Abstract
Occult pathological T3a (pT3a) upstaging in cT1b–T2a clear cell renal cell carcinoma (ccRCC) correlated with poor prognosis and necessitated modifications in surgical planning. However, predicting it preoperatively remains challenging. In this multicenter study involving 1661 patients across five institutions and the KiTS23 dataset, RENALNet, a 3D deep learning framework trained on nephrographic-phase CT, was developed and validated. RENALNet outperformed radiomics models, further enhancing diagnostic accuracy when combined with radiologists of varying experience. Grad-CAM visualizations concentrated on anatomically significant invasion sites, improving interpretability. Risk scores derived from RENALNet were found to correlate with Ki-67 proliferation indices and effectively stratified 5-year progression-free survival, demonstrating both biological and prognostic relevance. Transcriptomic analysis revealed that high RENALNet risk was associated with gene expression signatures enriched in pathways such as epithelial–mesenchymal transition, IL6–JAK–STAT3 signaling, and PI3K–Akt signaling, highlighting its link to tumor aggressiveness. RENALNet thus offers a biologically interpretable framework for risk stratification in ccRCC, supporting surgical decision-making and advancing the integration of radiogenomic deep learning into precision oncology.
Similar content being viewed by others
Introduction
Clear cell renal cell carcinoma (ccRCC) comprises approximately 70% of all renal malignancies, with up to 70% of cases diagnosed at an early stage1,2. Accurate staging is essential for guiding surgical decisions and optimizing oncologic outcomes1. Nephron-sparing surgery (NSS) is the preferred treatment for clinical T1 tumors, particularly those smaller than 3 cm3. Reflecting this, the European Association of Urology (EAU) Guidelines Panel recently revised the cT1a definition, lowering the size threshold from 4 cm to 3 cm4. Additionally, NSS is increasingly considered for tumors ≤ 10 cm in select cases, depending on tumor complexity and surgical feasibility5. However, 10–20% of patients initially diagnosed with localized disease are later upstaged to pathological T3a (pT3a) at final pathology after surgery6,7. pT3a upstaging, characterized by invasion of perirenal or sinus fat, pelvicalyceal system, or extending into the renal veins or their segmental branches, is associated with a significantly higher risk of recurrence and mortality, and often requires radical nephrectomy (RN)8,9.
Although conventional CT and MRI are essential for preoperative RCC staging, they have limited sensitivity in detecting microscopic invasion. The reported accuracy of CT for identifying perinephric fat invasion ranges from 32% to 64%10, and conditions like inflammation or fibrosis may mimic invasion. While MRI provides superior soft-tissue contrast, it is hindered by longer scan times, higher costs, and limited availability. Existing predictive models have largely concentrated on broad staging categories11,12 or specific invasion patterns, such as perinephric fat invasion13, but often lack interpretability, limiting their clinical utility. Given that pT3a tumors extend beyond the renal parenchyma, integrating both peritumoral and intratumoral data could improve predictive accuracy14. These challenges highlight the need for accurate, interpretable tools to predict occult pT3a upstaging in cT1b–T2a ccRCC.
Advancements in radiomics and deep learning present new avenues to address this challenge. Radiomics enables the extraction of high-dimensional, quantitative features from CT images, capturing tumor morphology and heterogeneity15. Meanwhile, deep learning models can autonomously identify complex imaging patterns related to tumor biology, eliminating the need for manual feature engineering16. Despite the increasing use of these techniques in renal cancer research17,18,19, few studies have specifically targeted pT3a upstaging in cT1b–T2a ccRCC, and even fewer have incorporated interpretability or biological validation. Integrating imaging-based predictions with molecular and prognostic data can offer essential biological insights and facilitate clinical translation.
This multicenter study developed and validated RENALNet (Residual Enhancement Niche Attention for Lesion Classification Network), an interpretable deep learning framework for the noninvasive prediction of occult pT3a upstaging in cT1b–T2a ccRCC, demonstrating robust and superior performance compared to radiomics-based models. Grad-CAM visualizations revealed that RENALNet consistently focused on anatomically relevant invasion sites, enhancing interpretability and providing clinically actionable insights. Further integrative analyses showed that high RENALNet risk scores were associated with elevated Ki-67 proliferation indices and transcriptomic signatures, including epithelial–mesenchymal transition (EMT) and IL6–JAK–STAT3 signaling, thereby linking imaging-derived risk with tumor aggressiveness. These critical findings were validated across four external clinical cohorts and transcriptomic data from TCGA-KIRC, confirming their robustness. This integrative framework not only supports precise and individualized surgical planning in ccRCC but also exemplifies how radiogenomic deep learning can bridge imaging, biology, and outcomes to advance precision oncology.
