Abstract
Lamin B1 (LMNB1) is an intermediate filament protein that is an integral component of the nuclear lamina, a structure that is critical for nuclear organization and function. Mutations involving the lamin B1 gene cause the adult-onset demyelinating disorder, Autosomal Dominant Leukodystrophy (ADLD) which is charactered by increased lamin B1 expression. Increased LMNB1 expression is also associated with poorer outcomes in multiple cancer subtypes. Reducing LMNB1 is thus an attractive therapeutic pathway for ADLD and potentially other diseases. Here we present the results of a high throughput / high content screen (HTS/HCS) to identify small molecules that reduce LMNB1 levels. Approximately 97,000 molecules were screened using an inducible mouse fibroblast model of LMNB1 overexpression that we have previously generated. Two small molecules, Pubchem CID 662896 and CID 5308648, were identified that reduced LMNB1 in a dose dependent manner without causing cellular toxicity and corrected nuclear abnormalities associated with LMNB1 overexpression, a hallmark of ADLD. CID 662896 also reduced LMNB1 levels in ADLD patient fibroblast samples, exhibited favorable “drug-like” physicochemical properties and crossed the blood brain barrier in mouse studies. While CID 662896 may be a promising candidate for ADLD therapy, further investigations are required to determine its mechanism of action and ability to target disease relevant cell types.
Similar content being viewed by others
Introduction
The nuclear lamina is a fibrous meshwork of intermediate filament proteins that is found adjacent to the inner nuclear envelope. It is an integral component of the nuclear envelope in all eukaryotic cells and was originally thought to play a purely structural role. However, the recent years have witnessed an explosion in our understanding of the functions of the nuclear lamina. Together with maintaining the structural integrity of the nucleus, it also plays an important role in essential cellular processes such as transcription, DNA replication and repair, and regulation of various epigenetic phenomena1,2. In addition, the nuclear lamina has been shown to play an important role in both normal and pathological aging3,4.
In vertebrates there are two main lamin subtypes – the A and B type lamins. The A type lamins (LMNA) include lamin A and lamin C and are splice isoforms encoded by a single gene lamin A/C. The B type lamins, lamin B1 and B2 (LMNB1 and LMNB2), are encoded by two independent genes1,5. Lamin B3 is a germline specific alternatively spliced isoform of the LMNB2gene. Type A and B lamins have distinct spatial localizations within the lamina and carry out many independent functions6,7.
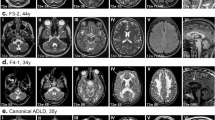
More than a dozen diseases have been associated with mutations in LMNA while diseases caused by B type lamin variants are much less common4,8. Diseases caused by mutations in the lamin genes are collectively known as laminopathies4. Two distinct monogenic disease phenotypes are caused by mutations involving the LMNB1gene. The first disease identified was Autosomal Dominant Leukodystrophy (ADLD), a fatal, adult onset, progressive neurological disease that is characterized by widespread CNS demyelination9,10. Most cases of ADLD are caused by tandem genomic duplications (ADLD-dup) involving the LMNB1 gene while a small subset is caused by genomic deletions upstream of LMNB111,12. Both these mutations are thought to cause increased CNS LMNB1 expression and are 100% penetrant i.e., all individuals with the mutation develop the disease10. The second pathological phenotype is that of microcephaly caused by de-novo mutations within LMNB1gene that are thought to disrupt nuclear integrity13.
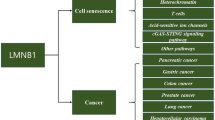
Apart from disorders directly caused by mutations in LMNB1, other pathological conditions including many cancers have been associated with alterations in LMNB1 levels8. Increased LMNB1 levels were associated with poorer prognosis in pancreatic cancer and renal cell carcinoma while they were associated positively with tumor stages, tumor sizes, and number of nodules in hepatocellular carcinoma14,15,16. Conversely, in breast cancer, the levels of LMNB1 declined with worsening clinical outcome and LMNB1 knockdown promoted gastric cancer cell proliferation and migration17,18. Overexpression of LMNB1 was also reported in ataxia telangiectasia (A-T) and Werner syndromes, which are associated with genetic instability, cancer predisposition, and premature aging19,20.
Given that LMNB1 overexpression underlies ADLD, other laminopathies, and cancers, we sought to carry out a high throughput screen (HTS) to identify small molecules that can reduce LMNB1 levels. The primary assay is based on the use of an inducible cell culture model of LMNB1 overexpression that we have previously described21. In this study we present the results of a high-content screen of ~ 97,000 small molecules from the NIH Molecular Libraries Small Molecule Repository (MLSMR) collection.
Results
Assay validation using the Library of Pharmacologically Active Compounds (LOPAC)
We previously developed and optimized a high throughput/high content screening assay for detection of LMNB1 overexpression and nuclear abnormalities21. The next phase in development was to validate the assay in a small-scale pilot library screen. The LOPAC library is a 1,280 compound collection of annotated biologically active small molecules that cover key drug targets and signaling pathways. Information from this library is used to learn about assay behavior under screening conditions, obtain guidance for establishing hit selection criteria, expected hit and confirmation rates, false positives, and susceptibility to assay interference. To fully exploit the information contained in the LOPAC pilot library, it was screened at three concentrations (30 µM, 10 µM, and 3 µM) in duplicate. Screening at multiple concentrations allows for potentially active compounds that at high concentrations are toxic or morphology-altering to be identified at one of the lower concentrations. Mouse TRE-FLAG-LMNB1MEFs were plated and treated as described in the Materials and Methods Section. Plates were imaged on an Arrayscan VTI for LMNB1 expression, texture, LMNA/C intensity, and nuclear shape morphology as previously described21. Assay performance was assessed by Linear Discriminant Analysis (LDA), which combines several single assay parameters that in aggregate describe the phenotype of positive (DOX induced) and negative (DOX withdrawal) controls, and by LMNB1 intensity as the primary biologically relevant parameter. As expected from assay development21, LDA separated MAX (DOX-induced) and MIN (DOX withdrawal) controls with Z’ values22 of 0.56 ± 0.10 (30 µM), 0.53 ± 0.11 (10 µM), and 0.56 ± 0.07 (3 µM) (Fig. S1A and table). Because we were ultimately interested in selecting compounds that reduce LMNB1 expression, we also established QC measures for LMNB1 intensity. As reported before21 and expected for an intensity-based screen in primary cells, this parameter was substantially more variable than LDA and presented with larger numbers of outliers (Fig. S1B and table). We therefore applied robust statistics using median and median absolute deviation (MAD) as recommended by Haelewyn et al.23. The LMNB1 intensity parameter separated MIN and MAX controls with robust Z-prime (rZ’) values of 0.52 ± 0.12 (30 µM), 0.40 ± 0.15 (10 µM), and 0.45 ± 0.05 (3 µM) and signal-to-backgrounds (S/B) above 2.9 (Fig. S1, table). These data indicated that despite the complex nature and inherent variability of primary cells, the LMNB1 parameter would be suited to establish a primary hit criterion. Although the LMNB1 intensity parameter did not perform as well as LDA, there was a clear correlation (r2 = 0.93) between the two parameters (Fig. S1C).
We then proceeded to establish an activity criterion for hit selection. Using the robust Z-prime calculation for LMNB1 intensity as guidance we chose an initial activity cutoff of < median—3 MAD for each concentration tested (Fig. 1A). Applying this statistics-based threshold, the number of compounds identified at each concentration were 57 (30 µM), 26 (10 µM), and 16 (3 µM) (Fig. 1B). As might be expected, the 30 µM plates had a higher hit rate (57 compounds, 4.1%) compared to the 10 µM concentration (26 compounds, 2.0%) and the 3 µM plate (16 compounds, 1.1%) (Fig. 1B). Whereas these hit rates would be too high for HTS, they were purposefully accepted here because one important function of pilot libraries is to develop a hit selection scheme that in large scale HTS identifies the most promising compounds with the potential to be validated in follow-up assays. Application of the median – 3 MAD selection criterion to all concentrations yielded 92 active conditions, which were subjected to visual confirmation of archived scan images for selective reduction of LMNB1 without appreciable cell loss. This analysis yielded 57 unique compounds (some compounds hit at multiple concentrations), resulting in a hit rate of 4.5%, of which 53 were commercially available and repurchased (Fig. 1B).
