Abstract
Precise evaluation of bone gain after maxillary sinus augmentation is critical for optimizing implant therapy yet manual measurements remain time consuming. This study validated a fully automated deep learning system named SA-ai to quantify bone augmentation. A paired CBCT dataset from 85 patients was used to train and test the system which integrates a 2D U-Net for sinus contour and a 3D V-Net for maxilla segmentation. The system achieved a Dice coefficient of 93.2% and registration RMSE of 1.046 mm. Clinical validation against manual measurements showed excellent agreement for bone volume (ICC = 0.993) and other parameters. Bias analysis confirmed measurement stability while workflow efficiency improved over 20-fold compared to manual methods. This registration subtraction paradigm delivers an automated and objective solution for longitudinal monitoring of bone graft volume including one-stage implant cases potentially standardizing clinical evaluation of post augmentation bone dynamics.
Similar content being viewed by others
Introduction
Insufficient alveolar bone height in the posterior maxilla presents a common challenge in oral and maxillofacial surgery, particularly for implant dentistry1. Due to factors, such as periodontitis, post-extraction alveolar ridge resorption, and maxillary sinus pneumatization, an estimated 33.3 to 54.2% of patients require maxillary sinus augmentation (SA) for bone grafting prior to implant placement2,3. SA involves surgically accessing the maxillary sinus and elevating its Schneiderian membrane, bone graft material is then placed into the space created between the sinus floor and the elevated membrane4. This procedure, which is essential for obtaining adequate bone volume to ensure the primary stability and long-term success of dental implants, is currently considered the gold standard for managing vertical bone deficiency in the posterior maxilla5. Clinically, SA may be performed as a one-stage procedure (simultaneous implant placement and bone grafting) or a two-stage protocol (implant placement after graft healing)6. Regardless of the approach, one of the key determinants of success is precise control of volumetric resorption and long-term stability of the grafted material during healing7,8. All grafting materials inevitably undergo varying degrees of resorption and remodeling. Accurately understanding and quantifying these volumetric changes is critically instructive for selecting appropriate graft materials, anticipating and compensating for expected resorption, and ultimately optimizing surgical outcomes9.
Cone-beam computed tomography (CBCT), owing to its relatively low radiation dose, high spatial resolution, and three-dimensional reconstruction capability, has become the imaging modality of choice for post-operative evaluation following SA—particularly for assessing volumetric changes of graft materials10. Despite these diagnostic advantages, substantial methodological challenges remain in achieving precise and reproducible 3D volumetric quantification of bone graft materials11. To date, no internationally standardized workflow exists for measuring post-SA graft volume using CT or CBCT imaging12. Current techniques rely predominantly on manual or semi-manual contouring of graft boundaries across serial tomographic slices13. These approaches are time-consuming and labor-intensive, and they are highly susceptible to inter- and intra-observer variability, potentially compromising measurement accuracy and reliability14. These inherent limitations underscore an urgent need for more efficient and objective analytical tools.
Deep learning (DL), a rapidly advancing frontier in artificial intelligence, has demonstrated potential in computer vision, especially in medical image analysis15. From early convolutional neural networks (CNNs) to more recent Transformer-based architectures, DL models have consistently achieved state-of-the-art performance in tasks, such as anatomical recognition, segmentation, and lesion detection16. Indeed, DL-based methods have been successfully applied to the segmentation of multiple oral and maxillofacial anatomical structures, including teeth17,18,19, the Maxilla and mandible19,20,21, mandibular nerve canal22,23, and the maxillary sinus1,24,25. Investigators have also begun applying DL to address the challenge of post-operative bone augmentation assessment following SA, giving rise to a mainstream “direct segmentation” paradigm. This paradigm aims to train neural networks to directly identify and delineate graft material contours on CBCT images. For example, Tao et al.26 employed a 3D Attention V-Net model to, for the first time, automatically, accurately, and efficiently segment graft material after SA, achieving a Dice coefficient of 90.36%, with segmentation speed far surpassing that of human experts—compellingly demonstrating the promise of DL in this application. However, their study explicitly excluded clinically common one-stage (simultaneous implant and graft) cases and required manual extraction of the region of interest (ROI)26, limiting clinical applicability. Xi et al.27 adopted an advanced Swin-UPerNet architecture to directly segment graft material in CBCT images containing metallic implant artifacts, likewise showing strong performance (Dice coefficient 84.9%). Notably, during healing and remodeling, graft material density, texture, and morphology undergo continuous transformation, resulting in dynamic temporal evolution of its radiographic characteristics (e.g., Hounsfield unit distribution)28,29. DL models are highly sensitive to such “domain shift” phenomena. A direct segmentation model trained on immediate post-operative CBCT data may experience pronounced performance degradation when confronted with grafts whose imaging features have substantially changed at 6 or 12 months. Moreover, the interface between graft material and native sinus floor or lateral wall often appears indistinct or exhibits gradual density transitions; after extended healing intervals, this boundary ambiguity further complicates accurate delineation—especially for models trained largely on grayscale intensity cues—leading to difficulty in segmenting subtle or poorly defined regions29. Consequently, for the clinically critical objective of reliable longitudinal monitoring, the direct segmentation approach possesses intrinsic fragility, and a novel, more robust technical paradigm is urgently needed.
To systematically address these multifaceted challenges, the present study proposes and validates a DL-based automated intelligent maxillary sinus and graft analysis system (SA-ai). The system innovatively integrates two synergistic neural network architectures: (1) a U-Net–like 2D network dedicated to precise segmentation of the maxillary sinus contour from CBCT images; and (2) a 3D V-Net designed for robust three-dimensional segmentation of the pre-operative maxilla and the post-operative region containing graft material. Leveraging a registration–subtraction strategy, the system provides a more comprehensive and stable solution. Through coordinated operation of these two models, the system efficiently and accurately segments pre- and post-operative CBCT scans, thereby enabling quantitative measurement of sinus volume, maxillary bone, and grafted augmentation. Critically, to enhance clinical relevance—especially for the common simultaneous implant (one-stage) scenario—a specialized post-processing workflow is integrated into the system to determine net bone gain, i.e., the integrated graft volume after excluding implant-occupied space. By delivering reliable quantitative data, this intelligent sinus segmentation and graft quantification system has the potential to optimize treatment planning and objectively assess augmentation outcomes.
Results
Baseline Performance of the SA-ai System
The proposed SA-ai system achieved outstanding segmentation performance across all evaluation metrics on the independent test dataset, reaching a level of clinically relevant accuracy suitable for automated volumetric analysis. Comprehensive assessment demonstrated consistently high performance on multiple complementary metrics, indicating robust and reliable segmentation capability. To justify selection of the 3D V-Net architecture, four DL architectures were compared on the same test set (Table 1, Fig. 1). According to Table 1, 3D V-Net outperformed all alternatives on every metric: its Dice coefficient (93.2 ± 1.34%) exceeded those of the MS-D network (92.6 ± 1.70%), 3D U-Net (85.4 ± 3.67%), and Res-UNet (69.3 ± 3.34%). Figure 1 illustrates maxillary segmentation (pre-operative CBCT) for one representative patient, where 3D V-Net achieved the highest anatomical fidelity. To strictly evaluate the robustness and generalization capability of the SA-ai system, a comprehensive multi-dimensional performance analysis was conducted (Fig. 2). The overall distribution of segmentation metrics (Fig. 2a–f) revealed a high degree of accuracy and consistency. The scatter overlays indicate that the vast majority of cases clustered tightly around the median, with minimal outliers. Crucially, given the challenge of “domain shift” caused by post-operative remodeling, we assessed the longitudinal stability across three time points (T0, T1, and T2). As shown in Fig. 2g–l and the radar chart (Fig. 2m), the model performance remained remarkably stable without significant degradation from pre-operative to post-operative stages. Additionally, the density distribution (Fig. 2n) and per-patient performance analysis (Fig. 2o) demonstrated that the system maintains robust performance across heterogeneous anatomical variations. The temporal consistency plot (Fig. 2p) tracks individual patient trajectories, confirming that segmentation quality did not fluctuate significantly for the same subject over time. Figure 2q summarizes the overall and time-specific segmentation effects of the maxilla by the SA-ai system, showing no statistically significant differences in Dice (p = 0.295), IoU (p = 0.525), or other metrics at different time points (p > 0.05).