Results
Patient characteristics
The detailed workflow of patient selection is presented in Fig. 1. A total of 1661 patients with cT1b–T2a ccRCC were finally included (median age 59.9 years [range, 24–85]; 66% male). The development dataset comprised 788 patients from Tongji Hospital (January 2015–June 2022) and Xiangyang Central Hospital (July 2021–July 2024), who were randomly assigned in an approximately 7:3 ratio to a training cohort (n = 551) and a testing cohort (n = 237). Four external validation cohorts comprised 296 patients from Jiangsu Provincial People’s Hospital, 234 from Henan Cancer Hospital, 187 from Shanxi Bethune Hospital, and 156 from the KiTS23 dataset.
Study enrollment process flowchart.
Baseline demographic and clinical characteristics were well balanced across cohorts (Table 1). In the training cohort (n = 551), 60 (10.9%) patients were pathologically upstaged to pT3a. Upstaging rates in the testing cohort (n = 237) and four external validation cohorts (n = 296, 187, 234, and 156) were 11.0%, 7.1%, 23.0%, 10.7%, and 14.7%, respectively.
Radiomics model performance
Three classifiers were developed using peritumoral features (Radiomics_Peritumor), intratumoral features (Radiomics_Tumor), and their combination (Radiomics_Combined). In the testing cohort, the Radiomics_Combined model demonstrated the highest performance (AUC, 0.800 [95% CI, 0.709–0.892]; accuracy, 0.899; specificity, 0.994), although sensitivity remained low (0.154), consistent across all radiomics models. In all external validation cohorts, Radiomics_Combined consistently outperformed models based on single-region features. For instance, in Validation Cohort IV, the AUC of Radiomics_Combined was 0.751, compared to 0.664 for Radiomics_Tumor and 0.685 for Radiomics_Peritumor.
Although DeLong’s test did not demonstrate statistically significant differences between Radiomics_Combined and Radiomics_Tumor across cohorts, the consistently higher AUC values of the combined model suggest an added benefit of integrating intra- and peritumoral features (Fig. 2 and Tables 2 and 3).
A Area under the receiver operating characteristic (ROC) curves (AUC) of three radiomics-based models and the RENALNet deep learning model across the testing and four external validation cohorts. B Confusion matrices of the RENALNet model in the testing cohort and four external validation cohorts.
Deep learning model performance
RENALNet achieved perfect convergence in the training cohort (AUC, 1.00; sensitivity, 100%; specificity, 89.1%; accuracy, 99.5%). In the internal testing cohort, performance remained robust (AUC, 0.818; sensitivity, 53.8%; specificity, 99.4%; accuracy, 85.2%). Across multicenter external validation cohorts, RENALNet achieved AUCs ranging from 0.696 to 0.764 and accuracies between 76.1% and 88.5%.
Compared with radiomics models, RENALNet consistently outperformed all three radiomics-based models in terms of AUC, and exhibited substantially higher sensitivity (33.3%–46.5% vs. 7.0%–28.6%), improving detection of occult pT3a disease, albeit with moderately lower specificity (80.4%–92.7%). DeLong’s test confirmed a significantly higher AUC for RENALNet than Radiomics_Combined in the training cohort (p = 0.017), with non-significant differences in the testing and external validation cohorts (all p > 0.05) (Fig. 2 and Tables 2 and 3).
Human–AI collaboration improves diagnostic accuracy
A multi-reader study involving junior, attending, and senior radiologists demonstrated the incremental diagnostic value of RENALNet when combined with radiologist assessments. Logistic regression models combining each radiologist’s binary classification with RENALNet scores consistently enhanced diagnostic accuracy. For the junior radiologist, AUCs improved from 0.619 to 0.761, 0.556 to 0.770, 0.660 to 0.734, and 0.611 to 0.716 across Validation Cohorts I–IV, respectively. Similar improvements were observed for attending and senior radiologists. In Validation Cohort I, for example, the senior radiologist’s AUC increased from 0.760 to 0.855 (p = 0.024), with sensitivity rising from 0.571 to 0.714. DeLong’s test confirmed statistical significance (p < 0.05) in most comparisons. Although specificity slightly decreased in some instances, overall accuracy remained stable or improved, underscoring the synergistic benefit of combining human expertise with deep learning outputs (Table 4).
Model interpretability and visualization
For the Radiomics_Combined model, SHAP (Shapley Additive exPlanations)20 analysis identified 40 contributing features, including 30 peritumoral and 10 intratumoral. Six of the top ten predictors were peritumoral, with peritumor_shape_Maximum3DDiameter ranked the highest. Additional key peritumoral features included peritumor_glcm_Correlation, peritumor_ngtdm_Coarseness, peritumor_ngtdm_Busyness, peritumor_firstorder_Minimum, and peritumor_glcm_Idn, suggesting that complex geometric and textural patterns in adjacent tissue provide strong predictive signals. Among intratumoral features, tumor_shape_MinorAxisLength and tumor_glrlm_RunLengthNonUniformity contributed complementary structural information. SHAP dependence analysis revealed nonlinear interactions among top features, where intratumoral texture contributions varied as a function of peritumoral geometry, highlighting that the model captures not only additive effects but also cross-regional interactions indicative of subtle morphological changes associated with local invasion (Fig. 3a, b).
