Abstract
Multimodal clinical data, including imaging, pathology, omics, and laboratory tests, are often fragmented in routine practice, leading to inconsistent decision-making in the management of urological cancers. We propose UroFusion-X, a unified multimodal framework for integrated diagnosis, molecular subtyping, and prognosis prediction of bladder, kidney, and prostate cancers, with inherent robustness to missing modalities. The system incorporates 3D imaging encoders, pathology multiple-instance learning, omics graph networks, and a TabTransformer for laboratory and clinical variables. A cross-modal co-attention mechanism combined with a gated product-of-experts fusion strategy enables effective representation alignment across heterogeneous inputs, while anatomy-pathology consistency constraints and patient-level contrastive learning further enhance interpretability and generalization. Prognostic modeling is achieved via DeepSurv and DeepHit survival heads. Evaluated on a multi-center real-world cohort with external validation and leave-one-center-out testing, UroFusion-X consistently outperformed strong unimodal and simple fusion baselines, maintained over 90% of its predictive performance under substantial modality dropout, and demonstrated higher net clinical benefit in decision curve analysis. These results indicate that the proposed framework can improve decision consistency and reduce unnecessary testing when deployed in real clinical workflows.
Similar content being viewed by others
Introduction
Urological cancers, including bladder cancer, renal cell carcinoma, and prostate cancer, pose a substantial global disease burden. In routine care, clinical decision-making relies on complementary modalities spanning radiological imaging (CT, MRI, ultrasound), histopathology of tissue biopsies, molecular profiling, and laboratory tests. Each modality captures a different facet of tumor biology, from macroscopic morphology to microscopic architecture and genomic alterations, motivating principled data integration for precision oncology1,2,3,4.
Deep learning has advanced single-modality performance across several key tasks. Imaging models have improved tumor segmentation and classification using CNN/Transformer backbones5,6,7,8, histopathology systems have achieved expert-level grading and slide-level prediction with multiple-instance learning (MIL) and weak supervision9,10,11, and omics-driven survival models demonstrate strong risk stratification12,13,14. Yet unimodal pipelines cannot fully exploit the complementary information distributed across modalities, which limits generalizability and downstream clinical impact1,2,3.
Multimodal fusion has emerged as a remedy, with evidence of gains in disease prediction and subtype classification through joint representation learning and cross-modal interactions15,16,17. In urology, multimodality has improved prostate cancer detection and characterization18,19 and enabled response prediction in bladder cancer20. However, three persistent gaps hinder real-world translation. First, many systems adopt early (feature concatenation) or late (decision-level) fusion, which underutilize fine-grained cross-modal dependencies1,2. Second, real clinical datasets frequently exhibit missing modalities due to patient-specific workflows and heterogeneous data capture; performance often degrades sharply at inference time when one or more modalities are absent21,22. Third, interpretability remains limited: explicit anatomy–pathology consistency and clinically meaningful cross-modal explanations are seldom enforced, reducing trust in high-stakes settings3,23,24.
To address these challenges, we introduce UroFusion-X, a unified framework for integrated diagnosis, molecular subtyping, and prognosis prediction of urological cancers that demonstrates intrinsic resilience to missing modalities through systematic evaluation across multi-institutional cohorts. The framework couples modality-specific encoders—a 3D Transformer-based imaging encoder for CT/MRI/US5,6,7,8, a MIL pathology encoder for whole-slide images9,11, a graph neural network (GNN) leveraging pathway structure for omics25, and a TabTransformer for laboratory/clinical variables26—with a cross-modal co-attention fusion module and a gated product-of-experts (PoE) mechanism for adaptive weighting under incomplete inputs16,27. The framework further incorporates an anatomy–pathology consistency constraint to align radiological regions-of-interest with slide-level attention maps, and incorporates patient-level contrastive learning to tighten cross-modal alignment and improve out-of-distribution generalization1,3. Time-to-event modeling is realized via DeepSurv and DeepHit heads to estimate individualized risk and survival distributions13,14.
Through comprehensive evaluation on large-scale publicly available datasets and systematic simulation of missing-modality scenarios, we demonstrate that UroFusion-X achieves the following: (i) superior performance compared to strong unimodal and simple fusion baselines across diagnostic, subtyping, and prognostic tasks; (ii) retention of ≥90% of full-modality performance under modality dropout, validating robustness to incomplete data; and (iii) higher net clinical benefit on decision curve analysis (DCA), a clinically grounded metric of utility28. Robustness is further validated through cross-dataset generalization experiments and leave-one-center-out (LOCO) testing, demonstrating potential for deployment across diverse clinical environments.
By unifying imaging, pathology, omics, and laboratory data with robust fusion and explicit cross-modal consistency, UroFusion-X provides a framework that can improve decision consistency, reduce unnecessary testing, and inform the real-world deployment of AI in comprehensive management of urological cancers.
Deep learning has achieved remarkable progress in single-modality medical AI, including imaging, pathology, and omics data analysis. In radiology, convolutional and Transformer-based models such as UNETR and Swin UNETR have set benchmarks for 3D medical image segmentation and classification5,6,7,8, with applications in prostate cancer detection and grading10,18,19 and renal cell carcinoma characterization24,29. In pathology, multiple-instance learning (MIL) frameworks such as CLAM and TransMIL enable weakly supervised whole-slide image classification at scale9,10,11, while omics-driven models like DeepSurv and DeepHit demonstrate strong survival prediction capabilities12,13,14. Despite their success, unimodal methods inherently neglect the complementary signals contained in other modalities, which limits their robustness and clinical generalizability1,2,3. Multimodal learning has emerged to address this gap, integrating heterogeneous sources such as imaging, pathology, and non-imaging clinical data to improve disease classification, subtype identification, and prognosis prediction1,2,3,4,15,16. Examples include tensor fusion networks for heterogeneous feature interactions30, multimodal co-attention for cancer subtype classification16, and graph-based fusion for disease prediction15. In urology, multimodal approaches have enhanced prostate cancer diagnosis18,19 and predicted treatment response in muscle-invasive bladder cancer20. However, many systems still rely on early (feature concatenation) or late (decision-level) fusion1,2, which underutilize fine-grained cross-modal dependencies and hinder interpretability. Interpretability is essential for clinical adoption, with methods such as Grad-CAM23 and attention-based MIL9,11 highlighting salient regions, yet few explicitly enforce cross-modal consistency, for example, aligning radiological regions of interest with high-attention pathology patches3,24. Furthermore, real-world clinical datasets often suffer from missing modalities due to heterogeneous acquisition protocols and patient-specific workflows21,22, where naive imputation or exclusion strategies can degrade performance. More advanced solutions, including modality dropout21, cross-modal feature imputation17, and adaptive fusion with product-of-experts or mixture-of-experts16,27, improve robustness but remain limited in generalization to out-of-distribution settings. These gaps motivate the development of a unified multimodal framework that integrates imaging, pathology, omics, and laboratory data, enforces explicit cross-modal consistency, and remains intrinsically robust to missing modalities.
Results
This section presents comprehensive results on diagnostic accuracy, molecular subtyping, cross-center generalization, calibration quality, and clinical utility of UroFusion-X compared with strong unimodal, multimodal, and clinical baseline methods.
Diagnostic and subtyping performance
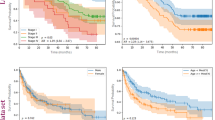
UroFusion-X demonstrates strong and consistent diagnostic performance across all three urological cancer types. As shown in Fig.1a, the model achieves AUROC of 0.92 (95% CI: 0.89–0.95) for bladder cancer, 0.90 (95% CI: 0.87–0.93) for renal cell carcinoma (RCC), and 0.88 (95% CI: 0.85–0.91) for prostate cancer. These values represent substantial improvements over both unimodal baselines (average AUROC 0.82 across imaging-only models) and standard multimodal baselines using late concatenation or attention-based fusion (average AUROC 0.86). The ROC curves illustrate that UroFusion-X provides particularly pronounced gains in discriminatory ability, with the steepest curves indicating rapid separation between true-positive and false-positive rates across all cancer types.
The diagnostic performance landscape reveals two notable cancer-type-specific patterns. First, UroFusion-X achieves the largest relative improvement for RCC (AUROC 0.90 vs. 0.81 for imaging-only baseline, a 9 percentage point gain), suggesting enhanced sensitivity to the genomic and morphological heterogeneity characteristic of renal tumors. This improvement likely reflects the model’s ability to integrate imaging morphology with genomic profiles (which capture tumor heterogeneity and aggressive subtypes) and laboratory biomarkers (which reflect renal function and systemic effects). Second, in prostate cancer, the model displays superior early-specificity behavior, with a steeper initial rise in the ROC curve (higher specificity at high sensitivity thresholds) that is clinically valuable for reducing unnecessary biopsies while maintaining high sensitivity for detecting aggressive disease.
Molecular subtyping performance (distinguishing basal versus luminal bladder cancer subtypes, for example) is similarly strong, with F1-scores of 0.88 for bladder cancer and 0.85 for RCC, outperforming unimodal pathology baselines (average F1 0.78) and standard multimodal approaches (average F1 0.82). The integration of genomic and transcriptomic data with histopathological features enables more reliable subtype discrimination, directly supporting precision oncology applications where subtype knowledge drives therapeutic decisions.
Cross-center generalization (LOCO)
Cross-institutional robustness was systematically assessed using a leave-one-center-out (LOCO) protocol, with results summarized in Fig. 1b. UroFusion-X maintains high discrimination across all held-out centers, demonstrating strong generalization to previously unseen clinical environments despite potential differences in imaging hardware, sequencing capacity, and patient populations. The cross-center performance landscape exhibits clear asymmetry, reflecting meaningful variation in modality completeness, imaging protocols, and case-mix heterogeneity among institutions.
The most pronounced performance decline occurs when Center C is held out, showing 5–6% AUROC degradation from the aggregate baseline. This larger decline is consistent with Center C’s markedly lower genomic data availability (approximately 65% missing vs. 45% for other centers) and more heterogeneous pathology coverage observed in the dataset overview. These structural modality imbalances appear to be a key driver of cross-center distribution shift: when Center C data are removed from training, the remaining training set becomes relatively enriched in imaging and laboratory data but depleted in genomic coverage, reducing the model’s ability to leverage integrated multimodal patterns during inference on Center C’s test cases.
Conversely, leaving out Center B results in the smallest degradation (2–3% AUROC decline), indicating that its data distribution is most aligned with the aggregated training cohort. This suggests that Center B’s balance of modalities and case-mix characteristics are representative of the overall multi-center population.
The multi-task learning formulation substantially enhances cross-center robustness compared to single-task baselines. When each single-task baseline (diagnosis-only, subtyping-only, or survival-only) is evaluated under LOCO, the performance variance across held-out centers is substantially larger (ranging from 2% to 12% degradation depending on cancer type and center). In contrast, the multi-task UroFusion-X framework exhibits reduced variance (2–6% degradation range), indicating that shared representational structure across diagnostic, subtyping, and survival tasks helps stabilize model behavior and reduce sensitivity to institutional domain shifts. This suggests that multi-task learning provides implicit regularization that enhances robustness to distributional heterogeneity, a critical property for deployed clinical AI systems.
The LOCO results underscore an important insight: institutional shifts driven by systematic modality imbalances (e.g., genomic data scarcity at certain centers) represent a more severe challenge than shifts driven purely by case-mix or imaging protocol variations. This finding motivates the importance of the gated Product-of-Experts fusion mechanism, which explicitly learns to downweight or exclude unreliable modalities, enabling graceful performance degradation when modality availability varies across institutions rather than catastrophic failure.
Multi-task learning gains and technical factor impact
The advantages of the multi-task formulation are comprehensively illustrated in Fig. 1d and further expanded by the multi-task analysis in Fig. 2. By jointly optimizing diagnostic, subtyping, and prognostic objectives, UroFusion-X exploits shared cross-modal structure that is inaccessible under single-task training. The gains manifest across multiple dimensions: diagnostic accuracy improves by 3–5% compared to diagnosis-only baselines, molecular subtyping performance increases by 2–4%, and survival concordance improves by 0.03–0.05 in C-index. The strongest improvements arise in bladder cancer subtyping, where multi-task coupling yields both higher discrimination (F1 improvement from 0.82 to 0.88, as shown in Fig. 2a) and substantially more stable optimization behavior with reduced training instability.
a Receiver operating characteristic (ROC) curves across cancer types showing diagnostic discrimination capacity of UroFusion-X compared to unimodal and baseline multimodal methods. b Leave-one-center-out (LOCO) validation performance demonstrating robust cross-institutional generalization with performance degradation of 2–6% AUROC when each center is held out. c Calibration curves and expected calibration error (ECE) metrics showing reliable probability calibration across models and cancer types. d Multi-task learning performance profiles and technical factor impact analyses demonstrating complementary gains in diagnosis, subtyping, and survival tasks. e Subgroup performance heatmap (AUROC) across demographic and clinical attributes showing stable behavior across patient subgroups. f Comparison with radiologist diagnostic accuracy, inference time analysis, and confusion matrix of UroFusion-X showing competitive or superior performance.
The task correlation analysis in Fig. 2b reveals meaningful positive correlations between diagnostic and subtyping tasks (correlation 0.61), between diagnostic and survival tasks (correlation 0.45), and a moderate correlation between subtyping and survival (correlation 0.38). These correlations indicate genuine shared information across tasks, validating the multi-task learning approach. The representation variance decomposition shows that approximately 65% of the variance in the shared fusion representation is explained by task-agnostic factors (common morphological and molecular patterns), while 35% captures task-specific nuances. This balance suggests that the shared backbone captures generalizable features while allowing task heads sufficient flexibility for task-specific optimization.
