Abstract
Homologous recombination deficiency (HRD) can lead to genomic instability, increased cancer susceptibility, and enhanced sensitivity to DNA-targeting therapies. Although radiomics has been used for various medical applications, its application in animal studies remains largely unexplored, primarily due to the typically limited availability of preclinical data. In this study, we applied a state-of-the-art foundation model (FM) on preclinical computed tomography (CT) images in mice, aiming to: (i) distinguish HRD status within isogenic xenografts, and (ii) predict differential therapeutic responses of CP-506, a novel hypoxia-activated DNA-crosslinking agent. The dataset comprises micro-CT scans of 307 mice with balanced HRD status, collected both before and after CP-506 or control treatment. The FM demonstrated robust HRD classification performance, achieving an AUC of 0.88 on the test set, which significantly outperformed the handcrafted radiomics and supervised deep learning (sDL). The highest AUC (0.93) was achieved in the consensus subgroup (71%) between sDL and FM. Additionally, HRD-related features predicted DNA damage and growth delay following the treatment. Interpretability analysis indicated the important role of texture heterogeneity in HRD classification. Therefore, these results suggest that FM successfully overcomes the data scarcity in animal studies and enables HRD classification and treatment response prediction from preclinical CT imaging.
Similar content being viewed by others
Introduction
Homologous recombination (HR) is a crucial DNA repair pathway for high-fidelity repair of double-strand DNA breaks1. A deficiency in this pathway, termed homologous recombination deficiency (HRD), results in the accumulation of unrepaired DNA damage and can lead to genomic instability and cancer development. HRD is often associated with, but not limited to, germline mutations in key genes and is linked to the pathogenesis of multiple cancers2,3,4,5. It has been extensively investigated in breast and high-grade serous ovarian cancers, where BRCA mutations are clinically actionable2,6. Germline and somatic alterations in HR repair genes, including BRCA and Fanconi anemia pathway (e.g., FANCA, FANCD2), have also been identified in prostate and colorectal cancers, as well as other genitourinary and gastrointestinal tumors7,8,9. HRD tumors present with higher spatial and temporal heterogeneity, complicating tumor diagnosis10,11. Due to the inability of HRD tumors to repair DNA damage, DNA-targeted therapies, such as PARP inhibitors and chemotherapies, lead to increased cell death and cause better treatment outcomes12,13,14,15,16. Therefore, the detection of HRD can guide cancer treatment decisions and improve responses to targeted therapies that induce DNA damage.
Current HRD detection involves tests to identify mutations in HR repair genes, the presence of genomic scars and signatures, as well as functional assays15,17,18,19,20. However, these methods, which typically rely on invasive biopsies, present challenges in terms of high cost, long processing times, and their inability to dynamically map HRD status in space and over time. Alternatively, another approach is to examine how the genomic deficiencies translate to the radiological tumor phenotype21. A deep learning approach was used to analyze pathology tissue slides, therefore, to identify associated morphological patterns and enable the detection of HRD22. While this approach is innovative and promising, it remains invasive and time-consuming.
Radiomics, a non-invasive analytical technique that extracts quantitative features from medical imaging modalities, has been investigated for its potential to capture genetic changes and mutation signatures in various cancers23,24. For example, emerging evidence shows a correlation between radiomic features observed in magnetic resonance imaging (MRI) and TP53 mutations in lower-grade gliomas25. Other computed tomography (CT) image features have also been shown to effectively predict BRCA mutations in epithelial ovarian cancer26. Several approaches have applied radiomics in animals, but this field remains largely unexplored, primarily due to the typically limited availability of preclinical data27. To overcome such challenges, pre-trained models have been developed to leverage prior knowledge and enhance radiomic analysis. Recent studies have demonstrated that foundation models (FMs) based on self-supervised deep learning can advance cancer imaging biomarker discovery and show a strong association with the biological basis of gene expression28,29. Compared to traditional supervised learning, FMs require less data for fine-tuning in downstream tasks, and exhibit greater stability in terms of test-retest scenario and resilience to perturbations28. On the basis of previous radiomic studies in animals, we extend this line of work by applying FMs on animal data for identifying HRD status30,31,32,33,34,35.
As a clinically indispensable modality, CT imaging has been employed to predict responses to cancer therapies by capturing tumor phenotype characteristics, such as size and morphology36,37. However, to the best of our knowledge, no studies have specifically focused on identifying HRD-related features from CT imaging to predict therapeutic outcomes26,38,39. Given the established role of HRD in influencing treatment responses and the rich information contained in CT images, we hypothesized that CT-based FMs could identify HRD tumors on a preclinical dataset and become a biomarker of treatment efficacy29.
Therefore, the objectives of this study are twofold: (i) to detect HRD based on CT images using an FM across various xenografts and mutation types and compare the results to handcrafted radiomics (HCR) and supervised deep learning (sDL); and (ii) to assess HRD-related CT imaging features for predicting treatment outcomes. To achieve these goals, we have set up mouse models with balanced HRD and HR proficient (HRP) xenografts. CT scans were performed both before and after CP-506 and control treatment, and predictive models were trained to detect HRD and evaluate treatment efficacy on DNA damage and growth delay after CP-506 therapy.
Results
Data structure and characteristics
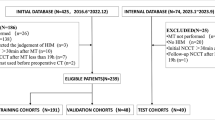
614 CT scans of isogenic xenografts from 307 mice were collected before and after CP-506 (i.e., hypoxia-activated prodrug with an alkylating warhead) or control treatment (Figs. 1–2, Supplementary Table S1). After excluding 71 scans due to impaired or missing data files (n = 53), damaged files (n = 4), partial tumor coverage (n = 13), or movement artefacts (n = 1), 543 scans were used for further analysis. These scans were classified into two categories: no-treatment scans (n = 414, i.e., pre- and post-treatment scans from the control group and pre-treatment scans from the treatment group), and treatment scans (n = 129, i.e., post-treatment scans from the treatment group). All no-treatment scans were split into a training set (n = 287) and a test set (n = 127). Characteristics (e.g., mean region of interest [ROI] volume, mean ROI intensity, cell line, and mutations) showed no significant differences between the training and test datasets (all p > 0.05, Table 1).
307 mice were divided into four groups based on isogenic xenografts (HRP vs. HRD) and treatment (CP-506 vs. vehicle) stratification. CT scans were collected before and after treatment. No-treatment scans (pre- and post-scans from the control group and pre-treatment scans from the treatment group) were split into a training set (70%) and a test set (30%). HRP homologous recombination proficiency, HRD homologous recombination deficiency, HRP homologous recombination proficiency, CT computed tomography.
a Study design for animal experiment: mice bearing isogenic xenografts (HRD/HRP) were subjected to CP-506 (i.e., hypoxia-activated DNA-crosslinking agent) and control (vehicle) treatments. In the main experiment, CT scans were acquired before and after treatment, and DNA damage intensity was quantified in tumor tissues using γ-H2AX immunohistochemical staining 48 h post-treatment. In a parallel experiment, tumor volumes were measured regularly until tumors quadrupled in volume (event-free survival, EFS)51. b Radiomic pipeline for HRD classification and treatment response prediction: A pre-trained foundation model (FM) was employed to derive high-dimensional features from the CT scans. A shallow classifier was utilized to determine HRD tumor status. Subsequently, HRD-related features extracted from the final layer of the classifier were used to predict treatment responses (i.e., DNA damage and EFS).
