Abstract
The rapid expansion of biomedical datasets generated by international research programs across Africa highlights an urgent need for advanced data science expertise to translate these resources into impactful interventions. We designed an innovative training model for a data science research program funded through the National Institutes of Health Data Science for Health Discovery and Innovation in Africa initiative. The two-phase training model combined foundational instruction, integrating machine learning with traditional statistical methods, followed by intensive group- and competition-based learning. The model was tailored to leverage and analyze multimodal data warehouses from completed research projects. The inaugural datathon united trainees from 14 African countries, leveraging data sources for a large cohort study on malaria. Our pedagogical strategy bridged traditional statistical methods with analog machine learning approaches, illustrated through case studies on regression and image analysis. Each datathon team initiated a research project that culminates in a scientific manuscript. This paper details our complete datathon learning model, including its core competencies, learning objectives, and evaluation metrics, offering a comprehensive resource for researchers seeking to implement similar programs. The datathon framework provides a practical platform for advancing data science skills, fostering multidisciplinary research, and maximizing the impact of biomedical data resources.
Similar content being viewed by others
Introduction
Sub-Saharan African countries are severely affected by the world’s most devastating infectious diseases, including malaria, HIV/AIDS, tuberculosis, and neglected tropical diseases. In Mali, malaria remains a major public health concern, with over 90% of its population residing in malaria-endemic areas1. A critical gap in current malaria control strategies is the limited availability of quality data and data science expertise to guide the implementation and evaluation of interventions. Recent studies have revealed a significant underrepresentation of African populations in local biomedical data sources, a disparity with profound implications in the era of artificial intelligence2,3.
The University of Sciences, Techniques and Technologies of Bamako, Mali (USTTB) has led training, infrastructure, and research programs sponsored by the National Institutes of Health (NIH) for decades4. One example is the West Africa Center of Excellence for Data Science Research, Education, and Training (WACE-DSRE) program. This program was launched in 2023 as part of the NIH Data Science for Health Discovery and Innovation in Africa (DS-I Africa) initiative5. DS-I Africa is a large collaborative network of research and education programs that leverages data science technologies to transform biomedical and public health research in Africa6. An earlier NIH training program, the West African Center of Excellence for Global Health Informatics Training, built bioinformatics and data science capacity by fostering genetics research to address global health issues in Africa as part of the Human Heredity and Health in Africa (H3Africa) initiative7. The African Center of Excellence in Bioinformatics and Data-Intensive Science of Bamako, Mali (ACE-B) is one of two such centers on the African continent that facilitate research and training through advanced data science cyberinfrastructure8.
A flagship research initiative in Mali is the West Africa International Center of Excellence for Malaria Research (WA-ICEMR)9. This program is one of eight regional centers within the global network of International Centers of Excellence for Malaria Research (ICEMRs), established in malaria-endemic regions to generate knowledge, develop tools, and produce evidence-based strategies that strengthen malaria control efforts10,11,12. For over 15 years, the WA-ICEMR program and other international partnerships have generated a breadth of complex clinical and biomedical data sets across the human, vector, and parasite domains, leading to rich data availability for the broader scientific community to support malaria control interventions13. These data sources, coupled with those generated through routine disease surveillance, offer considerable opportunities for applying data science to make discoveries that catalyze innovation in diagnosing and treating diseases in West Africa. Integrating biomedical data sources into training programs provides empirical data and inputs that strengthen and refine training models. However, these opportunities remain underutilized due to critical shortages of skilled researchers and professionals with data science expertise who can organize and analyze these sources to drive meaningful improvements in public health outcomes.
The WACE-DSRE aims to expand multidisciplinary data science and analytical capacity in West Africa. The program is a multi-institutional research education partnership of investigators at USTTB, Gamal Abdel Nasser University of Conakry (UGANC), and Tulane University5. Leveraging training, research, and infrastructure partnerships, the WACE-DSRE sought to enhance data science research capacity in West Africa through comprehensive training, tailored content, and intensive mentoring in data science. The program specifically focused on multidisciplinary team building across multiple institutions to build data science capacity in the subregion.
The signature training component of the WACE-DSRE is its innovative datathon program, which builds on hackathon learning principles. Hackathons have become a hallmark of Silicon Valley’s innovation culture through short-duration, competition-based approaches for generating groundbreaking solutions14,15,16,17. These events bring together computer programmers to collaborate intensively with one another to meet a shared goal within a short timeframe18. Using hackathon principles, datathons were introduced in 2016 to assemble data scientists, statisticians, and clinical experts to perform sound, reproducible analyses16,19. Datathons organize groups into teams to promote competitive problem-solving for data-driven challenges20. Like hackathons, datathons utilize written and oral communication to convey the timing, schedule, goals, and theme of the training, and typically award prizes to competition winners21. Datathon training has been shown to foster positive leadership principles and increase the likelihood of research projects reaching completion20,22. Datathons have been increasingly used in healthcare; for instance, they were used to develop techniques for predicting infectious disease outbreaks in response to the COVID-19 pandemic23,24. Datathon events have also been utilized in critical care training and proposed as training models in neurosurgery25. These events are typically short-term training mechanisms. Short-term training has been identified as a crucial complement to traditional degree programs to meet the growing demand for data science expertise26.