Assay validation by LOPAC pilot library screen for reduction in LMNB1 protein levels. TRE-FLAG-LMNB1 MEF were used to screen the LOPAC at three concentrations (30 µM, 10 µM, and 3 µM) in duplicate. The cells were incubated for 48 h with library compounds in DOX containing medium. LMNB1 intensity was normalized to MAX/MIN controls. A. LMNB1 intensity graphs for each concentration tested compounds (blue), + DOX MAX control (yellow), and DOX withdrawal MIN control (red). Dotted line showing median of controls and solid lines represent median ± 3 MAD. Orange arrow indicates LMNB1 intensity threshold gate (median – 3 MAD). B. Detailed hit selection overview.
LOPAC hit confirmation
After identifying potential hits from the LOPAC pilot library screen, the next step was to confirm activity in ten-point two-fold dose–response curves in the primary screening assay, covering a concentration range from 100 µM – 200 nM. Plates were stained for LMNB1, LMNA/C, and nuclei as described in Materials and Methods, and analyzed for expression levels of LMNB1 and LMNA/C and cell numbers to investigate selectivity and toxicity. Surprisingly, only three compounds (SB202190, SP600125, and camptothecin) showed dose-dependent decreases in LMNB1 levels. (S)-( +)-camptothecin, a topoisomerase inhibitor and antineoplastic agent, showed an incomplete dose–response curve and cellular toxicity (data not shown). SB202190 and SP600125, a p38 MAPK and c-JNK/SAPK inhibitor, respectively, selectively reduced LMNB1 over LMNA/C with no or low toxicity, although the maximum response with SP600125 was small and the curve not sigmoidal (Fig. 2A). Because published reports had suggested an involvement of MAPKs in LMNB1 regulation (Barascu et al., 2012), we confirmed the activity of SB202190 and SP600125 in Western blots as non-fluorescent orthogonal assays (Fig. 2B, C). Whereas SP600125 did not confirm, consistent with its shallow dose curve in the immunofluorescence assay (Fig. 2A and data not shown), SB202190 concentration-dependently reduced both human-FLAG-LMNB1 and endogenous mouse LMNB1 (Fig. 2B, C).
LOPAC hit confirmation in TRE-FLAG-LMNB1 MEF. To validate the activity of LOPAC hits, TRE-FLAG-LMNB1 MEFs were treated in DOX containing medium for 48 h with ten-point, two-fold dose–response curves (in triplicate). Cells were stained for LMNB1, LMNA/C, nuclei counterstained with Hoechst 33342 and imaged on the Arrayscan VTI HCS reader. A. Dose–response in the primary HCS assay. Hits were analyzed for LMNB1 intensity (green), LMNA/C intensity (red), and cell numbers (Cell Survival, blue). Graphs show the averages ± SD from three independent biological repeats. B. Western blot confirmation. TRE-FLAG-LMNB1 MEF were treated for 48 h with the indicated concentrations of SB202190 and lysates immunoblotted with anti-LMNB1 and anti-beta actin antibodies. A representative blot is shown. Markers on the left indicate molecular weights in kDa. C. Quantification of compound response. Y-axes show LMNB1 / actin intensity ratios normalized to DMSO controls. Data show mean + SD of three independent biological repeats. P-values were computed by one-way ANOVA with Holm-Šídák's multiple comparisons test.
Collectively, the data document that the LMNB1 HCS assay can identify compounds that selectively and dose-dependently reduce LMNB1 in the TRE-LMNB1-MEF cell line. The data also suggest that the chemical space for compounds that dose-dependently reduce LMNB1 without cellular toxicity and confirm in orthogonal assays is small, and that low hit rates in HTS could be expected.
Primary HTS
Based on the results from the LOPAC pilot screen, we next screened 96,724 compounds from the NIH MLSMR collection in TRE-FLAG-LMNB1 MEF at a single concentration of 30 µM, anticipating an overall low confirmed hit rate. The screen consisted of 14 runs, with each run containing about twenty 384-well microplates (Fig. S2A,B). As in the LOPAC pilot library, assay performance was assessed by screening statistics of LDA and LMNB1 intensity22,23. As expected from assay development21 and the results from the LOPAC screen, LDA separated MAX (DOX-induced) and MIN (DOX withdrawal) controls with Z’ values of 0.55 ± 0.08, and LMNB1 with rZ’ values of 0.33 ± 0.13 (Fig. S2C,D). There was a good correlation between LDA and LMNB1 intensity (Fig. S2E). Two runs had Z’ values < 0.5 (LDA) and < 0.2 (LMNB1) and were eliminated from further analysis.
We then established a primary hit criterion based on plate statistics, namely a robust z-score of < −4 (i.e., plate median – 4 MAD), which amounted to a 72% reduction of LMNB1 levels (Fig. 3A). This resulted in a primary hit rate of 403 compounds (0.42%), which was reduced to 62 compounds (0.06% hit rate) after eliminating agents that also affected levels of LMNA/C or caused cell loss (Fig. 3B). Visual examination of archived scan images documented that the majority of hits that selectively reduced levels of LMNB1 without causing cell loss were scored as strong reducers of LMNB1 (Fig. 3B, red spheres). Forty-one compounds were commercially available and repurchased for dose–response confirmation studies.
Primary HTS. A. Scatter plot showing normalized LMNB1 intensities with runs 2 and 5 removed. Yellow; DMSO controls (MAX); red, doxycycline withdrawal controls (MIN), blue, library compounds. Horizontal lines show median ± 3MAD based on plate data, and a hit selection cutoff (median −4MAD, corresponding to a 72% reduction in LMNB1 (28% of control). B. Graphical representation of hit triage. 3D plot shows all compounds that passed the primary selection criterion (% LMNB1 of plate control < median – 4MAD; horizontal dashed line). Vertical dashed line indicates toxicity gate (90% cell survival). LMNA/C intensity is denoted by sphere size (smaller spheres indicate reduction of LMNA/C). Colored spheres show annotations from visual scoring of LMNB1 staining intensity (red, strong reduction; green, partial reduction).
HTS hit confirmation studies
The 41 commercially available compounds were obtained through MolPort (Table S1) and reconstituted in DMSO as described in Materials and Methods for dose–response evaluations. Nine compounds showed at least some activity, although six had very shallow curves (maximum reduction of LMNB1 ~ 30%; data not shown). One compound, MolPort-001–493–770 (Pubchem SID: 3716220, CID: 645580) showed very strong reduction of LMNB1, which was an artifact of image analysis (compound formed blue fluorescent precipitates). Two compounds, Molport-000–794–856 (Pubchem SID: 861659, CID: 662896) and MolPort-007–643-161 (Pubchem SID: 7967135, CID: 5308648) showed dose-dependent reductions in LMNB1 but not LMNA/C and lacked cellular toxicity. The latter two compounds were then obtained from multiple sources and/or vendors (Fig. S3, table) for re-analysis. All batches were analyzed for dose–response in the primary screen assay with little differences in potency (Fig. S3). Identity and purity of CID 662896 and CID 5308648 were ascertained by analytical data from vendors, and resynthesized batches of CID 662896 were re-confirmed in-house (Fig. S3B). Mean EC50 values for LMNB1 reduction were 8.6 ± 1.8 and 8.7 ± 2.6 µM, respectively, for CID 662896 and CID 5308648, with no changes in LMNA/C expression or cytotoxicity (Fig. 4A, B). These results were further confirmed by Western blots as an orthogonal assay not relying on fluorescence microscopy, although CID 5308648 was significantly less potent that CID 662896 (Fig. 4C-F).
Dose–response confirmation of CID 662896 and CID 5308648 in TRE-FLAG-LMNB1 MEF by HCS and Western blot. A. and B. Confirmation of compound activity and selectivity in the primary HTS assay. Each data point represents the average of the indicated number of independent repeats ± SD. EC50 values for LMNB1 were obtained by a four-parameter logistic regression fitted through pooled data from all experiments. CID 662896 is a diastereomeric mixture showing two peaks on a chiral column (not shown); its absolute configuration has not been unambiguously assigned yet. CID 5308648 is a racemate. C. Representative Western blot of TRE-FLAG-LMNB1 MEF treated with CID 662896. D. Quantification of CID 662896 response. E. Representative Western blots of TRE-FLAG-LMNB1 MEF treated with CID 5308648. Markers on the left indicate molecular weights in kDa. F. Quantification of CID 5308648 response. LMNB1 / actin intensity ratios were normalized to DMSO controls. Data in D. and F. show mean ± SD of the indicated number of independent biological repeats. P-values were computed by one-way ANOVA with Holm-Šídák's multiple comparisons test.