HRD classification performance across multiple models
For each isogenic xenograft, HRD status was separately predicted using three models: an HCR model, an sDL model, and an FM. Predicted probabilities were calibrated using Platt scaling (Supplementary Fig. S1 and Table S2). On the training set, the HCR demonstrated an area under the curve (AUC) ranging from 0.77 to 0.79, and the sDL model demonstrated an AUC between 0.71 and 0.78 across different CT energies. An sDL initialized with pre-trained weights from MedicalNet showed no significant improvement compared to sDL (p > 0.05, Supplementary Table S3–S4). The FM consistently outperformed HCR and sDL, achieving an AUC between 0.89 and 0.90 and an accuracy of ca. 0.85 (p < 0.05, Fig. 3a, Table 2). Additionally, the FM showed significantly higher sensitivity and specificity compared to the HCR and sDL models (p < 0.05, Table 2). On the test set, the FM maintained its superior performance, with an AUC ranging from 0.87 to 0.88, closely mirroring its training performance (Fig. 3b, Table 3). Notably, FM was least influenced by CT energy levels (p > 0.05) and consistently outperformed the HCR and sDL models on both the training and test datasets across all CT energy levels (i.e., 40 kVp CT, 80 kVp CT, and combined CT) (Fig. 3a, b). Details of the HCR model, including feature selection, feature importance analysis, and ablation experiments comparing multiple machine learning classifiers, are provided in Supplementary Fig. S2 and Tables S5–S8.
a ROC curves on the training set comparing the performance of the handcrafted radiomics (HCR), supervised deep learning (sDL), and foundation model (FM) for HRD classification across 40 kVp CT, 80 kVp CT, and combined CT datasets. b Same as (a), but on the test set. c AUC values as a function of training set size for each model. Lines represent mean AUCs; shaded areas (a, b) and error bars (c) indicate 95% confidence intervals (CIs). * p < 0.05, ** p < 0.01.
We assess consensus as the subset of cases in which both models predict the same HRD status, offering insight into uncertainty between models37,40. sDL and FM achieved a consensus rate of 71%, and demonstrated the highest AUC of 0.93 (95% CI: 0.87–0.98) in combined CT. The consensus between HCR and FM ranged from 61 to 67% (AUC: 0.89–0.91), while the consensus between sDL and FM was slightly higher (65–71%) and showed a higher AUC of 0.91–0.93.
To assess the effect of training data size on model performance, we retrained the models using 100, 50, 20, and 10% of the training set. The HCR model consistently exhibited the lowest and most variable AUCs, dropping below 0.5 when trained on only 10% of the data, indicating its high sensitivity to dataset size. Both the sDL model and the FM demonstrated an approximately linear AUC decline as training data size decreased (Fig. 3c). When using more than 10% of the training data, the FM consistently outperformed the sDL model.
HRD classification performance of foundation model
Based on the superior performance of the FM, we focused specifically on its predictive results in the following analyses. Among the HRD and HRP isogenic xenografts, the predicted HRD probabilities by FM (40 kVp CT) were significantly higher in the true HRD group than in the true HRP group on the test dataset (median [interquartile range, IQR]: HRD, 0.70 [0.45–0.79]; HRP, 0.21 [0.16–0.35]; p < 0.0001; Fig. 4a, left). This result indicates an alignment between the model prediction and the true HRD status. Similar results were observed when investigating the different tumor types separately. In the LNCaP AR xenografts, HRD probabilities differed significantly by gene mutation: FANCA-/- (n = 15; 0.75 [0.64–0.79]) and FANCD2-/- (n = 17; 0.67 [0.43–0.79]) xenografts, both classified as HRD, showed significantly higher predicted HRD probabilities compared to parental xenografts (classified as HRP, n = 16; 0.18 [0.14–0.21]; p < 0.0001; Fig. 4a, middle). No significant difference was observed between FANCA-/- and FANCD2-/- xenografts (p > 0.05). Similar results were observed in the HCT116 and DLD-1 xenografts: BRCA2-/- xenografts (classified as HRD) had significantly higher predicted HRD probabilities compared to both parental and DNA-PKcs-/- xenografts (classified as HRP) (HCT116: p < 0.0001; DLD-1: p < 0.05; Fig. 4a). Together, these results show that the predicted HRD probability was higher in the true HRD group compared to the true HRP group, and no significant differences were observed within the HRD group.
a Predicted HRD probabilities for true HRP and HRD groups across all tumors (left) and individual tumor types (LNCaP AR, HCT116, and DLD-1). b Confusion matrices show classification performance in all tumors and within individual tumor types. * p < 0.05, **** p < 0.0001, ns non-significant.
The quantitative performance of the FM for HRD prediction across different tumor types is illustrated in Fig. 4b. Overall, FM demonstrated robust predictive ability, with an AUC of 0.88 (95% confidence interval [CI], 0.82–0.93), sensitivity of 66%, and specificity of 87%. Among the three tumor types, the highest performance was observed in LNCaP AR xenografts (AUC, 0.96; sensitivity, 69%; specificity, 100%), followed by HCT116 xenografts (AUC, 0.94; sensitivity, 81%; specificity, 90%). Performance was lower in DLD-1 xenografts (AUC, 0.72; sensitivity, 44%; specificity, 69%).
HRD-related features predicted DNA damage with foundation model
A total of 86 tumor tissue samples were obtained from both the treatment and control groups 48 h post-treatment. After excluding four mice without pre-treatment scans, 82 mice were included in the DNA damage intensity (DDI) prediction (training set: n = 57; test set: n = 25). No significant difference in DNA damage (γ-H2AX) was observed overall between the true HRP and HRD subgroups in the control group (p > 0.05; Fig. 5a). Comparatively, after CP-506 treatment, DNA damage was significantly higher in the true HRD subgroup compared to the true HRP subgroup both overall and within each tumor cell line (Overall: p < 0.001; LNCaP AR: p < 0.01; HCT116: p < 0.05; DLD-1: p < 0.01; Fig. 5a), indicating a differential treatment response based on HRD status. To assess whether pre-treatment CT-based HRD classification could directly stratify subsequent CP-506-induced DNA damage, we compared DDIs between the predicted HRP and predicted HRD subgroups. After treatment, DDI was significantly higher in the predicted HRD subgroup than in the predicted HRP subgroup overall and within the LNCaP AR and HCT116, whereas no difference was observed in DLD-1 (Overall: p < 0.001; LNCaP AR: p < 0.01; HCT116: p < 0.05; DLD-1: p > 0.05; Fig. 5b).
a DNA damage intensities (DDIs) stratified by true HRP and HRD subgroups across control and CP-506 treatment groups. b DDIs stratified by predicted HRP and HRD subgroups from pre-treatment CT scans across the control and CP-506 treatment groups. c ROC curves for DNA damage classification (high vs. low). d Predicted probabilities of DNA damage in the true high and low DNA damage subgroups. e Confusion matrices show the model’s prediction performance for DNA damage (high vs. low) on the training and test datasets. * p < 0.05; ** p < 0.01; *** p < 0.001; ****p < 0.0001; ns. non-significant.
For DDI prediction, damage levels were dichotomized as high (≥ median) or low (< median). A logistic regression model trained to predict DDI achieved an AUC of 0.92 (95% CI, 0.83–1.00) on the training dataset and 0.75 (95% CI, 0.63–0.87) on the test dataset (Fig. 5c). Predicted probabilities differed significantly between the true low and high DNA damage subsets in both the training (p < 0.0001, Fig. 5d) and test datasets (p < 0.05, Fig. 5d), supporting the model’s capacity to distinguish between DNA damage levels. Specificity was high across datasets (96% for training, 100% for test), while sensitivity was higher on the training set (100%) than on the test set (50%, Fig. 5e).