Internet search interest for datathons has recently surged (reaching a Google Trends search score of 99 in 2024). Yet, a notable gap remains in the published literature detailing their protocols and practical implementation, and to our knowledge, none have covered data science training in West Africa. A recent PubMed search with the keywords “data science” and “training” yielded 3,864 hits. However, repeating this search with “West Africa” as an additional search term yielded just 11 results. We postulated that combining datathon training approaches with data sources and infrastructure through historical research projects could be used to build multidisciplinary and international data science capacity in West Africa. In this paper, we describe our novel datathon framework to assist the scientific community in enhancing and expanding data science training in African contexts.
Methods
We developed the datathon training model in response to a call for data science research education proposals from the NIH DS-I initiative, launching the program in 2023. Designed to be more interactive and multidisciplinary than our previous research training workshops, we integrated foundational instruction with an intensive, trainee-led competition on data science research projects. The program utilized research data from a large longitudinal cohort study conducted through the WA‑ICEMR program. All methods were conducted in accordance with relevant guidelines and regulations. Experimental protocols for all of the data sources used in the datathon projects were reviewed and approved by the Institutional Review Boards (IRBs) of the USTTB (FWA00001769), University Cheikh Anta Diop in Dakar, Senegal (FWA00002691), Medical Research Council Unit in The Gambia (FWA00006873), and Tulane University (FWA00002055) before participant enrollment in the longitudinal cohort study. Informed consent for the use and publication of anonymized cohort data was obtained from all participants at the time of enrollment. The USTTB Ethics Committee determined that analyses of de-identified, aggregate datathon evaluation results were exempt from additional ethics review.
Conceptual framework
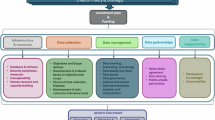
The datathon framework was divided into four main stages: a pre-datathon phase, a foundational phase, the datathon event, and a follow-up period. Each stage of the training was characterized by its specific length, modality, purpose, and deliverables, as shown in Fig. 1.
Overview of the WACE-DSRE datathon training model. WACE-DSRE = West Africa Center of Excellence for Data Science Research Education and Training program. The schematic illustrates the structure and key components of the datathon training phases. The first component consisted of instructor-led instruction (hybrid modality), and the second was intensive datathon training (in-person modality). Participants could attend the foundational training remotely, without needing to attend the in-person phase of the datathon training. Trainees for the in-person datathon training were required to complete the foundational training (either in-person or remotely).
The pre-datathon foundation training was a 5-day hybrid format that accommodated up to 50 trainees, focusing on data source acclimatization and foundational coding skills. Certificates were awarded to participants upon completion of the pre-datathon training. The datathon phase lasted an additional 5 days and was conducted in person with up to 15 trainees. The event centered on idea and hypothesis generation, coding, analysis, and presentation and validation of results. The datathon component also awarded certificates upon completion. The follow-up stage consisted of weekly progress meetings with the datathon teams to develop manuscripts as deliverables. Meetings were on a rotational basis, where each team presented its progress once every five weeks.
Participant recruitment and travel support
The target participants were graduate students, early-career researchers, and professionals at affiliate research institutions and consortia. Selected participants typically specialized in biological sciences, public health, clinical research, epidemiology, data science, or bioinformatics. Recruitment was conducted via electronic announcements sent via e-mail to research and consortium affiliates. Applicants were evaluated using a structured rubric designed to assess their alignment with program objectives and their stated expectations. Selection criteria included current professional position, nationality, field of study or research, program expectations, and demonstrated level of expertise in data science. All applicants were required to submit a résumé or curriculum vitae. Additional details on the trainee recruitment process are provided in Supplementary File 1, Table S1, and the datathon training announcement is available in Supplementary File 2. Fifteen participants were awarded monetary stipends to cover all travel and lodging expenses for both the foundational and datathon phases.
Phase 1: Foundational, instructor-led training
The foundational training modules were designed to build fundamental skills in computer programming and data analytics. These skills were then utilized for project development during the datathon phase of the training. We established the foundational competencies for the program aligned with Bloom’s Revised Taxonomy27. A detailed mapping of competencies to learning objectives for the foundational training component is presented in Table 1.
