Abstract
Retinoic acid (RA) signaling through CRABP2 and FABP5 differentially influences therapeutic responses and drug resistance in anaplastic thyroid cancer (ATC), yet regulatory mechanisms remain unclear. Our study reveals low CRABP2 and high FABP5 expression in ATC cell lines (THJ-11T/16T/21T), with RA alone showing no growth inhibition but RA/gemcitabine (GEM) combination exhibiting synergistic effects. ZEB1 transcriptionally regulates both proteins, where RA/GEM treatment alters its promoter binding pattern—reducing FABP5 association while enhancing CRABP2 binding. Co-regulators KAT2B and EP300 compete for ZEB1’s CP domain during transcriptional regulation. Notably, RA/GEM modulates m6A modifications: suppressing CRABP2 and enhancing FABP5 methylation through METTL3/ALKBH5 interactions, thereby reducing mRNA stability. Clinical analysis of 286 thyroid cancer specimens confirmed variable CRABP2/FABP5 expression. Crucially, ZEB1 and ALKBH5 overexpression potentiated GEM/RA efficacy against ATCs. These findings identify m6A-mediated post-transcriptional regulation and ZEB1-driven transcriptional dynamics as key determinants of the combined effects of GEM and RA in ATCs, providing new targets for combination therapy optimization.
Similar content being viewed by others
Introduction
Thyroid cancer is the most prevalent endocrine malignancy and is responsible for the majority of deaths related to endocrine cancers1,2. Most thyroid cancers, including papillary and follicular types, arise from follicular epithelium with a well-differentiated phenotype, resulting in a more favorable prognosis following thyroidectomy2,3. However, Surveillance, Epidemiology, and End Results (SEER) data published in April 2021 reported a 5-year survival rate of only 7% for patients with anaplastic thyroid cancer (ATC) across all SEER stages, accounting for ~40% of all thyroid cancer-related deaths4. To prevent tumor recurrence, postoperative treatments such as radiotherapy, chemotherapy, or their combination have been used. Additionally, the cell differentiation enhancer all-trans retinoic acid (RA) has been used to increase radiosensitivity in ATC patients5. Clinical data indicate that these treatment options yield varying outcomes5,6. For example, RA-induced redifferentiation has been shown to inhibit the invasive behavior of thyroid cancer7. Gemcitabine (GEM)-loaded liposomes are significantly more cytotoxic to ATC cells than the free drugs are, and RA has been shown to be safe and tolerable when used with GEM as a matrix-targeting agent to increase the intensity of chemotherapy8,9.However, RA therapy is insufficient in patients with metastatic thyroid cancer10. Given the clinical application of RA-based redifferentiation therapy, it is crucial to thoroughly assess its suitability for treating ATC thoroughly.
Owing to their tendency to exhibit growth arrest and sensitivity to anticancer therapies11, cancer cells undergoing redifferentiation have been utilized to overcome resistance to radioactive iodine in cancer treatment12. However, as noted above, treatment outcomes vary considerably for reasons that remain unknown. RA signaling is transduced through two classical pathways mediated by cellular retinoic acid binding protein 2 (CRABP2) or fatty acid binding protein 5 (FABP5), resulting in different cellular responses13. CRABP2 transports RA to the nuclear retinoid receptor bound to the retinoic acid X receptor and subsequently binds to regulatory regions of target genes involved in apoptosis, differentiation and cell cycle arrest14. Conversely, RA signaling through the FABP5/peroxisome proliferator-activated receptor (PPAR) pathway promotes cancer cell survival by activating cancer-associated genes15. In this context, the expression status of CRABP2 and FABP5 is critical in determining the fate of cancer cells treated with RA. Alternatively, an imbalance in the expression patterns of CRABP2 and FABP5 may lead to different treatment outcomes. However, there are currently no reports on the in vitro or in vivo expression patterns of these two RA-related factors in ATCs.
In recent years, transcriptional regulation and N6-methyladenosine (m6A) modification have received particular attention because of their critical roles in cancer, particularly their ability to influence tissue homeostasis and tumor progression by affecting mRNA expression levels16,17. Studies from our laboratory have shown that transcription factors and m6A can concurrently regulate the expression of one gene: the transcription factor homeobox A13 (HOXA13) promotes the expression of solute carrier family 3 member 2 (SLC3A2), an essential antioxidant component of the tumor cell membrane, whereas m6A modification mediated by YTH N6-methyladenosine RNA binding protein C2 (YTHDC2) leads to the degradation of HOXA13 mRNA, thereby inhibiting SLC3A2 expression18,19. In addition, research by other teams has also confirmed that transcription factors and m6A can coregulate the same gene. For example, increased m6A levels mediate the upregulation of the transcription factor nuclear factor I/B (NFIB), which in turn promotes the transcription of epithelial-to-mesenchymal transition (EMT)-related factors leading to metastasis20. However, there is currently limited research on the effects of transcriptional regulation and m6A modification on CRABP2 and FABP5. Clarifying the molecular mechanisms underlying the regulation of CRABP2 and FABP5 could guide improvements in RA related therapy for thyroid cancer.
The current study aimed to measure and manipulate the expression levels of CRABP2 and FABP5 in RA-resistant ATC cell lines through various experimental methods to identify the underlying reasons for the lack of response to RA treatment in ATC cells and how GEM may alter this response. Additionally, we explored the molecular mechanisms by which RA and GEM affect ATCs, focusing on the transcriptional and m6A regulation of CRABP2 and FABP5. Immunohistochemical staining based on tissue microarrays was used to analyze the expression patterns of CRABP2 and FABP5 in cancerous and noncancerous thyroid surgical samples and assess their potential correlation with RA sensitivity.
Results
Imbalanced CRABP2 and FABP5 levels in ATC cells
CRABP2 and FABP5 expression varies across various ATC cell lines. Here, we selected six ATC cell lines, namely 8505 C, Cal62, HTH7, THJ-11T, THJ-16T, and THJ-21T, and assessed the protein levels of CRABP2 and FABP5. We found that, compared with the other three cell lines, THJ-11T, THJ-16T, and THJ-21T had lower levels of CRABP2 and higher levels of FABP5 (Fig. 1A), and lower sensitivity to RA treatment (Fig. 1B and Supplementary Fig. 1A). ICC and immunofluorescence (IF) staining revealed that CRABP2 expression was very low (faintly positive) in THJ-11T, THJ-16T, and THJ-21T cells, whereas FABP5 expression was distinctly high (strongly positive) in these ATC cell lines (Fig. 1C, D). In agreement with the above ICC and IF staining findings, quantitative real-time polymerase chain reaction (qRT-PCR) and western blot analyses revealed that CRABP2 was expressed at a low level in THJ-11T cells and nearly undetectable in THJ-16T and THJ-21T cells; whereas FABP5 was highly expressed in these three cell lines (Fig. 1E, F). Collectively, these data suggested that in THJ-11T, THJ-16T, and THJ-21T cells, CRABP2 levels were low, whereas FABP5 levels were high, and that RA treatment did not affect the expression of CRABP2 or FABP5.
A WB analysis of CRABP2 and FABP5 expression in the 8505 C, Cal62, HTH7, THJ-11T, THJ-16T and THJ-21T cell lines. B CCK8 assays were performed on 8505 C, Cal62, HTH7, THJ-11T, THJ-16T, and THJ-21T cells cultured normally (N) or treated with 0.2% DMSO or 10 μM RA for 48 h (RA). C Immunocytochemical staining of CRABP2 and FABP5 in the three cell lines with no treatment (N) or with 48 h of 10 μM RA treatment (RA). D Immunofluorescence staining of CRABP2 and FABP5 in the three cell lines with no treatment (N) or with 48 h of 10 μM RA treatment (RA). Green, CRABP2; red, FABP5. Merge, overlapping green and red staining. qRT-PCR (E) and WB analyses (F) of CRABP2 and FABP5 expression in THJ-11T, THJ-16T, and THJ-21T cells with no treatment (N) or with 10 μM RA treatment for 48 h (RA). The molecular weights of CRABP2 and FABP5 were 16 and 15 kDa, respectively. Band density was quantified by ImageJ and normalized to the level of GAPDH as the loading control. N.S., nonsignificant. Scale bar, 5 mm. The data in A and B were analyzed by one-way ANOVA. The data in (C–F) were analyzed by t tests. Images from WB, ICC staining and IF staining experiments are representative of three independent experiments. *p < 0.05, **p < 0.01 indicate statistical significance.
