Abstract
Swine dysentery (SD) presents considerable challenges to both animal welfare and pork industry sustainability. Control and prevention of SD rely on antibiotics and non-vaccine biosecurity practices. Host defense peptides (HDPs) have emerged as promising alternatives to treat and prevent such health concern. This study investigated the effects of porcine β-defensin 5 (pBD-5) and its potential host cytotoxicity, metabolic influence, gene expression modulation and direct antimicrobial activity on Brachyspira hyodysenteriae growth in vitro. pBD-5 does not directly inhibit B. hyodysenteriae growth or significantly alters the metabolic activity or membrane integrity of host cells, indicating no significant cytotoxicity at the tested concentrations. Host transcriptome sequencing revealed a reduction in the number of differentially expressed genes in cells exposed to B. hyodysenteriae following pBD-5 treatment, when compared to the pathogen alone, suggesting an immunomodulatory effect. Pathway analysis revealed the downregulation of immune-related pathways, including IL-17, toll-like receptor (TLR), and NOD-like receptor signalling pathways, upon pBD-5 exposure. Conversely, metabolic pathways such as ribosome, protein digestion and absorption, and renin-angiotensin system were upregulated by pBD-5 treatment, hinting at a role in producing and conserving energy during the challenge. While this study offers insights into the immunomodulatory effects of pBD-5, further research is necessary to elucidate its precise mechanisms and potential applications as an alternative treatment for infectious diseases.
Similar content being viewed by others
Introduction
There is a global effort to reduce antimicrobial use in human and veterinary medicine1, which acknowledges a current health challenge: the emergence of antimicrobial-resistant bacterial strains in human and veterinary hospitals as we enter the post-antibiotic era1. Novel technologies are being developed to minimize or replace antibiotics2, including those based on host defense peptides (HDPs). Swine possess a variety of HDPs, including cathelicidin3, natural killer (NK)-lysin4, hepcidin5, liver expressed antimicrobial peptide-2 (LEAP-2)5, peptidoglycan recognition proteins (PGRPs)6 and β-defensins7. These naturally expressed innate immune molecules are important modulators of host-microbiome-pathogen interactions8, as they possess both or either antimicrobial and immunomodulatory properties8. Furthermore, the microbiome composition influences HDP production and activity9,10, establishing a bidirectional relationship critical for maintaining homeostasis. For instance, in mice, gut microbiota-derived molecules, such as aryl hydrocarbon receptor (AHR) ligands and butyrate, were shown to promote interleukin-22 (IL-22) secretion by innate lymphoid cells, which in turn stimulate the expression of murine β-defensin 14 (mBD14) in pancreatic endocrine cells11. In swine, butyrate and its analogs act as potent inducers of multiple—though not all—HDP genes in monocytes, macrophages, and epithelial cells12,13. These findings illustrate the conserved role of microbiota-derived metabolites in modulating HDP expression across species14. β-defensins, the largest and most studied family of HDPs in pigs, exhibit direct antimicrobial mechanisms that involve microbial aggregation, membrane pore formation, and interference with cell wall synthesis and protein folding15. Immunomodulation involves activation of membrane receptors and the up or downregulation of the canonical NF-kB pathway and the NOD-like receptor signalling pathway16,17.
Swine dysentery is characterized by mucohemorrhagic diarrhea and colitis in pigs. This disease has several implications for animal welfare and dramatic economic losses to pig farmers18. It is caused by Brachyspira hyodysenteriae (B. hyodysenteriae), B. hampsonii or B. suanatina, oxygen-tolerant bacteria that colonize the large intestine of pigs18. Prevention, control, and treatment of swine dysentery rely exclusively on antibiotics and biosecurity measures, as vaccines are unavailable18. As one of the most consumed animal proteins globally, food security is directly linked to pork production19.
In previous work investigating the early pathogenesis of swine dysentery in porcine colonic explants18, we combined transcriptomic and histopathological analyses to identify putative resilience factors. We observed that pigs with a resilient phenotype, characterized by fewer Brachyspira-related microscopic lesions, exhibited increased transcript levels from an unannotated genomic region on chromosome 15 between nucleotides 38,096,027 and 38,099,470 (Sus scrofa 11.1), Ensemble ID ENSSSCG00000032503. This region encodes a peptide with high homology (> 80%) to β-defensins and is located near other porcine β-defensin genes. Based on its sequence and potential biological significance, we hypothesized that this peptide might contribute to disease resilience. To investigate, we synthesized the peptide and named it porcine β-defensin 5 (pBD-5).
The purpose of this study was to describe the discovery of pBD-5 and investigate whether it can mitigate the infection by B. hyodysenteriae in vitro. Specific objectives included the evaluation of the potential direct antimicrobial activity, transcriptomic response of an immortalized porcine epithelial cell model, and mitochondrial and lactase dehydrogenase release response of the cells to the tested dose.
Materials and methods
Immortalized intestinal cell culture
Intestinal porcine epithelial cells 1 (IPEC-1), ACC705, were purchased from Leibniz Institute DSMZ—German Collection of Microorganisms and Cell Cultures, Braunschweig, Germany. Cells were cultured in DMEM/HAMs F12 supplemented with 5% foetal calf serum (FCS), 16 mM 4-(2-hydroxyethyl)-1-piperazineethansulfonic acid (HEPES) and 5 ng/mL of epidermal growth factor (all from Invitrogen, Mississauga, ON) was used as culture medium. Cells were incubated at 39 °C in an atmosphere of 5% CO2 and 95% relative humidity. Cells were grown to > 90% confluency prior to exposure to the treatments outlined below. In all experiments, cells were seeded with a density of 105 cells/cm2.
Peptide synthesis and preparation of the different concentrations for incubation
The mature peptide of the novel porcine β-defensin (pBD-5, Table 1) was chemically synthesized through solid-phase peptide synthesis (Kinexus, Vancouver, BC). Cysteine residues were protected by acetamidomethyl to prevent the formation of disulfide bridges. Lyophilized peptide was reconstituted on molecular biology grade water (Fisher BioReagents) at a concentration of 1 M and stored at -80 °C until further diluted for use, as outlined below.