Each module prioritized practical, hands-on learning by presenting key topics, followed by interactive exercises utilizing computer applications. The applications selected for the datathon training purposely included those that were open source or freely available. Requisite applications for the program were available across Microsoft Windows and Apple macOS operating systems. For the training, ACE-B provided a teaching computer laboratory equipped with personal computers and projectors, and the trainees also maintained personal laptop computers. The foundational phase was a hybrid (in-person and remote) learning model. Successful program completion for the foundation phase was determined by attending at least four out of the five days of training (80% attendance). Upon successful completion of the foundational training, trainees were awarded certificates of completion.
Data science pedagogical approaches
Because most of our selected participants were familiar with traditional statistics, our instructional strategy for introducing artificial intelligence (AI) and machine learning (ML) focused on bridging the gap between statistical concepts and ML approaches. We defined traditional statistics as analytical techniques to apply sample data to draw conclusions about a broader population. We introduced fundamental AI and ML terminology and highlighted real-world applications. Traditional statistics and AI/ML approaches were then connected by mapping their terminologies, helping trainees leverage their existing knowledge as a foundation for understanding AI and ML concepts. Table 2 provides a comparison of select characteristics for traditional analytical methods and their corresponding ML analogs, as explored in our foundational training modules.
Case scenarios were presented to demonstrate how traditional statistics or machine learning approaches can be used to achieve an analytical objective. Next, the key criteria for choosing between these methodologies were examined, emphasizing the pivotal role of training data and cross-validation techniques as key differentiators between statistics and AI/ML analytical approaches. The strengths of AI/ML models were then underscored, particularly their capacity to capture complex nonlinear relationships, which often result in predictive accuracy beyond what is achieved with traditional statistical methods. We provide our initial training presentation for introducing AI in the biomedical sciences at https://github.com/jshaffer1000/Datathon-training-materials.
Phase 2: Datathon training approach, competencies, and learning objectives
On the first day of the training, trainers provided introductory presentations outlining the datathon and their goals and expectations. Next, the datathon instructors introduced the data sources and a description of the project objectives. Trainees then led activities in hypothesis generation, data analysis, and presentation of findings. Large health data sources from completed research projects were shared with trainees using the Google Drive (Mountain View, California) file-sharing application, and the trainers introduced the data structure and content. Participants were then organized into five groups of three participants. Teams were balanced by data science and clinical expertise to the extent possible, using information provided in the applications for participating in the program. One participant was chosen to represent each group. Next, trainers posed potential research questions that could be studied using the available data resources. The groups then generated research questions and hypotheses through roundtable discussions. Trainers supported the team groups during the hypothesis generation phase and ensured that their topics did not overlap with those from competing teams. Throughout the training period, team leaders presented their progress daily over five days toward testing their research hypotheses to the panel of judges. After the event, participants provided final presentations and reports summarizing their key findings. The competencies for the datathon training phase, along with their corresponding Bloom’s taxonomy levels, learning objectives, and signature assessments, are listed in Table 3.
The learning objectives encompassed competencies focused on idea and hypothesis generation from secondary analyses of multimodal biomedical data sets. At least one signature assessment was typically performed on each day of the training.
Award structure and long-term follow-up
A panel of expert scientific judges evaluated and scored each project based on criteria for scientific quality and methodological rigor using an evaluation rubric. Follow-up meetings were held biannually following the event, with the overarching objective of supporting each group in producing a single scientific manuscript draft within one year following the datathon training activities. Prizes were distributed according to a tiered system, recognizing the first-, second-, and third-highest scored projects.
Results
The inaugural datathon event was held at ACE-B in Bamako, Mali, from May 13 to 24, 2024. There were 92 applicants, of whom 49 were selected to participate in the foundational training. Fifteen participants were selected to attend in-person at the ACE-B training site for the program’s foundational and datathon phases. The remaining 34 selected participants participated remotely in the foundational part of the training. The applicant and selected trainee characteristics are shown in Table 4.
There was a higher representation of male than female applicants (77.2%, 71/92 versus 22.8%, 21/92). However, the proportion of female participants did not significantly differ across the three groups (p = 0.638). Most trainees maintained a master’s degree or were enrolled in a master’s degree program (53.1%, 26/49). All of the foundational participants (49/49) and datathon participants reported at least basic data science expertise. Intermediate training was the most common data science expertise level reported among the hybrid participants in the foundation phase (49%, 24/49). All of the trainees participating in person were from the Central African region (100%, 15/15). There were significant differences across the three comparison groups for the highest degree achieved (p = 0.0003), data science experience (p = 0.020), and profession (p = 0.035).
Pre-datathon foundational training
The foundational training covered advanced computer programming modules for data management and analysis of health outcomes using AI/ML approaches. The three-step pedagogical foundational training approach introduced an analytical topic in the context of traditional statistics. This topic was then explored by comparing AI/ML approaches with traditional statistical methods and highlighting their distinguishing characteristics and strengths. Our case studies for AI/ML regression and imagery analysis are illustrated in Fig. 2.