HRD-related features predicted survival with foundation model
Pre-treatment scans were utilized for the survival analysis, maintaining the same data split as used in the HRD classification task (training set: n = 201, test set: n = 82). Because mice in this imaging experiment were sacrificed after treatment in order to assess DNA damage, event-free survival (EFS) times for these animals were synthetically generated from a parallel tumor growth experiment (see “Methods”). Upon CP-506 treatment, EFS was significantly longer in the true HRD groups compared to the HRP groups (p < 0.0001, Supplementary Fig. S3). Using HRD-related features, a Cox model was trained to predict tumor growth risk score, and mice were stratified into high- and low-risk groups. In this exploratory analysis, Kaplan–Meier (KM) curves showed a significant difference in EFS between the two groups (p < 0.0001, Fig. 6a, left), with a hazard ratio of 0.25 (95% CI, 0.23–0.27) and a concordance index (C-index) of 0.69 (95% CI, 0.63–0.77). Across three tumor types, KM curves showed similar significant EFS differences between high- and low-risk groups, with hazard ratios ranging from 0.24 to 0.28. The C-index was 0.69 (95% CI, 0.57–0.79) in LNCaP AR, 0.73 (95% CI, 0.58–0.89) in HCT116, and 0.59 (95% CI, 0.49–0.78) in DLD-1 xenografts. Subsequently, the intersection of risk groups and predicted HRD classes (yielded four groups: high-risk/predicted HRD, high-risk/predicted HRP, low-risk/predicted HRD, and low-risk/predicted HRP) revealed distinct survival patterns (p < 0.0001, Fig. 6b), with the low-risk/predicted HRD group exhibited the longest EFS. This trend was consistent across cell lines, with more pronounced separation in LNCaP AR and HCT116, and weaker discrimination observed in DLD-1.
a EFS curves for low- and high-risk groups, overall and for individual tumor types (LNCaP AR, HCT116, and DLD-1). b EFS curves for combined stratification by risk group and predicted HRD status: low-risk/predicted HRP, low-risk/predicted HRD, high-risk/predicted HRP, and high-risk/predicted HRD groups. Risk scores were predicted by a multivariate Cox proportional hazards regression model using HRD-related features from foundation model (FM). Low and high-risk groups were stratified based on the median risk score on the training set and validated on the test set.
Model interpretability
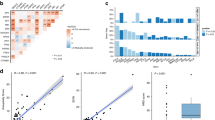
Using the regression concept vector (RCV) method, we first identified which radiomic features were effectively learned by the FM. After excluding highly volume-correlated features (n = 63), we analyzed the remaining features (n = 309) for their contribution to HRD classification. The distribution of R² values (Fig. 7b) showed that most features (n = 269, 87%) exhibited positive learnability (R² > 0), with 62 classified as very weak (0 ≤ R² < 0.2), 83 as weak (0.2 ≤ R² < 0.4), 87 as moderate (0.4 ≤ R² < 0.6), 31 as strong (0.6 ≤ R² < 0.8), and 6 as very strong (0.8 ≤ R² < 1). In contrast, 40 features exhibited negative R² values, indicating they were not captured by the FM and therefore did not contribute to model predictions. We then evaluated the contributions (bidirectional relevance [Br] scores) of learned radiomic features to HRD classification in the FM (Fig. 7a). Feature contributions varied between original and Laplacian of Gaussian (LoG)-filtered images, without a consistent pattern across filter types. The analysis identified the top 10 highest-impact features from those with at least strong learnability (Fig. 7b): Coarseness (LoG-2, Br = −1.000; LoG-1, Br = −0.852; LoG-3, Br = −0.801; original, Br = −0.719), GrayLevelNonUniformity (LoG-2, Br = −0.887), ClusterShade (LoG-1, Br = 0.864), ZonePercentage (LoG-3, Br = −0.821), Skewness (LoG-1, Br = 0.645), RootMeanSquare (LoG-1, Br = −0.644), and Busyness (original, Br = 0.638). Furthermore, we compared the feature contributions between the HCR and FMs. In the HCR model, eight out of twelve features showed consistent directionality with FM in contributions to HRD classification (Fig. 7c).
a Heatmap of feature contributions (Br) across different filters (original, LoG-1, LoG-2, LoG-3) and feature types. Colors indicate the direction of contribution to HRD classification (positive or negative). Features excluded due to high correlation with volume (Spearman’s ρ > 0.8) or poor learnability (R² < 0) are marked. b Joint distribution of feature contributions (Br) versus the coefficient of determination (R²) for all learned features, with the top 10 features (identified by the highest absolute bidirectional relevance (Br) scores and R² > 0.6) highlighted. Histograms show the distributions of Br and R² values. c Consistency comparison of feature contributions between the handcrafted radiomics (HCR) model (SHAP values) and the foundation model (normalized Br) on selected HCR features. Eight of twelve features showed consistent contribution directions for HRD classification. HRD homologous recombination deficiency, Br bidirectional relevance, LoG Laplacian of Gaussian filter, SHAP Shapley additive explanations.
Discussion
In this study, we applied the foundation model to predict HRD from preclinical CT images of isogenic xenografts. The FM demonstrated strong predictive performance for HRD status, which outperformed the HCR and sDL models. Furthermore, the HRD-related CT features identified by FM accurately predicted both DNA damage and EFS following CP-506 therapy. These results remained robust across LNCaP AR, HCT116, and DLD-1 isogenic xenografts. Therefore, the findings emphasize that the FM can effectively identify HRD-related information from preclinical CT images, establishing it as a valuable marker for both HRD detection and treatment response prediction. This non-invasive, automatic, state-of-the-art approach for HRD detection holds potential value to inform clinical decision-making.
Tumor heterogeneity manifests across various spatial scales, including the molecular, cellular, and radiological levels10,11. While directly measuring genetic mutations at the molecular or cellular level is highly accurate, it requires biopsies or invasive procedures18. Comparatively, capturing these features from diagnostic imaging offers a more accessible, non-invasive alternative. However, the extent to which imaging modalities can accurately reflect tumor phenotype and genotype is still being actively investigated. Previous work from our lab has shown that image features causally reflected genetic factors, and these features can be captured by CT-based radiomics24,41,42,43. However, the attempts to predict tumor genotypes using clinical imaging modalities have yielded inconsistent results across different diseases. For example, magnetic resonance imaging–based models predicting TP53 mutations have shown AUC values ranging from 0.74 to 0.95 across breast, prostate, and glioma cancers25,44,45. In high-grade serous ovarian cancer, some studies reported associations between BRCA mutation status and CT features26,46, while others found CT texture features associated with overall and progression-free survival but not significantly linked to BRCA mutations38,47,48,49. These inconsistencies may be due to the variations in tumor types, scanning protocols, and modeling methods. This variability was also observed in our findings, as the FM outperformed HCR and sDL, but its performance in DLD-1 was lower than in LNCaP AR and HCT116. One potential biological explanation is that DLD-1 is MMR-deficient (biallelic MSH6 loss, MSI) and exhibits an atypical DNA-repair capacity that can alter lesion processing50. In fact, in vitro experiments confirmed that DLD-1 cells require higher concentrations of the DNA-crosslinking agent CP-506 to achieve comparable cell kill, indicating a lower intrinsic sensitivity to therapy despite BRCA2 loss51. This reduced sensitivity was also reflected in our in vivo experiment: when treated with an identical dose of 600 mg/kg, DLD-1 xenografts exhibited less relative difference in DNA damage accumulation 48 h post-treatment between HRD and HRP groups compared to LNCaP AR and HCT116 cell lines (Fig. 4a). Overall, this suggests that HRD prediction models should be fine-tuned and evaluated for each cancer type, adapting to its unique microenvironment.