Inoculum preparation and inoculation
Brachyspira hyodysenteriae was cultured as described before in Rubin et al.20. Briefly, brain heart infusion broth (BHI) with 5% (v/v) fetal calf serum (Gibco, Carlsbad, CA), 5% (v/v) sheep’s blood (Gibco, Carlsbad, CA), and 1% (w/v) glucose was used. Broth cultures were incubated in glass vials with magnetic stir bars anaerobically at 39 °C for 48 h with constant stirring.
IPEC-1 cell preparations (n = 3 flasks/treatment) were inoculated with: control (sterile BHI), Brachy (log phase pure culture of Brachyspira hyodysenteriae), pBD + Brachy (log phase pure culture of Brachyspira hyodysenteriae and 40 µM of pBD-5), pBD (40 µM of novel pBD-5), and LPS (100 µg of Escherichia coli O:127 B:8 LPS, Sigma, St. Louis, USA).
All treatments were co-incubated with IPEC-1 cells in the same conditions as described above, for a total of 8 h.
Transcriptome sequencing
Total RNA was extracted from pelleted cells (centrifuged at 2000 g for 15 min) using a commercial kit (Qiagen RNeasy, Mini Kit QIAGEN, Mississauga, ON, Canada) following the manufacturer’s instructions. Extracted RNA integrity, concentration and purity were assessed using the Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA) and Nanodrop 2100 spectrophotometer (Thermo Scientific, Wilmington, DE, USA). rRNA were depleted from 400 ng of total RNA using a commercial kit (Ribo-Zero rRNA Removal Kits, Meta-Bacteria, Epicentre, Madison, WI, USA). Residual RNA was cleaned up using the Agencourt RNACleanTM XP Kit (Beckman Coulter, Indianapolis, IN, USA) and eluted in water. cDNA synthesis was achieved with the NEBNext RNA First Strand Synthesis and NEBNext Ultra Directional RNA Second Strand Synthesis Modules (New England Biolabs, Pickering, ON, Canada). Library preparation was performed using NEBNext Ultra II DNA Library Prep Kit for Illumina (New England Biolabs, Pickering, ON, Canada). Libraries were quantified using the Quant-iT™ PicoGreen® dsDNA Assay Kit (Life Technologies, Burlington, ON, Canada) and the Kapa Illumina GA with Revised Primers-SYBR Fast Universal kit (Kapa Biosystems Wilmington, MA, USA). Average size fragment was determined using a LabChip GX (PerkinElmer, Hopkinton, MA, USA) instrument. Individually indexed libraries were sequenced in four lanes on the Illumina HiSeq 2000 system at the McGill University, Genome Quebec Innovation Centre (Quebec, Canada) to obtain high-quality, 100-bp paired-end reads (average phred quality score ≥ 36).
Transcriptome data analysis
Sequencing data, up to the raw-counts stage, were analyzed using the GenPipes pipeline21. FastQ files containing raw reads were trimmed using Trimmomatic and filtered for quality22. Filtered, high-quality reads were aligned to the Sus scrofa genome (Ensembl Sscrofa 11.1) using the STAR aligner 2-passes mode23. Samtools (v1.124) was used to sort the BAM alignment files and to convert them into SAM format. These files were used as input for HTSeq-count, to generate a matrix with the number of reads per gene25. The raw-counts matrix was imported into the iDEP (v0.86) framework for downstream analysis26, which included sample filtering (at least 0.5 read counts per million, CPM, in at least one sample), count normalization (variance stabilizing transformation, VST, as per the DESeq2 package), exploratory data analysis (k-mer clustering). Pathway analysis was performed using gene set enrichment analysis (GSEA) and the KEGG database27.
Metabolic activity assay
IPEC-1 cells metabolic activity was evaluated using the water-soluble tetrazolium salt (WST-1) Cell Proliferation Reagent (Roche, Canada). Approximately 5 × 104 cells were seeded in a 96-well plate and incubated at 37 °C with 5% CO2 until 80% confluence was achieved. Following overnight incubation, the medium in each well was substituted with 90 µL of fresh cell culture medium as described above, and 10 µL of one of the following treatments: molecular biology water (control) (n = 5), LPS (100 µg of Escherichia coli O:127 B:8 LPS, Sigma, St. Louis, USA) (n = 5), pBD-5 resuspended in molecular water (40 µM) (n = 5), or LPS (100 µg) + pBD-5 (40 µM) (n = 5). After an 8-hour incubation period, cell proliferation reagent (WST-1) was added to each well (10 µL/well) and the 96-well plate was incubated for an additional 4 hours at 37 °C with 5% CO2. At the end of this period, absorbance was measured at 440 nm and 680 nm, the higher serving only to remove background signal.
Cytotoxicity assay
The same experimental design as described for the metabolic activity assay was used for this assay. As a positive control group cells exposed to a lysis buffer (Life Technologies Inc., Burlington, Canada) was included as well (n = 5). Treatment groups exposed to Brachyspira were not evaluated for IPEC-1-LDH release, as B. hyodysenteriae releases the enzyme as part of its anaerobic metabolic pathway28. The CyQUANT LDH Cytotoxicity kit was utilized following the manufacturer’s instructions (Life Technologies Inc., Burlington, Canada).
Antimicrobial activity assay
B. hyodysenteriae growth was assessed via quantitative PCR at two time points: immediately after inoculation (T0) and after 48 h (T48). The following treatments were used: negative control (n = 9), consisted of 9 mL of BHI supplemented with 1 mL of FBS, 100 µL of sterile broth, and 1 mL of molecular biology water; positive control (n = 9), comprised 9 mL of BHI supplemented with 1 mL of FBS, 100 µL of B. hyodysenteriae broth, and 1 mL of molecular biology water; defensin treatments, (each n = 9), each treatment included 9 mL of BHI supplemented with 1 mL of FBS, 100 µL of B. hyodysenteriae broth, and 1 mL of pBD-5 at concentrations of 0.2516 µM, 2.516 µM, and 251.6 µM.
