Abstract
Highly variable response shown by individuals against mosquito-borne infections suggests that host genetic factors play an important role in determining mosquito-borne disease onset. Therefore, it is necessary to determine the genetic risk of these diseases in specific populations. The current study aimed to determine the percentage of individuals in the general population carrying mosquito-borne disease susceptibility and protection-related variants. This study initially aggregated mosquito-borne disease susceptibility and protection-related variants from all publically available data and literature. Afterward, the allele frequency was calculated 1009 genetic variants of 366 genes associated with susceptibility and protection to estimate the global prevalence in multiple ethnicities (Middle Eastern, Ashkenazi Jewish, European (Non-Finnish), Latino/Admixed American, South Asian, East Asian, European (Finnish), North Asian, Southeast Asian, African American, and Swedish population). Furthermore, the cumulative allele frequency of all susceptibility and protection-related variants was calculated in diverse ethnic groups and the relationship with mosquito-borne disease-associated morbidity and mortality was examined to determine whether results are consistent with founder effect in these populations. Two prioritized genetic variants of IL-10 (rs1800871) and FcγRIIA (rs1801274) were examined in the Tehsil Haripur population to assess the genetic risks linked to susceptibility and protection against mosquito-borne diseases. The findings of this study revealed overlapping genes most implicated in mosquito-borne disease linked with susceptibility and protection across different ethnic ancestries. In the available sample size, the percentage of TC and TT genotypes in IL-10 genetic variant (rs1800871) was 12% and 88%, respectively and GA and GG genotypes in FcγRIIA(rs1801274) genetic variant were 6% and 94% respectively. Based on statistical analysis, the percentage allele frequency of IL-10 (rs1800871) variant was 0.2112% and the FcγRIIA (rs1801274) variant is 0.1128% in the current study. Additionally, this study reflects that screening of genetic variants associated with susceptibility and protection in a population gives better insights into organizing public health awareness campaigns to control diseases.
Similar content being viewed by others
Introduction
Mosquito-borne diseases (MBDs) cause serious public health concern globally. The primary pathogens transmitted by mosquitoes are chikungunya virus (CHIKV), zika virus (ZIKV), dengue virus (DENV), west nile virus (WNV), yellow fever virus (YFV), rift valley fever virus (RVFV), Japanese encephalitis virus (JEV), Plasmodium spp and Wuchereria bancrofti1. MBDs have one of the highest burdens in number of cases of morbidity and mortality annually2,3,4. Roughly half of the world’s population is at risk of MBDs5. The European Centre for Disease Prevention and Control as of October 2023, reported that over 4.2 million cases and over 3,000 dengue-related deaths have been reported from 79 countries/territories globally (https://www.ecdc.europa.eu/en/dengue-monthly). Furthermore, across the Asian population, like Bangladesh, Sri Lanka, Thailand, and Malaysia have experienced a surge in cases of MBDs6. Asia has been a hotspot of DENV and CHIKV infections mainly due to its dense human population, unplanned urbanization, and poverty7. However, the number of malarial deaths globally remains over 30-fold higher than those from DENV infection, although this ratio is less extreme if one only considers disease burden outside Africa8,9. DENV infection has resulted in important socioeconomic burdens placed on different countries around the world, including those in Latin America, especially Brazil10. More than a hundred thousand DENV infection cases are diagnosed in India annually, and about half of the country’s population carries DENV infection specific antibodies11. CHIKV infection, which mainly invades tropical and subtropical regions, is one of the recently emerging pathogens associated with severe morbidity in humans12. Since CHIKV infection arrived in Asia, it has become widely established and caused major outbreaks like the one in India with more than 1.4 million cases in 200613. The presence of ZIKV infection antibodies has been reported in Pakistan, indicating active circulation of the virus in the Sindh region. ZIKV infection is a potential cause of seasonal dengue-like illnesses observed in the country14. The high urban population density and abundance of competence vectors may result in a high risk of local transmission following the introduction of ZIKV infection through travel in Punjab, Pakistan15. In dengue-endemic countries, the recent epidemics of CHIKV infection and WNV disease have created panic among the public and are thought to provoke an outbreak of ZIKV infection in Pakistan16.
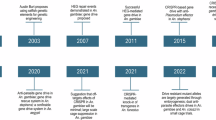
MBDs transmission depends on complex interactions between the environment and the susceptibility, exposure, and adaptive capacity of populations. Variations in climatic factors, including temperature and precipitation which modulate the spatiotemporal distribution of vectors, hosts, and pathogens17. The impact of climate change on the incidence, transmission season, duration, and spread of vector-borne diseases characterizes a serious problem18. Low- and middle-income countries with higher population density, poor healthcare systems, rapid unplanned urbanization, and global warming-induced changing climatic factors are particularly vulnerable to DENV infection19 (Fig. 1).
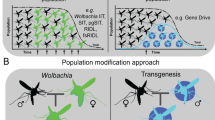
Transmission of MBDs occurs following a mosquito (Aedes aegypti or Aedes albopictus, Culex, and Anopheles mosquitoes) bite. MB infection mostly replicates in the skin, fibroblasts, and a few in liver cells, and disseminates to the liver, muscle, joints, lymphoid tissue (lymph nodes and spleen), small intestine, thymus, one marrow, kidney, and brain. The target cells are indicated for each tissue. Immunogenetic pathways are involved against MBD infections. There is evidence of MBD-specific adaptive immunity (that is, T cell and antibody-mediated responses).
Genome-Wide Association Studies (GWAS) have been conducted across various populations including African, European, Asian European, and Asian populations. The application of GWAS to African populations could offer valuable insights into the genetic pathways underlying resistance to malaria, as well as shed light on the genetic origins of related diseases20. Disease prevalence and severity exhibited variations across populations. Many studies provides robust evidence that genetic variation in human populations contributes to susceptibility to infectious diseases21. The majority of GWASs have focused on patients of European and Southeast Asian descent, with insufficient representation of studies involving individuals from other ancestral backgrounds22. Jian-Wen et al.. conducted a large GWAS on Systemic Lupus Erythematosus (SLE) in a Chinese Han population, analyzing 1,047 cases and 1,205 controls using Illumina Human610-Quad BeadChips. Replication of 78 single nucleotide polymorphisms (SNPs) in two additional cohorts (3,152 cases and 7,050 controls) identified nine new susceptibility loci (ETS1, IKZF1, RASGRP3, SLC15A4, TNIP1, 7q11.23, 10q11.22, 11q23.3, and 16p11.2) and confirmed seven previously reported risk loci23. The HLA-DR2 (DRB1*1501) and HLA-DR3 (DRB1*0301) class II genes have been found to consistently associate with SLE in many European populations, with a twofold relative risk conferred by each allele24. Similarly, GWAS from European (4,036 cases and 6,959 controls) and Chinese (1,659 cases and 3,398 controls) cohorts studies revealed that these populations share more than half of known SLE-associated genetic loci25. Furthermore, a recent study on SLE progression has identified a locus pointing to involvement of the central nervous system in disease outcome as opposed to the enrichment for immunological-related signals observed for disease susceptibility26.
Khor et al. identified genes that significantly associated with dengue shock syndrome (DSS) including MHC and MICB in the chromosome 6 and the PLCE1 on chromosome 10 region27. Susceptibility to infection and many other human diseases (including diabetes and ischaemic heart disease) arises from the complex interaction of environmental and host genetic factors21. According to Karin et al. the availability of more data is expected to reveal risk variants that are rare in specific populations, shedding light on the shared genetic susceptibility across diverse ancestries22. Furthermore, another study highlighted the genetic polymorphisms associated with the immune response to DENV infection. Fang et al. identified several genetic variants modulating host immune responses. This study highlighted the role of specific alleles in determining disease severity28. Similarly, Oliveira et al. conducted a meta-analysis on genes associated with dengue fever (DF). The study revealed certain SNPs linked to disease risk across different ethnic groups. This trans-ethnic consistency suggests that shared genetic variants may underlie susceptibility to DENV infection. These studies warrant further investigations to comprehend the genetic architecture of MBDs29. Patro et al. demonstrated that polygenic risk scores (derived from GWAS) can predict the risk of dengue hemorrhagic fever (DHF) across multiple ancestries. This emphasizes the need for an integrated approach in predicting disease risk30. In addition to DENV, CHIKV and ZIKV infections have also been subjects of GWAS investigations31. The concept of genetic susceptibility revolves around whether specific alleles consistently elevate or diminish susceptibility across various regions of the world32. Genetic variations play a crucial role in influencing susceptibility to DHF33. The DENV genetic heterogeneity (serotype and genotype) adds another dimension to the public health challenge due to the increased risk of disease severity (secondary and tertiary infection)34. In certain populations, specific genetic variants are associated with severe complications of DENV infection. Family-based analysis of whole-exome sequencing (WES) provides a high detection rate for prevalent and rare variations35. Among neurological conditions, the authors Tan et al. focused on disease survival as well as cognitive or motor decline. In one of the largest studies, researchers have identified three novel loci associated with Parkinson’s disease progression36. In Crohn’s disease, a study has identified four loci for disease progression, indicating distinct genetic contribution from disease susceptibility37.
This study initially aggregated MBDs susceptibility and protection-related variants from all publicly available information. Further, we propose to develop a framework of methods that can prioritize overlapping and unique genes and their variants to explore new associations with MBDs. The current study expanded our knowledge base regarding the most prevalent genetic variations associated with disease in the local Haripur population. To the best of our knowledge, this study reflects that screening of genetic variants associated with susceptibility and protection in a population gives better insights into organizing public health awareness campaigns to control diseases.
