Abstract
Pseudomonas aeruginosa is a common pathogen of a wide range of nosocomial infections in humans, as well as foodborne illnesses. The current study focuses on the molecular genotyping of P. aeruginosa recovered from human, water, and food specimens. A total of 350 samples, fifty samples each from human, tap water, fish swamp, chicken meat, minced meat, raw milk, and hospital surface from the Menoufiya governorate, Egypt. P. aeruginosa was detected in 14.28%, including human (26%), tap water (18%), fish swamp (18%), chicken meat (12%), minced meat (10%), raw milk (16%), and hospital surface (0%). The results of testing of 50 P. aeruginosa isolates against sixteen antibiotics revealed a relatively high antibiotic resistance for Amoxicillin (100%), Erythromycin (98%), Cephradine (90%), Colistin (82%), Oxytetracyclin, (79%), Chloramphenicol (70%), Doxycycline (70%), and Kanamycin (62%) and high susceptibility for Imipenem (96%), Apramycin (94%), Amikacin (90%), Norfloxacin (78%), Sulphamethoxazol (86%), Enrofloxacin (64%), and Ofloxacin (60%). Furthermore, PCR was successfully amplified for the toxA, exoS, and oprL virulence genes at 396, 118, and 504 bp, respectively, as well as amplifying the ermB, pelA, blaTEM, and tetA resistance genes at 639, 786, 516, and 570 bp, respectively. The dendrogram investigation by ERIC-PCR of 10 clinical P. aeruginosa isolates revealed two main clusters and 10 different ERIC-PCR patterns. The presence of P. aeruginosa isolates in food may represent a potential public health concern, with the need for further epidemiological studies, as well as whole-genome sequencing and correlations of P. aeruginosa in water, food samples, and human infections.
Similar content being viewed by others
Introduction
Pseudomonas aeruginosa is an abundant environmental pathogen found in soil and water. It is a main reason for both cystic fibrosis and nosocomial infections in humans1. Definitely, drinking water is consistently regarded as an important environmental habitat for P. aeruginosa2. In addition, severe illnesses resulted from the consumption of contaminated milk products3. Many Pseudomonas species, mostly P. aeruginosa, have been incriminated as the main relevant agents of numerous illnesses in both veterinary and medical concerns4.
Globally, serious nosocomial infections and malignant syndromes are caused by invasive and opportunistic P. aeruginosa, with significant increases in deaths and morbidity rates in humans5. Hence, P. aeruginosa is an obligate aerobe pathogen that can synthesize arginine, proliferate in anaerobic conditions, and this makes it the most plentiful organism on earth6. This organism has been associated with various disease syndromes, including urinary tract infections, respiratory tract infections, pneumonia, and wound infections7. In addition, McCabe-Sellers and Beattie8 reported P. aeruginosa as a causal agent of numerous food-borne illnesses. Additionally, P. aeruginosa is primarily a food contaminant and spoilage organism, with occasional involvement in food-associated infections as a result of the consumption of contaminated food and poor hygiene procedures9.
Interestingly, the emergence of resistant P. aeruginosa isolates to various antibiotics has led to widespread infections in numerous organs such as the respiratory, urinary, and nervous tissues, resulting in substantial morbidity and deaths, and emerging as a nosocomial pathogen, particularly in immunocompromised patients, posing a life-threatening complaint10,11. Furthermore, it was widely recognized that P. aeruginosa’s pathogenicity is largely triggered by various virulence elements5. P. aeruginosa pathogenicity is attributed to the secretion of various invasive, toxigenic extracellular components, which exert substantial effects on the disease severity12. Moreover, the virulence activity is related to many determinants comprising enzymes such as monooxygenase, proteases, alginate, phospholipases, pyocyanin pigment, as well as three secretory systems and toxins, which interfere with the host cellular alleyways13. Additionally, other virulence elements such as exotoxins, proteases, elastases, alginate, phenazine that are encoded by toxA, toxR, lecA, plcH, lasB, and phzA1 genes, respectively, and diverse pigments, thus contribute a role in the disease severity12,14.
Furthermore, P. aeruginosa can create biofilms in a variety of sites, reducing antibiotic treatment effectiveness, which in turn motivates the development of chronic illnesses15. This pathogen is considered the extremely adaptable microorganisms that have the capability to persevere in different environments through its involved genomic elements.
Another critical aspect of P. aeruginosa complications is the emergence of antimicrobial resistance (AMR), which frequently produces obstacles in controlling infections, as well as developing rapid resistance as a result of inherent and developed resistance elements due to abuse or misuse of antimicrobials16. Multidrug-resistant P. aeruginosa multiply by establishing biofilms, which can result in harmful nosocomial infections in hospitalized patients13. Antibiotics become less effective against bacteria that produce biofilms, which is a crucial outcome in the combat against P. aeruginosa strains that produce biofilms17.
Genetic characterization through molecular typing methods offers quick and reliable tools that are useful for tracking the origins and subsequently preventing P. aeruginosa infections across the drinking water chain1. Interestingly, molecular typing is a tool for genetic assortment that determines the crucial phenotypic characters such as virulence determinants, pathogenicity, specificity, and AMR18. Enterobacterial Repetitive Intergenic Consensus Polymerase Chain Reaction (ERIC-PCR) is a Gram-negative enteric bacteria-specific repetitive extragenic palindromic PCR (rep-PCR) technique that aims for highly specific repetitive sequence components19. An Australian study used ERIC-PCR for genotyping of P. aeruginosa isolates and to study their epidemiology20. In a previous survey, ERIC-PCR was reported as a superior tool for classifying P. aeruginosa isolates than other molecular tools21. The use of repeated intergenic consensus sequences is the primary basis for the ERIC PCR method, providing more sensitive results for bacterial typing than phenotypic assays22.