Quantification of LMNB1-induced nuclear abnormalities by high content texture analysis
LMNB1 overexpression results in characteristic nuclear abnormalities manifesting themselves as blebs, invaginations, and folding (Fig. 5A) cf. (Nmezi et al., 2020). These nuclear abnormalities are a direct functional consequence of LMNB1 overexpression and are observed in brains of ADLD patients. Measurements of nuclear abnormalities therefore present a biologically relevant functional readout for LMNB1 overexpression. We were able to quantify these nuclear abnormalities by high-content analysis of nuclear texture and morphology on a PerkinElmer Opera Phenix plus HCS reader with Harmony 5.1 using SER (spot, edge, ridge) based texture measurements, and a machine-learning algorithm to classify abnormal nuclei (Fig. 5B, red objects). CID 662896 and CID 5308648 inhibited LMNB1 expression and corrected associated nuclear abnormalities with essentially identical EC50 values for both parameters (Fig. 5C).
CID 662896 and CID 5308648 correct LMNB1-induced associated nuclear abnormalities. A. TRE-FLAG-LMNB1 MEF were exposed to doxycycline for three days, leading to elevated LMNB1 levels and characteristic nuclear abnormalities (+ DOX vs. -DOX) (cf. Nmezi, 2020). B. Images of DOX stimulated cells in the presence of vehicle (DMSO, + DOX), CID 662896, or CID 5308648 were acquired on an Opera Phenix Plus high-content reader using a 20X objective and analyzed for nuclear expression of LMNB1 and associated nuclear abnormalities by a machine-learning texture (SER texture) algorithm in Harmony 5.1. Left panels, original fluorescence micrographs; right panels, images with algorithm applied. Green nuclei, normal; red nuclei, abnormal). C. Quantification of nuclear abnormalities. Each data point is the average of four independent biological replicates ± SD. EC50 values were calculated for each replicate by a four-parameter logistic equation and averaged.
Activity of CID 662896 and CID 5308648 in fibroblasts from ADLD patients
We then tested the two prioritized compounds in ADLD patient cells by Western blot. Figure 6 shows that CID 662896 significantly and dose-dependently reduced levels of endogenous LMNB1 in ADLD patient fibroblasts. In contrast, CID 5308648 showed only a minor and non-significant reduction trend (Fig. 6).
Activity of prioritized hits in ADLD patient cells. Cultured fibroblasts from ADLD patients were treated for 48 h with compounds. Lysates were immunoblotted with anti-LMNB1 and anti-beta actin antibodies. A and B. Representative Western blot of ADLD cells treated with CID 662896 or CID 5308648, respectively. Markers on the left indicate molecular weights in kDa. C and D. Quantification of CID 662896 and CID 5308648 response. LMNB1 / actin intensity ratios were normalized to DMSO controls. Data in C. and D. show mean ± SD of the indicated numbers of independent biological repeats. P-values were computed by one-way ANOVA with Holm-Šídák's multiple comparisons test.
Physicochemical properties and in silico ADMET predictions
Both CID 662896 and CID 5308648 have favorable “drug-like” physicochemical properties (FAFDrugs4 analysis accessed December 2020; https://fafdrugs4.rpbs.univ-paris-diderot.fr/24) (Fig. S5 and data not shown). This includes no pan assay interference compounds (PAINS) or toxicity flags, no Lipinski violations, and no potential reactivity or promiscuity using the Eli-Lilly MedChem Rules toolkit that is used for open drug discovery projects25and the Pfizer 3/7526and GSK 4/40027physchem rules, which flag compounds for potential reactivity and liabilities in future optimization studies. Both compounds have good predicted oral absorption28and blood–brain-barrier (BBB) permeability based on total polar surface area (TPSA)29 (Fig. S4A). Both were predicted by a computational algorithm to cross the BBB (https://www.cbligand.org/BBB; last accessed October 2021) using OpenbabelFP2 fingerprints30 (Fig. S4B). Neither compound had any confirmed active flags in PubChem, ZINC, or STITCH databases, despite having been tested in > 800 biochemical and cellular assays (Databases last accessed September 2021). The only potential concern is that CID 5308648, a thioether, could be oxidized to the corresponding sulfoxide and sulfone in vivo.
Blood brain barrier permeability studies
Because ADLD is a CNS disorder, we next investigated whether CID 662896 and CID 5308648 could potentially be BBB permeable and performed a parallel artificial membrane permeability assay (PAMPA). PAMPA is widely used in the early phase of drug discovery to predict passive diffusion of drug molecules across phospholipid membranes as it is a cost-effective and robust method with good reproducibility and correlation with in vivo drug uptake31.
The assay was performed by Creative Bioarray, Inc. using a Millipore#MAIPN4550 transwell plate containing artificial membranes coated with brain lipid extract. After a four-hour incubation, the sandwich plate was separated and compound concentrations in donor and acceptor solutions were determined by LC/MS–MS. Positive and negative controls were propranolol (high permeability) and atenolol (low permeability). Samples were run in duplicates; tolbutamide served as internal standard. Compounds were quantified by LC–MS/MS analysis based on peak area as described in Materials and Methods. Both compounds crossed the artificial sandwich membrane, suggesting they would be CNS penetrant (Fig. S5).
In vivo brain penetration studies
Based on the results from the PAMPA assay, we performed a short pharmacokinetic study in mice to assess plasma levels and to investigate BBB permeability. We moved forward with CID 662896 because it consistently showed larger maximum reductions in LMNB1 levels in TRE-FLAG-LMNB1 MEF (Fig. 4), confirmed in ADLD patient cells (Fig. 6), and because it was available from a variety of vendors and sources. The compound was also resynthesized by two chemical suppliers (Vitascreen and Enamine, Fig. S4A) according to a published synthesis route32. Compound was administered i.p. and plasma samples taken at 0.5, 1, and 3 h. Samples were analyzed by LC–MS/MS as described in Materials and Methods. CID 662896 had peak plasma concentrations of 158, 95, and 27 µM at 0.5, 1, and 3 h after injection, respectively, and was detectable in brain tissue, with plasma-to-brain ratios of 0.108 – 0.138, documenting it had the ability to cross the BBB in mice (Fig. S6).A molecule is commonly deemed CNS penetrant if its brain-to-plasma concentration ratio (Cb:Cp) is > 0.04 using nonperfused brain tissue as cerebral blood volume approximates 4% of total brain volume33, although this is only an approximation, and free drug levels neeed to be determeined experimentally (for example by microdialysis). Nonetheless, a brain-to-plasma ratio of 0.1 is deemed moderately brain penetrant34.
Preliminary structure–activity relationship (SAR) studies based on CID 662,896
We performed a limited SAR study based on the CID 662896 structure. Specifically, we tested three analogs from the small molecule library of the University of Pittsburgh’s Center for Methodologies and Library Development (CMLD), as well as several commercially available analogs (Table S2). Most compounds were inactive, precluding detailed conclusions; however, some structural elements were shown to be essential. Notably, major structural modifications of the annulated triazole abolished activity (UPCMLDCBRC1040264, UPCMLDCBRC1049772). Removing the amine substituent on the triazole ring also completely depleted activity (CID 663139) and additionally caused solubility problems. Interestingly, also not tolerated was an oxidation of the tetrahydropyrimidine ring to a dihydropyrimidine, indicating that flattening of the heterocyclic scaffolds interferes with target binding (Table S2, UPCMLDCBRC1002139). These findings suggested that the 4H,5H,6H,7H-[1,2,4]triazolo[1,5-a]pyrimidine core was essential for activity in this scaffold. Furthermore, this core structure was quite sensitive to changes in the substitution patterns on the 5- and 7-positions. The o-chlorophenyl group in CID 662896 conveys a unique potency advantage, possibly by restricting the rotation of this aromatic group vs. the heterocyclic core through atropoisomeric interactions (compare with CID 3389508, CID 2948225 and CID 3154582). However, it is not sufficient to convey activity. Remarkably, CID 3154587 as well as CID 663139 are both inactive, in spite of the presence of the o-chlorophenyl substituent in these structures, and CID 3154585 showed a limited response at the highest concentration tested. Whereas, accordingly, most analogs tested were inactive, one (CID 3512343) retained some activity. In the latter analog, the o-chlorophenyl substituent is replaced with a simple phenyl ring, but a relatively bulky o-ethoxyphenyl substituent is present in the southern region of the compound, likely also causing restricted rotation. A definitive SAR analysis will require testing of a larger compound collection and a systematic modification of substituents on the 4H,5H,6H,7H-[1,2,4]triazolo[1,5-a]pyrimidine core structure.