HRD classification models hold promise for patient stratification to maximize treatment benefit and reduce unnecessary treatment-induced toxicity43. For example, PARP inhibitors effectively target HRD tumors by the induction of synthetic lethality, i.e., simultaneous disruption of two DNA repair pathways resulting in cell death52. Genomic HRD assays are used as diagnostics to select patients who are most likely to benefit from PARP inhibitor maintenance therapy20,51,53. Therefore, optimizing thresholds for high specificity in our non-invasive model ensures treatment is specific to HRD patients. In addition, HRD tumors show enhanced sensitivity to DNA-damaging therapeutics, especially alkylating agents such as platinum-based chemotherapies. The mono-therapeutic efficacy of CP-506, a hypoxia-activated alkylating agent, has been shown to be more enhanced in HRD xenografts compared to HRP xenografts51. Therefore, the model requires high sensitivity to capture all potential HRD cases16. Additionally, the model remains robust across different CT energy levels, which simplifies imaging acquisition and reduces dependency on specific CT parameters.
HRD tumors have been shown to respond exceptionally well to PARP inhibitors and CP-50651,54,55. This supports that endogenous mutational processes might play a crucial role in shaping clinical outcomes. Multiple prospective cohorts have demonstrated that HRD status is an independent predictor of response to PARP inhibitors, with HRD-positive patients experiencing markedly longer progression-free survival than HRD-negative patients15,56. Previous studies show that biology-guided deep learning models for mutation detection can predict prognosis and immunotherapy response in gastric cancer57,58. Other studies have shown the correlation between CT image features and prognostic outcomes38,49, but it remained unclarified whether these CT image features reflected the HRD-related information. Our study, to the best of our knowledge, is the first one to use imaging-based HRD detection models to predict potential cancer treatment outcomes.
We demonstrated that HRD-related features captured in CT images can effectively predict treatment outcomes at both the microscopic and macroscopic levels. At the microscopic level, the model accurately predicts DNA damage, reflecting the biological mechanisms underlying CP-506 treatment. At the macroscopic level, it also assesses potential tumor survival, providing insight into how CP-506 treatment impacts overall tumor progression. This dual capability offers a comprehensive view of treatment efficacy, capturing both cellular responses and global tumor effects. However, it is important to note that the DNA damage intensity may be influenced by several factors. First, DDI was derived from cross-sectional histological analysis, which captures damage only in a single tumor slice and may therefore miss the full extent of damage throughout the tumor. Second, previous work has shown that tumor hypoxia, cellular reductase activity, and tissue pharmacokinetics can modulate CP-506 activation and consequently the magnitude and distribution of DNA damage59.
HRD, associated with genomic instability, often manifests as a more heterogeneous and disorganized texture. Previous studies have shown that radiomic features can effectively capture underlying tissue heterogeneity60,61. Consistent with this, our interpretability analysis of the foundation model also highlighted the essential role of texture heterogeneity in HRD classification. Using RCV, we found that the ten most important features in the foundation model all reflect the degree of texture heterogeneity, with greater heterogeneity linked to a higher probability of HRD status. For example, the feature “coarseness”—where a lower value indicates a more heterogeneous local texture-was a consistent contributor to HRD classification across different filters. In contrast, coarseness was excluded from the HCR model during feature selection due to collinearity. The foundation model offers potential advantages by capturing novel features that contribute to HRD classification, including characteristics that may not be predefined or captured by traditional HCR models.
However, there are several limitations in our study. Firstly, this study is based on preclinical xenograft models, which enabled image and histological-level validation but do not capture the full biological and clinical heterogeneity of human cancers. Secondly, the survival analysis, which relies on synthetic event times from the parallel experiment, is intended only to illustrate the potential prognostic value of HRD-related CT features. This analysis should be examined explicitly in future studies using directly observed survival data. Thirdly, our work is limited to a small number of prostate and colorectal cell lines with a modest sample size, which reduces power for subgroup analyses and may introduce cell-line-specific signals. Imaging was acquired using a micro-CT under fixed settings; differences between micro-CT and clinical CT (spatial resolution, contrast, reconstruction) may limit direct translatability and require further evaluation on human datasets. Finally, variation in training strategy, dataset size, and domain may affect transferability. Larger, multi-center clinical validation is therefore required.
In conclusion, we validated the efficiency of foundation models in preclinical data and successfully predicted HRD status and treatment responses using CT images. By extracting critical features directly from preclinical images, our method provides a non-invasive, early-stage diagnostic tool for HRD detection. However, before this approach can be translated into clinical practice, it needs to be validated using clinical data. If successfully validated, this method could serve as a non-invasive biomarker to identify patients most likely to benefit from alkylating agents or other therapies, both of which show enhanced efficacy in HRD tumors. Specifically, for CP-506, a hypoxia-activated alkylating agent, HRD radiomic signatures derived from foundation models could be complemented with hypoxia signatures to further improve patient stratification. This combined approach has the potential to enhance personalized treatment strategies and improve clinical outcomes. In a broader context, we have demonstrated that radiomics studies utilizing pretrained foundation models and synthetic data can be conducted on preclinical data despite its inherent data scarcity.
Methods
Experiment and CT image collection
This study was conducted at Maastricht University, involving 307 mice isogenic xenografts derived from three cancer cell lines: LNCaP AR (androgen-resistant prostate adenocarcinoma; NOD.Cg-PrkdcSCIDIl2rgtm1Wjl/SzJ; male; n = 132), HCT116 (colon carcinoma; NU-Foxn1 nu/nu; female; n = 105), and DLD-1 (colorectal adenocarcinoma; BALB/c nu/nu; male; n = 70). Each cancer cell line included both HRD and HRP isogenic xenografts. Specifically, the LNCaP AR xenografts consisted of parental (HRP, n = 45), FANCA-/- (HRD, n = 38), and FANCD2-/- (HRD, n = 49) isogenic xenografts; the DLD-1 xenografts included parental (HRP, n = 34) and BRCA2-/- (HRD, n = 36) isogenic xenografts; and the HCT116 xenografts included parental (HRP, n = 35), DNA-PKcs-/- (HRP, n = 36), and BRCA2-/- (HRD, n = 34) isogenic xenografts (Fig. 1). Mice were randomly stratified to either the control or the treatment group, ensuring the balanced distribution of HRD and HRP types between groups (Fig. 2a). Upon reaching a tumor starting volume (SV) of ca. 200 mm3 (mean ± s.e.m: 223.9 ± 4.7 mm3), the treatment group received CP-506, a hypoxia-activated prodrug with DNA-alkylating metabolites, at a dose of 600 mg/kg (QD1; intraperitoneally), while the control group received vehicle treatment (water for injection)51. Body weights and tumor dimensions were monitored at least three times per week throughout the study. Animal experiments were approved by the Centrale Commissie Dierproeven (AVD1070020198905) and conducted according to the institutional guidelines of Maastricht University. Detailed information of the mouse models is provided in the Supplementary Table S1.
CT scans were acquired before and after treatment using the X-Rad 225Cx cone beam micro-CT scanner. Pre-treatment scans were obtained prior to treatment once tumors reached the starting volume, while post-treatment CT scans were acquired at 6, 24, 48, and 72 h after the CP-506/vehicle dose injection. For each scan, mice were sedated with isoflurane inhalation (induction 4%, maintenance 2.5%) and positioned head supine on the scanning table. Sequential CT images were acquired at two different energy levels (40 and 80 kVp), with a reconstructed voxel size consistently maintained at 0.1 mm³.