RA resistance of ATC cell lines
We observed that RA could not significantly regulate the activity of caspase 3/7 or the growth of THJ-11T, THJ-16T, and THJ-21T cells (Supplementary Fig. 1B–D). The results from 3-dimensional (3D) spheroid cultures also showed that RA treatment could not induce additional cell death in the cell spheroids (Supplementary Fig. 1E, F). We then constructed THJ-16T-based nude mouse xenograft models with Dimethyl sulfoxide (DMSO) or RA treatment. On Day 18, RA (dose: 1.5 μg/kg) was administered once daily for next 18 days. We found that compared with the control treatment, the RA treatment did not influence tumor growth (Supplementary Fig. 1G, H). These data suggested that RA cannot independently inhibit the growth of THJ-11T, THJ-16T, and THJ-21T cells.
GEM treatment increases CRABP2 expression and RA sensitivity
We further determined the IC50 values of GEM in the aforementioned ATC cell lines and found that, consistent with the response to RA treatment, THJ-11T, THJ-16T, and THJ-21T cells presented lower IC50 values than 8505 C, Cal62, and HTH7 cells did (Supplementary Fig. 2). Since the combination of GEM and RA is more effective than RA alone21,22, we used a combination of RA and GEM to treat THJ-11T, THJ-16T, and THJ-21T cells. CCK8 assays revealed that, in comparison with negative control and RA-treated cells, both GEM- and GEM + RA- treated cells exhibited significant growth inhibition after 48 h of treatment (Fig. 2A, B). ICC and IF staining revealed that CRABP2 expression increased, whereas FABP5 expression decreased after GEM treatment, and these trends were further strengthened after treatment with GEM in combination with RA (Fig. 2C, D). The results from 3D spheroid cultures also revealed that GEM could induce cell death in THJ-16T cell spheroids, and that RA could further increase the ability of GEM to induce cell death (Fig. 2E). We observed that GEM could induce the activity of caspase 3/7 in THJ-11T, THJ-16T, and THJ-21T cells, and the effect of the combined GEM and RA treatment was more significant (Fig. 2F). We then constructed THJ-16T-based nude mouse xenograft models with the following treatments: Group 1: DMSO treatment; Group 2: RA treatment; Group 3: GEM treatment; and Group 4: combined GEM and RA treatment. Each animal received a daily intraperitoneal dose of RA (1.5 μg/kg), GEM (100 mg/kg), or RA (1.5 μg /kg) plus GEM administered 4 h later (100 mg/kg) for 18 days starting on Day 18. Compared with the control group (Group 1), the RA treatment group (Group 2) did not exhibit inhibited tumor growth, whereas the GEM treatment group (Group 3) presented significantly inhibited tumor growth. Compared with Group 3, the combined treatment with GEM and RA (Group 4) resulted in a significant reduction in tumor volume (Fig. 2G, H). These data indicated that RA promotes the tumor inhibitory role of GEM in THJ-11T, THJ-16T, and THJ-21T cells.
A CCK8 assays were performed on THJ-11T, THJ-16T, and THJ-21T cells cultured normally (N) or treated with 0.2% DMSO, 10 μM RA for 48 h (RA), 5 μM gemcitabine for 48 h (GEM), or 5 μM gemcitabine and 10 μM RA for 48 h (GEM + RA). B Morphological findings of the three cell lines after the same treatment as described in A. Insets, normally cultured cells. Scale bar, 5 mm. C Immunocytochemical staining of CRABP2 in the three cell lines cells after the same treatment as described in A. Insets, normally cultured cells. Scale bar, 5 mm. D Immunofluorescence staining of CRABP2 and FABP5 in the three cell lines cells after the same treatment as described in A. CRABP2 and FABP5 proteins are green and red, respectively. Scale bar, 5 mm. E 3D spheroids were generated with THJ-16T cells. After the formation of 3D spheroids, cell death was measured by staining with SYTOX green following the treatments as described in A. Representative images and graphed data are shown; scale bar, 50 μm. F Relative Caspase 3/7 activity was measured in the three cell lines cells after the same treatment as described in (A). G, H THJ-16T cell-based CDX mouse models. Representative images of tumors at the end of the experiments, in which the mouse were administered DMSO, RA (1.5 μg/kg), GEM (100 mg/kg), or RA (1.5 μg/kg) plus GEM 4 h later (100 mg/kg) for 18 days starting on Day 18 (n = 5/group) are shown in panel G, and the tumor growth curves are shown in panel H. N.S., nonsignificant. The data in (A, C, D and F) were analyzed by one-way ANOVA. The data in H were analyzed by two-way ANOVA. Images of the HE staining, ICC staining, IF staining and 3D spheroids are representative of three independent experiments. *p < 0.05, **p < 0.01 indicate statistical significance.
Combined GEM and RA treatment induces a shift in Zinc Finger E-Box Binding Homeobox 1 (ZEB1) binding from the FABP5 promoter to the CRABP2 promoter
We further explored the regulatory mechanisms of CRABP2 and FABP5 transcription. First, using the LASAGNA-Search database (http://biogrid-lasagna.engr. uconn.edu/lasagna_search/), we found that the transcription factors ZEB1 and Sp1 transcription factor (SP-1) are able to bind to the promoters of CRABP2 and FABP5 simultaneously (Fig. 3A). Chromatin immunoprecipitation (ChIP)- polymerase chain reaction (PCR) experiments confirmed that ZEB1 and SP-1 could bind to the CRABP2 and FABP5 promoters and that the binding of ZEB1 and SP1 to the CRABP2 promoter was increased by GEM treatment and further increased by the combined use of GEM and RA, whereas the binding of ZEB1 and SP1 to the FABP5 promoter showed the opposite trend. In addition, the binding of ZEB1 to the target gene was more obviously regulated by GEM and RA than the binding of by SP-1 was, and thus, we focused mainly on the regulatory effect of ZEB1 on CRABP2 and FABP5 in the subsequent experiments (Fig. 3B). The electrophoretic mobility shift assay (EMSA) results further demonstrated that the binding of ZEB1 to CRABP2 was increased by GEM treatment and further increased by the combined use of GEM and RA, whereas the binding of FABP5 to ZEB1 showed the opposite trend (Fig. 3C). The luciferase activities of the WT CRABP2 and FABP5 promoters could be regulated by GEM and GEM combined with RA, but when the binding site of ZEB1 was mutated, the luciferase activities of the two promoters were no longer regulated (Supplementary Fig. 3A). A previous study revealed that Yes1 Associated Transcriptional Regulator (YAP) can increase the transcription of ZEB123. Here, we found that knocking out YAP inhibited the binding of ZEB1 to the CRABP2 promoter, whereas overexpressing YAP increased the binding of ZEB1 to the FABP5 promoter (Fig. 3D). Transcriptional activation is regulated by histone modifications, such as methylation and acetylation24,25. We used the STRING database (https://cn.string-db.org,version 11.5; confidence score ≥ 0.4; medium confidence) to predict the interactions between ZEB1 and key enzymes of histone modifications, including methylation and acetylation. The results showed that ZEB1 is most likely to interact with E1A binding protein P300 (EP300) and lysine acetyltransferase 2B (KAT2B) (Fig. 3E). In addition, ChIP experiments revealed increased enrichment of EP300 and KAT2B with increased levels of the hallmark of open chromatin related to EP300: H3K27Ac, as well as the hallmark of open chromatin related to KAT2B: H3K9Ac and H3K122Ac, within the region around the ZEB1 motif of the CRABP2 promoter, which started at 4 h after GEM + RA treatment. In contrast, we observed a reduced enrichment of EP300 and KAT2B with decreased levels of H3K27Ac, H3K9Ac and H3K122Ac within the region around the ZEB1 motif of the FABP5 promoter, which started at 3 h after GEM + RA treatment (Fig. 3F). The signals from the IgG control were weak and did not depend on GEM + RA processing of the CRABP2 and FABP5 promoters; in addition, the “−2k” or “+2k” regions around the CRABP2 and FABP5 promoters presented low recruitment (Supplementary Fig. 3B), supporting the specificity data from the “ZEB1” region. These results confirmed that ZEB1 can regulate the transcription of CRABP2 and FABP5 and that combined treatment with GEM and RA can lead to a decrease in binding between the ZEB1 and FABP5 promoters, with ZEB1 instead binding to the CRABP2 promoter.
