Abstract
Immunotherapy has emerged as an effective treatment for breast cancer, making the exploration of novel immune-related biomarkers of paramount importance. A vital aspect of this exploration is the investigation into the subtyping of tumor-associated macrophages (TAMs). While polarity proteins within TAMs can shift their functional status, the impact of polarity proteins on inflammation-related signaling in tumor cells and their subsequent influence on the tumor microenvironment (TME) remains elucidated. We discovered that RICH1, functioning as a tumor suppressor molecule in breast cancer, significantly increased the infiltration of pro-inflammatory M1-like TAMs within TME in 4T1 tumor-bearing mice. Furthermore, the conditioned medium from RICH1-overexpressing 4T1 cells promoted M1-like polarization in vitro by stimulating the secretion of IFN-γ and other cytokines. Mechanistically, high expression of RICH1 in breast cancer cells facilitated the ubiquitination degradation of RhoA through binding with TRIM21 and enhancing the interaction between TRIM21-RhoA, thereby inhibited the phosphorylation of STAT3, up-regulated the production and secretion of IFN-γ, consequently induced M1-like polarization of macrophages. Our findings reveal that RICH1 plays a crucial role in promoting pro-inflammatory TAMs infiltration in breast cancer through modulation of inflammatory signaling. These results suggest that RICH1 could serve as an immune-related biomarker and a key contributor to the formation of immune-active microenvironments, with potential applications in combination immunotherapy strategies.
Similar content being viewed by others
Introduction
Immunotherapy has emerged as a highly effective treatment for breast cancer, complementing traditional chemoradiotherapy1. To address challenges related to immunotherapy insensitivity, clinicians categorize breast cancer based on various tumor microenvironment (TME) characteristics, such as triple negative breast cancer (TNBC) classification, including Lehmann classification2, FUSCC classification3, and classification based on the spatial distribution of CD8+ TILs (tumor-infiltrating lymphocytes)4, and the immunomodulatory subtype SNF2 in the SNF classification of Luminal breast cancer5. These classifications not only reveal the heterogeneity within the TME but also optimize the tailoring of individualized immunotherapy for breast cancer patients6. The exploration on the elements of typing is the process of identifying novel immune-related biomarkers and targets of breast cancer.
As the primary part of inflammatory infiltration in TME, tumor-associated macrophages (TAMs) are the most dominant type of immune cells7. To understand the spatial diversity of tumor-infiltrating immune cells within the intra-tumor and intra-stromal regions, it is imperative to meticulously examine TAM subtypes and their associated functions8. Investigating the equilibrium between the anti-tumor pro-inflammatory TAMs and the pro-tumor anti-inflammatory TAMs is of paramount importance and can serve as a foundational framework for novel research on immunotherapeutic agents9. Explaining immunoinflammatory signaling pathways can shed light on the mechanisms of macrophage reprogramming and repolarization in the TME, providing valuable insights into the orchestration of the immune-activating milieu in breast cancer.
TAMs subtype polarization in TME is influenced by stimulating factors like interferon-gamma (IFN-γ) and related immunomodulators, promoting the shift of macrophages toward proinflammatory M1 phenotypes10. JAK-STAT (Janus kinase-signal transducers and activators of transcription) signaling, essential in tumor-driven immune escape and inflammation, triggers a complex network of signals involving chemokines, growth factors, and IFN-related receptors11. Numerous studies have confirmed that the activation of NF-κB (nuclear factor-κB) and STAT3 in TME accelerates tumorigenesis by shaping the identity and functions of infiltrating macrophages and other immune cells12.
Dysregulation of cell polarity proteins and loss of apical–basal polarity are closely linked to tumor development13. These proteins help maintain epithelial integrity and also modulate immune cell functions such as polarization, activation, and cytokine production, thereby influencing the balance between antitumor and protumor responses in the tumor microenvironment14,15. There is also increasing evidence that polarity proteins participate in inflammation-associated signaling in both tumor and immune cells16, underscoring their relevance in cancer biology. RICH1, also known as ARHGAP17 (Rho GTPase Activating Protein 17), is a polarity-associated protein that contributes to apical-basal polarity maintenance through interactions with polarity complexes. It also contains a RhoGAP (Rho GTPase activating protein) domain that inactivates Rho family GTPases, including RhoA, Cdc42, and Rac1, thereby influencing cytoskeletal organization and cell motility17,18,19. And in previous study of our research group, RICH1 has been reported as a tumor suppressor molecule in breast cancer20.
Excessive Rho pathway activation has been implicated in the upregulation of STAT3 and NF-κB signaling, promoting a proinflammatory tumor microenvironment21. For instance, RacGAP promotes IL-6-driven leukemia cell differentiation via STAT3 activation22, while RhoGEFs in profibrotic macrophages enhance STAT3 phosphorylation through RhoA-dependent mechanisms23. Given these findings, we hypothesize that RICH1 expression in breast cancer cells may reset the pro-inflammatory phenotype of macrophages by altering cytokine secretion-a process potentially mediated by crosstalk between Rho GTPase signaling and JAK-STAT pathways. Therefore, our interest lies in exploring how RICH1 might impact the TME by modulating inflammatory molecular signals in breast cancer cells. In the present study, RICH1 regulates inflammatory signaling and immune infiltration in TME through Rho GTPase-STATs pathway, which will provide the basis for research on related signaling targets and combined immunotherapy.
Results
High RICH1 is associated with abundant pro-inflammatory M1-like macrophages infiltration and better prognosis in breast cancer
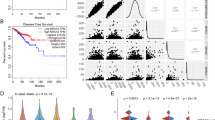
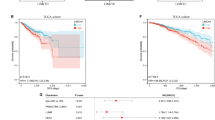
Transcriptome data from 1097 breast cancer samples in the TCGA database were analyzed using gene set enrichment analysis (GSEA) for Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways. The top 10 RICH1-related pathways, ranked by normalized enrichment score (NES), included several immune-related pathways such as the chemokine signaling pathway, T/B cell receptor signaling pathway, and natural killer cell-mediated cytotoxicity (Fig. 1A). To explore the relationship between RICH1 and immune cell infiltration in the TME of breast cancer, we used the CIBERSORT deconvolution algorithm to estimate the proportions of 22 immune cell types in TCGA samples. Differential analysis of immune cell components revealed that the RICH1-high group exhibited higher infiltration levels of macrophages M1 (p < 0.01), activated CD4 memory T cells (p < 0.01), and naive B cells, while having lower levels of macrophages M2 (p < 0.001) and resting mast cells (p < 0.01) compared to the RICH1-low group (Fig. 1B). Breast cancer patients with higher RICH1 had longer overall survival (OS) from datasets GSE42568 and GSE103091 in GEO database. In TNBC patients, high expression of RICH1 also correlated with a longer OS, which could be seen in GSE31519 (Fig. S1A). A meta-analysis across these three datasets confirmed that high RICH1 expression is a positive prognostic indicator for breast cancer (Fig. 1C). The relation of TAMs with RICH1 expression and the survival of patients with different types of TAMs infiltration (Fig. S1B, C) also suggests high RICH1 is associated with higher M1 infiltration and better prognosis in breast cancer. To further clarify the correlation between RICH1 expression and TAMs infiltration in TME, the spatial distribution of TAMs was detected through spatial transcriptomic analysis. Multiple genetic subclones in the breast cancer sample were identified, and the distribution of these clusters showed significant regional differences (Fig. 1D). Furthermore, at the boundaries of regions with high RICH1 expression, there was a notable accumulation of CXCL9+ and CXCL11+ pro-inflammatory M1-like TAMs. However, no similar aggregation trend was observed for CD68+ or CD163+ labeled TAMs (Fig. 1E and S1D).
A RICH1-related KEGG pathways of NES top10 enriched by GSEA. B Violin plot of proportion of tumor-infiltrating immune cells compared in the samples with different expression of RICH1 obtained from TCGA database. C A meta-analysis conducted based on datasets GSE42568, GSE103091 and GSE31519. D The clusters using spatial transcriptomic UMAP analysis for the breast cancer sample (Block ID: 1168993F from the 10x Genomics Visium platform) with significant regional differences. E Spatial distribution of RICH1 expression, CD68+ and CXCL9+ labeled macrophages.
High RICH1 promotes M1-like TAMs infiltration in TME of 4T1 tumor-bearing mice
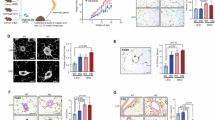
To investigate the effects of RICH1 expression in breast cancer cells on tumor growth and macrophage infiltration in vivo, 4T1 breast cancer cells derived from Balb/c mice were engineered to overexpress RICH1 (4T1-RICH1 OE) or serve as negative controls (4T1-NC) using RICH1/NC-targeted lentivirus. These cells were orthotopically transplanted into female Balb/c mice (Fig. S2A). After 21 days, tumors in the 4T1-RICH1 OE group showed significantly reduced volume and weight compared to the 4T1-NC group (Fig. 2A and Table S1). The weight of mice, immunohistochemistry (IHC) staining, and score of RICH1 in two groups were shown in Fig. S2B, C and Table S1. These findings indicate that high RICH1 expression suppresses tumor growth in 4T1 tumor-bearing mice. Flow cytometry analysis revealed differences in TAM infiltration within the TME of these mice.