After CT image collection, no-treatment scans (pre- and post-scans from the control group and pre-scans from the treatment group) were stratified into a training set (70%) and a test set (30%). Notably, both pre- and post-treatment scans of an individual mouse in the control group were assigned collectively to the same dataset to avoid information leaking. The treatment scans (i.e., post-scans from the treatment group) were used to investigate the impact of CP-506-induced DNA damage on the classification performance as an explorative objective. Therefore, the post-scans from the treatment group were designated as an interference test set with details provided in the Supplementary Figs. S3–S6.
CT ROI segmentation
To obtain tumor ROIs, CT images were delineated using a semi-automated segmentation pipeline. First, tumors in twenty scans were manually delineated by two operators (L.S. and J.J.) to train a DynUNet segmentation model with three-fold cross-validation on 40 kVp, 80 kVp and combined CT; for each energy level, the best model per fold (based on the Dice Similarity Coefficient) generated contours that were averaged into a “voting contour,” yielding twelve candidate contours per scan. These candidates were then corrected and jointly reviewed by four operators (S.K., L.S., J.v.d.L., and R.B.) to produce the final tumor ROIs.
HRD classification models
Handcrafted radiomics
The CT-based HCR pipeline was developed using the training dataset and their associated ROI masks. We extracted 93 features from the original CT images, including first-order statistics (n = 18), gray level co-occurrence matrix (GLCM, n = 24), gray level size zone matrix (GLSZM, n = 16), gray level dependence matrix (GLDM, n = 14), gray level run length matrix (GLRLM, n = 16), and neighboring gray tone difference matrix (NGTDM, n = 5). Additionally, we applied LoG filtering with kernel sizes of 0.1, 0.2, and 0.3 mm to generate filtered versions of the same 93 features for each kernel size. In total, 372 (i.e., 93\(\times\)4) features were extracted representing both the original and filtered image characteristics across different spatial scales. All features were extracted using PyRadiomics with a default bin width of 25 Hounsfield Units for discretizing the gray levels in CT images.
After feature extraction, we performed the feature selection using five-fold cross-validation (CV) on the training set to retain the most relevant features: (i) all features were z-score normalized; (ii) constant features, and those highly correlated with volume (Spearman’s correlation coefficient > 0.8) were excluded; (iii) highly inter-correlated features (Spearman) were clustered based on a correlation threshold; (iv) within each cluster, the feature with the highest univariate AUC for the HRD status was retained; (v) Lasso regression with regularization was applied to further reduce the feature set. The above feature selection was performed independently on the 40 and 80 kVp CT datasets, yielding two distinct feature sets. Additionally, all these selected features were concatenated to create a third, combined feature set. A support vector machine (SVM) classifier with regularization was trained within each CV fold to predict HRD probability for each input image. Hyperparameter tuning, including the correlation threshold, regularizations, and kernel type of the classifier, was conducted using grid search based on AUC. For comparison, we also trained additional machine-learning classifiers, including logistic regression and random forests. The trained classifiers of each fold in the CV were applied to the test set to predict HRD probabilities, and these predictions were averagely ensembled across the five folds to generate the final prediction.
Supervised deep learning approaches
A supervised ResNet-50 model was trained from scratch using 40 and 80 kVp CT scans separately, and an additional combined model was trained with a two-channel input formed by concatenating the two CT scans. To mitigate overfitting due to the limited dataset size, the number of channels in each intermediate ResNet-50 layer was reduced to one-fourth of its original size, lowering model complexity. Additionally, to enable effective training in our small dataset, an extensive data augmentation pipeline was applied, including random rotation, flipping, translation, shifting, zooming, and Gaussian noise. Cross-entropy loss was used during training, with the Adam optimizer. The initial learning rates were set to 10⁻³ for 40 kVp CT, 10⁻⁴ for 80 kVp CT, and 3 × 10⁻⁴ for combined CT models. These rates were reduced exponentially with a decay factor of 0.11/200. The L1 regularization coefficient was set at 10⁻⁵. Five-fold CV was also applied, and models were trained over 1000 epochs. The model with the highest validation AUC for each fold was retained. To assess whether pretraining on relatively large datasets affects performance, we also fine-tuned a ResNet-50 initialized with MedicalNet weights using identical data splits and training settings. Its performance was reported in Supplementary results and Table S3-S4.
Foundation model
We utilized an FM based on a ResNet-50 encoder, which has been pre-trained on a comprehensive human CT dataset due to its strong generalization to out-of-distribution tasks and significant associations with cancer biomarkers28,62. We adapted this pre-trained FM as a feature extractor and built upon it by adding a three-layer multilayer perceptron (MLP) as classifier28. For the 40 and 80 kVp CT scans, the masked raw images were input into the ResNet-50 encoder to extract 4096-dimensional feature representations. The MLP classifier then reduced these feature dimensions through three layers, sequentially compressing them from 512 to 64, and finally to 2 dimensions for the classifier output. The first two layers used LeakyReLU activation functions, while the final layer employed SoftMax to produce one-hot probabilities63. For the combined CT model, the single-energy features from 40 and 80 kVp were concatenated, resulting in an 8192-dimensional feature vector for the MLP classifier. To enhance model robustness, we applied data augmentation techniques, including random rotations, flipping, and translations28. Consistent with the HCR model, training and testing were performed using five-fold CV. During training, each model was trained for 1000 epochs, with weights updated using the Adam optimizer with a learning rate of 10⁻⁶ and cross-entropy loss. The model with the highest validation AUC in each fold was retained, and these models were ensembled to generate the final HRD prediction (Fig. 2b).
DNA damage quantification
CP-506 is a hypoxia-activated prodrug that releases cytotoxic metabolites in hypoxic tumor regions, leading to DNA interstrand crosslinks and double-strand breaks20,51. DNA damage can be detected within 6 h of administration, but part of this early damage is repairable. By 48 h post-treatment, the remaining γ-H2AX signal predominantly reflects persistent, unrepaired DNA damage59. DNA damage intensity (DDI) was therefore selected as a metric for assessing cellular treatment response. To evaluate DDI across different genotypes, tumors were collected from mice 48 h post-treatment from both treatment groups. Immunohistochemical staining of γ-H2AX, a well-established marker for DNA double-strand breaks, was performed on formalin-fixed and paraffin-embedded tumor sections according to the protocol previously described (Representative immunohistochemistry images see Supplementary Fig. S7)64. Tumor sections were scanned using a Precipoint M8 microscope with a 20x objective. A DynUNet model was used to automatically segment the vital areas of the histological tumor section51. These segmentations were reviewed and manually corrected by L.S., who was blinded to treatment labels, using ImageJ 1.54f65. Quantification of γ-H2AX staining was then performed using QuPath version 0.4.366. Staining vectors were first set on one entire tumor section per tumor cell line. Tissue boundaries and individual cells were detected using the simple tissue detection and positive cell detection functions, with parameters optimized, ensuring similar settings for the analysis of each isogenic model per tumor cell line. The γ-H2AX staining intensity per detected tumor cell within the vital tumor regions was averaged to provide a DDI value per isogenic tumor. To reduce inter-cell-line variation in staining intensity and nuclear density, DDI values were normalized by dividing the mean DDI of the control group from the same tumor cell line.