Immediately after collecting 1 mL at T0 and T48, samples were centrifuged at 8000 rpm for 5 min, and 800 µL of the supernatant were removed and discarded from each tube. The remaining 200 µl in the tube were immediately frozen at -80° C for DNA extraction. Bacterial DNA was extracted using DNeasy Blood & Tissue kit (Qiagen). After DNA extraction, the samples were submitted for concentration and purity evaluation (260/280) using a spectrophotometer (NanoDrop 2000c, Thermo Fisher Scientific).
Since colonies are not visible in agar, B. hyodysenteriae was quantified through qPCR20. Each reaction consisted of a mastermix SsoAdvanced Universal SYBR Green Supermix (BioRad, Mississauga, ON, Canada), 2 µL of DNA and primer at 200 nM with a final volume of 25 µL per reaction. Reactions were run in duplicates on a 96-well plate, sealed with an optical adhesive cover before reading in a qPCR system. The thermocycling profile consisted of 95 °C for 3 min for denaturation followed by 40 cycles of 95 °C for 15 s, 63 °C for 15 s, 72 °C for 15s, and 76 °C for 15 s. The final stage included an analysis of the melting curve, verifying the presence of a single peak in each well. Every plate for qPCR analysis included a no-template control and plasmid standard curve ranging in concentration from 100 to 107 copies/reaction.
Statistical analyses
WST-1, LDH and antimicrobial activity assays were conducted using Stata Statistical Software version 17 (StataCorp LLC, College Station, TX, 2019). Normality was assessed using the Shapiro-Wilk test, and equality of variances was evaluated using Levene’s test. To determine whether pBD, LPS, or their combination (LPS + pBD) had an impact on optic density measures in comparison to the control group, we performed an analysis of variance (ANOVA).
Differential gene expression analysis was performed using the DESeq2 package, applying a false discovery rate (FDR) threshold of < 0.1, determined by the Benjamini-Hochberg method, and a fold-change cutoff of > 2. For pathway analysis, we utilized pre-ranked gene set enrichment analysis (GSEA) on the subset of differentially expressed genes with a fold-change > 2, focusing on Sus scrofa genes. Gene sets were derived from the GO biological processes and KEGG pathways databases. Pathways with an FDR < 0.1 were considered significant, while genes with an FDR > 0.9 were excluded from the analysis.
We also conducted a pairwise analysis among the Control, Brachy and pBD + Brachy groups, using a false discovery rate (FDR) threshold of < 0.1, also determined by the Benjamini-Hochberg method, and a fold-change cutoff of > 3. We did this to potentially identify gene markers of resilience in the group exposed to pBD-5.
Results
Overview of sequencing data
The total number of high-quality reads was 78,100,000, averaging 5,210,000/sample, resulting in 12,445 gene transcripts retained for analysis (at least 0,5 counts per million, CPM).
Differentially expressed genes
Treatments with the greatest differences in the number of differentially expressed genes were LPS versus Brachy, LPS versus pBD + Brachy, Brachy versus pBD, Brachy versus Control, and pBD + Brachy versus Control (Fig. 1).
The bar plot illustrates the number of differentially expressed genes across the specified contrasts. Total number of genes upregulated in each contrast are represented in red, while downregulated genes are shown in blue. Contrasts include: LPS versus Brachy, LPS versus pBD + Brachy, Brachy versus pBD + Brachy, LPS versus pBD, Brachy versus pBD, pBD + Brachy versus pBD, LPS versus Control, Brachy versus Control, pBD + Brachy versus Control, and pBD versus Control. A dotted-line box highlights the contrast “Brachy versus Control,” and a solid-line box emphasizes “Brachy versus pBD + Brachy,” as these contrasts are central to the discussion in this manuscript.
Exposure of intestinal cells to B. hyodysenteriae caused significant differences compared to the Control group (Fig. 1). However, co-exposure of host cells to pBD-5 and B. hyodysenteriae (pBD + Brachy) reduced the number of genes differentially expressed between the groups (Fig. 1).
Upon contrasting the expression results of the Control group with those of the pBD group, our analysis revealed that only one gene was significantly impacted: upregulation of DNA Damage Inducible Transcript 4 (DDIT4) gene in the pBD-exposed group (fold change = 1.912, P = 0.026).
Interestingly, we identified five genes with fold-changes greater than 3 (Rho Guanine Nucleotide Exchange Factor 10 Like, ARHGEF10L; Solute Carrier Family 38 Member 3, SLC38A3; Heat Shock Protein Family A (Hsp70) Member 1 Like, HSPA1L; Thioredoxin Interacting Protein, TXNIP; and Lipopolysaccharide Binding Protein, LBP) that were uniquely downregulated and three genes (ENSSSCG00000041859, ENSSSCG00000046276, and ENSSSCG00000047131) that were uniquely upregulated in the pBD + Brachy group compared to the Brachy group (Fig. 2). These genes were not differentially expressed when comparing the pBD + Brachy to the Control group (Fig. 2). While the biological implications of these gene expression changes are explored in detail in the discussion section, the observed regulation patterns suggest a potential role for pBD-5 in modulating immune responses (LBP, LAG3, HAMP), stress-related pathways (HSPA1L, TXNIP, ARHGEF10L), and pathogen resilience mechanisms (MMP25, ANPEP, KLHL40, VWA5B2, SPTBN5, DISP2).