Materials and methods
In the current study, we employed a two-step approach to calculate allele frequencies of genetic variants related to susceptibility and protection among MBDs. Firstly, we used a computational pipeline to retrieved genetic variants, process, and calculated allele frequencies of these variants from publicly available whole genome datasets, capturing a comprehensive global perspective. Secondly, we validated our findings by performing Amplification refractory mutation system PCR (ARMS-PCR) on prioritized genetic variants to determine allele frequencies in a local population. ARMS-PCR, also known as allele-specific amplification allows to identify gene barcodes with single-nucleotide resolution and has become a widely used method in clinical practice38. ARMS-PCR is more sensitive than Sanger sequencing with its detection limit of about 1%. It is a very simple method for detecting any known mutations by using sequence-specific PCR primers that allow amplification only when the target al.lele is present within the sample39. It is based on the principle that PCR amplification is inefficient or completely refractory if there is a mismatch between the 3’ terminal nucleotide of a primer and its template sequence. This technique employs two primer pairs to amplify two alleles, one wild-type allele and one mutant allele in a single PCR reaction40. This dual approach ensured that our results for global, Asian, and South Asian populations were consistent and applicable at a local level, providing g robust validation for our computational analysis.
Data retrieval and processing
To understand the genetic spectrum of MBDs, a comprehensive search was performed to identify all previously reported disease related genetic variants associated with MBDs. Data was retrieved from Google Scholar, PubMed, Science Direct and Scopus databases using the following combinations of search terms: genetic polymorphism, genetic susceptibility, genetic predisposition, genetic determinants, susceptibility, protection, severity, single nucleotide polymorphism, mutations, variants, variations, mutants, MBDs, genetic susceptibility and protection against dengue, chikungunya, rift valley fever, zika, west nile virus, yellow fever, lymphatic filariasis and malaria infections/diseases (Fig. 2).
Schematic representation of prioritization pipeline. Screening and identification of genetic variants related to susceptibility and protection associated with MBDs.
Calculation of allele frequency from available whole genomes
Following the screening of shared and divergent genetic variants, the subsequent step involved calculating the allele frequency of each variant present in lymphatic filariasis (LF), JEV, malaria, DENV, ZIKV, CHIKV, RVF, WNV and YF infections. While GWAS has advanced our understanding of the genetic architecture of MBDs, post-GWAS functional genomic analyses are required to prioritize genes that modulate disease susceptibility and nominate candidate genes for further functional validation through ARMS-PCR.
To accomplish this, we utilized various population-specific databases known for their ability to estimate allele frequencies across diverse populations and datasets. We analyzed the available whole genome from various publicly available next-generation sequencing databases that aggregates exome and genome sequencing information from a wide variety of large-scale projects and stratify allele counts by ethnicity, including Genome Asia 100k (http://www.genomeasia100k.com), IndiGen (https://clingen.igib.res.in/indigen), SAGE (South Asian Genome), PGG.Han41, GenomeAsia 100 K Project42(https://browser.genomeasia100k.org/), Vietnamese Genetic Variation Database (https://genomes.vn/), SweGen Dataset43, NyuWA Chinese Population Variant Database (http://bigdata.ibp.ac.cn/NyuWa_variants/searchact.php#search-denote)44, gnomAD v3 (Genome Aggregation Database), and PopHuman (http://pophuman.uab.cat). These freely accessible databases provide large-scale genetic datasets from diverse populations to estimate allele frequencies associated with specific genes and their variants. By utilizing multiple resources, we aimed to ensure comprehensive coverage of various populations and regions. This allele frequency data contributes to our understanding of the genetic landscape of these diseases, helping to identify population-specific variations and potential correlations between genetic variants and disease susceptibility or protection.
Screening of prioritized genetic variants in the local population
To confirm and corroborate the initial analysis, preliminary validation of the two prioritized genetic variants was performed using ARMS-PCR. The sample size comprised fifty participants, all residents of Tehsil Haripur, Khyber Pakhtunkhwa (KPK), Pakistan. Preliminary screening of the IL-10 (rs1800871) and the FcγRIIA (rs1801274) genetic variants were performed. Ethical approval for this study was obtained from the Research Ethics/Bioethics Committee of the Directorate of Advanced Studies & Research Board (ASRB) at the University of Haripur, KPK, Pakistan. The study was conducted in accordance with ASRB’s relevant guidelines and regulations. Informed consent was obtained from all study participants. This analytical observational study was conducted from December 2022 to August 2023.
Inclusion and exclusion criteria
A random sampling technique was used to collect samples. Individuals aged ≥ 18 years, regardless of gender, with a permanent address of Tehsil Haripur were included in this study. Individuals who had been diagnosed with a genetic disease and those under 18 years of age were excluded. Venous whole blood was collected in EDTA tubes using a sterile disposable syringe. 05 mL whole blood was collected and transported at 04 °C, after which the sample was stored at 4 °C for further analysis.
DNA extraction and quantification
Genomic DNA was isolated from 200 µL of blood using the Gene JET Genomic DNA Purification Catalog no: K0721, (JETTM Thermofisher Genomic DNA Purification and Isolation kit). Genomic DNA was quantified using a nanodrop (OPTIZEN NanoDrop) spectrophotometer. All samples were stored at 4 °C until analysis. High-quality intact genomic DNA with an optical density ratio of 260/280 ~ 1.8 and 260/230 > 1.5 were used for further analysis.
ARMS-PCR amplification
We designed ARMS-PCR primers through Primer1 software (http://primer1.soton.ac.uk/primer1.html), and A-Plasmid Editor software45. Furthermore, UCSC Genome Browser (genome.ucsc.edu) was used to check the specificity and sensitivity of designed primers by running in silico PCR (Table 1).
PCR amplifications were performed in 20µL reactions containing 10 µL 2X PCR Taq Master Mix (amaR OnePCR, Cat. No. SM213-0250), 1 µL genomic DNA (1 ng/µL) template, 2 µL of 3.3 µM of tetra primers (o.5 µL for each primer), and 7 µL nuclease-Free water. The PCR regimen was as follows: initial denaturation at 95 °C for 5 min, followed by 33 cycles for 30 s at 94 °C for 01 min, 01 mint at 62–64 °C, 01 min at 72 °C, and then a final extension for 5 min at 72 °C, and finally the PCR products were maintained at 4 °C in the end. PCR products were resolved on a 2.5% agarose gel with 1× Tris–acetate-ethylenediaminetetraacetic acid (TAE) buffer, and then subjected to electrophoresis at 110 V for 30 min. The agarose gel was stained with 0.5 µg/mL ethidium bromide. Bands were visualized under UV light by Automatic Gel Imaging and Analysis System (LABTRON) and the molecular weight of the PCR products was determined by comparing them with the100 base pairs (bps) DNA marker (Cat.no FNB-500304, Fine Biotech Life Sciences).
Data analysis
The Hardy-Weinberg equation (p2 + 2pq + q2 = 1)46 was used to calculate the expected genotype frequencies for heterozygous (2pq) and homozygous (p2) genotypes, using the minor and total allele counts, where p is equal to the cumulative minor allele frequency (cMAF) of causally related variants and q is (1 − p).
Results
Identification of genetic variants showing pleiotropy for MBDs
309 genes and 931 genetic variants associated with susceptibility and 63 genes with 78 genetic variants related to protection against nine MBDs including DENV, CHIKV, ZIKV, RVF, WNV, YF, JEV, malaria and LF infections were identified through a literature search. Moreover, the analysis revealed a total of 1009 genetic variants of 366 genes associated with both susceptibility and protection against MBDs as shown in Table 2 (Fig. 3).
Representation of pleiotropic genes and their variants related to susceptibility and protection among dengue, malaria, CHIKV, RVF, WNV, YF, JEV and LF diseases.
Identification of pleiotropic genes related to susceptibility and protection in MBDs
Across various search platforms, a collective analysis has identified 21 genes with 49 variants linked with susceptibility and protection against DENV infection (Fig. 3), (Supplementary table S2) in which 27 variants of 08 genes were associated with protection and 22 variants of 13 genes were associated with DENV disease susceptibility as shown in Table 2. A total of 117 genes and 362 variants linked with susceptibility and protection against malaria including 94 genes and 336 variants related to susceptibility and 23 genes with 26 variants linked with protection against malaria as shown in Fig. 3, Table 2 and Supplementary table S5. In WNV disease a total of 99 genes and 388 variants related to susceptibility and protection were obtained, in which 95 genes and 388 variants were associated with susceptibility and 04 genes were linked with protection against WNV disease as shown in Fig. 3, Table 2 and Supplementary table S7. Among ZIKV disease a total of 26 genes and 32 variants were associated with susceptibility and protection against ZIKV infection including 25 genes and 32 variants related to susceptibility and 01 genes linked with protection against ZIKV disease as shown in Fig. 3, Table 2 and Supplementary table S9. In RVF a total of 19 genes and 70 variants were associated with susceptibility and protection, in which 16 genes and 58 variants linked to susceptibility and 03 genes, and 12 variants associated with protection against RVF as shown in Fig. 3, Table 2 and Supplementary table S6. LF infection related studies revealed that 21 genes and 37 variants were associated with disease susceptibility as shown in Fig. 3, Table 2 and Supplementary table S4. Furthermore, JEV disease related studies showed 17 genes, and 16 variants associated with susceptibility and protection including 13 variants of 14 genes were linked with susceptibility and 03 variants of 04 genes related to protection against JEV infection as shown in Fig. 3, Table 2 and Supplementary table S3. In CHIKV disease a total of 24 genes and 53 variants were associated with susceptibility and protection including 16 genes and 45 variants linked with susceptibility and 08 genes, and 08 variants related to protection against disease as shown in Fig. 3, Table 2 and Supplementary table S1. A total of 29 genes and 02 variants were identified against YF related to disease susceptibility and protection in which 12 genes and 02 variants were linked with the protection of the disease and 17 genes were related to susceptibility to the YF disease as shown in Fig. 3, Table 2 and Supplementary table S8.