A SHapley Additive exPlanations (SHAP) summary plot showing the global contribution of radiomic features in the Radiomics_Combined model. B SHAP dependence plots for the top three contributing features, illustrating their marginal effects on the predicted risk of pT3a upstaging. C Representative gradient-weighted class activation mapping (Grad-CAM) heatmaps generated by RENALNet for patients with various invasion patterns.
For RENALNet, Grad-CAM21 visualization frequently emphasized anatomically relevant regions at the tumor–fat interface (perirenal or renal sinus fat) and along the renal vein, which are key sites where perirenal/sinus invasion and venous tumor thrombus are evaluated in routine practice. In cases with multifocal or extensive invasion, several of these regions were qualitatively activated on the heatmaps, suggesting that the model captured complex local invasion patterns that were compatible with the pathological reports (Fig. 3c).
Biological and prognostic validation
Patients stratified by RENALNet risk (cutoff = 0.5) showed significantly higher Ki-67 indices in the high-risk group across the training, testing, and two external validation cohorts (all p < 0.05). RENALNet risk positively correlated with Ki-67 in all cohorts (Fig. 4a, b), suggesting that the model-derived risk reflects tumor proliferative activity.
A Comparison of Ki-67 indices between high-risk and low-risk groups (cutoff = 0.5) in the training, testing, and two external validation cohorts (Validation II and III), showing significantly elevated Ki-67 levels in the high-risk group across all cohorts. B Spearman correlation analysis showing significant positive associations between RENALNet risk scores and Ki-67 indices across all cohorts, supporting the link between imaging-derived risk and tumor proliferative activity. C Kaplan–Meier analysis of 5-year progression-free survival (PFS) stratified by RENALNet-derived risk groups using an optimal cutoff determined by maximally selected rank statistics, revealing significantly poorer PFS in the high-risk group (p < 0.001).
Progression-free survival (PFS) was assessed in 246 patients with follow-up data. The optimal RENALNet cutoff, identified through maximally selected rank statistics, stratified patients into groups with significantly different 5-year PFS (p < 0.001, Fig. 4c), further supporting its prognostic value for long-term outcomes.
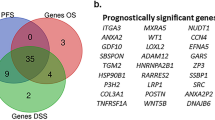
To investigate the molecular basis of RENALNet risk, differential expression analysis was performed with respect to continuous risk scores. Genes showing significant correlations were identified as differentially expressed genes (DEGs), with MUC17, FOXI1, IRS4, and KLK1 exhibiting the strongest positive associations (Fig. 5a). These genes are functionally linked to EMT and PI3K–Akt signaling, emphasizing their potential roles in promoting tumor aggressiveness and facilitating advanced staging in renal cell carcinoma. KEGG analysis further confirmed that these DEGs are enriched in pathways such as PI3K–Akt signaling and cytokine–cytokine receptor interaction, both implicated in EMT and tumor–microenvironment remodeling, offering mechanistic insight into ccRCC progression (Fig. 5b). Complementary GSEA confirmed positive enrichment of EMT (NES = 1.703, adjust p = 0.002), IL6–JAK–STAT3 signaling (NES = 1.589, adjust p = 0.046), and allograft rejection (NES = 1.676, adjust p = 0.003) with increasing RENALNet risk scores (Fig. 5c–e). Collectively, these findings indicate that higher model-predicted risk is associated with the activation of invasive and pro-inflammatory programs, consistent with the biology of advanced-stage RCC.
A Volcano plot showing differentially expressed genes (DEGs) significantly correlated with RENALNet risk, with representative top-ranked genes labeled. B Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis of DEGs, highlighting pathways such as PI3K–Akt signaling and cytokine–cytokine receptor interaction. C, D Gene set enrichment analysis (GSEA) results. The bubble plot C illustrates the significance, enrichment direction, and gene set size of hallmark pathways, while the heatmap D shows normalized enrichment scores (NES) for representative pathways. GSEA enrichment plots showing positive associations of epithelial–mesenchymal transition (EMT) (E) and IL6–JAK–STAT3 signaling F with higher RENALNet risk.
Discussion
Accurately identifying pT3a upstaging preoperatively in ccRCC remains challenging, as conventional imaging often fails to detect microscopic extrarenal invasion, leading to understaging and suboptimal surgical decisions. In this study, RENALNet, a 3D deep learning model, was developed and externally validated to noninvasively predict pT3a upstaging in cT1b–cT2a ccRCC. RENALNet consistently outperformed radiomics-based models across internal and external cohorts, and further enhanced radiologist performance, particularly among less experienced readers. Importantly, RENALNet-derived risk was correlated with Ki-67 proliferation indices and 5-year PFS, while transcriptomic analysis revealed enrichment of invasive and pro-inflammatory programs such as EMT and IL6–JAK–STAT3 signaling. These findings suggest that RENALNet captures imaging signatures linked to both clinical aggressiveness and molecular pathways, bridging radiologic risk assessment with tumor biology and patient outcomes.