A The volume and weight of the tumors compared in the mice injected with 4T1-RICH1 OE and 4T1-NC cells. B CD45 positive subpopulation represents immune cells in TME of 4T1 tumor-bearing mice. C Flow cytometry analysis and representative images of F4/80+CD86+ subpopulations and F4/80+CD163+ subpopulations in CD45 positive cells, in which the proportions indicate M1-like and M2-like TAMs percentages, respectively. Immunofluorescence assays marked F4/80+iNOS+ M1-like D and F4/80+CD206+ M2-like E TAMs for frozen sections of 4T1 tumors. F Representative images of CD11b+F4/80+ macrophages in CD45+ cells by flow cytometry analysis. G Representative images and proportions of CD86+ or NOS2+ or TNFα+ subpopulations in CD45+CD11b+F4/80+ cells by flow cytometry analysis. H The volume and weight of the tumors were compared in the 4T1-RICH1 OE/NC cells bearing mice with clophosome or control liposomes intraperitoneal injection. Abbreviation: P values in (A, C, G, and H) were calculated using unpaired two-tailed Student’s t tests. P value in line chart of A was calculated using two-way ANOVA tests. *P < 0.05; **P < 0.01; ***P < 0.001.
It was observed that the proportion of CD45+F4/80+CD86+ M1-like TAMs was significantly lower in the 4T1-NC group compared to the 4T1-RICH1 OE group. Conversely, the proportion of CD45+F4/80+CD163+ M2-like TAMs exhibited the opposite trend, although the differences were not substantial (see Fig. 2B, C and Table S2). Immunofluorescence assays were conducted on frozen sections of 4T1 tumors to demonstrate increased F4/80+iNOS+ M1-like infiltration in the 4T1-RICH1 OE group (Fig. 2D). However, the differences in F4/80+CD206+ M2-like infiltration between the two groups were not statistically significant (Fig. 2E). Further analysis of CD45+CD11b+F4/80+CD86+, NOS2+, and TNFα+ M1-like TAMs using flow cytometry revealed greater M1-like TAM infiltration in tumors from the 4T1-RICH1 OE group compared to the 4T1-NC group (Fig. 2F, G and Table S2). Similarly, high RICH1 expression inhibited tumor growth and enhanced M1-like TAM infiltration in SUM159 tumor-bearing mice, consistent with the results observed in the 4T1 model (Fig. S2D, E and Tables S2 and 3).
To further elucidate whether the effect of RICH1 for tumor inhibition is mediated through its impact on macrophage infiltration, an in vivo experiment involving macrophage depletion was performed. Clophosome could trigger irreversible apoptosis in macrophages, eliminating them via caspase-dependent pathways. The results demonstrated that the disparities in tumor volume and weight between the 4T1-RICH1 OE and 4T1-NC groups in the clophosome group were significantly smaller than those observed in the control liposomes group (see Figs. 2H and S3A and Table S4). Additionally, whether in the mouse spleen (Fig. S3B) or in the tumor (Fig. S3C), after treatment with clophosome, F4/80+CD11b+ labeled macrophages in CD45+ immune cell population were mostly cleared. These findings suggest that high RICH1 expression enhances M1-like TAM infiltration within the TME of tumor-bearing mice, contributing to tumor growth inhibition.
Conditioned medium of 4T1-RICH1 OE promotes M1-like polarization in vitro
To determine whether RICH1 influences M1 macrophage infiltration in the TME through breast cancer cell signaling, we conducted an in vitro study using conditioned medium from 4T1-RICH1 OE and 4T1-NC cells to polarize RAW264.7 macrophages. Immunofluorescence assays demonstrated that a higher proportion of RAW264.7 cells cultured in conditioned medium from 4T1-RICH1 OE were polarized towards the F4/80+iNOS+ M1-like macrophage phenotype compared to those cultured in 4T1-NC conditioned medium (Fig. 3A). Furthermore, a substantial number of RAW264.7 cells in both 4T1-RICH1 OE and 4T1-NC conditioned media exhibited polarization towards the F4/80+CD206+ M2-like macrophage phenotype, and the differences were not significant (Fig. 3B). The macrophage subtypes polarized by the conditioned medium were verified again by PCR. The relative mRNA expression levels of inflammatory cytokines in RAW264.7 cells showed that the mRNA levels of M1 markers, including IL12 (Interleukin-12), IL6, IL1b, TNFa (tumor necrosis factor alpha), and CXCL10 (C-X-C motif chemokine ligand 10), were upregulated in RAW264.7 cells polarized by conditioned medium from 4T1-RICH1 OE, while M2 marker expression (IL10 and TNFb) showed only minor differences (Fig. 3C and Table S5). Flow cytometry results confirmed these findings, with a higher proportion of CD86+ and CD80+ M1-like macrophages observed in the 4T1-RICH1 OE-conditioned medium compared to 4T1-NC. Lipopolysaccharide (LPS)-treated cells served as a positive control (Fig. 3D, E and Table S5).
Immunofluorescence assays marked F4/80+iNOS+ M1-like A and F4/80+CD206+ M2-like B RAW264.7 polarized by conditioned medium of 4T1-RICH1 OE/NC. C Relative mRNA expression level of inflammatory cytokines as macrophage M1 or M2 phenotypic markers in polarized RAW264.7. D, E Flow cytometry analysis, representative images including peak map and proportions of CD86+ or CD80+ subpopulations in polarized RAW264.7. F The volume and weight of the tumors were compared in the mice injected with 4T1-WT mixed with the polarized peritoneal-derived macrophages. G The proportions of F4/80+CD86+ subpopulations and F4/80+CD163+ subpopulations in CD45 positive cells of TME in the above tissues by flow cytometry analysis. Abbreviation: LPS lipopolysaccharide. P values in (C, E, F and G) were calculated using unpaired two-tailed Student’s t tests. *P < 0.05; **P < 0.01; ***P < 0.001.
We further evaluated the functional status of peritoneal-derived macrophages polarized using conditioned medium. To test their impact on tumor progression, these polarized macrophages were co-transplanted with wild-type 4T1 cells (4T1-WT) into mice. Results showed that in the group where macrophages polarized by 4T1-RICH1 OE-conditioned medium were mixed with 4T1-WT cells, both tumor volume and weight were significantly reduced compared to the control group (Figs. 3F and S4A and Table S6). The classification of TAMs infiltration in TME of mice bearing tumors was characterized by flow cytometry, as shown in Figs. 3G and S4B and Table S2. These findings indicate that RICH1 overexpression in tumor cells promotes the polarization of macrophages toward a functional M1-like TAM phenotype, contributing to tumor suppression.
RICH1 overexpression in breast cancer cells promotes macrophages toward M1-like polarization by secreting IFN-γ and chemokines
The role of elevated RICH1 expression in promoting M1-like macrophage polarization was confirmed. However, further investigation is required to identify the specific cytokines secreted by 4T1-RICH1 OE cells that drive this polarization. A comparative analysis of chemokine mRNA expression between 4T1-RICH1 OE and 4T1-NC revealed significantly higher levels of IFNG (Interferon-gamma), CXCL9, CXCL10, and CXCL11 in 4T1-RICH1 OE cells—cytokines known to induce M1 polarization. Conversely, expression levels of IL4 and IL13, which promote M2 polarization, were lower in 4T1-RICH1 OE cells (Fig. 4A and Table S7). To confirm these findings, we assessed the secretion levels of IFN-γ, CXCL9, IL-4, and IL-13 through enzyme linked immunosorbent assay (ELISA) between 4T1-RICH1 OE and 4T1-NC. Only the differences in IFN-γ and CXCL9 secretion were observed (Fig. 4B and Table S7). To further explore these findings, HC11 mammary epithelial cell lines with RICH1 knockout (HC11-RICH1 K/O1/2) were generated using CRISPR/Cas9 and monoclonal selection (Figs. 4C and S5). These cells, derived from the mammary tissue of pregnant Balb/c mice, exhibited significantly lower secretion levels of IFN-γ and CXCL9 compared to HC11-NC cells (Fig. 4D and Table S7).