Identification of common genes (susceptibility and protection) among MBD and their overlap
A total of 67 overlapping genes and variants related to susceptibility and protection among MBDs were identified as common among two or more diseases (Table 3) and (Fig. 4). IL-1β gene was common in malaria and JEV diseases. TNFα gene overlapping across malaria, DENV, ZIKV, CHIKV, JEV, and WNV diseases. Similarly, FcγRIIA/CD32 gene overlaps in malaria and DENV diseases. Moreover, the G6PD gene was shared between malaria, DENV, and ZIKV infections, whereas the HLA gene was common to malaria, ZIKV, and WNV diseases. The TLR4 gene overlaps across malaria, DENV, and LF diseases. Similarly, the TLR9 gene was shared among malaria, DENV, JEV and WNV diseases. The TIRAP gene was present in both malaria and WNV diseases (Table 3) and (Fig. 4).
Overlapping genes associated with disease susceptibility and protection against MBDs. Genes depicted in red are those shared among only two MBDs. Genes depicted in black are those shared by more than two MBDs (DENV, malaria, ZIKV, RVF, YF, CHIKV, LF, JEV, and WNV diseases).
The IRF1 gene was shared among malaria, WNV, and YF diseases. The IL-4 gene overlaps in malaria and ZIKV infections, while the IL-12B gene was shared among malaria, ZIKV, and JEV diseases. Additionally, the ICAM1 gene was found in malaria, DENV, and JEV diseases. The IFN-γ gene was common to malaria, DENV, JEV, and WNV diseases. Moreover, the MyD88 gene was shared across malaria, CHIKV, RVF, and WNV diseases (Table 3) and (Fig. 4). The IL-8 gene overlaps in malaria and DENV diseases. Similarly, the IL-10RA gene was shared between malaria and LF diseases. Additionally, the ADAR gene overlaps in malaria and YF, while the IDO1 gene was shared between DENV and YF diseases. Moreover, the MDA5 gene overlaps in YF and CHIKV diseases, and the C6 gene was present in YF, ZIKV, JEV, and DENV diseases. Furthermore, the RNF2 gene overlaps in ZIKV and RVF, while the MIR605 gene was shared between DENV and ZIKV infections. Additionally, the LIF gene was common to DENV and ZIKV infections, and the NOS3 gene overlaps in DENV and ZIKV infections. Lastly, the NOS2 gene was shared among malaria, DENV and ZIKV infections. Similarly, the CCL2 gene was common in DENV, ZIKV, JEV, and WNV disease. Additionally, the CXCL8 gene was associated with DENV and ZIKV diseases, while CXCL10 overlaps in DENV, ZIKV, and WNV diseases. The CCR2 gene was shared between ZIKV and JEV infections, and CCR5 was found in ZIKV, WNV and JEV diseases. Moreover, OAS2 was involved in DENV, CHIKV, and WNV infections, and OAS3 was found in DENV, CHIKV, and WNV diseases. IFITM3 was associated with DENV, WNV, RVF, and YF, while JAK1 shared among DENV and WNV diseases. CD2 was common in ZIKV and WNV diseases, and STAT1 overlaps in YF and WNV diseases. CTLA4 linked with WNV and LF, while TLR3 was shared among RVF, ZIKV, CHIKV, WNV, JEV, and DENV diseases. Furthermore, MBL2 was associated with DENV, WNV, and LF diseases, and IRF7 was common among YF and WNV infections. TIRAP overlaps in DENV and WNV infections, and VDR associated with DENV and WNV diseases. Lastly, OAS1 was common among DENV, CHIKV, and WNV diseases, while CD209 is implicated in DENV, ZIKV, CHIKV, WNV, and JEV infections (Table 3) and (Fig. 4).
The TLR8 gene overlaps in DENV, CHIKV, WNV, RVF, and JEV diseases. Similarly, the CXCR3 gene was shared between DENV and WNV diseases, while the CD40LG gene was associated with DENV and WNV diseases. The RNO3 gene overlaps in RVF and ZIKV infections, and the IL-6 gene was shared among RFV, ZIKV, CHIKV, and DENV diseases. Moreover, the TLR7 gene was common in RVF, ZIKV, CHIKV, WNV, JEV, and DENV diseases. The TLR2 gene overlaps in DENV and LF diseases, while the DC-SIGN gene overlaps among DENV and CHIKV infections. Additionally, the CLEC5A gene overlaps in malaria and DENV diseases, and the VDR gene was shared between DENV and WNV diseases. The JAK1 gene was common in DENV and WNV diseases, while the MICB gene overlaps in DENV and ZIKV infections. The IL-4R gene was shared between malaria and DENV infections, and the CXCL10 gene overlaps in DENV, ZIKV, and WNV diseases. The ICAM-1 gene overlaps in malaria, DENV and JEV infections, while the IL-17 gene was shared among malaria and DENV diseases. Furthermore, the IL-8 gene was shared between malaria and DENV diseases, and the HLA-DRB1 and HLA-DQA1 genes overlap in CHIKV and WNV diseases. Across diseases such as DENV, ZIKV, malaria, CHIKV, WNV, RVF, YF, LF and JEV infections the genes IL-10, TLR7, TLR3, TNFα, TLR8, CD209, CCL2, TLR9, MyD88, IFN-γ, IL-6, C6, and IFITM3 showed the maximum combination and co-occurrence, suggesting substantial interrelatedness across these diseases at the genetic level as mentioned in Table 3 and Fig. 4.
Identification of unique genes (susceptibility and protection) among MBD showing no overlap
A total of 205 unique genes and their variants related to susceptibility and protection against MBDs (malaria, WNV, ZIKV, RVF, DENV and CHIKV diseases) were identified as shown in Table 4 and Fig. 5. In malaria a total of 80 distinct genes and their variants related to susceptibility and protection were obtained including IL-13, RTN3, HBB, ATP2B4, FREM3, USP38, GYPE A, ABO, MARVELD3, HBA1, HBA2, HbC, HbE, GYPB, TLR5, TNFPD, LTA, BAT2, SLC22A4, SLC22A5, LOC441108, IL-5, RAD50, KIF3A, SEPT8, ANKRD43, IL-12 A, IL-12RB1, IL-4, IL4 VNTR, IL-1 A, CD36, CD37, CD31, PECAM-1, CR1, CD14, IL-17RC, TRIM, DARC, GBP7, IL17RD, TLR1, CTL4, IL20RA, CFTR, NOD1, TRIM5, IL-22, SPTB, ADCY9, IL-4R, ADORA2B, EMR1, GNAS, DERL3, IL-18, FPN, HLA-F, TPI1, ADRB2, BF, SCO1, HbS, FcγRIIB, FcγRIIC, FcγRIIIA, NF-kB1, FcγR, RNASE 3, ABCB1, ADORA2A, GRK5, STAT6, TNFRSF18, FOXO3A, CD40L, IRF1, IL17RE, and ABO GTG. A total of 64 genes and their variants related to susceptibility and protection related to WNV disease were found, including ANPEP, SCN1A, RFC1, HLA-DRB1, HLA-DQA1, HLA-DPA1, DQA1*01:02, CD58, FcγRIIA, RNASEL, EIF2AK2, IFIH1, PRKRA, CD28, ICOS, PDCD1, CD80, CD86, PTX3, NFKB1, CASP3, GZMA, IFNGR1, IRF5, INDO, TRAM1, JAK2, PDCD1L1, PDCD1L12, IFNB1, IFNA, TRAF1, PRF1, IFIT2, IFIT1, IFITM2, IFITM1, CD81, TRAF6, CREB3L1, RARRES3, FADD, PRKRIR, IL-10RA, CLEC4C, IRAK4, STAT2, TBK1, OASL, ISGF3G, GZMB, IFI27, SOCS1, CCL3, SOCS3, TICAM1, TNFSF14, CLEC4M, IRF3, CD40, IFNGR2, MX1, ICOSLG, and SAT1 as shown in Table 4 and Fig. 5.
Worldwide allele frequency distribution of susceptibility and protection-related genetic variants associated with MBDs and the genetic variants exhibiting the highest allele frequencies across multiple ethnic groups.
In ZIKV disease a total of 32 genes and their variants associated with susceptibility and protection were distinctly identified including DISP3, IL-12RB2, SPG7, RYR1, CAPN3, Fig. 4, COL2A1, CRB1, FGFR3, FLNB, SHH, FBN2, TNXB, KMT2A, CHD4, COL4A2, ZNF423, DNM2, LMNB2, IFNAR, TP53, LILRB1, LILRB2, HLAG, HLA-G, MDM2, VEGFA, PTGS2, TREM1, IFNR1, CXCR1 and TFRC genes. A total of 12 genes and their variants related to susceptibility and protection were uniquely found in RVF including RVFS-1, RVFS-2, RVFS-3, RNO9, CHF, DDX58, MVS, TRIF, RNO3, IFITM3, ALDH2, and IL28B. In DENV disease, a total of 09 genes along with their variants related to susceptibility and protection were distinctly identified, including CXCL11, TAP1, PLEC1, IL-6R, GYPA, CXCR3, IFNL1, RXRA, and BAK1. Additionally, a total of 06 genes and variants related to susceptibility and protection were uniquely found in CHIKV disease, including OAS, CRP, COMP, IL-2R, IL-1RN, and MIF as shown in Table 4 and Fig. 5.
Calculation of allele frequency of the overlapping of genes and their genetic variants in population-specific databases
Aiming to assess allele frequency of worldwide genetic variants related to susceptibility and protection among DENV, ZIKV, CHIKV, LF, JEV, malaria, RVF, YF and WNV diseases. Overall, 366 genes with 1009 genetic variants associated with susceptibility and protection genes were screened. These variants were stratified according to the ethnical group of the individuals carrying population sequencing data from IndiGen, SAGE, PGG.Han, GenomeAsia 100 K Project, GenomeAsia 100 K, Vietnamese Genetic Variation Database, SweGen Dataset, NyuWA Chinese Population Variant Database, PopHuman, and gnomAD database (Africans, East Asians, South Asians, Latinos, Europeans (non-Finnish, and Finnish), based on the information provided in the gnomAD database (version 3.1; all data are based on GRCh37/hg19) and bioinformatics algorithm to calculate allele frequency of these variants among multiple ethnics group as shown in Fig. 5.