To better understand the modeling strategies, RENALNet was compared with handcrafted radiomics models. Among the radiomics models, combining intratumoral and peritumoral features yielded higher predictive performance than using either region alone, in line with previous research22 emphasizing the diagnostic value of the peritumoral microenvironment in identifying early extrarenal invasion. SHAP analysis confirmed that the top-ranking features were predominantly from the peritumoral region, and SHAP dependence plots revealed nonlinear interactions between tumor and peritumor features. These results highlight that handcrafted models can capture complementary information from adjacent tissue compartments but remain limited by predefined feature sets.
In contrast, RENALNet demonstrated clear advantages over handcrafted radiomics models. By learning complex spatial representations directly from volumetric CT data, RENALNet achieved higher AUCs and significantly improved sensitivity across diverse datasets. Furthermore, RENALNet enhanced human interpretation: when integrated with radiologists’ assessments, diagnostic AUCs improved consistently across all experience levels, with the substantial improvement observed in junior readers (AUC increased from 0.619 to 0.761; p = 0.010). These results underscore RENALNet’s dual strength in providing more accurate automated predictions while simultaneously elevating radiologist performance. Grad-CAM visualizations further supported interpretability by consistently focusing on anatomically relevant sites of various invasions, providing intuitive evidence for preoperative decision-making.
Beyond diagnostic utility, RENALNet outputs demonstrated meaningful associations with tumor biology and clinical outcomes. RENALNet risk was significantly positively correlated with Ki-67 proliferation indices, with high-risk patients (based on a 0.5 probability cutoff) consistently showing higher Ki-67 levels across multiple cohorts. Additionally, RENALNet stratification effectively distinguished patients with significantly different 5-year PFS (75.0% in the high-risk group vs. 94.1% in the low-risk group, p < 0.001). At the transcriptomic level, genes associated with high risk, including MUC17, FOXI1, IRS4, and KLK1, are functionally involved in EMT and PI3K–Akt signaling, both of which are known to drive tumor aggressiveness23. KEGG and GSEA analyses further revealed enrichment of PI3K–Akt signaling, cytokine–cytokine receptor interaction, EMT, and IL6–JAK–STAT3 signaling, indicating activation of pro-inflammatory and invasive programs24. Although our radiogenomic analyses were performed using the RENALNet risk score, the regions highlighted by Grad-CAM provide a biologically plausible link between the CT patterns and the enriched pathways. Grad-CAM maps tended to focus on key anatomical sites of invasion, including capsular disruption, early perirenal fat or sinus invasion, and renal vein involvement, which is compatible with an EMT- and cytokine-driven pro-invasive microenvironment. These qualitative correspondences suggest that the image patterns exploited by RENALNet may partially reflect underlying invasive programs. Although these radiogenomic relationships are associative and derived from bulk RNA sequencing without spatial transcriptomic alignment, they collectively indicate that RENALNet identifies radiogenomic correlates of tumor aggressiveness and reinforce its biological interpretability and prognostic value.
The clinical implications of the RENALNet risk score also warrant consideration, especially for choosing between RN and NSS in cT1b–cT2a tumors. In our cohorts, a higher RENALNet-predicted risk of occult pT3a was associated with higher Ki-67 indices and worse 5-year PFS, suggesting that the score can serve as a preoperative biomarker of high-risk tumor phenotype. In practice, RENALNet could be used as a triage tool: patients with anatomically suitable tumors and low predicted risk may be prioritized for NSS, whereas those with high predicted risk may warrant a lower threshold for considering RN when anatomy is complex, and, if NSS is still chosen because of impaired baseline renal function, more intensive postoperative oncologic surveillance. The probability thresholds used in this study were derived from retrospective data and should be validated prospectively before being used to guide treatment decisions.
Several key aspects underscore the strength and translational potential of our work. First, RENALNet was trained and validated on a large, multicenter cohort with four external datasets, demonstrating its robust generalizability across diverse scanners, protocols, and patient populations. Second, by directly comparing radiomics and deep learning approaches, this study uncovered the diagnostic value of peritumoral handcrafted features while also establishing the advantages of end-to-end representation learning. Third, interpretability was prioritized through SHAP and Grad-CAM analyses, ensuring both transparency and clinical trust. Fourth, and crucially, RENALNet risk was linked not only to pathological upstaging but also to transcriptomic pathways and survival outcomes, enabling multi-scale phenotyping that connects imaging signatures with molecular biology and clinical prognosis. This integrative approach positions RENALNet as more than just a diagnostic tool; it bridges between radiology, genomics, and patient outcomes in precision oncology.
However, this study has several limitations. First, as the datasets were retrospectively collected, prospective validation in real-world clinical workflows is still needed. Second, CT scans were acquired on different scanners using heterogeneous acquisition parameters across centers; although resampled to a uniform voxel spacing and processed with a standardized pipeline, such variability may still influence model performance and limit generalizability. Third, the model was developed using only nephrographic-phase CT; incorporating multiphase or multimodal data may further enhance its performance. Fourth, this work focused solely on ccRCC, and extending the approach to other histologies or endpoints, such as treatment response, will be necessary to broaden its clinical applicability. Finally, mechanistic interpretations linking attention maps and transcriptomic pathways remain constrained by the lack of spatially resolved molecular data.