Performance heat map of different architectures for maxilla segmentation.
a–f Box plots displaying the overall distribution of six key segmentation metrics (Dice, IoU, Precision, Sensitivity, HD95, and ASD) across the test dataset (n=51 scans). The box represents the interquartile range, the horizontal line indicates the median, whiskers extend to the data range, and scattered dots represent individual cases. g–l Performance metrics stratified by time point (T0, T1, T2), showing consistent accuracy across preoperative and postoperative stages. m Radar chart demonstrating the high overlap of performance metrics across T0, T1, and T2. n Probability density distribution of Dice coefficients fitted with a Kernel Density Estimation (KDE) curve. o Per-patient mean Dice scores sorted by rank. p Longitudinal consistency plot tracking Dice trajectories for individual patients across time points. The flat red line with error bars represents the mean trajectory ± standard deviation. q Statistical summary table comparing performance across time points. p > 0.05 indicate no significant performance differences in segmentation quality between time points.
The rigid registration algorithm exhibited robust alignment between pre- and post-operative CBCT scans, achieving the precision required for volumetric change analysis. Based on independent validation using 40 patients with manually annotated anatomical landmarks, the registration algorithm achieved an RMSE of 1.046 ± 0.389 mm, ranging from 0.600–2.343 mm. The MAE was 0.937 mm, reflecting consistent alignment quality across different anatomical regions. The multiresolution Mattes mutual information strategy proved highly effective in this clinical context, providing reliable foundation for subsequent volumetric analysis. Figure 3 displays the spatial distribution of 3D distance errors after registration between pre- and post-operative maxillae.
Color scale transitions from blue (0.0 mm error) to red (≥10.0 mm error); deep red corresponds to augmented bone within the sinus.
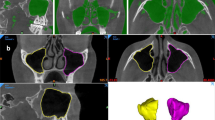
Figure 4 shows the modular workflow of the SA-ai system for automatically segmenting the maxilla before and after SA surgery and finally segmenting the bone material using the paired-subtraction method. Figure 5 illustrates detailed comparative analyses of manual annotations versus AI-driven segmentation in three representative cases: Case 1 (a, b) post-operative 9 months; Case 2 (c, d) immediate post-operative; Case 3 (e, f) post-operative 6 months with two implants placed simultaneously. Mean registration errors for the three cases were 0.662 mm, 0.740 mm, and 0.853 mm, all within a submillimeter range, confirming consistent performance across post-operative time points (immediate, 6 months, 9 months). Most regions display blue–green tones, indicating concentration of errors within 0.5~1.5 mm; higher deviations appear only in a few anatomically complex zones (white arrows), further supporting accuracy and stability.
a Comparative display of pre- and post-operative maxilla segmentations: red is pre-operative maxilla; green is post-operative maxilla (including implants). b Comparison of pre- and post-operative segmentations: upper row shows original (pre-registration) relative positioning; lower row shows precise rigid alignment. c Implant identification and graft volume correction: green (a) is automatically segmented maxillary sinus; yellow (b) is net graft augmentation region after exclusion of implant volume via Boolean operation; red arrows are portions of implants protruding into the sinus cavity.
a, c, e Multi-planar views (coronal, sagittal, and axial) showing the superposition of expert manual annotations (red contour) and SA-ai automated segmentation (green contour). White arrows indicate areas of minor discrepancy. b, d, f Corresponding 3D distance error heat maps quantifying the spatial deviation between the manual and automated surface meshes. The color scale transitions from blue (0.0 mm error) to red (>3.0 mm error).
Comparative Analysis Between SA-ai Automated and Manual Measurements
To validate the clinical accuracy of the SA-ai system, automated measurements from the test set (n = 17 patients) were compared against the averaged manual measurements derived from three independent, experienced implantologists (n = 3 evaluators). All eight key parameters demonstrated inter-observer Intraclass Correlation Coefficient (ICC) values > 0.900. Table 2 presents detailed comparative results between automated and manual measurements on the test cohort, all parameters demonstrated strong linear correlations (Pearson’s r ranged from 0.772 to 0.994) and achieved ‘good’ or ‘excellent’ reliability (all ICC ≥ 0.750). Bone Volume (BV) achieved the highest reliability (ICC = 0.986) and correlation (r = 0.994), directly validating the precision of the core 3D “registration–subtraction” framework. Implant protrusion length (IPL) showed excellent agreement (ICC = 0.983; r = 0.986). Other key parameters also showed high reliability, including sinus bone augmentation height (SBAH) and residual bone height (RBH). Collectively, these findings demonstrate that the SA-ai system delivers not only superior volumetric assessment but also precise measurement of key linear and anatomical indices.
Analysis of Systematic Measurement Bias
Despite a strong correlation, the automated and manual measurement values showed statistically significant differences in almost all parameters (all p < 0.05). The only exception was the lateral wall angle (A) (p = 0.096). The Cohen’s d values for all parameters ranged from 0.117 to 0.373, falling within the ‘negligible’ to ‘small’ range. Bland-Altman analysis (Fig. 6) was therefore employed to characterize the nature of these differences for all parameters. All parameters exhibited a consistent, systematic negative bias—SA-ai measurements were slightly lower than the averaged manual measurements. The magnitude of bias varied by parameter type. Implant-related 3D measurement (IPL) showed the smallest bias (−0.234 mm). Linear height metrics (RBH is −0.424 mm; SBAH is −0.332 mm) and volume (BV is −117.205 mm³) also showed relatively small deviations. Width (SW is −0.886 mm; SBAW is −0.581 mm; SBAL is −1.044 mm) and angular measurement (A is −2.746°) biases were comparatively more pronounced. Notably, over 95% of all data points for every parameter lay within the 95% Limits of Agreement (LoA), indicating minimal random error despite systematic bias and confirming highly stable, predictable behavior of the SA-ai system.
a–h correspond to RBH, SW, A, IPL, BV, SBAL, SBAW, and SBAH. Blue dashed line: mean bias; red dashed lines: 95% LoA.
To elucidate underlying mechanisms and interdependency patterns of discrepancies, a bias-based correlation framework was constructed (Fig. 7). Among 28 possible bias pairings, only one exhibited statistical significance (p < 0.05), indicating 96.4% independence among parameter biases. The significant positive correlation was between sagittal augmentation length bias (SBAL_bias) and coronal augmentation height bias (SBAH_bias) (r = 0.398, p = 0.017). A near-significant negative association was observed between sinus width bias (SW_bias) and lateral wall angle bias (A_bias) (r = −0.313, p = 0.092).