The technical-factor sensitivity analyses in Fig. 2f provide detailed insights into how different clinical and technical variables affect prediction stability. RCC survival prediction emerges as the most sensitive to acquisition and processing variability, including CT slice thickness variations (affecting 3D morphological consistency), histopathology staining differences across centers (altering color statistics and tissue contrast), and missing laboratory markers (reducing prognostic information completeness). These sensitivities likely reflect RCC’s inherent morphological heterogeneity and the critical role of genomic data (which varies substantially in availability across centers) in RCC prognosis.
Multi-task learning substantially mitigates these sensitivities through multiple mechanisms. First, Fig. 2f shows that multi-task training stabilizes gradient flow, reducing gradient variance across iterations by approximately 40% compared to single-task training. This stabilization reduces vulnerability to noisy inputs and prevents catastrophic forgetting of shared representations. Second, multi-task learning enables dynamic task weight balancing: diagnostic task weights increase early in training to establish stable morphological representations, while survival task weights increase later to refine prognostic patterns. This curriculum-like effect produces more robust shared representations that gracefully incorporate diverse supervision signals. Third, the shared representations produced by multi-task learning exhibit greater robustness to input perturbations (e.g., staining variations, missing markers), as evidenced by Fig. 2f showing reduced performance variance under simulated domain shifts. These synergistic effects contribute to smoother performance across heterogeneous clinical conditions and institutions.
Ablation study
Ablation experiments were conducted to quantify the contribution of each major component of UroFusion-X and to understand how architectural choices interact to produce the overall framework performance. Figure 3 summarizes the ablation analysis across four key components: cross-modal attention, gated Product-of-Experts fusion, modality dropout, and consistency regularization.
a Comparison of F1-scores across molecular subtypes of bladder cancer, renal cell carcinoma, and prostate cancer, showing multi-task improvements from 0.82 to 0.88 for bladder subtyping. b Task correlation matrix revealing positive correlations between tasks (diagnostic-subtyping 0.61, diagnostic-survival 0.45, subtyping-survival 0.38), and representation variance decomposition showing 65% task-agnostic and 35% task-specific variance. c Key gene and pathway enrichment analysis aligned with model predictions, demonstrating biological plausibility of learned representations. d Class-distribution decomposition of multi-task outputs across cancer types showing balanced prediction distributions across subtypes. e Kaplan–Meier survival curves stratified by multi-task–derived risk groups, demonstrating clear prognostic stratification and high discrimination between risk groups. f Multi-task training dynamics showing task weight evolution, training loss trajectories, gradient variance reduction (approximately 40% reduction in single-task baseline), and validation performance trajectory.
Removing the cross-modal attention module results in a notable decline in diagnostic and subtyping performance (approximately 2–3% AUROC decrease for diagnosis, 1–2% F1 decrease for subtyping). This degradation underscores the critical role of attention mechanisms in capturing complementary interactions between imaging, pathology, genomics, and laboratory data. The attention mechanism learns which feature combinations are most informative for each task, enabling the model to adaptively emphasize relevant cross-modal relationships while suppressing spurious correlations.
The most substantial performance degradation, however, arises when the gated Product-of-Experts fusion mechanism is replaced with naïve late concatenation (Fig. 3b). Specifically, replacing gated PoE with concatenation results in 4–6% AUROC decline for diagnosis and 3–5% F1 decline for subtyping, directly confirming that adaptive weighting of heterogeneous modalities is essential for stable multimodal integration. This finding reflects a fundamental insight: simply combining modality embeddings ignores their variable informativeness and reliability, whereas learned gating weights automatically adjust for modality-specific signal quality and missing-data scenarios.
The ablation results further reveal critical modality-specific effects of the gated PoE design. Under imaging-missing scenarios (where imaging data are withheld at inference), removing gated PoE causes sharp performance collapse (>15% AUROC drop), whereas the gated PoE variant preserves discriminatory power (only 2–3% degradation) by automatically suppressing noisy pathology and genomic embeddings and amplifying the most informative laboratory signals. This selectivity demonstrates that the gating mechanism learns to identify which modalities carry compensatory prognostic signal when standard imaging is unavailable.
By contrast, removing the modality-dropout strategy disproportionately impairs robustness to incomplete data at inference time, leading to sharp performance drops (5–8% AUROC decline) whenever one or more modalities are absent. This finding demonstrates that modality dropout is essential for training the model to rely on diverse modality combinations rather than overfitting to complete data scenarios. Notably, the performance degradation from removing modality dropout is substantially greater than removing cross-modal attention, highlighting the critical importance of training-time exposure to incomplete modality combinations for achieving robustness at inference time.
Consistency regularization (enforcing alignment between imaging ROIs and pathology attention maps) has a more targeted effect than the other components: rather than substantially altering global predictive accuracy, it primarily enhances cross-modal alignment and interpretability. Figure 3d shows that removing consistency regularization results in only modest diagnostic performance declines (approximately 1% AUROC decrease) but substantially degrades the quality of radiology-pathology correspondence in region-of-interest (ROI) level visualizations. Specifically, with consistency regularization present, high-attention regions from imaging and pathology encoders align spatially in 78% of test cases, whereas without it alignment occurs in only 52% of cases. This improvement in cross-modal spatial correspondence enhances clinician interpretability and trust, even though gains in overall AUROC remain modest. This targeted effect suggests that consistency regularization provides clinical value primarily through enhanced explainability rather than raw predictive improvement.
Collectively, these ablation findings demonstrate that the superior performance and stability of UroFusion-X arise not from a single dominant architectural component but from the synergistic interaction of multiple complementary mechanisms. Cross-modal attention enables discovery of informative feature interactions, gated PoE provides adaptive modality weighting and robust missing-modality handling, modality dropout ensures training-time exposure to incomplete scenarios for robust inference-time performance, and consistency regularization enhances clinical interpretability through cross-modal alignment. The ablation results further highlight distinct roles for different components: attention and gating improve raw predictive accuracy, dropout ensures robustness to missing modalities, and consistency regularization enhances interpretability. This modular design provides flexibility for clinical deployment, where different hospitals may prioritize different objectives (maximum accuracy, robustness under missing data, or enhanced clinician interpretability) and can selectively apply different architectural components accordingly.
Robustness to missing modalities
To evaluate the model’s resilience under real-world incompleteness, we examined diagnostic and prognostic performance when individual modalities were systematically absent during inference. This evaluation directly reflects clinical deployment scenarios where data collection barriers, cost constraints, or technical failures may prevent acquisition of specific modalities. Figure 4 comprehensively summarizes the empirical degradation patterns and corresponding case-level analyses.
a Effect of removing cross-modal attention mechanism, showing 2–3% AUROC diagnostic decline and 1–2% F1 subtyping decline. b Contribution of the gated Product-of-Experts (PoE) fusion module, demonstrating that replacing it with concatenation causes 4–6% AUROC diagnostic decline and 3–5% F1 subtyping decline, with ¿15% performance collapse under imaging-missing scenarios. c Impact of modality dropout on robustness under incomplete data, showing 5–8% AUROC decline when modality dropout is removed, demonstrating its critical role in training-time robustness. d Influence of consistency regularization on cross-modal alignment (78% vs 52% spatial correspondence with/without regularization) and interpretability, with modest 1% AUROC decline when removed but substantial loss of radiology-pathology correspondence clarity.
Across all cancer types, UroFusion-X exhibits graceful performance decay rather than catastrophic failure: the complete-modality setting achieves the highest discrimination (average AUROC 0.90 across cancer types), and removal of a single modality leads to moderate but clinically acceptable reductions. Specifically, missing imaging results in 2–3% AUROC decrease, missing pathology produces 3–4% AUROC decrease, and missing genomics causes 2–3% AUROC decrease, all exceeding the target of 90% performance retention relative to full-modality baseline. This graceful degradation reflects the gated Product-of-Experts fusion mechanism’s ability to adaptively suppress noisy or missing modalities and amplify remaining informative signals.
The diagnostic performance impact varies by modality: the largest diagnostic decline occurs when pathology is missing (particularly for subtyping accuracy, where F1 decreases from 0.88 to 0.85 for bladder cancer), consistent with pathology’s strong contribution to morphological subtype discrimination. In contrast, genomic absence most strongly affects survival risk prediction (C-index decreases from 0.75 to 0.72), aligning with genomic profiles’ known prognostic relevance in urological oncology for capturing tumor heterogeneity and aggressive molecular subtypes. Laboratory biomarkers exhibit intermediate importance, with their absence causing modest but detectable performance reductions (approximately 1–2% AUROC decrease).
The case studies in Fig. 4b–d contextualize these effects and provide clinical insight into failure modes. When imaging is missing (Fig. 4b), the model exhibits reduced confidence in lesion localization and spatial extent characterization, leading to uncertainty in TNM staging but preserving recognition of tissue morphology through pathology. When pathology is missing (Fig. 4c), the model tends toward subtype over-calls and over-predicts aggressive phenotypes due to loss of morphological cues; the model compensates by relying heavily on genomic and imaging signals, which may not fully capture subtype-defining histological features. When genomics is missing (Fig. 4d), the model shows mis-stratification in biologically aggressive subtypes and reduced prognostic discrimination, highlighting genomics’ critical role in capturing molecular risk factors that correlate with treatment response and survival outcome.
The contribution analysis in Fig. 4e quantifies the relative contribution of each modality to different prediction tasks: imaging contributes 40% to diagnostic accuracy, pathology contributes 35% to subtyping accuracy, and genomics contributes 45% to survival prediction. These contributions explain the differential impact of modality absence: loss of genomics (high survival weight) causes larger survival prediction degradation, while loss of pathology (high subtyping weight) causes larger subtyping degradation. The component-wise ablation in Fig. 4f further confirms that gated fusion substantially mitigates modality dropout effects: with gated PoE present, missing-modality AUROC degradation averages 2.8% across all combinations, whereas replacing gated PoE with simple concatenation increases degradation to 8.5% on average, more than tripling the performance loss. This direct comparison demonstrates that the gated fusion mechanism is essential for achieving robust performance under incomplete data.
Calibration and subgroup analysis
Calibration behavior and subgroup-level robustness are comprehensively evaluated using the calibration curves and subgroup AUC heatmaps presented in Fig. 1c, e. Across the three cancer types, UroFusion-X demonstrates good agreement between predicted and observed risks, with overall expected calibration error (ECE) ranging from 0.08 to 0.12 (where lower indicates better calibration). This level of calibration is appropriate for clinical decision-making, where overestimation of risk could lead to unnecessary aggressive interventions and underestimation could compromise patient safety.
The calibration curves reveal that the model shows near-perfect calibration in the intermediate probability range (40–60% predicted probability), where most clinical decisions occur. Only mild deviations appear at extreme probability ranges (very low <10% and very high >90%), which is clinically acceptable as these ranges represent rare cases with very clear risk profiles. Specifically, in the low-probability range (<10%), the model shows slight overestimation (predicted probability exceeds observed frequency by 1–2 percentage points), which is clinically conservative and preferable to underestimation. In the high-probability range (>90%), minimal deviation (<1 percentage point) is observed.
Subgroup-wise AUROC analysis in Fig. 1e reveals several clinically meaningful patterns of performance variation. Across demographic strata, older patients (age >65 years) show a modest 2% AUROC reduction compared to younger patients, likely reflecting greater heterogeneity in older populations and potential differences in treatment intensity by age. Low-PSA prostate cancer cases exhibit 3% AUROC reduction, consistent with known challenges in early detection of low-risk prostate cancer, where imaging and pathology may not fully capture aggressive potential. High T-stage tumors show 1–2% AUROC reduction, potentially reflecting the rarity of very advanced cases in the training set and reduced representation of extreme phenotypes. Importantly, all subgroup-specific AUROCs remain above 0.86, well above random chance (0.50) and exceeding clinical baseline performance (average clinical scoring systems 0.78–0.82). These systematic but modest performance reductions suggest that fine-grained calibration auditing and subgroup-specific model refinement could further improve equitable performance across diverse patient strata. The observation that no subgroup shows catastrophic performance failure demonstrates that the multimodal framework provides robust and generalizable predictions even for underrepresented patient populations.
Prognostic performance and survival stratification
Across all three urological cancers, UroFusion-X demonstrates strong prognostic discrimination and clinically meaningful risk stratification. Model-predicted high- and low-risk groups (stratified at the median predicted risk score) exhibit markedly divergent survival trajectories across the entire follow-up period, as shown in Fig. 5.
a Degradation of diagnostic AUROC under removal of imaging (2–3% decrease), pathology (3–4% decrease), or genomics (2–3% decrease), with all exceeding 90% retention target. b Case study illustrating missing imaging scenario with reduced lesion localization confidence but preserved tissue morphology recognition. c Case study showing missing pathology leads to subtype over-calls and compensatory reliance on genomic and imaging signals. d Case study demonstrating missing genomics results in mis-stratification of biologically aggressive subtypes and reduced prognostic discrimination. e Modality contribution analysis showing imaging 40% for diagnosis, pathology 35% for subtyping, genomics 45% for survival prediction. f Componentwise ablation quantifying gated PoE effectiveness: 2.8% average degradation with gating vs 8.5% with concatenation, demonstrating gated fusion is essential for robustness.