In conclusion, RENALNet addresses a critical unmet need in cT1b–T2a ccRCC by providing an accurate and interpretable preoperative prediction of occult pT3a upstaging. The model outperformed radiomics-based approaches, enhanced radiologist performance, and demonstrated biological relevance through correlations with Ki-67 indices, survival outcomes, and transcriptomic pathways. These findings establish RENALNet as a biologically interpretable framework for patient risk stratification, with the potential to assist surgical decision-making and advance the integration of radiogenomic deep learning into precision oncology.
Methods
Study design and patient cohorts
This retrospective, multicenter study was conducted in accordance with the Declaration of Helsinki and the TRIPOD (Transparent Reporting of a Multivariable Prediction Model for Individual Prognosis or Diagnosis) guideline. The study protocol was approved by the Medical Ethics Committees of Tongji Hospital, Tongji Medical College, Huazhong University of Science and Technology (TJ-IRB202502100), Xiangyang Central Hospital, Affiliated Hospital of Hubei University of Arts and Science (2025-097), Shanxi Bethune Hospital, Shanxi Academy of Medical Sciences (YXLL-2025-144), the Affiliated Cancer Hospital of Zhengzhou University & Henan Cancer Hospital (2025-237-001), and the First Affiliated Hospital of Nanjing Medical University (2022-SR-408). The requirement for informed consent was waived by the ethics committees due to the retrospective nature of the study. Data were collected from five hospitals and the publicly available KiTS23 dataset. Eligible patients were ≥ 18 years of age, had pathologically confirmed ccRCC, and underwent preoperative contrast-enhanced abdominal CT within one month prior to surgery. All included tumors were surgically resected, and patients underwent either RN or NSS. For institutional cohorts, patients were excluded if their tumor size was < 3 cm or > 10 cm, if there was any radiological or pathological evidence of regional lymph node or distant metastasis (N1 or M1), if the preoperative CT reports or clinical documentation described definite perirenal or renal sinus fat invasion, renal vein or vena cava thrombus, or adjacent organ invasion, if CT image quality was poor, or if they had received any preoperative therapy. For the public KiTS23 cohort, clinical TNM stage and radiology reports were not available, and the recorded pathological T stage did not provide T3 subclassification. We therefore applied the same size criteria (3–10 cm), excluded cases with T4 or N1/M1 disease, and used the remaining non-metastatic, pathologically confirmed ccRCC cases as an external validation cohort.
CT image acquisition and processing
All patients underwent contrast-enhanced abdominal CT scans before surgery. Corticomedullary and nephrographic phases were acquired at 25–28 s and 65–70 s, respectively. Nephrographic-phase images were used for all subsequent analyses. All images were converted to NIfTI format and resampled to a uniform voxel spacing of 0.5 × 0.5 × 1 mm using trilinear interpolation to ensure spatial consistency. Image orientation was standardized across datasets to eliminate variations due to scanning direction. Kidneys, tumors, and perirenal adipose tissue were first segmented using an nnU-Net framework pretrained on KiTS2325 and then applied to local data. Segmentation outputs were manually reviewed and refined by two radiologists (G.M.M. and L.S.C., with 3 and 6 years of experience in abdominal CT, respectively) using ITK-SNAP software. These final masks served as the basis for defining the intratumoral and peritumoral regions used in subsequent radiomic and deep learning analyses. In cases with multiple tumors in a single kidney, the largest lesion was selected based on morphological criteria. The overall image processing pipeline is illustrated in Fig. S1.
Clinicopathologic characteristics
Pathological grade and Ki-67 indices were extracted from the original pathology reports at each center. In most cases, pathological grade followed the WHO/ISUP system; a small subset of earlier cases was reported using the Fuhrman system. In all centers, Ki-67 was assessed and reported by local pathologists as the percentage of tumor nuclei showing positive staining according to institutional practice.
PFS was ascertained from electronic medical records and defined as the time from surgery to the first documented postoperative recurrence or progression, including local recurrence, regional lymph node metastasis, or distant metastasis. Patients without documented progression were censored at the date of their last clinical or imaging follow-up.
Radiomic analysis
Radiomic features were extracted from intratumoral and peritumoral regions based on the kidney, tumor, and perirenal adipose masks described above. The intratumoral region was defined as the entire tumor volume on nephrographic-phase CT. The peritumoral region was defined as a three-dimensional 5-mm morphological dilation of the tumor mask, selected after comparative experiments with 3-, 5-, and 7-mm peritumoral rings (Supplementary Fig. S2 and Table S1). A total of 107 original features were extracted using PyRadiomics26, including shape, first-order statistics, and texture features, and normalized using z-score transformation. Feature selection was performed using a mutual information–based SelectKBest approach (k = 20–50). Four machine learning classifiers—logistic regression, support vector machine, decision tree, and random forest—were trained and optimized via grid search and five-fold cross-validation27. SHAP analysis was used to interpret the contributions of individual features to the predictions.