A LogFC of mRNA expression level of chemokines, which polarize macrophages towards M1 or M2 in 4T1-RICH1 OE/NC. B The secretion level of IFN-γ, CXCL9, IL-4, and IL-13 was evaluated through ELISA between 4T1-RICH1 OE/NC. C The PCR products by RICH1 primers were analyzed by electrophoresis in agarose gels stained with ethidium bromide, which were extracted from HC11-RICH1 K/O1/2/ NC established using CRISPR/Cas9 RICH1/NC-targeted lentivirus and picking monoclonal. D The secretion level of IFN-γ, CXCL9, IL-4, and IL-13 was evaluated through ELISA between HC11-RICH1 K/O1/2/ NC. E The volume and weight of the tumors compared in the mice injected with 4T1-RICH1 OE shIFNG/4T1-RICH1 OE. F The proportions of CD86+ subpopulations in CD45+CD11b+F4/80+ cells of TME in the above tissues by flow cytometry analysis. Abbreviation: FC fold change, ELISA enzyme linked immunosorbent assay, PCR polymerase chain reaction. P values in (A, B, D, E and F) were calculated using unpaired two-tailed Student’s t tests. *P < 0.05; **P < 0.01; ***P < 0.001.
To further investigate the involvement of IFN-γ in breast cancer with high RICH1 expression, we orthotopically transplanted 4T1-RICH1 OE cells with IFNG knockdown (4T1-RICH1 OE shIFNG, see Fig. S6A) into female Balb/c mice. After a 21-day period, we observed a notable increase in both tumor volume and weight in the 4T1-RICH1 OE shIFNG group compared to the 4T1-RICH1 OE group (as depicted in Figs. 4E and S6B and Table S8). The 4T1-RICH1 OE shIFNG group exhibited a reduced population of M1-like TAMs in comparison to the 4T1-RICH1 OE group, as demonstrated by flow cytometry in Figs. 4F and S6C and Table S2. These findings imply that, in breast cancer with high RICH1 expression, IFN-γ secretion plays a pivotal role in shaping the immune-activated TME, contributing to increased M1-like TAM infiltration and tumor suppression. In addition, the role of IFN-γ in the immune microenvironment was evaluated for a potential link to hypoxia by measuring HIF-1α expression level in 4T1 tumor sections. A modest decrease in HIF-1α expression was observed in the RICH1-overexpressing group relative to controls by IHC (as shown in Fig. S6D).
RICH1 up-regulates the expression level of IFN-γ through binding with TRIM21 and inhibiting the activation of RhoA-STAT3 pathway
To elucidate the mechanism by which RICH1 enhances IFN-γ expression, we performed co-immunoprecipitation (Co-IP) followed by mass spectrometry on 4T1-RICH1 OE cells to identify proteins interacting with RICH1 (as shown in Fig. S7A). Among the identified candidates, tripartite motif-containing protein 21 (TRIM21), a gene associated with innate immunity, was highlighted (Fig. 5A and Table S9). Given that STAT3 plays a role in modulating the levels of chemokines, including IFN-γ, and the predictive interaction network involving RICH1, STAT3, and other immune-related molecules (as shown in Fig. 5B), it is necessary to verify the expression levels of RhoA, STAT3, and TRIM21 in response to RICH1 up-/down-regulation. The interaction between RICH1 and TRIM21 was confirmed through Western blotting following Co-IP and is presented in Figs. 5C and S7B, C. Furthermore, versus 4T1-NC, we observed a decrease in the total RhoA level, a reduction in the phosphorylation of STAT3 (p-STAT3), and an increase in the IFN-γ level in 4T1-RICH1 OE, while the corresponding results showed in HC11-RICH1 K/O1/2/ NC cell lines (Fig. 5D). IHC staining of consecutive paraffin sections from transplanted tumors revealed that the overexpression of RICH1 significantly inhibited the expression of RhoA and p-STAT3 while increasing the expression of IFN-γ (as illustrated in Fig. 5E and Table S10). These findings suggest that the up-regulation of IFN-γ by RICH1 may depend on the inhibition of the RhoA-STAT3 pathway, potentially mediated through its interaction with TRIM21.
A The top lists RICH1 binding proteins in MASS analysis (IP RICH1 in RICH1 overexpressed 4T1 cells), including RICH1 and its functionally related proteins, and innate immune-related TRIM21. B The protein-protein interaction network between RICH1, TRIM21, and RhoA-STAT3 identified by STRING database. RICH1, as a Rho GTPase activating protein, interacts with RhoA, and STAT3 was reported interacting with RhoA, TRIM21, and IFNG in previous studies. C Western blotting after Co-IP analysis for confirmation of interaction between RICH1-TRIM21 in 4T1 cells. D Detection of main above molecules after RICH1 overexpression and knockout by Western blotting. E IHC staining and score for RICH1, RhoA-STAT3, and IFN-γ of same fields on serial paraffin sections of 4T1 transplanted tumors. All the experiments were repeated three times independently with similar results. Abbreviation: Co-IP co-immunoprecipitation, MASS mass spectrometry, ARHGAP17 gene symbol for RICH1, HC heavy chain, IHC immunohistochemistry.
The binding of RICH1-TRIM21 makes a crucial role in the ubiquitin degradation of RhoA and non-activating of STAT3
To gain further insight into how the interaction between RICH1 and TRIM21 regulates the expression of RhoA, we assessed the RhoA half-lives in HC11-RICH1 K/O1/ NC cell lines and 4T1-RICH1 OE/NC. Our findings indicate that the presence of RICH1 reduces the half-life of RhoA, suggesting that RICH1 disrupts the stability of RhoA (as shown in Figs. 6A and S7D and Table S11). Furthermore, the reduction in RhoA levels could be reversed in cells expressing RICH1 when treated with MG132 or bortezomib. In the 4T1-RICH1 OE group, where RhoA levels are constitutively suppressed, MG132 or bortezomib treatment partially restored RhoA expression. Similarly, in HC11-NC cells with baseline low RhoA, MG132 or bortezomib increased RhoA protein abundance, consistent with proteasome-dependent degradation (as illustrated in Figs. 6B and S7E). To determine if RhoA is degraded through the ubiquitination pathway, we conducted immunoprecipitation experiments. In cells with RICH1 expression (4T1-RICH1 OE and HC11-NC), RhoA ubiquitination was significantly elevated compared to low-RICH1 controls, demonstrating that RICH1 promotes RhoA ubiquitination (as demonstrated in Figs. 6C and S7F). Furthermore, after confirming the regulation of RhoA degradation by RICH1 expression, we investigated whether the E3 ubiquitin ligase TRIM21 plays a role in this process. We validated the interaction between RhoA and TRIM21 through Western blotting following Co-IP. The results indicated that the binding interaction between RhoA and TRIM21 was more pronounced in 4T1-RICH1 OE than in the control lacking RICH1 (as depicted in Fig. 6D). This implies that the presence of RICH1 is pivotal for the binding of RhoA-TRIM21 and the subsequent ubiquitination degradation of RhoA. Additionally, the knockdown of TRIM21 by siRNA could abolish RICH1 overexpression-induced ubiquitination degradation of RhoA, down-regulation of RhoA and p-STAT3, and up-regulation of IFN-γ significantly (Figs. 6E and S7F). These results establish a mechanistic cascade wherein RICH1 recruits TRIM21 to ubiquitinate RhoA, targeting it for proteasomal degradation.
A Detection of RhoA expression after CHX (100 μg/ml) contacting HC11 cells with gradient time by Western blotting and quantification of RhoA protein levels determined using Image J, normalized to GAPDH. B Detection of RhoA expression after MG132 (5 μM) contacting 4T1 and HC11 cells with 8 h by Western blotting. C The ubiquitination of RhoA was examined by IP of RhoA and Western blotting with anti-ubiquitin antibody after ubiquitin-HA was transfected into RICH1 overexpression and knockout cells. D Western blotting after Co-IP analysis for confirmation of interaction between RhoA-TRIM21 in 4T1 cells with RICH1 overexpression. E Detection of the ubiquitination of RhoA and protein levels of RICH1, TRIM21, and RhoA-STAT3 by Western blotting after inhibiting the expression of TRIM21 in RICH1 overexpressed 4T1 cells. F, G Detection of the active form of RhoA (RhoA-GTP) in above cells by the pull-down assay. All the experiments were repeated three times independently with similar results. H, I The visualized image of molecular docking between RICH1 (yellow), TRIM21 (blue), and RhoA (green). J The volume and weight of the tumors compared in the mice injected with 4T1-RICH1 OE shTRIM21/4T1-RICH1 OE. K The proportions of CD86+ subpopulations in CD45+CD11b+F4/80+ cells of TME in the above tissues by flow cytometry analysis. Abbreviation: CHX cycloheximide, Co-IP co-immunoprecipitation, HC heavy chain. P values in (J and K) were calculated using unpaired two-tailed Student’s t tests. *P < 0.05; ***P < 0.001.
