Abstract
Rare diseases affect an estimated 300–400 million people worldwide, yet individual conditions remain underdiagnosed and poorly characterized due to low prevalence and limited clinician familiarity. Computational phenotyping offers a scalable approach to improving rare disease detection, but algorithm development is constrained by scarce high-quality labeled data. Expert-labeled datasets from chart reviews and registries are highly accurate but limited in scope, whereas labels derived from electronic health records (EHRs) provide broader coverage but are often noisy or incomplete. To efficiently leverage both sources, we propose WEST (WEakly Supervised Transformer) for rare disease diagnosis and subphenotyping from EHRs. At its core, WEST employs a weakly supervised transformer trained on a limited set of expert-validated labels and extensive probabilistic silver-standard labels—derived from structured and unstructured EHR features—that are iteratively refined across training rounds to improve model calibration. We evaluate WEST on two rare pulmonary conditions using EHR data from Boston Children’s Hospital and show that it outperforms existing methods in phenotype classification, identification of clinically relevant subphenotypes, and prediction of disease progression. By reducing reliance on manual annotation, WEST enables label-efficient representation learning that supports accurate rare disease diagnosis and reveals deeper clinical insights from routine EHR data.
Similar content being viewed by others
Introduction
Rare diseases are broadly defined as conditions affecting fewer than 1 in 2000 people in any World Health Organization region or, in the United States, as those affecting fewer than 200,000 people1,2. While individually uncommon, rare diseases collectively impose a substantial public health burden, affecting an estimated 300–400 million people worldwide1,3. In the United States alone, approximately 30 million individuals—10% of the population—are living with a rare disease, a prevalence comparable to that of type 2 diabetes3,4.
Despite their widespread impact, rare diseases—encompassing more than 7000 distinct conditions—remain exceptionally difficult to diagnose. Many clinicians encounter some of these conditions only once, if ever, in their careers, limiting familiarity with their diverse clinical presentations5,6. These challenges contribute to the so-called “diagnostic odyssey”—a years-long process marked by inconclusive tests, repeated specialist referrals, and frequent misdiagnoses—that many rare disease patients experience7. On average, patients see between three and ten physicians and wait 4–7 years before receiving an accurate diagnosis1,5,8. Such delays hinder timely treatment, elevate the risk of preventable complications, and contribute to premature mortality9,10,11. The burden is especially severe in pediatrics, as 70% of rare diseases manifest in childhood and 30% of affected children die before age five1,12. There is therefore an urgent need for more accurate and timely diagnosis to improve outcomes and quality of life for rare disease patients across the lifespan.
These diagnostic challenges are particularly pronounced in rare pulmonary diseases, which are notoriously difficult to identify due to their symptomatic overlap with more common respiratory conditions. Up to one-third of individuals initially diagnosed with asthma are later found to have been misdiagnosed, with their symptoms instead attributable to less prevalent comorbid conditions13,14. Pulmonary hypertension (PH), a progressive disorder characterized by mean pulmonary arterial pressure ≥20 mmHg, often presents with nonspecific symptoms such as breathlessness and hypoxia—features that closely mimic asthma15,16,17. This overlap frequently delays recognition of PH until irreversible vascular damage has occurred18. Severe asthma, a distinct and high-burden phenotype requiring high-dose inhaled corticosteroids along with a second controller to prevent it from becoming uncontrolled, poses a similarly complex diagnostic challenge19,20. Despite accounting for more than one-third of asthma-related deaths, severe asthma remains under-recognized, and its clinical heterogeneity further complicates timely diagnosis and effective management21. Together, these pitfalls underscore the limitations of relying solely on clinical expertise and highlight the need for data-driven approaches capable of detecting subtle, multi-dimensional disease patterns often missed in routine practice.
Efforts to consolidate rare disease cases into condition-specific registries have yielded important clinical and epidemiologic insights, yet most registries remain too small and narrowly focused to support comprehensive, generalizable disease characterization22,23,24. Because registry inclusion typically requires a confirmed diagnosis, patients with atypical presentations or unrecognized disease—those most essential for building representative datasets—are systematically excluded. The widespread adoption of electronic health records (EHRs) has helped address these limitations, enabling rare disease research at scale by capturing a broader and more heterogeneous spectrum of clinical presentations than traditional registries. EHRs contain rich longitudinal data in both structured (e.g., diagnosis, medication, and procedure codes) and unstructured (e.g., free-text notes) formats, documenting each patient’s diagnostic trajectory—including misdiagnoses and testing patterns that trace where patients with rare diseases are missed along the diagnostic pathway 25.