Synthetic data of tumor growth
To investigate whether HRD influences post-treatment survival, tumor growth data are important. However, since the mice were sacrificed after the treatment to assess DNA damage, tumor growth could not be measured. To address this, we conducted a parallel animal experiment and sampled tumor growth data in the main experiment. This approach assumes that xenografts under identical experimental conditions follow the similar tumor growth. In this parallel experiment, 151 mice bearing the same types of isogenic xenograft were treated with CP-506 (600 mg/kg, QD5) and vehicle, to replicate the main experimental treatment as previously described16. Tumor size was measured using a Vernier caliper at regular intervals until each tumor quadrupled from its starting volume (T4xSV). Tumors that did not reach this threshold until the end of the experiment were considered censored. We then fitted a Weibull distribution, commonly used to model biological time-to-event data, to the time for each xenograft type using maximum likelihood estimation. Based on the fitted distributions, we sampled tumor quadruple time for each mouse in the main experiment and used it as time-to-event for the survival analysis. As an exploratory analysis, this synthetic EFS was used only for demonstrating the potential value of HRD features for survival prediction and does not represent observed survival outcomes in the main experiment.
HRD-related features for predicting DNA damage
Due to the limited tissue sample size and variability from cross-sectional cuts and staining processes, we binarized the DDI to increase the signal-to-noise ratio. Specifically, we used the median DDI to dichotomize the DDI into categories that reflected high (i.e., DDI larger than or equal to the median value) and low DNA (i.e., DDI smaller than the median value) damage. To examine whether pre-treatment HRD-related features predict treatment-induced DNA damage, we implemented a logistic regression classifier with l2-regularization to predict DDI categories. The inputs to the logistic regression model included treatment status (0 = control, 1 = treatment) and the 64-dimensional HRD-related feature vector. These features were extracted from the final layer of the MLP in the FM using pre-treatment scans. The training and test sets followed the same partitioning as in the HRD classification. Nested CV was applied to obtain a reliable estimate of classification performance. The outer loop used five-fold CV, consistent with the HRD classification model, to evaluate overall performance. Within each outer loop, we employed leave-one-out cross-validation (LOOCV) to fine-tune the regularization parameter to prevent overfitting. The trained classifier in each outer loop was applied to the test set for prediction. The training and test performance were finally averaged across outer loops to obtain the model evaluation metrics.
HRD-related features for predicting survival
To predict tumor treatment response, we performed a survival analysis using EFS, which was defined as T4xSV. First, EFS was compared between HRD and HRP groups using KM survival analysis. Next, the relationship between HRD-related features and EFS was assessed using a multivariate Cox proportional hazards regression model, generating individual risk scores. Based on these scores in the training set, mice were stratified into high- and low-risk groups (i.e., above or below median risk scores) in both the training and test sets. Finally, risk groups were combined with predicted HRD classes to create four prognostic subgroups: low-risk/predicted HRP, low-risk/predicted HRD, high-risk/predicted HRP, and high-risk/predicted HRD. Survival across these subgroups was analyzed using KM curves. To quantify prediction stability, the EFS data were resampled 100 times. For each resampling iteration, the Cox model was retrained on the training set and applied on the test set. Meanwhile, the same training and test partition, and five-fold CV as in the HRD classification were applied. The KM curves were obtained by averaging across all iterations and CV folds.
Statistical analysis and model evaluation
To compare characteristics between the training and test sets, the Mann–Whitney U test was utilized for continuous variables, and the chi-squared test was employed for categorical variables. The performances of the HRD tumor classification models were evaluated using the ROC curve, AUC, accuracy, sensitivity, and specificity. Calibration was performed on all classification models using Platt scaling on the validation set and applied to the test set. Calibration was evaluated with reliability diagrams, the Brier score, and expected calibration error (ECE). The cutoff thresholds for the probability of HRD status were determined using the maximum sensitivity and specificity on the training set. The threshold for the ensembled probability on the test set was set to the mean thresholds across each fold. Model consensus was defined as the subset of cases in which both models produced identical binary predictions. We reported both the consensus rate and the AUC computed on this consensus subset. All 95% CIs of the above-mentioned metrics were calculated by 1000-time bootstrapping. The DeLong tests were applied to the ROC curves on the test set of the HRD classification to assess method differences. To determine if the HRD model’s predictions significantly differ across various mutation types within each cell line, the predictions were compared using the Mann-Whitney U test. The Fisher’s exact test was applied to evaluate whether the confusion matrix of the HRD predictions was significantly above the chance level (50%).
Similarly, the Mann–Whitney U test was used to determine the significance of DNA damage between treatment groups. For the DDI classification model, the Mann–Whitney U test was employed to determine the significance between predicted low and high DNA damage groups, and the Fisher’s exact test was used to assess the significance of the model’s confusion matrix.
To evaluate the prognostic performance of the Cox proportional hazards model, the hazard ratio between high-risk and low-risk groups was calculated, and the log-rank test was performed to assess statistical significance. The C-index was computed to quantify the Cox model’s discriminative ability in predicting EFS. To assess the significance of EFS for intersected stratification of risk groups and predicted HRD classes, a multivariate log-rank test was performed.
False discovery rate correction was applied using the Benjamini–Hochberg method when multiple comparisons were performed67. All statistical analyses were performed with SciPy 1.11.3. p values below 0.05 were considered statistically significant.
Model interpretability
Given the interpretability of radiomic features, we adapted a methodology by using continuous radiomic features to explain the decision-making of FM68. A gradient-based regression concept vector (RCV) method69 was used to answer two key questions: (i) which radiomic features were learned by the FM, and (ii) whether these learned radiomic features contributed to HRD classification. RCV employs least squares linear regression by mapping the 4096-dimensional foundation features to each radiomic feature. The coefficient of determination (R²) quantifies the feature learnability of the FM with respect to each radiomic feature. An R² value between 0 and 1 indicates that the FM has learned the radiomic feature, with values closer to 1 showing stronger learning. A negative R² means the feature has not been learned. Subsequently, RCV generates the bidirectional relevance (Br) score to measure the impact of each radiomic feature on HRD classification. The absolute Br score reflects the feature’s contribution strength: a positive Br score means higher radiomic feature values increase the likelihood of HRD classification, while a negative Br score indicates the opposite. The RCV includes the same features in the HCR model, with all constant, shape, and highly volume-correlated features excluded.
Furthermore, to explore whether the radiomic features have similar impacts in both the HCR and FMs, we performed a sign comparison of feature contributions between the two models. Specifically, Shapley Additive Explanations (SHAP) were used to quantify feature importance in the HCR model, while Br scores were used for the FM70. Both SHAP values and Br scores were averaged across CV folds and then normalized between −1 and 1.
Data availability
The CT and histopathology data will be publicly accessible upon publication: [https://huggingface.co/datasets/ShengKuang93/HRD_CT_Preclinical].
Code availability
All scripts will be publicly accessible on the GitHub upon publication: https://github.com/ShengKuangCN/HRD_CT_Preclinical.
References
Lord, C. J. & Ashworth, A. BRCAness revisited. Nat. Rev. Cancer 16, 110–120 (2016).
Bell, D. et al. Integrated genomic analyses of ovarian carcinoma. Nature 474, 609–615 (2011).
Koboldt, D. C. et al. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
Ren, L. et al. Pan-cancer analysis of homologous recombination deficiency and homologous recombination repair–associated gene alterations in solid tumors from a large Asian cohort. BMC Cancer 25, 946 (2025).
Heeke, A. L. et al. Prevalence of Homologous Recombination–Related Gene Mutations Across Multiple Cancer Types. JCO Precis. Oncol. 1–13 https://doi.org/10.1200/PO.17.00286 (2018).
Devlin, M.-J. & Miller, R. Genomic testing in high-grade serous ovarian cancer: current options and future development. Int. J. Gynecol. Cancer 33, 358–363 (2023).
Pritchard, C. C. et al. Inherited DNA-repair gene mutations in men with metastatic prostate cancer. N. Engl. J. Med. 375, 443–453 (2016).
Amodio, V., Vitiello, P. P., Bardelli, A. & Germano, G. DNA repair-dependent immunogenic liabilities in colorectal cancer: opportunities from errors. Br. J. Cancer 131, 1576–1590 (2024).