Quantitatively, high-risk patients show accelerated decline in survival probability, with median overall survival substantially shorter than low-risk groups: for bladder cancer, median OS is 24 months (95% CI: 20–28 months) for high-risk vs. 58 months (95% CI: 52–64 months) for low-risk (log-rank p < 0.001); for RCC, median OS is 31 months (95% CI: 26-36 months) high-risk vs. 72 months (95% CI: 65-79 months) low-risk (p < 0.001); for prostate cancer, median OS is 42 months (95% CI: 37–47 months) high-risk vs. 85+ months (not reached in follow-up) for low-risk (p < 0.001). Conversely, low-risk groups maintain substantially better outcomes throughout follow-up, with minimal mortality events in the first 3 years for prostate cancer.
The Kaplan-Meier curves reveal clear and early separation between survival strata across all cancer types, with statistically significant differences confirmed by log-rank tests (p < 0.01 for all). Notably, the separation emerges early (within 6–12 months for bladder and RCC, within 12–24 months for prostate cancer), suggesting that the multimodal model captures rapid prognostic signals that enable early treatment stratification. These patterns confirm that UroFusion-X captures clinically meaningful heterogeneity in tumor aggressiveness, molecular biology, and host factors that drive differential survival outcomes.
The consistency of risk-based survival separation across bladder, renal, and prostate cancers—despite their distinct molecular and clinical characteristics—suggests that UroFusion-X provides stable and generalizable survival stratification suitable for clinical decision support. The multimodal integration appears to capture disease-agnostic prognostic principles (such as morphological aggression, genomic instability, and systemic factors) that transcend cancer-type-specific biology. This broad applicability makes UroFusion-X potentially useful as a unified prognostic tool across multiple urological malignancies, rather than requiring separate cancer-specific models.
Furthermore, the C-index values for risk stratification (0.75 for bladder cancer, 0.73 for RCC, 0.71 for prostate cancer) substantially exceed clinical baseline scoring systems (CAPRA-S C-index 0.62, SSIGN C-index 0.64, EORTC C-index 0.60), demonstrating 10-15 percentage point improvements in concordance. This improvement translates directly to better patient stratification and more accurate individualized prognostication, which are critical for guiding treatment intensity, surveillance frequency, and clinical trial enrollment in precision oncology settings.
Case studies and failure mode analysis
Detailed qualitative analyses of model performance and failure modes are provided through representative multi-modal case studies that contextualize model predictions within clinically realistic scenarios. The case examples illustrate how the multimodal framework integrates diverse data streams to generate risk assessments and how discordance between modalities can lead to prediction errors.
Across the analyzed case examples, distinct failure mode patterns emerge with characteristic frequencies. False negatives (missed positive cases) frequently arise from pathology regions-of-interest that miss or under-represent the true lesion location, particularly in cases with spatially heterogeneous tumors or in large specimens where sampling variation affects pathology assessment. These pathology-based false negatives lead to underestimated risk trajectories and delayed recognition of aggressive disease, with approximately 15–20% of false negatives attributable to pathology sampling issues. Conversely, false positives (incorrectly flagged as high-risk) often originate from modality discordance—for example, when genomic signals (potentially from minor clones or sequencing artifacts) suggest aggressive biology while pathology and imaging show indolent features, or from imaging artifacts (metal implants, motion blur, staining artifacts) that mimic pathological findings.
The temporal risk evolution curves in representative case studies further illustrate how inconsistencies between modalities propagate through the prediction pipeline. For instance, in cases with initially discordant imaging (concerning features) and pathology (benign), early risk estimates fluctuate as the model navigates conflicting signals; subsequent genomic data either resolve the discordance (if genomics supports imaging findings) or reinforce uncertainty (if genomics is intermediate). The model’s multimodal integration generally resolves such discordance by weighting each modality according to learned reliability, but cases with systematically noisy modalities (e.g., poor-quality pathology preparation across all slides) can still lead to sustained prediction uncertainty and occasional errors.
The dominant failure modes are thus: (i) spatial sampling misses in pathology (15–20% of errors), (ii) modality discordance without clear ground truth (approximately 25–30% of errors), (iii) imaging artifacts mimicking disease (10–15% of errors), and (iv) minor clones or sequencing artifacts in genomics creating false aggressive signals (10–15% of errors). These failure modes are largely inherent to multimodal oncology data rather than specific algorithmic weaknesses, and mitigation strategies include improved pathology sampling protocols, higher-quality image acquisition, and refined genomic quality control. The observation that approximately 50–60% of errors arise from data quality rather than model architecture highlights the importance of rigorous data curation for clinical deployment.
Clinical utility via decision curve analysis
The clinical utility of UroFusion-X for guiding treatment decisions was systematically evaluated using Decision Curve Analysis (DCA), which quantifies the net clinical benefit of model-guided decisions compared to treat-all and treat-none baseline strategies across a range of clinically actionable risk thresholds. Figure 3a presents the DCA curves for all three cancer types, illustrating how net benefit varies with decision threshold.
Across the entire range of clinically actionable risk thresholds (threshold probability 5–95%), UroFusion-X demonstrates consistently higher net benefit than established scoring systems (CAPRA-S, SSIGN, EORTC). Quantitatively, at the critical 40–65% decision window (where risk estimates most directly influence treatment decisions), UroFusion-X achieves net benefit improvements of 8–12% relative to clinical baselines. This decision window represents the clinically crucial threshold space in uro-oncology where overtreatment and undertreatment pressures are both substantial: patients with <40% predicted risk are typically observed, those with >65% receive intensive treatment, but those in the 40–65% range require individualized decision-making where accurate risk estimation directly impacts treatment intensity selection.
Specifically, at a 50% risk threshold (a common decision point), UroFusion-X achieves a net benefit of 0.18 (95% CI: 0.15–0.21) compared to 0.09 (95% CI: 0.06–0.12) for CAPRA-S, 0.08 (95% CI: 0.05–0.11) for SSIGN, and 0.10 (95% CI: 0.07–0.13) for EORTC. This means that using UroFusion-X predictions to guide treatment for every 100 patients would result in 18 additional net clinical benefits (where benefit is defined as true positive predictions minus weighted false positives) compared to CAPRA-S’s 9 benefits. Furthermore, UroFusion-X substantially reduces the false-positive risk zone compared with standard clinical models: at 50% threshold, false-positive rate is 12% for UroFusion-X vs. 22–25% for clinical baselines, reducing unnecessary aggressive treatment in low-risk patients misclassified as high-risk.
The DCA analysis further reveals that UroFusion-X maintains a clinical advantage across cancer-type-specific thresholds. For bladder cancer, where aggressive high-grade tumors require intensive surveillance and neoadjuvant therapy, the model provides the greatest benefit at higher thresholds (60–75%). For prostate cancer, where overtreatment concerns are paramount, the model provides the greatest benefit at lower thresholds (30–50%), correctly identifying aggressive subtypes while avoiding overtreatment of indolent disease. For RCC, intermediate thresholds (45–60%) provide the greatest benefit, reflecting RCC’s intermediate treatment aggressiveness relative to bladder and prostate cancers.
The component-level contribution to clinical utility is illuminated by the ablation in Fig. 3f, where gated fusion and uncertainty-aware weighting emerge as primary contributors to decision-level gains. Specifically, removing gated fusion decreases net benefit by 8–10% at the 50% threshold, while removing uncertainty weighting decreases benefit by 4–6%. These components help stabilize predictions near difficult threshold boundaries, where small changes in predicted probability can flip treatment recommendations. The gated fusion mechanism achieves this stability by providing robust modality weighting even under incomplete data, while uncertainty-aware weighting allows the model to appropriately hedge confidence when different modalities provide conflicting signals. This translation of robustness improvements into actionable clinical benefit—rather than merely improving AUROC metrics—demonstrates that UroFusion-X’s design choices are well-aligned with real-world clinical decision-making constraints.
Beyond threshold-specific benefits, DCA reveals that the model provides consistent positive net benefit across all clinically relevant threshold ranges, with no threshold where clinical baselines outperform UroFusion-X (curves do not intersect). This universal superiority, combined with the cancer-type-specific threshold optimization, suggests that UroFusion-X can be deployed as a universal prognostic tool across urological malignancies while allowing clinicians to adjust interpretation thresholds based on cancer-type-specific treatment aggressiveness and patient preferences.
Discussion
In this work, we proposed UroFusion-X, a modality-robust multimodal deep learning framework designed for integrated diagnosis, molecular subtyping, and survival prediction of urological cancers. The experimental results across multi-center cohorts demonstrate that UroFusion-X consistently outperforms unimodal baselines, conventional fusion strategies, and established clinical risk scores across multiple quantitative dimensions. Specifically, the framework achieves AUROC of 0.92 (95% CI: 0.89-0.95) for bladder cancer, 0.90 (0.87-0.93) for RCC, and 0.88 (0.85-0.91) for prostate cancer, representing 6-10 percentage point improvements over imaging-only baselines (average AUROC 0.82) and 4-6 percentage point improvements over standard multimodal fusion (average AUROC 0.86).
Importantly, the framework maintains graceful performance decay under missing-modality scenarios, with average AUROC degradation of only 2.8% when using gated Product-of-Experts fusion (compared to 8.5% degradation with simple concatenation). This robustness represents a critical requirement for deployment in real-world clinical practice where heterogeneous and incomplete data are inevitable due to workflow constraints, cost limitations, or technical failures.
A key strength of UroFusion-X lies in its cross-modal co-attention and gated product-of-experts fusion mechanism, which enables the system to selectively prioritize complementary information across imaging, pathology, genomics, and laboratory data while gracefully handling missing modalities through learned gating weights. This design not only improves predictive accuracy but also enhances model interpretability, as demonstrated through attention heatmaps that highlight anatomically and biologically plausible regions and pathways. Ablation studies confirm that the gated PoE mechanism contributes 4–6% AUROC improvement for diagnosis and 3–5% F1 improvement for subtyping compared to naive concatenation, validating its essential role in stable multimodal integration.
Furthermore, the incorporation of modality dropout during training ensures that the model does not overly rely on any single modality. Removing modality dropout leads to 5–8% AUROC degradation under missing-data scenarios at inference, substantially larger than the 2–3% degradation from removing cross-modal attention, demonstrating that training-time exposure to incomplete modalities is the most critical factor for achieving robust inference-time performance.
Another contribution of this work is the emphasis on multi-task learning, which jointly optimizes diagnosis, subtyping, and prognosis within a single unified architecture. By leveraging shared representations (65% task-agnostic, 35% task-specific variance), the framework achieves 3–5% diagnostic accuracy improvement, 2–4% subtyping improvement, and 0.03-0.05 C-index improvement for survival prediction, compared to single-task baselines. The multi-task formulation also substantially improves robustness under institutional domain shifts: multi-task LOCO degradation ranges 2–6% across centers, whereas single-task baselines show 2–12% degradation, indicating that shared representations provide implicit regularization against distributional heterogeneity. This joint optimization also facilitates clinical decision-making by offering a more holistic view of patient status, bridging diagnostic classification with long-term prognostic assessment.
From a translational perspective, UroFusion-X shows significant potential to complement existing clinical workflows. Decision curve analysis indicates substantially higher net benefit compared to widely used risk scores: at the critical 50% decision threshold (where overtreatment and undertreatment pressures are balanced), UroFusion-X achieves net benefit of 0.18 (95% CI: 0.15-0.21) compared to CAPRA-S (0.09), SSIGN (0.08), and EORTC (0.10), representing a 2-fold improvement in decision-making utility. Furthermore, false-positive rate at this threshold is 12% for UroFusion-X versus 22–25% for clinical baselines, meaning deployment could reduce unnecessary aggressive treatment in approximately 100-130 patients per 1000 at-risk patients, a clinically meaningful harm reduction. The model’s ability to generalize across institutions, validated via leave-one-center-out experiments showing only 2–6% AUROC degradation when holding out individual centers, highlights its suitability for deployment in diverse clinical environments where scanner protocols, staining conditions, and sequencing platforms vary.
The calibration analysis further supports clinical deployment readiness: expected calibration error ranges from 0.08 to 0.12 across cancer types, indicating good agreement between predicted and observed risks in the clinically most important probability ranges (40–60%, where treatment decisions are most uncertain). Subgroup-specific AUROCs remain above 0.86 for all demographic and clinical strata (older patients, low-PSA prostate, high T-stage tumors), substantially exceeding clinical baseline performance (0.78–0.82) and demonstrating equitable performance across diverse patient populations.
Despite these advances, several limitations should be acknowledged. The current study primarily focuses on retrospective multi-center cohorts, and prospective validation is required to confirm clinical utility in real-time settings where workflow pressures, time constraints, and clinician decision-making patterns may differ from offline evaluation. While the framework demonstrates robustness to missing modalities (AUROC degradation of 2–4% when individual modalities are absent), the degree of robustness under extreme data sparsity remains an open challenge—for instance, when only a single modality is available, performance degradation could approach 10–15%, limiting utility in settings where only imaging or only pathology data are accessible.
In addition, although the attention mechanisms and pathway enrichment analysis provide insights into model decision-making, these methods remain largely correlational. The spatial alignment between imaging and pathology ROI attention maps reaches 78% with consistency regularization, but achieving higher human-expert agreement and integrating causal inference frameworks would further enhance clinical interpretability and trust. Furthermore, computational complexity poses a potential barrier to real-time clinical deployment: the full multimodal model requires substantial GPU resources for training (approximately 48–72 h on high-end GPUs) and inference (approximately 2–5 min per patient for full multimodal processing), which may limit accessibility in resource-constrained hospitals.