Building on the breadth and depth of these data, machine-assisted clinical decision support is increasingly being integrated into research and healthcare workflows across a wide range of applications26,27,28,29. Within pulmonary medicine, such approaches have shown particular promise, supporting real-time imaging interpretation30,31, flagging high-risk patients for specialist referral32, and retrospectively identifying undiagnosed individuals for inclusion in registries and observational studies33. A central capability underlying many of these advances is the automated analysis of EHR data to infer patient state—such as presence or absence of a disease condition given data observed during a temporal period—often referred to as computational phenotyping. Traditionally, computational phenotyping has relied on rule-based algorithms that apply predefined logical criteria—such as specific diagnostic codes, medication patterns, or characteristic laboratory abnormalities documented in the medical record—to infer disease status34,35. Recently, unsupervised and weakly supervised phenotyping algorithms have emerged as more generalizable alternatives, leveraging rich clinical features and, when available, “noisy" but informative proxy labels36,37,38,39. While effective for well-characterized conditions with standardized coding systems, these methods often have unsatisfactory performance for rare diseases, which are clinically heterogeneous and may lack clearly codified diagnostic criteria40,41. Moreover, they do not support subphenotyping—the stratification of patients into clinically meaningful subgroups based on prognosis or treatment response—because they treat disease as a binary construct, overlooking heterogeneity in presentation and progression. As clinical care increasingly demands personalized diagnosis, risk assessment, and modeling of disease trajectories, there is a growing need to move beyond binary phenotyping toward richer diagnostic and subphenotyping frameworks that capture the full complexity of the patient’s longitudinal EHR profile.
To enable more expressive and scalable modeling of clinical data, machine learning (ML) and deep learning (DL) have emerged as powerful approaches for large-scale disease characterization and prediction40,42. A key development in this space is representation learning, which transforms heterogeneous, high-dimensional EHR data into low-dimensional embeddings that capture semantic, temporal, and contextual structure among medical concepts43. These concept-level representations—learned from both structured codes and unstructured narratives44—can be aggregated into patient-level embeddings that support a wide range of downstream tasks. Importantly, such representations leverage all available EHR data, eliminating the need for labor-intensive feature curation that captures only a small fraction of the information contained in each patient’s record45. By compressing the full EHR into rich summary embeddings with shared latent structure, representation learning enables multi-outcome assessment that extends well beyond traditional single-outcome risk scores46. This foundation allows patient embeddings to drive statistically efficient subphenotyping, disease trajectory modeling, and early detection of high-risk states47. Collectively, these advances motivate replacing hand-engineered rules with flexible ML/DL models that learn scalable, generalizable clinical representations suited to the complexity and diversity of modern healthcare data.
Despite their success in modeling common diseases, existing ML/DL methods often generalize poorly to rare disease contexts due to both data- and model-related challenges. From a data perspective, rare disease EHRs are inherently sparse, heterogeneous, and noisy48,49. Because these conditions are infrequently encountered and inconsistently documented, critical information is often missing or misclassified. For example, the presence of a diagnostic code or concept in the EHR does not necessarily indicate a confirmed diagnosis, as codes may be entered for billing purposes, used provisionally, or persist from outdated assessments50,51. Moreover, variation in documentation practices across providers and institutions further complicates the construction of accurate, large-scale training datasets, amplifying noise and bias in downstream analyses. From a modeling standpoint, most ML/DL approaches for EHR interpretation rely on supervised learning, which depends on large, high-quality labeled datasets—a resource rarely available in rare disease research52. Fully supervised models trained on small cohorts often overfit to narrow or imperfect labels that fail to capture the full spectrum of disease manifestations, limiting their generalizability and clinical utility. Collectively, these challenges have fueled growing interest in weakly supervised learning, which leverages large collections of noisy or partially labeled data to build more robust and data-efficient models in low-label settings.