Allele frequency in the Latino/Admixed American population
In the Latino/Admix American population, we calculated the allele frequency of variants related to susceptibility and protection. The analysis found that the IL-6 (rs1800795), OAS2 (rs15895), MICB (rs3132468), OAS3 (rs2285932), and TLR7 (rs179010) variants showed 0.8085%, 0.8271%, 0.8203%, 0.8326% and 0.8511% allele frequency respectively. Among these variants the TLR7 gene (rs179010) variants showed the highest allele distribution about (0.8511%), among the Latino/Admixed American population, while almost absent among other ethnic groups in the world like in South Asia, America, Oceania, Africa, West Eurasia, Southeast Asia, Northeast Asia) based on the data recorded in gnomAD. Furthermore, the G6PD gene (rs1050828) variant showed the least allele frequency about 0.004137% in the Latino/Admix American population as mentioned in Fig. 5.
Allele frequency in the African/African American population
In the African/African American population the analysis revealed that IL-6 (rs1800795), OAS2 (rs15895), OAS3 (rs2285932), CXCR3 (rs2280964), TLR7 (rs179010) and MICB (rs3132468) variants showed 0.927%, 0.9414%, 0.9481%, 0.83755%, 0.842%, and 0.8093% allele frequency respectively. Among these variants, the OAS3 variant (rs2285932) was highly prevalent in the African/African American population, with an allele frequency of 0.9481%, while this variant was absent in the South Asian population. Furthermore, in the Latino/Admix American population, the TLR4 gene variant (rs4986791) exhibited a lower allele frequency (0.01448%) as mentioned in Fig. 5.
Allele frequency in the Ashkenazi jewish population
In the Ashkenazi Jewish population, the variants of MICB (rs3132468), TLR3 (rs6552950), IL-10 (rs1800872), and TLR8 (rs5741883) gene have allele frequencies of 0.8093%, 0.8576%, and 0.7138%, respectively. Notably, TLR3 (rs6552950) showed a significantly high prevalence among Ashkenazi Jewish population about 0.8576%, while it was absent in the South Asian population. Furthermore, the G6PD gene variant (rs1050828) exhibited no allele frequency (0%) in the Ashkenazi Jewish population as mentioned in Fig. 5.
Allele frequency in the European (Finnish) population
In the European (Finnish) population, the allele frequencies of genetic variants including OAS3 (rs2285932), OAS2 (rs15895), TLR3 (rs6552950), OAS1 (rs10774671), TLR8 (rs5741883), and FcγRIIA (rs10774671) were observed at 0.7999%, 0.7825%, 0.7597%, 0.7366%, 0.7374%, and 0.7366%, respectively. Among these variants, the OAS3 variant (rs2285932) demonstrates a notable prevalence at 0.7999% in this population. Furthermore, it was observed that (0%) allele frequencies for both variants including CCL2 (rs1024611) and G6PD (rs1050828) within the European (Finnish) population as mentioned in Fig. 5.
Allele frequency in the European (non-Finnish) population
In the European (non-Finnish) population, the allele frequencies of the IL-10 (rs1800872), TLR3 (rs6552950), TLR8 (rs5741883), and MICB (rs3132468) variants were reported at 0.7646%, 0.7546%, 0.733%, and 0.743%, respectively. Notably, the IL-10 gene variant (rs1800872) demonstrates a significant prevalence with an allele frequency of 0.7999%. Additionally, the G6PD (rs1050828) variant exhibited the lowest allele frequency of about 0.0001946%, marking it as among the least frequent variants observed within this ethnic group as mentioned in Fig. 5.
Allele frequency in OTHERS population
In the context of the gnomAD (Genome Aggregation Database), “OTHER population” refers to individuals whose ancestry or ethnicity does not fall clearly into the predefined major ancestry groups. These groups typically include populations like African, Latino, East Asian, South Asian, and European47. In the population designated as OTHERS, the analysis indicates allele frequencies for the OAS3 (rs2285932), OAS2 (rs15895), and IL-10 (rs1800871) gene variants of 0.7109%, 0.7021%, and 0.7132%, respectively. Notably, the OAS3 variant (rs2285932) was more prevalent in this population, exhibiting an allele frequency of 0.7109%. Additionally, within the OTHERS population, the G6PD (rs1050828) variant showed a minimal allele frequency of 0.003005% as mentioned in Fig. 5.
Allele frequency in the American population
In American population the observed genetic diversity showed that TLR3 (rs6552950), MBL2 (rs1800450), OAS1 (rs1131454), CD209 (rs4804803), OAS3 (rs2285932), OAS2 (rs15895), TLR3 (rs3775291) and IL-6 (rs1800795) gene variants revealed 1.01838%, 1.16855%, 1.268665%, 1.36877%, 0.980769%, 0.980769%, 0.96832% and 0.980769% allele frequency respectively. Among these variants, the CD209 (rs4804803), was highly prevalent in the American population, with an allele frequency of 0.7109%. Furthermore, in the American population, the MyD88 (rs6853) variant exhibited no allele frequency about (0%) as mentioned in Fig. 5.
Allele frequency in the oceanian population
In the Oceanian population, genetic diversity analysis revealed that allele frequencies of 1% for OAS2 (rs15895), 1% for OAS3 (rs2285932), and 1.13526% for CD209 (rs4804803) gene variants. Notably, both OAS2 and OAS3 variants are highly prevalent among Oceanians, with an allele frequency of 0.7109% for each. Moreover, within this population, the TNF gene variants (rs1800629), TLR4 (rs4986791), and TLR9 (rs5743836) exhibited an absence of allele frequency (0%) as mentioned in Fig. 5.
Allele frequency in the African population
In African population the observed genetic diversity showed that CD209 (rs2287886), CD209 (rs4804803), OAS1 (rs10774671), OAS2 (rs15895), OAS3 (rs2285932) and MBL2 (rs1800450) gene variants exhibited 1.05717%, 1.095681%, 0.985576%, 0.985578%, 0.971153%, 0.941646% allele frequency respectively. Among these variants, the OAS2 and OAS3, both were highly prevalent in the African population, with an allele frequency of 0.7109%. Furthermore, TLR4 (rs4986791) exhibited the least allele frequency (0.004807%) in the African population compared with other ethnic groups as mentioned in Fig. 5.
Allele frequency in the West eurasian population
In the West Eurasian population, the observed genetic diversity showed that CD209 (rs2287886), OAS1 (rs10774671), and MBL2 (rs1800450) gene variants exhibited 0.9084963%, 0.847522%, and 0.786549% allele frequency respectively. Among these variants, the OAS2 and OAS3 were both highly prevalent in the West Eurasian population, with an allele frequency of 0.7109%. Furthermore, TLR4 (rs4986790) exhibited the least allele frequency (0.070175%) in the West Eurasian population compared with other ethnic groups as mentioned in Fig. 5.
Allele frequency in Sweden’s population
In Sweden’s population, the observed genetic diversity showed that TLR3 (rs6552950), OAS1 (rs10774671), CD209 (rs2287886), MICB (rs3132468) and FcγRIIA (rs10774671) gene variants exhibited 0.7425%, 0.6275%, 0.6265%, 0.714% and 0.713% allele frequency respectively. Among these variants, the TLR3 (rs6552950) was highly prevalent in the Sweden population, with an allele frequency of 0.7425%. Furthermore, TNF (rs361525) exhibited the least allele frequency (0.034%) in the Swedish population compared with other ethnic groups as mentioned in Fig. 5.
Allele frequency in the southeast Asian population
In the Southeast Asian population, the observed genetic diversity showed that IL-6 (rs1800795), OAS2 (rs15895), OAS3 (rs2285932), CTLA4 (rs231775) gene variants exhibited 0.988439%, 0.994219%, 0.91763%, and 0.992020 allele frequency respectively. Among these variants, the CTLA4 (rs231775) was highly p333revalent in the Southeast Asian population, with an allele frequency of 0.992020%. Furthermore, TLR9 (rs5743836) exhibited the least allele frequency (0.001445%) in the Southeast Asian population compared with other ethnic groups as mentioned in Fig. 6.
Genetic variants containing highest allele frequency in the South Asian population.
Allele frequency in the northeast Asian population
In Northeast Asian population the observed genetic diversity showed that CD209 (rs2287886), OAS1 (rs10774671), OAS1 (rs1131454), MBL2 (rs1800450), TLR3 (rs3775296), TLR3 (rs3775290), TLR3 (rs6552950), IL-6 (rs1800795), OAS2 (rs15895) and OAS3 (rs2285932) variants exhibited 1.36340%, 1.263845%, 1.214068%, 1.164291%, 1.16429%, 1.114514%, 1.014959%, 0.970085%, 0.962962% and 0.903133% allele frequencies respectively. Among these variants, the CD209 (rs2287886) was highly prevalent in the Northeast Asian population, with an allele frequency of 1.36340%, while this variant was absent in the South Asian population. Furthermore, TLR4 (rs4986790) exhibited the least allele frequency (0.004273%) in the Northeast Asian population compared with other ethnic groups as shown in Fig. 6.
Allele frequency in the east Asian population
In the East Asian population, the IL-6 (rs1800795), CXCL10 (rs3921), OAS2 (rs15895), OAS3 (rs2285932), TLR8 (rs5741883), MICB (rs3132468), and IL-4 (rs2243250) variants showed 1%, 0.9551%, 0.9974%, 0.9203%, 0.976%, 0.905% and 0.82% allele frequency respectively. Among these variants, the IL-6 (rs1800795) variant, was highly prevalent in the East Asian population, with an allele frequency of about 1%. Furthermore, in the East Asian population, the G6PD (rs1050828) variant exhibited about 0% allele frequency as mentioned in Fig. 6.