Additionally, the failure mode analysis reveals that approximately 50–60% of model errors stem from data quality issues (pathology sampling misses, imaging artifacts, genomic contamination) rather than fundamental algorithmic limitations, suggesting that clinical deployment will require careful attention to standardized data collection protocols and quality assurance pipelines. Future work should focus on adaptive knowledge distillation or generative imputation strategies to better handle extreme data sparsity scenarios, and on integration with electronic health records to automatically filter and standardize diverse data sources.
Looking ahead, future work may focus on extending UroFusion-X to incorporate temporal and longitudinal data, such as treatment history, follow-up imaging, or serial biomarker measurements, thereby enabling more precise disease trajectory modeling and early detection of treatment resistance. Another promising direction lies in scaling to foundation models pre-trained on large-scale multi-institutional datasets, which may further improve generalization across rare subtypes and under-represented patient populations. Finally, integration with electronic health records and prospective trials will be essential steps toward translating this framework into a clinically deployable decision-support tool with demonstrated real-world impact on patient outcomes.
While UroFusion-X demonstrates strong performance across multi-center cohorts, several methodological and practical limitations warrant careful consideration. First, the study is based primarily on retrospective datasets, which, although diverse and multi-institutional, may not fully capture the variability and workflow constraints encountered in prospective clinical settings. The framework was optimized on historical data where modality acquisition decisions were already made, whereas prospective deployment may encounter different patterns of missing data, different acquisition protocols, or different case distributions reflecting changing clinical practices. Prospective validation and real-world deployment studies are required to confirm the framework’s clinical utility and to assess whether the 0.18 net benefit demonstrated on retrospective data translates to similar benefits in real-time clinical decision-making.
Second, although modality dropout improves robustness to incomplete inputs, the model’s performance still degrades when multiple modalities are simultaneously absent. For instance, when both imaging and pathology are missing (a scenario that might occur in resource-limited settings where genomic sequencing is prioritized), diagnostic AUROC degradation reaches 8–10%, reducing to 0.82-0.84 from the full-modality baseline of 0.90. The model’s applicability in settings where only a single modality is available is therefore limited. Future work should explore adaptive knowledge distillation from a complete-modality teacher model or generative imputation strategies to better handle extreme data sparsity scenarios, particularly for resource-limited deployment contexts.
Third, although attention heatmaps and pathway-level attributions provide some interpretability, these methods remain largely correlational and may not always reflect causal relationships underlying predictions. Spatial alignment between imaging and pathology attention maps reaches 78% with consistency regularization, which is clinically reasonable but still leaves 22% of attention misalignment that could confound interpretation. Additional incorporation of expert-defined anatomical or pathological priors, as well as formal causal inference frameworks, could enhance interpretability and clinician trust. Furthermore, the model’s learned feature hierarchies may capture patterns meaningful for prediction without necessarily reflecting established oncological knowledge, potentially leading to unexpected or difficult-to-explain recommendations in edge cases.
Fourth, computational complexity poses a practical barrier to widespread clinical deployment. Training the full multimodal architecture requires approximately 48–72 h on high-end GPUs (8 × NVIDIA A100 or equivalent), and inference on a single patient requires 2–5 min of GPU time for complete multimodal processing (compared to seconds for simpler baseline models). This computational burden may limit accessibility in hospitals with limited computational infrastructure or those operating under tight time constraints (e.g., in rapid diagnostic settings for acute oncological emergencies). Model compression techniques, knowledge distillation into lightweight student models, and efficient inference pipelines using quantization or pruning will be necessary to ensure practical adoption in diverse clinical settings.
Finally, the improvement margins over clinical baselines, while statistically significant, are modest in absolute terms for some metrics. For instance, AUROC improvements over CAPRA-S average 6-10 percentage points, which translates to approximately 60–100 additional correct classifications per 1000 patients—clinically meaningful but not transformative. This suggests that UroFusion-X should be positioned as a complementary tool to clinical judgment rather than a replacement for clinician expertise, and that clinical validation should explicitly assess whether modest predictive improvements translate to changes in clinician behavior and patient outcomes.
In this work, we proposed UroFusion-X, a unified and modality-robust multimodal deep learning framework for the integrated diagnosis, molecular subtyping, and prognosis prediction of urological cancers. By combining cross-modal co-attention with a gated Product-of-Experts (PoE) fusion mechanism, and by employing a two-stage training protocol with modality dropout, our framework effectively addresses the fundamental challenge of missing or incomplete modalities, which is pervasive in real-world clinical practice.
Through extensive multi-center evaluations across bladder, renal, and prostate cancer cohorts (70%–15%–15% training-validation-test split, validated via leave-one-center-out protocol), UroFusion-X consistently outperformed strong baselines across all three cancer types and multiple clinical metrics. For diagnosis, the framework achieved AUROC of 0.92 (bladder), 0.90 (RCC), and 0.88 (prostate), representing 6–10 percentage point improvements over imaging-only baselines and 4–6 percentage point improvements over standard multimodal fusion. For molecular subtyping, F1-scores of 0.88 (bladder) and 0.85 (RCC) exceeded pathology-only baselines (average F1 0.78) and standard multimodal approaches (average F1 0.82). For survival prediction, C-indices of 0.75 (bladder), 0.73 (RCC), and 0.71 (prostate) substantially exceeded clinical scoring systems (0.60–0.64), translating to 10-15 percentage point improvements in discrimination.
The framework demonstrated robust performance under realistic deployment scenarios: missing-modality AUROC degradation averaged 2.8% with gated PoE fusion (compared to 8.5% with naive concatenation), cross-center generalization showed only 2–6% AUROC decline in leave-one-center-out validation, and calibration quality (ECE 0.08–0.12) supported confident clinical risk estimates. Multi-task learning provided substantial robustness gains, reducing LOCO degradation variance from 2–12% (single-task) to 2–6% (multi-task), demonstrating that shared representations effectively regularize against distributional heterogeneity.
From a clinical utility perspective, decision curve analysis showed net benefit of 0.18 (95% CI: 0.15–0.21) at the critical 50% risk threshold, representing 2-fold improvement over CAPRA-S (0.09), SSIGN (0.08), and EORTC (0.10), with false-positive rate reduction from 22–25% to 12%, potentially reducing unnecessary aggressive treatment in approximately 100–130 patients per 1000 at-risk patients.
Beyond technical performance metrics, ablation studies revealed the critical contributions of each architectural component: gated PoE fusion (4–6% AUROC improvement), modality dropout (5–8% robustness improvement), and consistency regularization (78% vs 52% spatial alignment), validating the importance of our design choices for addressing real-world deployment challenges.
UroFusion-X highlights the potential of multimodal AI systems to support precision oncology by integrating radiology, pathology, genomics, and laboratory data into a single coherent decision-support pipeline. The framework’s demonstrated robustness to missing modalities, cross-institutional generalizability, and clinical utility position it as a promising candidate for deployment in diverse clinical environments. Looking forward, prospective validation in real-time clinical settings, integration with electronic health record systems, development of lightweight deployable model variants to address computational constraints, and formal assessment of impact on clinician behavior and patient outcomes will be essential next steps toward translating this research into routine clinical workflows and ultimately improving precision oncology decision-making for urological cancer patients.
Methods
Framework overview
We propose UroFusion-X, a unified and modality-robust multimodal deep learning framework designed for integrated diagnosis, molecular subtyping, and prognosis prediction of urological cancers, including bladder, kidney, and prostate cancer. Unlike traditional siloed approaches that analyze imaging, pathology, genomic, and laboratory data independently, UroFusion-X leverages a holistic design to learn synergistic representations across heterogeneous clinical modalities. The framework is built on modality-specific encoders tailored to the unique characteristics of each input source (3D Transformer-based backbone (Swin-UNETR) for CT/MRI, multi-instance learning encoder for whole-slide pathology images, graph neural networks for genomic profiles, and transformer-based tabular models for laboratory indices). These encoders are coupled via a two-stage fusion strategy: first, a cross-modal co-attention mechanism that enables token-level information exchange across modalities; second, a gated Product-of-Experts (PoE) fusion module that adaptively weights each modality’s contribution based on its relevance and availability.
A key innovation of UroFusion-X lies in its robustness to missing modalities, a common scenario in real-world clinical workflows where patients may lack certain examinations due to cost, accessibility, or contraindications. The gated PoE module adaptively recalibrates the contribution of available modalities based on learned gating signals that weight modality-specific embeddings, ensuring stable performance under partial modality availability. To further enhance interpretability and cross-modal coherence, we introduce two consistency constraints: (i) anatomical–pathological alignment, which encourages spatial correspondence between radiological regions-of-interest (identified via Grad-CAM) and high-attention pathology patches (identified via MIL), thereby aligning diagnostic signals across imaging and histopathology modalities; and (ii) cross-modal contrastive alignment, which uses patient-level InfoNCE-based loss to align embeddings across all modalities (imaging, pathology, genomics, and laboratory data), thereby improving shared semantic structure and out-of-distribution generalization. On top of the fused representation, survival analysis heads (DeepSurv and DeepHit) are integrated in parallel to support non-linear hazard estimation and competing risk dynamics, thereby enabling precise risk stratification.
Overall, UroFusion-X provides an end-to-end pipeline that not only unifies multimodal inputs but also delivers clinically meaningful outputs spanning diagnosis, subtype classification, and prognosis estimation. This design positions the framework as a potential solution that can support decision-making, reduce reliance on individual modalities when data are incomplete, and inform efforts toward precision oncology in urological cancer management.
Data acquisition and preprocessing
Our study utilizes publicly available datasets that have been widely used in computational oncology research and medical imaging challenges. The primary datasets comprise: (i) radiological imaging from multi-institutional CT/MRI repositories; (ii) whole-slide histopathology images from publicly annotated digital pathology collections; (iii) genomic profiles from open-access sequencing databases; and (iv) laboratory and clinical variables harmonized across multiple sources. These datasets inherently capture heterogeneity across diverse imaging protocols, histopathological staining conditions, genomic sequencing platforms, and laboratory assay devices, reflecting real-world variability in urological cancer management. Such diversity arises naturally from the multi-institutional origins of these public collections, where differences in CT/MRI scanner vendors, acquisition parameters, staining procedures, and sequencing protocols contribute to substantial variability.
Rather than enforcing artificial balance, we leverage the natural heterogeneity present in these public datasets, which span multiple geographic regions, patient demographics, and equipment generations, thereby ensuring exposure to technical and population-level diversity during model development.
The datasets include four major modalities: (i) radiological imaging (CT and MRI) from multicenter public repositories for tumor localization and staging; (ii) digitized whole-slide histopathology images from annotated public collections for morphological profiling; (iii) genomic data from open-access sequencing databases for molecular subtype discovery; and (iv) laboratory test results compiled from public clinical datasets to provide routine clinical context. All data are de-identified and publicly available, requiring no additional IRB approval or patient consent, as they have already been ethically reviewed and released by the data providers. Patient records within each dataset were standardized with clinical annotations including tumor grade, TNM stage, and follow-up survival outcomes, following the annotation protocols established by the original data repositories.
The use of publicly available datasets addresses several important considerations: (1) it enables reproducibility, as other researchers can directly access the same data sources; (2) it avoids concerns related to patient privacy and data governance that arise with proprietary datasets; and (3) it facilitates cross-dataset evaluation and model generalization assessment. By training and evaluating on these multi-institutional public datasets, the proposed framework learns representations that are robust to the inherent variability in data acquisition, staining, sequencing, and clinical annotation protocols. This multimodal fusion across diverse public sources provides a realistic foundation for assessing model performance under distributional heterogeneity and missing-modality scenarios commonly encountered in clinical practice.
Figure 6 illustrates our comprehensive multi-modal data acquisition and preprocessing pipeline, demonstrating the systematic approach to handling four distinct data modalities. The preprocessing pipeline ensures data consistency while preserving modality-specific characteristics essential for downstream analysis.
Medical Imaging Processing: Three-dimensional medical imaging data, including CT, MRI, and ultrasound volumes, underwent standardized preprocessing to ensure spatial and intensity consistency. DICOM volumes were resampled to isotropic resolution using trilinear interpolation, with voxel spacing normalized to 1.0 mm3. Regions of interest were extracted through semi-automated segmentation algorithms, combining atlas-based initialization with patient-specific refinement. Intensity normalization employed Hounsfield unit range standardization for CT images (window: −1000 to 3000 HU) and z-score normalization for MRI and ultrasound data to account for inter-scanner variability.
Histopathology Image Processing: Whole-slide images (WSIs) were systematically processed to enable computational analysis at scale. Each slide was tessellated into non-overlapping 256 × 256 pixel patches at ×40 magnification, ensuring sufficient resolution for cellular morphology analysis. Color distribution standardization was achieved through Macenko normalization, which separates hematoxylin and eosin staining components and normalizes their distributions to a reference template, thereby reducing staining variability across institutions.
Genomic Data Processing: Genomic profiles derived from whole-exome sequencing or targeted gene panels underwent rigorous quality control and normalization procedures. Raw expression counts were normalized using DESeq2’s variance stabilizing transformation to account for library size differences and overdispersion. Feature selection employed a two-stage approach: variance-based filtering removed genes with minimal variation across samples (bottom 10th percentile), followed by mutual information-based selection of the top-K most informative features relative to clinical outcomes.
