eluted ligand data. Nucleic Acids Res. 48, W449–W454 (2020).
Shey, R. A. et al. In-silico design of a multi-epitope vaccine candidate against onchocerciasis and related filarial diseases. Sci. Rep. 9, 4409 (2019).
Araf, Y. et al. Immunoinformatic Design of a multivalent peptide vaccine against mucormycosis: Targeting FTR1 protein of major causative Fungi. Front. Immunol. 13, 863234 (2022).
Zhang, H., Lund, O. & Nielsen, M. The PickPocket method for predicting binding specificities for receptors based on receptor pocket similarities: application to MHC-peptide binding. Bioinforma Oxf. Engl. 25, 1293–1299 (2009).
Saha, S. & Raghava, G. P. S. Prediction of continuous B-cell epitopes in an antigen using recurrent neural network. Proteins 65, 40–48 (2006).
Doytchinova, I. A. & Flower, D. R. VaxiJen: a server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC Bioinform. 8, 4 (2007).
Sanami, S. et al. Design of a multi-epitope vaccine against cervical cancer using immunoinformatics approaches. Sci. Rep. 11, 12397 (2021).
Gupta, S. et al. In Silico Approach for Predicting toxicity of peptides and proteins. PLoS ONE. 8, e73957 (2013).
Dhanda, S. K., Gupta, S., Vir, P. & Raghava, G. P. S. Prediction of IL4 inducing peptides. Clin. Dev. Immunol. 263952 (2013). (2013).
Dhanda, S. K., Vir, P. & Raghava, G. P. Designing of interferon-gamma inducing MHC class-II binders. Biol. Direct. 8, 30 (2013).
Bui, H. H. et al. Predicting population coverage of T-cell epitope-based diagnostics and vaccines. BMC Bioinform. 7, 153 (2006).
Gasteiger, E. et al. Protein Identification and Analysis Tools on the ExPASy server. in The Proteomics Protocols Handbook (ed Walker, J. M.) 571–607 (Humana, Totowa, NJ, doi:https://doi.org/10.1385/1-59259-890-0:571. (2005).
Hon, J. et al. SoluProt: prediction of soluble protein expression in Escherichia coli. Bioinformatics 37, 23–28 (2021).
Magnan, C. N., Randall, A. & Baldi, P. SOLpro: accurate sequence-based prediction of protein solubility. Bioinformatics 25, 2200–2207 (2009).
Magnan, C. N. et al. High-throughput prediction of protein antigenicity using protein microarray data. Bioinforma Oxf. Engl. 26, 2936–2943 (2010).
Kouza, M., Faraggi, E., Kolinski, A. & Kloczkowski, A. The GOR Method of protein secondary structure prediction and its application as a protein aggregation prediction Tool. in Prediction of Protein Secondary Structure (eds Zhou, Y., Kloczkowski, A., Faraggi, E. & Yang, Y.) vol. 1484 7–24 (Springer New York, New York, NY, (2017).
Du, Z. et al. The trRosetta server for fast and accurate protein structure prediction. Nat. Protoc. 16, 5634–5651 (2021).
Xu, D. & Zhang, Y. Improving the physical realism and Structural Accuracy of Protein Models by a two-step Atomic-Level Energy minimization. Biophys. J. 101, 2525–2534 (2011).
Heo, L., Park, H. & Seok, C. GalaxyRefine: protein structure refinement driven by side-chain repacking. Nucleic Acids Res. 41, W384–W388 (2013).
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Wiederstein, M. & Sippl, M. J. ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res. 35, W407–W410 (2007).
Flory, P. J. Theory of Elastic mechanisms in Fibrous proteins. J. Am. Chem. Soc. 78, 5222–5235 (1956).
Craig, D. B. & Dombkowski, A. A. Disulfide by Design 2.0: a web-based tool for disulfide engineering in proteins. BMC Bioinform. 14, 346 (2013).
Ponomarenko, J. et al. ElliPro: a new structure-based tool for the prediction of antibody epitopes. BMC Bioinform. 9, 514 (2008).