To further explore potential differences in RhoA activation in cells with or without RICH1 expression and their impact on STAT3 phosphorylation, we conducted a pull-down assay to assess the active form of RhoA (RhoA-GTP). The results demonstrated that the levels of both total RhoA and RhoA-GTP were lower in cells expressing RICH1 compared to those lacking RICH1 (Fig. 6F). Moreover, even when total RhoA down-regulation was prevented in cells with TRIM21 knockdown, the level of RhoA-GTP remained low in the presence of RICH1 (Fig. 6G). These findings suggest a dual role for RICH1: promoting TRIM21-mediated ubiquitin-driven degradation of RhoA and enhancing RhoA-GTP hydrolysis, thereby leading to RhoA deactivation and subsequent suppression of STAT3 phosphorylation.
To understand how TRIM21 binds to RICH1 and RhoA, molecular docking simulations were performed. These simulations indicated the concrete binding sites shown as Figs. 6H, I and S8A, B. Additionally, van der Waals forces and Pi-Pi interactions resulted in a binding energy score of −14.8 kcal/mol for RICH1-TRIM21 and −3.1 kcal/mol for RhoA-TRIM21 (Fig. S8C, D).
To verify whether disrupting TRIM21 abrogates the tumor-suppressive effects and pro-inflammatory TAM infiltration induced by RICH1 overexpression, we established TRIM21 knockdown in 4T1-RICH1 OE cells (denoted as 4T1-RICH1 OE shTRIM21) via lentiviral transduction (Fig. S9A) and subsequently generated an orthotopic transplantation mouse model. Compared to the 4T1-RICH1 OE group, the 4T1-RICH1 OE shTRIM21 group exhibited a significant increase in both tumor volume and weight (Figs. 6J and S9B, and Table S12). Flow cytometry analysis further revealed a reduced proportion of CD86+ TAMs in the 4T1-RICH1 OE shTRIM21 group relative to the 4T1-RICH1 OE controls (Figs. 6K and S9C and Table S2).
The correlation with M1 infiltration and RICH1 expression in breast cancer patients-the potential biomarkers of application with ICIs
To confirm the association between M1 infiltration and RICH1 expression in clinical samples, IHC staining of RICH1 and multiplex immunohistochemistry/immunofluorescence (mIHC/IF) of tumor immune microenvironment were conducted in 22 TNBC samples. The findings revealed a greater presence of CD68+HLA-DR+ M1-like TAMs in the TME of patients with high RICH1 expression compared to those with low RICH1 expression. Additionally, the group with a high ratio of M1 to total TAMs exhibited a higher RICH1 score (as depicted in Fig. 7A, B and Table S13; clinical patient information is provided in Table S14).
A The mIHC/IF of tumor immune microenvironment including tumor cells (PanCK), macrophages (CD68, HLA-DR), CD8+ T cells and natural killer cell (CD56) from TNBC samples. B IHC staining of RICH1, mIHC/IF of CD68+HLA-DR+ M1-like TAMs and the differential analysis of RICH1 expression in the groups with different proportion of M1/total TAMs from 22 TNBC samples. C The volume and weight of the tumors compared in the mice injected with 4T1-RICH1 OE/NC cells with 0.1 mg/kg sintilimab (anti-PD-1 mAb) or control normal sodium intraperitoneally injected 1 day, 8 days, 12 days, and 15 days after tumor cell implantation. D Representative peak images and proportions of CD86+ subpopulations in CD45+CD11b+F4/80+ cells of TME in the above tissues by flow cytometry analysis. E Schematic model of the role of RICH1 regulating M1-like TAMs infiltration in breast cancer. Abbreviation: ICI immune checkpoint inhibitor, mIHC/IF immunohistochemistry/immunofluorescence, TNBC triple negative breast cancer, PD-1 programmed cell death protein 1, NS normal sodium. P values in (B–D) were calculated using unpaired two-tailed Student’s t tests. *P < 0.05; **P < 0.01; ***P < 0.001; ns not significant.
To investigate whether RICH1 expression reduces tumor load via the “pro-inflammatory macrophages-T cell activation-tumor cell apoptosis” immune axis, RICH1 overexpression promoted the infiltration of CD45⁺CD3⁺CD8⁺ T cells and increased Granzyme B⁺ cytotoxic T cell populations (Fig. S10A and Table S2). Consistent with enhanced T cell activation, we observed a concurrent increase in Caspase 8 activity and enhanced Cleaved Caspase‑8/Pan-Cytokeratin co-staining, which correlated with elevated CD8⁺ T cell accumulation around apoptotic tumor cells (Fig. S10B–D).
Based on the observed alterations in the immune microenvironment across both human and mice samples, we further performed an omics-level investigation to validate whether RICH1 intervention elicits corresponding pathway changes. Transcriptomic profiling and microenvironment-focused KEGG analysis in RICH1-overexpressing versus control 4T1 cells confirmed enrichment of key immune-related pathways, including TNF, JAK-STAT, and NF-κB signaling (Fig. S11A–C), along with the RICH1-associated cGMP-PKG pathway. These results corroborate that RICH1 modulates immune pathways consistent with the phenotypic changes observed in vivo.
To explore the therapeutic implications of RICH1 in immune modulation, the anti-programmed cell death protein 1 (PD-1) monoclonal antibody (mAb), sintilimab, was administered to mice bearing 4T1-RICH1 OE/NC tumors. Anti-PD-1 mAb treatment resulted in a significant reduction in tumor volume and weight, with the most pronounced inhibition observed in the 4T1-RICH1 OE group receiving anti-PD-1 mAb therapy (as shown in Figs. 7C and S12A and Table S15). Additionally, the 4T1-RICH1 OE group treated with anti-PD-1 mAb showed the highest infiltration of M1-like TAMs among all experimental groups, as demonstrated by flow cytometry analysis (Figs. 7D and S12B and Table S2). These findings suggest that RICH1 may serve as an immune-related biomarker and a potential therapeutic target for combination strategies involving immune checkpoint inhibitors in TNBC.
Discussion
In our previous study20, we discovered that RICH1, known for its interaction with polarity complexes, is associated with a favorable prognosis and the inhibition of cancer stem cell-like properties in breast cancer. As a member of the RhoGAP family, RICH1 plays a potential role in shaping the immune microenvironment of breast cancer by regulating inflammatory signaling, as preliminarily revealed in this study using bioinformatics. In previous studies, abnormal expression of polarity proteins in tumor induced alteration of immune and inflammation-related signals24,25. Galvez et al.26 reported that the deletion of aPKCζ within the Par complex increased IL-6/STAT3-mediated pro-inflammatory signals, promoting the proliferation of lung cancer cells. Furthermore, Par3/aPKCλ-driven activation of STAT3 in inflammatory tumors affects the infiltration of IL-4-receptor-positive macrophages, promoting the formation of a tumor-permissive immune microenvironment16. Our current study is the first to demonstrate that the overexpression of RICH1 in breast cancer cells leads to increased infiltration of M1-like TAMs in TME, resulting in tumor inhibition in mouse models. This polarization of macrophages towards the M1 phenotype induced by RICH1 has also been confirmed in vitro. Prior investigations into the relationship between polarity proteins and macrophage polarization mainly relied on the use of small molecule agonists targeting polarity proteins27. In terms of RICH1 and its connection to inflammation-related signals, research has shown that RICH1 deficiency in mice led to enhanced intestinal epithelial cell permeability. Furthermore, the up-regulation of TNF and CXCL2 expression in tissues was observed exclusively in RICH1-deficient mice after treatment with dextran sulfate sodium, a known inducer of severe colitis28. Yang et al.29 reported that ARHGAP family genes are associated with pro-tumor immune microenvironment characteristics, such as exhausted T cell infiltration, in bladder cancer. In summary, the association between RICH1 and inflammatory factors was not clear in reported studies, and further elucidation of immune-related biological functions of RICH1 will provide new biomarkers and targets for combined immunotherapy in breast cancer.
This study identifies IFN-γ as the central cytokine driving RICH1-mediated M1 macrophage polarization. As established in the literature, IFN-γ and its induced chemokines CXCL9/10/11 promote M1-like macrophage polarization10. However, prior studies have mainly focused on immune cell-derived IFN-γ or tumor cell responses to IFN signaling30. In contrast to previous studies, our investigation reveals that RICH1 in tumor cells exerts an influence on their own secretion of IFN-γ, consequently leading to pro-inflammatory macrophage polarization in TME. Mechanistically, RICH1 upregulation in breast cancer cells may augment IFN-γ secretion, which both reinforces tumor-intrinsic IFN pathway activation and cytokine production, and directly polarizes macrophages toward a pro-inflammatory phenotype, thereby instigating a cascade amplification of immune activation within the TME.