Source and service layer credits for satellite imagery: Esri, DigitalGlobe, GeoEye, Earthstar, Geographics, CNES/Airbus DS, USDA, USGS, AEX, Getmapping, Aerogrid, IGN, IGP, swisstopo, and the GIS User Community.
Pedagogical training examples comparing traditional and AI/ML analog approaches. AI artificial intelligence, ML Machine learning, RMSE root mean square error. Panel (a): Stepwise demonstration using a relational data set. Step 1 introduced the problem with a single data set. Step 2 applied ordinary linear regression, and Step 3 transitioned to an AI/ML learning workflow by splitting the data for training and evaluation of the regression model. Panel (b): Application to satellite imagery. Step 1 presented the source satellite image. Step 2 employed manual digitization (considered as a traditional statistics approach), and Step 3 demonstrated the automated segmentation of housetops using AI/ML techniques.
The first case study explored the differences between traditional and ML regression techniques. Ordinary linear regression was demonstrated with a single dataset. The distinguishing factor for ML was expressed in terms of splitting data into training and test sets (see Fig. 2a). We then performed cross-validation using the training and test sets to illustrate the ML approach. Both analyses were performed using R (version 4.5.0, R Foundation for Statistical Computing, Vienna, Austria) and Python (version 3.12, Python Software Foundation, Wilmington, Delaware) algorithms to provide participants with experience in both computing interfaces. We provide the R and Python code for generating the sample data sets and performing the data set splitting, regression analyses, and cross-validation in a GitHub repository at https://github.com/jshaffer1000/Datathon-training-materials.
The second case study shifted focus to image analysis. Here, the traditional analytical approach involved manually digitizing satellite imagery. Manual digitization was then contrasted with ML-based approaches, where unsupervised ML classification methods were applied to automate and streamline the analytical processes (see Fig. 2b). The ArcGIS Online (ESRI, Redlands, CA) and QGIS (QGIS Development Team) applications were used to prepare the maps and perform the spatial ML analyses.
Datathon training
Among the 49 participants in the foundational phase, 15 trainees also participated in the in-person datathon training. These participants were divided into five teams of three members, balancing expertise across the groups in computer programming, epidemiology, and clinical fields. Each team nominated and appointed a leader, with female investigators leading two of the five teams (40%). Trainees were acclimated to the data sources through a training segment at the beginning of the in-person phase of the event. The data were obtained from a large malaria cohort study conducted between 2010 and 2017 under the WA-ICEMR program, funded by the NIH.28. The study was a rolling cohort design with 7,708 individuals and 893 households across four sites in three countries. The WA-ICEMR data sources for the datathon training are characterized in Fig. 3.
Overview of datathon data sources, study site characteristics, and data structures. Panel (a): Number of participants by study site (Dangassa and Dioro, Mali; Gambissara, The Gambia; and Thies, Senegal). Panel (b): Passive case detection encounters. Panel (c): Number of enrolled households. Panel (d): Number of entomological encounters. Panel e: Temporal distribution of passive case detection encounters. Panel (f): Hierarchical structure of the data sets. Low, Medium, and High denote the historical levels of P. falciparum endemicity for each study site.
The data sets represented three rural study sites in West Africa (Dangassa, Mali; Dioro, Mali; and Gambissara, The Gambia) and one urban site (Thies, Senegal). The parent study’s overall objective was to compare the study sites regarding the intensity and prevalence of malaria infection, vector mosquito population dynamics, insecticide resistance, and disease pathogenesis. The hierarchical data levels included geographic area (household geocoordinates), household, mother, child, individual, and vector (mosquito). The study data were collected using cross-sectional biannual surveys for active case detection. Passive case detection was performed year-round at local public health units. The primary outcome for both active and passive case detection was the occurrence of Plasmodium (P.) falciparum malaria. The data covered time periods before and after the adoption of several new malaria intervention policies, providing a rich resource for evaluating these interventions. A detailed description of the WA-ICEMR data sources has been described previously11.
Data were curated before implementing the training, and a data warehouse was organized in relational formats as Comma-Separated Value (CSV) files. Data were shared with trainees using the Google Drive file-sharing application. Data dictionaries, collection instruments, and metadata files were shared with trainees in Portable Document Format (PDF).
Final datathon team research topics and long-term follow-up
The final topics put forth for each of the five study teams were drivers of persistent malaria infection, spatiotemporal distribution of malaria during the dry season, temporal and household factors, drivers of clinical malaria in children aged under two years, and asymptomatic P. falciparum carriage in the dry season. Each team concluded the datathon with a final presentation of their results and a completed written report of the findings. Training certificates were awarded to the trainees upon program completion and were formally delivered by USTTB administration. Long-term follow-up was performed by matching each datathon group with a scientific mentor, and weekly teleconference meetings were held to assess progress toward a completed manuscript draft for each team. These meetings were on a rotational basis such that each team presented once every five weeks. Each of the manuscripts was in progress at the time this paper was completed.