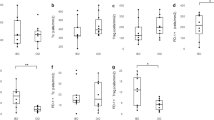
Venn diagram illustrating differentially expressed genes (DEGs) across three experimental groups: pBD + Brachy-Control (red), Brachy-pBD + Brachy (blue), and Brachy-Control (green). Up (a) and downregulated (b) DEGs are contrasted by color, where Red vs. Blue represents DEGs between pBD + Brachy-Control and Brachy-pBD + Brachy; Red vs. Green: DEGs between pBD + Brachy-Control and Brachy-Control; and Blue vs. Green: DEGs between Brachy-pBD + Brachy and Brachy-Control. Genes uniquely upregulated or downregulated in Brachy-Control compared to Brachy-pBD + Brachy are highlighted and asterisks denote statistically significant differences at alpha level of 0.05. Column heights represent mean values, while error bars indicate standard deviation (SD).
Gene set enrichment analysis (GSEA)
When contrasting the Control and pBD-only groups, pathway analysis revealed that the JAK-STAT signalling pathway (P < 0.0001) and cytokine-cytokine receptor interaction pathway (P < 0.0001) were among the most significantly enriched pathways (Fig. 3). Furthermore, we observed that NOD-like receptor signalling pathway (P = 0.041) was significantly upregulated when contrasting Brachy versus pBD + Brachy groups and the ribosome (P = 0.0055) and protein digestion and absorption pathways (P = 0.041) were significantly downregulated in the pBD-Brachy exposed group in comparation to the Brachy group (Fig. 4).
Significantly affected pathways when contrasting pBD versus Control groups. Pre-ranked GSEA approach with a pathway significance cut-off FDR = 0.1. The Kyoto Encyclopedia of Genes and Genomes (KEGG) database of genes was used. (a) Two pathways (nodes) are connected if they share 10% or more genes. Red and green represents up- and down-regulated pathways, respectively. Darker nodes are more significantly enriched gene sets. Bigger nodes represent larger gene sets. Thicker edges represent more overlapped genes. (b, c) Top differentially expressed genes in the most significantly enriched upregulated (red) or downregulated (green) pathways.
Top affected pathways when contrasting Brachy versus pBD-Brachy groups. Pre-ranked fgsea GSEA approach with a pathway significance cut-off FDR = 0.1. The Kyoto Encyclopedia of Genes and Genomes (KEGG) database of genes was used. (a) Two pathways (nodes) are connected if they share 10% or more genes. Red and green represents up- and down-regulated pathways, respectively. Darker nodes are more significantly enriched gene sets. Bigger nodes represent larger gene sets. Thicker edges represent more overlapped genes. (b, c, d, e, and f) Top differentially expressed genes in the most significantly enriched upregulated (red) or downregulated (green) pathways.
LDH-cytotoxicity and metabolic activity assays
No significant differences were observed among treatments regarding membrane integrity (P = 0.1044) or metabolic activity (P = 0.1958) compared to the control group (Fig. 5). A numerical trend is observed on the metabolic activity of host cells exposed to pBD-5 (Fig. 5b).
Assessment of membrane integrity (a) and metabolic response (b) in IPEC-1 cells following non-Brachyspira treatments after 8 h of incubation. Experimental conditions were tested with five replicates per treatment group. Column heights represent mean values, while error bars indicate standard deviation (SD).
Antimicrobial activity assay
There is no evidence that pBD-5 had an effect on B. hyodysenteriae growth independently of the concentration of pBD-5 used (P = 0.4361) (Fig. 6).
Direct antimicrobial activity of pBD-5 on B. hyo (B. hyodysenteriae) after 48 h of incubation. The figure represents the direct activity of various concentrations of pBD-5 (0.2516 µM, 2.516 µM, and 251.6 µM) on B. hyodysenteriae, expressed as fold change. Experimental conditions were tested with three replicates per treatment group, in three independent trials. Column heights represent mean values, and error bars indicate standard deviation (SD).
Discussion
This study aimed to explore the effects of pBD-5 on porcine intestinal cells and B. hyodysenteriae through immunomodulation and direct bacterial killing. We uncovered that pBD-5 mitigated mRNA level differences in IPEC-1 cells compared to the group solely exposed to B. hyodysenteriae. Additionally, we identified important immune-related pathways that were significantly enriched when cells were co-exposed to B. hyodysenteriae and pBD-5, when compared to B. hyodysenteriae alone. This included the IL-17, toll-like receptor (TLR), and NOD-like receptor signalling pathways. Our results suggest a potential immunomodulatory role of pBD-5 and are consistent with previous research on the regulatory mechanisms of pBDs to mitigate disease progression29,30,31,32,33. β-defensins modulate inflammation by inducing or repressing chemokine production34, promoting chemotaxis of specialized cells35, attenuating exacerbated nitric oxide (NO) production36, and ultimately, orchestrating responses of other immune and epithelial cells7,32,33,37. Interestingly, Huang et al.30 showed that the porcine β-defensin 2 (pBD-2) can bind to TLR4 receptors and downregulate the NFkβ signalling cascade, alleviating inflammation. In their model, the interference with this pathway ultimately lead to a reduced production of pro-inflammatory cytokines and chemokines. Similarly, pBD-5 also appears to downregulate the TLR4/NF-κB signaling pathway, exhibiting an anti-inflammatory profile. In the IL-17 signaling pathway, we observed that the genes C-X-C Motif Chemokine Ligand 8 (CXCL8, encoding interleukin-8 IL-8), TNF Alpha-Induced Protein 3 (TNFAIP3, encoding A20), NFKB Inhibitor Alpha (NFKBIA, encoding IκBα), C-X-C Motif Chemokine Ligand 2 (CXCL2, encoding macrophage inflammatory protein-2α MIP-2α), and C-X-C Motif Chemokine Ligand 10 (CXCL10, encoding interferon gamma-induced protein 10 IP-10) were downregulated compared to the group exposed solely to Brachyspira hyodysenteriae. Notably, the overexpression of IL-17 has been previously identified