During the exploration of the mechanism by which RICH1 up-regulates IFN-γ, the interaction between immune-related E3 ubiquitin ligase TRIM21 and RICH1 was discovered for the first time. While TRIM21 has been previously associated with innate immunity and autoimmune diseases like systemic lupus erythematosus and Sjogren’s syndrome, recent research has also implicated TRIM21 in tumor proliferation through the ubiquitination of cancer-related molecules31. Furthermore, TRIM21 plays a critical role in immunoinflammatory processes, particularly in the NF-κB and JAK-STAT pathways, where pivotal molecules like STAT3 are publicly recognized as regulatory factors for the production and secretion of IFN-γ11,32. For example, TRIM21 enhances JAK-STAT signaling via K63-linked ubiquitination of MAVS at Lys32533, and in keratinocytes, it stabilizes K17 through similar ubiquitination to drive STAT3 activation34, positioning TRIM21 as a therapeutic target in STAT3-driven cancers. Building on prior studies, RICH1, a member of the RhoGAP family, has been shown to activate RhoA and other Rho GTPase proteins19. Notably, RhoA-GTP has been linked to the promotion of STAT3 phosphorylation22. The RICH1-TRIM21 interaction integrates STAT3-mediated IFN-γ regulation, TRIM21-STAT3 association, and RhoA-dependent STAT3 phosphorylation. STRING database predictions confirmed RICH1/RhoA/STAT3/IFN-γ and TRIM21/STAT3 correlations. This study newly linked RICH1-TRIM21, bridging this network to explain RICH1’s control of IFN-γ levels. Consequently, our study unveils a previously undisclosed role for RICH1, acting as a tumor suppressor molecule in breast cancer. RICH1 promotes M1-like polarization of macrophages through a potential mechanism. In breast cancer cells with high RICH1 expression, RICH1 enhances the interaction between TRIM21 and RhoA, thereby facilitating the ubiquitination degradation of RhoA through binding with TRIM21, leading to the down-regulation of total RhoA protein levels. Simultaneously, RICH1 directly promotes the hydrolysis of RhoA-GTP, consequently reducing the levels of active RhoA. This process inhibits the phosphorylation of STAT3, up-regulates the production and secretion of IFN-γ, along with other cytokines, and ultimately induces M1-like polarization of macrophages (as illustrated in Fig. 7E).
In this study, we delved deeper into the relationship between RICH1, TRIM21, and RhoA. E3 ubiquitin ligase TRIM21, which interacts with RICH1 and facilitates the degradation of RhoA through the ubiquitination pathway, belongs to the TRIM family. Common domains among the more than 80 proteins in this family include the B-box domain, the C-C (coiled-coil) domain, and the RING domain with E3 ubiquitin ligase activity at the N-terminus31. The direct ubiquitination and degradation effect of TRIM21 on RhoA were initially reported in this study. Emerging studies establish that RING domain-containing E3 ligases, such as Cul3 and FBXL19, regulate RhoA stability through distinct mechanisms. Cul3 partners with BACURD adaptors to directly ubiquitinate and degrade RhoA, limiting its role in cytoskeletal remodeling35,36, while FBXL19 requires ERK2-mediated phosphorylation of RhoA to prime it for SCF complex-dependent degradation37. Building on these paradigms, we hypothesize that TRIM21-a RING-domain E3 ligase requiring oligomerization for activity38 may similarly target RhoA-GTP for ubiquitination. The results of molecular docking (the binding energy score of −14.8 kcal/mol for RICH1-TRIM21 and −3.1 kcal/mol for RhoA-TRIM21) and prior reports (e.g., TGM2-driven TRIM21-STAT1 dissociation via GTP binding39) suggest a mechanistic link: RICH1, a RhoGAP that hydrolyzes RhoA-GTP, might collaborate with TRIM21 to destabilize active RhoA. Specifically, RICH1’s GTPase-activating function may expose RhoA-GTP for recognition by TRIM21, whose RING domain is activated upon aggregation. This aggregation might be induced by RICH1 binding, enabling TRIM21 to polyubiquitinate RhoA-GTP and target it for proteasomal degradation. Critical to validating this model is defining the interacting domains, including TRIM21’s RING domain and RICH1’s RhoGAP domain. Future studies using truncation mutants will dissect these domain-specific roles, clarifying how RICH1-TRIM21 cooperation regulates RhoA-driven pathways in cancer.
In addition to the RhoA-mediated impact of RICH1 on STAT3 phosphorylation, as reported in this study, and the evidence of RhoA-mediated STAT3 phosphorylation in previous studies22, it is worth considering whether RhoA-GTP might also influence molecular signaling in other members of the STAT family. Considering the pro-tumor inflammatory effects of STAT3 and STAT5, as well as the anti-tumor inflammatory effects of STAT1 within the STAT family40, it remains to be explored and established whether RICH1 induces inflammatory effects in STATs other than STAT3 through the mediation of RhoA. Additionally, the existence of a more extensive RhoA-STAT signaling network in breast cancer warrants further investigation. Our clinical findings, derived from a limited TNBC cohort, provide preliminary insights into RICH1’s role in immune modulation but warrant validation in larger cohorts. While rigorous methodology strengthens internal validity, the small sample size and exclusive focus on TNBC limit generalizability to other breast cancer subtypes (e.g., Luminal, HER2+), which may exhibit distinct RhoA/STAT3 signaling dynamics. Particularly, more studies are emphasizing the importance of immunomodulatory effects in Luminal breast cancer5. Future studies incorporating diverse subtypes and expanded cohorts will clarify broader applicability.
In terms of clinical significance, this study employed sintilimab, an anti-PD-1 monoclonal antibody (mAb), to intervene in breast cancer formation in mice. The results indicated that breast cancer samples with high RICH1 expression exhibited a more robust response to ICI treatment than those with low expression. This heightened response could be attributed to the regulation of tumor-suppressive macrophages. While this study utilized murine cell lines to explore RICH1/TRIM21-mediated RhoA regulates pro-inflammatory TAM infiltration, we acknowledge potential interspecies divergence in tumor biology. Murine models, though invaluable for mechanistic interrogation under controlled conditions, may incompletely recapitulate human tumor microenvironments, particularly in immune signaling. Future validation in patient-derived xenografts or human macrophage-tumor cocultures will strengthen clinical relevance. In a mouse model of lung cancer xenografts, Wang et al.41 reported that sintilimab demonstrated a potent anti-tumor effect by increasing the secretion of IFN-γ and the presence of CD8+ T cells. Furthermore, aside from the immunomodulatory effect of ICI on CD8+ T cells in TME of breast cancer, recent studies have unveiled the interplay between ICI, CD8+ T cells, and TAMs. Emerging evidence indicates that pro-inflammatory macrophages modulate anti-tumor immunity by depleting or suppressing effector CD8+ T cell populations42,43. Consequently, the application of ICI and the modulation of RICH1 expression effectively inhibited tumor growth and significantly increased the infiltration of M1-like TAMs in this study. This phenomenon is likely due to the synergistic effect of RICH1 on M1 polarization, combined with the immunomodulatory function of ICI. However, given that ICI primarily enhances the activity of CD8+ T cells, it is important to investigate RICH1’s influence on T cells and the potential synergistic effects between T cells and macrophages. Our results suggested RICH1 overexpression enhanced the infiltration of CD8⁺ T cells and increased Granzyme B⁺ cytotoxic T cell frequency, and enhanced co‑localization of Caspase 8 with PanCK, which spatially correlated with CD8⁺ T cell accumulation around apoptotic tumor cells. These results suggest that RICH1 enhances pro‑inflammatory macrophage polarization, which in turn potentiates T cell‑mediated tumor apoptosis and antitumor immunity. Together with prior evidence that ICIs augment CD8⁺ T cell activity, our data support a model wherein the synergy between anti‑PD1 and RICH1 expression is initiated by macrophages and executed by T cells. Future studies should further define how RICH1 shapes the breast cancer immune landscape and elucidate bidirectional T cell–macrophage crosstalk. RICH1 expression in breast cancer cells remodels the immune microenvironment by polarizing macrophages toward a pro-inflammatory phenotype, suggesting that targeting RICH1 could amplify anti-tumor immunity. Building on the previous study and our findings, we will propose a dual-action immunotherapy in the future: a fusion antibody combining RICH1/TRIM21, enhancing macrophage-driven inflammation. To maximize efficacy, this strategy could synergize with PD-1/PD-L1 inhibitors, who reinvigorate cytotoxic T cells. Preclinical validation in RICH1-high models will prioritize biomarker-driven patient stratification, bridging mechanistic insights into RICH1-RhoA-STAT3 crosstalk with actionable immunotherapy design.