The excessive existence of different food products of animal origin in Egypt without following the health inspection rules has led to contamination by several bacterial species, including P. aeruginosa. Consequently, this study spot highlights on the isolation rate, antimicrobial susceptibility, and application of ERIC-PCR for genetic characterization of P. aeruginous recovered from human and food products, as well as highlights the public health threat and the portability of resistance genes transmitting to humans via the food chain. This study uniquely highlights the potential risk of P. aeruginosa transmission through the food chain by integrating data from food products, hospital environmental surfaces, and human sources within the same regional setting. To the best of our knowledge, this study represents the first report of ERIC-PCR genotyping of P. aeruginosa isolates recovered from food and human samples in Menoufiya Governorate. In addition to this regional novelty, the study’s originality also lies in its combined sampling strategy, which integrates food products, hospital environmental surfaces, and human sources within the same investigation.
Results
Isolation rate of P. aeruginosa from food and human specimens
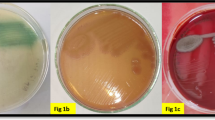
Examination of 350 samples from human pus (50), tap water (50), fish swab (50), chicken meat (50), minced meat (50), raw milk (50), and hospital surfaces (50) revealed that P. aeruginosa was isolated with a 50 (14.28%) isolation rate. All positive samples showed characteristic growth appearance on cetrimide agar medium. As described in Table 1, the prevalence of P. aeruginosa isolated from human, tap water, fish swamp, chicken meat, minced meat, raw milk, and hospital surface was 26, 18, 18, 12, 10, 16, and 0%, respectively.
Biochemical and virulence features of the P. aeruginosa isolates
Next, we confirmed the isolated P. aeruginosa isolates through biochemical and enzymatic activity. The results revealed that all isolates were oxidase positive, TSI (red/red) (K/K) gas negative, urease test negative, indole negative, citrate positive, and catalase positive. Also, all P. aeruginosa isolates that gave characteristic colony morphology in cetrimide agar (yellow-green to blue color) were tested for their pathogenicity by culturing on Congo red medium. The results demonstrated that the biofilm phenotypes accounted for 30% (n = 15), being distributed as follows: 33.33% (n = 5) strong biofilm; 46.6% (n = 7) moderate biofilm; 20% (n = 3) weak biofilm, while 70% of isolates (n = 35) were recognized as non-biofilm producers. Collectively, all P. aeruginosa isolates of food and human origin were found to carry potential virulence activities.
Antimicrobial profile of P. aeruginosa isolates from food products and human samples
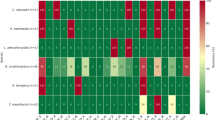
The antimicrobial susceptibility of tested P. aeruginosa isolates from food products and human origin. The results in Table 2 revealed a high level of antibiotic resistance as follows: amoxicillin, erythromycin, cephradine, colistin, oxytetracyclin, chloramphenicol, Doxycycline, and Kanamycin with 100, 98, 90, 82, 79, 70, 70, and 62%, respectively. Meanwhile, Imipenem, apramycin, amikacin, norfloxacin, sulphamethoxazole, enrofloxacin, and ofloxacin exhibited high susceptibility with 96, 94, 90, 78, 86, 64, and 60%, respectively. Together, the tested P. aeruginosa isolates displayed a high degree of AMR from food products and human origin.
Phenotypic resistance profile of P. aeruginosa from food and human samples
Fifty P. aeruginosa isolates showed multidrug resistance (MDR) against 16 different antibiotics from ten antimicrobial groups. The results of the MDR profile are presented in Table 3. Notably, all 16 antibiotics from ten antimicrobial groups were found to be resistant by only one isolate. Additionally, one isolate exhibited resistance against 15 antibiotics from nine different antimicrobial groups. Eleven P. aeruginosa isolates displayed resistance against 14 different antibiotics from eight different antimicrobials. Nine P. aeruginosa isolates demonstrated resistance to 11 antibiotic groups from seven different antimicrobials. Another 9 P. aeruginosa isolates displayed resistance to 8 antibiotics from six antimicrobials.
Besides, we observed four P. aeruginosa isolates displaying MDR to 7 antibiotics from five antimicrobial classes. Only two and five P. aeruginosa were found to be MDR to four and three antimicrobials, respectively. Collectively, almost all of the P. aeruginosa 42 (84%) isolates from human animal food origin showed MDR with an AMR index ranging from 0.25 to 1.
Molecular characterization of virulence and antibiotic resistance genes of P. aeruginosa isolates
Regarding the molecular characterization of the P. aeruginosa strains, certain virulence and antibiotic resistance genes were tested among the selected 15 P. aeruginosa strains, which were biofilm producers. The PCR method successfully amplified 396, 118, and 504 nucleotide fragments of the toxA, exoS, and oprL virulence genes of P. aeruginosa strains, respectively (Supplementary Fig. 1–3). In addition to successfully amplifying 639, 786, 516, and 570 nucleotide fragments of the ermB, pelA, blaTEM, and tetA genes of P. aeruginosa isolates, respectively (Supplementary Fig. 4–7). The prevalence of virulence and resistance genes in the tested isolates from different sources is shown in Table 4.