in the cascade of mucus production during the pathogenesis of B. hyodysenteriae in pigs38. The authors demonstrated that elevated IL-17 levels were linked to increased mucus secretion during the colitis process38. There is evidence that human β-defensin 2 (hBD-2) suppress the production of IL-17 in T cells while enhancing the production of IFN-γ and IL-1039. Here we propose that pBD-5 mitigate B. hyodysenteriae-associated exacerbated inflammatory response during the onset of infection. This hypothesis, however, needs further research, as the measurement of proinflammatory mediators such as IL-8, TNF-α, IL-6, IL-1β, and IL-12 would better clarify the mechanisms of action. NOD-like receptor signaling pathway was found to be downregulated in the pBD-5 + Brachy group, when compared to B. hyodysenteriae alone. This finding provides further indication of how pBD-5 interacts with host innate immunity receptors. NOD-like receptors are intracellular pattern recognition receptors that play a crucial role in activating the inflammasome response40. Inflammasome formation ultimately leads to the activation of caspase-1, which cleaves and activates pro-inflammatory cytokines such as IL-1β and IL-1840,41, and regulates key innate responses42,43,44,45. In this context, our results once again suggest that pBD-5 plays a role at downregulating potentially exacerbated immune responses during the early stage of infection by B. hyodysenteriae. When comparing the enriched biological processes of the group solely exposed to pBD-5 with the control group, the JAK-STAT and cytokine-cytokine receptor interaction pathways were found to be the top upregulated pathway in the group solely exposed to pBD-5 compared to the control group. These pathways are crucial transcription regulators of various immunity-related genes, such as interferon regulatory factor 1 (IRF1), interferon gamma receptor 1 (IFNγR1), interleukin-12 receptor beta 1 (IL12RB1), signal transducer and activator of transcription 1 (STAT1) and janus kinase 2 (JAK2) gene46. The activation of this pathway is important for the proper functioning of the immune system and its upregulation can help to mount an effective immune response against infections through regulating cytokine-dependent pathways46,47.
Investigating resilience marker genes is important for understanding the molecular mechanisms that enhance cellular defense against pathogens48,49. Here we found that treating IPEC-1 cells with pBD-5 during a B. hyodysenteriae challenge resulted in 21 differentially expressed genes (DEGs) with a minimum fold-change of 3. Among these, 8 genes were restored to their baseline expression levels by pBD-5, while 13 were upregulated compared to cells exposed solely to B. hyodysenteriae. Rho Guanine Nucleotide Exchange Factor 10 Like (ARHGEF10L), one of the genes regulated to basal levels during the challenge with B. hyodysenteriae, was identified by Liepelt et al.50 as associated with poorer prognosis in sepsis patients when upregulated. Similarly, Heat Shock Protein Family A (Hsp70) Member 1 Like (HSPA1L), a heat shock protein gene, which was shown to be involved in the pathogenesis of SARS-CoV-2 infection51 and in modulating cytokine concentrations and clinical outcomes after major trauma52, was fetched to basal levels of expression after pBD-5 exposure. Thioredoxin Interacting Protein (TXNIP) is one of the genes involved in modulating oxidative stress, regulating redox homeostasis, and activating the inflammasome53,54,55. Under oxidative stress, the TXNIP protein dissociates from thioredoxin (TRX) and binds to the NLRP3 inflammasome, leading to the activation and release of IL-1β55. Experiments with TXNIP downregulation and knockout mice confirmed its important role in NLRP3 inflammasome activation55. In B. hyodysenteriae infection, pBD-5 treatment helped in restraining the expression of TXINP, making it a candidate for further investigations. Lipopolysaccharide Binding Protein (LBP), an essential recognizing and clearing component of lipopolysaccharides (LPS), as evidenced by both in vitro and in vivo studies56,57,58, was also identified as downregulated to basal levels by pBD-5. While LPS is not a major structural component of B. hyodysenteriae, LBP expression may still be induced by lipooligosaccharide (LOS), the main endotoxin forming the cell wall of B. hyodysenteriae59. The upregulation above basal levels of Matrix Metallopeptidase 25 (MMP25), Kelch Like Family Member 40 (KLHL40), Von Willebrand Factor A Domain Containing 5B2 (VWA5B2), Spectrin Beta, Non-Erythrocytic 5 (SPTBN5), Radial Spoke Head 6 Homolog A (RSPH6A), Hepcidin Antimicrobial Peptide (HAMP), Lymphocyte Activating 3 (LAG3), Dispatched RND Transporter Family Member 2 (DISP2), Alanyl Aminopeptidase, Membrane (ANPEP), and Testis Expressed 52 (TEX52), alongside the Ensembl-annotated genes ENSSSCG00000041859, ENSSSCG00000046276, ENSSSCG00000047720, ENSSSCG00000038331, ENSSSCG00000047131, and ENSSSCG00000044657, may represent an even more relevant mechanism of protection against Brachyspira hyodysenteriae. MMP25, a metalloproteinase, may facilitate extracellular matrix remodeling and immune cell recruitment, critical for tissue repair in infection60,61. HAMP, an antimicrobial peptide, likely restricts bacterial access to iron, a key nutrient, by modulating host iron homeostasis62,63. LAG3, another key player in immune modulation, regulates T-cell responses, which could help balance antimicrobial activity and limit excessive inflammation64,65. The remaining genes, including ANPEP, KLHL40, VWA5B2, SPTBN5, DISP2, are less clearly linked to infection or defensin-mediated mechanisms, and their upregulation may reflect indirect effects, such as cellular stress, signaling cascades, or compensatory responses. While in this paragraph we have potentially identified genes associated with increased resilience induced by pBD-5, their involvement is likely much more complex and require extensive investigation as comparations here were made across different species.