Allele frequency in the south Asian population
In the Southeast Asian population, the observed genetic diversity showed that IL-10 (rs1800871), IL10 (rs1800872), CXCL10 (rs3921), CCR5 (rs1799987), CCR5 (rs1799988), CCR5 (rs1800023), OAS1 (rs1131454), OAS1 (rs10774671), OAS2 (rs15895), OAS2 (rs1732778), OAS3 (rs2285932), TLR3 (rs6552950), IL-6 (rs1800795), ICAM-1 (rs5498), TLR9 (rs187084), TLR9 (rs187084), IL-4 (rs2243250), ICAM1 (rs5498) and FcγRIIA (rs1801274) genetic variants exhibited 4.688657%, 4.19014%, 4.55585%, 4.31062%, 4.29173%, 3.22456%, 4.77668%, 5.84959%, 6.20148%, 3.07543%, 6.61853%, 5.38533%, 6.15577%, 3.80109%, 2.48923%, 2.48923%, 2.64448%, 3.80109% and 2.83783% allele frequency respectively. Among these variants, the OAS2 (rs15895) was highly prevalent in the Southeast Asian population, showing the highest allele frequency of 6.20148%. Furthermore, in the South Asian population, the G6PD (rs1050828) variant exhibited minimum allele frequency about (0.000367%) mentioned in Fig. 6).
Validation of prioritized genetic variants against MBDs among the Haripur population
Genetic variants prevalent in the South Asian population were examined within the local Haripur population. Two specific variants from IL-10 (rs1800871) and FcγRIIA (rs1801274) mentioned were analyzed on the local population to validate our study findings. In this study, allele and genotype frequencies for IL-10 (rs1800871) and FcγRIIA (rs1801274) were examined in a sample of 50 participants according to the inclusion-exclusion criteria defined in the methodology section. For IL-10 (rs1800871), the genotype distribution showed that the wild-type (TT) genotype and heterozygous genotype (TC) present in the local Haripur population were 88% and 12% respectively in the available sample size. Similarly, for FcγRIIA (rs1801274) variant, the frequencies of wild-type (GG) and heterozygous GA genotypes were 6% and 94%, respectively in the available sample size. Statistical analysis revealed an allele frequency of 0.1128% for the FcγRIIA variant and 0.2112% for IL-10 variant as shown in Table 5 and Fig. 7.
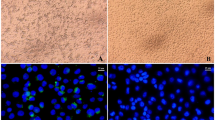
(A) ARMS-PCR using tetra primers to study genetic variations in the FcγRIIA and IL-10 gene. In IL-10 (rs1800871) the outer primers (OF and OR) amplified a 607 bp product. The IF primer generated a wild allele with an amplicon size of 380 bp while the IR primer generated a mutated allele with an amplicon size of 272 bp. (B) The outer primers (OF and OR) for FcγRIIA (rs1801274) amplified a 492 bp product. The IF primer generated a wild allele with an amplicon size of 316 bp while the IR primer generated a mutated allele with an amplicon size of 269 bp. Representative PCR products obtained by ARMS are interpreted from an agarose gel. Lane (L) presents the DNA ladder that is 100 base pairs long.
Discussion
The varied genetic architectures across ethnic groups highlight the importance of region-specific studies, as genetic factors may differentially influence disease pathogenesis. South Asia’s unique genetic diversity offers both challenges and opportunities for population-based genetic studies48,49. Advancements since the Human Genome Project, including SNP analysis, DNA sequencing, and high-throughput genotyping, have enabled large-scale GWAS that have revolutionized our understanding of complex diseases50,51,52. In MBDs research, GWAS have been instrumental in identifying genetic variations in mosquito vectors, human hosts, and pathogens. These studies have been instrumental in helping us to understand more about disease pathogenesis and various associations between parasite and host genetic factors51,52.
In the current study, we systematically identified candidate genes, and their genetic variants associated with both susceptibility and protection against MBDs, focusing on their distribution across diverse ethnic groups. Our analysis revealed notable differences in allele frequencies, highlighting population-specific genetic resilience. African populations and their descendants exhibited greater protection against severe MBDs compared to Southeast and Northeast Asians. Similarly, individuals of European ancestry and neighboring regions showed genetic protection against DF, whereas Southeast and Northeast Asians were more susceptible29.
Moreover, MBD-related genetic variants were aggregated from publicly available datasets and then subjected to broader analysis to prioritize overlapping and unique genes using a structured framework. This approach facilitated the categorization of gene-variants (overlapping and unique) among MBDs. Also, it helped in identification of their associations across MBDs. After prioritization based on overlapping genes and their variants among MBDs, we calculated allele frequencies of these variants across multiple global populations, including Middle Eastern, Ashkenazi Jewish, European (Non-Finnish and Finnish), Latino/Admixed American, South Asian, East Asian, North Asian, Southeast Asian, African American, and Swedish populations. By examining allele frequencies in this diverse set of populations, we captured a comprehensive global perspective on genetic susceptibility and protection against MBDs.
Our results revealed that TLR7 (rs179010) variant have highest allele frequency of about 0.8511% in Latino/Admixed American population and which reflects that this population linked with disease susceptibility against CHIKV, DENV, JEV, RVF and WNV diseases as shown in Supplementary tables S1, S2, S3, S6 and S7. OAS3 (rs2285932) variant exhibited highest allele frequency of about of 0.9481% in African/African American population, which reflect that this population is susceptible to CHIKV and WNV diseases as mentioned in Supplementary tables S2 and S7.
In the Ashkenazi Jewish population, the TLR3 genetic variant (rs6552950) exhibited the highest allele frequency of about 0.8576% among all studied variants. This observation suggests a potential genetic predisposition in these populations, correlating with increased susceptibility to diseases including CHIKV, DENV, JEV, RVF, WNV, and ZIKV infections as mentioned in Supplementary tables S1, S2, S3, S6, S7 and S9. The OAS3 genetic variant (rs2285932) demonstrates the highest allele frequency of about 0.7999% in the European (Finnish) population. The highest allele frequency of this variant showed that this population might be susceptibility to CHIKV, DENV, and WNV diseases, emphasizing population-specific genetic predispositions that influence responses to viral infections as shown in Supplementary tables S1, S2 and S7.
Furthermore, our results revealed that the IL-10 (rs1800872) variant showed highest allele frequency in the European (non-Finnish) population and its associations with susceptibility to various infectious diseases. With an allele frequency of 0.7999%, it was one of the most prevalent variants in this population. This polymorphism allele frequency revealed that this population increased susceptibility to DENV, JEV, WNV, malaria and ZIKV infections as mentioned in Supplementary tables S2, S3, S7, S5 and S9. The OAS3 (rs2285932) variant demonstrates the highest allele frequency within the European (Finnish) population and OTHERS population. This observation suggests a potential genetic predisposition in these populations, correlating with increased susceptibility to diseases such as CHIKV, DENV, and WNV diseases. The heightened frequency of this variant may indicate a population-specific genetic factor influencing disease outcomes or responses to viral infections as shown in Supplementary tables S1, S2 and S7.
The CD209 (rs4804803) variant was highly prevalent in the American population, with an allele frequency of 0.7109%. This variant has been strongly associated with increased susceptibility to various viral infections, including CHIKV, DENV, JEV, WNV, and ZIKV diseases as mentioned in Supplementary tables S1, S2, S3, S7 and S9. Its high prevalence might play a significant role in shaping immune responses and disease outcomes within this population. In the Oceanian, African and West Eurasian population, the OAS2 variant (rs15895) and OAS3 (rs2285932) variant exhibited the highest allele frequencies of about 0.7109%. Among these, the OAS3 (rs2285932) variant showed a stronger association with susceptibility to diseases such as CHIKV, DENV and WNV diseases as shown in Supplementary tables S1, S2 and S7. On the other hand, the OAS2 variant (rs15895) was linked to disease susceptibility against DENV, CHIKV, and WNV diseases as well, emphasizing the potential role of these genetic markers in influencing the immune response and vulnerability to these infections as mentioned in Supplementary tables S2, S1 and S7.
Sweden’s population revealed that TLR3 (rs6552950) variant have highest allele frequency among all variants which showed that this population is susceptible to DENV, CHIKV, JEV, RVF, WNV and ZIKV infections as mentioned in Supplementary tables S2, S1, S3, S6, S7 and S9. The CTLA4 (rs231775) variant was highly prevalent in the Southeast Asian population, with an allele frequency of 0.992020%. This variant allele frequency revealed that this population was more susceptible to various MBDs, including JEV and WNV diseases as shown in Supplementary tables S3 and S7. In Northeast Asian population the CD209 (rs2287886) was highly prevalent, with an allele frequency of 1.36340% showed that this population was more susceptible against CHIKV, DENV, JEV, ZIKV and WNV diseases as shown in Supplementary tables S1, S2, S3, S9 and S7 as well their potential role of this genetic variant in influencing the immune response and vulnerability to these infections. The IL-6 (rs1800795) variant revealed an allele frequency of about 1% in the East Asian population. This variant provides protection against infections caused by several viruses, including DENV, ZIKV, RVFV, and CHIKV infections as shown in Supplementary tables S2, S9, S6 and S1.