A The LASAGNA database was used to predict interactions between the transcription factors of CRABP2 and FABP5 with moderate confidence. B ChIP-PCR experiment showing the binding of ZEB1 or SP-1 to CRABP2/FABP5 in cells treated with or without RA or GEM as indicated for 48 h. C EMSA were used to assess DNA-protein interactions in THJ-16T cell nuclear extracts incubated with the ZEB1-CRABP2 probe and ZEB1-FABP5 probe from cells pretreated with or without RA or GEM as indicated for 48 h. D A ChIP-PCR experiment revealed the binding of CRABP2/FABP5 to ZEB1 in WT or YAP-/- THJ-16T cells. E Analysis of the interaction between ZEB1 and histone methylation/acetylation enzymes with the STRING database. F ChIP experiments revealed modifications of H3K27Ac, H3K9Ac, and H3K122Ac, as well as enrichment of KAT2B and ZEB1, in designated regions around the CRABP2 and FABP5 promoters in THJ-16T cells treated with GEM + RA for a specified period of time. The data are shown as the means ± SD from three independent experiments. Images from the ChIP and EMSA experiments are representative of three independent experiments. *p < 0.05, **p < 0.01 indicate statistical significance.
The central part (CP) domain of ZEB1 is crucial for the binding of KAT2B and EP300 to ZEB1
We further explored the molecular mechanisms by which KAT2B and EP300 bind to ZEB1. ZEB1 contains three key domains: an N-terminal zinc-finger cluster (NZF), a CP, and a C-terminal zinc-finger cluster (CZF)23. We found that when the CP domain was deleted, KAT2B and EP300 no longer bound to ZEB1, whereas deleting the two ZF domains separately or simultaneously did not affect the binding of KAT2B and EP300 to ZEB1 (Fig. 4A, B). The results of the proximity ligation assay (PLA) and IF experiments further demonstrated the requirement of the CP domain for the binding of KAT2B and EP300 to ZEB1 (Fig. 4C, D). Coimmunoprecipitation (Co-IP) experiments demonstrated that the overexpression of EP300 inhibited the binding between ZEB1 and KAT2B (Fig. 4E). These results indicated that KAT2B and EP300 competitively bind to the CP domain of ZEB1.
A Co-IP assay with anti-Myc antibodies in ZEB1-FLAG- and WT-KAT2B-Myc- overexpressing THJ-16T cells followed by IB. B Co-IP assay with anti-Myc antibodies in ZEB1-FLAG- and WT-EP300-Myc- overexpressing THJ-16T cells followed by IB. C, D PLA and IF staining experiments were used to assess the binding requirements of the CP domain in ZEB1 to KAT2B and EP300. Scale bar, 50 µm. E Co-IP experiments demonstrated that the influence of EP300 overexpression on the interaction between ZEB1 and KAT2B. The data are shown as the means ± SD from three independent experiments. Images from the IB, PLA and IF staining experiments are representative of three independent experiments. *p < 0.05, **p < 0.01 indicate statistical significance.
m6A modification decreases the mRNA stability of CRABP2 and FABP5
Next, we investigated whether the m6A modifications of CRABP2 and FABP5 are influenced by GEM and RA. First, we found that GEM inhibited the m6A modification of CRABP2 and promoted the m6A modification of FABP5 and that these effects were further increased by the combination of GEM and RA (Fig. 5A). Both CRABP2 and FABP5 interacted with m6A METTL3 and the demethylase AlkB homolog 5 (ALKBH5) (Fig. 5B). GEM inhibited the binding of METTL3 and CRABP2 mRNAs but promoted the binding of ALKBH5 and CRABP2 mRNAs. These trends were further promoted by the combined use of GEM and RA. However, the binding of FABP5 mRNA to METTL3 and ALKBH5 showed the opposite trend (Fig. 5C). METTL3 overexpression inhibited the expression of CRABP2 and FABP5 and decreased their stability, whereas ALKBH5 upregulated the expression of CRABP2 and FABP5 and increased their stability (Fig. 5D, E). Because m6A modifications are very abundant in the 3’UTR and are likely to affect RNA stability25, we next used SRAMP online software to predict the most likely m6A modification sites in the 3’UTR regions of CRABP2 and FABP5. The results revealed that the 895–899 bp region, the 918–922 bp region of CRABP2, and the 487-491 bp region of FABP5 were most likely to undergo m6A modification (Fig. 5F). We constructed mutants of m6A modification sites in the 3’UTR regions of CRABP2 and FABP3 (Fig. 5G), and found that the two predicted modification sites of CRABP2 might both be modified, and only when both sites were mutated was there no m6A modification in the CRABP2 3’UTR; the predicted FABP5 site might also be modified, and mutation of this site abolished the m6A modification of the FABP5 3’UTR (Fig. 5H). These results indicated that CRABP2 and FABP5 simultaneously undergo m6A modification mediated by METTL3 and removal by ALKBH5. The m6A modification of CRABP2 can be inhibited by the combined use of GEM and RA, while the m6A modification of FABP5 can be promoted by the combined use of GEM and RA, which are modifications that inhibit the stability and expression of mRNAs. We also revealed the specific m6A modification sites.
A RIP experiments were conducted on THJ-16T cells treated with 5 μM GEM and 10 μM RA for 48 h with anti-m6A antibodies and IgG. B RIP experiments were conducted on THJ-16T cells with anti-METTL3, anti-METTL14, anti-FTO, and anti-ALKBH5 antibodies to investigate their interaction with FABP5 or CRABP2. C RIP experiments were conducted on THJ-16T cells treated with 5 μM GEM and 10 μM RA for 48 h with anti-METTL3 andanti-ALKBH5 antibodies to investigate their interaction with FABP5 or CRABP2. D qRT-PCR experiments on THJ-16T cells overexpressing METTL3, METTL14, FTO or ALKBH5. E CRABP2 and FABP5 mRNA stability was measured in THJ-16T cells overexpressing ALKBH5 and METTL3 after ActD treatment for the indicated times. F SRAMP online software for predicting m6A sites in the 3’UTRs of CRABP2 and FABP5. G Mutants with m6A modification sites in the 3’UTRs of CRABP2 and FABP5 were presented. Red fonts represented the mutant bases. H WT-CRABP2-3’UTR, Mut1-CRABP2-3’UTR, Mut2-CRABP2-3’UTR, Mut1+Mut2-CRABP2-3’UTR, WT-FABP5-3′UTR, or Mut-FABP5-3’UTR were overexpressed in CRABP2- or FABP5-knockout THJ-16T cells. The enrichment of m6A in the CRABP2 or FABP5 3′UTR was calculated as the percentage of the input RNA via RNA-IP with the m6A antibodies. N.S., nonsignificant. The data are shown as the means ± SD from three independent experiments. *p < 0.05, **p < 0.01 indicate statistical significance.
Analysis of specific domains of involved in binding among CRABP2 and FABP5 mRNAs and METTL3 and ALKBH5 proteins
We further studied the binding among CRABP2 mRNA, FABP5 mRNA, the METTL3 protein and the ALKBH5 protein. First, we used the catRAPID database (http://s.tartaglialab.com/page/catrapid_group) to predict the key regions that might be related to binding, and the key regions are listed in Table 1. We combined the closer regions predicted to have binding potential into a longer region, including four segments (P#1 to P#4) on the METTL3 proteins, four segments (P#1 to P#4) on the ALKBH5 proteins, three segments (R#1 to R#3) on CRABP2 mRNA and three segments (R#1 to R#3) on the FABP5 mRNA, and designed the corresponding mutants (Table 1). RNA pulldown experiments revealed that the R#2 region of CRABP2 was essential for its binding to METTL3, whereas its R#3 region was essential for its binding to ALKBH5; the R#2 region of FABP5 was essential for its binding to both METTL3 and ALKBH5 (Fig. 6A). In addition, both RNA immunoprecipitation (RIP) and photoactivatable ribonucleoside-enhanced crosslinking and immunoprecipitation (PAR-CLIP) experiments revealed that the P#3 domain of METTL3 and the P#1 domain of ALKBH5 were essential for their binding to CRABP2 mRNA and that the P#1 domain of METTL3 and the P#2 domain of ALKBH5 were essential for their binding to FABP5 mRNA (Fig. 6B, C).