Geographic representation and language profile of the training cohort
Most applicants for the training program resided in Mali or Guinea (n = 14 and n = 12 participants in the foundational training for Mali and Guinea, respectively). In total, participants in either the foundational or datathon component of the program represented 14 different nationalities, as illustrated in Fig. 4.
Geographic and linguistic nationality of participants. Panel (a): Geographic distribution of participants, with shading indicating the number of participants from each country. Panel (b): Map showing the primary language of each country represented in the datathon, categorized as English, French, Italian, Portuguese, Spanish, or other languages. Panel (c): Percentages of participants reporting at least some language barriers during the training; responses to this question were unavailable for two participants.
Sixteen of the 28 participants (57.1%) who responded to the question about language barriers reported experiencing them, making it the most frequently cited challenge in our program evaluation (see Fig. 4c). While the primary working language for most participants was French, the datathon training was primarily conducted in English. Our trainers and the datathon team leaders delivered English-to-French translation summaries, lasting approximately 30 min, at the end of each training day.
Discussion
To the best of our knowledge, this work is the first of its kind to describe and evaluate datathon models for data science research education in Africa. The highly interactive training model fosters multidisciplinary team learning among participants from diverse nationalities, emphasizing short, intensive sessions tailored for data science research. Our training approach combined a foundational phase of data science training with an intensive, competitive training component. Our pedagogical approach focused on introducing AI/ML concepts by matching them with analogous traditional statistical methods, which achieved strong participant engagement. We equipped trainees with access to rich, real-world data from historical research projects, which we believe is an ideal use for maximizing the impact of archived biomedical data sources. The program attracted a strong and capable pool of applicants, each bringing at least basic proficiency in data science. While the two-week training period was sufficient for carrying out the foundational and datathon activities, we found the one-year follow-up period fell short for the study groups to produce deliverable manuscripts. Optimizing the long-term training might call for extending the follow-up period beyond one year or considering more practical short-term deliverables, such as presentations at scientific conferences.
In the context of this study, traditional statistical methods have been successfully employed for the WA-ICEMR cohort data1,10. We aimed to leverage familiarity with traditional statistics to introduce ML modeling approaches. Previous analyses of the WA-ICEMR data sources have primarily focused on association studies rather than predictive modeling. Machine learning methods have been shown to provide better predictive estimates in modeling, which can be used independently or in conjunction with traditional statistical methods. The differences between traditional statistics and ML methods were more pronounced for image analysis. For example, we applied ML to analyze satellite imagery, which had considerable advantages over manual approaches with respect to processing time and accuracy.
The in-person phase of our datathon training was designed as a trainee-led instructional experience. These events share similarities with Montessori learning models. While Montessori learning was traditionally used in childhood education, it is increasingly adopted in higher education contexts29. Here, students choose their own work topics and are given uninterrupted work cycles. These Montessori approaches correspond to our datathon teams choosing their own research topics and working intensively on their projects over a fixed time period. Montessori learning is tailored to children’s strengths, which we view as bringing individual, diverse expertise into group settings. Unlike conventional scientific workshops, datathon training approaches are especially conducive to team building, fostering participant collaboration and camaraderie. Most of our trainees informally reported that they found the friendly competition enjoyable and motivating, resulting in high levels of engagement throughout the event. We purposely incorporated multimodal, hierarchical data sources that required thoughtful data linkage to stimulate idea and hypothesis generation across complex data structures. These complexities encouraged participants to explore research questions from geographical, household, individual, or vector perspectives. The approach provided valuable training on data management and linkage. Our program also provided training on linking the WA-ICEMR data sources with external, publicly available sources, as data warehousing skills are fundamental to data science.
A critical gap that was addressed through our datathon training was uniting multidisciplinary and regional teams to organize and analyze data over a short time period. Regional approaches to problem-solving offer an opportunity to unite multidisciplinary groups of collaborators and consider regional health outcomes from diverse perspectives. The datathons fostered networking and communication across multiple countries, which often spawn growth in collaborative research. Applying the datathon model here to other areas in Africa is perhaps most suitable in a university or hospital environment capable of maintaining core infrastructure and teaching computer laboratories. This project benefited from its implementation in a university-based research environment with such capacity. Its generalizability to other settings likely depends on maintaining comparable environments that align trainees around specific health outcomes and encourage them to regard health challenges in other countries as shared responsibilities. It is important that the research education efforts translate to local capacity building. The WA-ICEMR has routinely collaborated with the Mali National Malaria Control Program (NMCP) in capacity building and the delivery of preventive interventions10. These collaborations have bolstered local surveillance systems and reduced duplication in capacity-building efforts. Several of our trainees were also part of the ongoing WA-ICEMR project, allowing them to apply the datathon skills to support the project and its collaborative efforts with the NMCP.