Attempts to identify putative targets by computational methods failed due to a lack of activity data in public databases. CID 662896 did not have any reported bioactivities (Pubchem, (pubchem.ncbi.nlm.nih.gov) last accessed 9.25.2023, 892 bioassays) or STITCH (http://stitch.embl.de, last accessed 9.25.2023). We attempted a computational chemical similarity ensemble approach similar to35. Briefly, we obtained a list of analogs by running a live 2D similarity search on PubChem using the default Tanimoto threshold of 0.9. The search identified 402 compounds that are structurally similar to CID 662896. We queried the STITCH database (stitch.embl.de) for murine and human targets that are known with medium confidence (i.e., STITCH confidence score of at least 400 out of 1000) to be acted upon by these compounds. No such targets were identified.
Discussion
ADLD is a devastating, late onset demyelinating disease for which no treatments exist. Because the disease is caused by duplication of the LMNB1gene, resulting in overexpression of LMNB1 in the brain, we conducted an HTS/HCS of the LOPAC library for validation purposes, and ~ 97K compounds from the NIH MLSMR collection using an inducible cell culture model21 of LMNB1 overexpression in hopes of identifying non-toxic, selective reducers of LMNB1. This is, to our knowledge, the first large scale screen for non-toxic LMNB1 reducing compounds.
LOPAC pilot library screening
The LOPAC library is a collection of annotated bioactive compounds that is used to glean information about assay behavior under screening conditions, guidance for establising hit selection criteria, expected hit and confirmation rates, false positives, and susceptibility to assay interference. We purposefully set a loose criterion for activity and found that despite a high initial hit rate, only three compounds confirmed in dose–response. One of them (camptothecin) was an antineoplastic agent. Two compounds (SB202190 and SP600125) were further examined and one (SB202190) confirmed in Western blots. This suggested that in HTS a stringent cutoff for LMNB1 might miss potentially interesting compounds, and that the primary hit criterion for LMNB1 levels should not be set excessivly high. To avoid too large numbers of primary hits for secondary assays, we increased the cutoff for LMNB1 reduction only slightly and added additional gates for LMNA/C expression and apoptotic nuclear morphology to screen out non-selective and toxic agents. The results from the LOPAC further suggested low hit and confirmation rates in HTS.
Besides their utility as validation tools, the use of annotated bioactives presents an opportunity to uncover novel mechanisms or support prior hypotheses presented in the literature. The top hit from LOPAC, SB202190, is a selective p38 MAP kinase inhibitor. This was consistent with an earlier report that genetic ablation of p38 MAPK leads to reduced levels of LMNB1 protein19. Lamin levels decrease during apoptosis and senescence36. Consistent with this, the screen identified camptothecin, an antineoplastic agent that has been reported to reduce LMNB levels in HL-60 cells as a result of phosphorylation by PKC and subsequent apoptosis37. Another example for an antineoplastic agent that reduces LMNB1 levels was presented in a recent paper by Giorgio et al.38, who identified alvespimycin in a 717-compound screen for small molecules that reduce LMNB1. Alvespimycin is a geldanamycin analog that targets heat shock protein 90 (HSP90) and, like other geldanamycin analogs, is an anticancer agent. Surprisingly, Giorgio et al. found that alvespimycin reduced LMNB1 but did not report toxicity. We attempted to replicate their results in our cell model but observed significant cellular toxicity (data not shown). We speculate that this discrepancy may be due to our cell models being different. In the context of laminopathies such as ADLD, compounds that indirectly reduce LMNB1 as a result of cell death are not useful for therapeutic development as these diseases are chronic conditions that require life-long therapy in patients whose health is already compromised. Our screen therefore aimed at discovering non-toxic reducers of LMNB1 while eliminating non-specific toxicants.
Challenges and outcomes from the large scale screen
As expected from the pilot library screen, primary hit rates in HTS and confirmation rates in dose–response were low. After setting stringent gates for LMNA/C and cell survival, followed by visual examination of archived scan images, the number of primary hits was 62/97,000 (0.06% hit rate). Of these, only nine compounds showed any activity in dose–response (15%), and only two had full dose-curves with reasonable maximum reductions in LMNB1. These two compounds (CID 662986 and CID 5308648), however, corrected LMNB1-induced nuclear abnormalities, had drug-like physicochemical properties, crossed the BBB in PAMPA assays, and were subsequently validated in fibroblasts from ADLD patients. Both compounds reduced LMNB1 levels in the TRE-FLAG-LMNB1 MEFs; however, only CID 662896 significantly reduced levels of LMNB1 in ADLD patient fibroblasts (Fig. 6) and was prioritized for further validation studies. To exclude the possibility that the reduction of LMNB1 by CID 662896 was secondary to reduced proliferation or cellular toxicity, we evaluated growth characteristics of ADLD patient fibroblasts. When cells were plated at low density and followed over time, CID 662896 did not affect cell proliferation (Fig. S7A). Morphology measurements after a two-day exposure to test agents (as in the primary assay) did not show any effect of CID 662896 on cell numbers, nuclear morphology, or cell size (Fig. S7B). In contrast, known cytotoxic LMNB1 reducing agents camptothecin and etoposide reduced cell numbers and caused cells to enlarge, indicating cellular toxicity (Fig. S7B). There have been contrasting reports regarding the role of lamin B1 in senescence and cell proliferation and it is unclear if a reduction of lamin B1 actually drives senescence or whether this reduction is a secondary consequence36,39,40. Collectively, our results document that the loss of LMNB1 by CID 662896 is not secondary to growth inhibition or cytotoxicity.
In the ADLD patient fibroblasts, CID 662,896 also affected levels of LMNB2 and LMNA/C, although their loss was less pronounced than that of LMNB1 and highly variable (Fig. S8). Further studies, using an alternative, more sensitive methodology than Western blots (e.g., targeted mass spectroscopy) will be required to definitively answer the question of selectivity. CID 662896 was subsequently resynthesized on a gram scale and shown to cross the BBB in mice. Therefore the screen, despite its low hit rates, yielded at least one high-quality agent that is worthy of further study in more disease relevant cellular models as well as mouse models of ADLD.
Utility and limitations of LDA in HTS
An open question in the drug discovery community is the utility of dimensionality reduction methods in HTS. We used LDA in assay development because of prior reports41and our own experience with complex phenotypical assays42. LDA robustly tests the ability of cells to respond to reduction of LMNB1 and correction of nuclear abnormalities21. It is thus a suitable means to assess the limits and quality of the assay with regards to positive and negative controls, as is customary for HTS43. Whereas for controls, LDA and lamin B1 were highly correlated (r-square > 0.9; Fig. S1C and S2E), LMNB1 intensity as the primary, biologically relevant parameter separated controls with a much lower level of statistical significance (Fig. S1A,B and S2A-D). With compounds, LDA and lamin B1 levels were still positively correlated, but with much lower statistical significance (r-square 0.238, Fig. S9). This is perhaps not surprising as LDA “forces” separation of defined classes into the best dimension. While this procedure separates known classes very well, it can also result in unknown samples being less accurately classified. We therefore investigated the value of LDA in hit selection by comparing LMNB1 intensities to LDA, visual confirmation, and ultimately, dose–response.