In conclusion, our findings demonstrate that RICH1 interacts with TRIM21 to suppress the RhoA-STAT3 signaling pathway, thereby promoting the infiltration of pro-inflammatory M1-like TAMs in breast cancer. These insights highlight the potential of RICH1 as an immune-related biomarker and a key contributor to the formation of immune-active microenvironments, with potential applications in combination immunotherapy strategies. Furthermore, this study underscores the clinical value of utilizing RICH1 to refine the precise classification of breast cancer based on distinct tumor microenvironment characteristics.
Methods
Patients and datasets
RNA sequencing data of samples from 1097 breast cancer patients in the TCGA database (http://portal.gdc.cancer.gov) were extracted. In addition, three datasets with breast cancer RNA expression data were obtained from Gene Expression Omnibus (GEO) database (http://www.ncbi.nlm.nih.gov/geo), including GSE42568, GSE103091, and GSE31519, in which the survival data are available. Formalin-fixed paraffin-embedded (FFPE) surgical tissue sections were from 22 TNBC patients admitted to the First Affiliated Hospital of Xi’an Jiaotong University, who didn’t suffer from other malignancy or received radiotherapy or chemotherapy. The patients in this study had signed consent from the Ethics Committee of the First Affiliated Hospital of Xi’an Jiaotong University (Ethics Approval Number: 2020-G-173), and the study was conducted in conformity to the Declaration of Helsinki.
Gene set enrichment analysis (GSEA) and estimates of immune cells
GSEA was performed by using GSEA version 4.1.0 (http://software.broadinstitute.org/gsea/index.jsp), and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis was obtained through gene sets from the Broad Institute (http://ftp.broadinstitute.org://pub/gsea/gene_sets/c2.cp.kegg.v6.2.symbols.gmt).
The CIBERSORT deconvolution algorithm was used to estimate the fraction of 22 immune cell types in each breast cancer sample44, which was calculated via the online calculator (https://cibersort.stanford.edu/), and the CIBERSORT results were filtered by p < 0.05 to obtain more accurate simulated results.
Analysis for spatial transcriptomics
Spatial transcriptomics data of breast cancer tissue (Block ID: 1168993F) were obtained from the Visium platform (https://www.10xgenomics.com/platforms/visium). Seurat clustering default algorithms (FindNeighbors and FindClusters) were used to define tissue regions, and gene expression of tumor tissues was combined with spatial information to generate detailed spatial omics maps. Cluster-specific gene expression patterns were detected using the Seurat differential search tool (FindAllMarkers), adjusting the number of clusters to the maximum. BulkSignalR and project-specific R scripts were conducted for gene spatial expression analysis45, and the spatial distribution of immune cells was also defined by corresponding gene signatures.
Cell culture and reagents
4T1, RAW264.7, and HC11 cells were purchased from Beijing Union Medical College Hospital cell resource-sharing platform. SUM159 cells were a gift from Dr. Jianjun He. 4T1 was cultured in RPMI 1640 medium with 10% fetal bovine serum (FBS) and 1% penicillin-streptomycin (P-S). SUM159 was cultured in DMEM/F12 medium with 10% FBS and 1% P-S. RAW264.7 was grown in DMEM/high glucose medium with 10% FBS and 1% P-S. HC11 was cultured in RPMI 1640 medium with 10% FBS, 1% sodium pyruvate, and 1% P-S. Medium, FBS, and P-S were purchased from Corning. All the cell lines were cultured in a humidified incubator at 37 °C, 5% CO2. Antibodies against RICH1 (ab229221, dilution 1:500), F4/80 (ab90247, dilution 1:50), iNOS (ab178945, dilution 1:100), TRIM21 (ab207728, dilution 1:1000) were purchased from Abcam. Antibodies against RhoA (sc-418, dilution 1:500) and ubiquitin (sc-8017, dilution 1:1000) were purchased from Santa Cruz. Antibody against IFN-γ was purchased from Abmart (T56671, dilution 1:1000). Antibodies against STAT3 (#9139, dilution 1:500), phosphor-STAT3Tyr705 (#9145, dilution 1:1000), Cleaved Caspase‑8 (#8592, dilution 1:500), and Pan-Cytokeratin (#4545, dilution 1:200) were purchased from Cell Signaling Technology. Antibodies against CD8 (#14-0808-82, dilution 1:200) were purchased from Invitrogen. HRP-labeled GAPDH (HRP-60004, dilution 1:10000) and antibody against CD206 (60143-1-Ig, dilution 1:1000) were from Proteintech.
Vector construction and cell transfection
Mouse RICH1 overexpression plasmid vector and packed lentivirus were obtained from Hanbio (Shanghai, China). 4T1 cells were infected with lentivirus for RICH1 overexpression/negative control (RICH1 OE/NC) as MOI (multiplicity of infection) 10, and stable RICH1 overexpressed cells were selected under the deal with medium containing neomycin (MedChemExpress, USA) for a week. SUM159-RICH1 OE/NC were infected as our previous study16. Mouse CRISPR/Cas9 RICH1 knockout/NC-targeted lentivirus was obtained from Hanbio. HC11 cells were infected with lentivirus for RICH1 knockout/NC (RICH1 K/O/ NC) as MOI 20, and 2 monoclonal RICH1 knockout cell lines were picked by limiting dilution cloning in the medium containing puromycin (MedChemExpress). RICH1 overexpressed 4T1 cells were infected with lentivirus for shRNA (small hairpin RNA)-IFNG (obtained from Hanbio) as MOI 10 under the selection of medium containing puromycin. All cells were infected using virus transfection reagents polybrene (Hanbio). In addition, mouse TRIM21 siRNA molecules were obtained from GenePharma (Shanghai, China) which were transfected into cells using Lipofectamine 2000 (Invitrogen, Carlsbad, CA). All sequences were listed in Table S16.
In vivo tumor experiments
All animal studies were conducted in accordance with the guidelines provided by the Animal Ethics Committee of Xi’an Jiaotong University (Ethics Approval Number: 2020-G-173). Six-week female Balb/c mice were obtained from the Laboratory Animal Center of Xi’an Jiaotong University and housed in a special pathogen-free animal facility. All mice were randomly assigned to either the experimental or control group, with six mice per group. For orthotopic transplantation, 5 × 105 4T1 cells diluted in 200 μl PBS were injected into the fourth mammary fat pad on the right side. 5 × 105 SUM159 cells diluted in 100 μl PBS were transplanted into 4 weeks female Balb/c nude mice. For macrophage depletion in vivo experiment, clophosome (FormuMax, F70101C-NF, 20 mg/mL) was used, which could disrupt liposomal membranes, releasing the water-soluble prodrug chlorophosphonic acid into cells. Upon reaching its minimal effective concentration, chlorophosphonic acid triggers irreversible apoptosis in macrophages, eliminating them via caspase-dependent pathways. Each mouse in the macrophage depletion group or the control group was intraperitoneally injected with 200 μL clophosome or the control liposomes for the first time on the day 0 of tumor transplantation, and then injected with 100 μL twice a week.
Mouse peritoneal derived macrophages were induced by intraperitoneal injection of zymosan A (Sigma) into 8 weeks female Balb/c mice. For the function of the peritoneal derived macrophages polarized by conditioned medium, 5 × 105 4T1 cells mixed with 1.5 × 106 macrophages diluted in 200 μl PBS were orthotopically transplanted. For PD-1 treatment, 0.1 mg/kg sintilimab was intraperitoneally injected 1 day, 8 days, 12 days, and 15 days after tumor cell implantation, and the control group was peritoneal injection with the same amount of normal sodium. The mice were euthanized via inhalation anesthesia with carbon dioxide after 21 days or when the longest tumor diameter exceeded 1.5 cm. The tumor tissues were removed, after which tumor weights were measured and tumor volumes were calculated by the formula (a × b2)/2, where a is the major tumor axis, and b is the minor tumor axis.
Immunohistochemistry (IHC)
The FFPE tissue sections were roasted at 60 °C for 4 h, deparaffinized in xylene, and hydrated in gradient concentration of ethanol. Tissue slides were performed in sodium citrate buffer (PH = 9.0) at the microwave oven subsequently, for antigen recovery. IHC kit (Solarbio, SP0041), including 3% hydrogen peroxide and goat serum albumin were applied, and the slides were incubated with primary antibodies overnight at 4 °C. Next day, the biotin-labeled homologous secondary antibody and horseradish peroxidase-labeled streptomycin ovalbumin in IHC kit were applied on the slides. Then, DAB (diaminobenzidine) staining, hematoxylin staining, hydrochloric ethanol solution differentiation, returning to blue in ammonia, dehydration in gradient ethanol, and transparency in xylene were followed to handle the slides. We selected 10 fields of each tissue section randomly for semi-quantitative scoring as reported previously20, and the average IHC score of each sample was counted.