Bias correlation matrix of the SA-ai system.
Longitudinal Stability of Measurement Performance
To assess temporal stability, measurement performance at immediate post-operative (T1) and follow-up (T2) time points was compared (Table 3). BV reliability remained excellent and improved slightly at T2 (ICC = 0.994; 95% CI [0.985, 0.998]) compared to T1 (ICC = 0.991; 95% CI [0.979, 0.997]), with correlation coefficients stable at 0.994 at both time points. For the linear augmentation parameters (SBAL, SBAW, SBAH), all reliability scores (ICC) remained at ‘good’ or ‘excellent’ levels at both time points (ICC range: 0.759–0.914), and correlation coefficients for all correlation coefficients ranged from 0.767 to 0.937, confirming high longitudinal consistency.
Time Efficiency: SA-ai Automated Versus Manual Measurement
A quantitative comparison of time requirements for paired CBCT analysis per patient demonstrated substantial efficiency gains with the SA-ai system (Table 4). Completing a full evaluation (pre-operative plus two post-operative time points), including graft segmentation and all parameter measurements, required 74.82 ± 16.61 min using the manual workflow, versus only 3.40 ± 1.88 min with the fully automated pipeline—an over 20-fold improvement (p < 0.001). Each workflow stage showed similar acceleration: manual graft segmentation required 24.68 ± 8.05 min versus 0.73 ± 0.36 min for SA-ai; manual landmark localization and parameter measurement required 50.14 ± 25.69 min versus 2.67 ± 1.55 min for automation.
Discussion
Accurate quantification of post-SA grafted bone volume is central to evaluating surgical success and predicting the long-term stability of implants. However, this process has long faced a fundamental imaging challenge: on CBCT images, the boundary between graft material and native maxillary sinus bony walls is often indistinct due to gradual density transitions and partial volume effects. After several months of healing and remodeling, this blurring becomes even more pronounced. In this study, the successfully developed and validated SA-ai system achieved a maxilla segmentation Dice score of 93.2 ± 1.34% and clinically acceptable registration accuracy (1.046 ± 0.389 mm). In comparison with manual measurements, all eight key parameters showed ICC ≥ 0.75, with the core bone volume measurement reaching excellent agreement (ICC = 0.993), while analytical efficiency was improved more than 20-fold. Accordingly, by introducing an innovative “registration–subtraction” strategy together with multiple key technical components, this study provides a powerful solution to a long-standing clinical problem.
To address the technical challenges of segmenting graft materials in SA, this study proposes an innovative registration–subtraction strategy. The SA-ai system uniquely integrates two synergistic neural networks: a 2D U-Net for maxillary sinus contour segmentation and a 3D V-Net for pre- (T0) and post-operative (T1) maxillary bone segmentation. Mainstream “direct segmentation” approaches—such as recent work by Tao et al.26 and Xi et al.27 —essentially frame the task as a pattern recognition problem, i.e., training a DL model to delineate the graft material. However, the graft is a structure whose morphology and radiodensity evolve over time as it remodels and integrates with host bone28,29. These temporal changes challenge the robustness of pattern-based algorithms across different healing stages, limiting their suitability for reliable longitudinal monitoring. In contrast, the SA-ai system reframes the task from an unstable pattern recognition problem into a highly robust subtraction problem. Rather than attempting to identify the unstable graft, the system segments the maxillary bone—a structure with consistently clear and stable anatomical boundaries—at both T0 and T1. After high-precision 3D image registration aligns the pre- and post-operative maxillary models, the augmented bone region is precisely derived via a Boolean subtraction operation. This methodological shift transfers the segmentation target from a boundary-blurred, feature-variable “unknown” (the graft) to a well-bounded, morphologically stable “constant” (the maxilla), thereby circumventing the intrinsic bottlenecks of directly segmenting the graft material. This boundary ambiguity, stemming from biological integration and partial volume effects, is an inherent challenge that cannot be resolved by conventional imaging protocols, or by simple digital post-processing.
More importantly, this paradigm provides inherent advantages for longitudinal studies. Dynamic volumetric monitoring of graft material is clinically essential for understanding resorption and remodeling patterns. A model trained on immediate postoperative CBCT images may underperform at 6 or 12 months when graft radiographic characteristics have substantially evolved26. The SA-ai system is less vulnerable to such temporal degradation because global maxillary contours remain recognizable regardless of internal graft remodeling. Thus, registration–subtraction is not only a tactical solution to initial segmentation ambiguity but also a superior strategy for reliable long-term monitoring of graft volumetric stability, supporting comparative evaluation of different graft materials.
To ensure the system’s performance is both robust and capable of generalization, our methodology was designed to leverage the unique structure of the available cohort. While the total patient number was 85, the dataset possesses significant temporal depth. Each patient provided up to three longitudinal CBCT volumes (T0, T1, and T2), resulting in a training set that was rich with over 200 distinct anatomical scans. This paired data is crucial for validating the stability required for longitudinal monitoring. Instead of standard k-fold cross-validation, we utilized a stringent “patient-wise” split to create an independent test set (17 cases). In addition, a dataset of 120 healthy patients was additionally used for the U-Net model for 2D maxillary sinus segmentation. This guarantees data-split reproducibility and ensures that the evaluation assesses the SA-ai system’s ability to successfully generalize to entirely new patient anatomies.
Effective subtraction-based measurement in the proposed system rests on two pillars: (1) accurate segmentation of the foundational object (the maxilla) and (2) precise inter-temporal registration. Comparative evaluations against MS-D network, 3D U-Net, and Res-UNet architectures demonstrated that 3D V-Net achieved superior performance, validating the architectural choice. On an independent test dataset, maxillary segmentation attained a Dice coefficient of 93.2 ± 1.34%, surpassing the mean accuracy (DSC 90.71%) reported in a recent systematic review30, and matching state-of-the-art DL systems validated on large multicenter datasets (maxillary DSC 93.0% ~ 94.1%)19 and studies employing multi-scale dense architectures (jaw DSC 93.4%)31. Compared with earlier classical machine learning approaches—e.g., prior-guided random forest methods (maxillary DSC 91%)—the DL architecture used here shows clear advantages. Notably, performance may degrade on especially challenging datasets (DSC as low as 80%)32; however, the SA-ai system maintained a high 93.0% Dice score in the complex postoperative context containing graft materials, with no significant decrease compared to the pre-operative state, underscoring its robustness. Its performance approaches that of models focused on finer anatomical substructures (e.g., cancellous maxilla, DSC 95.57%)21 or optimized workflows where AI-assisted expert refinement yields DSC values of 96.0–99.3%33. Boundary localization was likewise strong, with an ASD of 0.5 ± 0.27 mm.
For image registration, the multi-resolution Mattes mutual information algorithm was validated through independent assessment using 40 patients with manually annotated anatomical landmarks, achieving an RMSE of 1.046 ± 0.389 mm. While the average precision slightly exceeded 1 mm, considering the complexity of maxillary anatomy and potential anatomical changes between pre- and post-operative scans, this precision level still meets clinical requirements. In oral implantology, 1–2 mm registration accuracy is generally considered safe for planning34, while sub-millimeter alignment meets navigation-grade requirements35,36. In a 3D craniofacial landmark study, Liberton et al. reported that reliable 3D measurements typically require RMSE within 1 – 2 mm, with manual landmarking of 61 points yielding a mean RMSE of 1.21 mm37. The independent landmark-based validation provides a more objective assessment of registration accuracy, avoiding potential bias from AI segmentation errors that could affect surface-based metrics. Given anatomical complexity and subtle post-surgical changes between scans, the SA-ai registration performance satisfies clinical expectations and supports accurate subsequent volumetric differencing—critical for longitudinal graft assessment. Compared with end-to-end “black box” models, the modular design (segmentation–registration–calculation) confers enhanced transparency and interpretability.