ERIC-PCR for genotyping of P. aeruginosa isolates
Supplementary Fig. 8 shows ten clinical P. aeruginosa isolates amplified by ERIC-PCR bands that ranged from three to seven bands. Moreover, the molecular fragment size of the PCR products ranged from 100 to 2000 bp, with a common fragment size range, 100–1000 bp, and also 1000–2000 bp, and similarity index ranges (0.18–0.86). The results of dendrogram analysis of ten P. aeruginosa isolates were categorized into 2 main clusters. Moreover, 10 different ERIC patterns were generated. Cluster 1 was divided into 2 sub-clusters, the first one containing 4 serotypes from (water, milk, chicken, and fish) sources. The second sub-cluster contains 3 serotypes from different sources (human and water). Cluster II contains 3 serotypes of different sources (human, minced meat, and fish), as shown in Fig. 1.
Discussion
P. aeruginosais a frequent and common hospital-related pathogen, demonstrative in pneumonia, urinary tract, and post-surgical infections. According to Weiner et al.23, about 7% of health illnesses are due to P. aeruginosa. Our findings reported an overall isolation rate of 14.28% of P. aeruginosa from food and human specimens, particularly from human, tap water, fish swamp, chicken meat, minced meat, raw milk, and hospital surface, with rates of 26%,18%, 18%, 12%, 10%, 16%, and 0%, respectively. Similar international findings reported an overall P. aeruginosa prevalence rate of 16.2% − 23% in intensive care units, with predominance of the respiratory source24. Moreover, similar finding was reported from different human samples from burns, wounds, urine, and throat swabs, with prevalence rates of 22%, 9.5%, 6.9%, and 6.6%, respectively, out of 156 clinical samples.
Concerning the predominance of P. aeruginosa among food products, lower prevalence of P. aeruginosa was recorded in fresh minced meat (5%), fresh sausage (2%), and frozen beef burger (1%) samples in a previous Egyptian study25. In addition, Jawher and Hasan26 reported the predominance of P. aeruginosain chicken, beef, and mutton meat, with detection rates of 22%, 14%, and 6%, respectively. Similarly, in a West African survey, Benie et al.27 found a high prevalence of P. aeruginosa in bovine meat (53.04%), fresh fish (37.69%), and smoked fish (23.57%). These findings collectively indicate widespread contamination of P. aeruginosa across various meat types, including camel meat (80%)28, chicken meat (46.7%)29, frozen imported meat (6.67%)30, and retail meat (3%)31. In a similar line, the current study found that P. aeruginosawas prevalent in raw milk at a rate of 16% while these results were higher than those obtained by Benie et al.27 from subclinical mastitis milk (5.4%) in Pengal.
Meanwhile, our results had a lower prevalence than those obtained by Aziz et al.32, who recovered 70% and 24% from milk and milk tanks, respectively. In an Egyptian study, Ibrahim et al.33 showed that P. aeruginosa was prevalent in milk and dairy products, with a prevalence rate of 22.5% as well as rates of 48%, 16%, 18%, and 8% from raw milk, yoghurt, soft cheese, and Karish cheese, respectively. In addition, our findings reported that the prevalence of P. aeruginosafrom tap water was 18%. A reported study by Rezaloo et al.34 revealed a 20% recovery rate of P. aeruginosa from bovine frozen meat and lower in sausage (2%). The disparity in the spreading and prevalence of P. aeruginosa between different studies may be recognized by the ability of adaptation of P. aeruginosa isolates to the adverse ecological and growth conditions35.
Noteworthy, the testing of 50 P. aeruginosastrains revealed a high level of antibiotic resistance profile for Amoxicillin, Erythromycin, Cephradine, Colistin, Oxytetracyclin, Chloramphenicol, Doxycycline, and Kanamycin, while high susceptibility for Imipenem, Apramycin, Amikacin, Norfloxacin, Sulphamethoxazole, Enrofloxacin, and Ofloxacin. This was in consistent with Rezaloo et al.34, who detected high resistance rates against ampicillin, penicillin, tetracycline, cefoxitin, gentamicin, and clindamycin, with rates of 89.65%, 86.2%, 82.75%, 37.93%, 34.48%, and 31.03%, respectively. Additionally, CDC36 recommended that P. aeruginosa strains emerge with phenotypic resistance patterns to various antimicrobial classes, mainly β-lactams, tetracycline, aminoglycosides, macrolides, and quinolones, causing complicated diseases and high mortality, with severe economic losses, due to high-cost treatment and control measures. Together, the antimicrobial susceptibility results recorded that P. aeruginosa isolates displayed higher antibiotic resistance from food and human origin. Moreover, similar results were obtained by Badr et al.37 revealed P. aeruginosa resistance to tetracycline, doxycycline, nalidixic acid, lincomycin, neomycin, trimethoprim-sulfamethoxazole, and chloramphenicol, and sensitivity to levofloxacin, norfloxacin, ciprofloxacin, colistin sulphate and gentamicin, and streptomycin. Furthermore, P. aeruginosastrains exhibited MDR between 1999 and 2009, as reported by Williams et al.38, who also found that approximately 86% of sepsis-related deaths in pediatric units were attributed to P. aeruginosa.
Our findings described that almost all P. aeruginosa isolates displayed phenotypic resistance against various antibiotics from eight antimicrobial groups. This was in contact with Woolhouse and Ward39, who recorded that P. aeruginosaisolates were likely resistant to various drugs as a result of the unselective practice of several antibiotics in food-producing animals and humans. Besides, Sebola et al.40 observed P. aeruginosa resistance to β-lactam classes, predominantly to penicillin and cephalosporins.