Among emerging DL architectures, transformers have shown particular promise for modeling EHR data due to their ability to capture complex temporal dependencies and long-range relationships across irregular clinical events53,54. Models such as BEHRT55, Med-BERT56, RatchetEHR57, and Foresight58 have demonstrated state-of-the-art performance across predictive and classification tasks in both structured and unstructured data. These advances underscore the potential of transformer-based models to learn expressive patient representations that support diagnosis, risk prediction, and subphenotyping. Yet despite this promise, existing transformer approaches remain constrained by the same supervised learning paradigm that limits other ML/DL applications in rare diseases. Most require large volumes of clean labeled data to train effectively, which restricts their utility in data-limited, noisy, or label-scarce environments. As a result, their potential for rare disease detection and broad diagnostic modeling remains only partially realized.
To address this gap, we propose a weakly supervised transformer (WEST) framework for learning robust patient representations from EHR data in low-label, high-noise settings. Our end-to-end pipeline integrates a small set of expert-validated gold-standard labels with a much larger pool of silver-standard labels from real-world EHRs, which are refined iteratively through self-training. By updating weak supervision within the training loop rather than relying on fixed or single-pass pseudo-labels, WEST jointly learns contextual code representations and their aggregation into patient-level embeddings that support multiple downstream tasks—including phenotype classification and subphenotype clustering—providing a label-efficient training paradigm for EHR-based modeling and enabling evaluation of representation quality beyond a single predictive endpoint. Importantly, the framework performs effectively even when only positive gold-standard cases are available, making it well suited for rare diseases where registries exist but exhaustive chart review is infeasible. We demonstrate the value of this paradigm through two case studies, using pulmonary hypertension and severe asthma as motivating examples to evaluate WEST’s learned patient representations across multiple downstream tasks. Within this scope, we observe improved disease status classification and the identification of clinically meaningful patient subgroups in rare disease cohorts at Boston Children’s Hospital. Complementing these analyses, we conduct targeted ablation studies to assess the roles of iterative label refinement, transformer-based modeling, and supervision efficiency, providing insight into WEST’s additive value relative to existing ML and DL baselines and examining how its learned representations may support modeling beyond phenotype identification in heterogeneous clinical populations.
Results
We evaluated the WEST framework on two rare pulmonary diseases—PH and severe asthma—using EHR data from Boston Children’s Hospital. For each disease, the model was trained and validated independently using disease-specific EHR cohorts and labels curated by board-certified physicians with subspecialty expertise in PH and severe asthma.
Data curation
For both PH and severe asthma, we constructed disease-specific cohorts by first identifying at-risk patient populations from the EHR data at Boston Children’s Hospital. The at-risk PH cohort comprised 14,305 randomly selected patients with PheCode 415.2 (indicative of potential PH), while the severe asthma cohort comprised 7822 randomly selected patients with International Classification of Diseases, 10th Revision (ICD-10) codes beginning with J45 (indicative of asthma of any severity).
Gold-standard cohorts consisted of patients with confirmed disease status, established either through expert chart review or enrollment in a disease-specific registry. Diagnostic criteria for gold-standard chart review are provided in Section S2 of the Supplementary Materials. The gold-standard PH cohort comprised 531 patients, with 106 (20%) set aside for validation and testing, while the severe asthma cohort comprised 248 patients, with 99 (40%) set aside. Within the validation and testing subsets, the PH cohort included 37 negative and 69 positive cases, whereas the severe asthma cohort included 47 negative and 52 positive cases. These held-out patients were further split into two equally sized cross-validation folds—one used for validation (model checkpoint selection) and the other for testing (performance evaluation). Final performance metrics were averaged across cross-validation folds.
The silver-standard cohorts comprised the remaining at-risk patients whose phenotype status had not been definitively adjudicated, totaling 13,774 for PH and 7575 for severe asthma. Initial probabilistic labels \({y}_{i}^{silver}\in (0,1)\) were assigned to these patients using the Knowledge-driven Online Multimodal Automated Phenotyping (KOMAP) algorithm36.
For PH, KOMAP was applied to codified EHR features, including PheCode diagnoses, RxNorm medications, LOINC laboratory tests, and Clinical Classifications Software (CCS) procedure codes. For severe asthma, KOMAP was applied to natural language features extracted from clinical notes using Narrative Information Linear Extraction (NILE)59. From these codes and concepts, we designated PheCode:415.2 for PH and CUI:C0581126 for severe asthma as the target phenotypes, where CUI denotes a Concept Unique Identifier from the Unified Medical Language System (UMLS). For representation learning, we mapped the codified EHR data in the PH cohort to pre-trained Multisource Graph Synthesis (MUGS) embeddings60 and the natural language processing (NLP)-derived features in the severe asthma cohort to pre-trained Online Narrative and Codified Feature Search Engine (ONCE) embeddings36. This selection reflected both computational feasibility and the distinct data modalities of the two cohorts, while also demonstrating the flexibility of the WEST framework to accommodate different pre-trained embedding sources.