Reilly, N. M., Novara, L., Di Nicolantonio, F. & Bardelli, A. Exploiting DNA repair defects in colorectal cancer. Mol. Oncol. 13, 681–700 (2019).
Cunnea, P. et al. Spatial and temporal intra-tumoral heterogeneity in advanced HGSOC: Implications for surgical and clinical outcomes. Cell Rep. Med. 4, 101055 (2023).
von Wahlde, M.-K. et al. Intratumor heterogeneity of homologous recombination deficiency in primary breast cancer. Clin. Cancer Res. 23, 1193–1199 (2017).
Zhang, L. et al. Impact of homologous recombination deficiency on outcomes in patients with triple-negative breast cancer treated with carboplatin-based neoadjuvant chemotherapy: secondary analysis of the NeoCART randomized clinical trial. JCO Precis. Oncol. e2200337 https://doi.org/10.1200/PO.22.00337 (2023).
O’Connor, M. J. Targeting the DNA damage response in cancer. Mol. Cell 60, 547–560 (2015).
Lord, C. J. & Ashworth, A. PARP inhibitors: synthetic lethality in the clinic. Science 355, 1152–1158 (2017).
Miller, R. E. et al. ESMO recommendations on predictive biomarker testing for homologous recombination deficiency and PARP inhibitor benefit in ovarian cancer. Ann. Oncol. 31, 1606–1622 (2020).
van der Wiel, A. M. A. et al. Selectively targeting tumor hypoxia with the hypoxia-activated prodrug CP-506. Mol. Cancer Ther. 20, 2372–2383 (2021).
Davies, H. et al. HRDetect is a predictor of BRCA1 and BRCA2 deficiency based on mutational signatures. Nat. Med. 23, 517–525 (2017).
Hoppe, M. M., Sundar, R., Tan, D. S. P. & Jeyasekharan, A. D. Biomarkers for homologous recombination deficiency in cancer. JNCI J. Natl. Cancer Inst. 110, 704–713 (2018).
Zhang, M. et al. Inferring homologous recombination deficiency of ovarian cancer from the landscape of copy number variation at subchromosomal and genetic resolutions. Front. Oncol. 11, 772604 (2021).
van der Wiel, A. M. A. et al. Homologous recombination deficiency scar: mutations and beyond—implications for precision oncology. Cancers 14, 4157 (2022).
Foulkes, W. D. & Polak, P. Probing the relevance of BRCA1 and BRCA2 germline pathogenic variants beyond breast and ovarian cancer. JNCI J. Natl. Cancer Inst. djae184 https://doi.org/10.1093/jnci/djae184 (2024).
Lazard, T. et al. Deep learning identifies morphological patterns of homologous recombination deficiency in luminal breast cancers from whole slide images. Cell Rep. Med. 3, 100872 (2022).
Aerts, H. J. W. L. et al. Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach. Nat. Commun. 5, 4006 (2014).
Panth, K. M. et al. Is there a causal relationship between genetic changes and radiomics-based image features? An in vivo preclinical experiment with doxycycline inducible GADD34 tumor cells. Radiother. Oncol. J. Eur. Soc. Ther. Radiol. Oncol. 116, 462–466 (2015).
Zhang, X. et al. Radiomics strategy for molecular subtype stratification of lower-grade glioma: detecting IDH and TP53 mutations based on multimodal MRI. J. Magn. Reson. Imaging 48, 916–926 (2018).
Liu, M., Ge, Y., Li, M. & Wei, W. Prediction of BRCA gene mutation status in epithelial ovarian cancer by radiomics models based on 2D and 3D CT images. BMC Med. Imaging 21, 180 (2021).
A clinically relevant computed tomography (CT) radiomics strategy for intracranial rodent brain tumour monitoring | Scientific Reports. https://www.nature.com/articles/s41598-024-52960-1.
Pai, S. et al. Foundation model for cancer imaging biomarkers. Nat. Mach. Intell. 6, 354–367 (2024).
Zhang, S. & Metaxas, D. On the challenges and perspectives of foundation models for medical image analysis. Med. Image Anal. 91, 102996 (2024).
O’Farrell, A. C. et al. Implementing systems modelling and molecular imaging to predict the efficacy of BCL-2 inhibition in colorectal cancer patient-derived xenograft models. Cancers 12, 2978 (2020).
Montoya, C. et al. Predicting response to immunotherapy in non-small cell lung cancer- from bench to bedside. Front. Oncol. 13, 1225720 (2023).
Mihaylov, I. B. et al. Toward prediction of abscopal effect in radioimmunotherapy: pre-clinical investigation. PLoS ONE 16, e0255923 (2021).
Brown, K. H. et al. Characterisation of quantitative imaging biomarkers for inflammatory and fibrotic radiation-induced lung injuries using preclinical radiomics. Radiother. Oncol. 192, 110106 (2024).
Müller, J. et al. Radiomics-based tumor phenotype determination based on medical imaging and tumor microenvironment in a preclinical setting. Radiother. Oncol. J. Eur. Soc. Ther. Radiol. Oncol. 169, 96–104 (2022).
Drayson, O. G. G., Montay-Gruel, P. & Limoli, C. L. Radiomics approach for identifying radiation-induced normal tissue toxicity in the lung. Sci. Rep. 14, 24256 (2024).
Gong, J. et al. A short-term follow-up CT based radiomics approach to predict response to immunotherapy in advanced non-small-cell lung cancer. OncoImmunology 11, 2028962 (2022).
Primakov, S. P. et al. Automated detection and segmentation of non-small cell lung cancer computed tomography images. Nat. Commun. 13, 3423 (2022).
Meier, A. et al. Association between CT-texture derived tumor heterogeneity, outcomes, and BRCA mutation status in patients with high grade serous ovarian cancer. Abdom. Radiol. N.Y. 44, 2040–2047 (2019).
Wu, Y. et al. CT-based radiomics predicts HRD score and HRD status in patients with ovarian cancer. Front. Oncol. 14, 1477759 (2024).
Beuque, M. P. L. et al. Combining deep learning and handcrafted radiomics for classification of suspicious lesions on contrast-enhanced mammograms. Radiology 307, e221843 (2023).
Leijenaar, R. T. H. et al. External validation of a radiomic signature to predict p16 (HPV) status from standard CT images of anal cancer patients. Sci. Rep. 13, 7198 (2023).
Leijenaar, R. T. et al. Development and validation of a radiomic signature to predict HPV (p16) status from standard CT imaging: a multicenter study. Br. J. Radiol. 91, 20170498 (2018).
Rios Velazquez, E. et al. Somatic mutations drive distinct imaging phenotypes in lung cancer. Cancer Res. 77, 3922–3930 (2017).
Chen, R. et al. Association of pathological features and multiparametric MRI-based radiomics with TP53-mutated prostate cancer. J. Magn. Reson. Imaging 60, 1134–1145 (2024).
Sun, K., Zhu, H., Chai, W. & Yan, F. TP53 mutation estimation based on MRI radiomics analysis for breast cancer. J. Magn. Reson. Imaging 57, 1095–1103 (2023).
Cao, Y. et al. CT-based radiomics nomogram analysis for assessing BRCA mutation status in patients with high-grade serous ovarian cancer. Acta Radio. 64, 2802–2811 (2023).
Avesani, G. et al. CT-based radiomics and deep learning for BRCA mutation and progression-free survival prediction in ovarian cancer using a multicentric dataset. Cancers 14, 2739 (2022).