Rey, J., Murail, S., de Vries, S., Derreumaux, P. & Tuffery, P. PEP-FOLD4: a pH-dependent force field for peptide structure prediction in aqueous solution. Nucleic Acids Res. 51, W432–W437 (2023).
Zhou, P., Jin, B., Li, H. & Huang, S. Y. HPEPDOCK: a web server for blind peptide–protein docking based on a hierarchical algorithm. Nucleic Acids Res. 46, W443–W450 (2018).
Binkowski, T. A. & CASTp Computed Atlas of Surface Topography of proteins. Nucleic Acids Res. 31, 3352–3355 (2003).
Laskowski, R. A. et al. Structural summaries of PDB entries. Protein Sci. 27, 129–134 (2018).
Abraham, M. J. et al. High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2. GROMACS, 19–25 (2015).
Lindorff-Larsen, K. et al. Improved side‐chain torsion potentials for the Amber ff99SB protein force field. Proteins Struct. Funct. Bioinforma. 78, 1950–1958 (2010).
Ahamed, A. et al. Nonsynonymous mutations in VEGF receptor binding domain alter the efficacy of bevacizumab treatment. J Cell. Biochem. jcb. 30515 https://doi.org/10.1002/jcb.30515 (2024).
Mir, S. A., Meher, R. K. & Nayak, B. Molecular modeling and simulations of some antiviral drugs, benzylisoquinoline alkaloid, and coumarin molecules to investigate the effects on Mpro main viral protease inhibition. Biochem. Biophys. Rep. 34, 101459 (2023).
Bhattacharya, K. et al. Multi-epitope Vaccine Design against Monkeypox Virus via Reverse Vaccinology Method exploiting Immunoinformatic and Bioinformatic approaches. Vaccines 10, 2010 (2022).
Ahn, J. H. Seungjin Choi, & Jong-Hoon Oh. A new way of PCA: integrated-squared-error and EM algorithms. in IEEE International Conference on Acoustics, Speech, and Signal Processing vol. 5 V-777–80 (IEEE, Montreal, Que., Canada, 2004). vol. 5 V-777–80 (IEEE, Montreal, Que., Canada, 2004). (2004).
Rapin, N., Lund, O., Bernaschi, M. & Castiglione, F. Computational immunology meets Bioinformatics: the Use of Prediction Tools for molecular binding in the Simulation of the Immune System. PLoS ONE. 5, e9862 (2010).
Riveau, G. et al. Safety and efficacy of the rSh28GST urinary schistosomiasis vaccine: a phase 3 randomized, controlled trial in Senegalese children. PLoS Negl. Trop. Dis. 12, e0006968 (2018).
Santini-Oliveira, M. et al. Schistosomiasis vaccine candidate Sm14/GLA-SE: phase 1 safety and immunogenicity clinical trial in healthy, male adults. Vaccine 34, 586–594 (2016).
Shilling, P. J. et al. Improved designs for pET expression plasmids increase protein production yield in Escherichia coli. Commun. Biol. 3, 214 (2020).
Akhtar, M., Rashid, S. & Al-Bozom, I. A. PD – L1 immunostaining: what pathologists need to know. Diagn. Pathol. 16, 94 (2021).
Bailly, C., Thuru, X. & Quesnel, B. Soluble programmed death Ligand-1 (sPD-L1): A Pool of circulating proteins implicated in Health and diseases. Cancers 13, 3034 (2021).
Perry, E. et al. Fragment-based screening of programmed death ligand 1 (PD-L1). Bioorg. Med. Chem. Lett. 29, 786–790 (2019).
Testa, J. S. & Philip, R. Role of T-cell epitope-based vaccine in prophylactic and therapeutic applications. Future Virol. 7, 1077–1088 (2012).
Ahmad, S. M., Svane, I. M. & Andersen, M. H. The stimulation of PD-L1-specific cytotoxic T lymphocytes can both directly and indirectly enhance antileukemic immunity. Blood Cancer J. 4, e230–e230 (2014).