Another critical aspect of P. aeruginosa is the presence of numerous virulence and antimicrobial resistance genes within its genome, as well as its ability to form biofilms, which enhance its resistance and pathogenicity41. In our study, the PCR successfully amplified 396, 118, and 504 nucleotide fragments of the toxA, exoS, and oprL virulence genes of P. aeruginosastrains. Additionally, Rezaloo et al.34 showed that exoS, lasA, exoU, lasB, plcH, and algD were the most virulence-detected genes from meat and meat products origin, with rates of 75.86%, 68.96%, 58.62%, 51.72%, 48.27%, and 44.82%, respectively, and this highlights that the consumption of such contaminated food products poses a serious public health threat. Furthermore, Badr et al.37 applied PCR for efficient recognition of the oprL gene at 504 bp fragments. Moreover, Al-Wrafy et al.42 described exotoxin A, designated as the tox A gene, which is a defensive component that plays a prospective role in P. aeruginosa pathogenicity through its cytotoxic activity. The virulence activity is essential to collaborate during P. aeruginosa establishment and biofilm activity, permitting further infection, and enhancing more pathogenic effects.
In addition, our results revealed successful amplification of 639, 786, 516, and 570 nucleotide fragments of the ermB, pelA, blaTEM, and tetA genes of P. aeruginosa strains. Similar findings found that blaDHA, blaCTX-M, and blaSHV were the main commonly identified resistance elements with rates of 93.1%, 83.65%, and 48.27%, respectively34. Similarly, Ibrahim et al.33 reported that P. aeruginosa harbored the resistance genes blaTEM, blaSHV, and ermB, with a multiple antibiotic resistance (MAR) index of 0.50.
It’s worth mentioning that the ERIC-PCR method is a rapid screening, PCR-based tool based on the number of ERIC sequences, as well as a genetic pointer for bacterial genotyping43. Therefore, ERIC PCR is a trustworthy, low-cost method for genotyping of P. aeruginosa44. Furthermore, Eladawy et al.45 applied ERIC-PCR for P. aeruginosa genotyping and revealed 99 different ERIC patterns with 70% similarity that were categorized into 8 main clusters, and thus indicated the significance of recognizing and regulating the possible causes of P. aeruginosa infections. Furthermore, an Iranian study tested 99 MDR P. aeruginosa strains by REIC-PCR, eight clusters, and 50 single genetic copies were produced, with high heterogeneity among isolates from burn infections46. Nevertheless, next-generation sequencing will permit capturing of the whole genome of the bacterium and offer a definitive purpose. However, this might not be available/possible under many conditions.
Conclusion, limitations, and recommendations
P. aeruginosa is a noteworthy healthcare-associated pathogen in humans and food consumers, causing mortalities. This current study spotlights the relatively high recovery rate of P. aeruginosa in human, water, and many food samples, and thus suggests its threat in the aspect of public health. In addition, almost all P. aeruginosa isolates displayed high levels of drug resistance for many antimicrobial drugs that constitute a significant challenge to food safety and human consumers, as well as limiting the selection of effective antibiotics. The existence of diverse virulence genetic elements among screened P. aeruginosa isolates suggests their critical role in the pathogenicity and disease severity. Furthermore, our study showed the high polymorphism and genetic variety among P. aeruginosa isolates from food and human origin, with relatively different progenitors, which revealed different infection sources. Further surveys using phenotypic and genotypic approaches are mandatory to explore the relationship between P. aeruginosa phenotypic resistance, virulence, and biofilm activity. Also, a better understanding of the epidemiology of antibiotic resistance transmission to humans would help to reduce the transmission of P. aeruginosa infections between patients and the environment and food-borne outbreaks. Conversely, whole-genome sequencing of P. aeruginosa is essential for accurate typing of clinical isolates; however, it remains unavailable in many laboratories.
Finally, this study highlights the importance of continuous surveillance to detect biofilm-forming and antimicrobial-resistant P. aeruginosa isolates from food sources. We recommend enhancing hygiene and sanitation practices throughout the food production and handling chain, as well as routine monitoring of high-risk foods and food-contact surfaces. Implementation of robust antimicrobial stewardship programs is also essential to reduce potential public health risks. Future research should involve larger sample sizes and broader molecular characterization to better understand the epidemiology, resistance mechanisms, and potential transmission pathways of food-associated P. aeruginosa. In hospital settings, strict infection control measures, including environmental sanitation, water system monitoring, adherence to hand hygiene protocols, antimicrobial stewardship, and routine surveillance, are critical to limit the spread of P. aeruginosa and reduce healthcare-associated infections. We also acknowledge that our hospital sampling, limited to surface swabs, may underestimate the true prevalence of P. aeruginosa.
Materials and methods
Ethics approval and consent to participate
Human samples and hospital surface swabs were collected by medical staff at Sadat City Hospital in Menoufia Governorate, which reviewed and approved the study, in accordance with the regulations of the Egyptian Ministry of Health and under full medical supervision. Written informed consent was obtained from all patients prior to specimen collection. In addition, various food samples, including water, fish, chicken, meat, and milk, were collected from local markets in the study area around the hospital. All animal-handling procedures, as well as samples’ collection and disposal, were performed according to the regulations of the Institutional Animal Care and Use Committee (IACUC) with oversight of the faculty of Veterinary Medicine, University of Sadat City, Egypt (Ethical approval number: VUSC-010-1-25).