Evaluation metrics
We first assessed the classification performance of the WEST pipeline on labels not used during training. Evaluation was performed using two-fold cross-validation, computing the area under the receiver operating characteristic curve (AUC), F1 score, positive predictive value (PPV), and specificity for each fold and averaging across folds. Sensitivity was fixed at 80% to enable fair and stable comparison across methods at a clinically meaningful detection threshold, reflecting a practical balance between case detection and false-positive burden in low-prevalence rare disease screening. To quantify uncertainty in model performance, 95% confidence intervals for AUC, F1 score, PPV, and specificity were estimated using non-parametric bootstrapping with 1000 resamples on patient-level predictions. WEST performance was compared against five classification baselines:
-
(1)
Count: Binary labels derived by thresholding the frequency of the target concept appearing in each patient’s EHR.
-
(2)
KOMAP: Binary labels obtained by thresholding the initial silver-standard probabilities generated by KOMAP36.
-
(3)
XGBoost: A supervised gradient-boosted trees classifier61.
-
(4)
Transformer (silver = gold): A transformer trained by treating all silver-standard labels as gold-standard, without any iterative updates or data augmentation.
-
(5)
Transformer (gold only): A fully supervised transformer trained exclusively on gold-standard labels.
As additional ablation studies, we examined two aspects of gold-standard supervision. First, we varied the number of gold-standard labels used for training, gradually increasing the labeled set from 25 to 400 examples. Second, we modified WEST to train without any gold-standard negative labels, simulating a setting where no confirmed negatives are available, and all negative training samples are drawn from the silver-standard cohort.
We next evaluated whether the learned patient representations captured clinically meaningful heterogeneity. Among patients with known disease status who were excluded from model training, we assessed whether their embeddings could effectively separate true positive from true negative cases. To visualize this separation, we applied t-distributed stochastic neighbor embedding (t-SNE) to the WEST embeddings of held-out patients and compared the resulting visualization with one derived from term frequency-inverse document frequency (TF-IDF) embeddings, a widely used baseline feature engineering approach62,63,64.
For subphenotype discovery, we focused on patients that the model classified as having positive disease status. The WEST embeddings for these patients were first reduced in dimensionality using principal component analysis (PCA), retaining components that together explained at least 90% of the variance. We then applied k-means clustering to these reduced embeddings to identify latent structures within the representation space corresponding to potential patient subgroups. To visualize the resulting subgroups, we generated t-SNE plots showing the separation of k-means-derived clusters among patients predicted to be disease positive. Finally, we assessed the prognostic relevance of these clusters. For PH, we compared survival distributions using Kaplan-Meier curves. For severe asthma, we estimated hazard ratios (HRs) for recurrent clinical signs and symptoms indicative of disease severity across clusters. Recurrent events were modeled using a Cox proportional hazards model in the Andersen-Gill formulation, which accounts for multiple episodes per patient and within-patient correlation65. Signs and symptoms were identified from the EHR using UMLS Concept Unique Identifiers (CUIs): dyspnea (C0013404), tachypnea (C0231835), bronchospasm (C4552901), low oxygen (C0242184, C1963140, C0700292, C4061338), respiratory failure (C4552651), and status asthmaticus (C0038218).
Pulmonary hypertension
The WEST pipeline trained with both positive and negative gold-standard PH labels achieved the highest overall classification performance—including AUC, F1 score, PPV, and specificity—across all baseline methods (Table 1). Even when trained without gold-standard negative labels, WEST still exceeded the performance of all baselines.
Figures 1 and S1 demonstrate that WEST performance increased steadily with the number of gold-standard training labels. Notably, with as few as 100 labels, WEST matched or outperformed all baseline methods, and continued to improve as more labels were added.
Curves show a AUC and b F1 score with 95% confidence intervals for PH as the number of gold-standard training labels increases. Metrics are averaged across two cross-validation folds. The horizontal black dashed line indicates the best-performing baseline model, Transformer (gold only).
In Fig. 2, we examine model performance across iterative rounds of silver-label refinement for the PH cohort. Performance metrics—including AUC, F1 score, PPV, and specificity—consistently improved from Round 1 to Round 2, reflecting the benefit of updating noisy silver-standard labels with model-generated probabilities. By Round 3, performance curves stabilized, indicating convergence of the refinement process. Accordingly, we report Round 2 results throughout the manuscript.