Mishra, S. Designing of cytotoxic and helper T cell epitope map provides insights into the highly contagious nature of the pandemic novel coronavirus SARS-CoV-2. R Soc. Open. Sci. 7, 201141 (2020).
Khan, M. S. et al. Immunoinformatics design of B and T-cell epitope-based SARS-CoV-2 peptide vaccination. Front. Immunol. 13, 1001430 (2022).
Kobayashi, H. et al. Identification of naturally processed helper T-cell epitopes from prostate-specific membrane antigen using peptide-based in vitro stimulation. Clin. Cancer Res. Off J. Am. Assoc. Cancer Res. 9, 5386–5393 (2003).
Hirata-Nozaki, Y. et al. PD-L1-specific helper T-cells exhibit effective antitumor responses: new strategy of cancer immunotherapy targeting PD-L1 in head and neck squamous cell carcinoma. J. Transl Med. 17, 207 (2019).
Galanis, K. A. et al. Linear B-Cell Epitope Prediction for in Silico Vaccine Design: a performance review of methods available via command-line interface. Int. J. Mol. Sci. 22, 3210 (2021).
Ahmad, T. A., Eweida, A. E. & Sheweita, S. A. B-cell epitope mapping for the design of vaccines and effective diagnostics. Trials Vaccinol. 5, 71–83 (2016).
Dittmer, U. et al. Role of Interleukin-4 (IL-4), IL-12, and Gamma Interferon in primary and vaccine-primed Immune responses to friend retrovirus infection. J. Virol. 75, 654–660 (2001).
Teixeira, L. K. et al. IFN-γ production by CD8 + T cells depends on NFAT1 transcription factor and regulates th differentiation. J. Immunol. 175, 5931–5939 (2005).
Wu, C. et al. IFN-γ primes macrophage activation by increasing phosphatase and tensin homolog via downregulation of miR-3473b. J. Immunol. 193, 3036–3044 (2014).
Xu, J. & Zhang, Y. How significant is a protein structure similarity with TM-score = 0.5? Bioinformatics 26, 889–895 (2010).
Heydari Zarnagh, H., Hassanpour, K. & Rasaee, M. J. Constructing chimeric Antigen for Precise Screening of HTLV-I infection. Iran. J. Allergy Asthma Immunol. 14, 427–436 (2015).
Abdi, S. A. H. et al. Multi-epitope-based vaccine candidate for Monkeypox: an in Silico Approach. Vaccines 10, 1564 (2022).
Samanta, A., Alam, S. S. M., Ali, S. & Hoque, M. Analyzing the interaction of human ACE2 and RBD of spike protein of SARS-CoV-2 in perspective of Omicron variant. EXCLI J. 21Doc610 ISSN 1611–2156 (2022). https://doi.org/10.17179/EXCLI2022-4721
Sanami, S. et al. In silico design of a multi-epitope vaccine against HPV16/18. BMC Bioinform. 23, 311 (2022).
Saha, S., Vashishtha, S., Kundu, B. & Ghosh, M. In-silico design of an immunoinformatics based multi-epitope vaccine against Leishmania donovani. BMC Bioinform. 23, 319 (2022).
Lin, Z. et al. A PD-L1-Based Cancer Vaccine elicits Antitumor Immunity in a mouse Melanoma Model. Mol Ther. - Oncolytics. 14, 222–232 (2019).
Chen, X., Zaro, J. L. & Shen, W. C. Fusion protein linkers: property, design and functionality. Adv. Drug Deliv Rev. 65, 1357–1369 (2013).
Athanasiou, E. et al. A poly(Lactic-co-Glycolic) Acid Nanovaccine based on chimeric peptides from different Leishmania infantum proteins induces dendritic cells maturation and promotes peptide-specific IFNγ-Producing CD8 + T cells essential for the Protection against Experimental Visceral Leishmaniasis. Front. Immunol. 8, 684 (2017).