Study area
This studied area is Sadat City, which is one of the Menoufia Governorate centers, and considered as common of the industrialized areas in Egypt. The area is located about 94 km northwest of Cairo, between 30.3811◦ N and 30.5266◦ E, and it is considered one of the first-generation new urban publics in Egypt. Most inhabitants reside in rural areas47. The area is considered one of the manufacturing areas where the government emphasizes farming, factories of food manufacturing, and packaging. Therefore, the food industry is a major business for the residents in the study area.
Samples collection, preparation, and processing
Fifty samples from each (water, minced meat, chicken, milk, fish, hospital surface, and human pus from open wounds), with a total of 350 samples, were gathered from December 2022 to April 2023. The selection of 50 samples per category was primarily guided by sample availability during the study period and practical feasibility, while ensuring equal representation across all sample types to allow meaningful comparative analysis.
Fish and chicken used in this study were purchased from local markets in Sadat City, Egypt. All food product samples were collected using a systematic random sampling approach from multiple local markets across the study area to enhance representativeness and minimize selection bias. The samples were placed in clean polyethylene plastic bags under a septic condition and transported in cooled conditions at 4 °C in a heat-insulated ice box as recommended by APHA48. While the human samples were collected from hospitals by medical staff under physician supervision, as recommended by Rice et al.49. Following, the samples were transported to the laboratory in cooled conditions at 4 °C for further microbiological laboratory investigation.
Each gram of samples/ml was transported into 10 ml of a sterile buffer peptone water supplemented with Novobiocin 10 mg/500 ml, then homogenized, and incubation was done at 37 °C for 18 ± 2 h under suitable conditions as described by APHA48. The human pus samples were further processed, homogenized, and incubated at 37 °C for 18 h in nutrient broth, as follows by Rice et al.49.
Isolation and identification of P. aeruginosa
All the prepared specimens were streaked on selective medium Cetrimide-nalidixic acid agar within 8 h and incubated at 37 °C for 24 h. After 24 h incubation, the identified colonies were then subcultured on citramide agar for 24 ± 2 h at 37 ᵒC50. The positive colonies were then picked up for further subculturing on nutrient agar for the purification of colonies. The pure colonies were confirmed by morphological and biochemical identification, including oxidase test, Indole, Citrate, TSI, Catalase, H2S production, and Haemolytic activity according to Cheesbrough51.
Antimicrobial profile of P. aeruginosa
The method of Kirby–Bauer disk diffusion was used for the recovered P. aeruginosa isolates to evaluate their antimicrobial susceptibility. Adjustment to 0.5 McFarland was used for the prepared bacterial suspension. All antimicrobial disks of different antimicrobial classes (Oxoid Ltd., UK) were used for this test.
The following antibiotics were used: 30 µg Oxytetracycline (TE), 30 µg Doxycycline (DO), 5 µg Ofloxacin(O), 5 µg Enrofloxacin (EN), 10 µg Norfloxacin (NOR), 5 µg Ciprofloxacin (CIP), 15 µg Erythromycin (E), 30 µg Amikacin (AK), 30 µg Kanamycin (K), 30 µg Chloramphenicol (C), 10 µg Apramycin (A), 10 µg Imipenem (IPM), 30 µg Amoxicillin (AX), 30 µg Cephradine (CE), 25 µg Sulfamethoxazole (SXT), and 25 µg Colistin (CO). The antimicrobial results were assessed according to the guidelines of CLSI52 by measuring the inhibition zone and recording it as resistant, intermediate, or susceptible.
Molecular identification of P. aeruginosa isolates
The DNA of P. aeruginosa isolates was extracted using the commercial kits QIAamp DNA Mini Kit (Qiagen, Hilden, Germany). Using specific primers, some virulence and resistance genes of P. aeruginosa isolates were detected, as listed in Table 5. The PCR reaction was performed in a 50 µL volume composed of 25 µL of PCR Master Mix (Qiagen, Germany), 6 µL target DNA, 1 µL of each primer (10 pmol/µL), and deionized water was completed to up to 50 µL. The PCR reaction was conducted in a thermal cycler (Applied Biosystems, Foster, CA, USA). The following protocol was used: Amplification conditions composed of an initial denaturation at 94 °C for 5 min, followed by 35 cycles of amplification (secondary denaturation at 94 °C for 30 s, annealing at 50 °C for 40 s, and extension at 72 °C for 45 s) and final extension at 72 °C for 10 min. After that, 1.5% gel electrophoresis was prepared for the PCR products (15 µL) and examined under UV light.
Genotyping and analysis of P. aeruginosa by enterobacterial repetitive intergenic consensus polymerase chain reaction (ERIC-PCR)
ERIC fingerprinting data were transformed into a binary code depending on the presence or absence of each band. For the construction of dendrograms, both the unweighted pair group method with arithmetic average (UPGMA) and Ward’s hierarchical clustering routine were applied. Also, cluster and dendrogram analyses were performed using SPSS, version 2253,54. Similarity index between examined specimens was calculated using the online search tool (https://planetcalc.com/1664/).
Dendrogram analysis for Pseudomonas aeruginosa isolates by ERIC-PCR.
Data availability
Datasets generated during and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Wood, S. J., Kuzel, T. M. & Shafikhani, S. H. Pseudomonas aeruginosa: Infections, animal modeling, and therapeutics. Cells 12, 199. https://doi.org/10.3390/cells12010199 (2023).
Ruiz-Roldán, L. et al. Occurrence of Pseudomonas spp. In Raw vegetables: molecular and phenotypical analysis of their antimicrobial resistance and virulence-related traits. Int. J. Mol. Sci. 22, 12626. https://doi.org/10.3390/ijms222312626 (2021).