To validate clinical accuracy, SA-ai outputs were compared with manual measurements from three experienced implantologists. All eight key parameters showed strong linear correlations and achieved good or excellent reliability. The 95% confidence intervals for both correlation and ICC were narrow, demonstrating high concordance. The core BV metric achieved a near-perfect ICC of 0.993 (95% CI [0.987, 0.996]). Bland-Altman analysis revealed a consistent negative bias, i.e., SA-ai yielded slightly lower values versus manual averages. While superficially suggestive of underestimation, deeper examination indicates that AI measurements are more objective. Manual contouring in ambiguous boundary zones particularly under partial volume effects invites a “tendency to include,” leading to mild over-segmentation. In contrast, voxel-wise Boolean operations in SA-ai are deterministic and immune to such subjective inflation. Thus, the systematic bias likely reflects a stable distinction between machine objectivity and human interpretive subjectivity. Narrow 95% LoA further confirm minimal random error and highly stable measurement behavior.
A correlation network analysis was employed to deconstruct the internal structure of measurement bias. Of 28 possible inter-parameter bias pairings, only one exhibited statistical significance (p < 0.05), indicating that 96.4% of parameter biases were mutually independent. This independence confirms that errors do not propagate or amplify across dimensions, ensuring robustness in multi-parametric assessment. A significant positive correlation was noted between SBAL_bias and SBAH_bias (r = 0.398, p = 0.017), suggesting a potential shared geometric tendency: overestimation of SBAL with overestimation of SBAH. This observation provides a concrete direction for future geometric fidelity optimization.
Monitoring the longitudinal volume change of bone graft materials is a core part of the post-SA assessment, which is directly related to the long-term success rate of implants and the precise formulation of clinical decisions. Clinical studies have shown that there are significant differences in the absorption rates of different graft materials at 6 months and 12 months after surgery. The absorption rate of autogenous bone can reach 23.1%, while that of composite graft materials is 18.9%38. Moreover, some research has found that the volume absorption of bone graft materials within 12 months can be as high as 43.7 ± 19.0%39. Such dramatic time-dependent changes highlight the clinical urgency of establishing a reliable longitudinal monitoring system. However, existing manual measurement methods have poor consistency at different time points, and the intra-observer variability increases over time29. More crucially, the current mainstream “direct segmentation” AI methods face a serious “domain shift” problem. A model trained on immediate post-operative CBCT images may experience a significant decline in performance when dealing with grafts whose imaging features have changed significantly after 6 or 12 months of healing28. This makes it difficult for an AI system trained at a single time point to undertake the reliable longitudinal monitoring task. The results of the time-effect analysis in this study fully confirm this technical advantage. The ICC value of the volume measurement parameter at time point T2 is actually better than that at time point T1, and the correlation coefficient remains 0.994 at both time points. Research shows that significant absorption of bone graft materials mainly occurs in the first 6 months after surgery, and the absorption rate slows down significantly thereafter40. The SA-ai system can provide stable and reliable measurement data during this critical period, providing precise quantitative evidence for clinical decisions, such as whether additional bone augmentation is needed and when to perform the second-stage surgery. In addition, the temporal robustness of the SA-ai system is particularly suitable for identifying high-risk patients with abnormal bone resorption. The SA-ai system can detect cases deviating from the normal absorption pattern at an early stage, providing clinicians with an opportunity for timely intervention, thereby improving the long-term retention rate of implants and patient satisfaction.
This study indicates that the high correlation between the volume biases of the SA-ai system at T1 and T2 demonstrates a high degree of temporal consistency and predictability of systematic bias, providing a scientific basis for establishing time-point-specific correction coefficients. This predictable bias characteristic enables the SA-ai system not only to provide objective measurement results but also to establish standardized prediction models for the absorption patterns of different graft materials, thus enabling the formulation of personalized treatment plans based on evidence-based medical evidence. Although the ICC values for the linear bone augmentation parameters (SBAL, SBAW, SBAH) showed a slight decrease in reliability at time point T2, all remained at good or excellent levels. The correlation coefficients of all parameters across both time points remained high, confirming the stability of the SA-ai system in long-term longitudinal studies. A systematic review study pointed out that there are significant differences in volume stability among different graft materials (autogenous bone, allogeneic bone, xenogeneic bone, artificial bone)41. However, the high consistency of the SA-ai system did not show significant differences due to different bone graft materials. For example, in Case 3 shown in Fig. 4, freeze-dried bone allograft (FDBA), which has relatively good new bone formation ability, was used, and the average surface distance deviation was only 0.853 mm. However, due to the large differences in the sample sizes of patients with the three types of bone materials included in the test set, this study did not conduct an analysis on this.
Processing time per patient (paired one pre-operative and two post-operative CBCT scans) was quantitatively compared between the SA-ai system and traditional manual workflows. Manual processing required 74.82 ± 16.61 min on average, whereas the fully automated SA-ai pipeline required only 3.40 ± 1.88 min—an efficiency gain exceeding 20-fold (p < 0.001). Although this improvement is slightly less dramatic than isolated task accelerations reported in recent “direct segmentation” studies—e.g., Tao et al.26 reduced a single segmentation from 19.15 min to 7.2 s; Xi et al.27 reduced manual 1390 s (~23 min) to 19.28 s—the present system evaluates the entire clinical analytical continuum (registration, segmentation, parameter extraction). Crucially, SA-ai introduces unprecedented standardization and objectivity to SA evaluation. Manual measurement is time-consuming and susceptible to observer experience, fatigue, and variability, impairing reproducibility. Automation eliminates inter-observer variability, enforcing uniform measurement criteria.
This study successfully resolves the challenge of automated volumetric analysis in simultaneous implant placement (one-stage) cases. Prior automated studies excluded such cases because metallic implants produce severe streak and beam-hardening artifacts that disrupt segmentation, and implant volume cannot be easily separated from graft volume. This limitation restricted clinical applicability, given the prevalence of one-stage SA. Recently, Xi et al.27 addressed this frontier challenge using a Swin-UPerNet architecture for direct segmentation in the presence of metal artifacts, achieving an 84.9% Dice score, underscoring the difficulty. The SA-ai system employs high-density thresholding and connected component analysis to specifically identify and segment implants protruding into the sinus cavity. The implant volume is then subtracted from the preliminary total augmentation volume to yield a net bone gain. The automated measurement of IPL exhibited excellent agreement with manual values (ICC = 0.983; r = 0.986), directly validating the accuracy of the volumetric correction algorithm.