Boehm, K. M. et al. Multimodal data integration using machine learning improves risk stratification of high-grade serous ovarian cancer. Nat. Cancer 3, 723–733 (2022).
Vargas, H. A. et al. Radiogenomics of high-grade serous ovarian cancer: multireader multi-institutional study from the Cancer Genome Atlas Ovarian Cancer Imaging Research Group. Radiology 285, 482–492 (2017).
Lózsa, R. et al. DNA mismatch repair protects the genome from oxygen-induced replicative mutagenesis. Nucleic Acids Res. 51, 11040–11055 (2023).
Schuitmaker, L. et al. Deficiencies in the Fanconi anemia or the homologous recombination pathway enhance the antitumor effects of the novel hypoxia-activated prodrug CP-506. Preprint at https://doi.org/10.1101/2025.05.21.655302 (2025).
Rose, M., Burgess, J. T., O’Byrne, K., Richard, D. J. & Bolderson, E. PARP inhibitors: clinical relevance, mechanisms of action and tumor resistance. Front. Cell Dev. Biol. 8, 564601 (2020).
Callens, C. et al. Shallow whole genome sequencing approach to detect Homologous recombination deficiency in the PAOLA-1/ENGOT-OV25 phase-III trial. Oncogene 42, 3556–3563 (2023).
Moore, K. et al. Maintenance olaparib in patients with newly diagnosed advanced ovarian cancer. N. Engl. J. Med. 379, 2495–2505 (2018).
van der Wiel, A. Exploiting tumor DNA repair status with the novel hypoxia-activated DNA alkylating agent CP-506. Mol. Cancer Ther. 20, 2372–2383 (2021).
Homologous recombination deficiency testing to inform patient decisions about niraparib maintenance therapy for high-grade serous or endometrioid epithelial ovarian cancer: a health technology assessment. Ont. Health Technol. Assess. Ser. 23, 1–188 (2023).
Jiang, Y. et al. Biology-guided deep learning predicts prognosis and cancer immunotherapy response. Nat. Commun. 14, 5135 (2023).
Jiang, Y. et al. Non-invasive tumor microenvironment evaluation and treatment response prediction in gastric cancer using deep learning radiomics. Cell Rep. Med. 4, 101146 (2023).
Jackson-Patel, V. et al. Tissue pharmacokinetic properties and bystander potential of hypoxia-activated prodrug CP-506 by agent-based modelling. Front. Pharmacol. 13, 803602 (2022).
Khorrami, M. et al. Changes in CT radiomic features associated with lymphocyte distribution predict overall survival and response to immunotherapy in non–small cell lung cancer. Cancer Immunol. Res. 8, 108–119 (2020).
Liu, T. et al. Radiomic signatures reveal multiscale intratumor heterogeneity associated with tissue tolerance and survival in re-irradiated nasopharyngeal carcinoma: a multicenter study. BMC Med. 21, 464 (2023).
He, K., Zhang, X., Ren, S. & Sun, J. Deep Residual Learning for Image Recognition. in 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 770–778 https://doi.org/10.1109/CVPR.2016.90 (2016).
Xu, B., Wang, N., Chen, T. & Li, M. Empirical evaluation of rectified activations in convolutional network. Preprint at https://doi.org/10.48550/arXiv.1505.00853 (2015).
Yaromina, A. et al. Overcoming radioresistance with the hypoxia-activated prodrug CP-506: A pre-clinical study of local tumour control probability. Radiother. Oncol. 186, 109738 (2023).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Bankhead, P. et al. QuPath: open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Methodol. 57, 289–300 (1995).
Salahuddin, Z., Woodruff, H. C., Chatterjee, A. & Lambin, P. Transparency of deep neural networks for medical image analysis: a review of interpretability methods. Comput. Biol. Med. 140, 105111 (2022).
Graziani, M., Andrearczyk, V. & Müller, H. Regression concept vectors for bidirectional explanations in histopathology. In International Workshop on Machine Learning in Clinical Neuroimaging 124–132 (2018).
Lundberg, S. M. & Lee, S.-I. A unified approach to interpreting model predictions. in Proc. 31st International Conference on Neural Information Processing Systems 4768–4777 (Curran Associates Inc., 2017).
Acknowledgements
This work was supported by the ERC PoC Reverse the Advantage, HYPOXIMMUNO, Dutch Cancer Society (KWF Kankerbestrijding 14449/2021-PoC). Authors acknowledge financial support from the European Union’s Horizon research and innovation program under grant agreement: ImmunoSABR n° 733008, CHAIMELEON n° 952172, EuCanImage n° 952103, IMI-OPTIMA n° 101034347, RADIOVAL (HORIZON-HLTH-2021-DISEASE-04-04) n°101057699, EUCAIM (DIGITAL-2022-CLOUD-AI-02) n°101100633, GLIOMATCH n° 101136670, AIDAVA (HORIZON-HLTH-2021-TOOL-06) n°101057062, REALM (HORIZON-HLTH-2022-TOOL-11) n° 101095435.
Author information
Authors and Affiliations
Contributions
S.K., L.S., L.J.D., H.C.W., and P.L. conceived the idea of the article. A.v.d.W., S.K., L.S., J.v.d.L., N.L., R.B., and J.J. participated in the data acquisition and delineation. S.K., L.S., M.W., and Z.S. implemented the analysis. S.K., M.W., and L.S. drafted the manuscript. All authors edited the manuscript. A.Y., L.J.D., H.C.W, and P.L. supervised the work. P.L. approved the submitted version and has agreed both be personally accountable for the author’s own contributions and to ensure that questions related to the accuracy or integrity of any part of the work, even ones in which the author was not personally involved, are appropriately investigated, resolved, and the resolution documented in the literature.
Corresponding author
Ethics declarations
Competing interests
P.L.: none related to the current manuscript; outside of current manuscript: grants/sponsored research agreements from Radiomics SA, Convert Pharmaceuticals SA, and LivingMed Biotech srl. He received a presenter fee and/or reimbursement of travel costs/consultancy fee (in cash or in kind) from Astra Zeneca, BHV srl & Roche. P.L. has/had minority shares in the companies Radiomics SA, Convert pharmaceuticals SA, Comunicare SA, LivingMed Biotech srl and Bactam srl. PL is co-inventor of two issued patents with royalties on radiomics (PCT/NL2014/050248 and PCT/NL2014/050728), licensed to Radiomics SA; one issued patent on mtDNA (PCT/EP2014/059089), licensed to ptTheragnostic/DNAmito; one granted patent on LSRT (PCT/ P126537PC00, US patent No. 12,102,842), licensed to Varian; one issued patent on Radiomic signature of hypoxia (U.S. Patent 11,972,867), licensed to a commercial entity; one issued patent on Prodrugs (WO2019EP64112) without royalties; one non-issued, non-licensed patents on Deep Learning-Radiomics (N2024889) and three non-patented inventions (softwares) licensed to ptTheragnostic/DNAmito, Radiomics SA and Health Innovation Ventures). P.L. confirms that none of the above entities were involved in the preparation of this paper. H.C.W. has, outside of the current manuscript, minority shares in the company Radiomics SA. L.J.D. has, outside of the current manuscript, minority shares in the companies Convert Pharmaceuticals SA and LivingMed Biotech srl. L.J.D. is co-inventor of a granted patent on LSRT (PCT/ P126537PC00, US patent No. 12,102,842), licensed to Varian. The other authors do not have a competing interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kuang, S., Schuitmaker, L., Wu, M. et al. Predicting homologous recombination deficiency and treatment responses using a computed tomography-based foundation model: a preclinical study. npj Precis. Onc. 10, 121 (2026). https://doi.org/10.1038/s41698-026-01322-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-026-01322-3