In South Asian population, the OAS2 (rs15895) variant exhibited the highest allele frequencies of about 0.7109%. This variant allele frequency revealed that this population might be more susceptible to CHIKV, DENV, and WNV diseases as mentioned in Supplementary tables S1, S2 and S7 as well, emphasizing the potential role of this genetic variant in influencing the immune response and vulnerability to these infections. Furthermore, we explored the correlation of observed allele frequencies with available data on morbidity and mortality rates of MBDs in different populations. The presence of MIC-B and PLCE1 allelic variants in in African ancestry are linked with the severity of DENV infection in individuals53,54. Another study showed the involvement of TNF, IFNG and IL-10 genetic variation with increased susceptibility to DHF in the Brazilian population55. In Malaysian population genetic variants of HLA may increase the risk of DHF in DENV infected patients33. Mark Loeb et al. identified genetic variants of 3 genes (RFC1, SCN1A, ANPEP) that potentially implicated in susceptibility to WNV disease neurological complications in United States and Canadian populations56. Our findings suggest that the observed patterns align with the founder effect, where population-specific genetic adaptations may influence disease outcomes57.
While our study encompassed a broad range of populations, we placed particular emphasis on the South Asian population, given the region’s significant burden of emerging infectious diseases and its critical role in global public health. The South Asian region has been a focal point for diseases with pandemic potential, posing substantial public health and economic challenges. In South Asian population we calculated the allele frequency 1009 genetic variants of 366 genes among these variants 19 were revealed the highest allele frequency in this population. The observed genetic diversity in this population revealed that IL-10 (rs1800871), IL10 (rs1800872), CXCL10 (rs3921), CCR5 (rs1799987), CCR5 (rs1799988), CCR5 (rs1800023), OAS1 (rs1131454), OAS1 (rs10774671), OAS2 (rs15895), OAS2 (rs1732778), OAS3 (rs2285932), TLR3 (rs6552950), IL-6 (rs1800795), ICAM-1 (rs5498), TLR9 (rs187084), TLR9 (rs187084), IL-4 (rs2243250), ICAM1 (rs5498) and FcγRIIA (rs1801274) genetic variants exhibited 4.688657%, 4.19014%, 4.55585%, 4.31062%, 4.29173%, 3.22456%, 4.77668%, 5.84959%, 6.20148%, 3.07543%, 6.61853%, 5.38533%, 6.15577%, 3.80109%, 2.48923%, 2.48923%, 2.64448%, 3.80109% and 2.83783% allele frequency respectively. These findings suggests that in South Asian region the observed allele frequencies of these variants showed that this population is most susceptible against most of the MBDs including DENV, ZIKV, RFV, WNV, CHIKV, LF, malaria and JEV infections.
In order to validate our study findings, we selected two genes and their variants (related to susceptibility and protection) that were more prevalent in studies related to South Asian populations, underscoring the importance of region-specific genetic insights in understanding and mitigating MBDs risk. FcγRIIA and IL10 genes play an important role in immune responses and pathogenesis of malaria and DENV infection that were more prevalent in Pakistani population. They have been shown to influence individual susceptibility and the overall immune landscape of infections such as DENV, malaria, and ZIKV58,59,60,61,62. Immune-modulating genes may function synergistically or antagonistically in populations with unique genetic backgrounds, altering the dynamics of infection spread and severity62.
ARMS PCR analysis of the genomic samples in local Haripur population were performed for IL-10 (rs1800871) and FcγRIIA (rs1801274) alleles. Based on our findings, the TT genotype of IL-10 (rs1800871) represented a homozygous normal allele, indicating a typical genomic pattern that confers protection against MBDs. In contrast, the TC genotype was associated with increased susceptibility to MBDs. While for FcγRIIA (rs1801274), the GG genotype is related to protection, with the GA genotype was related to susceptibility against MBDs. In our sample of local population, it was shown that the TT genotype of IL-10 (rs1800871) was predominant (88%) in local population, whereas the TC genotype was less common (12%). Similarly, for FcγRIIA (rs1801274), the GG genotype was most prevalent (94%), with the GA genotype being a minority (6%). This pattern suggests that the local population is somewhat protected against MBDs due to minimum allele frequency against MBDs. These patterns are consistent with larger results in European and African region, implying that specific genotypes give either vulnerability or protection, which may differ between groups due to evolutionary and environmental variables.
IL-10 is an anti-inflammatory cytokine that plays an essential role in dampening immune responses during infection to avoid excessive tissue damage63. Although its functions in the regulation of inflammatory pathways are protective, but the immunosuppressive properties of IL-10 impaired the host’s ability to mount effective immune defenses, allowing pathogens perhaps to persist or replicate unchecked64. Genetic polymorphism in IL-10 (rs1800871), have been substantially linked to greater susceptibility to DENV infection. Moreover, in the Mexican population, the IL-10 (rs18001871) variant indicates the there is an independent relationship between TC and CC alleles with DENV disease susceptibility. This study highlights the relationship between DENV disease and the IL-10 (rs18001871) variant65. This variant affect the expression levels of IL-10, leading to an imbalanced immune response that promote virus persistence and exacerbate the severity of the disease65. Thus, IL-10 is a potential biomarker for predicting disease prognosis not only in DENV infection but also in other viral diseases58. Furthermore, the rs1800871 polymorphism in IL-10 gene is associated with clinical malaria in children and IL-10 production66, and IL-10 genetic variation with increased susceptibility to DHF in the Brazilian population55. A study in Mozambique found higher IL-10 levels in cord blood cells exposed to infected erythrocytes, linked to the presence of the IL-10 rs1800871 G allele. This association remained significant after adjusting for various confounders, including maternal factors and malaria interventions66. The rs1800871 polymorphism has been found to have contradictory associations, with some cases indicating increased risk and others suggesting protection against mosquito-borne infections. Specifically, the GG genotype of rs1800871 was linked with protection compared to CC, indicating a reduced risk of severe complications from MBDs67. Another study reported that IL10 variants associated with increased IL-10 production elevate the risk of clinical malaria in children. Elevated levels of circulating IL-10 have been observed in patients with cerebral, severe, and moderate malaria. Similar findings from pre-clinical models indicate that African children with severe anemia exhibit lower plasma IL-10 levels compared to those with moderate anemia or cerebral malaria, underscoring the role of IL-10 in preventing severe anemia66,68,69.
FcγRIIA plays a significant role in the recognition and response of the immune system to pathogen-antibody complexes, especially regarding antibody-dependent enhancement. This receptor mediates immune effector functions, including phagocytosis, antibody-dependent cytotoxicity, and the stimulation of inflammatory pathways70. The FcγRIIA gene variation of rs1801274 has been linked to immune regulation during infections, increasing vulnerability to illnesses against DENV diseases71. The H131R polymorphism, have been linked in altering the affinity of the receptor to immune complexes and thus susceptible to a range of vector-borne diseases, such as malaria and DENV infections72,73.
In the context of malaria, FcγRIIA variants have been associated with variable disease severity, depending on the allele; the resultant ADE-infectivity enhancement enhances virus entry into host cells, promoting higher viral replication and inflammation74. The FcγRIIA gene polymorphism (rs1801274), having R131H, was found to be protective against blood-stage malaria infections75. Similarly, the coding mutation is FcγRIIA-H131R (rs1801274), which impacts the activation of monocytes through FcγRIIA, leading to alterations in IgG recognition and the phagocytosis of IgG parasites73,75. Another study from Pakistan revealed that mutation in the FcγRIIA gene (rs1801274) affects receptors where immunoglobin G shows binding71. The mutated A allele of the FcγRIIA gene rs1801274 has been associated with protection against malaria, indicating the significant role of FcγRIIA in susceptibility to the disease61.
FcγRIIA has also been associated, with DENV infection, with an increased risk for the development of severe outcomes of DENV infection including DHF and DSS, when pre-existing antibodies against this virus are present within the host71,76,77. In the Vietnamese population genetic variant of FcγRIIA R131H (rs1801274) in patients with DSS showed a significant association with recessive protection of the R allele78. Another study in Mexican population revealed that genetic polymorphism in the FcγRIIA variant rs1801274 was associated with individuals carrying the histidine allele, whether homozygous (GG) or heterozygous (GA), showing protection from symptomatic DENV infection compared to those with the asymptomatic genotype. Additionally, homozygous variants of the FcγRIIA gene were less likely to result in severe DENV infection79. Individuals with the R allele and R/R genotype are protected from symptomatic DENV infection, while those with the H allele and H/H genotype are associated with both DF and DHF. This highlights rs1801274 as a crucial genetic determinant for severe DENV infection80. Another study revealed that genetic susceptibility to DF is associated with FcγRIIA gene polymorphism in Pakistani population81. Similarly, the FcγRIIA rs1801274 gene variant is a crucial factor in susceptibility and protection against CHIKV infection. The FcγRIIA gene variation plays a key role in regulating immune responses during CHIKV infection, determining whether an individual is susceptible or resistant to the virus82. It has been found that the significant genetic variation rs1801274 within the FcγRIIA gene regulates the immune response to CHIKV infection83. Furthermore, CHIKV disease susceptibility is connected to the CD209 gene’s rs4804803 GG genotype, OAS and CD209 genes may affect CHIKV infection symptomatology84.
Our current investigations further strengthen the significance of these IL-10 polymorphisms in modifying host responses and emphasize their predictive potential for disease prognosis in diverse geographic regions and populations. A better understanding of the genetic basis of immune responses in MBDs is crucial for developing targeted, personalized health therapies. The implementation of more effective public health measures, such as novel vaccinations and antiviral treatments can be designed to alter these immune pathways to enhance disease outcomes.
Implications and contributions
This study lays the groundwork for understanding the genetic architecture of MBDs in South Asian populations, particularly in areas with high illness burden. Identifying protective and susceptibility variations can help drive public health efforts such as targeted screening and population-specific treatments. Furthermore, this study emphasizes the need of incorporating genetic data into disease prevention strategies to increase their efficacy and sustainability. Population genomics could therefore be used to help develop effective new interventions, as well as to support the efficient deployment of existing disease control measures through resistance surveillance. Population genomics will play an important role in future vaccine development efforts by mapping differences in vaccine target polymorphism profiles across populations and identifying immunogenic protein targets that are not highly polymorphic in any geographical region. Furthermore, incorporating genetic screening into public health policies may allow for more effective risk prediction and mitigation of MBDs, while research into the socioeconomic and cultural obstacles to genetic testing adoption might guarantee that such treatments are more widely applicable and accepted.