Despite strong performance, several limitations warrant future investigation. First, although this study utilized a paired 3D dataset consisting of 85 patients, this sample size is relatively small for deep-learning applications, and the dataset was sourced from a single center. Coupled with the limitation to the lateral window technique, this may restrict the generalizability of the research findings across different CBCT devices, scanning protocols, ethnic groups, or graft materials. Specifically, our model was trained on data from a single vendor (Planmeca ProMax) using fixed settings (Standard reconstruction kernel and specific exposure parameters). Variations across different manufacturers or changes in imaging protocols result in significant shifts in image grayscale distribution and noise characteristics. This Domain Shift phenomenon is known to severely degrade the performance of deep learning models, meaning the achieved Dice score of 93.2% cannot be directly guaranteed when inputting data from unfamiliar hardware or clinical protocols. Future validation must therefore prioritize multi-center, multi-vendor datasets to fully assess cross-protocol robustness. Second, although the system incorporates specialized algorithms for metal implants, streak and beam-hardening artifacts can still degrade surrounding maxillary bone image quality, indirectly impacting primary segmentation. Third, performance in cases with mucosal thickening, polyps, or cystic pathology remains insufficiently validated. While 3D V-Net demonstrated superiority here, broader multi-center datasets involving more complex pathological and anatomical variations are needed to evaluate cross-architecture adaptability and optimize overall system robustness.
This study successfully developed and validated a deep learning–based SA-ai system that, for the first time, enables fully automated, high-precision analysis of augmented bone volume after SA on CBCT imaging. By introducing an innovative registration–subtraction paradigm, the system circumvents the longstanding technical bottleneck of directly segmenting graft materials with blurred or evolving boundaries. Importantly, through dedicated algorithmic design, the system also overcomes the challenge of one-stage cases with simultaneous implant placement, substantially broadening its clinical applicability.
Meanwhile, the SA-ai system represents a significant step toward intelligent, precision-oriented peri-implant assessment. Its >20-fold efficiency gain and standardization of measurement workflows position it to reshape clinical practice, particularly by enabling integration into commercial CBCT workstations via plugins or APIs. This would make standardized longitudinal monitoring feasible in routine clinical practice, supporting optimized treatment and objective evaluation. Further clinical deployment as a Software as a Medical Device will depend on regulatory approval, which necessitates larger-scale, multi-center prospective clinical trials to rigorously validate its robustness and accuracy. Upon successful validation, SA-ai holds promise to become a standardized tool for postoperative evaluation of SA, contributing meaningfully to improved implant success and patient satisfaction.
Methods
Study Design and Ethics Approval
The design and implementation of the present study were conducted in accordance with the Declaration of Helsinki. The study protocol and the use of clinical case data have been approved by the Medical Ethics Committee of Zhejiang Provincial People’s Hospital (Approval No. QT2025124). No participant personal information irrelevant to this research will be disclosed throughout the study. Informed consent was waived by the Medical Ethics Committee of Zhejiang Provincial People’s Hospital due to the retrospective nature of the study.
Construction of the Dataset
To train the 2D U-Net–based maxillary sinus segmentation model, a dataset of 6000 two-dimensional images was compiled from 120 healthy patients (50 images per patient) without maxillary sinus lesions. These scans were acquired with a Planmeca ProMax (Planmeca Oy, Helsinki, Finland) and all images were reconstructed using the Standard reconstruction kernel. During the CBCT acquisition, the patient was required to maintain an upright position with their head stabilized. The Frankfort horizontal plane was kept parallel to the floor, and the subject was instructed to keep their head and tongue still throughout the scan. CBCT with a voxel size of 0.15 mm, 15-bit grayscale, and a 12 × 9 cm field of view (FOV). Each image contained the maxillary sinus region and was manually annotated by three experienced clinicians using LabelMe software (v5.5.0, Massachusetts Institute of Technology, United States) to create precise segmentation masks. Preprocessing included grayscale intensity adjustment and subsequent normalization to a target range of [0, 255] using the 5 and 99th percentile values calculated across all 6000 images as bounds. To increase data diversity and enhance model generalization, horizontal flipping was applied as a data augmentation strategy, effectively expanding the training sample space.
The 3D dataset consisted of paired CBCT scans from 85 patients who underwent SA at Zhejiang Provincial People’s Hospital. Each patient provided a set of temporally-matched CBCT images. For this longitudinal analysis, we defined three time points: T0 was the pre-operative baseline scan, T1 was the immediate post-operative scan (completed within 1 week after surgery), and T2 was the follow-up scan. Since this was a retrospective study, the time interval for the T2 scan was not fixed. The time range of the T2 scan was from 5 to 12 months after surgery, with an average follow-up time of 7.3 ± 2.1 months. All scans were acquired using the same CBCT device and reconstruction settings as described in 2D dataset. This cohort included both unilateral and bilateral SA procedures, for the purpose of analysis, each augmented maxillary sinus was treated as an independent sample. All cases were treated using the lateral window approach with particulate grafting materials, including deproteinized bovine bone mineral (DBBM), deproteinized porcine bone mineral (DPBM), and freeze-dried bone allograft (FDBA), combined with simultaneous implant placement following a one-stage protocol.
The inclusion criteria were: (1) The availability of a complete set of paired CBCT scan images, including one pre-operative (T0) and two post-operative (T1 and T2) scans, all of which have high diagnostic quality and completely cover the target anatomical area; (2) treatment with standard lateral window SA technique; (3) Adult patients (age ≥ 18 years) who were deemed systemically healthy and suitable for implant surgery. Cases were excluded if they presented with: non-diagnostic image quality (e.g., pronounced motion or severe metal artifacts); pre-existing sinus pathology (e.g., acute sinusitis, cysts); uncontrolled systemic diseases known to affect bone metabolism (e.g., uncontrolled diabetes, bisphosphonate therapy); or incomplete anatomical coverage in the CBCT’s field of view.
The dataset was randomly divided at the patient-wise into a training set (68 cases, 80%) and an independent test set (17 cases, 20%), ensuring that all data (including T0, T1, and T2 scans) from a single patient resided exclusively in one set. Baseline patient characteristics and CBCT acquisition parameters are summarized below (Table 5).
Three experienced implantologists and two medical image processing engineers collaboratively performed the 3D annotations, meticulously delineating the pre-operative maxilla and the post-operative region containing graft material. Preprocessing included clipping voxel intensities to the 0–2500 HU range, followed by normalization. To balance anatomical fidelity with training efficiency and inference speed, all CBCT volumes were resampled isotropically to 0.3 × 0.3 × 0.3 mm³.
Deep Learning Framework
To identify the optimal core architecture for this task, four network types were comparatively evaluated on the same dataset: 3D V-Net19, MS-D network42, Res-UNet43, and 3D U-Net (https://github.com/wolny/pytorch-3dunet). Based on comprehensive assessment of segmentation accuracy and boundary localization, 3D V-Net was selected as the primary 3D segmentation backbone. The automated segmentation system employs two distinct deep learning architectures for 2D and 3D tasks.
Maxillary sinus (2D) segmentation: A 2D U-Net–like encoder–decoder architecture with five convolutional levels in the encoder progressively extracts features from low-level texture to high-level semantics. The decoder restores spatial resolution via upsampling, producing a pixel-wise probability map of the sinus region. Skip connections fuse deep semantic context with shallow detail features, enabling precise boundary localization.
Maxilla and graft (3D) segmentation: A 3D V-Net architecture—an extension of U-Net to volumetric convolutions—directly processes 3D medical image volumes using 3D kernels. The encoder path consists of alternating stacked 3D convolutions and pooling operations for hierarchical feature abstraction while reducing spatial resolution. The decoder applies transposed convolutions to recover resolution and integrates skip connections from corresponding encoder stages. Residual modules at each convolutional stage mitigate vanishing gradients, and Parametric ReLU (PReLU) activations enhance nonlinear representational capacity. The network outputs a voxel-wise probability map indicating class membership. Figure 8 shows a flowchart of the study implementation.