A RNA pulldown experiments were used to investigate the specific structural domains responsible for the binding of the CRABP2 (R#1, R#2, R#3) and FABP5 (R#1, R#2, R#3) mRNAs binding to the METTL3 and ALKBH5 proteins. B RIP experiments were performed to investigate the specific structural domains responsible for the binding of the METTL3 (P#1, P#2, P#3, and P#4) and ALKBH5 (P#1, P#2, P#3, and P#4) proteins to the CRABP2 and FABP5 mRNAs. C PAR-CLIP experiments were used to investigate the specific structural domains responsible for the binding of the METTL3 (P#1 and P#3) and ALKBH5 (P#1 and P#2) proteins to the CRABP2 and FABP5 mRNAs. The data are shown as the means ± SD from three independent experiments. Images from the WB experiments are representative of three independent experiments.
Concurrent ZEB1 and ALKBH5 expression increases the sensitivity of ATC cells to GEM and RA
ZEB1 and ALKBH5 led to downregulation of CRABP2 and upregulation of FABP5 after GEM and RA treatment, and deletion of the CP domains of ZEB1, which bound to KAT2B and EP300, and the P#2 domain of ALKBH5, which bound to FABP5, reduced this tendency, which was increased when ZEB1 and ALKBH5 were simultaneously overexpressed (Fig. 7A, B). Therefore, we hypothesized that ZEB1 and ALKBH5 may increase the sensitivity of ATC to GEM and RA treatment. Indeed, the inhibitory effect of GEM and RA on tumor growth was more pronounced after ZEB1 was overexpressed; whereas deletion of the CP domain did not increase the inhibitory effect of GEM and RA on tumor growth (Fig. 7C). Similarly, overexpression of ALKBH5 resulted in a more pronounced inhibitory effect of GEM and RA on tumor growth; and when the P#2 domain was deleted, the inhibitory effect of GEM and RA on tumor growth was no longer increased. We also found that the overexpression of ALKBH5 and ZEB1 further increased the inhibitory effects of GEM and RA on tumor growth (Fig. 7D). Next, we also confirmed that overexpression of ZEB1 and ALKBH5 could increase the activity of caspase 3/7 after GEM and RA treatment, and that GEM and RA treatment more significantly inhibited soft agar colony formation, but deletion of the CP domain or P#2 domain could not; furthermore, simultaneous overexpression of ALKBH5 and ZEB1 further increased the above effects (Fig. 7E–H). Nude mouse xenograft experiments revealed that GEM and RA significantly reduced tumor volume, and this effect could be further increased by overexpressing of ZEB1 and ALKBH5 (Fig. 7I–K). The overexpression of ZEB1 and ALKBH5 prolonged the overall survival (OS) of CDX-loaded mice (Fig. 7L). Collectively, the above results confirmed that simultaneous overexpression of ZEB1 and ALKBH5 can promote the therapeutic effect of GEM and RA on ATC.
A, B WB analyses using anti-FLAG, anti-HA, anti-CRABP2 and anti-FABP5 antibodies in ZEB1-FLAG, Del-CP-ZEB1, ALKBH5-HA, Del-P#2-ALKBH5 and ALKBH5 + ZEB1-overexpressing THJ-16T cells with or without 5 μM GEM and 10 μM RA (GEM + RA) treatment for 48 h. C, D CCK8 assays were performed on THJ-16T cells overexpressing ZEB1, Del-CP-ZEB1, ALKBH5, Del-P#2-ALKBH5 and ALKBH5 + ZEB1 following the same treatments described in (A, B). E, F Caspase 3/7 activity was measured in THJ-16T cells overexpressing ZEB1, Del-CP-ZEB1, ALKBH5, Del-P#2-ALKBH5 and ALKBH5 + ZEB1 following the same treatment described in (A, B). G, H Representative images from soft agar colony formation assays of THJ-16T cells overexpressing ZEB1, Del-CP-ZEB1, ALKBH5, Del-P#2-ALKBH5 and ALKBH5 + ZEB1 following the same treatment described in (A, B) are shown. Colony diameter was measured and graphed on the right; scale bar, 100 µm. I WB analyses using anti-ZEB1 and anti-ALKBH5 antibodies in ZEB1- and ALKBH5-overexpressing THJ-16T cells following the same treatment described in (A, B). J, K Mouse with a CDX of THJ-16T cells overexpressing ALKBH5 and ZEB1 were treated with or without RA (1.5 μg/kg) plus GEM administered 4 h later (100 mg/kg) for 18 days starting on Day 18. Representative images of the tumors at the end of the experiments were shown in (J), and the tumor growth curves were shown in (K). L Survival of mouse bearing xenografts formed by THJ-16T cells with or without ZEB1 and ALKBH5 knockout was monitored after inoculation for the indicated number of days. Mouse were administrated DMSO or RA (1.5 μg/kg) plus GEM 4 h later (100 mg/kg) for 18 days starting on Day 18 (n = 5/group). The data in (C–H) were analyzed by one-way ANOVA. The data in (K, L) were analyzed by two-way ANOVA. Images in (G, H) are representative of three independent experiments. *p < 0.05, **p < 0.01 indicate statistical significance.
Variable CRABP2 and FABP5 expression in thyroid samples
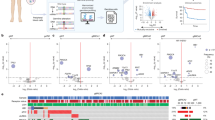
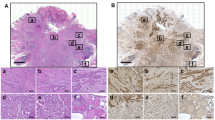
A total of 286 thyroid specimens were classified into four pathological subtypes, noncarcinoma (N), adenoma (A), differentiated thyroid carcinoma (DTC), and ATC (Supplementary Table 2). The patterns of CRABP2 and FABP5 expression in the four thyroid tissue groups were analyzed by tissue microarray-based immunohistochemical staining. The results revealed highly variable CRABP2 and FABP5 expression patterns not only among the four thyroid tissue groups but also among individual cases within each of the histological groups. Four CRABP2 and FABP5 expression patterns in thyroid samples were first summarized according to the presence or absence of their proteins in the immunolabelling: CRABP2(−)/FABP5(−), CRABP2(+)/FABP5(+), CRABP2(+)/FABP5(−), and CRABP2(-)/FABP5(+) (Fig. 8A). The CRABP2(+)/FABP5(+) pattern was then divided into C ≈ F (the levels of CRABP2 and FABP5 are close), C ↓ /F↑ (the level of CRABP2 is lower than that of FABP5) and C ↑ /F↓ (the level of CRABP2 is higher than that of FABP5). In N tissues, the majority of samples were CRABP2(-)/FABP5(-) (88%). In A tissues, the predominant pattern was CRABP2(-)/FABP5(+) (62%). In DTC tissues, the most common subtype was CRABP2( + )/FABP5(+) with a C ↓ /F↑ expression pattern (39%), followed by CRABP2(-)/FABP5(+) samples (31%) and CRABP2( + )/FABP5(+) samples with a C ≈ F expression pattern (19%). In ATC tissues, the primary subtype was CRABP2(-)/FABP5(-) (50%), with a significant proportion of CRABP2(-)/FABP5(+) samples (30%) also observed (Fig. 8B).
The staining patterns were scored as “−” if no immunolabeling was observed in the samples and “+” if distinct staining was generally observed. A CRABP2 and FABP5 expression patterns in noncancerous tissues and anaplastic thyroid cancer samples. BCRABP2 and FABP5 expression rates in the N, A, DTC, and ATC samples.
Discussion
ATC is a well-known lethal malignancy due to its aggressive behavior and distant metastasis4. To prevent tumor recurrence, postoperative treatments are used, with a combination of radiotherapy and RA being widely used, as RA increases the uptake of radioactive iodine in ATC cells by inducing redifferentiation26,27. However, the therapeutic effects of this treatment have not been promising, as patients receiving 131I therapy showed a reduction in Tg levels, accompanied by adverse effects such as dry skin and lips28 and a proportion of DTC patients have impaired 131I uptake capacity29. Therefore, re-evaluating the anti-ATC efficacy of RA to achieve more reliable treatment is of clinical value. Recent findings clearly indicate that a portion of ATC cell lines are resistant to RA. These findings further suggest that RA-based ATC therapy should be approached with caution in clinical settings unless the underlying mechanisms leading to changes in RA sensitivity in ATC patients are clearly identified, to avoid the risk of poor efficacy. Alternatively, a comprehensive investigation into the molecular determinants of RA sensitivity in ATC cells could help personalize ATC treatment. In this context, the three RA-resistant ATC cell lines used here constitute an ideal in vitro model to address this issue.