We believe there is a considerable opportunity to use datathons as a platform to promote data sharing. While providing trainees with data sources using file-sharing applications was straightforward, establishing formal data repositories is needed to encourage sharing and integrating local, personal, and external data sources into the training. These repositories should be capable of accommodating historical data sources that are often siloed, unstandardized, or inaccessible. One example is ClinEpiDB, an open-access platform for epidemiological studies13 that will ultimately serve as a repository for the data used here. Such initiatives might also allow for trainee investigator data sources with data-sharing agreements arranged before their use in the training activities.
Because French is widely spoken across West Africa, we carefully considered whether to conduct our training in English or French. As is common in scientific research and publication, scientific research education programs are often delivered in a single language, such as English. Ultimately, we chose English instruction to better align with our goals of fostering research collaboration and developing scientific manuscripts primarily written in English. Train-the-trainer strategies might leverage participants with strong English proficiency to mentor peers using their local languages. Much of our training was interactive and driven by data management, computer programming, and visualization, which likely reduced language barriers. The selected software applications were available in multiple languages, providing additional support for non-English speakers. Most of our participants, however, reported a language barrier during the training. Language barriers are typically among the most common challenges for implementing short-term training programs for international collaborative research. West Africa has thousands of spoken languages, with French, English, and Arabic being the most commonly spoken30. Emerging AI translation technologies have considerable potential for reducing language barriers in research education, and some researchers even suggest that AI may ultimately eliminate the need to learn multiple languages31. With their exponentially increasing availability, implementing these tools in training program settings is becoming increasingly feasible. Freely available tools such as ChatGPT, Google Translate, and Microsoft translation tools are now widely used in classroom settings32. Because data science training is highly interactive, it may be better equipped to address inaccuracies that occur with AI-based real-time translation than less interactive research fields. While our current training program employed routine translation by bilingual investigators, it was impractical to implement in real-time. AI translation tools are now more widely accessible, but to our knowledge, they have not been routinely used or evaluated for data science research education programs. We provide a hypothetical mapping of AI technologies to the steps in our datathon event process in Supplementary File 1, Table S2.
While our female participation rates were consistent with and surpassed global trends, males were more likely to apply to and participate in our training program. Notably, 40% (2/5) of our datathon teams were led by female investigators, highlighting an opportunity to use datathons to foster female research leadership. Identifying historical and current barriers to research participation is needed to reflect the general population’s perspective and promote growth in female mentorship. Only 17% of countries reportedly have a balanced distribution between males and females in research; globally, women account for fewer than 30% of fractionalized authorships33. Providing additional funds for childcare during the datathon training period might be considered to increase female participation. Including female scientific awards within training mechanisms might also enhance female participation. Some examples are the NIH fellowships and fellowships at the United Nations Educational, Scientific, and Cultural Organization34.
The training activities faced several limitations. The main challenges for carrying out the datathon training centered around coordinating travel logistics, particularly those arising from administrative delays and visa processing. Also, some participants were not fully fluent in English, which may have affected engagement and comprehension. Additionally, this study was limited in its capacity to evaluate the program’s long-term impact, which we believe is significant. Measuring the impact of short-term training programs is limited through traditional evaluation surveys, which often fail to capture deeper learning outcomes. To address this limitation, each training group was required to perform research that culminated in a scientific manuscript. The eventual publication of these papers will serve as a more tangible and meaningful indicator of the program’s long-term success.
Conclusions
Datathon training models are rapidly emerging and powerful tools for advancing multidisciplinary data science research education. The intensive, team-oriented, and short-term nature of datathon training makes it an ideal complement to research programs. Harnessing historical data sources and maximizing their use through datathon training fosters research collaboration and promotes data sharing. Scientific literature that explores novel, scalable training approaches plays a crucial role in shaping the development of data science expertise in Africa.
Data availability
Data sources referenced in this paper are detailed within the main article and supplementary information. The R and Python code used for package installation, data splitting, and linear regression analyses is publicly available in a GitHub repository at https://github.com/jshaffer1000/Datathon-training-materials. This repository also includes an introductory artificial intelligence training module and will continue to feature additional materials and updates from ongoing datathon training activities. The WA-ICEMR data set used for the datathon training will ultimately be accessible through ClinEpiDB at https://clinepidb.org.