Method framework diagram of SA-ai model training.
Training Protocol
For the 2D U-Net model, training was conducted using the RMSprop optimizer with the BCEWithLogitsLoss as the objective function. This loss function implicitly combines a Sigmoid activation with binary cross-entropy, making it well suited for pixel-wise binary classification tasks. (Loss function omitted; to be inserted).
In this expression, \(\sigma ({x}_{i})\) denotes the sigmoid function and \({y}_{i}\) the ground-truth label. The training configuration employed a batch size of 4, an initial learning rate of 0.001, and 500 epochs. To enhance robustness and generalization, on-the-fly data augmentation was applied, including Gaussian noise injection, image sharpening, contrast enhancement, and random rotation.
The 3D V-Net model was trained using stochastic gradient descent (SGD) with Dice Loss as the principal objective.
Here, \({p}_{i}\in [\mathrm{0,1}]\) represents the predicted probability, \({g}_{i}\in \mathrm{[0,1]}\) the ground-truth label, and ε = 1 × 10⁻⁵ a smoothing constant to avoid division by zero. Training parameters included a batch size of 1, an initial learning rate of 0.01, and 2000 epochs. PReLU activations were employed throughout the network. Dice loss is particularly advantageous in medical image segmentation because it directly optimizes spatial overlap between prediction and reference, thereby mitigating class imbalance—a frequent issue in medical imaging.
Image Registration Algorithm
To enable quantitative comparison between preoperative and postoperative scans, a comprehensive rigid registration pipeline was implemented using the Simple ITK library. The workflow systematically traversed the input directory to identify and pair pre-operative (fixed) and post-operative (moving) nii.gz images. As a preprocessing step, all images were cast to Float32 to ensure numerical precision and prevent overflow during optimization.
Initialization used Mattes mutual information as the similarity metric with 50 histogram bins. The choice of 50 bins balanced detail preservation with runtime performance; analytic gradients combined with the regular step optimizer promoted stable convergence. Mattes mutual information was selected for its robustness and suitability for multimodal alignment scenarios in medical imaging. Optimization employed a regular step gradient descent strategy with a learning rate of 1.0, a minimum step length of 1 × 10⁻⁴, a maximum of 400 iterations, and a relaxation factor of 0.5. The transformation model was initialized as a 3D rigid transformation (rotation and translation only):
In this formulation, \(R\in {SO}(3)\) is the rotation matrix and t ∈ R3 is the translation vector. The initial transform was centered using the geometric centers of the two volumes to accelerate convergence.
A three-level multiresolution scheme balanced accuracy and efficiency. The shrink factors of [4, 2, 1] represent the downsampling ratios applied at each resolution level, where the original images are reduced by factors of 4, 2, and 1, respectively, to create a pyramid of progressively higher resolution representations. Gaussian smoothing with standard deviations of [2, 1, 0] (in voxel units) was applied at corresponding levels to reduce noise and prevent aliasing artifacts. This coarse-to-fine refinement strategy reduced susceptibility to local minima while preserving final precision. The final transform was applied to the moving image using linear interpolation.
Mattes mutual information was chosen due to its proven performance in medical registration.
In this expression, \(p(k,l)\) denotes the joint histogram distribution, whereas \({p}_{f}(k)\) and \({p}_{m}(l)\) represent the marginal distributions of the fixed and moving images, respectively. B-spline–based histogram estimation was employed to reduce noise and discretization artifacts; sparse sampling reduced computational load; and analytic derivatives avoided numerical approximation, thereby improving optimization efficiency.
Volumetric Analysis and Clinical Measurements
The calculation of augmented bone volume is achieved through a fully automated, multi-stage Registration–Subtraction Pipeline designed to overcome volumetric ambiguity, particularly in one-stage implant cases. The process begins after segmentation yields binary masks for the pre-operative maxilla (\({M}_{T0}\)), the post-operative maxilla (\({M}_{T1/T2}\)), and the implanted volume (\({M}_{{Implant}}\)). Spatial alignment is then performed by rigidly registering the moving image and its associated masks (\({M}_{T1/T2}\) and \({M}_{{Implant}}\)) to the fixed T0 space using the Multi-resolution Mattes Mutual Information algorithm. Voxel alignment during this transformation employs linear interpolation (SimpleITK sitkLinear) to maintain sub-voxel precision and boundary mapping consistency.
The net graft volume (\({V}_{{graft}}\)) is derived through a multi-stage Boolean subtraction sequence. This sequence defines the gross augmentation as \({M}_{{Gross}}={M}_{T1/T2}^{{\prime} }\setminus {M}_{T0}\) (retaining only the newly present volume), which intrinsically handles partial overlap by operating on the difference between the registered models. For one-stage cases, the metallic implants are identified through voxel intensity thresholding at the 99th percentile of CT values and connected component analysis, with the extracted mask intersected with the maxillary sinus segmentation to isolate high-density structures within the sinus cavity. This allows for the critical Implant Masking step, where the implant volume is explicitly excluded via a final Boolean difference (\({M}_{{Graft}}={M}_{{Gross}}\backslash {M}_{{Implant}}^{{\prime} }\)), thereby yielding a true volumetric measurement of the augmented bone/graft material only. The final net graft volume is calculated as:
where Vtotal denotes the recorded volumetric difference, and \({V}_{{implant}(i)}\) the volume of the i-th implant. The resulting volumetric masks undergo post-processing optimization, including the removal of small, isolated regions via morphological filtering (min_voxels < 200) to ensure regional continuity and the application of Nearest Neighbor resampling to maintain the integrity of the binary label data.
Beyond volume, the system computes multiple clinically relevant parameters via coordinate-based analysis. By determining the 3D centroids of sinus-entering implants, standardized coronal slices are extracted. On these normalized 2D slices, automatically anchored anatomical landmarks (including the lowest point of the sinus floor) enable the extraction of 2D parameters, such as SBA (site-specific sinus bone augmentation height/width), RBH (residual bone height at the implant site), SW (sinus width), and A (lateral wall angle relative to a reference line). The 3D parameter IPL (implant protrusion length into the sinus) is directly derived from the volumetric segmentation.
System Evaluation
Model performance was comprehensively evaluated on an independent test dataset using standard segmentation and registration metrics. For segmentation, the Dice Similarity Coefficient (DSC) was used to quantify volumetric overlap for segmentation:
where \({TP}\) denotes true positives, \({FP}\) false positives, and \({FN}\) false negatives.
Intersection over Union (IoU) was defined as the ratio of intersection to union:
Volume was calculated as:
In this expression, \({N}_{{voxel}}\) is the number of voxels within the segmentation mask, and Δxi denotes voxel spacing along the x, y, and z axes (0.3 mm). Precision and sensitivity provided a balanced assessment of classification accuracy. The 95th percentile Hausdorff Distance (HD95) captured boundary outlier performance, while Average Surface Distance (ASD) quantified mean boundary localization error.