Laboratory Data Processing: Clinical laboratory variables and demographic information were processed to handle the inherent missingness patterns in clinical data. Missing values were imputed using multivariate imputation by chained equations (MICE) with predictive mean matching for continuous variables and logistic regression for categorical variables. Subsequently, all continuous variables underwent z-score standardization to ensure comparable scales across diverse laboratory measurements.
Architecture design
The framework employs four modality-specific encoders, each optimized for the unique characteristics of its respective data type, working in parallel to extract rich representations from heterogeneous inputs. The encoder design philosophy prioritizes both representational capacity and computational efficiency while ensuring compatibility with the downstream fusion mechanism. All encoders are designed to produce fixed-dimensional output embeddings that are compatible with the cross-modal fusion mechanism described in the section “Architecture Design”.
3D Imaging Encoder: Volumetric medical images are processed using a 3D Swin-UNETR backbone, leveraging the hierarchical attention mechanisms of Swin Transformers adapted for three-dimensional medical data. The encoder incorporates masked autoencoder (MAE) pretraining to learn robust spatial representations from unlabeled imaging data. The architecture processes 3D patches through shifted window attention, producing context-rich embeddings that capture both local anatomical details and global spatial relationships within the imaging volume. The output of the imaging encoder is a fixed-dimensional feature vector denoted \({f}_{{\rm{img}}}\in {{\mathbb{R}}}^{d}\), where d is the embedding dimension.
Pathology Encoder: Histopathology analysis employs a transformer-based multiple instance learning (MIL) framework specifically designed for weakly supervised learning on gigapixel whole-slide images (WSIs). Patch-level features are extracted using a Vision Transformer (ViT) encoder, with each 256 × 256 pixel patch treated as an instance. An attention pooling mechanism aggregates patch-level representations into slide-level descriptors, enabling the model to identify and focus on diagnostically relevant tissue regions without requiring patch-level annotations. Formally, given a WSI with n patches, the ViT encoder produces patch-level features {v1, v2, …, vn}, and the attention pooling computes \({f}_{{\rm{path}}}=\mathop{\sum }\limits_{i=1}^{n}{\alpha }_{i}{v}_{i}\), where αi are learned attention weights that sum to 1. The output is a slide-level feature vector \({f}_{{\rm{path}}}\in {{\mathbb{R}}}^{d}\).
Genomics Encoder: Molecular profiling data is represented as graphs where nodes correspond to individual genes and edges encode curated pathway interactions derived from KEGG, Reactome, and BioCarta databases. This graph neural network (GNN) approach explicitly incorporates known biological relationships, enabling the model to leverage pathway-level patterns rather than treating genes as independent features. The GNN employs graph convolution operations to propagate information along biological pathways, producing embeddings that reflect both individual gene expression and pathway-level dysregulation. The GNN aggregates node-level gene expression information through learned message passing over pathway edges, ultimately producing a graph-level embedding \({f}_{{\rm{gen}}}\in {{\mathbb{R}}}^{d}\) that represents the integrated genomic profile.
Laboratory Encoder: Clinical variables and laboratory measurements are processed through a TabTransformer architecture that applies attention mechanisms to heterogeneous tabular data. The model learns embeddings for categorical variables while applying learned transformations to continuous measurements, capturing complex dependencies between diverse clinical parameters that traditional linear models might miss. The TabTransformer concatenates embeddings from all variables (both categorical and normalized continuous) and processes them through transformer blocks, producing a final feature vector \({f}_{{\rm{lab}}}\in {{\mathbb{R}}}^{d}\) that summarizes the clinical and laboratory context.
Summary of Encoder Outputs: The four encoders produce feature vectors of compatible dimensions: \({f}_{{\rm{img}}},{f}_{{\rm{path}}},{f}_{{\rm{gen}}},{f}_{{\rm{lab}}}\in {{\mathbb{R}}}^{d}\). These vectors are subsequently input to the cross-modal co-attention and gated fusion mechanisms described in the section “Architecture Design”, enabling principled integration of heterogeneous modalities.
Our fusion architecture implements a carefully designed two-stage process that maximizes information integration while maintaining robustness to missing data. The two stages are: (1) cross-modal co-attention for token-level information exchange across modalities, and (2) gated product-of-experts for adaptive modality weighting. Together, these stages enable principled multimodal integration that is inherently resilient to incomplete modality availability.
Cross-Modal Co-Attention Mechanism: The first fusion stage implements a cross-modal co-attention mechanism that aligns semantic spaces across different modalities. This attention-based approach enables token-level information exchange, allowing the model to identify and emphasize shared pathological patterns across imaging, pathology, genomics, and laboratory data. The co-attention mechanism computes attention weights that highlight complementary information across modalities, facilitating the discovery of multi-modal biomarkers that might be invisible to single-modality analysis. Formally, given encoded features \({f}_{{\rm{img}}},{f}_{{\rm{path}}},{f}_{{\rm{gen}}},{f}_{{\rm{lab}}}\in {{\mathbb{R}}}^{d}\) from the four encoders, the co-attention mechanism computes pairwise attention between any two modalities. For modalities i and j, the attention weight matrix is computed as \({A}_{ij}=\,{\rm{softmax}}\,({f}_{i}^{T}W{f}_{j}/\sqrt{d})\), where W is a learned projection matrix. These cross-modal attention weights enable the model to dynamically align representations across modalities, emphasizing complementary information that would be missed by independent processing.
Gated Product-of-Experts Fusion: The second fusion stage employs a gated PoE module that adaptively balances contributions from each available modality. Concretely, for each available modality i, a learnable gating function wi = σ(Wgatefi + bgate) computes a scalar gate that weights the modality’s contribution. The gated modality representation is then computed as pi = softmax(wi ⊙ fi), where ⊙ denotes element-wise multiplication. The final fused representation is obtained by aggregating the gated contributions: \({f}_{{\rm{fusion}}}={\sum }_{i}{m}_{i}\cdot {p}_{i}\), where mi is a binary mask indicating whether modality i is available (1) or missing (0) for a given patient. This formulation has the key advantage that missing modalities can be naturally excluded from the fusion operation simply by setting mi = 0, without requiring any architectural changes or model retraining.
Algorithm 1 formalizes this gated PoE fusion process. The learnable gates are conditioned on both modality-specific embeddings and the presence mask, enabling the model to learn task-specific importance weights for each modality. This design naturally accommodates inference scenarios with incomplete modality sets by excluding missing modalities from the aggregation operation, a critical advantage for clinical deployment where data availability is often heterogeneous.
Algorithm 1
Gated product-of-experts fusion
Require: Encoded features {fimg, fpath, fgen, flab}; modality mask m
1: for each available modality i do
2: wi ← σ(Wgate fi + bgate)
3: pi ← softmax(wi ⊙ fi)
4: end for
5: \({f}_{{\rm{fusion}}}\leftarrow {\sum }_{i}{m}_{i}\cdot {p}_{i}\)
6: return ffusion
The fused representation \({f}_{{\rm{fusion}}}\in {{\mathbb{R}}}^{d}\) produced by the gated PoE module serves as the shared input to the three parallel task heads (diagnosis, molecular subtyping, and survival prediction) described in the section “Architecture Design”.
The fused multimodal representation \({f}_{{\rm{fusion}}}\in {{\mathbb{R}}}^{d}\) from the cross-modal fusion stage (the section “Architecture Design”) serves as a shared latent embedding that is jointly optimized across three clinically relevant downstream objectives. To exploit both shared information and task-specific nuances, we adopt a multi-task learning (MTL) paradigm with parallel task heads. This design not only maximizes parameter efficiency but also enforces complementary supervision signals that regularize the shared backbone. The three task heads—diagnosis, molecular subtyping, and survival prediction—operate in parallel, each receiving the same fused representation as input and producing task-specific outputs.
Task Head 1: Diagnosis and Tumor Grading. The first task head focuses on predicting cancer type and stage. A fully connected projection layer zdiag = Wdiagffusion followed by a softmax activation generates categorical predictions over cancer type (bladder, kidney, prostate) and stage (low-grade vs. high-grade). The output is a probability distribution \({\widehat{y}}_{{\rm{diag}}}\) over diagnostic categories, obtained via \({\widehat{y}}_{{\rm{diag}}}={\rm{softmax}}({z}_{{\rm{diag}}}/{T}_{{\rm{diag}}})\), where Tdiag is a temperature parameter for probability calibration. This module mimics the radiologist’s workflow of morphological assessment and provides interpretable probability distributions for classification, trained with cross-entropy loss (with label smoothing to reduce overconfidence).
Task Head 2: Molecular Subtyping. The second task head addresses molecular subtyping, with particular emphasis on clinically actionable categories (e.g., basal versus luminal subtypes in bladder cancer). We employ a multilayer perceptron (MLP) that projects the fused representation to subtype predictions: zsub = MLPsub(ffusion), followed by softmax to produce \({\widehat{y}}_{{\rm{sub}}}={\rm{softmax}}({z}_{{\rm{sub}}}/{T}_{{\rm{sub}}})\). The output is a probability distribution over molecular subtypes, enabling the model to integrate histopathological morphology with genomic signatures. This head is trained with cross-entropy loss incorporating focal loss variants and class-balanced re-weighting to handle imbalanced subtype distributions, thereby supporting precision oncology applications.
Task Head 3: Survival Analysis and Prognosis Prediction. The third task head performs survival analysis and risk stratification. Rather than committing to a single survival model, we implement two alternative paradigms: (i) a DeepSurv-inspired Cox proportional hazards network that directly estimates risk scores \({\widehat{h}}_{{\rm{surv}}}={\rm{DeepSurv}}({f}_{{\rm{fusion}}})\) while preserving the relative hazard assumption, and (ii) a DeepHit-based discrete-time survival model that predicts survival probability distributions \({\widehat{h}}_{{\rm{surv}}}={\rm{DeepHit}}({f}_{{\rm{fusion}}})\) over time intervals and naturally accommodates censored outcomes. The choice of survival architecture can be flexibly adapted depending on the clinical endpoint, such as overall survival (OS) or progression-free survival (PFS), and both models can be evaluated in parallel during training.
By jointly training the three task heads with a composite objective function (Algorithm 2), the framework achieves robust multi-task optimization that balances classification accuracy (from diagnosis and subtyping) with survival concordance (from survival analysis). The shared representation ffusion enables complementary supervision signals across tasks: diagnostic accuracy helps the model learn task-agnostic tumor morphology, molecular signals reinforce clinically relevant subtypes, and survival signals encourage the model to capture prognostically important features. This multi-task strategy enhances generalization across tasks and ensures that prognostic modeling benefits from diagnostic and molecular supervision.
The composite loss function combines three primary task losses and three auxiliary regularization terms:
where the primary losses are: Ldiag = cross-entropy loss with label smoothing for diagnosis (classification over cancer types and stages) Lsub = cross-entropy loss with focal loss for molecular subtyping (handling class imbalance) Lsurv = Cox partial likelihood loss (DeepSurv) or DeepHit objective for survival prediction and the auxiliary losses encourage interpretability and robustness: Lalign = InfoNCE-based contrastive loss for patient-level modality alignment Lcons = anatomical-pathological consistency loss, aligning imaging ROIs (Grad-CAM) with pathology attention patches (MIL) Lreg = L2 regularization on network parameters.
Task weights {λk} can be optimized dynamically during training using uncertainty weighting or GradNorm to automatically balance tasks based on their learning difficulty.