Flow cytometry
For marking TAMs of tissues, the fresh tumor tissues were digested and sieved, followed with removing red blood cells to make single cell suspension diluted in PBS. PerCP/Cyanine5.5-conjugated CD45 antibody (Biolegend, #103132), PE/Cyanine7-conjugated F4/80 antibody (Biolegend, #123113), FITC-conjugated CD86 antibody (Biolegend, #105005) and PE-conjugated CD163 antibody (Biolegend, #155307) were used to detect CD45+F4/80+CD86+ or CD45+F4/80+CD163+ subpopulations, and the above antibodies with APC-conjugated CD11b antibody (Biolegend, #101212), PE-conjugated NOS2 antibody (Biolegend, #696805), Brilliant Violet 421-conjugated TNFα antibody (Biolegend, #506327), FITC-conjugated CD80 antibody (Biolegend, #104705) were used to detect CD45+CD11b+F4/80+CD86+ or NOS2+ or TNFα+ or CD80+ subpopulations. APC/Cyanine7-conjugated CD3 antibody (Biolegend, #100221), APC-conjugated CD4 antibody (Biolegend, #100411), FITC-conjugated CD8a antibody (Biolegend, #100705), and PE-conjugated Granzyme B antibody (Biolegend, #396405) were used to detect CD3⁺CD4⁺/CD8⁺Granzyme B⁺ T cells subpopulations. Flow cytometry controls included isotype-matched antibodies and single-color stains to ensure specificity and resolve spectral overlap. The marked single cell suspension was analyzed by Canto II Flow Cytometer (BD), and the data were processed using FlowJo 10.6.2. For marking different phenotypes of macrophages polarized by conditioned medium, RAW264.7 cells under the deal with conditioned medium of 4T1 cells for 48 h were collected and resuspended with PBS. The positive control group was under the treatment of 1 μg/mL LPS, then F4/80+CD86+ subpopulations were detected and analyzed. In the process described above, <1 × 106 cells in each flow tube were incubated with antibodies of recommended concentration at 37 °C for 30 min in dark, and permeabilization following with fixation were processed before intracellular cytokine staining.
Immunofluorescence and mIHC/IF
For slides of mouse tumor tissues that had been frozen, embedded with OCT, and sectioned, 4% paraformaldehyde and 5% bovine serum albumin (BSA) were applied for fixation and blocking separately. Slides were incubated with specific primary antibodies overnight at 4 °C, followed by fluorescent-labeled secondary antibodies (Proteintech) next day, and DAPI was used for DNA staining. Image acquisition was performed by Leica confocal laser scanning microscopy. Except for the above procedures, RAW264.7 cells adhering to slides were permeabilized with 0.2% Triton X-100.
The FFPE tissue sections were performed for multiplex immunohistochemistry/immunofluorescence (mIHC/IF) staining, and the estimated proportions of tumor-infiltrating immune cells were conducted as we reported previously46.
RNA isolation, real-time PCR, and RNA sequencing
Total RNA from cell lines was extracted using the RNeasy mini kit (Qiagen, Valencia, CA) and then converted to cDNA using the PrimeScriptTM RT Master Mix (Takara). For detecting the mRNA expression levels, real-time PCR was conducted with the SYBR-Green kit (Takara). The primers used for real-time PCR in this study were listed in Table S17 and obtained from Sangon Biotech. Following extraction with Trizol reagent, RNA quality was assessed on an Agilent Bioanalyzer 2100 system. Sequencing libraries were prepared from qualified RNA via poly-A selection and cDNA synthesis, followed by cluster generation and 150 bp paired-end sequencing on an Illumina Novaseq platform.
Western blotting
Whole-cell lysates were prepared by RIPA buffer, then proteins were separated with SDS/PAGE gel and transferred to PVDF membranes, which were incubated overnight with primary antibodies. HRP-conjugated secondary antibodies (Cell Signaling Technology) were incubated, and chemiluminescent signals were detected by ECL (Millipore) in the next day.
ELISA and pull-down assay
The concentrations of IFN-γ, CXCL9, IL-4, and IL-13 in the cell medium were measured using enzyme linked immunosorbent assay (ELISA) kits specific for mouse IFN-γ, CXCL9, IL-4, and IL-13 (Multi Sciences, China). RhoA-GTP in cells was pulled down and detected using Rho activation assay biochem kit (Cytoskeleton, #BK036) in accordance with the manufacturer’s protocol. Caspase 8 activity in tumor homogenates was measured using ELISA kits (BestBio, BB-4107).
Co-immunoprecipitation (Co-IP) and mass spectrometry
Total protein of cells was extracted by NP-40 lysis buffer (20 mM Tris-HCl, 150 mM NaCl, 0.5% NP-40, 20% glycerol) containing protease inhibitor and phosphatase inhibitors (Roche, NJ, USA). The lysates were immediately centrifuged at 12 × 103 rpm, 4 °C for 20 min, and then were incubated with magnetic beads to prepare bead-antigen-antibody complexes using 4 μg antibody and 50 μl protein A/G magnetic beads (Thermo Fisher, 10007D). The final binding protein was obtained by washing with elution buffer, then protein complexes were boiled and prepared to Western blotting.
For mass spectrometry analysis of potential RICH1-binding proteins, the eluted samples were separated with 10% SDS-PAGE and stained with colloidal coomassie. The obtained gel lanes were cut into defined pieces, de-stained, and trypsinized. The resulting peptide solutions were extracted, subjected to mass spectrometry (Q ExactiveTM Plus, ThermoFisher Scientific, USA), processed using Proteome Discoverer 2.1 for secondary mass spectrum data. When we verified the co-immunoprecipitation of RICH1-TRIM21, the HRP-conjugated secondary antibody in Western blotting was conformation specific for not binding to IgG heavy chain (Cell Signaling Technology #5127).
Ubiquitination assay
Plasmid expressed ubiquitin-HA was transiently infected to cells by Lipofectamine 2000 (Add 2 μg of plasmid to 500 μL of serum-free medium and let it stand for 5 minutes). After 24 h, the cells were treated with 5 μmol/L MG132 (Sigma) for 8 h. Then, Co-IP was conducted with RhoA antibody for NP-40 lysates of the cells, and Western blotting was applied for detecting ubiquitinated-RhoA.
Prediction for molecular docking
Using cluspro (https://cluspro.org/) to butt joint of protein, the model imports PDBePISA (https://www.ebi.ac.uk/pdbe/pisa/) to calculate the binding energy, patterns, and good structure drawing. Pymol (https://pymol.affinitycn.cn) was used to draw 3D and 2D maps. Ligplot 2.2.9 was used to draw 2D maps to show docking sites and interaction residues.
Statistical analysis
All experiments in vitro were repeated at least three times, and at least 5 mice were set up in each group for all animal experiments. All data were analyzed with Graphpad Prism 8. These results were described as the mean ± S.D. The differences between two groups were compared using Student’s two-sided t-test. Differences of survival between different groups were compared using Kaplan–Meier analysis followed with log-rank test. The correlation between RICH1 expression levels and clinical characteristics of TNBC patients’ samples was analyzed using Fisher’s exact test. P < 0.05 was considered statistically significant.
Correlation analyses between RICH1 levels and immune cell infiltration were performed as follows: (1) Public Dataset Analysis: RNA-seq data were stratified into high- and low-RICH1 expression groups. Immune cell proportions were inferred using CIBERSORT and TIMER, followed by differential analysis (Wilcoxon rank-sum test) and Pearson correlation analysis. (2) Experimental Validation: In breast cancer patient tissue specimens, macrophage infiltration was quantified via mIHC/IF. In murine tumor models, macrophage populations were assessed by flow cytometry. Group comparisons (high vs. low RICH1) were analyzed using Student’s t test or Mann–Whitney U test, with statistical significance set at P < 0.05 for all analyses.
Data availability
All datasets are publicly available in the corresponding repositories via their respective accession numbers. RNA sequencing data of samples from TCGA database and three datasets from Gene Expression Omnibus (GEO) database, including GSE42568, GSE103091, and GSE31519, were publicly extracted. Spatial transcriptomics data of breast cancer tissue (Block ID: 1168993F) were publicly available at the Visium platform. Bioinformatics analyses were performed using established R packages, with no custom scripts utilized. Raw experimental data are provided in the Supplementary Information.
References
Heater, N. K., Warrior, S. & Lu, J. Current and future immunotherapy for breast cancer. J. Hematol. Oncol. 17, 131 (2024).
Lehmann, B. D. et al. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J. Clin. Investig. 121, 2750–2767 (2011).
Zhao, S. et al. Molecular subtyping of triple-negative breast cancers by immunohistochemistry: molecular basis and clinical relevance. oncologist 25, e1481–e1491 (2020).
Gruosso, T. et al. Spatially distinct tumor immune microenvironments stratify triple-negative breast cancers. J. Clin. Investig. 129, 1785–1800 (2019).