Whereas LDA and lamin B1 levels were largely correlated with unknowns, many of the visually confirmed compounds (Fig. S9, red markers) had high LDA values, indicating that the use of LDA as a primary hit selection criterion could lead to a large number of false negatives. (Fig. S9, upper left quadrant). We therefore asked if LDA could be useful as a secondary triage tool. Among hits that passed the initial LMNB1 intensity gate and were visually confirmed, LDA only flagged a small number of hits as positive (Fig. S9, lower left quadrant). This could mean that while LDA might potentially miss a number of potentially interesting compounds, it could also reduce the number of hits that do not confirm in dose–response. In support of the latter, we found that only a small number of visually apparent hits repeated in dose–response. This was consistent with the results from the LOPAC pilot library, which had suggested that dose–response confirmation rates would likely be very low. Importantly, the two compounds with unambiguous dose–response curves (CID 662896 and CID 5308648), were correctly flagged by LDA (Fig. S9, lower left quadrant). Therefore, using LDA as an additional hit selection criterion would have dramatically reduced the number of hits that despite being scored as visually confirmed did not confirm in dose–response. These positive results notwithstanding, using LDA as a hit selection tool appears to be risky, mainly due to the potential of false negatives, its aforementioned propensity to misclassify unknowns. Our conclusion at this time is that LDA is a powerful tool to define the limits and HTS performance of a complex phenotypic assay, but that its utility as a hit selection tool is questionable. More examples of using LDA in HTS hit selection are needed to answer this question, and other methods of dimensionality reduction should be explored.
Future directions
Whereas CID 662896 and CID 5308648 are potential tool compounds, further study is needed to elucidate their mechanism(s) of action. The regulation of lamin levels is complex and still not fully understood. Therefore, tools to uncover mechanisms of LMNB1 regulation would enable generation of novel testable hypotheses, and unveil possible targets for intervention. At the present time, however, we have no information about possible targets for our lead compound CID 662896, and target identification studies will be needed to fully exploit it as a chemical probe in LMNB1 biology.
In summary, we have identified a small molecule that reduces levels of LMNB1 without gross toxicity, rigorously validated its activity in a mouse and human cell model of ADLD, established a preliminary SAR profile based on an assay of selected analogs, and shown that it has the ability to cross the BBB in mice. This compound is now positioned for proof-of-principle pharmacodynamic (PD) studies in a mouse model of ADLD we have previously developed44,45. If successful, this would be the first study to show that it is possible to reduce levels of LMNB1 in the brain with small molecules, which would represent a milestone in the possible development of LMNB1-reducing compounds into potential experimental therapeutics.
Materials and methods
Reagents
Primary antibodies for HTS: Lamin B1 (ab16048) and Lamin B2 (151735) were from abcam (Cambridge, United Kingdom) and Lamin A/C (47775) was from Cell Signaling Technologies (Danvers, MA). Secondary antibodies: Cy3 AffiniPure Donkey Anti- Rabbit IgG (H + L) (711–165-152) and Cy5 AffiniPure Donkey Anti- Mouse IgG (H + L) (715–175-150) were both from Jackson ImmunoResearch Laboratories Inc. (West Grove, PA). Nuclear stain Hoechst 33,342, Trihydrochloride, Trihydrate (H1399) and Alexa 555-conjugated phalloidin (A34055) were from Invitrogen (Waltham, MA). Doxycycline hyclate (DOX, D-9891), Formaldehyde 37% (252549), Bovine Serum Albumin lyophilized powder (BSA, A2153), and Triton X-100 (T8787) were all from Millipore Sigma (Burlington, MA). Assay plates 384-well PS Black Collagen Type 1 CellCoat from Greiner Bio-One (Monroe, NC) or PhenoPlate 384-well collagen-coated (6057700) from Revvity (Waltham, MA). Automation Tips for PerkinElmer Janus MDT (PK-384-R) were from Axygen (Union City, CA), Bravo tips (AGI-30.NS) from Fluotics (New York, NY), and Agilent BioTek MultiFlo FX/EL406 Dispenser Cassettes (7170011) from Agilent BioTek (Winooski VT).
Cell culture
Primary mouse embryonic fibroblasts (MEFs) generated from TRE-FLAG-LMNB1 (TRE-FLAG-LMNB1 MEF); Rosa-rtTAmice21 were cultured in high glucose Dulbecco’s modified Eagle’s medium (DMEM; 11965126 Gibco Fisher, Waltham, MA) with 10% fetal bovine serum (FBS; MT35010CV Corning, Corning, NY), 1% penicillin–streptomycin (15–140-122 Cytivia Hyclone, Logan, UT), and 2 mM L-glutamine (25–030–081 Cytivia Hyclone, Logan, UT) in a humidified 37 °C, 5% CO2 incubator.
Primary skin fibroblasts from autosomal dominant leukodystrophy (ADLD) patients were isolated as described12. ADLD patient cells were cultured with the same growth medium and incubator conditions as described above for the TRE-FLAG-LMNB1 MEF.
Library of Pharmacologically Active Compounds (LOPAC) Pilot Library Screen
The LOPAC library, a collection of 1280 biologically annotated compounds, was obtained from Sigma-Aldrich (cat # LO4200) and maintained at the UPDDI in 384 well assay ready plates (2 µl of 10 mM stocks in DMSO) under temperature and humidity controlled conditions, as recommended46,47. For the pilot library screen, TRE-FLAG-LMNB1 MEF were placed into medium containing 2 µg/mL DOX for three days to induce Lamin B1 expression. On the day of experiment, cells were trypsinized and plated (4,000 cells/ 30 µl) into thin bottom 384-well collagen-coated plates (Greiner Bio-One). Cells in columns 2—23 were plated in medium + DOX and cells in columns 1 and 24 were plated in medium without DOX (DOX withdrawal control). Plates were incubated overnight in a humidified 37 °C, 5% CO2 incubator. The LOPAC library was tested at three concentrations (30 µM, 10 µM, and 3 µM). Assay ready LOPAC daughter plates were thawed and reconstituted in 45 µl medium + DOX (Microflo, BioTek), creating an intermediate dilution plate of 426 µM at 4.3% DMSO. From the intermediate dilution plate, three treatment plates at 3X the final assay concentration (90 µM, 30 µM, and 9 µM/3% DMSO) were prepared with the Janus MDT (PerkinElmer). To each treatment plate positive controls (3% DMSO + 2 µg/mL DOX) were added to columns 2 and 23 and negative controls (3% DMSO, no DOX) were added to columns 1 and 24 (n = 32 each) with an electronic multichannel pipettor (ThermoFisher Matrix). Plates were centrifuged at 50 × g for one minute, and 15 µl transferred to assay plates with a Janus MDT for final concentrations of 30 µM, 10 µM, and 3 µM (1% DMSO) in duplicate. After forty-eight hours at 37 °C, cells were fixed with 15 µl of 16% formaldehyde for 15 min at room temperature. The fixative solution was aspirated with an ELx405 plate washer (BioTek) and plates washed three times with 30 µl PBS. Cells were permeabilized with 15 µl blocking buffer (0.3% Triton X-100 and 1% BSA in PBS, 0.2 µm filtered) for one hour at RT, followed by addition of 15 µl primary antibody solution containing lamin B1 (1:2000, rabbit) and lamin A/C (1:1000, mouse) antibodies in blocking buffer. Plates were incubated in primary antibody solution at 4 °C overnight, washed three times with PBS, and stained with 15 µl secondary antibodies (Cy3 donkey anti-rabbit (1:500, Lamin B1), Cy5 donkey anti-mouse (1:500, Lamin A/C), and Hoechst 33,342 (10 µg/ml) in blocking solution for one hour, washed three times with PBS, sealed, and stored at 4 °C until imaging.
Imaging and analysis
Plates were scanned on an Arrayscan VTI (Cellomics, Thermo Fisher Scientific, Waltham, MA) using a 20X 0.75NA objective and a Semrock BGRFR (Blue-Green–Red-Far Red) multiband filter set. High content analysis of LMNB1 expression, texture, nuclear shape morphology, and linear discriminant analysis (LDA) were performed exactly as described previously21, except that the LDA parameter set was reduced from 27 to eight (QE_AvgIntenCh2, KS_Norm_AvgIntenCh2, Percent_Outlier_AvgIntenCh2, MEAN_ObjectShapeP2ACh1, MEAN_AvgIntenCh2, MEAN_TotalIntenCh2, MEAN_VarIntenCh2, and MEAN_ObjectShapeLWRCh1) to avoid overfitting.
Statistical analysis
HTS data were normalized to MIN (DOX withdrawal) and MAX (+ DOX) controls. Unless otherwise noted, data were analyzed by one-way ANOVA with Holm-Šídák's multiple comparisons test in Graph Pad Prism (Ver 10). p-values < 0.05 were considered significant.