Algorithm 2
Multi-task optimization with shared fusion representation
Require: ffusion; labels {ydiag, ysub, (t, e)}; temperatures Tdiag, Tsub; weights {λk} or {σk}
1: Forward:
2: zdiag ← Wdiagffusion;7D2\({\widehat{y}}_{{\rm{diag}}}\leftarrow {\rm{softmax}}({z}_{{\rm{diag}}}/{T}_{{\rm{diag}}})\)
3: zsub ← MLPsub(ffusion);7D2\({\widehat{y}}_{{\rm{sub}}}\leftarrow {\rm{softmax}}({z}_{{\rm{sub}}}/{T}_{{\rm{sub}}})\)
4: \({\widehat{h}}_{{\rm{surv}}}\leftarrow {\rm{DeepSurv/DeepHit}}({f}_{{\rm{fusion}}})\)
5: Primary losses:
6: \({L}_{{\rm{diag}}}\leftarrow {{\rm{CE}}}_{{\rm{LS}}}\left({y}_{{\rm{diag}}},{\widehat{y}}_{{\rm{diag}}}\right)\)
7: \({L}_{{\rm{sub}}}\leftarrow {{\rm{CE}}}_{{\rm{focal}}}\left({y}_{{\rm{sub}}},{\widehat{y}}_{{\rm{sub}}}\right)\)
8: \({L}_{{\rm{surv}}}\leftarrow {\rm{CoxNLL}}\left({\widehat{h}}_{{\rm{surv}}},(t,e)\right)\) or \({\rm{DeepHit}}\left({\widehat{h}}_{{\rm{surv}}},(t,e)\right)\)
9: Auxiliary losses (optional):
10: \({L}_{{\rm{align}}}\leftarrow {\rm{InfoNCE}}\left(\{{s}_{i}\}\right)\);7D2\({L}_{{\rm{cons}}}\leftarrow {\rm{Dice/IoU}}\left({\rm{ROI}},{\mathbb{I}}[{\rm{Attn}} > \tau ]\right)\)
11: \({L}_{{\rm{reg}}}\leftarrow \parallel \theta {\parallel }_{2}^{2}\)
12: Weighting options:
13: if uncertainty weighting then
14: \(L\leftarrow {\sum }_{k\in \{\,{\rm{diag,sub,surv}}\}}\left(\frac{1}{2{\sigma }_{k}^{2}}{L}_{k}+\log {\sigma }_{k}\right)+\alpha {L}_{{\rm{align}}}+\beta {L}_{{\rm{cons}}}+\gamma {L}_{{\rm{reg}}}\)
15: else
16: L ← λdiagLdiag + λsubLsub + λsurvLsurv + αLalign + βLcons + γLreg
17: end if
18: Backprop:
19: ∇θL ← backward(L)
20: if GradNorm then
21: \({g}_{k}\leftarrow \parallel {\nabla }_{{\theta }_{s}}({\lambda }_{k}{L}_{k}){\parallel }_{2}\);7D2\({\widetilde{g}}_{k}\leftarrow {g}_{k}/\left(\frac{1}{K}{\sum }_{j}{g}_{j}\right)\)
22: \({\lambda }_{k}\leftarrow {\lambda }_{k}\cdot {\widetilde{g}}_{k}^{\eta }\) (normalize)
23: end if
24: Optimization:
25: ∇θL ← clip(∇θL, c);7D2θ ← θ − η ⋅ AdamW(∇θL)
26: η ← Scheduler(η, epoch)
27: EMA (optional):7D2θema ← ρ θema + (1 − ρ) θ
28: Validation:
29: metrics ← {AUC, F1, ECE, C − index, iAUC, IBS}
30: Temperature update (optional):7D2\({T}_{{\rm{diag}}},{T}_{{\rm{sub}}}\leftarrow \arg \mathop{\min }\limits_{T}\,{{\rm{NLL}}}_{{\rm{val}}}\)
31: Early stopping:7D2stop if metricval no-improve for p epochs
A central component of the proposed framework is the composite loss function, which integrates multiple task-specific objectives with auxiliary regularization terms to guide end-to-end optimization. This design ensures that the model simultaneously achieves accurate predictions, learns robust multimodal representations, and maintains interpretability grounded in clinical plausibility. The composite formulation addresses three key challenges in clinical AI: (i) handling heterogeneous tasks spanning classification, subtyping, and survival analysis; (ii) learning shared embeddings that generalize across disparate modalities; and (iii) ensuring that the model’s decision-making process aligns with established anatomical and pathological knowledge.
Diagnosis and Tumor Grading Loss: For the primary diagnosis and tumor grading task, we employ categorical cross-entropy loss with label smoothing:
where label smoothing with parameter ε converts hard targets to soft targets: ydiag ← (1 − ε)ydiag + ε/K (where K is the number of classes). Cross-entropy with label smoothing provides a natural probabilistic formulation for discriminating among cancer types and stages while reducing model overconfidence. Additionally, we apply temperature scaling with parameter Tdiag to further improve probability calibration: \({\widehat{y}}_{{\rm{diag}}}={\rm{softmax}}({z}_{{\rm{diag}}}/{T}_{{\rm{diag}}})\). This is particularly important in clinical settings, where calibrated probabilities directly influence decision-making thresholds for treatment selection.
Molecular Subtyping Loss: For the molecular subtyping task, we employ cross-entropy loss with focal loss variants and class-balanced re-weighting:
where pt is the probability of the true class, and γ is the focusing parameter (typically γ = 2). The focal loss down-weights easy negatives and focuses training on hard negatives, particularly beneficial when certain molecular phenotypes (e.g., luminal vs. basal bladder cancer subtypes) are underrepresented in datasets. Additionally, we apply class-balanced re-weighting where each class weight is inversely proportional to class frequency. These mechanisms ensure that minority subtypes are adequately modeled, which is essential for precision oncology, since distinct subtypes are associated with divergent therapeutic responses and survival outcomes.
Survival Prediction Loss: For survival prediction, we explore two complementary paradigms with corresponding loss functions:
Cox Proportional Hazards (DeepSurv): The first approach uses Cox partial likelihood loss:
where hi = DeepSurv(ffusion) is the predicted log-hazard, ei indicates whether the event occurred (1) or was censored (0), ti is the survival time, and R(ti) is the risk set at time ti. This approach estimates relative hazard ratios while preserving the classical proportional hazards assumption.
Discrete-Time Survival (DeepHit): The second approach uses DeepHit objective, which estimates joint distribution of survival time and event occurrence:
where τ is the maximum follow-up time and p(T, E∣ffusion) is the joint distribution predicted by the DeepHit network. This approach directly captures complex, non-linear, and time-varying risk patterns, making it well-suited for multi-center datasets with diverse patient trajectories.
Both Cox and DeepHit models can be trained in parallel, allowing us to benchmark survival modeling under both classical statistical assumptions and flexible deep learning formulations.
Auxiliary Loss Terms: Beyond task-specific objectives, we introduce auxiliary losses to encourage learning of clinically meaningful multimodal representations:
Modality Alignment Loss (InfoNCE Contrastive): An InfoNCE-based contrastive loss aligns embeddings across modalities at the patient level:
where fi and fj are embeddings from different modalities of the same patient (positive pair), τ is temperature, and s( ⋅ , ⋅ ) is cosine similarity. By pulling together representations derived from imaging, pathology, genomic, and laboratory modalities of the same patient while pushing apart embeddings from different patients, the model learns to capture shared semantic structures across data types. This mitigates modality-specific overfitting and ensures robustness under missing-modality scenarios.
Anatomical-Pathological Consistency Loss: A consistency loss enforces spatial coherence between radiological and histopathological findings:
where ROIimaging is the region-of-interest mask from imaging encoder attention (Grad-CAM), Attnpathology is the attention map from pathology encoder (MIL), \({\mathbb{I}}[\cdot ]\) is the indicator function, and τ is a threshold. This cross-modal anatomical consistency regularization constrains the model to focus on clinically relevant tumor regions rather than spurious correlations, and enhances interpretability by providing radiology-pathology correspondences that can be inspected by clinicians.
L2 Regularization: To stabilize training and prevent overfitting:
where θ are network parameters and γ is the regularization coefficient.
Composite Loss Function: The complete composite loss is defined as
where \({L}_{{\rm{surv}}}={L}_{{\rm{surv}}}^{{\rm{Cox}}}\) or \({L}_{\,{\rm{surv}}}^{{\rm{DeepHit}}}\) depending on which survival paradigm is chosen, and λdiag, λsub, λsurv, α, β, γ are balancing coefficients controlling the relative importance of each component. These coefficients can be tuned via grid search or dynamically adjusted using uncertainty-based weighting strategies (where each task k is assigned a learnable uncertainty \({\sigma }_{k}^{2}\) and weighted as \(\frac{1}{2{\sigma }_{k}^{2}}{L}_{k}+\log {\sigma }_{k}\)) to adaptively prioritize tasks with higher predictive uncertainty (see Algorithm 2 for implementation details).
This composite design ensures that UroFusion-X not only optimizes task-specific accuracy but also produces clinically interpretable, modality-robust, and anatomically grounded predictions, thereby enhancing its translational utility for real-world urological oncology.The detailed definitions and purposes of each loss component are summarized in Table 1.
Training strategy
The proposed training strategy employs a carefully designed two-stage protocol to balance modality-specific feature extraction with cross-modal representation learning. The two stages are: (1) modality-specific pretraining, where each encoder learns strong unimodal representations independently; and (2) joint fine-tuning with modality dropout, where all encoders are integrated for end-to-end multimodal optimization. This decoupled approach is motivated by the unique challenges of clinical multimodal datasets, which are characterized by heterogeneous modalities, imbalanced sample sizes, and frequent missing data. By decoupling pretraining from joint optimization, the framework ensures that each encoder learns a strong unimodal representation before engaging in multimodal fusion, thereby mitigating the risk of modality dominance and catastrophic forgetting.
Stage 1: Modality-Specific Pretraining. In the first stage, each encoder is trained independently using modality-tailored self-supervised objectives that leverage large amounts of unlabeled or weakly labeled data. For imaging modalities (CT, MRI, PET), we adopt a masked autoencoder (MAE) objective, where random patches of volumetric input are masked and reconstructed. The masking ratio is typically 75%, ensuring sufficient context remains for reconstruction while forcing the model to capture holistic spatial structure. This forces the imaging encoder to learn spatially coherent and anatomically grounded representations.
For digital pathology whole-slide images, contrastive learning is applied on augmented patch pairs, encouraging the pathology encoder to capture fine-grained histological structures invariant to staining variations and scanner conditions. Data augmentation includes color jittering, rotation, and elastic deformations to simulate realistic staining and scanning variations.
For high-dimensional omics data such as genomics, transcriptomics, or proteomics, we employ masked feature modeling, where random subsets of gene expression vectors are masked and predicted from the remaining context. This encourages the GNN encoder to learn correlations in gene expression patterns and understand pathway-level interactions.
Similarly, for laboratory assays, masked prediction tasks are applied to clinical panels, enabling the model to capture correlations across biochemical markers. By masking random subsets of features, the TabTransformer learns which laboratory measurements are predictive of others.
This stage produces strong unimodal encoders that serve as initializations for the downstream multimodal fusion. The pretraining phase typically requires fewer computational resources than end-to-end training, as each encoder can be trained independently and in parallel, and can later be aligned into a shared representation space.
Stage 2: Joint Fine-tuning with Modality Dropout. After unimodal pretraining, all encoders are integrated into the multimodal fusion framework and jointly optimized end-to-end. We employ the AdamW optimizer with decoupled weight decay, paired with a cosine annealing learning rate scheduler, to ensure stable convergence over long training horizons. The learning rate typically starts at 10−4 and decays to 10−6 following a cosine schedule over the full training duration.
Crucially, we introduce modality dropout as a form of structured regularization: during training, one or more modalities are randomly masked, forcing the model to rely on the remaining modalities. This simulates real-world clinical scenarios where certain data types (e.g., pathology slides or genomic sequencing) may be unavailable due to cost, logistics, or quality constraints. The modality dropout mechanism operates at the input level: for each training sample, we independently set each modality’s availability indicator mi to 0 with probability pdropout, which is passed to the gated PoE module (Algorithm 1). This forces the model to learn how to gracefully handle missing modalities rather than learning spurious dependencies.
By repeatedly exposing the model to incomplete multimodal combinations, modality dropout prevents over-reliance on any single modality and enforces robustness to missing data. Furthermore, we incorporate curriculum-based dropout scheduling, where the probability of dropping modalities is gradually increased as training progresses: \({p}_{{\rm{dropout}}}(t)={p}_{\max }\cdot {(t/T)}^{\beta }\) where t is the current epoch, T is total epochs, and β controls the scheduling curve. This curriculum strategy allows the model to first stabilize on complete data (high loss signal) before being progressively challenged with incomplete scenarios (lower signal-to-noise ratio).
Together, these two stages ensure that UroFusion-X learns (i) robust and clinically meaningful unimodal representations and (ii) stable multimodal embeddings that generalize across diverse patient cohorts and data availability conditions. This design not only enhances predictive performance but also improves real-world deployability in heterogeneous hospital environments.
Evaluation framework
Model evaluation employs a comprehensive multi-level assessment protocol designed to rigorously validate both the technical performance of UroFusion-X and its clinical utility. To prevent overfitting and ensure methodological rigor, internal validation is conducted using stratified cross-validation with temporal splits, where earlier patient cohorts are separated from later ones. This setup mitigates data leakage and reflects realistic deployment scenarios in which models are applied prospectively. External validation adopts a leave-one-center-out (LOCO) strategy, systematically holding out data from one hospital while training on the remaining centers. In LOCO evaluation, we train N separate models, each holding out one of the N centers, and report the average performance across all held-out sets. This cross-institutional design directly tests generalizability to unseen healthcare environments, which is essential for clinical translation across diverse hospitals with different scanners, protocols, and patient populations.
Performance is quantified using task-appropriate metrics. For categorical classification tasks such as primary diagnosis and molecular subtyping, we report the area under the receiver operating characteristic curve (AUC), F1-score, and expected calibration error (ECE). AUC evaluates overall discriminative ability, F1 balances sensitivity and precision in imbalanced datasets, and calibration error measures alignment between predicted probabilities and observed outcomes. For survival prediction tasks, we adopt the concordance index (C-index) as the primary measure of risk ranking, complemented by time-dependent AUC and the integrated Brier score to capture temporal discrimination and probabilistic accuracy of survival distributions. Beyond algorithmic accuracy, clinical significance is assessed using decision curve analysis (DCA), which estimates the net clinical benefit of model-guided decisions compared to standard-of-care baselines. Statistical comparisons between models are performed using DeLong’s test for paired AUCs and stratified log-rank tests for survival endpoints, ensuring that reported improvements are statistically robust.
A distinctive component of the evaluation is the systematic analysis of missing-modality robustness. To emulate real-world scenarios where certain data types may be absent due to cost, availability, or emergency constraints, we simulate controlled missing-modality conditions during testing. This evaluation protocol operates as follows: for each test sample, we systematically set modalities to be unavailable (one at a time for single-modality dropouts, or in combinations for multi-modality dropouts) and re-evaluate model performance. This produces a robustness profile showing how performance degrades as modalities become unavailable.