To provide objective registration accuracy evaluation independent of AI segmentation results, a manual landmark-based validation method was employed. Three experienced implantologists with over 5 years of SA experience independently annotated 10 stable anatomical landmarks on 40 paired CBCT images from the test dataset. These landmarks were selected according to specific criteria: (1) clearly identifiable in both pre- and post-operative images; (2) distributed across key anatomical regions of the maxilla; (3) spatially dispersed to comprehensively assess registration quality across different anatomical zones. The registration accuracy was evaluated by calculating the three-dimensional Euclidean distance between the corresponding landmark points before and after registration, including the root mean square error (RMSE), mean absolute error (MAE), as well as descriptive statistical indicators, such as the maximum distance, minimum distance, and standard deviation. All computations were executed on an NVIDIA RTX 4060 GPU with 8 GB memory to reflect practical performance under typical clinical hardware constraints. To justify architectural choices, four deep learning architectures (3D V-Net, MS-D network, Res-UNet, and 3D U-Net) were comparatively evaluated on the same test set.
Manual Measurement Validation
To validate the accuracy and clinical reliability of the SA-ai system, automated measurements were systematically compared with conventional manual measurements. A stratified analytical framework was adopted to reflect the distinct clinical implications of different parameters. Bone volume measurements (T1 and T2 BV) were analyzed at the sinus level (n = 19), as each unilateral maxillary sinus was treated as an anatomical unit representing the patient’s overall augmentation outcome. Other parameters were analyzed at the implant level (n = 30), reflecting site-specific anatomical conditions. This stratification aligned statistical methodology with clinical practice.
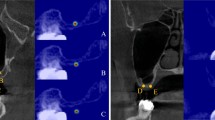
To ensure measurement quality, a detailed Standard Operating Procedure specified each parameter’s definition, anatomical landmark criteria, and procedural cautions, guaranteeing standardization and reproducibility. Manual assessments were independently performed in Mimics 21.0 (Materialize, Belgium) by three oral implantologists each with more than five years of experience in maxillary SA. The same CBCT datasets were imported into Mimics with uniform window and level settings to ensure consistent visualization. Key anatomical landmarks were manually identified on coronal, sagittal, and axial planes, relying on detailed anatomical knowledge and clinical experience. Measurement procedures (illustrated in Fig. 9) were as follows:
Manual measurement methods of various indicators in pre- and postoperative CBCT for SA.
Statistical Methods
The three observers performed measurements independently; measurement time for graft segmentation and other parameters was recorded. Reproducibility and reliability were subsequently evaluated, and averaged values were used to mitigate inter-observer variability.
Multiple statistical methods were applied to assess the agreement between the automated and manual methods. First, the Shapiro-Wilk test was used to evaluate the normality of paired differences. Subsequently, depending on the results of the normality test, a paired t-test (p > 0.05) or the non-parametric Wilcoxon signed-rank test (p ≤ 0.05) was employed to assess systematic differences. Pearson correlation analysis was used to quantify the strength of the linear association. Meanwhile, the effect size for all comparisons was calculated using Cohen’s d. The ICC (Two-way mixed effects, consistency) was used to assess reliability. Based on the guidelines established by Koo & Li44, ICC values were interpreted as follows: < 0.500 (Poor), 0.500–0.750 (Moderate), 0.750–0.900 (Good), and >0.900 (Excellent). Bland-Altman analysis was conducted to evaluate agreement and bias by visualizing the mean-difference scatter distribution. To further clarify the potential mechanisms of differences and patterns of interdependence, a correlation framework based on bias was constructed. The bias for each parameter was defined as (SA-ai measurement value-mean manual measurement value). The correlation patterns among parameter biases were analyzed to reveal potential systematic tendencies.
All statistical analyses were performed using SPSS 27.0 and Python 3.8 (including libraries, such as matplotlib, seaborn, pyvista, and scipy). A two-tailed p < 0.05 was considered statistically significant.
Data availability
The datasets generated and/or analyzed during the current study are not publicly available due to the use of clinical data from real patients but are available from the corresponding author on reasonable request. The underlying code for this study and training/validation datasets is not publicly available for proprietary reasons.
Code availability
The underlying code for this study and training/validation datasets is not publicly available for proprietary reasons.
References
Choi, H. et al. Deep learning-based fully automatic segmentation of the maxillary sinus on cone-beam computed tomographic images. Sci. Rep. 12, 14009 (2022).
Tomruk, C., Sençift, M. & Capar, G. Prevalence of sinus floor elevation procedures and survival rates of implants placed in the posterior maxilla. Biotechnol. Biotechnol. Equip. 30, 134–139 (2025).
Seong, W. J. et al. Prevalence of sinus augmentation associated with maxillary posterior implants. J. Oral. Implantol. 39, 680–688 (2013).
Jordi, C., Mukaddam, K., Lambrecht, J. T. & Kühl, S. Membrane perforation rate in lateral maxillary sinus floor augmentation using conventional rotating instruments and piezoelectric device-a meta-analysis. Int. J. Implant Dent. 4, 3 (2018).
Testori, T., Weinstein, T., Taschieri, S. & Wallace, S. S. Risk factors in lateral window sinus elevation surgery. Periodontol. 2000 81, 91–123 (2019).
Lyu, M., Xu, D., Zhang, X. & Yuan, Q. Maxillary sinus floor augmentation: a review of current evidence on anatomical factors and a decision tree. Int. J. Oral. Sci. 15, 41 (2023).
Jing, L. & Su, B. Resorption rates of bone graft materials after crestal maxillary sinus floor elevation and its influencing factors. J. Funct. Biomater. 15, 133 (2024).
Raghoebar, G. M., Onclin, P., Boven, G. C., Vissink, A. & Meijer, H. J. A. Long-term effectiveness of maxillary sinus floor augmentation: a systematic review and meta-analysis. J. Clin. Periodontol. 46, 307–318 (2019).
Simonsen, J. O. et al. Radiographic changes in the maxillary sinus following closed sinus augmentation. J. Periodontol. https://doi.org/10.1002/jper.11376 (2025).
Ohe, J. Y. et al. Volume stability of hydroxyapatite and β-tricalcium phosphate biphasic bone graft material in maxillary sinus floor elevation: a radiographic study using 3D cone beam computed tomography. Clin. Oral. Implants Res. 27, 348–353 (2016).
Okada, T., Kanai, T., Tachikawa, N., Munakata, M. & Kasugai, S. Long-term radiographic assessment of maxillary sinus floor augmentation using beta-tricalcium phosphate: analysis by cone-beam computed tomography. Int. J. Implant Dent. 2, 8 (2016).
Kwon, J. J. et al. Automatic three-dimensional analysis of bone volume and quality change after maxillary sinus augmentation. Clin. Implant Dent. Relat. Res. 21, 1148–1155 (2019).
Shujaat, S. et al. Emergence of artificial intelligence for automating cone-beam computed tomography-derived maxillary sinus imaging tasks. a systematic review. Clin. Implant Dent. Relat. Res. 26, 899–912 (2024).
Gardiyanoğlu, E., Ünsal, G., Akkaya, N., Aksoy, S. & Orhan, K. Automatic segmentation of teeth, crown-bridge restorations, dental implants, restorative fillings, dental caries, residual roots, and root canal fillings on orthopantomographs: convenience and pitfalls. Diagnostics 13, 1487 (2023).
Dayarathna, S. et al. Deep learning based synthesis of MRI, CT and PET: review and analysis. Med. Image Anal. 92, 103046 (2024).
Gao, Y. et al. Medical image segmentation: a comprehensive review of deep learning-based methods. Tomography 11, 52 (2025).