Abbreviations
- ACE:
-
African centers of excellence in bioinformatics and data-intensive science
- ACE-B:
-
African center of excellence in bioinformatics and data-intensive science of Bamako, Mali
- AI:
-
Artificial intelligence
- AI/ML:
-
Artificial intelligence and machine learning
- COVID-19:
-
Coronavirus disease 2019
- CSV:
-
Comma-separated value
- DS-I Africa:
-
Data science for health discovery and innovation in Africa
- GIS:
-
Geographic information systems
- H3Africa:
-
Human heredity and health in Africa
- ML:
-
Machine learning
- NIAID:
-
National institute of allergy and infectious diseases
- NIH:
-
National institutes of health
- PDF:
-
Portable document format
- QGIS:
-
Quantum geographic information systems
- REDCap:
-
Research electronic data capture
- USTTB:
-
University of sciences, techniques and technologies of Bamako, Mali
- WACE-DSRE:
-
West Africa center of excellence for data science research education and training program
- WA-ICEMR:
-
West Africa international centers of excellence for malaria Research
- UGANC:
-
Gamal abdel nasser university of Conakry
References
Doumbia, S. et al. A decade of progress accelerating malaria control in Mali: Evidence from the West Africa International Center of Excellence for Malaria Research. Am. J. Trop. Med. Hyg. 107, 75–83. https://doi.org/10.4269/ajtmh.21-1309 (2022).
Gao, Y., Sharma, T. & Cui, Y. Addressing the challenge of biomedical data inequality: An artificial intelligence perspective. Annu. Rev. Biomed. Data Sci. 6, 153–171. https://doi.org/10.1146/annurev-biodatasci-020722-020704 (2023).
Adebamowo, C. A. et al. The promise of data science for health research in Africa. Nat. Commun. 14, 6084. https://doi.org/10.1038/s41467-023-41809-2 (2023).
Shaffer, J. G. et al. Expanding research capacity in Sub-Saharan Africa through informatics, bioinformatics, and data science training programs in Mali. Front. Genet. 10, 331. https://doi.org/10.3389/fgene.2019.00331 (2019).
DS-I Africa: Data Science for Health Discovery and Innovation in Africa. West Africa Center of Excellence for Data Science Research Education (WACE-DSRE). https://dsi-africa.org/project/36 (2025).
National Institutes of Health. Harnessing Data Science for Health Discovery and Innovation in Africa (DS-I Africa). https://commonfund.nih.gov/harnessing-data-science-health-discovery-and-innovation-africa-ds-i-africa (2024).
Human Heredity and Health in Africa. West African Center of Excellence for Global Health Bioinformatics Research Training. https://h3africa.org/index.php/west-african-center-of-excellence-for-global-health-bioinformatics-research-training/ (2025).
African Center of Excellence in Bioinformatics & Data Science (ACE-B). https://aceb-mali.org/ (2025).
National Institute of Allergy and Infectious Diseases. West Africa International Center of Excellence for Malaria Research. https://www.niaid.nih.gov/research/west-africa-icemr.
Doumbia, S. et al. The West Africa ICEMR partnerships for guiding policy to improve the malaria prevention and control. Am. J. Trop. Med. Hyg. 107, 84–89. https://doi.org/10.4269/ajtmh.21-1330 (2022).
Shaffer, J. G. et al. Development of a data collection and management system in West Africa: Challenges and sustainability. Infect. Dis. Poverty 7, 125. https://doi.org/10.1186/s40249-018-0494-4 (2018).
National Institute of Allergy and Infectious Diseases. ICEMR Program Overview. https://www.niaid.nih.gov/research/icemr-program-overview.
Ruhamyankaka, E. et al. ClinEpiDB: an open-access clinical epidemiology database resource encouraging online exploration of complex studies. Gates Open Res 3, 1661. https://doi.org/10.12688/gatesopenres.13087.2 (2019).
Irani, L. Hackathons and the making of entrepreneurial citizenship. Sci. Technol. Hum. Values 40, 799–824. https://doi.org/10.1177/0162243915578486 (2015).
Richterich, A. Hacking events: Project development practices and technology use at hackathons. Convergence 25, 1000–1026. https://doi.org/10.1177/1354856517709405 (2019).
Aboab, J. et al. A “datathon” model to support cross-disciplinary collaboration. Sci. Transl. Med. 8, 333ps338. https://doi.org/10.1126/scitranslmed.aad9072 (2016).
Moseley, E. T., Hsu, D. J., Stone, D. J. & Celi, L. A. Beyond open big data: Addressing unreliable research. J. Med. Internet Res. 16, e259. https://doi.org/10.2196/jmir.3871 (2014).
Merriam-Webster. hackathon. https://www.merriam-webster.com/dictionary/hackathon (2022).
Celi, L. A. et al. Datathons and software to promote reproducible research. J. Med. Internet Res. 18, e230. https://doi.org/10.2196/jmir.6365 (2016).
Braune, K. et al. Interdisciplinary online hackathons as an approach to combat the COVID-19 pandemic: Case study. J. Med. Internet Res. 23, e25283. https://doi.org/10.2196/25283 (2021).