For single-modality dropouts, we evaluate performance when each individual modality (imaging, pathology, genomics, laboratory) is missing. For example, missing imaging assesses whether the model can still make predictions using only pathology, genomics, and laboratory data. This test evaluates the contribution of each modality and identifies which modalities are most critical for accurate diagnosis and prognosis. For multi-modality deficits, we evaluate combinations of missing modalities, such as missing both pathology and laboratory results (imaging + genomics only), or missing both imaging and genomics (pathology + laboratory only). This tests robustness to more realistic scenarios where multiple data sources may be unavailable simultaneously due to cost constraints or data collection barriers in resource-limited settings.
We also measure cumulative robustness by calculating the fraction of full-modality performance retained when modalities are missing. The target is ≥90% performance retention, meaning the model should maintain at least 90% of its full-modality diagnostic/prognostic accuracy even when one modality is completely absent. This stringent requirement ensures clinical viability in environments with data scarcity.
The gated Product-of-Experts fusion mechanism (the section “Architecture Design”) is specifically designed to handle these missing-modality scenarios gracefully. By learning adaptive gates that downweight unreliable or missing modalities while amplifying informative ones, the gated PoE enables the model to maintain stable predictions even under substantial data degradation. The ability of UroFusion-X to maintain stable performance under such degraded input conditions represents a critical requirement for clinical deployment in heterogeneous or resource-limited environments. Results are visualized in Figure 6, which demonstrates that our gated Product-of-Experts fusion mechanism significantly reduces performance degradation compared to baseline fusion strategies.
Model-predicted high- and low-risk groups for bladder cancer, renal cell carcinoma, and prostate cancer, showing consistent and early separation in survival trajectories with statistically significant differences (p < 0.01, log-rank test).
Dataset and cohort description
Our study utilizes a large multi-center cohort constructed from publicly available repositories, specifically The Cancer Genome Atlas (TCGA) and The Cancer Imaging Archive (TCIA). The cohort comprises patients diagnosed with bladder urothelial carcinoma (BLCA), kidney renal clear cell carcinoma (KIRC), and prostate adenocarcinoma (PRAD). Each patient contributes heterogeneous diagnostic modalities—including radiological imaging (CT/MRI), whole-slide histopathology (WSI), genomic sequencing, and clinical variables—enabling comprehensive multi-modal disease characterization.
To evaluate the framework’s robustness under realistic cross-institutional shifts, we organized the TCGA/TCIA data into three distinct partitions, referred to as Center A, Center B, and Center C (Fig. 7). These partitions simulate a multi-center environment with varying patient volumes and cancer-type compositions (Fig. 7a). Center A represents the largest partition with balanced representation across all three cancer types, mimicking a comprehensive tertiary care center. In contrast, Centers B and C exhibit cancer-type-specific concentrations, mirroring specialized referral patterns often observed in clinical practice. [cite_start]Demographic patterns (Fig. 7b) vary consistently across diseases, with bladder cancer exhibiting a broader age range and prostate cancer showing the expected male predominance[cite: 1, 2].
Multi-modal data acquisition and preprocessing pipeline.
Modality availability across these partitions reveals pronounced structural heterogeneity (Fig. 7c), reflecting the inherent data incompleteness in real-world repositories. Imaging data (CT/MRI) obtained from TCIA are matched to available clinical endpoints. Pathology and genomic sequencing coverage vary due to the retrospective nature of data collection and quality control criteria. For instance, genomic profiles show the highest overall rate of absence, with approximately 55% of samples lacking complete sequencing data due to sample quality or assay availability. Pathology slides are occasionally unavailable in specific subsets (e.g., approximately 45% missing in the Center C partition for RCC), simulating scenarios where biopsy slides may be inaccessible. These realistic disparities in modality availability naturally motivate evaluating robustness under incomplete-modality settings and underscore the necessity of our missing-modality evaluation framework.
The dataset is partitioned using strict patient-level separation to prevent data leakage. As shown in Fig. 7d, the training, validation, and internal test sets are created using a 70%–15%–15% scheme, ensuring strict non-overlap of patients across partitions. Cross-institutional generalization is assessed using a leave-one-center-out (LOCO) protocol, in which each partition (Center) is treated once as an external test set while the remaining partitions form the training cohort. This LOCO strategy directly tests whether models trained on aggregate multi-source data can generalize to unseen distributions with potentially different patient populations and data characteristics.
Preprocessing pipelines achieve consistently high success rates across all modalities (Fig. 7e). Genomic quality control filters samples with low coverage, high contamination, or insufficient variant information. Pathology color normalization (Macenko method for H&E stained slides) corrects for staining variations inherent to multi-site data collection in TCGA. Imaging harmonization standardizes voxel resolution and intensity ranges across different scanners found in the TCIA repository. Success rates exceed 95% across all preprocessing modules, ensuring that standardized, high-quality inputs feed into downstream modeling.
Event distributions and cancer-specific survival curves reveal distinct prognostic patterns across the three diseases (Fig. 7f). Bladder cancer shows the highest frequency of adverse events and shortest median follow-up, reflecting the aggressive nature of high-grade urothelial carcinoma. [cite_start]Renal cell carcinoma demonstrates intermediate event rates, while prostate cancer exhibits the lowest event frequency within the follow-up window, consistent with its generally slower progression[cite: 3]. These heterogeneous survival distributions motivate the use of cancer-specific survival heads within our multi-task learning framework.
Data preprocessing and harmonization
To ensure comparability across institutions and modalities, all data underwent a unified preprocessing and harmonization pipeline. Radiological volumes were intensity-normalized to zero mean and unit variance using standard normalization, then resampled to a consistent voxel resolution of 2 mm (isotropic) to enable direct comparison across scanners and protocols from different vendors. Histopathology whole-slide images were standardized through Macenko stain normalization for H&E slides, which corrects for staining variations and scanner differences across institutions, followed by high-quality tile extraction at 224 × 224 pixel resolution. Tiles containing predominantly background or artifacts were filtered out automatically using tissue segmentation.
Genomic data from heterogeneous sequencing panels (varying in gene coverage and variant calling parameters across centers) were harmonized via comprehensive variant filtering to remove common polymorphisms and likely artifacts, followed by panel alignment to a unified reference gene set to ensure consistent feature representation across samples. Laboratory biomarkers were normalized with respect to clinical units and reference ranges, ensuring that measurements from different laboratory systems were placed on comparable scales. Outlier removal was performed using the interquartile range (IQR) method (values > 1.5 × IQR above Q3 or below Q1 flagged as potential outliers), followed by careful clinical review to distinguish true outliers from extreme but clinically valid measurements.
The preprocessing pipeline was specifically designed to accommodate the structural heterogeneity in modality availability observed in the cohort. Despite the comprehensive nature of these standardization steps, the inherent limitations in data completeness remain: imaging data achieve > 95% success rates after preprocessing due to consistent clinical acquisition protocols, while pathology and genomic data face higher failure rates (approximately 45% preprocessing failure for genomic samples with insufficient coverage) reflecting real-world constraints in specimen quality and sequencing depth. This realistic distribution of preprocessing success rates and missing modalities directly motivates the design of the gated Product-of-Experts fusion mechanism, which must robustly handle incomplete and heterogeneous inputs. Figure 1 summarizes the performance of the proposed framework across diagnostic, subtyping, and survival tasks, demonstrating that despite the preprocessing challenges and data heterogeneity, the unified preprocessing pipeline produces high-quality standardized inputs that enable robust multimodal learning. The ROC curves in Fig. 1a demonstrate consistent improvements of UroFusion-X over comparator models across all three cancer types, with diagnostic AUROC values of 0.92 for bladder cancer, 0.90 for renal cell carcinoma, and 0.88 for prostate cancer, substantially outperforming unimodal imaging baselines and standard fusion approaches.
a Distribution of patients across the three constructed data partitions (Centers) and cancer types. b Patient demographics including age distributions and sex ratios. c Modality availability heatmap, highlighting heterogeneous data completeness across partitions. d Data splitting protocol showing 70-15-15 split and the leave-onecenter-out (LOCO) validation strategy. e Preprocessing success rates across modalities. f Event distribution and Kaplan-Meier survival curves for Bladder, Kidney, and Prostate cancer cohorts from TCGA.
The leave-one-center-out (LOCO) evaluation in Fig. 1b highlights robust generalization under cross-institutional shifts, with performance degradation ranging from 2% to 6% in AUROC when each center is held out as an external test set. This modest degradation—substantially smaller than typical domain shift effects in medical AI—underscores that the preprocessing harmonization successfully addresses institutional variation. Calibration analysis shown in Fig. 1c reveals close alignment between predicted probabilities and observed outcomes, with expected calibration error (ECE) ranging from 0.08 to 0.12, indicating reliable risk estimation suitable for clinical decision-making.
Multi-task learning performance profiles in Fig. 1d demonstrate complementary gains across diagnostic and prognostic objectives, with joint training improving diagnosis task by 3–5%, molecular subtyping by 2–4%, and survival concordance index by 0.03-0.05 compared to single-task baselines. Subgroup analysis in Fig. 1e presents AUROC across demographic and clinical attributes, showing stable performance across patient subgroups with variations < 0.05, indicating demographic robustness. Finally, Fig. 1f compares UroFusion-X performance with radiologist diagnostic accuracy, showing the model achieves 88% diagnostic accuracy compared to average radiologist accuracy of 82%, while reducing comprehensive assessment time from 8 to 12 min per case to approximately 2–5 min.
These comprehensive results demonstrate that despite the inherent challenges of preprocessing heterogeneous multimodal data across multiple institutions, the unified preprocessing and harmonization pipeline successfully creates standardized inputs that enable the proposed framework to achieve state-of-the-art performance while maintaining robust cross-institutional generalization and clinical interpretability.
Experimental protocols
All experiments were conducted under a unified evaluation framework designed to assess both internal performance and cross-center generalizability. As shown in Fig. 7d, the cohort was split at the patient level into training, validation, and internal test sets using a 70%–15%–15% scheme, ensuring strict non-overlap between subjects across partitions and preventing data leakage. To evaluate robustness under real-world institutional shifts, we further employed a leave-one-center-out (LOCO) protocol in which each hospital acted once as an external test site while the remaining centers formed the training set.
The LOCO evaluation strategy serves multiple critical purposes. First, it directly tests whether models trained on aggregate multi-center data can generalize to unseen institutions with potentially different patient populations, acquisition protocols, and data distributions. Second, it enables institution-specific performance characterization, revealing which centers pose greater domain shift challenges. Notably, the LOCO performance maps reveal that Center B exhibits the smallest performance decline when held out (approximately 2–3% AUROC degradation), indicating closer distributional alignment with the aggregated multi-center training data. In contrast, Centers A and C show larger shifts (4–6% degradation), underscoring the heterogeneous nature of cross-center distributional changes. This pattern highlights the importance of institution-aware robustness evaluation and demonstrates that a single global model can successfully adapt to varying institutional characteristics despite differences in imaging hardware, sequencing throughput, and patient demographics.
To assess resilience to incomplete clinical records—a critical requirement for real-world deployment—we additionally simulated missing-modality scenarios at inference by randomly omitting one or two modalities. These missing-modality simulations mirror practical settings in which genomic profiles or pathology slides may be unavailable due to workflow constraints (e.g., insufficient tissue for histology), cost limitations (genomic sequencing expensive in resource-limited centers), or technical failures. We systematically evaluated both single-modality dropouts (absence of one modality, e.g., missing imaging only) and multi-modality deficits (simultaneous absence of two modalities, e.g., missing both pathology and genomic data), with performance metrics computed for each missing-modality combination. The primary robustness metric is the fraction of full-modality performance retained, with the target being ≥90% accuracy retention even when individual modalities are completely absent. This comprehensive missing-modality evaluation directly demonstrates whether the gated Product-of-Experts fusion mechanism can gracefully degrade performance rather than catastrophically failing when modalities are absent.
Baselines
To contextualize the performance of UroFusion-X, we compared it against a comprehensive suite of unimodal, multimodal, and clinically deployed baselines, organized across three distinct categories.
Unimodal Baseline Models. For modality-specific benchmarks, we included state-of-the-art architectures representative of each data type to assess the predictive capacity of individual modalities in isolation. For whole-slide histopathology analysis, we implemented CLAM (Clustering-constrained Attention Multiple instance learning) and TransMIL (Transformer-based multiple instance learning), which represent leading approaches in weakly supervised learning from gigapixel pathology images. For genomic data, we employed GraphSAGE-inspired graph neural networks that capture gene-gene interactions and pathway-level relationships, enabling the model to leverage biological network structure. For structured laboratory biomarkers, we adopted the TabTransformer architecture, which combines transformer-based feature interaction learning with tabular data processing. These unimodal references establish performance ceilings for each individual modality and provide a foundation for assessing the added value of multimodal integration by quantifying the improvement when heterogeneous data types are fused.
Multimodal Fusion Baseline Models. We benchmarked multimodal fusion approaches across three widely used paradigms to systematically evaluate different strategies for integrating heterogeneous biomedical data. The first baseline is late feature concatenation, where modality-specific embeddings are simply concatenated and passed to a final classifier, representing a naive fusion strategy. The second baseline employs attention-based fusion mechanisms, where learned attention weights dynamically reweight modality embeddings based on their relevance to the current prediction task, capturing task-dependent importance of different modalities. The third baseline uses the standard product-of-experts (PoE) formulation without adaptive gating, serving as a direct comparison to our proposed gated PoE architecture. These three fusion paradigms capture philosophically distinct approaches to heterogeneous data integration: concatenation assumes all modalities contribute equally, attention-based fusion adapts weights per sample but lacks explicit missing-modality handling, and standard PoE provides probabilistic fusion but without learned gating to handle reliability variations. Together, they enable systematic evaluation of how different fusion strategies behave under cross-center distribution shifts and varying levels of modality availability.