Interestingly, we also observed that pBD-5 modulates metabolic modulation pathways. Oxidative phosphorylation, ribosome and steroid hormone biosynthesis pathways were found to be downregulated among cells exposed to our novel pBD-5. We hypothesize that pBD-5 exposure under non-challenging circumstances may help boost energy reserves and direct energy towards mitigating infections by activating immune pathways involved in producing chemokines and cytokines. This hypothesis is supported by evidence that immune cells need to rapidly adapt their metabolism to respond to infections, and this process may involves reducing mitochondrial oxidative phosphorylation activity66,67,68. Our analysis of mitochondrial activity in groups exposed to pBD-5 did not yield any statistically significant differences. However, there is a noticeable numerical difference between the groups exposed to pBD-5. Longer incubation period may have been beneficial in this case, as well as larger sample size.
In this study, exposure of B. hyodysenteriae to various concentrations of pBD-5 did not demonstrate direct antimicrobial activity on bacterial growth. We attribute this lack of efficacy to the chemical synthesis approach used. β-defensins are known to exhibit greater antimicrobial activity when they contain disulfide bonds69. These bonds are crucial for the proper folding and structural integrity of the peptide, which in turn enhances its ability to interact with and disrupt bacterial membranes69. We synthesized pBD-5 with its cysteine residues protected by acetamidomethyl groups. This protection prevents the formation of disulfide bonds, resulting in a linear peptide conformation that lacks the structural features necessary for potent antimicrobial activity. The linear form of pBD-5 was likely unable to adopt the structure required for optimal interaction with bacterial cell membranes, leading to its reduced direct antimicrobial activity.
Most of our key findings are based on gene set enrichment analyses, which presents a limitation as these results are primarily associative. Thus, while informative, these findings require further experimental validation to confirm direct causal links and ensure robustness. Secondly, our study had a relatively small number of replicates per treatment and employed IPEC-1 cells, which are derived from the porcine small intestine, which is not the target site for B. hyodysenteriae. These cells were used given the lack of commercially available immortalized swine colonic cells. Despite this, they did present as suitable models for our study, as we could see a clear distinction expression pattern across the groups exposed to Brachyspira. Lastly, our study did not measure cell viability before and after exposure to B. hyodysenteriae treatments, which would be important to verify if Brachyspira-treatments played a role at inducing cell damage or apoptosis, and to measure if pBD-5 may have mitigated this effect at some level.
Future research on pBD-5 may investigate its interactive role with toll-like and NOD-like receptors, as well as the cascades involved in the production of inflammatory cytokines and chemokines. Testing different chemical synthesis approaches such as the ones that involve disulfide bonds formation, would be valuable to confirm if pBD-5 may have a direct antimicrobial activity as well, or if its immunomodulatory activity can be enhanced.
pBD-5 was found to reduce the number of differentially expressed genes in IPEC-1 cells exposed to B. hyodysenteriae, and to downregulate the IL-17, toll-like receptor, and NOD-like receptor signalling pathways, suggesting that pBD-5 plays a role in immunomodulation. Additionally, pBD-5 modulates cell metabolism. These findings provide insights into the potential mechanism of action of pBD-5 in mitigating exacerbated inflammatory response in the early stage of B. hyodysenteriae infection. Further in vivo research is needed to clarify its use as an alternative treatment for infectious diseases where antibiotics are the only available option.
Data availability
The data that support the findings of this study are not openly available due to reasons of sensitivity and are available from the corresponding author upon reasonable request. Data are located in controlled access data storage at the University of Saskatchewan.
References
Tang, K. L. et al. Restricting the use of antibiotics in food-producing animals and its associations with antibiotic resistance in food-producing animals and human beings: A systematic review and meta-analysis. Lancet Planet. Health 1, e316–e327 (2017).
Angst, D. C., Tepekule, B., Sun, L., Bogos, B. & Bonhoeffer, S. Comparing treatment strategies to reduce antibiotic resistance in an in vitro epidemiological setting. Proc. Natl. Acad. Sci. U. S. A. 118, 1–7 (2021).
Javed, A. et al. Antimicrobial and immunomodulatory activities of porcine cathelicidin Protegrin-1. Mol. Immunol. 173, 100–109 (2024).
Lin, Q. et al. Functional characterization of porcine nk-lysin: a novel immunomodulator that regulates intestinal inflammatory response. Molecules 26, (2021).
Sang, Y., Ramanathan, B., Minton, J. E., Ross, C. R. & Blecha, F. Porcine liver-expressed antimicrobial peptides, hepcidin and LEAP-2: Cloning and induction by bacterial infection. Dev. Comp. Immunol. 30, 357–366 (2006).
Sang, Y., Ramanathan, B., Ross, C. R. & Blecha, F. Gene silencing and overexpression of porcine peptidoglycan recognition protein long isoforms: Involvement in β-defensin-1 expression. Infect. Immun. 73, 7133–7141 (2005).
Elahi, S. et al. The host defense peptide beta-defensin 1 confers protection against Bordetella pertussis in newborn piglets. Infect. Immun. 74, 2338–2352 (2006).
Meade, K. G. & O’Farrelly, C. Β-Defensins: Farming the microbiome for homeostasis and health. Front. Immunol. 10, 1–20 (2019).
Koleva, P. et al. CD71 + erythroid cells promote intestinal symbiotic microbial communities in pregnancy and neonatal period. Microbiome 12, 1–23 (2024).
Wu, J., Ma, N., Johnston, L. J. & Ma, X. Dietary nutrients mediate intestinal host defense peptide expression. Adv. Nutr. 11, 92–102 (2020).
Miani, M. et al. Gut microbiota-stimulated innate lymphoid cells support β-Defensin 14 expression in pancreatic endocrine cells, preventing autoimmune diabetes. Cell. Metab. 28, 557–572e6 (2018).
Zeng, X. et al. Induction of porcine host defense peptide gene expression by short-chain fatty acids and their analogs. PLoS One 8, 1–8 (2013).
Whitmore, M. A., Li, H., Lyu, W., Khanam, S. & Zhang, G. Epigenetic regulation of host defense peptide synthesis: Synergy between histone deacetylase inhibitors and DNA/Histone methyltransferase inhibitors. Front. Immunol. 13, 1–12 (2022).