Deep learning classifier
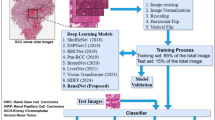
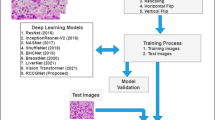
A 3D convolutional neural network, RENALNet, was developed to capture both intratumoral and peritumoral spatial features associated with pT3a upstaging (Fig. 6). RENALNet consists of three stages, each comprising a convolutional block, a residual block, and dual attention modules (channel and spatial). The convolutional block incorporates 3D convolution, batch normalization, and ReLU activation. Residual connections ensure the preservation of feature flow and support stability in deep learning models28. To enhance feature representation, channel and spatial attention mechanisms were integrated29. Channel attention reweights features using global average pooling and multilayer perceptrons, while spatial attention generates attention maps through max and average pooling to highlight key spatial regions. A mask-guided attention module30 incorporates anatomical priors, further focusing the model on tumor-relevant areas. This design refines the model’s focus on clinically significant regions, improving its ability to detect subtle signs of local invasion. The final layer includes global average pooling and a sigmoid-activated fully connected layer to output the probability of upstaging.
RENALNet architecture for predicting occult pT3a upstaging in clinical localized clear cell renal cell carcinoma.
Compared with conventional 3D ResNet/DenseNet, RENALNet replaces pure residual or dense feature propagation with a hierarchical attention design that combines convolutional block attention modules (CBAM) and mask-guided attention to focus on peritumoral and tumor-adjacent regions relevant to pT3a upstaging. A schematic comparison of the Dense-Block, Res-Block, CBAM-Block, and Basic-Block is provided in Supplementary Fig. S3.
Input volumes were preprocessed to a fixed size of 64 slices of 224 × 224 pixels, windowed to 40/400 HU, and normalized. Data augmentation, including horizontal flips (p = 0.5) and random rotations (±10°, p = 0.3), was performed to increase variability while maintaining anatomical consistency. A mini-batch size of 16 with balanced sampling ensured each batch included sufficient pT3a cases. Two sampling strategies were employed: a non-repeating balanced sampler ensured each batch contained at least one positive case, while a ratio-balanced batch sampler (positive ratio = 0.2) allowed repeated inclusion of minority samples.
RENALNet was trained using the Adam optimizer (initial learning rate = 1 × 10−4, weight decay = 1 × 10−5), a cosine annealing schedule (min LR = 1 × 10−6, 100 epochs), and Focal Loss31 (α = 0.8, γ = 2.0). Validation AUC was monitored at each epoch to guide training and prevent overfitting. Model development was conducted in Python 3.10.0 with PyTorch 2.1.0 and CUDA 12.1 for GPU acceleration.
The reader study and RENALNet-assisted diagnosis
To assess clinical utility, a reader study was conducted across four external validation cohorts. Three radiologists with varying experience levels (junior, 3 years; attending, 6 years; senior, 22 years) independently reviewed nephrographic-phase CT images to assess whether occult pT3a upstaging was present and recorded a binary judgement, blinded to the RENALNet output. Logistic regression models then integrated each radiologist’s binary assessment with RENALNet’s probability output; the performance of these combined models is reported as the “RENALNet-assisted” condition. Diagnostic performance was compared between radiologists alone and the RENALNet-assisted condition, with statistical significance evaluated using the DeLong test.
Model interpretation
Grad-CAM was applied to the final convolutional layer of RENALNet to generate 3D attention heatmaps. Forward and backward gradients were combined to compute voxel-wise importance scores, which were subsequently overlaid onto the CT slices. As RENALNet processes volumetric inputs (64 × 224 × 224 voxels), the resulting heatmaps retained spatial continuity in three dimensions. These attention maps were displayed in a multi-slice format, allowing radiologists to visually interpret the regions that contributed most to the model’s predictions across adjacent slices. Two abdominal radiologists qualitatively reviewed the Grad-CAM maps together with the corresponding CT images and pathology reports to assess whether the highlighted regions were located near anatomically relevant invasive sites.
Model evaluation
A three-step postprocessing pipeline was employed during external validation. First, Bayesian calibration of RENALNet’s raw probabilities was performed using dataset-specific priors to generate calibrated confidence intervals. Next, k-nearest neighbor smoothing (k = 5, smoothing factor = 0.3) was applied in feature space to reduce prediction noise. Finally, bootstrap-based ensemble inference was utilized by training five RENALNet instances on 80% of bootstrap resamples and aggregating their outputs via weighted averaging. This framework addresses class imbalance, provides uncertainty estimates, and supports efficient batch processing of large-scale data.