Classification performance across refinement rounds for the PH cohort. Points denote cross-validated estimates and error bars reflect 95% confidence intervals for a AUC, b F1 score, c PPV, and d specificity. Improvements from Round 1 to Round 2 and stabilization thereafter indicate convergence of the silver-label updating procedure.
As shown in Fig. 3, the WEST embeddings achieved clearer latent-space separation between confirmed PH-positive and PH-negative cases excluded from model training than did TF-IDF embeddings.
t-SNE plots compare the separability of patients using a TF-IDF embeddings and b WEST embeddings. Each blue circle represents a confirmed PH-negative patient and each red circle a confirmed PH-positive patient. WEST embeddings show clearer separation between disease states.
The WEST pipeline identified 1977 patients with PH. Clustering of predicted PH-positive patient embeddings revealed two clinically meaningful subgroups: a Slow Progression cluster (n = 1099) and a Fast Progression cluster (n = 878) (Fig. 4). Kaplan-Meier survival analysis showed a significant difference in 5-year mortality between the two clusters (log-rank p = 0.013; Fig. 5).
t-SNE plot of patient embeddings colored by progression subphenotypes derived from k-means clustering. Each red circle represents a slow progressor (n = 1099) and each blue circle a fast progressor (n = 878) among patients predicted to be PH-positive.
Kaplan-Meier survival curves show 5-year survival probability for Slow Progression (red line) and Fast Progression (blue line) clusters identified by k-means on WEST embeddings. Shaded regions represent 95% confidence intervals. The difference between curves was significant (log-rank p = 0.013).
Severe asthma
Table 2 presents classification metrics across methods for the severe asthma phenotype. Again, when trained with both positive and negative gold-standard labels, WEST achieved the highest AUC, PPV, and specificity, outperforming all baselines.
Patients classified by WEST as having severe asthma demonstrated substantially higher risks for multiple markers of disease severity compared with patients classified as non-severe. Significant associations were observed for recurrent status asthmaticus (HR = 55.30, 95% CI: 43.93–69.61, p < 0.0001) and respiratory failure (HR = 3.19, 95% CI: 2.05–4.97, p < 0.0001). Additional elevated risks were observed for recurrent low-oxygen events (HR = 2.66, 95% CI: 2.05–3.45, p < 0.0001), tachypnea (HR = 3.67, 95% CI: 3.17–4.26, p < 0.0001), bronchospasm (HR = 3.49, 95% CI: 2.20–5.55, p < 0.0001), and dyspnea (HR = 2.97, 95% CI: 2.66–3.32, p < 0.0001), which, when occurring frequently, indicate poorer asthma control (Fig. 6).
Hazard ratios and 95% confidence intervals (horizontal lines) are shown for six clinical indicators of asthma severity, based on recurrent event analyses. Each panel represents a separate comparison: a severe versus non-severe asthma and b high versus low exacerbator clusters among individuals with severe asthma. Vertical dashed lines indicate a hazard ratio of 1 (no association).
Again, among confirmed severe asthma-positive and -negative cases held out from model training, latent-space separation was more distinct when using WEST embeddings than with TF-IDF (Fig. 7).
t-SNE plots compare the separability of patients using a TF-IDF embeddings and b WEST embeddings. Each blue circle represents a confirmed severe asthma-negative patient and each red circle a confirmed severe asthma-positive patient. WEST embeddings show clearer separation between disease states.
Among 582 patients predicted to have severe asthma, k-means clustering identified a Low Exacerbator cluster (n = 209) and a High Exacerbator cluster (n = 373) (Fig. 8). Patients in the High Exacerbator cluster had higher risk of recurrent status asthmaticus (HR = 2.35, 95% CI: 1.91–2.91, p < 0.0001), respiratory failure (HR = 2.68, 95% CI: 1.31–5.47, p = 0.0068), low oxygen events (HR = 1.54, 95% CI: 1.05–2.28, p = 0.0291), and tachypnea (HR = 1.41, 95% CI: 1.11–1.79, p = 0.0050) compared with the Low Exacerbator cluster (Fig. 6).
t-SNE plot of patient embeddings colored by subphenotypes identified by k-means clustering. Each red circle represents a low exacerbator (n = 209) and each blue circle a high exacerbator (n = 373) among patients predicted to have severe asthma.