Livingston, B. et al. A rational strategy to Design Multiepitope Immunogens based on multiple th Lymphocyte Epitopes. J. Immunol. 168, 5499–5506 (2002).
Bahrami, A. A. et al. Silico Approaches and Computational Designmulti-epitopepiimmunogenicoproteinrotein. Int. Rev. Immunol. 38, 307–322 (2019).
Sanches, R. C. O. et al. Immunoinformatics Design of Multi-epitope peptide-based vaccine against Schistosoma mansoni using transmembrane proteins as a target. Front. Immunol. 12, 621706 (2021).
Khan, M. T. et al. Immunoinformatics and molecular modeling approach to design universal multi-epitope vaccine for SARS-CoV-2. Inf. Med. Unlocked. 24, 100578 (2021).
Moin, A. T. et al. Immunoinformatics Approach to Design Novel Subunit vaccine against the Epstein-Barr Virus. Microbiol. Spectr. 10, e01151–e01122 (2022).
Moin, A. T. et al. In-silico formulation of a next-generation polyvalent vaccine against multiple strains of monkeypox virus and other related poxviruses. PLOS ONE. 19, e0300778 (2024).
Farhani, I. et al. Designing a multi-epitope vaccine against the SARS-CoV-2 variant basedon an Immunoinformatics Approach. Curr. Comput. Aided Drug Des. 20, 274–290 (2024).
Ahmad, S. et al. Computational design of a multi-epitope vaccine candidate against Langya henipavirus using surface proteins. J. Biomol. Struct. Dyn. 1–18. https://doi.org/10.1080/07391102.2023.2258403 (2023).
Ullah, A. et al. An in Silico multi-epitopes Vaccine Ensemble and characterization against nosocomial Proteus penneri. Mol. Biotechnol. https://doi.org/10.1007/s12033-023-00949-y (2023).
Ahmad, S. et al. In silico design of a novel multi-epitope vaccine against HCV infection through immunoinformatics approaches. Int. J. Biol. Macromol. 267, 131517 (2024).
Ghaffari-Nazari, H. et al. Improving Multi-epitope long peptide vaccine potency by using a strategy that enhances CD4 + T help in BALB/c mice. PLOS ONE. 10, e0142563 (2015).
Rahman, M. M., Masum, M. H. U., Talukder, A. & Akter, R. An in silico reverse vaccinology approach to design a novel multiepitope peptide vaccine for non-small cell lung cancers. Inf. Med. Unlocked. 37, 101169 (2023).
Jørgensen, N. G. et al. Peptide vaccination against PD-L1 with IO103 a Novel Immune Modulatory Vaccine in multiple myeloma: a phase I first-in-human trial. Front. Immunol. 11, 595035 (2020).
Acknowledgements
The authors are thankful to the Department of Biological Sciences, Aliah University, Kolkata, India for providing the necessary facilities for this work. SSMA acknowledges the Council of Scientific & Industrial Research (CSIR) Govt. of India for financial assistance in the form of Senior Research Fellowship (SRF). AS acknowledge Swami Vivekananda Merit Cum Means Scholarship for providing financial support.
Author information
Authors and Affiliations
Contributions
S.S.M.A: Data curation, Methodology, Visualization, Investigation, Software, Formal analysis, Validation, Writing – original draft; S.A.M: Methodology, Visualization, Investigation, Software, Formal analysis, Validation, Writing – original draft; A.S.: Methodology, Visualization, Investigation; B.N.: Software, Investigation, Validation; S.A.: Formal analysis, Validation, Writing – review and editing; M.H.: Conceptualization, Supervision, Validation, Writing – review and editing; All authors have agreed with the manuscript and provided their consent for publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mahafujul Alam, S., Mir, S.A., Samanta, A. et al. Immunoinformatics based designing of a multi-epitope cancer vaccine targeting programmed cell death ligand 1. Sci Rep 15, 12420 (2025). https://doi.org/10.1038/s41598-025-87063-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-87063-y