The addition of GEM affected the efficacy of RA, leading to significant growth inhibition and altering the epigenetic silencing of CRABP2. In RA-resistant cell lines with predictive “resistance” characteristics (CRABP2(-)/FABP5(+)), RA monotherapy did not exhibit any growth inhibitory effects. However, when combined with GEM, this situation changed completely. Both in vitro and in vivo, the combined action of RA and GEM resulted in the growth inhibition of ATC cells. There are reports that increased expression of L1 cell adhesion molecule (L1CAM) in ATC cell lines increases chemosensitivity to GEM30. Therefore, it may be worth considering the use of RA and GEM combination therapy for ATC treatment, along with gene editing of L1CAM.
Through database searches, we found that the transcription factors ZEB1 and SP-1 can bind to the promoters of both CRABP2 and FABP5. ZEB1 is a transcription factor that is extensively involved in gene regulation and play a key role in EMT, thereby promoting cancer progression31. SP-1 regulates the expression of target genes by binding to GC-rich sequences in promoters or enhancers32. SP-1 participates in various biological processes, including cell cycle regulation, stress responses, development, and tumor invasion27,33. In tumors, ZEB1 and SP-1 have a close relationship. SP-1 can act as a transcription factor by binding to the ZEB1 promoter, promoting ZEB1 expression34, and ZEB1 expression is positively correlated with SP-1 expression35. Additionally, ZEB1 and SP-1 can coregulate E-cadherin, thus modulating the EMT of lung epithelial cells36. In this study, we found that ZEB1 and SP-1 simultaneously regulate the transcription of CRABP2 and FABP5. Since ZEB1 has a more significant regulatory role, we further investigated the molecular mechanism of ZEB1-mediated transcription: the CP domain of ZEB1 competitively bound to KAT2B and EP300, both of which promoted ZEB1-mediated transcription of CRABP2 and FABP5. Under baseline conditions, ZEB1 mainly bound to the FABP5 promoter, but after the addition of GEM and RA, ZEB1 switched to binding the CRABP2 promoter and promotes its expression. Previous studies have shown that ZEB1 plays a critical role in various cancers, particularly in metastasis and drug resistance. However, relatively few studies have investigated the role of ZEB1 in thyroid cancer. Our research demonstrated that ZEB1 enhances the sensitivity of thyroid cancer to GEM and RA treatment by regulating the expression of CRABP2 and FABP5, with ZEB1 overexpression further amplifying the therapeutic effects. These findings suggested that ZEB1 could be a critical factor in improving the responsiveness of thyroid cancer patients to treatment. Moreover, overexpression of METTL3 in papillary thyroid cancer (PTC) and ATC cells improved the efficacy of anti-PD-1 therapy in a humanized peripheral blood mononuclear cell (huPBMC)-NCG mouse model37; our study also revealed that overexpression of the m6A demethylase ALKBH5 can enhance the therapeutic effects of GEM and RA. Therefore, the regulatory mechanism of CRABP2 and FABP5 expression identified in this study may provide new therapeutic strategies for the treatment of thyroid cancer.
YAP is the core transcriptional coactivator of the Hippo signaling pathway, and its aberrant activation is considered one of the driving factors in the development and progression of various tumors38. YAP primarily activates downstream target genes through the transcriptional enhancer factor associated domain (TEAD) family proteins, regulating mechanisms such as the cell cycle, metabolic reprogramming, and immune evasion, thereby increasing the survival and invasive capabilities of tumor cells39,40. However, other molecular mechanisms cannot be ruled out, including the recent discovery that YAP can promote polyamine biosynthesis by activating the transcription of the rate-limiting enzyme ornithine decarboxylase 1 (ODC1), which in turn facilitates the hypusination of eukaryotic translation initiation factor 5 A (eIF5A), supporting the translation of the transcriptional repressor lysine-specific demethylase 1 (LSD1), leading to the downregulation of numerous tumor suppressor genes41. Previous studies have also shown that YAP interacts with ZEB1. Our laboratory revealed that in LUAD, ZEB1 and YAP interact via the zinc finger cluster domain of ZEB1 and the WW domain of YAP, and that YAP can increase ZEB1-mediated promotion of the ferroptosis determinant LPCAT3, increasing tumor cell sensitivity to ferroptosis23. Here, we further confirmed the interaction between YAP and ZEB1 and demonstrated that the activation of YAP-related pathways can increase the sensitivity of thyroid tumors to GEM and RA treatments. Previous studies have also shown that high YAP expression increases tumor cell sensitivity to ferroptosis42. These studies collectively confirm that high YAP expression can provide more targets and methods for tumor therapy.
This study investigated CRABP2 and FABP5 expression patterns across four distinct thyroid tissue types: N, A, DTC, and ATC. Our findings revealed tissue-specific expression profiles that prompted further exploration into therapeutic implications. Subsequent investigations will use primary cell cultures and PDX models to examine how differential expression patterns of CRABP2/FABP5, regulated by ZEB1/METTL3/ALKBH5, influence therapeutic responses to GEM and RA in various thyroid cancer subtypes. These findings align with the literature demonstrating that CRABP2-high thyroid cancer cells are sensitive to RA43,44.
If our hypothesis that tumors with high CRABP2 and low FABP5 expression exhibit increased sensitivity to the combined GEM/RA therapy and that elevated expression of regulatory molecules such as ZEB, METTL3, and ALKBH5 further promotes therapeutic sensitivity to GEM/RA treatment is confirmed, this would provide critical insights into the molecular mechanisms underlying the drug response in thyroid cancer. Such confirmation could significantly advance precision medicine through the development of companion diagnostics, optimization of targeted therapies, and implementation of personalized treatment algorithms.
The clinical translation of CRABP2/FABP5 biomarkers involves several key aspects. First, patient stratification and therapeutic decision-making can be implemented through immunohistochemistry (IHC) staining or RT-qPCR assays for molecular subtyping. Patients can be stratified into GEM + RA-sensitive (CRABP2(+)/FABP5(-) or CRABP2 ↑ /FABP5 ↓ ) and GEM + RA-resistant (CRABP2(-)/FABP5(+) or CRABP2 ↓ /FABP5 ↑ ) groups. Further subdivision into ZEB/METTL3/ALKBH5 high/low expression subgroups (e.g., ZEB(+)/METTL3(+)/FABP5(+) as a sensitive subgroup) can be performed, enabling the implementation of tailored therapeutic strategies on basis of these molecular profiles. Sensitive subgroups may benefit from GEM + RA therapy combined with radiotherapy, whereas resistant subgroups may require alternative regimens.
Combination therapy strategies offer a promising approach for addressing resistant cases. These strategies may include the integration of GEM + RA therapy with FABP5 inhibitors (small molecules or monoclonal antibodies) and epigenetic modulators (activators of ZEB/METTL3/FABP5) to increase the sensitivity of tumors to GEM and RA treatment. Prioritizing clinical trials comparing combination therapies and standard care is essential for validating these approaches.
Prognostic evaluation and dynamic monitoring represent additional critical components. High-risk profiles characterized by CRABP2(-)/ FABP5(+) may require intensified surveillance, whereas favorable prognosis cases with CRABP2 ↑ / FABP5↓ may be managed with routine monitoring. The implementation of liquid biopsy protocols for serial ctDNA analysis of expression changes and real-time treatment adjustment on the basis of molecular evolution could revolutionize patient management.
Finally, targeted drug development and precision implementation strategies are crucial for advancing the field. This includes strategic pipeline development of FABP5-specific inhibitors (small molecules/antibody-drug conjugates), RA formulation optimization for increased bioavailability, and CRABP2, ZEB1, METTL3, and FABP5 expression activators (epigenetic modifiers). Clinical trial design should incorporate enrichment strategies for resistant populations, adaptive trial designs with biomarker-driven arm allocation, and the codevelopment of diagnostic-therapeutic packages.
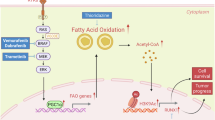
In conclusion, this study identified that the high expression of CRABP2 and low expression of FABP5 in ATCs are associated with increased sensitivity to combination therapy with GEM and RA. Furthermore, following GEM and RA treatment, the sensitivity of specific types of ATC cells to drug therapy can be increased through ZEB1-dependent transcriptional regulation and m6A-mediated mRNA stability (Fig. 9). The elucidation of this mechanism suggests a transformative approach in thyroid cancer management through the integration of molecular diagnostics with mechanism-based therapeutics. Successful implementation requires coordinated efforts across molecular pathology, pharmaceutical development, and clinical trial infrastructure, ultimately paving the way for more effective and personalized thyroid cancer treatment strategies.