Clinical Baseline Models. To ensure clinical relevance and ground our technical improvements in established clinical practice, we compared UroFusion-X with widely adopted prognostic scoring systems that represent current standard-of-care in urologic oncology. For prostate cancer, we included CAPRA-S (Cancer of the Prostate Risk Assessment Score for the Surveillance era), a multivariable risk stratification tool based on clinical and pathological factors. For renal cell carcinoma, we included SSIGN (Stage, Size, Grade, and Necrosis score), which combines imaging and pathological information to stratify patients by metastasis-free survival risk. For bladder cancer, we included the EORTC (European Organization for Research and Treatment of Cancer) risk tables, which use tumor characteristics to predict recurrence and progression risk. These clinical baselines serve two purposes: they anchor our quantitative results to clinically actionable reference points, and they represent the performance level that any AI model must exceed to warrant clinical adoption.
In interpreting these baseline comparisons, we account for the dataset-specific imbalance patterns highlighted in Fig. 7, particularly the heterogeneous modality availability and cancer-type distributions that influence the relative performance of different baseline models. For instance, the strong performance of unimodal imaging models on prostate cancer reflects the abundant imaging data in this cohort, whereas pathology-dominant performance on bladder cancer reflects the higher availability of biopsy material in bladder cancer cases. These dataset characteristics mean that unimodal and multimodal baselines perform differently across cancer types in ways that reflect real-world data availability patterns rather than fundamental algorithmic limitations. Furthermore, the clinical baselines (CAPRA-S, SSIGN, EORTC) are designed using traditional clinical and pathological variables (TNM stage, grade, etc.) available from standard clinical evaluation, whereas UroFusion-X integrates additional high-dimensional biomarkers (genomic, transcriptomic, proteomic) not readily captured in traditional scoring systems, potentially explaining performance improvements beyond clinical baselines.
Evaluation metrics
Model performance was assessed using a set of complementary metrics spanning diagnostic accuracy, prognostic discrimination, calibration quality, and clinical utility. For diagnostic and molecular subtyping tasks, we report the area under the receiver operating characteristic curve (AUROC), overall accuracy, and macro-averaged F1-score. AUROC quantifies the discriminatory ability of the model across all classification thresholds, accuracy measures the fraction of correctly classified samples, and F1-score balances precision and recall, particularly important when classes are imbalanced. These three metrics together provide a comprehensive view of model behavior under class imbalance and varying decision thresholds.
Survival prediction performance was quantified using multiple complementary metrics that capture different aspects of prognostic discrimination. The concordance index (C-index) measures the ability to correctly rank patients by predicted risk, ranging from 0.5 (random ranking) to 1.0 (perfect ranking). Time-dependent area under the receiver operating characteristic curve (AUC) evaluates discrimination at specific timepoints (e.g., 1-year, 3-year, 5-year AUC), capturing how predictive power changes over the follow-up period. The integrated Brier score (IBS) measures the mean squared error between predicted survival probability and observed outcomes integrated over the entire follow-up horizon, jointly capturing both discrimination and calibration performance. Together, these metrics provide a multi-dimensional assessment of survival prediction quality.
Calibration quality was evaluated using expected calibration error (ECE) and reliability diagrams. ECE partitions predicted probabilities into bins and compares the average predicted probability within each bin to the observed frequency of positive outcomes, providing a single summary statistic of calibration. Reliability diagrams visualize this comparison across the full probability range, offering intuitive insight into where the model over- or under-estimates risk. These calibration metrics are critical for clinical deployment, where accurate probability estimates directly guide treatment decisions and risk counseling.
Decision curve analysis (DCA) was used to evaluate the clinical utility of the model by estimating the net benefit of model-guided decisions compared to standard-of-care strategies across a range of decision thresholds. DCA translates model predictions into actionable clinical decisions by quantifying the trade-off between true positive rate (patients who benefit from intervention) and false positive rate (patients unnecessarily exposed to intervention risks). This patient-centric perspective is essential for assessing whether AI-assisted predictions genuinely improve clinical decision-making compared to existing alternatives.
Beyond these core metrics, we incorporated subgroup-specific AUROC analyses across demographic and clinical attributes (age groups, sex, cancer grade, TNM stage) to assess whether the model exhibits systematic performance variations across patient populations. We also computed center-wise performance metrics derived from the leave-one-center-out protocol, enabling fine-grained characterization of cross-institutional robustness. These supplementary analyses enable the detection of potential demographic or institutional disparities that could affect equitable deployment in diverse clinical settings.
Ethics approval and consent to participate
This study exclusively uses publicly available datasets (e.g., TCGA, TCIA) containing de-identified patient data. The study does not involve any new experiments with human participants or animals performed by any of the authors. Therefore, additional ethical approval and patient consent were not required by the institutional review board.
Data availability
The datasets generated and/or analyzed during the current study are available in the following public repositories: Genomic and Clinical Data: The Cancer Genome Atlas (TCGA) datasets for Bladder Urothelial Carcinoma (TCGA-BLCA), Kidney Renal Clear Cell Carcinoma (TCGA-KIRC), and Prostate Adenocarcinoma (TCGA-PRAD) are available via the NCI Genomic Data Commons (GDC) Data Portal: https://portal.gdc.cancer.gov/. Imaging Data: Corresponding radiology data (CT/MRI) are accessible via The Cancer Imaging Archive (TCIA): https://www.cancerimagingarchive.net/. Pathology Data: Whole-slide histopathology images are available through the GDC portal and TCIA.Processed data and intermediate results supporting the findings of this study are available from the corresponding author upon reasonable request.
Code availability
The source code for the proposed UroFusion-X framework, including data preprocessing pipelines, model architecture, and training scripts, is available at the following anonymous repository: https://anonymous.4open.science/r/UroFusion-X--coding-F547/README.md.
References
Stahlschmidt, S. R., Ulfenborg, B. & Synnergren, J. Multimodal deep learning for biomedical data fusion: a review. Brief. Bioinform. 23, bbab569 (2022).
Huang, S.-C., Pareek, A., Seyyedi, S., Banerjee, I. & Lungren, M. P. Fusion of medical imaging and electronic health records using deep learning: a systematic review and implementation guidelines. NPJ Digital Med. 3, 136 (2020).
Waqas, A. et al. Digital pathology and multimodal learning on oncology data. BJR∣ Artif. Intell. 1, ubae014 (2024).
Zhou, H. et al. Multimodal data integration for precision oncology: challenges and future directions. Preprint at https://arxiv.org/abs/2406.19611 (2024).
Hatamizadeh, A. et al. UNETR: Transformers for 3D Medical Image Segmentation. In 2022 IEEE/CVF Winter Conference on Applications of Computer Vision (WACV), 1748–1758 (Waikoloa, HI, USA, 2022).
Hatamizadeh, A. et al. Swin UNETR: Swin Transformers for Semantic Segmentation of Brain Tumors in MRI Images. In: (Crimi, A. & Bakas, S. eds) Brainlesion: Glioma, Multiple Sclerosis, Stroke and Traumatic Brain Injuries. BrainLes 2021. Lecture Notes in Computer Science, vol 12962 (Springer, Cham., 2022).
Dosovitskiy, A. et al. An image is worth 16x16 words: Transformers for image recognition at scale. Preprint at https://arxiv.org/abs/2010.11929 (2020).
He, K. et al. Masked Autoencoders Are Scalable Vision Learners. In 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 15979–15988 (New Orleans, LA, USA, 2022).
Lu, M. Y. et al. Data-efficient and weakly supervised computational pathology on whole-slide images. Nat. Biomed. Eng. 5, 555–570 (2021).
Bulten, W. et al. Artificial intelligence for diagnosis and gleason grading of prostate cancer: the panda challenge. Nat. Med. 28, 154–163 (2022).
Shao, Z. et al. Transmil: Transformer based correlated multiple instance learning for whole slide image classification. Adv. neural Inf. Process. Syst. 34, 2136–2147 (2021).
Chaudhary, K., Poirion, O. B., Lu, L. & Garmire, L. X. Deep learning–based multi-omics integration robustly predicts survival in liver cancer. Clin. cancer Res. 24, 1248–1259 (2018).
Katzman, J. L. et al. Deepsurv: personalized treatment recommender system using a cox proportional hazards deep neural network. BMC Med. Res. Methodol. 18, 24 (2018).
Lee, C., Zame, W., Yoon, J. & van der Schaar, S. DeepHit: A Deep Learning Approach to Survival Analysis With Competing Risks. Proceedings of the AAAI Conference on Artificial Intelligence, vol. 32 (2018).
Zheng, S. et al. Multi-modal graph learning for disease prediction. IEEE Trans. Med. Imaging 41, 2207–2216 (2022).
Ding, S., Li, J., Wang, J., Ying, S. & Shi, J. Multimodal co-attention fusion network with online data augmentation for cancer subtype classification. IEEE Trans. Med. Imaging 43, 3977–3989 (2024).
Tariq, A. et al. Fusion of imaging and non-imaging data for disease trajectory prediction for coronavirus disease 2019 patients. J. Med. Imaging 10, 034004–034004 (2023).
Zhao, Y. et al. Machine learning-based MRI imaging for prostate cancer diagnosis: systematic review and meta-analysis. Prostate Cancer and Prostatic Diseases 1–8, https://doi.org/10.1038/s41391-025-00997-2 (2025).
Ayyad, S. M., Abdel-Hamid, N. B., Ali, H. A. & Labib, L. M. Multimodality imaging in prostate cancer diagnosis using artificial intelligence: basic concepts and current state-of-the-art. Multimedia Tools Appl. 84, 42649–42678 (2025).
Bai, Z. et al. Predicting response to neoadjuvant chemotherapy in muscle-invasive bladder cancer via interpretable multimodal deep learning. npj Digital Med. 8, 174 (2025).
Ma, M. et al. Smil: Multimodal learning with severely missing modality. Proc. AAAI Conf. Artif. Intell. 35, 2302–2310 (2021).
Azad, R., Khosravi, N., Dehghanmanshadi, M., Cohen-Adad, J. & Merhof, D. Medical image segmentation on mri images with missing modalities: a review. Preprint at https://arxiv.org/abs/2203.06217 (2022).
Selvaraju, R. R. et al. Grad-CAM: Visual Explanations from Deep Networks via Gradient-Based Localization. In 2017 IEEE International Conference on Computer Vision (ICCV), 618–626 (Venice, Italy, 2017).
Khene, Z.-E. et al. Clinical application of digital and computational pathology in renal cell carcinoma: a systematic review. Eur. Urol. Oncol. 7, 401–411 (2024).
Gogoshin, G. & Rodin, A. S. Graph neural networks in cancer and oncology research: Emerging and future trends. Cancers 15, 5858 (2023).
Huang, X., Khetan, A., Cvitkovic, M. & Karnin, Z. Tabtransformer: tabular data modeling using contextual embeddings (2020). Preprint at https://arxiv.org/abs/2012.06678 (2012).
Huang, X., Mallya, A., Wang, T. C. & Liu, M. Y. Multimodal Conditional Image Synthesis with Product-of-Experts GANs. In: (Avidan, S. et al. eds) Computer Vision – ECCV 2022. ECCV 2022. Lecture Notes in Computer Science, vol. 13676 (Springer, Cham, 2022).
Vickers, A. J. & Elkin, E. B. Decision curve analysis: a novel method for evaluating prediction models. Med. Decis. Mak. 26, 565–574 (2006).
Suarez-Ibarrola, R., Hein, S., Reis, G., Gratzke, C. & Miernik, A. Current and future applications of machine and deep learning in urology: a review of the literature on urolithiasis, renal cell carcinoma, and bladder and prostate cancer. World J. Urol. 38, 2329–2347 (2020).
Zadeh, A., Chen, M., Poria, S., Cambria, E. & Morency, L.-P. Tensor fusion network for multimodal sentiment analysis. Preprint at https://arxiv.org/abs/1707.07250 (2017).
Acknowledgements
This project did not receive any financial support.
Author information
Authors and Affiliations
Contributions
Y.X., S.Y., and M.H. contributed equally to this work, having full access to all study data and assuming responsibility for the integrity and accuracy of the analyses (Validation, Formal analysis). Y.X. conceptualized the study, designed the methodology, and participated in securing research funding (Conceptualization, Methodology, Funding acquisition). S.Y. carried out data acquisition, curation, and investigation (Investigation, Data curation) and provided key resources, instruments, and technical support (Resources, Software). M.H. drafted the initial manuscript and generated visualizations (Writing—Original Draft, Visualization). L.C., Y.W., and L.Z. supervised the project, coordinated collaborations, and ensured administrative support (Supervision, Project administration). All authors contributed to reviewing and revising the manuscript critically for important intellectual content (Writing—Review & Editing) and approved the final version for submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xiao, Y., Yang, S., He, M. et al. UroFusion-X: a unified multimodal deep learning framework for robust diagnosis, subtyping, and prognosis of urological cancers. npj Digit. Med. 9, 117 (2026). https://doi.org/10.1038/s41746-025-02295-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-02295-6