LOPAC dose–response confirmation
Selected hits from the LOPAC pilot library screen were repurchased as powders and reconstituted to 10 mM DMSO stock solutions. LOPAC master dose–response plates containing 10-point dose–response curves (in triplicate) were created by serially diluting 25 µl 10 mM stocks into 25 µl DMSO 1:2 with an Agilent Bravo liquid handler. Multiple sets of daughter plates were stamped out by transferring 2 µl stock solution to fresh microplates. Plates were sealed and stored at −20 °C. For the dose–response confirmation assay, TRE-FLAG-LMNB1 MEF were plated (4,000 cells/30 µl) into thin bottom 384-well collagen-coated plates as described above for the LOPAC screen, and allowed to adhere overnight in a humidified 37 °C, 5% CO2 incubator. The next day, compound daughter plates were thawed and reconstituted in 65 µl medium + DOX creating a 3X treatment plate (300 µM/3% DMSO). Positive and negative controls were added to the treatment plates as described above for the LOPAC library (n = 32 each), for a final concentration range of 100 µM – 200 nM (1% DMSO). Incubation, processing steps, fixation, antibody staining, imaging, and analysis were the same as the LOPAC screen.
Primary HTS
Using the LOPAC pilot library screen as a guide, a 96,724 compound subset of the NIH Molecular Libraries Small Molecule Repository (MLSMR) compound library48was screened at a single concentration of 30 µM. The MLSMR collection was established through the NIH Molecular Libraries and Imaging (MLI) Roadmap and a copy of the library is maintained by the UPDDI as assay-ready daughter plates containing 2 µl 10 mM DMSO stocks in columns 3–22 under temperature and humidity controlled conditions as recommended46,47. Compounds in daughter plates were reconstituted in 65 µl medium + DOX, diluted to 90 µM/3% DMSO and controls added as above. Treatment, staining, imaging, and analysis were the same as described above for the LOPAC screen.
HTS dose–response confirmation
Hits from the primary HTS were repurchased as powder and reconstituted as 10 mM DMSO stock solutions. HTS master dose–response plates were prepared as half-log 10-point dose–response curves (in triplicate) by serially diluting 25 µl 10 mM stocks into 50 µl DMSO. Daughter plates were stamped out by transferring 2 µl to new microplates, sealed, and stored at −20 °C until use. The HTS dose–response confirmation assay was performed as described above for the LOPAC dose–response confirmation. Imaging and analysis was performed on the ArrayScan VTI as described above, or on an Opera Phenix Plus high content reader (Revvity, Waltham, MA) with a 20 × air objective and laser lines and emission filters for Hoechst (ex405/em435-480 nm), Cy3 (ex561/em570-630 nm), and Cy5 (ex640/em650-760 nm) fluorophores. Analysis of nuclear abnormalities was performed on the Opera Phenix Plus images in Harmony 5.1 using a machine learning algorithm (Phenologic™). This algorithm permits automatic detection of two classes of objects by a linear classifier. The algorithm is trained by a human observer on positive (+ DOX) and negative (DOX withdrawal) samples. Briefly, images were segmented and analyzed for intensity and texture based on LMNB1 staining. The SER (saddle, edge, ridge, valley, dark, and bright) texture algorithm in Harmony applies texture filters and generates numerical outputs for each. We purposefully used an unnormalized version, which also takes into account intensities and is useful when intensity measurements also contribute to the classification (as is the case here with LMNB1). This data was then used to build a linear classifier. The final output is the percentage of abnormal cells.
General LC–MS separation/analysis conditions
Identity and purity of repurchased compounds mostly relied on analytical data (LC–MS/NMR) supplied by vendors. For resynthesized CID 662896, purity was assessed in-house using an Agilent Technologies 1260 Infinity II LC at 220 nm UV absorption (Waters XBridge BEH C18 2.1 × 50 mm, 2.5 µm). A 10 min gradient elution with a MeCN/H2O/MeOH mobile phase containing 0.1% formic acid at a flow rate of 500 μL/min was used with an eluent ratio from 3:92:5 at 0–0.5 min to 93:2:5 at 4.0 min, back to 3:92:5 from 6.0 to 7.5 min. Samples of CID 662896 were analyzed with total ion count (TIC, positive ion mode) and UV (220 nm, 254 nm) detection, and confirmed the identity and purity of the compound.
Western blotting
Total protein was extracted from cells using T-PER buffer (Thermo Scientific) with a protease inhibitor cocktail (Roche). Protein was extracted from three independent biological replicates for each condition. 50 µg protein were loaded per well and separated by polyacrylamide gel electrophoresis. Gels were transferred to nitrocellulose membranes followed by blocking with Odyssey Blocking Buffer (Licor) in TBS for 1 h at room temperature. Membranes were incubated with specific primary antibodies overnight at 4 °C. Primary antibodies were diluted in 0.2% Tween 20/Licor TBS buffer. Secondary antibodies were incubated for 1 h at room temperature. Blots were imaged using an Odyssey CLx Imaging System (Licor). Primary antibodies used were anti-Lamin B1 (Abcam—ab16048), anti-Lamin B2 (abcam 151735), anti-Lamin A/C (Cell Signaling 47775), anti-beta-Actin (Thermo Fisher Scientific – MA5-15739) and anti-GAPDH (Thermo Fisher Scientific – MA5-15738). Secondary antibodies used were Anti-Rabbit IRDye 800CW (LI-COR Biosciences—926–32213) and anti-Mouse IRDye 680LT (LI-COR Biosciences—926–68020).
Parallel Artificial Membrane Permeability Assay (PAMPA)
The assay was performed by Creative BioArray, Inc. (Shirley, NY). In brief, the assay was conducted in a transwell plate (Millipore #MAIPN4550) separated by an artificial membrane (Poly(vinylidene fluoride)) coated with 1% lecithin/dodecane mixture. Donor solution containing compound was added to the donor plate and PBS was added to the acceptor plate. The plates were sandwiched together and incubated four hours. Following incubation, the plates were separated and compound concentration was quantified by LC–MS/MS analysis on peak area for the donor and acceptor solutions. Each sample was run in duplicate with propranolol and atenolol used as high permeability and low permeability controls respectively. Tolbutamide served as an internal standard. Permeability rates (Pe), which are a quantitative measurement of drug permeation, are given in nm/sec and were calculated as:
where:
[drug]equilibrium = ([drug]donor x VD + [drug]acceptor x VA) / (VD + VA).
VD = 0.15 mL; VA = 0.30 mL; Area = 0.28cm2; time = 14400 s.
[drug]acceptor = (Aa/Ai x DF)acceptor; [drug]donor = (Aa/Ai x DF)donor.
Aa/Ai: Peak area ratio of analyte and internal standard; DF: Dilution factor.
In vivo pharmacokinetics (PK) and brain penetration study
Mouse PK studies were performed by Touchstone Biosciences (Plymouth Meeting, PA), following the National Research Council guide for the care and use of laboratory animals to ensure proper care, use, and human treatment of the animals used in the study, adhering to generally accepted procedures for animal housing, dosing and blood sampling, and following AVMA guidelines for euthanasia. All procedures were approved by the Touchstone Biosciences Animal Research Review Panel. Results are reported in accordance with ARRIVE guidelines (https://arriveguidelines.org).
Male CD‐1 mice were fed a standard laboratory rodent diet and housed in individual cages on a 12-h light and 12-h dark cycle with room temperature maintained at 22 – 30 °C and relative humidity at 50 −20%. Animals are typically fasted overnight before dosing, with food returned after the last blood samples are obtained. Water was provided ad libitum throughout the study. The dosing solution of the test compound was prepared in 10%DMSO:40%PEG300:50%H2O. Animals (three mice per time point) were dosed via injection in the lower right quadrant of the abdomen for intraperitoneal administration at 20 mg/kg (20 ml/kg). All blood samples (30–50 μl per sample) were taken via appropriate vein (saphenous or submandibular vein) at 0.5, 1 and 3 h after dosing. Blood samples were collected in Greiner MiniCollect K2EDTA tubes, placed on ice, and within 30 min, centrifuged at 15,000 × g for 5 min to obtain plasma samples. All plasma samples were stored at –70°C until analysis. The whole brain was harvested, immediately, briefly rinsed in water, and blotted dry with a paper towel. The whole brain was weighed and three volumes of PBS (pH 7.4) added to one volume of each tissue sample, which was then homogenized by a tissue homogenizer until fine tissue particles were completely dispersed or emulsified. The tissue homogenate samples were stored at –70 °C until analysis.