Transcriptomic analysis
RNA-seq data from TCGA-KIRC (https://portal.gdc.cancer.gov/) were analyzed using DESeq2, with RENALNet risk modeled as a continuous covariate. Genes with adjusted p < 0.05 (Benjamini–Hochberg correction) were considered significantly associated. DEGs were visualized using volcano plots (ggplot2), classifying genes as upregulated (log2 fold change > 1, adjusted p < 0.05), downregulated (log2 fold change < −1, adjusted p < 0.05), or not significant. The top 10 up- and downregulated genes were annotated (ggrepel).
Pathway enrichment of significant genes was performed using clusterProfiler against KEGG, with all tested genes serving as the background. Gene set enrichment analysis (GSEA) was conducted with fgsea on genes ranked by log2 fold change, using the MSigDB Hallmark and KEGG collections (minSize = 15, maxSize = 500, 1000 permutations). Enrichment results were reported as normalized enrichment scores (NES), with an adjusted p < 0.05 considered significant. Representative pathways, including EMT and IL6–JAK–STAT3 signaling, were visualized using the enrichplot package.
Statistical analyses
Continuous variables were assessed for normality using the Shapiro–Wilk test and compared by Student’s t-test or Mann–Whitney U test as appropriate. Categorical variables were compared via the chi-square test. Model discrimination was evaluated using the area under the ROC curve (AUC), and ROC curves were compared with the DeLong test. PFS was estimated with Kaplan–Meier curves and compared using the log-rank test. The optimal RENALNet cutoff for PFS stratification was determined using maximally selected rank statistics. For transcriptomic analyses, differential expression was assessed with DESeq2, and p values were adjusted by Benjamini–Hochberg correction. KEGG and GSEA analyses were performed with clusterProfiler and fgsea, respectively, with significance set at adjusted p < 0.05. All analyses were conducted in Python 3.10.0 and R 4.4.1, with ROC and AUC computations using pROC and scikit-learn. A two-sided p < 0.05 was considered statistically significant.
Data availability
Public datasets used in this study are available as follows: the KiTS2023 dataset from the 2023 Kidney and Kidney Tumor Segmentation Challenge (https://kits-challenge.org/kits23/); the TCGA-KIRC imaging collection from The Cancer Imaging Archive (https://www.cancerimagingarchive.net/collection/tcga-kirc/), and matched RNAseq data from the NCI Genomic Data Commons (https://portal.gdc.cancer.gov/, project “TCGA-KIRC”). Other data are available from the corresponding author upon reasonable request.
Code availability
The codes used for model development and analysis are publicly available at (https://github.com/BobbyWang16/RENALNet).
References
Young, M. et al. Renal cell carcinoma. Lancet 404, 476–491 (2024).
Rose, T. L. & Kim, W. Y. Renal cell carcinoma: a review. JAMA 332, 1001–1010 (2024).
Powles, T. et al. Renal cell carcinoma: ESMO clinical practice guideline for diagnosis, treatment and follow-up. Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 35, 692–706 (2024).
Capitanio, U. et al. A renewal of the TNM staging system for patients with renal cancer to comply with current decision-making: proposal from the European Association of Urology guidelines panel. Eur. Urol. 83, 3–5 (2023).
Gülşen, M., Köse, E. & Özden, E. Laparoscopic partial nephrectomy tips and tricks. J. Urol. Surg. 12, 49–51 (2025).
Laguna, M. P. Re: incidence of T3a up-staging and survival after partial nephrectomy: size-stratified rates and implications for prognosis. J. Urol. 200, 695 (2018).
Ramaswamy, K. et al. Significance of pathologic T3a upstaging in clinical T1 renal masses undergoing nephrectomy. Clin. Genitourin. cancer 13, 344–349 (2015).
Veccia, A. et al. Upstaging to pT3a in patients undergoing partial or radical nephrectomy for cT1 renal tumors: a systematic review and meta-analysis of outcomes and predictive factors. Eur. Urol. Focus 7, 574–581 (2021).
Abou Elkassem, A. M. et al. Role of imaging in renal cell carcinoma: a multidisciplinary perspective. Radiogr. Rev. Publ. Radiol. Soc. North Am. Inc. 41, 1387–1407 (2021).
Elkassem, A. A., Allen, B. C., Sharbidre, K. G., Rais-Bahrami, S., Smith, A. D. Update on the Role of Imaging in Clinical Staging and Restaging of Renal Cell Carcinoma Based on the AJCC 8th Edition, From the AJR Special Series on Cancer Staging, AJR. Am. J Roentgenol. 217, 541–555 (2021).
Wen-Zhi, G. et al. Prediction of pathological staging and grading of renal clear cell carcinoma based on deep learning algorithms. J. Int. Med. Res. 50, 3000605221135163 (2022).
Fateh, S. M. et al. Renal cell carcinoma T staging: diagnostic accuracy of preoperative contrast-enhanced computed tomography. Mol. Clin. Oncol. 18, 11 (2023).
Ma, J. et al. Diagnostic performance of CT for extrarenal fat invasion in renal cell carcinoma: a meta-analysis and systematic review. Insights Into Imaging 16, 19 (2025).