Limitations and future research directions
This study is a preliminary effort focused on screening genetic variants associated with susceptibility and protection against MBDs. Due to the small sample size, the analysis was limited to a restricted number of individuals from the local population of Tehsil Haripur, which may not fully represent the genetic diversity of the entire region. Additionally, many other factors which contribute to genetic risk in population like environmental, behavioral, and other epidemiological factors influencing susceptibility to MBDs were not included in this study. Genetic research and screening programs require significant funding, technical expertise, and infrastructure, which are often lacking in regions where MBDs are endemic. This limits the feasibility of large-scale genetic risk assessments in resource-constrained areas. Despite these limitations, this study provides valuable insights and lays the groundwork for future research. It is the first study conducted in this region to explore genetic variants linked to MBDs, addressing an important gap in understanding the genetic basis of disease susceptibility and protection. These findings can serve as a baseline for larger-scale studies and contribute to the development of targeted strategies for disease prevention and control. We believe that this study more accurately predicted the genetic screenings in local population. In addition, this study was a meaningful study that could confirm that public data analysis is helpful toward predicting the variant spectrum of actual patients. To our knowledge, this is the first report to demonstrate such polygenic risk for MBDs. Our data provides valuable insights into how genetic variants influence susceptibility to MBDs. Additionally, it highlights the importance of leveraging population-specific data to inform and organize more effective public health awareness campaigns aimed at controlling these diseases.
Conclusion
Genetic research has played an instrumental role in the discovery of new biological pathways underpinning infectious diseases and the evaluation of new targets for therapeutic development. Public awareness campaigns can play a significant role in mitigating the impact of MBDs in vulnerable populations by increasing knowledge about prevention, early detection, and treatment options. It can be useful in promoting and even accelerating the implementation to enroll subjects for genome projects, whether they are for research or clinical applications, in different countries. The lack of knowledge on the techniques and usage of results from genetic testing leads to misconceptions, unwarranted fear, and rejection of technologies on prejudicial beliefs. Every aspect of education plays a key role since attitudes toward genetic testing are hugely influenced by religious, cultural, and moral views. As diversity in population genetics plays a significant role in viral pathogenesis, drug resistance, viral evolution, and immune escape, further studies must be conducted on worldwide population genetics to expedite the development of population-specific therapeutics to mitigate this worldwide challenge.
Data availability
All data supporting the findings of this study were included within the manuscript, and no external datasets were used. The data was provided in the main manuscript and the supplementary information files.
References
Sanfelice, V. Mosquito-borne disease and newborn health. Health Econ. 31 (1), 73–93 (2022).
Naik, B. R., Tyagi, B. & Xue, R-D. Mosquito-borne diseases in India over the past 50 years and their global public health implications: a systematic review. J. Am. Mosq. Control Assoc. 39 (4), 258–277 (2023).
Omodior, O., Luetke, M. C. & Nelson, E. J. Mosquito-borne infectious disease, risk-perceptions, and personal protective behavior among US international travelers. Prev. Med. Rep. 12, 336–342 (2018).
Chilakam, N. et al. Economic Burden of Mosquito-Borne diseases in low-and Middle-Income Countries: protocol for a systematic review. JMIR Res. Protocols. 12 (1), e50985 (2023).
Wang, G-H. et al. Combating mosquito-borne diseases using genetic control technologies. Nat. Commun. 12 (1), 4388 (2021).
Urmi, T. J. et al. Frequent outbreaks of dengue fever in south Asian countries—A correspondence analyzing causative factors and ways to avert. Health Sci. Rep. 6(10). (2023).
Dhimal, M. et al. Threats of Zika virus transmission for Asia and its Hindu-Kush Himalayan region. Infect. Dis. Poverty. 7, 1–7 (2018).
Ferguson, N. M. Challenges and opportunities in controlling mosquito-borne infections. Nature 559 (7715), 490–497 (2018).
Hossain, M. S., Noman, A. A., Mamun, S. & Mosabbir, A. A. Twenty-two years of dengue outbreaks in Bangladesh: epidemiology, clinical spectrum, serotypes, and future disease risks. Trop. Med. Health. 51 (1), 1–14 (2023).
Lessa, C. L. S., Hodel, K. V. S., Gonçalves, M. S. & Machado, B. A. S. Dengue as a disease threatening Global Health: a narrative review focusing on Latin America and Brazil. Trop. Med. Infect. Disease. 8 (5), 241 (2023).
Jagtap, S., Pattabiraman, C., Sankaradoss, A., Krishna, S. & Roy, R. Evolutionary dynamics of dengue virus in India. PLoS Pathog. 19 (4), e1010862 (2023).
Badar, N. et al. Epidemiological trend of Chikungunya outbreak in Pakistan: 2016–2018. PLoS Negl. Trop. Dis. 13 (4), e0007118 (2019).
Pialoux, G., Gaüzère, B-A., Jauréguiberry, S. & Strobel, M. Chikungunya, an epidemic arbovirosis. Lancet. Infect. Dis. 7 (5), 319–327 (2007).
imtiaz, K. et al. Zika Virus looming epidemic in Pakistan: seroprevalence findings by plaque reduction neutralization test in the Sindh Region of Pakistan. medRxiv 2023:2023.04. 27.23289241.
Iqtadar, S. et al. Screening of ZIKA virus infection among dengue-like illness patients with negative RT-PCR for dengue virus in Punjab–Pakistan. Pakistan J. Med. Sci. 37 (3), 721 (2021).
Imran, M. et al. Epidemiological trends of mosquito-borne viral diseases in Pakistan. Anim. Dis. 2 (1), 1–10 (2022).
Colón-González, F. J. et al. Projecting the risk of mosquito-borne diseases in a warmer and more populated world: a multi-model, multi-scenario intercomparison modelling study. Lancet Planet. Health. 5 (7), e404–e14 (2021).
Field, C. B. & Barros, V. R. Climate Change 2014–Impacts, Adaptation and Vulnerability: Regional Aspects (Cambridge University Press, 2014).
Gubler, D. J. Dengue, urbanization and globalization: the unholy trinity of the 21st century. Trop. Med. Health. 39 (4SUPPLEMENT), S3–S11 (2011).
Driss, A. et al. Genetic polymorphisms linked to susceptibility to malaria. Malar. J. 10, 1–10 (2011).
Burgner, D., Jamieson, S. E. & Blackwell, J. M. Genetic susceptibility to infectious diseases: big is beautiful, but will bigger be even better? Lancet. Infect. Dis. 6 (10), 653–663 (2006).
Demkova, K., Morris, D. L. & Vyse, T. J. Genetics of SLE: does this explain susceptibility and severity across racial groups? Rheumatology 62 (Supplement_1), i15–i21 (2023).
Han, J-W. et al. Genome-wide association study in a Chinese Han population identifies nine new susceptibility loci for systemic lupus erythematosus. Nat. Genet. 41 (11), 1234–1237 (2009).
Tsao, B. The genetics of human lupus in Wallace DJ (ed Hahn, B. H.) DJ and Dubois’ Lupus Erythematosus. Lippincott Williams & Wilkins A Wolters Kluwer Company, USA; (2002).
Shipman, L. New GWAS loci and insights into ancestry. Nat. Rev. Rheumatol. 12 (9), 499 (2016).
75 MCHAJIBYLSMLSAVPSK. Locus for severity implicates CNS resilience in progression of multiple sclerosis. Nature 619 (7969), 323–331 (2023).
Khor, C. C. et al. Genome-wide association study identifies susceptibility loci for dengue shock syndrome at MICB and PLCE1. Nat. Genet. 43 (11), 1139–1141 (2011).
Fang, X. et al. Genetic polymorphisms of molecules involved in host immune response to dengue virus infection. FEMS Immunol. Med. Microbiol. 66 (2), 134–146 (2012).
Oliveira, M. et al. Population genetics-informed meta-analysis in seven genes associated with risk to dengue fever disease. Infect. Genet. Evol. 62, 60–72 (2018).
Patro, A. R. K. et al. A 23 bp Indel Polymorphism in TLR2 Gene Enhances Inflammation and Disease Severity in Dengue. bioRxiv 239988. (2017).
Wei, Z. et al. Genome-wide association studies of HIV-1 host control in ethnically diverse Chinese populations. Sci. Rep. 5 (1), 10879 (2015).
Pare, G. et al. Genetic risk for dengue hemorrhagic fever and dengue fever in multiple ancestries. EBioMedicine 51. (2020).
Alshammari, M. K. et al. Genetic variants associated with dengue hemorrhagic fever. A systematic review and meta-analysis. J. Infect. Public Health (2024).
Raban, R., Gendron, W. A. & Akbari, O. S. A perspective on the expansion of the genetic technologies to support the control of neglected vector-borne diseases and conservation. Front. Trop. Dis. 3, 999273 (2022).
Alfares, A. et al. What is the right sequencing approach? Solo VS extended family analysis in consanguineous populations. BMC Med. Genom. 13, 1–8 (2020).
Tan, M. et al. Genome-wide determinants of mortality and motor progression in Parkinson’s disease. (2022).
Lee, J. C. et al. Genome-wide association study identifies distinct genetic contributions to prognosis and susceptibility in Crohn’s disease. Nat. Genet. 49 (2), 262–268 (2017).
Abi, A. & Safavi, A. Targeted detection of single-nucleotide variations: progress and promise. ACS Sens. 4 (4), 792–807 (2019).
Yang, L. et al. Evaluation of amplification refractory mutation system (ARMS) technique for quick and accurate prenatal gene diagnosis of CHM variant in choroideremia. Application Clin. Genet. 1–8. (2017).