Bioanalysis of PK samples
Plasma samples were prepared as follows. Three volumes of acetonitrile containing internal standard was added to one volume of plasma to precipitate proteins. Samples were centrifuged (3,000 × g for 10 min) and supernatant removed for analysis by LC–MS/MS. Calibration standards and quality controls were made by preparation of a 1 mg/mL stock solution and subsequently a series of working solutions in methanol : water (1:1, v/v) which were spiked into blank plasma to yield a series of calibration standard samples. All PK plasma samples were treated identically to the calibration standards. LC–MS/MS analysis was performed utilizing multiple reaction monitoring for detection of characteristic ions for each drug candidate, additional related analytes and internal standards. Tissue samples were prepared as follows. Three volumes of PBS (pH 7.4) was added to one volume of each tissue sample which was then homogenized to obtain each tissue homogenate sample. Subsequently, three volumes of acetonitrile containing internal standard was added to one volume of each tissue homogenate, the mixture vortexed and centrifuged (3,000 × g for 10 min), and supernatant removed for analysis by LC–MS/MS. Calibration standards were made by preparation of a 1 mg/ml stock solution and subsequently a series of working solutions in methanol : water (1:1, v/v) which were spiked into blank tissue homogenate to yield a series of calibration standard samples in the range of 1 ng/ml to 10 μg/ml. All PK tissue samples were treated identically to the calibration standards. LC–MS/MS analysis was performed utilizing multiple reaction monitoring for detection of characteristic ions for each drug candidate, additional related analytes and internal standard. Quantification was by Total Ion Chromatogam (TIC) after establishment of assay linearity range and lower limit of detection based on calibration standards.
Antiproliferative activity and expanded toxicity measurements
For assessment of antiproliferative activity, ADLD patient fibroblasts were plated at low density (400 cells/well) in 384 well plates, treated with vehicle (DMSO) or CID 662896 (100 µM) and growth monitored every day on the OPERA Phenix Plus using digital phase contrast live imaging. Cell numbers were enumerated by automated image analysis in Harmony 5.1. For expanded toxicity measurements, cells were plated at high density (4,000 cells/well) and treated with various concentrations of CID 662896, camptothecin, or etoposide for two days. Cells were stained with Hoechst 33342 and Alexa 555-conjugated phalloidin overnight, and imaged on the OPERA Phenix with a 20 × air objective and laser lines and emission filters for Hoechst (ex405/em435-480 nm) and Cy3 (ex561/em570-630 nm) channels. Cell numbers, nucleus size, and cell size were quantified in Harmony 5.1. Each experiment was performed three times in quadruplicate. Data are presented as the averages of the three biological repeats ± S.D.
Data availability
The data presented in this study are available upon request from the corresponding authors.
Abbreviations
- ADLD:
-
Autosomal Dominant Leukodystrophy
- BBB:
-
Blood–brain-barrier
- BSA:
-
Bovine serum albumin
- CMLD:
-
Center for Methodologies and Library Development
- Cy3:
-
Indocarbocyanine with three-carbon linker
- Cy5:
-
Indocarbocyanine with five-carbon linker
- DMSO:
-
Dimethyl sulfoxide
- DOX:
-
Doxycycline
- HCS:
-
High-content screening
- LC–MS:
-
Liquid chromatography/mass spectrometry
- LDA:
-
Linear discriminant analysis
- MAPK:
-
Mitogen activated protein kinase
- MEF:
-
Mouse embryonic fibroblasts
- PD:
-
Pharmacodynamics
- PK:
-
Pharmacokinetics
- RT:
-
Room temperature
- TRE:
-
Tetracycline responsive element
References
Wong, X., Melendez-Perez, A. J. & Reddy, K. L. The nuclear lamina. Cold Spring Harb. Perspect. Biol. https://doi.org/10.1101/cshperspect.a040113 (2022).
Stancheva, I. & Schirmer, E. C. Nuclear envelope: connecting structural genome organization to regulation of gene expression. Adv. Exp. Med. Biol. 773, 209–244. https://doi.org/10.1007/978-1-4899-8032-8_10 (2014).
Butin-Israeli, V., Adam, S. A., Goldman, A. E. & Goldman, R. D. Nuclear lamin functions and disease. Trends Genet. 28, 464–471. https://doi.org/10.1016/j.tig.2012.06.001 (2012).
Shin, J. Y. & Worman, H. J. Molecular Pathology of Laminopathies. Annu. Rev. Pathol. 17, 159–180. https://doi.org/10.1146/annurev-pathol-042220-034240 (2022).
Worman, H. J., Fong, L. G., Muchir, A. & Young, S. G. Laminopathies and the long strange trip from basic cell biology to therapy. J. clin. Invest. 119(7), 1825–1836 (2009).
Nmezi, B. et al. Concentric organization of A- and B-type lamins predicts their distinct roles in the spatial organization and stability of the nuclear lamina. Proc. Natl. Acad. Sci. U S A 116, 4307–4315. https://doi.org/10.1073/pnas.1810070116 (2019).
Worman, H. J. & Schirmer, E. C. Nuclear membrane diversity: underlying tissue-specific pathologies in disease?. Curr. Opin. Cell Biol. 34, 101–112. https://doi.org/10.1016/j.ceb.2015.06.003 (2015).
Evangelisti, C. et al. The wide and growing range of lamin B-related diseases: from laminopathies to cancer. Cell. Mol. Life Sci. 79, 126. https://doi.org/10.1007/s00018-021-04084-2 (2022).
Padiath, Q. S. & Fu, Y. H. Autosomal dominant leukodystrophy caused by lamin B1 duplications a clinical and molecular case study of altered nuclear function and disease. Methods Cell Biol. 98, 337–357. https://doi.org/10.1016/S0091-679X(10)98014-X (2010).
Padiath, Q. S. Lamin B1 mediated demyelination: Linking lamins. Lipids & Leukodystrophies. Nucl. 7, 547–553. https://doi.org/10.1080/19491034.2016.1260799 (2016).
Nmezi, B. et al. Genomic deletions upstream of lamin B1 lead to atypical autosomal dominant leukodystrophy. Neurol. Genet. 5, e305. https://doi.org/10.1212/NXG.0000000000000305 (2019).
Giorgio, E. et al. Analysis of LMNB1 duplications in autosomal dominant leukodystrophy provides insights into duplication mechanisms and allele-specific expression. Hum. Mutat. 34, 1160–1171. https://doi.org/10.1002/humu.22348 (2013).
Cristofoli, F. et al. De novo variants in LMNB1 cause pronounced syndromic microcephaly and disruption of nuclear envelope integrity. Am. J. Hum. Genet. 107, 753–762. https://doi.org/10.1016/j.ajhg.2020.08.015 (2020).
Li, L. et al. Lamin B1 is a novel therapeutic target of betulinic acid in pancreatic cancer. Clin. Cancer Res. 19, 4651–4661. https://doi.org/10.1158/1078-0432.CCR-12-3630 (2013).
Radspieler, M. M. et al. Lamin-B1 is a senescence-associated biomarker in clear-cell renal cell carcinoma. Oncol. Lett. 18, 2654–2660. https://doi.org/10.3892/ol.2019.10593 (2019).
Sun, S., Xu, M. Z., Poon, R. T., Day, P. J. & Luk, J. M. Circulating Lamin B1 (LMNB1) biomarker detects early stages of liver cancer in patients. J. Proteome. Res. 9, 70–78. https://doi.org/10.1021/pr9002118 (2010).
Wazir, U. et al. The clinicopathological significance of lamin A/C, lamin B1 and lamin B receptor mRNA expression in human breast cancer. Cell. Mol. Biol. Lett. 18, 595–611. https://doi.org/10.2478/s11658-013-0109-9 (2013).
Yu, Z. Y. et al. Lamin B1 deficiency promotes malignancy and predicts poor prognosis in gastric cancer. Neoplasma 67, 1303–1313. https://doi.org/10.4149/neo_2020_200109N33 (2020).