Xia, T. Y. et al. Predicting microvascular invasion in hepatocellular carcinoma using CT-based Radiomics Model. Radiology 307, e222729 (2023).
Guiot, J. et al. A review in radiomics: making personalized medicine a reality via routine imaging. Med. Res. Rev. 42, 426–440 (2022).
Chen, X. et al. Recent advances and clinical applications of deep learning in medical image analysis. Med. Image Anal. 79, 102444 (2022).
Uhlig, A. et al. Radiomics and machine learning for renal tumor subtype assessment using multiphase computed tomography in a multicenter setting. Eur. Radiol. 34, 6254–6263 (2024).
Li, S. et al. Incremental value of automatically segmented perirenal adipose tissue for pathological grading of clear cell renal cell carcinoma: a multicenter cohort study. Int. J. Surg. 110, 4221–4230 (2024).
Nie, P. et al. A CT-based deep learning radiomics nomogram outperforms the existing prognostic models for outcome prediction in clear cell renal cell carcinoma: a multicenter study. Eur. Radiol. 33, 8858–8868 (2023).
Nohara, Y., Matsumoto, K., Soejima, H., Nakashima, N. Explanation of machine learning models using improved Shapley additive explanation. In Proc. 10th ACM International Conference on Bioinformatics, Computational Biology and Health Informatics (Association for Computing Machinery, Niagara Falls, 2019).
Selvaraju, R. R. et al. Grad-CAM: visual explanations from deep networks via gradient-based localization. In Proc. 2017 IEEE International Conference on Computer Vision (ICCV), 618–626 (2017).
Huang, D. et al. Radiomics model based on intratumoral and peritumoral features for predicting major pathological response in non-small cell lung cancer receiving neoadjuvant immunochemotherapy. Front. Oncol. 14, 1348678 (2024).
Wu, Y. et al. Epigenetic and transcriptomic characterization reveals progression markers and essential pathways in clear cell renal cell carcinoma. Nat. Commun. 14, 1681 (2023).
Zegeye, M. M. et al. Activation of the JAK/STAT3 and PI3K/AKT pathways are crucial for IL-6 trans-signaling-mediated pro-inflammatory response in human vascular endothelial cells. Cell Commun. Signal. 16, 55 (2018).
Isensee, F., Jaeger, P. F., Kohl, S. A. A., Petersen, J. & Maier-Hein, K. H. nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat. Methods 18, 203–211 (2021).
van Griethuysen, J. J. M. et al. Computational radiomics system to decode the radiographic phenotype. Cancer Res. 77, e104–e107 (2017).
K, L., S, T., A, V., V, P. Feature selection for high dimensional data using supervised machine learning techniques. In Proc. IEEE International Conference on Big Data (Big Data), 3891–3894 (IEEE, 2022).
Uemura, T., Näppi, J. J., Hironaka, T., Kim, H., Yoshida, H. Comparative performance of 3D-DenseNet, 3D-ResNet, and 3D-VGG models in polyp detection for CT colonography. In Proc. Medical Imaging 2020: Computer-aided Diagnosis 736–741 (SPIE, 2020).
Liu, T. et al. Spatial channel attention for deep convolutional neural networks. Mathematics 10, 1750 (2022).
Li, Z. et al. A novel deep learning framework based mask-guided attention mechanism for distant metastasis prediction of lung cancer. In Proc. IEEE Transactions on Emerging Topics in Computational Intelligence 330–341 (IEEE, 2022)
Tran, G. S., Nghiem, T. P., Nguyen, V. T., Luong, C. M., Burie, J.-C. Improving accuracy of lung nodule classification using deep learning with focal loss. J. Healthc. Eng. 2019, 5156416 (2019).
Acknowledgements
The authors have no additional acknowledgements to declare. This study was supported by grants from the National Natural Science Foundation of China (Nos. 82502331, 82371942, and 62131009), Local Science and Technology Development Fund Project Guided by Central Government of China (No. YDZJSX 2025D074), and Xiangyang Science and Technology Program in Healthcare Field (No. 2022YL19A).
Author information
Authors and Affiliations
Contributions
S.C.L., Y.D.Z., J.R.Q., and Z.L. conceived and designed the study. S.C.L., K.W.H., and M.M.G. collected and curated the data. S.C.L., M.M.G., and Z.L. calibrated and reviewed the volumes of interest. C.H.W. and Z.L.Z. developed and validated the radiomics and deep-learning models. S.C.L., M.M.G, and W.N.Q. managed patient follow-up and data management. S.C.L. and C.H.W. prepared the figures and wrote the original draft of the manuscript. F.L., S.W., Y.D.Z., J.R.Q., and Z.L. reviewed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, S., Wang, C., Li, F. et al. Noninvasive prediction of occult pT3a upstaging in localized ccRCC with radiogenomic insights and prognostic relevance. npj Precis. Onc. 10, 104 (2026). https://doi.org/10.1038/s41698-026-01315-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-026-01315-2