Peng, B. et al. A novel and quick PCR-based method to genotype mice with a leptin receptor mutation (db/db mice). Acta Pharmacol. Sin. 39 (1), 117–123 (2018).
Gao, Y. et al. PGG. Han: the Han Chinese genome database and analysis platform. Nucleic Acids Res. 48 (D1), D971–D6 (2020).
The GenomeAsia 100. K Project enables genetic discoveries across Asia. Nature 576 (7785), 106–111 (2019).
Ameur, A. et al. SweGen: a whole-genome data resource of genetic variability in a cross-section of the Swedish population. Eur. J. Hum. Genet. 25 (11), 1253–1260 (2017).
Zhang, P. et al. NyuWa Genome Resource: Deep Whole Genome Sequencing Based Chinese Population Variation Profile and Reference Panel. bioRxiv. 2020:2020.11. 10.376574
Davis, M. W. & Jorgensen, E. M. ApE, a plasmid editor: a freely available DNA manipulation and visualization program. Front. Bioinf. 2, 818619 (2022).
Mayo, O. A century of hardy–Weinberg equilibrium. Twin Res. Hum. Genet. 11 (3), 249–256 (2008).
Chen, S. et al. A genomic mutational constraint map using variation in 76,156 human genomes. Nature 625 (7993), 92–100 (2024).
Wall, J. D. et al. South Asian medical cohorts reveal strong founder effects and high rates of homozygosity. Nat. Commun. 14 (1), 3377 (2023).
Miyashita, A., Kikuchi, M., Hara, N. & Ikeuchi, T. Genetics of Alzheimer’s disease: an east Asian perspective. J. Hum. Genet. 68 (3), 115–124 (2023).
Lan, N. T. P. & Hirayama, K. Host genetic susceptibility to severe dengue infection. Trop. Med. Health. 39 (4SUPPLEMENT), S73–S81 (2011).
Gibbs, R. A. et al. The international HapMap project. (2003).
Eleftherohorinou, H. et al. Pathway analysis of GWAS provides new insights into genetic susceptibility to 3 inflammatory diseases. PloS One. 4 (11), e8068 (2009).
Sierra, B. et al. OSBPL10, RXRA and lipid metabolism confer african-ancestry protection against dengue haemorrhagic fever in admixed cubans. PLoS Pathog. 13 (2), e1006220 (2017).
Halstead, S. B. et al. Haiti: absence of dengue hemorrhagic fever despite hyperendemic dengue virus transmission. Am. J. Trop. Med. Hyg. 65 (3), 180–183 (2001).
Santos, D. C. et al. Genomic ancestry as a risk factor for diabetic retinopathy in patients with type 1 diabetes from an admixed population: a nested case–control study in Brazil. Acta Diabetol. 57, 937–945 (2020).
Loeb, M. et al. Genetic variants and susceptibility to neurological complications following West Nile virus infection. J. Infect. Dis. 204 (7), 1031–1037 (2011).
Jiang, X. & Assis, R. Population-specific genetic and expression differentiation in europeans. Genome Biol. Evol. 12 (4), 358–369 (2020).
Tsai, T-T. et al. An emerging role for the anti-inflammatory cytokine interleukin-10 in dengue virus infection. J. Biomed. Sci. 20, 1–9 (2013).
Rodrigo, W. S. I., Jin, X., Blackley, S. D., Rose, R. C. & Schlesinger, J. J. Differential enhancement of dengue virus immune complex infectivity mediated by signaling-competent and signaling-incompetent human FcγRIA (CD64) or FcγRIIA (CD32). J. Virol. 80 (20), 10128–10138 (2006).
Ouma, C. et al. Functional haplotypes of fc gamma (Fcγ) receptor (FcγRIIA and FcγRIIIB) predict risk to repeated episodes of severe malarial anemia and mortality in Kenyan children. Hum. Genet. 131, 289–299 (2012).
Natama, H. M. et al. Genetic variation in the immune system and malaria susceptibility in infants: a nested case–control study in Nanoro, Burkina Faso. Malar. J. 20, 1–14 (2021).
Khandia, R. et al. Modulation of dengue/zika virus pathogenicity by antibody-dependent enhancement and strategies to protect against enhancement in Zika virus infection. Front. Immunol. 9, 597 (2018).
Rojas, J. M., Avia, M., Martín, V. & Sevilla, N. IL-10: a multifunctional cytokine in viral infections. J. Immunol. Res. 2017 (1), 6104054 (2017).
Howes, A., Stimpson, P., Redford, P., Gabrysova, L. & O’Garra, A. Interleukin-10: cytokines in anti-inflammation and tolerance. Cytokine Frontiers: Regulation of Immune Responses in Health and Disease. 327–352 (2014).
Monroy-Muñoz, I. E. et al. Genetic polymorphisms rs1800871 and rs1800872 of IL-10 gene are associated with dengue infection, especially with serotype 1 and DwoWS in Mexican population. Cytokine 166, 156194 (2023).
Zhang, G. et al. Interleukin-10 (IL-10) polymorphisms are associated with IL-10 production and clinical malaria in young children. Infect. Immun. 80 (7), 2316–2322 (2012).
Grijalva, A. et al. Interleukin 10 polymorphisms as risk factors for progression to Chagas disease cardiomyopathy: a case-control study and meta-analysis. Front. Immunol. 13, 946350 (2022).
Peyron, F. et al. High levels of circulating IL-10 in human malaria. Clin. Experimental Immunol. 95 (2), 300–303 (1994).
Lyke, K. et al. Serum levels of the proinflammatory cytokines interleukin-1 beta (IL-1β), IL-6, IL-8, IL-10, tumor necrosis factor alpha, and IL-12 (p70) in Malian children with severe Plasmodium Falciparum malaria and matched uncomplicated malaria or healthy controls. Infect. Immun. 72 (10), 5630–5637 (2004).
He, Y. et al. In vitro enhancement of Zika virus infection by preexisting West Nile virus antibodies in human plasma-derived immunoglobulins revealed after P2 binding site-specific enrichment. Microbiol. Spectr., e00758–e00724 (2024).
Mohsin, S. N. et al. Association of FcγRIIa polymorphism with clinical outcome of dengue infection: first insight from Pakistan. Am. J. Trop. Med. Hyg. 93 (4), 691 (2015).
Maiga, B. et al. Fc gamma receptor II a-H 131 R polymorphism and Malaria susceptibility in Sympatric ethnic groups, Fulani and Dogon of M ali. Scand. J. Immunol. 79 (1), 43–50 (2014).
Amiah, M. A., Ouattara, A., Okou, D. T., N’Guetta, S-P-A. & Yavo, W. Polymorphisms in fc gamma receptors and susceptibility to malaria in an endemic population. Front. Immunol. 11, 561142 (2020).
Nasr, A. et al. Significant differences in FcγRIIa, FcγRIIIa and FcγRIIIb genes polymorphism and anti-malarial IgG subclass pattern are associated with severe Plasmodium falciparum malaria in Saudi children. Malar. J. 20, 1–10 (2021).
Zhao, J. et al. Association between Fc-gamma receptor IIa (CD32) gene polymorphism and malaria susceptibility: a meta-analysis based on 6928 subjects. Infect. Genet. Evol. 23, 169–175 (2014).
Yamin, R. et al. Human FcγRIIIa activation on splenic macrophages drives dengue pathogenesis in mice. Nat. Microbiol. 8 (8), 1468–1479 (2023).
Wegman, A. D. et al. DENV-specific IgA contributes protective and non-pathologic function during antibody-dependent enhancement of DENV infection. PLoS Pathog. 19 (8), e1011616 (2023).
Coffey, L. L. et al. Human genetic determinants of dengue virus susceptibility. Microbes Infect. 11 (2), 143–156 (2009).
Noecker, C. A., Amaya-Larios, I. Y., Galeana-Hernández, M., Ramos-Castañeda, J. & Martínez-Vega, R. A. Contrasting associations of polymorphisms in FcγRIIa and DC-SIGN with the clinical presentation of dengue infection in a Mexican population. Acta Trop. 138, 15–22 (2014).
Bournazos, S., Gupta, A. & Ravetch, J. V. The role of IgG fc receptors in antibody-dependent enhancement. Nat. Rev. Immunol. 20 (10), 633–643 (2020).
Shih, C-M. et al. Association of TNF-α polymorphism with susceptibility to and severity of non-small cell lung cancer. Lung cancer. 52 (1), 15–20 (2006).
Gotay, W., Rodrigues, R. & Yaochite, J. Influence of host genetic polymorphisms involved in immune response and their role in the development of Chikungunya disease: a review. Braz. J. Med. Biol. Res. 56, e12557 (2023).
Dutta, S. K. & Tripathi, A. Association of toll-like receptor polymorphisms with susceptibility to Chikungunya virus infection. Virology 511, 207–213 (2017).
Tokuda, S. et al. The genetic basis for susceptibility to Rift Valley fever disease in MBT/Pas mice. Genes Immun. 16 (3), 206–212 (2015).
Acknowledgements
We are grateful to the Department of Medical Lab Technology, The University of Haripur, Haripur, Pakistan for their administrative support.
Author information
Authors and Affiliations
Contributions
Conceptualization, FMA and SNK. RB acquisition, analysis, and interpretation of the data. RB and FMA writing—original draft preparation. RB, MBA, FMA, SNK, AO, MM, WA, RP, AHS, MBA, TSA, SK, AN, writing, review and editing. FMA and SNK supervision, project administration and funding acquisition. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Basri, R., Aqeel, M.B., Awan, F.M. et al. An integrated approach to predict genetic risk for Mosquito-Borne diseases in the local Population of Tehsil Haripur, Khyber Pakhtunkhwa, Pakistan. Sci Rep 15, 3478 (2025). https://doi.org/10.1038/s41598-025-88095-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-88095-0