Abdurezak, A., Takele, B. & Walkite, F. Isolation, identification and antimicrobial resistance profiles of Salmonella from dairy farms in Adama and Modjo Towns, central Ethiopia. Eur. J. Med. Health Sci. 2, 1–11. https://doi.org/10.34104/ejmhs.02001011 (2020).
Gauthier, J., Charette, S. J. & Derome, N. Draft genome sequence of Pseudomonas fluorescens ML11A, an endogenous strain from brook charr with antagonistic properties against Aeromonas salmonicida subsp. salmonicida. Genome Announcements 5, 1. https://doi.org/10.1128/genomeA.01716-16 (2017).
Adlbrecht, C. et al. Efficacy, immunogenicity, and safety of IC43 Recombinant Pseudomonas aeruginosa vaccine in mechanically ventilated intensive care patients - a randomized clinical trial. Crit. Care. 24, 74. https://doi.org/10.1186/s13054-020-2792-z (2020). https://doi.org:.
Lederberg, J. Infectious history. Science 288, 287–293. https://doi.org/10.1126/science.288.5464.287 (2000). https://doi.org:.
Zakaria, Z. et al. Physicochemical composition, Microbiological quality and consumers’ acceptability of Raw and pasteurized locally produced goat milk. Malaysian J. Fundamental Appl. Sci. 16, 475–482 (2020).
McCabe-Sellers, B. J. & Beattie, S. E. Food safety: emerging trends in foodborne illness surveillance and prevention. J. Am. Diet. Assoc. 104, 1708–1717 (2004).
Voynow, J. A., Fischer, B. M. & Zheng, S. Proteases and cystic fibrosis. Int. J. Biochem. Cell. Biology. 40, 1238–1245. https://doi.org/10.1016/j.biocel.2008.03.003 (2008). https://doi.org:
Abdelrahman, D. N. et al. β-lactamases (blaTEM, blaSHV, blaCTXM-1, blaVEB, blaOXA-1) and class C β-lactamases gene frequency in Pseudomonas aeruginosa isolated from various clinical specimens in Khartoum State, Sudan: A cross sectional study. F1000Research 9, 774. https://doi.org/10.12688/f1000research.24818.3 (2021).
Pratama, I. G. W. T., Darwinata, A. E. & Tarini, N. M. A. Gene similarity examination Pseudomonas aeruginosa in intensive care unit (ICU) at Prof. Dr. dr. IGNG Ngoerah hospital using the ERIC-PCR method. Intisari Sains Medis. 14, 608–611. https://doi.org/10.15562/ism.v14i2.1656 (2023).
Sánchez-Diener, I. et al. Weighting the impact of virulence on the outcome of Pseudomonas aeruginosa bloodstream infections. Clin. Microbiol. Infect. 26, 351–357. https://doi.org/10.1016/j.cmi.2019.06.034 (2020). https://doi.org:.
Abdelaziz, A. A., Kamer, A. M. A., Al-Monofy, K. B. & Al-Madboly, L. A. Pseudomonas aeruginosa’s greenish-blue pigment pyocyanin: its production and biological activities. Microb. Cell. Fact. 22, 110. https://doi.org/10.1186/s12934-023-02122-1 (2023).
Liao, C., Huang, X., Wang, Q., Yao, D. & Lu, W. Virulence factors of Pseudomonas aeruginosa and antivirulence strategies to combat its drug resistance. Front. Cell. Infect. Microbiol. 12, 926758. https://doi.org/10.3389/fcimb.2022.926758 (2022).
Thi, M. T. T., Wibowo, D. & Rehm, B. H. Pseudomonas aeruginosa biofilms. Int. J. Mol. Sci. 21, 8671. https://doi.org/10.3390/ijms21228671 (2020).
Morales-Espinosa, R. et al. Pseudomonas aeruginosa strains isolated from animal with high virulence genes content and highly sensitive to antimicrobials. J. Global Antimicrob. Resist. 37, 75–80. https://doi.org/10.1016/j.jgar.2024.02.023 (2024). https://doi.org:.
Hoiby, N., Ciofu, O. & Bjarnsholt, T. Pseudomonas aeruginosa biofilms in cystic fibrosis. Future Microbiol. 5, 1663–1674. https://doi.org/10.2217/fmb.10.125 (2010). https://doi.org:.
Li, W., Raoult, D. & Fournier, P. E. Bacterial strain typing in the genomic era. FEMS Microbiol. Rev. 33, 892–916. https://doi.org/10.1111/j.1574-6976.2009.00182.x (2009). https://doi.org:.
Versalovic, J., Koeuth, T. & Lupski, R. Distribution of repetitive DNA sequences in eubacteria and application to finerpriting of bacterial Enomes. Nucleic Acids Res. 19, 6823–6831. https://doi.org/10.1093/nar/19.24.6823 (1991). https://doi.org:
Stehling, E. G., Leite, D. S. & Silveira, W. D. Molecular typing and biological characteristics of Pseudomonas aeruginosa isolated from cystic fibrosis patients in Brazil. Braz. J. Infect. Dis. 14, 462–467 (2010).
Hematzadeh, A. & Haghkhah, M. Biotyping of isolates of Pseudomonas aeruginosa isolated from human infections by RAPD and ERIC-PCR. Heliyon 7, e07967. https://doi.org/10.1016/j.heliyon.2021.e07967 (2021). https://doi.org:.