In a subset of ATC cells (including THJ-11T, THJ-16T, and THJ-21T cells), following combined treatment with GEM and RA, the ZEB1/EP300/KAT2B transcriptional complex relocates from the FABP5 promoter to the CRABP2 promoter, thereby increasing CRABP2 transcription. Concurrently, the binding of METTL3 and ALKBH5 shifts from CRABP2 mRNA to FABP5 mRNA, facilitating the degradation of FABP5. Collectively, these changes result in the upregulation of CRABP2 and the downregulation of FABP5, increasing the sensitivity of ATC cells to GEM and RA treatment.
Methods
Cell culture
Three human ATC cell lines—8505 C, Cal-62, and HTH7—were purchased from the Shanghai Institute of Cell Biology, Chinese Academy of Sciences. The THJ-11T, 16 T, and 21 T cell lines45 were generously provided by Professor Quentin Liu, Institute of Cancer Stem Cell, Dalian Medical University, as part of the general gifts from the Mayo Foundation for Medical Education and Research. The 8505 C, Cal-62, THJ-11T, 16 T, and 21 T cell lines were cultured in RPMI 1640 (#SH30809; HyClone, Logan, UT, USA) supplemented with 10% fetal bovine serum (FBS; #C0232; Gibco, Grand Island, NY, USA) at 37 °C in a 5% CO₂ atmosphere. HTH7 cells were grown in Dulbecco’s modified Eagle’s medium (DMEM; #SH30243; HyClone) supplemented with 10% FBS. The cells (5 × 10⁴/ml) were cultured in standard Petri dishes for RNA and protein extraction and seeded onto cell slides to prepare sufficient samples for various experimental purposes under identical conditions.
Drug treatments
RA (#1674004; Sigma, St Louis, MO, USA) and GEM (#Y0000675; Sigma) were dissolved in DMSO (#8418; Sigma) and diluted in culture medium to working concentrations of 10 μM and 5 μM, respectively, which have been shown to be effective in other cancer cell lines46,47. The cells were treated for 2 days and observed every 6 h. Cells cultured in medium with or without 0.2% DMSO served as controls. Cell viability was assessed with a CCK8 kit (#C0039; Beyotime, Haimen, Jiangsu, China), and caspase 3/7 activity was measured with the Caspase 3/7 Glo luciferase reagent (#G8981; Promega, Madison, WI, USA). Each experiment was repeated at least three times, and the data were statistically analyzed by ANOVA. For 3D spheroid cultures, cells were seeded onto basement membrane extract (#356234; BME; Corning, Corning, New York, USA) at a density of 1 × 10⁵ cells per well. Once spheroids were formed, the cells were treated with DMSO, RA, or GEM, respectively. Images were captured after staining with SYTOX™ Green (#S7020; Invitrogen, Carlsbad, CA, USA). For the anchorage-dependent colony formation assay, cells were seeded at a density of 6 × 103 per well in six-well plates containing 0.3% agarose. After two weeks, the number of colonies was counted under a microscope.
Animals
Upon arrival, thymus-free nude mice (n = 5/group, 4–5 weeks old, JieSiJie, Shanghai, China) were fed ad libitum and housed under optimal conditions, including a 12 h light/dark cycle. The mice were isolated in a specific pathogen-free (SPF) facility for one week before the start of the experiments. All experimental procedures were conducted in accordance with animal welfare guidelines, following the “3 R” principle (Replace, Reduce, Refine), and were approved by the Animal Ethics Committee of Shanghai Chest Hospital (KS24018). THJ-16T cells (5 × 10⁶) were resuspended in 200 μL of Hank’s balanced salt solution (HBSS; #SH30030; calcium-free, magnesium-free, phenol red-free; HyClone) and subcutaneously injected into the right abdomen of immunodeficient mice. Stock solutions of RA and GEM were prepared using DMSO as a solvent and subsequently diluted in saline prior to treatment. Palpable THJ-16T xenografts developed within 12–14 days. Animals with 18-day-old xenografts were randomly assigned to 4 groups (5 animals per group) for treatment: DMSO, RA, GEM, and RA plus GEM. Each animal received a daily intraperitoneal dose of RA (1.5 μg/kg), GEM (100 mg/kg), or RA (1.5 μg /kg) plus GEM administered 4 h later (100 mg/kg) for 18 days. Tumor development was monitored twice a week, and the animals were euthanized by lethal IP injection of 200 mg/mL pentobarbital sodium 36 days after injection. The tumor volume evaluated by measuring the tumor length and width (in millimeters) with a caliper. The tumor volume was calculated as 0.5 × L × W2 (L indicates length, whereas W indicates width)23.
Reagents and plasmids
CRABP2, FABP5, ZEB1, ALKBH5, YAP, KAT2B, EP300, methyltransferase-like 14 (METTL14) and FTO alpha-ketoglutarate dependent dioxygenase (FTO) expressing plasmids were purchased from BioVision (Shanghai, China). CRABP2-3’UTR and FABP5-3’UTR overexpression plasmids were purchased from BioVision (Shanghai, China). CRABP2, FABP5, methyltransferase-like 3 (METTL3), ALKBH5, and YAP knockout plasmids were constructed with the lentiCRISPR V2 vector. CRABP2, FABP5, ZEB1, METTL3 and ALKBH5 domain mutation plasmids were constructed by overlapping PCR and the PCR products were subsequently cloned and inserted into the pLVX-Puro vector. The primers used are listed in Supplementary Table 1.
IF staining, IHC staining, and western blotting (WB)
IF staining, IHC staining and WB were carried out following standard protocols. The primary antibodies used for IHC staining were anti-CRABP2 (1:250; #66468; Proteintech, Wuhan, China) and anti-FABP5 (1:50; #sc-365236; Santa Cruz Biotechnology [Santa Cruz], California, USA) antibodies. The primary antibodies used for IF were anti-CRABP2 (1:50; #66468; Proteintech), anti-FABP5 (1:50; #CL555-12348; Proteintech), anti-FLAG (1:100; #ab205606; Abcam, Hong Kong, China) and anti-Myc (1:50; #C6594; Sigma) antibodies. The primary antibodies used for WB were anti-GAPDH (1:1000; #51332; Cell Signaling Technology [CST], Boston, MA, USA), anti-CRABP2 (1:2500; #66468; Proteintech), anti-FABP5 (1:1000; #ab255276; Abcam), anti-FLAG (1:1000; #ab205606; Abcam), anti-HA (1:4000; #ab9110; Abcam), anti-Myc (1:1000; #ab32072; Abcam), anti-ZEB1 (1:100; #sc-515797; Santa Cruz) and anti-ALKBH5 (1:2000; #16837, Proteintech) antibodies.
Co-IP
Co-IP was carried out following standard protocols. Briefly, cell lysates were incubated overnight at 4 °C with the appropriate antibodies and protein A/G magnetic beads (#AP62L142; Life iLab, Shanghai, China). The following day, the beads were washed three times with Western/IP lysis buffer (#P0013; Beyotime), resuspended in SDS-PAGE loading buffer (#P0015; Beyotime), and subjected to SDS-PAGE for protein separation. The antibodies used for Co-IP included anti-KAT2B (1:100; #3378; CST), anti-ZEB1 (1:100; #sc-515797; Santa Cruz), anti-GAPDH (1:1000; #51332; CST), anti-FLAG (1:50; #ab205606; Abcam), and anti-Myc (1:50; #13987; CST) antibodies.
PLA
PLA was measured with the Duolink In Situ Red Starter Kit (#DUO92101; Sigma). Cells were seeded onto cell crawlers for 24 h. The cells were fixed with 4% paraformaldehyde, blocked with blocking buffer and incubated with primary antibodies at 4 °C overnight. On the third day, the indicated PLA probe solutions were added to each cell crawler slide and incubated at 37 °C for 1 h. Ligase conjugate solution was then added, and the samples were incubated at 37 °C for 30 min. After ligation, amplification polymerase solution was added to each well, and the samples were incubated at 37 °C for 100 min. The cells were then subjected to confocal microscopy. The primary antibodies used were anti-FLAG (1:500; #ab205606; Abcam) and anti-Myc (1:500; #2233; CST) antibodies.