Hex Technologies, Inc. Planning a Modern Datathon. https://hex.tech/blog/the-modern-datathon/ (2022).
Piza, F. M. T. et al. Assessing team effectiveness and affective learning in a datathon. Int. J. Med. Inform. 112, 40–44. https://doi.org/10.1016/j.ijmedinf.2018.01.005 (2018).
Luo, E. M. et al. MIT COVID-19 datathon: Data without boundaries. BMJ Innov. 7, 231–234. https://doi.org/10.1136/bmjinnov-2020-000492 (2021).
MIT COVID19 Challenge. MIT COVID19 Challenge. https://covid19challenge.mit.edu/.
Das, P. et al. Letter: Harnessing big data: The need for datathon research in neurosurgery. Neurosurgery 86, E402. https://doi.org/10.1093/neuros/nyz534 (2020).
Saporta, G. Training data scientists: A few challenges. Int. J. Data Sci. Anal. 6, 201–204. https://doi.org/10.1007/s41060-018-0114-1 (2018).
Anderson, L. K. DR (Addison Wesley Longman Inc, 2001).
National Institute of Allergy and Infectious Diseases. West Africa International Center of Excellence for Malaria Research,. https://www.niaid.nih.gov/research/west-africa-icemr.
Inside Higher Ed. Using Montessori Tactics in College Classes. https://www.insidehighered.com/opinion/career-advice/teaching/2023/07/05/benefits-using-montessori-principles-college-classes. (2023).
Provisio. Official Languages in Africa – An Analysis. https://provisioservices.com/languages-in-africa/ (2024).
Matsakis L. in The Atlantic (2024).
Microsoft. Microsoft Translator for Education. https://www.microsoft.com/en-us/translator/education/ (2024).
Larivière, V., Ni, C., Gingras, Y., Cronin, B. & Sugimoto, C. R. Bibliometrics: Global gender disparities in science. Nature 504, 211–213. https://doi.org/10.1038/504211a (2013).
United Nations Educational Scientific and Cultural Organization. Organization for Women in Science for the Developing World. https://owsd.net/.
Acknowledgements
The authors are grateful for the financial support for our training programs from the Fogarty International Center of the NIH. We thank the facilitators at the African Centers of Excellence in Bioinformatics and Data Intensive Science (ACE) program for developing and providing the teaching computer laboratories that support our research education programs. We are grateful to the administration at USTTB for their strong support and encouragement. Finally, we thank the datathon trainees for their participation and contributions.
Funding
This study was supported by the Fogarty International Center of the NIH under award numbers UE5 TW012526 and U2R TW010673. Trainee data were obtained from the West Africa International Center of Excellence for Malaria Research, supported by the NIH National Institute of Allergy and Infectious Diseases (NIAID) under grant number U19 AI089696 (2010–2017). The funders had no role in study design, data collection and analysis, decision to publish, or manuscript preparation.
Author information
Authors and Affiliations
Contributions
Conceptualization: SD, JGS, MW, COT, FK, OD, CC; Methodology: SD, JGS, JL, COT, FK, OD, CC; Software: JGS, JL, COT, FK, CC, NS; Validation: SD, JGS, MW, COT, FK, OD, CC; Formal analysis: SD, JGS, MW, COT, FK, OD, CC; Investigation: SD, JGS, MW, COT, FK, OD, CC, MK, NS; Resources: SD, JGS, MW, COT, AD, AHB; Data curation: JGS, FK, OD, CC; Writing—original draft: SD, JGS, MW, COT, FK, OD, CC; Writing—review and editing: All authors; Visualization: JGS, JL, COT, FK, OD, CC; Supervision: SD, JGS, MW, COT, FK, OD, CC, MK, NS, MD, MT; Project administration: COT; Funding acquisition: SD, JGS. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Ethical approval
Experimental protocols for all of the data sources used in the datathon projects were reviewed and approved by the Institutional Review Boards (IRBs) of the USTTB (FWA00001769), University Cheikh Anta Diop in Dakar, Senegal (FWA00002691), Medical Research Council Unit in The Gambia (FWA00006873), and Tulane University (FWA00002055) before participant enrollment in the longitudinal cohort study. Informed consent for the use and publication of anonymized cohort data was obtained from all participants at the time of enrollment. The USTTB Ethics Committee determined that analyses of aggregated datathon evaluation results were exempt from additional ethics review.
Competing interests
The authors declare that they have no competing interests. Figure 1 was created using BioRender.com.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Doumbia, S., Kane, F., Diabate, O. et al. Advancing data science research education in Africa through datathon-driven innovations. Sci Rep 16, 11527 (2026). https://doi.org/10.1038/s41598-026-41474-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-41474-7