Otokunefor, K., Ogugbue, C. & Fajoyomi, B. Enterobacterial repetitive intergenic consensus (ERIC) as a tool for genetic characterisation of bacterial isolates in Nigeria. Nigerian J. Biotechnol. 37, 122–128. https://doi.org/10.4314/njb.v37i1.13 (2020). https://doi.org:
Weiner, L. M. et al. Antimicrobial-resistant pathogens associated with healthcare-associated infections: summary of data reported to the National healthcare safety network at the centers for disease control and Prevention, 2011–2014. Infect. Control Hosp. Epidemiol. 37, 1288–1301. https://doi.org/10.1017/ice.2016.174 (2016).
Vincent, J. L. et al. Prevalence and outcomes of infection among patients in intensive care units in 2017. JAMA 323, 1478–1487. https://doi.org/10.1001/jama.2020.2717 (2020). https://doi.org:.
Salem, A., Ismail, A. & Marouf, H. Prevalence of Pseudomonas aeruginosa and its toxins in some meat products. Global J. Agric. Food Saf. Sci. 1, 39–50 (2014).
Jawher, I. M. & Hasan, M. G. Antibiotics resistance patterns of Pseudomonas aeruginosa isolated from meat at Mosul City retails. Iraqi J. Veterinary Sci. 37, 363–367 (2023).
Benie, C. et al. Prevalence and antibiotic resistance of Pseudomonas aeruginosa isolated from bovine meat, fresh fish and smoked fish. Archives Clin. Microbiol. 8, 1–9 (2017).
Mahato, A., Mahato, S., Dhakal, K. & Dhakal, A. Investigating the diversity of spoilage and food intoxicating bacteria from chicken meat of Biratnagar, Nepal. J. Bacteriol. Mycol. 7, 1141 (2020).
Benie, C. et al. Prevalence and diversity of Pseudomonas spp. Isolated from beef, fresh and smoked fish in Abidjan, Côte d’ivoire. J. Food Dairy. Technol. 4, 52–61 (2016).
Odoi, J. O. et al. Prevalence of colistin-resistant bacteria among retail meats in Japan. Food Saf. 9, 48–56. https://doi.org/10.14252/foodsafetyfscj.D-21-00002 (2021). https://doi.org:.
Osman, K., Orabi, A., Elbehiry, A., Hanafy, M. H. & Ali, A. M. Pseudomonas species isolated from camel meat: quorum sensing-dependent virulence, biofilm formation and antibiotic resistance. Future Microbiol. 14, 609–622. https://doi.org/10.2217/fmb-2018-0293 (2019).
Abdel Aziz, S. A. A., Mahmoud, R. & Mohamed, M. B. E. D. Control of biofilm-producing Pseudomonas aeruginosa isolated from dairy farm using Virokill silver nano-based disinfectant as an alternative approach. Sci. Rep. 12, 9452. https://doi.org/10.1038/s41598-022-13619-x (2022).
Ibrahim, M. M., Elsaied, E. I., Aal, A. E., Bayoumi, M. A. & S. F. & Prevalence of Pseudomonas aeruginosa in milk and some dairy products with reduction trials by some natural preservatives. J. Adv. Veterinary Res. 12, 434–438 (2022).
Rezaloo, M., Motalebi, A., Mashak, Z. & Anvar, A. Prevalence, antimicrobial resistance, and molecular description of Pseudomonas aeruginosa isolated from meat and meat products. J. Food Qual. 2022, 9899338. https://doi.org/10.1155/2022/9899338 (2022).
Sofy, A. R., Sharaf, A., Al Karim, A. G., Hmed, A. A. & Moharam, K. M. Prevalence of the harmful Gram-negative bacteria in ready-to-eat foods in Egypt. Food Public. Health. 7, 59–68 (2017).
CDC. Center of Diseases Control and Prevention (CDC). Multidrug-resistant Pseudomonas aeruginosa (2017).
Badr, H., Roshdy, H., El-Hafez, A., Farghaly, E. M. & A. & Prevalence, pathogenicity and antibiogram sensitivity of Pseudomonas aeruginosa isolated from diseased chickens. Assiut Veterinary Med. J. 62, 119–126. https://doi.org/10.21608/avmj.2016.170015 (2016).
Williams, F. N. et al. The leading causes of death after burn injury in a single pediatric burn center. Crit. Care 13, R183. https://doi.org/10.1186/cc8170 (2009).
Woolhouse, M. E. & Ward, M. J. Sources of antimicrobial resistance. Science 341, 1460–1461. https://doi.org/10.1126/science.1243444 (2013). https://doi.org:.
Sebola, D., Eliasi, U. L., Oguttu, J. W. & Qekwana, D. N. Antimicrobial resistance patterns of Pseudomonas aeruginosa isolated from canine clinical cases at a veterinary academic hospital in South Africa. J. S. Afr. Vet. Assoc. 91, 1–6. https://doi.org/10.4102/jsava.v91i0.2052 (2020).
Płókarz, D. & Rypuła, K. A one health perspective on the human-pets Pseudomonas aeruginosa transmission. Appl. Microbiol. 8, 227 (2022).
Al-Wrafy, F., Brzozowska, E., Górska, S. & Gamian, A. Pathogenic factors of Pseudomonas aeruginosa: the role of biofilm in pathogenicity and as a target for phage therapy. Adv. Hyg. Exp. Med./Postepy Higieny I Med. Doswiadczalnej 71, 78–91. https://doi.org/10.5604/01.3001.0010.3792 (2017).