ChIP
ChIP assays were conducted with kits obtained from CST (#9002; CST). In brief, the cells (2 × 10⁷) were cross-linked with 1% formaldehyde, washed with PBS buffer, and lysed. Following sonication, the protein-DNA complexes were incubated with antibody-bound protein G beads overnight at 4 °C. The next day, the DNA was eluted with 1% SDS and 0.1 mol/L NaHCO₃, followed by reverse cross-linking at 65 °C. The samples were then purified through phenol/chloroform extraction and ethanol precipitation, and analyzed by agarose gel electrophoresis prior to PCR. The primary antibodies used were anti-ZEB1 (1:50; #ab155249; Abcam), anti-SP-1 (1:50; #ab231778; Abcam), anti-KAT2B (1:50; #57625; CST), anti-H3K122Ac (1:100; #8173; CST), anti-H3K9Ac (1:50; #9649; CST), anti-EP300 (1:50; #54062; CST), anti-H3K27Ac (1:100; #8173; CST), anti-CRABP2 (1:50; #sc-166897, Santa Cruz), and anti-FABP5 (1:50; #sc-365236, Santa Cruz) antibodies and IgG (1:100; #3900; CST).
RT- qPCR
Total RNA was isolated with TRIzol reagent (#15596018; Ambion, Carlsbad, CA, USA) and reverse-transcribed into complementary DNA with the ABScript Neo RT Master Mix (#RK20433; ABclonal, Wuhan, China). RT-qPCR was performed with SYBR Green qPCR Mix (#RK21219; ABclonal). The primer sequences are provided in Supplementary Table 1.
Luciferase activity measurement
Luciferase activity was assessed with a dual-luciferase assay kit (#11402ES60; YEASEN, Shanghai, China) according to the manufacturer’s protocol. Firefly luciferase reporters (WT-CRABP2, Mut-CRABP2, WT-FABP5 and Mut-FABP5 promoter constructs) were cotransfected into cancer cells along with the Renilla plasmid with Lipofectamine™ 2000 Transfection Reagent (#11668019, Invitrogen). The cells were collected and lysed in passive lysis buffer after 48 h. The fluorescence intensity of the luciferase reporters was measured and normalized to Renilla luciferase activity.
EMSA
EMSA was conducted as described previously23. Nuclear proteins were extracted with a kit from Active Motif and incubated in reaction buffer with or without DNA competitors on ice prior to the addition of biotin-labeled probes (Sangon, Shanghai, China). The DNA-protein complexes were separated by electrophoresis on 5% native polyacrylamide gels and transferred to Immobilon-My+ membranes. The probes used are listed in Supplementary Table 1.
Protein and RNA interaction analyses, RNA pull-down, RIP, PAR-CLIP
The molecular basis underlying the interactions among CRABP2, FABP5, METTL3 and ALKBH5 interactions was predicted by catRAPID database48. The sequences for CRABP2, FABP5, METTL3, and ALKBH5 were entered into the respective fields, and upon clicking the submit button, the prediction results were displayed in a new window.
For RNA pull-down assays, synthesized CRABP2 and FABP5 sequences (both WT and mutants), along with their biotin-labeled antisense probes, were obtained from Sangon (details provided in Supplementary Table 1). Approximately 1 × 10⁷ cells were lysed and incubated with 3 μg of the biotinylated probes overnight at 4 °C. The biotin-RNA-protein complexes were captured with streptavidin magnetic beads (#88803; Thermo Fisher Scientific, Waltham, Massachusetts, USA) after incubation for an additional 4 h. After five washes with PBS, the streptavidin beads were boiled, and the eluates were analyzed by immunoblotting (IB) or proteomic analysis.
For the RIP assays, a Magna RIP Kit (#17-700; Merck Millipore, Billerica, MA, USA) was used. Briefly, the cell lysates were incubated with magnetic beads loaded with 5 μg of anti-METTL3 (#86132; CST); anti-METTL14 (#48699; CST), anti-FTO (#ab126605; Abcam), anti-ALKBH5 (#16837; Proteintech), anti-FLAG (#ab205606; Abcam), anti-HA (#ab9110; Abcam), or anti-m6A (#68055, Proteintech) antibodies or control IgG (#3900; CST) overnight at 4 °C. After proteinase K digestion, the remaining RNA was extracted with TRIzol reagent and analyzed by qPCR.
For PAR-CLIP, the cells were incubated with 4-thiouridine (4-SU; 250 μmol/L; #4509; Sigma) for 16 h, followed by crosslinking via irradiation with 365 nm ultraviolet (UV) light. The cells were then lysed in NP40 lysis buffer on ice and centrifuged at 18,000 × g for 15 min to collect the supernatant. This mixture was incubated with 600 μL of protein A Dynabeads conjugated with 15 μg of anti-FLAG (#ab205606; Abcam) and 15 μg of anti-HA (#ab9110; Abcam) antibodies for 2 h. After three washes with IP wash buffer, the beads were resuspended and boiled at 95 °C for 10 min. RNA was then recovered and analyzed by qPCR to assess FLAG or HA-bound circRNAs.
Tissue microarray-based immunohistochemical staining
After obtaining permission from the institutional review board and providing informed consent from patients, 286 archived, paraffin-embedded blocks of surgical thyroid samples were kindly provided for tissue microarray analysis by the Department of Clinical Pathology, the First Affiliated Hospital of Dalian Medical University and the Department of Laboratory Medicine, the Second Hospital of South China University of Technology without prejudice toward pathological diagnosis. Among these samples, 34 were diagnosed as A, 103 as well- DTCs, and 30 as ATCs. A total of 115 samples contained tumors surrounded by noncancerous thyroid tissues (N), with 4 cases of Hashimoto’s thyroiditis (HT). Tissue microarrays, with densities of 36–56 spots/cm², were constructed as described previously and sectioned into 5 μm slices for immunohistochemical analysis of CRABP2 and FABP5 expression and localization49. The same antibodies used for immunocytochemical staining were applied. Sections without primary antibody incubation served as background controls. On the basis of the labeling intensity, the staining results were assessed by two independent researchers and scored as negative (-) if no immunolabeling was detected, weakly positive (+) if the labeling was faint, moderately positive (++) if the for labeling was stronger, and strongly positive (+++) if the labeling was significantly stronger than samples scored as (++)9.
Statistical analysis
Differences between groups were analyzed by Student’s t-test, one-way ANOVA, and two-way ANOVA and the χ2 test. The correlation between two groups was evaluated by Spearman’s rank-correlation analysis. p < 0.05 was considered statistically significant.
Data availability
No datasets were generated or analysed during the current study.
Code availability
No codes were generated or analyzed during the current study.
References
Liu, Y. et al. Radioiodine therapy in advanced differentiated thyroid cancer: resistance and overcoming strategy. Drug Resist. Updat. 68, 100939 (2023).
Cabanillas, M., Mcfadden, D. & Durante, C. Thyroid cancer. Lancet 388, 2783 (2016).
Liu, N. et al. Subplatysmal or subfascial approach in totally endoscopic thyroidectomy has better postoperative efficacy for voice, sensory, swallowing symptoms and cosmetic result. Cohort study. Int J. Surg. 60, 22 (2018).
Lu, L. et al. Anaplastic transformation in thyroid cancer revealed by single-cell transcriptomics. J. Clin. Investig. 133, e169653 (2023).
Catalano, M. G. et al. Emerging molecular therapies of advanced thyroid cancer. Mol. Asp. Med 31, 215 (2010).
Li, Y. et al. Resveratrol suppresses the growth and enhances retinoic acid sensitivity of anaplastic thyroid cancer cells. Int. J. Mol. Sci. 19, 1030 (2018).
Elisei, R. et al. All-trans-retinoic acid treatment inhibits the growth of retinoic acid receptor beta messenger ribonucleic acid expressing thyroid cancer cell lines but does not reinduce the expression of thyroid-specific genes. J. Clin. Endocrinol. Metab. 90, 2403 (2005).
Fröhlich, E. & Wahl, R. Nanoparticles: promising auxiliary agents for diagnosis and therapy of thyroid cancers. Cancers13, 4063 (2021).
Kocher, H. M. et al. Phase I clinical trial repurposing all-trans retinoic acid as a stromal targeting agent for pancreatic cancer. Nat. Commun. 11, 4841 (2020).
Courbon, F. et al. Defective efficacy of retinoic acid treatment in patients with metastatic thyroid carcinoma. Thyroid 16, 1025 (2006).
Schmutzler, C. et al. Functional retinoid and thyroid hormone receptors in human thyroid-carcinoma cell lines and tissues. Int. J. Cancer 76, 368 (1998).
Kogai, T. & Brent, G. A. The sodium iodide symporter (NIS): regulation and approaches to targeting for cancer therapeutics. Pharm. Ther. 135, 355 (2012).