Xiang, B., Lu, J. & Yu, J. Evaluating tooth segmentation accuracy and time efficiency in CBCT images using artificial intelligence: a systematic review and meta-analysis. J. Dent. 146, 105064 (2024).
Jang, T. J., Kim, K. C., Cho, H. C. & Seo, J. K. A fully automated method for 3D individual tooth identification and segmentation in dental CBCT. IEEE Trans. Pattern Anal. Mach. Intell. 44, 6562–6568 (2022).
Cui, Z. et al. A fully automatic AI system for tooth and alveolar bone segmentation from cone-beam CT images. Nat. Commun. 13, 2096 (2022).
Fontenele, R. C. et al. Convolutional neural network-based automated maxillary alveolar bone segmentation on cone-beam computed tomography images. Clin. Oral. Implants Res. 34, 565–574 (2023).
Tian, Y. et al. Automatic jawbone structure segmentation on dental CBCT images via deep learning. Clin. Oral. Investig. 28, 663 (2024).
Ntovas, P., Marchand, L., Finkelman, M., Revilla-León, M. & Att, W. Accuracy of artificial intelligence-based segmentation of the mandibular canal in CBCT. Clin. Oral. Implants Res. 35, 1163–1171 (2024).
Jindanil, T., Marinho-Vieira, L. E., de-Azevedo-Vaz, S. L. & Jacobs, R. A unique artificial intelligence-based tool for automated CBCT segmentation of mandibular incisive canal. Dento Maxillofac. Radiol. 52, 20230321 (2023).
Altun, O. et al. Automatic maxillary sinus segmentation and pathology classification on cone-beam computed tomographic images using deep learning. BMC Oral. Health 24, 1208 (2024).
Morgan, N. et al. Convolutional neural network for automatic maxillary sinus segmentation on cone-beam computed tomographic images. Sci. Rep. 12, 7523 (2022).
Tao, B. et al. Deep learning-based automatic segmentation of bone graft material after maxillary sinus augmentation. Clin. Oral. Implants Res. 35, 964–972 (2024).
Xi, Y. et al. Automated segmentation of graft material in 1-stage sinus lift based on artificial intelligence: a retrospective study. Clin. Implant Dent. Relat. Res. 27, e13426 (2025).
Tabrizi, R., Sadeghi, H. M., Mohammadi, M., Barouj, M. D. & Kheyrkhahi, M. Evaluation of bone density in sinus elevation by using allograft and xenograft: a CBCT study. Int. J. Oral. Maxillofac. Implants 37, 114–119 (2022).
Al-Moraissi, E., Alhajj, W. A., Al-Qadhi, G. & Christidis, N. Bone graft osseous changes after maxillary sinus floor augmentation: a systematic review. J. Oral. Implantol. 48, 464–471 (2022).
Alahmari, M. et al. Accuracy of artificial intelligence-based segmentation in maxillofacial structures: a systematic review. BMC Oral. Health 25, 350 (2025).
Wang, H. et al. Multiclass CBCT image segmentation for orthodontics with deep learning. J. Dent. Res. 100, 943–949 (2021).
Chen, S. et al. Machine learning in orthodontics: introducing a 3D auto-segmentation and auto-landmark finder of CBCT images to assess maxillary constriction in unilateral impacted canine patients. Angle Orthod. 90, 77–84 (2020).
Nogueira-Reis, F. et al. Three-dimensional maxillary virtual patient creation by convolutional neural network-based segmentation on cone-beam computed tomography images. Clin. Oral. Investig. 27, 1133–1141 (2023).
Fokas, G., Vaughn, V. M., Scarfe, W. C. & Bornstein, M. M. Accuracy of linear measurements on CBCT images related to presurgical implant treatment planning: a systematic review. Clin. Oral. Implants Res. 29, 393–415 (2018).
Younis, H. et al. Accuracy of dynamic navigation compared to static surgical guides and the freehand approach in implant placement: a prospective clinical study. Head. Face Med. 20, 30 (2024).
Xu, Z. et al. Accuracy of dental implant placement using different dynamic navigation and robotic systems: an in vitro study. npj Digit. Med. 7, 182 (2024).
Liberton, D. K., Verma, P., Contratto, A. & Lee, J. S. Development and validation of novel three-dimensional craniofacial landmarks on cone-beam computed tomography scans. J. Craniofac. Surg. 30, e611–e615 (2019).
Peng, W. et al. Assessment of the autogenous bone graft for sinus elevation. J. Korean Assoc. Oral. Maxillofac. Surg. 39, 274–282 (2013).
Stricker, A. et al. Resorption of retromolar bone grafts after alveolar ridge augmentation—volumetric changes after 12 months assessed by CBCT analysis. Int. J. Implant Dent. 7, 7 (2021).
Lee, H. G. & Kim, Y. D. Volumetric stability of autogenous bone graft with mandibular body bone: cone-beam computed tomography and three-dimensional reconstruction analysis. J. Korean Assoc. Oral. Maxillofac. Surg. 41, 232–239 (2015).
Starch-Jensen, T., Deluiz, D., Vitenson, J., Bruun, N. H. & Tinoco, E. M. B. Maxillary sinus floor augmentation with autogenous bone graft compared with a composite grafting material or bone substitute alone: a systematic review and meta-analysis assessing volumetric stability of the grafting material. J. Oral. Maxillofac. Res. 12, e1 (2021).
Minnema, J. et al. Segmentation of dental cone-beam CT scans affected by metal artifacts using a mixed-scale dense convolutional neural network. Med. Phys. 46, 5027–5035 (2019).
Maji, D., Sigedar, P. & Singh, M. Attention res-UNet with guided decoder for semantic segmentation of brain tumors. Biomed. Signal Process. 71, 103077 (2022).
Koo, T. K. & Li, M. Y. A Guideline of Selecting and Reporting Intraclass Correlation Coefficients for Reliability Research. J. Chiropr. Med. 15, 155–163 (2016).
Acknowledgements
The authors gratefully acknowledge all experts and scholars who participated in this study for their valuable support and assistance in the successful completion of this research, and we would like to express our sincere gratitude to the Fund Committee of the Pioneer and Leading Goose Technology Project of Zhejiang Province. This work was supported by the Pioneer and Leading Goose Technology Project of Zhejiang Province [grant number 2024C03094].
Author information
Authors and Affiliations
Contributions
FY and XW were responsible for the design of the methodology, data analysis, implementation of manual measurements of clinical parameters for result verification, validation experiments, and the writing of the first draft. They are co-first authors and contributed equally to this study. YZ was responsible for the development and implementation of the deep-learning model and the writing of the first draft. XL was responsible for the collection, organization, and pre-processing of clinical data. YY and LM were responsible for the manual annotation and segmentation of CBCT images and the implementation of manual measurements of clinical parameters for result verification. YD and LW were responsible for the overall concept and design of the study, project supervision, acquisition of funding, and review and final revision of the manuscript. All authors participated in the critical review and revision of the manuscript, and read and approved the final submitted version.
Corresponding authors
Ethics declarations
Competing interests
All authors declare no financial or non-financial competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, F., Wu, X., Zhang, Y. et al. A deep learning based automated maxillary sinus segmentation and bone grafts analysis in CBCT images. npj Digit. Med. 9, 90 (2026). https://doi.org/10.1038/s41746-025-02275-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-02275-w