Liu, T., Sun, Z., Yang, Z. & Qiao, X. Microbiota-derived short-chain fatty acids and modulation of host-derived peptides formation: Focused on host defense peptides. Biomed. Pharmacother. 162, 114586 (2023).
Chen, R., bo, Zhang, K., Zhang, H., Gao, C. & Li, C. li. Analysis of the antimicrobial mechanism of porcine beta defensin 2 against E. coli by electron microscopy and differentially expressed genes. Sci. Rep. 8, 1–14 (2018).
Huang, C. et al. The intracellular interaction of porcine β-Defensin 2 with VASH1 alleviates inflammation via akt signaling pathway. J. Immunol. 208, 2795–2805 (2022).
Zhang, K. et al. Recombinant porcine beta defensin 2 alleviates inflammatory responses induced by Escherichia coli in IPEC-J2 cells. Int. J. Biol. Macromol. 208, 890–900 (2022).
Costa, M. O. & Harding, J. C. S. Swine dysentery disease mechanism: Brachyspira hampsonii impairs the colonic immune and epithelial repair responses to induce lesions. Microb. Pathog. 148, 104470 (2020).
Ryan, M. 0_1 Evaluating the economic benefits and costs of antimicrobial use in food-producing animals. OECD Food, Agric. Fish. Pap. 2, 6–39 (2019).
Rubin, J. E. et al. Reproduction of mucohaemorrhagic diarrhea and colitis indistinguishable from swine dysentery following experimental inoculation with ‘Brachyspira hampsonii’ strain 30446. PLoS One 8, (2013).
Bourgey, M. et al. GenPipes: An open-source framework for distributed and scalable genomic analyses. Gigascience 8, 1–11 (2019).
Bolger, A. M., Lohse, M. & Usadel, B. A flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Dobin, A. et al. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Anders, S., Pyl, P. T. & Huber, W. HTSeq-A python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Ge, S. X., Son, E. W., Yao, R. & iDEP An integrated web application for differential expression and pathway analysis of RNA-Seq data. BMC Bioinform. 19, 1–24 (2018).
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Witters, N. A. & Duhamel, G. E. Cell membrane permeability and mitochondrial dysfunction-inducing activities in cell-free supernatants from Serpulina hyodysenteriae serotypes 1 and 2. Comp. Immunol. Microbiol. Infect. Dis. 19, 233–244 (1996).
Huang, J. et al. Pigs overexpressing porcine β-defensin 2 display increased resilience to Glaesserella parasuis infection. Antibiotics 9, 1–12 (2020).
Huang, C. et al. Porcine beta-defensin 2 provides protection against bacterial infection by a direct bactericidal activity and alleviates inflammation via interference with the TLR4/NF-κB pathway. Front. Immunol. 10, 1–14 (2019).
Bowdish, D. M. E., Davidson, D. J. & Hancock, R. E. W. Immunomodulatory properties of defensins and cathelicidins. Curr. Top. Microbiol. Immunol. 306, 27–66 (2006).
Semple, F. & Dorin, J. R. β-Defensins multifunctional modulators of infection, inflammation and more? J. Innate Immun. 4, 337–348 (2012).
Dou, X. et al. TLR2/4-mediated NF-κb pathway combined with the histone modification regulates β-defensins and interleukins expression by sodium phenyl butyrate in porcine intestinal epithelial cells. Food Nutr. Res. 62, 1–13 (2018).
Koeninger, L. et al. Human β-Defensin 2 mediated immune modulation as treatment for experimental colitis. Front. Immunol. 11, 1–16 (2020).
Röhrl, J., Yang, D., Oppenheim, J. J. & Hehlgans, T. Human β-Defensin 2 and 3 and their mouse orthologs induce chemotaxis through interaction with CCR2. J. Immunol. 184, 6688–6694 (2010).
Lee, J. Y. et al. Identification of a cell-penetrating peptide domain from human beta-defensin 3 and characterization of its anti-inflammatory activity. Int. J. Nanomed. 10, 5423–5434 (2015).
Yang, D. et al. β-Defensins: Linking innate and adaptive immunity through dendritic and T cell CCR6. Sci. (80-). 286, 525–528 (1999).
Quintana-Hayashi, M. P. Neutrophil elastase and Interleukin 17 Brachyspira hyodysenteriae infection synergistically with the pathogen induce increased mucus transport speed and. 85, 1–16 (2017).
Kanda, N. et al. Human β-defensin-2 enhances IFN-γ and IL-10 production and suppresses IL-17 production in T cells. J. Leukoc. Biol. 89, 935–944 (2011).
Amarante-Mendes, G. P. et al. Pattern recognition receptors and the host cell death molecular machinery. Front. Immunol. 9, 1–19 (2018).
Kelley, N., Jeltema, D., Duan, Y. & He, Y. The NLRP3 inflammasome: An overview of mechanisms of activation and regulation. Int. J. Mol. Sci. 20, 3328 (2019).
Zhen, Y. & Zhang, H. NLRP3 inflammasome and inflammatory bowel disease. Front. Immunol. 10, 1–10 (2019).
Franchi, L., Warner, N., Viani, K. & Nuñez, G. Function of nod-like receptors in microbial recognition and host defense. Immunol. Rev. 227, 106–128 (2009).
Martinon, F., Pétrilli, V., Mayor, A., Tardivel, A. & Tschopp, J. Gout-associated uric acid crystals activate the NALP3 inflammasome. Nature 440, 237–241 (2006).
Dostert, C. et al. Innate immune activation through Nalp3 inflammasome sensing of asbestos and silica. Science 320, 674–677 (2008).
Seif, F. et al. The role of JAK-STAT signaling pathway and its regulators in the fate of T helper cells. Cell. Commun. Signal. 15, 1–13 (2017).