Gupta, S. et al. Molecular determinants of retinoic acid sensitivity in pancreatic cancer. Clin. Cancer Res. 18, 280 (2012).
Plantier, L. et al. Dysregulation of elastin expression by fibroblasts in pulmonary emphysema: role of cellular retinoic acid binding protein 2. Thorax 63, 1012 (2008).
An, Y. & Duan, H. The role of m6A RNA methylation in cancer metabolism. Mol. Cancer 21, 14 (2022).
Zhang, Y. et al. m(6)A RNA modification regulates innate lymphoid cell responses in a lineage-specific manner. Nat. Immunol. 24, 1256 (2023).
Ma, L. et al. Targeting SLC3A2 subunit of system X(C)(-) is essential for m(6)A reader YTHDC2 to be an endogenous ferroptosis inducer in lung adenocarcinoma. Free Radic. Biol. Med 168, 25 (2021).
Ma, L. et al. The m(6)A reader YTHDC2 inhibits lung adenocarcinoma tumorigenesis by suppressing SLC7A11-dependent antioxidant function. Redox Biol. 38, 101801 (2021).
Shu, F. et al. m6A Modification Promotes EMT and Metastasis of Castration-Resistant Prostate Cancer by Upregulating NFIB. Cancer Res 84, 1947 (2024).
Lu, M. D. et al. Dual inhibition of BRAF-MAPK and STAT3 signaling pathways in resveratrol-suppressed anaplastic thyroid cancer cells with BRAF mutations. Int. J. Mol. Sci. 23, 14385 (2022).
Yu, S. et al. Targeting CRABP-II overcomes pancreatic cancer drug resistance by reversing lipid raft cholesterol accumulation and AKT survival signaling. J. Exp. Clin. Cancer Res. 41, 88 (2022).
Hogg, S. J. et al. Targeting histone acetylation dynamics and oncogenic transcription by catalytic P300/CBP inhibition. Mol. Cell 81, 2183 (2021).
Cui, J. et al. LPCAT3 is transcriptionally regulated by YAP/ZEB/EP300 and collaborates with ACSL4 and YAP to determine ferroptosis sensitivity. Antioxid. Redox Signal 39, 491 (2023).
Sankar, A. et al. Histone editing elucidates the functional roles of H3K27 methylation and acetylation in mammals. Nat. Genet. 54, 754 (2022).
Shen, C. et al. m(6)A-dependent glycolysis enhances colorectal cancer progression. Mol. Cancer 19, 72 (2020).
Zhang, Y. & Wang, J. Targeting uptake transporters for cancer imaging and treatment. Acta Pharm. Sin. B 10, 79 (2020).
Wilms, C. et al. Glutaredoxin 2 promotes SP-1-dependent CSPG4 transcription and migration of wound healing NG2 glia and glioma cells: enzymatic Taoism. Redox Biol. 49, 102221 (2022).
Nikitski, A. V., Condello, V., Divakaran, S. S. & Nikiforov, Y. E. Inhibition of ALK-signaling overcomes STRN-ALK-induced downregulation of the sodium iodine symporter and restores radioiodine uptake in thyroid cells. Thyroid 33, 464 (2023).
Mu, Z. et al. Risk stratification for radioactive iodine refractoriness using molecular alterations in distant metastatic differentiated thyroid cancer. Chin. J. Cancer Res. 36, 25 (2024).
Kim, K. S. et al. Aberrant l1 cell adhesion molecule affects tumor behavior and chemosensitivity in anaplastic thyroid carcinoma. Clin. Cancer Res. 18, 3071 (2012).
Mosquera, N. et al. Uncovering cellular retinoic acid-binding protein 2 as a potential target for rheumatoid arthritis synovial hyperplasia. Sci. Rep. 8, 8731 (2018).
Kennedy, M. W. et al. Sp5 and Sp8 recruit β-catenin and Tcf1-Lef1 to select enhancers to activate Wnt target gene transcription. Proc. Natl. Acad. Sci. USA 113, 3545 (2016).
Kim, H. et al. In vitro and in vivo anti-tumor efficacy of krill oil against bladder cancer: Involvement of tumor-associated angiogenic vasculature. Food Res. Int. 156, 111144 (2022).
Jiang, Y. et al. Oncogenic HBXIP enhances ZEB1 through Sp1 to accelerate breast cancer growth. Thorac. Cancer 9, 1664 (2018).
Lee, J., Jung, E., Gestoso, K. & Heur, M. ZEB1 Mediates Fibrosis in Corneal Endothelial Mesenchymal Transition Through SP1 and SP3. Investig. Ophthalmol. Vis. Sci. 61, 41 (2020).
Zhang, X. et al. Transcriptional repression of E-cadherin in nickel-exposed lung epithelial cells mediated by loss of Sp1 binding at the promoter. Mol. Carcinog. 61, 99 (2022).
Ning, J. et al. METTL3 inhibition induced by M2 macrophage-derived extracellular vesicles drives anti-PD-1 therapy resistance via M6A-CD70-mediated immune suppression in thyroid cancer. Cell Death Differ. 30, 2265 (2023).
Zhou, W. et al. Role of YAP signaling in regulation of programmed cell death and drug resistance in cancer. Int. J. Biol. Sci. 20, 15 (2024).
Pearson, J. et al. Binary pan-cancer classes with distinct vulnerabilities defined by pro- or anti-cancer YAP/TEAD activity. Cancer Cell 39, 1115 (2021).
Dey, A., Varelas, X. & Guan, K. L. Targeting the hippo pathway in cancer, fibrosis, wound healing and regenerative medicine. Nat. Rev. Drug Discov. 19, 480 (2020).
Li, H. et al. YAP/TAZ drives cell proliferation and tumour growth via a polyamine-eIF5A hypusination-LSD1 axis. Nat. Cell Biol. 24, 373 (2022).
Wu, J. et al. Intercellular interaction dictates cancer cell ferroptosis via NF2-YAP signalling. Nature 572, 402 (2019).
Liu, C. L. et al. CRABP2 is associated with thyroid cancer recurrence and promotes invasion via the integrin/FAK/AKT pathway. Endocrinology 163, bqac171 (2022).
Liu, X. et al. Resveratrol reverses retinoic acid resistance of anaplastic thyroid cancer cells via demethylating CRABP2 gene. Front. Endocrinol.10, 734 (2019).
Silvis, A. M., Mccormick, M. L., Spitz, D. R. & Kiningham, K. K. Redox balance influences differentiation status of neuroblastoma in the presence of all-trans retinoic acid. Redox Biol. 7, 88 (2016).
Maftouh, M. et al. Synergistic interaction of novel lactate dehydrogenase inhibitors with gemcitabine against pancreatic cancer cells in hypoxia. Br. J. Cancer 110, 172 (2014).
Zhang, X. et al. Endogenous glutamate determines ferroptosis sensitivity via ADCY10-dependent YAP suppression in lung adenocarcinoma. Theranostics 11, 5650 (2021).
Bonhomme, B. et al. Molecular pathology of anaplastic thyroid carcinomas: a retrospective study of 144 cases. Thyroid 27, 682 (2017).
Zhang, P. et al. PIAS3, SHP2 and SOCS3 expression patterns in cervical cancers: relevance with activation and resveratrol-caused inactivation of STAT3 signaling. Gynecol. Oncol. 139, 529 (2015).
Acknowledgements
This work was supported by Shanghai Rising Star Program (22QA1408400, 22QA1408300), Project of Clinical Research Supporting System, Clinical Medicine First-class Discipline, Talent Training Plan of Shanghai Chest Hospital (to Xiao Zhang), Excellent Talents Nurture Project of Shanghai Chest Hospital (2021YNZYY01). The author extends sincere gratitude to Professor Hong Li and Professor Jia Liu from Dalian Medical University for their invaluable guidance in the preparation of this paper.
Author information
Authors and Affiliations
Contributions
X.T., K.P., T.Z., D.S. and X.Z. wrote the main manuscript text. K.P., S.Y., J.W., Q.M., and L.M. searched for information. X.T., B.J. and Y.M. prepared figures 1-2. K.P., T.Z. and L.M. prepared figures 3-4. X.T., D.S., and X.Z. prepared figures 5-9. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tian, X., Peng, K., Zhao, T. et al. ZEB1 and ALKBH5 coregulate CRABP2 and FABP5 and jointly determine the sensitivity of anaplastic thyroid cancers to gemcitabine and retinoic acid. npj Precis. Onc. 9, 334 (2025). https://doi.org/10.1038/s41698-025-01073-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-025-01073-7