Ranjbar, R. & Ghazi, F. M. Antibiotic sensitivity patterns and molecular typing of Shigella sonnei strains using ERIC-PCR. Iran. J. Public. Health. 42, 1151–1157 (2013).
Najem, S. A. A. & Shubbar, E. E. Molecular analysis of clinical isolated Pseudomonas aeruginosa by ERIC PCR. Int. J. Health Sci. 6, 10144–10151. https://doi.org/10.53730/ijhs.v6nS1.7203 (2022). https://doi.org:.
Eladawy, M., El-Mowafy, M., El-Sokkary, M. M. & Barwa, R. Antimicrobial resistance and virulence characteristics in ERIC-PCR typed biofilm forming isolates of P. aeruginosa. Microb. Pathog. 158, 105042. https://doi.org/10.1016/j.micpath.2021.105042 (2021).
Khosravi, A. D., Hoveizavi, H., Mohammadian, A., Farahani, A. & Jenabi, A. Genotyping of multidrug-resistant strains of Pseudomonas aeruginosa isolated from burn and wound infections by ERIC-PCR. Acta Cirurgica Brasileira. 31, 206–211. https://doi.org/10.1590/S0102-865020160030000009 (2016).
CAPMAS. The result of fifth economic Census 2017–2018. Total Egypt’s economic activity and governorates (2018).
APHA. Compendium of Methods for the Microbiological Examination of Foods, 3rd edn. (American Public Health Association, 1991).
Rice, M. et al. Child and maternal factors that influence child blood pressure in preschool children: an exploratory study. Appl. Nurs. Res. 31, 117–120. https://doi.org/10.1016/j.apnr.2016.01.008 (2016). https://doi.org:.
Quinn, P., Carter, M., Markey, B. & Carter, G. Clinical Veterinary Microbiology, 109–135 (Wolfe publishing, 1994).
Cheesbrough, M. District Laboratory Practice in Tropical Countries Part II, Low Price Edition, 193–194 (2000).
CLSI. Performance standards for antimicrobial disk susceptibility tests; Approved Standard, 12th edn (Clinical and Laboratory Standards Institute, 2012).
Hunter, J. E. & Schmidt, F. L. Methods of meta-analysis: Correcting Error and Bias in Research Findings (Sage Publications, Inc., 1990).
IBM. Discovering Statistics Using IBM SPSS Statistics (Sage Publications Ltd, 2013).
Matar, G. M., Ramlawi, F., Hijazi, N., Khneisser, I. & Abdelnoor, A. M. Transcription levels of Pseudomonas aeruginosa exotoxin A gene and severity of symptoms in patients with otitis externa. Curr. Microbiol. 45, 350–354. https://doi.org/10.1007/s00284-002-3703-z (2002).
Xu, J., Moore, J. E., Murphy, P. G., Millar, B. C. & Elborn, J. S. Early detection of Pseudomonas aeruginosa–comparison of conventional versus molecular (PCR) detection directly from adult patients with cystic fibrosis (CF). Ann. Clin. Microbiol. Antimicrob. 3, 21. https://doi.org/10.1186/1476-0711-3-21 (2004).
Winstanley, C. et al. Genotypic and phenotypic characteristics of Pseudomonas aeruginosa isolates associated with ulcerative keratitis. J. Med. Microbiol. 54, 519–526. https://doi.org/10.1099/jmm.0.46005-0 (2005). https://doi.org:.
Nguyen, L. C. et al. Genetic analysis of genes involved in synthesis of modified 4-amino-4, 6-dideoxyglucose in Flagellin of Pseudomonas syringae pv. tabaci. Mol. Genet. Genomics. 282, 595–605. https://doi.org/10.1007/s00438-009-0489-8 (2009). https://doi.org:
Ghadaksaz, A., Fooladi, A. A. I., Hosseini, H. M. & Amin, M. The prevalence of some Pseudomonas virulence genes related to biofilm formation and alginate production among clinical isolates. J. Appl. Biomed. 13, 61–68. https://doi.org/10.1016/j.jab.2014.05.002 (2015).
Colom, K. et al. Simple and reliable multiplex PCR assay for detection of blaTEM, blaSHV and blaOXA–1 genes in Enterobacteriaceae. FEMS Microbiol. Lett. 223, 147–151. https://doi.org/10.1016/S0378-1097(03)00306-9 (2003). https://doi.org:.
Randall, L., Cooles, S., Osborn, M., Piddock, L. & Woodward, M. J. Antibiotic resistance genes, integrons and multiple antibiotic resistance in thirty-five serotypes of Salmonella enterica isolated from humans and animals in the UK. J. Antimicrob. Chemother. 53, 208–216. https://doi.org/10.1093/jac/dkh070 (2004).
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB). This research received no external funding.
Author information
Authors and Affiliations
Contributions
Conceptualization: WSM and EEA; Methodology: WSM, EEA, HFE, RMA, and MMH; Validation: WSM, EEA, HFE, and MMH; Formal analysis: WSM, EEA, HFE, RMA, and MMH; Investigation: WSM, EEA, HFE, RMA, and MMH; Data Curation: WSM, EEA, HFE, RMA, and MMH; Writing - Original Draft: RMA and MMH; Writing - Review & Editing: WSM, EEA, and HFE; Supervision: WSM and EEA. All the authors contributed to the final manuscript and approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mousa, W.S., Abdeen, E.E., El-Gendy, H.F. et al. Prevalence and genotyping of Pseudomonas aeruginosa from food and human sources. Sci Rep 16, 7179 (2026). https://doi.org/10.1038/s41598-026-37559-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-37559-y