Barascu, A. et al. Oxidative stress induces an ATM-independent senescence pathway through p38 MAPK-mediated lamin B1 accumulation. EMBO J. 31, 1080–1094. https://doi.org/10.1038/emboj.2011.492 (2012).
Donadille, B. et al. Partial lipodystrophy with severe insulin resistance and adult progeria Werner syndrome. Orphanet. J. Rare Dis. 8, 106. https://doi.org/10.1186/1750-1172-8-106 (2013).
Nmezi, B. et al. Development and optimization of a high-content analysis platform to identify suppressors of lamin B1 overexpression as a therapeutic strategy for autosomal dominant leukodystrophy. SLAS Discov. 25, 939–949. https://doi.org/10.1177/2472555220915821 (2020).
Zhang, J. H., Chung, T. D. Y. & Oldenburg, K. R. A simple statistical parameter for use in evaluation and validation of high throughput screening assays. J. Biomol. Screen. 4(2), 67–73 (1999).
Haelewyn, J., Iversen, P. W. & Weidner, J. R. Addressing unusual assay variability with robust statistics. SLAS Discov. 26, 1291–1297. https://doi.org/10.1177/24725552211038379 (2021).
Lagorce, D., Sperandio, O., Galons, H., Miteva, M. A. & Villoutreix, B. O. FAF-Drugs2: free ADME/tox filtering tool to assist drug discovery and chemical biology projects. BMC Bioinform. 9, 396. https://doi.org/10.1186/1471-2105-9-396 (2008).
Bruns, R. F. & Watson, I. A. Rules for identifying potentially reactive or promiscuous compounds. J. Med. Chem. 55, 9763–9772. https://doi.org/10.1021/jm301008n (2012).
Hughes, J. D. et al. Physiochemical drug properties associated with in vivo toxicological outcomes. Bioorg Med Chem Lett 18, 4872–4875. https://doi.org/10.1016/j.bmcl.2008.07.071 (2008).
Gleeson, M. P. Generation of a set of simple, interpretable ADMET rules of thumb. J Med Chem 51, 817–834. https://doi.org/10.1021/jm701122q (2008).
Veber, D. F. et al. Molecular properties that influence the oral bioavailability of drug candidates. J Med Chem 45, 2615–2623. https://doi.org/10.1021/jm020017n (2002).
Van de Waterbeemd, H. In silico models to predict oral absorption. In comprehensive medicinal chemistry II (eds John B. Taylor David J. Triggle) 669–697 (Elsevier, 2007).
Liu, H. et al. AlzPlatform: an Alzheimer’s disease domain-specific chemogenomics knowledgebase for polypharmacology and target identification research. J. Chem. Inf. Model 54, 1050–1060. https://doi.org/10.1021/ci500004h (2014).
Müller, J., Esso, K., Dargo, G., Konczol, A. & Balogh, G. T. Tuning the predictive capacity of the PAMPA-BBB model. Eur. J. Pharm. Sci. 79, 53–60. https://doi.org/10.1016/j.ejps.2015.08.019 (2015).
Pyatakov, D. A. et al. Diversity oriented synthesis of polycyclic heterocycles through the condensation of 2-Amino[1,2,4]triazolo[1,5-a]pyrimidines with 1,3-diketones. J. Org. Chem. 80, 10694–10709. https://doi.org/10.1021/acs.joc.5b01908 (2015).
Shaffer, C. L.Chapter 4 - Defining neuropharmacokinetic parameters in CNS drug discovery to determine cross-species pharmacologic exposure–response relationships. In annual reports in medicinal chemistry 45 (ed John E. Macor) 55–70 (Academic Press, 2010).
Reichel, A. The role of blood-brain barrier studies in the pharmaceutical industry. Curr. Drug Metab. 7, 183–203. https://doi.org/10.2174/138920006775541525 (2006).
Wang, Z., Liang, L., Yin, Z. & Lin, J. Improving chemical similarity ensemble approach in target prediction. J. Cheminform. 8, 20. https://doi.org/10.1186/s13321-016-0130-x (2016).
Shimi, T. et al. The role of nuclear lamin B1 in cell proliferation and senescence. Genes Dev. 25, 2579–2593. https://doi.org/10.1101/gad.179515.111 (2011).
Shimizu, T., Cao, C. X., Shao, R. G. & Pommier, Y. Lamin B phosphorylation by protein kinase C alpha and proteolysis during apoptosis in human leukemia HL60 cells. J. Biol. Chem. 273, 8669–8674. https://doi.org/10.1074/jbc.273.15.8669 (1998).
Giorgio, E. et al. A high-content drug screening strategy to identify protein level modulators for genetic diseases: A proof-of-principle in autosomal dominant leukodystrophy. Hum. Mutat. 42, 102–116. https://doi.org/10.1002/humu.24147 (2021).
Dreesen, O. et al. Lamin B1 fluctuations have differential effects on cellular proliferation and senescence. J. Cell. Biol. 200, 605–617. https://doi.org/10.1083/jcb.201206121 (2013).
Yang, S. H. et al. An absence of both lamin B1 and lamin B2 in keratinocytes has no effect on cell proliferation or the development of skin and hair. Hum. Mol. Genet. 20, 3537–3544. https://doi.org/10.1093/hmg/ddr266 (2011).
Kümmel, A. et al. Integration of multiple readouts into the z’ factor for assay quality assessment. J. Biomol. Screen. 15(1), 95–101. https://doi.org/10.1177/1087057109351311 (2010).
Shun, T., Gough, A. H., Sanker, S., Hukriede, N. A. & Vogt, A. Exploiting analysis of heterogeneity to increase the information content extracted from fluorescence micrographs of transgenic zebrafish embryos. Assay Drug Dev. Technol. 15, 257–266. https://doi.org/10.1089/adt.2017.793 (2017).
Malo, N., Hanley, J. A., Cerquozzi, S., Pelletier, J. & Nadon, R. Statistical practice in high-throughput screening data analysis. Nat. Biotechnol. 24, 167–175 (2006).
Rolyan, H. et al. Defects of lipid synthesis are linked to the age-dependent demyelination caused by Lamin B1 overexpression. J. Neurosci. 35, 12002–12017. https://doi.org/10.1523/JNEUROSCI.1668-15.2015 (2015).
Heng, M. Y. et al. Lamin B1 mediates cell-autonomous neuropathology in a leukodystrophy mouse model. J. Clin. Invest. 123, 2719–2729. https://doi.org/10.1172/JCI66737 (2013).
Lane, S. J. et al. Defining and maintaining a high quality screening collection: the GSK experience. Drug Discov. Today 11, 267–272. https://doi.org/10.1016/S1359-6446(05)03730-X (2006).
Blaxill, Z., Holland-Crimmin, S. & Lifely, R. Stability through the ages: the GSK experience. J. Biomol. Screen. 14(5), 547–556. https://doi.org/10.1177/1087057109335327 (2009).
Austin, C. P., Brady, L. S., Insel, T. R. & Collins, F. S. NIH molecular libraries initiative. Science 306, 1138–1139. https://doi.org/10.1126/science.1105511 (2004).
Acknowledgements
We thank Harold Takyi, Taber S. Maskrey, and Desirae L. Crocker for technical assistance, and Stacey Sukoff Rizzo at the University of the Pittsburgh MODEL-AD preclinical testing core for helpful discussions. This work was supported by NIH grants R33NS106087 to AV/QSP and R01NS095884, R01NS126193, 1R21NS131906 and R21NS104384 to QSP. The project used shared instrumentation that was acquired with National Institutes of Health grant S10OD028450.
Funding
Namytional Institutes of Health,R33NS106087,R01NS095884,R01NS126193,R21NS131906,R21NS104384
Author information
Authors and Affiliations
Contributions
AV and QSP conceived the study; LLV, FL, BN, GRB, RL, and AV performed experiments; LLV, TOS, AG, TL, AV, and QSP analyzed and interpreted data; PW and TL provided scientific input and guidance; LLV, QSP, and AV wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Vollmer, L.L., Liu, F., Nmezi, B. et al. A high throughput, high content screen for non-toxic small molecules that reduce levels of the nuclear lamina protein, Lamin B1. Sci Rep 15, 7314 (2025). https://doi.org/10.1038/s41598-025-91546-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-91546-3