Jin, X. et al. Molecular classification of hormone receptor-positive HER2-negative breast cancer. Nat. Genet. 55, 1696–1708 (2023).
Wu, S. Y., Wang, H., Shao, Z. M. & Jiang, Y. Z. Triple-negative breast cancer: new treatment strategies in the era of precision medicine. Sci. China Life Sci. 64, 372–388 (2021).
Coussens, L. M. & Werb, Z. Inflammation and cancer. Nature 420, 860–867 (2002).
Wu, K. et al. Redefining tumor-associated macrophage subpopulations and functions in the tumor microenvironment. Front. Immunol. 11, 1731 (2020).
Mantovani, A. & Sica, A. Macrophages, innate immunity and cancer: balance, tolerance, and diversity. Curr. Opin. Immunol. 22, 231–237 (2010).
Schroder, K., Hertzog, P. J., Ravasi, T. & Hume, D. A. Interferon-gamma: an overview of signals, mechanisms and functions. J. Leukoc. Biol. 75, 163–189 (2004).
Groner, B. & von Manstein, V. Jak Stat signaling and cancer: opportunities, benefits and side effects of targeted inhibition. Mol. Cell. Endocrinol. 451, 1–14 (2017).
Sica, A., Allavena, P. & Mantovani, A. Cancer related inflammation: the macrophage connection. Cancer Lett. 267, 204–215 (2008).
Halaoui, R. & McCaffrey, L. Rewiring cell polarity signaling in cancer. Oncogene 34, 939–950 (2015).
Mastrogiovanni, M., Di Bartolo, V. & Alcover, A. Cell polarity regulators, multifunctional organizers of lymphocyte activation and function. Biomed. J. 45, 299–309 (2022).
Barreda, D. et al. The scribble complex PDZ proteins in immune cell polarities. J. Immunol. Res. 2020, 5649790 (2020).
Vorhagen, S. et al. Shared and independent functions of aPKCλ and Par3 in skin tumorigenesis. Oncogene 37, 5136–5146 (2018).
Wells, C. D. et al. A Rich1/Amot complex regulates the Cdc42 GTPase and apical-polarity proteins in epithelial cells. Cell 125, 535–548 (2006).
Beck, S., Fotinos, A., Lang, F., Gawaz, M. & Elvers, M. Isoform-specific roles of the GTPase activating protein Nadrin in cytoskeletal reorganization of platelets. Cell. Signal. 25, 236–246 (2013).
Kobayashi, Y. et al. The role of NADRIN, a Rho GTPase-activating protein, in the morphological differentiation of astrocytes. J. Biochem. 153, 389–398 (2013).
Tian, Q. et al. RICH1 inhibits breast cancer stem cell traits through activating kinases cascade of Hippo signaling by competing with Merlin for binding to Amot-p80. Cell Death Dis. 13, 71 (2022).
Crosas-Molist, E. et al. Rho GTPase signaling in cancer progression and dissemination. Physiol. Rev. 102, 455–510 (2022).
Campbell, H. et al. ∆133p53 isoform promotes tumour invasion and metastasis via interleukin-6 activation of JAK-STAT and RhoA-ROCK signalling. Nat. Commun. 9, 254 (2018).
Luo, C. et al. Inhibition of Rho GEFs attenuates pulmonary fibrosis through suppressing myofibroblast activation and reprogramming profibrotic macrophages. Cell Death Dis. 16, 278 (2025).
Bergstralh, D. T. & St Johnston, D. Epithelial cell polarity: what flies can teach us about cancer. Essays Biochem. 53, 129–140 (2012).
Diaz-Meco, M. T. & Moscat, J. The atypical PKCs in inflammation: NF-κB and beyond. Immunol. Rev. 246, 154–167 (2012).
Galvez, A. S. et al. Protein kinase Czeta represses the interleukin-6 promoter and impairs tumorigenesis in vivo. Mol. Cell. Biol. 29, 104–115 (2009).
Chen, L. et al. PAR2 promotes M1 macrophage polarization and inflammation via FOXO1 pathway. J. Cell. Biochem. 120, 9799–9809 (2019).
Lee, S. Y. et al. Arhgap17, a RhoGTPase activating protein, regulates mucosal and epithelial barrier function in the mouse colon. Sci. Rep. 6, 26923 (2016).
Yang, C. et al. Transcriptomic analysis identified ARHGAP family as a novel biomarker associated with tumor-promoting immune infiltration and nanomechanical characteristics in bladder cancer. Front. Cell Dev. Biol. 9, 657219 (2021).
Villarino, A. V., Kanno, Y. & O’Shea, J. J. Mechanisms and consequences of Jak-STAT signaling in the immune system. Nat. Immunol. 18, 374–384 (2017).
Alomari, M. TRIM21 - A potential novel therapeutic target in cancer. Pharmacol. Res. 165, 105443 (2021).
Zou, S. et al. Targeting STAT3 in cancer immunotherapy. Mol. Cancer 19, 145 (2020).
Xue, B. et al. TRIM21 Promotes innate immune response to RNA viral infection through Lys27-linked polyubiquitination of MAVS. J. Virol. 92, e00321–18 (2018).
Yang, L. et al. E3 ligase Trim21 ubiquitylates and stabilizes keratin 17 to induce STAT3 activation in psoriasis. J. Investig. Dermatol. 138, 2568–2577 (2018).
Chen, Y. et al. Cullin mediates degradation of RhoA through evolutionarily conserved BTB adaptors to control actin cytoskeleton structure and cell movement. Mol. Cell 35, 841–855 (2009).
Panek, J. et al. A cullin-RING ubiquitin ligase promotes thermotolerance as part of the intracellular pathogen response in Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 117, 7950–7960 (2020).
Wei, J. et al. A new mechanism of RhoA ubiquitination and degradation: roles of SCF(FBXL19) E3 ligase and Erk2. Biochim. Biophys. Acta 1833, 2757–2764 (2013).
Miller, L. V. C. et al. Co-opting templated aggregation to degrade pathogenic tau assemblies and improve motor function. Cell 187, 5967–5980.e5917 (2024).
Zhang, L. et al. Cytosolic TGM2 promotes malignant progression in gastric cancer by suppressing the TRIM21-mediated ubiquitination/degradation of STAT1 in a GTP binding-dependent modality. Cancer Commun. 43, 123–149 (2023).
Thomas, S. J., Snowden, J. A., Zeidler, M. P. & Danson, S. J. The role of JAK/STAT signalling in the pathogenesis, prognosis and treatment of solid tumours. Br. J. Cancer 113, 365–371 (2015).
Wang, J. et al. Durable blockade of PD-1 signaling links preclinical efficacy of sintilimab to its clinical benefit. mAbs 11, 1443–1451 (2019).
Waibl Polania, J. et al. Antigen presentation by tumor-associated macrophages drives T cells from a progenitor exhaustion state to terminal exhaustion. Immunity 58, 232–246.e236 (2025).
Luo, F. et al. The BCL-2 inhibitor APG-2575 resets tumor-associated macrophages toward the M1 phenotype, promoting a favorable response to anti-PD-1 therapy via NLRP3 activation. Cell. Mol. Immunol. 21, 60–79 (2024).
Pagès, F. et al. Effector memory T cells, early metastasis, and survival in colorectal cancer. N. Engl. J. Med. 353, 2654–2666 (2005).
Villemin, J. P. et al. Inferring ligand-receptor cellular networks from bulk and spatial transcriptomic datasets with BulkSignalR. Nucleic Acids Res. 51, 4726–4744 (2023).
Zhou, Y. et al. Correlation between immune-related genes and tumor-infiltrating immune cells with the efficacy of neoadjuvant chemotherapy for breast cancer. Front. Genet. 13, 905617 (2022).
Acknowledgements
We thank The Cancer Genome Atlas (TCGA) and Gene Expression Omnibus (GEO) for providing transcriptomics and clinicopathological data. We thank Ruiqi Wang and He Chen for their help with tumor section preparation. We thank National Natural Science Foundation of China (This study was supported by Nos. 82002794, 82173277, and 82303462).
Author information
Authors and Affiliations
Contributions
Y.Z. conceived the project and conducted all experiments. L.Z.Z., J.Z., and R.R. were responsible for clinical sample collection. H.G. and J.Y. analyzed the data. B.W. and L.Y.S. provided support with experimental techniques. P.J.L. and Q.T. supervised all experiments. Y.Z. and Y.J. constructed the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhou, Y., Gao, H., Shan, L. et al. RICH1 enhances pro-inflammatory TAM infiltration in breast cancer via promoting TRIM21-mediated ubiquitination of RhoA and inhibiting STAT3 phosphorylation. npj Precis. Onc. 10, 49 (2026). https://doi.org/10.1038/s41698-025-01252-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-025-01252-6