OShea, J. J., Gadina, M. & Schreiber, R. D. Cytokine signaling in 2002: New surprises in the Jak/Stat pathway. Cell 109, 121–131 (2002).
Bai, X. et al. Investigating the genetic architecture of disease resilience in pigs by genome-wide association studies of complete blood count traits collected from a natural disease challenge model. BMC Genom. 22, 1–15 (2021).
Nery da Silva, A., Silva Araujo, M., Pértille, F. & Zanella, A. J. How epigenetics can. Enhance pig welfare? Animals 12, 32 (2021).
Liepelt, A. et al. Differential gene expression in circulating CD14 + monocytes indicates the prognosis of critically ill patients with sepsis. J. Clin. Med. 9, 1–22 (2020).
Muhammad, J. S., Sharif-Askari, S., Cui, N., Hamad, Z. G., Halwani, R. & M. & SARS-CoV-2 infection-Induced promoter hypomethylation as an epigenetic modulator of heat shock protein A1L (HSPA1L) gene. Front. Genet. 12, 1–10 (2021).
Schröder, O. et al. Heat shock protein 70 genotypes HSPA1B and HSPA1L influence cytokine concentrations and interfere with outcome after major injury. Crit. Care Med. 31, 73–79 (2003).
Pan, M., Zhang, F., Qu, K., Liu, C. & Zhang, J. T. X. N. I. P. A double-edged sword in disease and therapeutic outlook. Oxid. Med. Cell. Longev. (2022).
Choi, E. H., Park, S. J. & TXNIP A key protein in the cellular stress response pathway and a potential therapeutic target. Exp. Mol. Med. 55, 1348–1356 (2023).
Zhou, R., Tardivel, A., Thorens, B., Choi, I. & Tschopp, J. Thioredoxin-interacting protein links oxidative stress to inflammasome activation. Nat. Immunol. 11, 136–140 (2010).
Knapp, S., Florquin, S., Golenbock, D. T. & Poll, T. Van Der. The LPS-Induced lung inflammation in vivo 1. J. Immunol. (2006).
Sun, Y. et al. Bovine recombinant lipopolysaccharide binding protein (BRLBP) regulated apoptosis and inflammation response in lipopolysaccharide-challenged bovine mammary epithelial cells (BMEC). Mol. Immunol. 65, 205–214 (2015).
Zweigner, J., Gramm, H. J., Singer, O. C., Wegscheider, K. & Schumann, R. R. High concentrations of lipopolysaccharide-binding protein in serum of patients with severe sepsis or septic shock inhibit the lipopolysaccharide response in human monocytes. Blood 98, 3800–3808 (2001).
Herbst, W., Schneider, S., Baljer, G. & Barth, S. A. An update of Brachyspira hyodysenteriae serotyping. Res. Vet. Sci. 111, 135–139 (2017).
He, L. et al. The immunomodulatory role of matrix metalloproteinases in colitis-associated cancer. Front. Immunol. 13, (2023).
Premadasa, L. S. et al. Cannabinoid enhancement of lncRNA MMP25-AS1/MMP25 interaction reduces neutrophil infiltration and intestinal epithelial injury in HIV/SIV infection. JCI Insight 8, 1–22 (2023).
Krepuska, M. et al. Bone marrow stromal cell-derived hepcidin has antimicrobial and immunomodulatory activities. Sci. Rep. 14, 1–13 (2024).
Barroso, C. et al. The era of antimicrobial peptides: use of Hepcidins to prevent or treat bacterial infections and Iron disorders. Front. Immunol. 12, 1–17 (2021).
Tian, X. et al. The upregulation of LAG-3 on T cells defines a subpopulation with functional exhaustion and correlates with disease progression in HIV-infected subjects. J. Immunol. 194, 3873–3882 (2015).
Huang, C. T. et al. Role of LAG-3 in regulatory T cells. Immunity 21, 503–513 (2004).
O’Neill, L. A. J. & Pearce, E. J. Immunometabolism governs dendritic cell and macrophage function. J. Exp. Med. 213, 15–23 (2016).
Olson, G. S. et al. Type I interferon decreases macrophage energy metabolism during mycobacterial infection. Cell. Rep. 35, 109195 (2021).
Andrews, J. T., Voth, D. E., Huang, S. C. C. & Huang, L. Breathe in, breathe out: Metabolic regulation of lung macrophages in host defense against bacterial infection. Front. Cell. Infect. Microbiol. 12, 1–8 (2022).
Ma, Y. et al. Key role of disulfide bridges in the antimicrobial activity of beta-defensin from Olive Flounder. Int. J. Pept. Res. Ther. 26, 291–299 (2020).
Acknowledgements
The authors gratefully acknowledge financial support from NSERC for funding the research project. ANF was supported by a University of Saskatchewan Harvey Graduate Scholarship as well as the WCVM Research Trust Graduate Student Scholarship and the Large Animal Clinical Sciences - CGPS 75th & SSF Award. We also recognize the contributions and hard work of undergraduate and graduate students who collaborated on antimicrobial activity assays, including Sulove Koirala, Sam Nixon, Jessica Barbosa, Lucie Amblard, Allyssa Cloet, Barbara Cortez, Maite Almeida, and Fernanda Facioli. Special thanks to the Glass and Media Preparation team, particularly Melinda Wyshynski and Ana Rodriguez, at the WCVM for their fast and high-quality services in supporting our research agenda.
Author information
Authors and Affiliations
Contributions
C.T.Y. and A.N.F. conducted the experiments and collected the data. A.N.F. analyzed the data. A.N.F. drafted the first manuscript version, including figured. M.O.C. reviewed the methodology used, the data and data analyses and the manuscript drafts. All authors reviewed the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Finatto, A.N., Yang, C. & de Oliveira Costa, M. Porcine β-defensin 5 (pBD-5) modulates the inflammatory and metabolic host intestinal response to infection. Sci Rep 15, 7568 (2025). https://doi.org/10.1038/s41598-025-90688-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-90688-8