Discussion
In this study, we introduce WEST, a weakly supervised transformer framework that integrates a limited set of expert-validated annotations with iteratively refined silver-standard labels to support data-efficient modeling of rare diseases from EHR data. Across PH and severe asthma case studies, WEST consistently outperformed rule-based and ML/DL baselines, while also generating patient representations that more clearly separated disease states and revealed clinically meaningful subgroups within each cohort. To contextualize WEST’s methodological contributions, we highlight two key sources of novelty relative to existing approaches: (1) WEST’s iterative weak-supervision strategy, which establishes a label-efficient transformer training paradigm distinct from prior clinical phenotyping frameworks, and (2) WEST’s ability to learn latent disease representations that expose clinically meaningful structure beyond primary labels. In this section, we compare WEST directly to related work, underscoring where and how the framework meaningfully extends existing phenotyping and representation learning methods.
WEST builds on a growing body of work demonstrating that ML/DL models can be trained effectively from imperfect labels when supported by weak supervision. Prior frameworks such as Snorkel66 and FixMatch67, along with recent surveys of weakly supervised learning68, show that noisy or partially labeled data can be transformed into reliable supervision signals through pseudo-labeling (i.e., generating silver-standard labels from weak heuristics) and denoising (i.e., refining those labels using model- or rule-based corrections). Within clinical informatics, a parallel line of work has leveraged surrogate signals to generate silver-standard labels for EHR-based phenotype modeling. Unsupervised approaches such as PheNorm37, MAP69, and PheVis70 derive probabilistic phenotype scores from diagnostic codes, NLP concepts, and healthcare utilization features. A related class of methods—including KOMAP36; Automated Phenotype Routine for Observational Definition, Identification, Training and Evaluation (APHRODITE)38; and weakly semi-supervised DL (WSS-DL)39—extend this regime by incorporating concept embeddings, curated anchor features, or neural architectures to further improve label quality and phenotyping accuracy in low-annotation settings. Collectively, these efforts demonstrate that combining a small number of curated labels with large quantities of noisy or weak labels can substantially improve generalization and provide an effective foundation for label-efficient clinical modeling.
However, none of these frameworks integrate weak supervision with transformer-based modeling or perform iterative refinement of silver-standard labels within the training loop. WEST’s novelty lies in its operationalization of silver-standard label refinement as a self-training (pseudo-labeling) paradigm, where model-predicted probabilities for unlabeled examples are reused as soft labels in subsequent training rounds. While prior weakly supervised phenotyping approaches leverage noisy or probabilistic labels, these labels are typically generated during preprocessing and treated as fixed during downstream training. In contrast, WEST embeds probabilistic silver-standard labels directly within a transformer-based training loop and updates them iteratively as model representations improve. This yields an end-to-end framework that jointly refines supervision and learns contextual patient representations from multimodal EHR data—including diagnoses, procedures, medications, and NLP-derived concepts—while remaining highly label-efficient. By coupling iterative pseudo-label refinement with a transformer backbone, WEST captures long-range dependencies across irregular clinical events 53,54, enabling representations that encode disease evolution, progression patterns, and clinical context beyond what static features or single-pass weak-label pipelines can recover.
In addition to the expressiveness of the transformer architecture, ablation studies demonstrate that WEST’s training paradigm contributes substantial additional performance gains. Across PH and severe asthma, WEST improved AUC over baseline transformers by +0.05 to +0.09 points, reflecting the consistent benefits of its data augmentation and iterative label-refinement strategy beyond architectural capacity alone (Tables 1 and 2). Moreover, when we vary the number of gold-standard labels (Fig. 1), WEST maintains superior performance and remains robust even with as few as 100 expert-labeled examples, underscoring the advantages of label-efficient weak supervision and iterative refinement. Together, these findings indicate that WEST provides a principled way to exploit unlabeled data and learn expressive patient representations even when high-quality labels are scarce. This advantage is especially important in rare disease settings, but it also extends to more common diseases where labeled data remain a bottleneck. The training paradigm itself is broadly applicable and continues to benefit from richer supervision, as reflected by the performance gains observed in Fig. 1.
Beyond identifying primary disease status in PH and severe asthma, WEST produced patient embeddings that clustered to reveal latent structure associated with meaningful clinical heterogeneity. In the PH cohort, these embeddings separated patients into Slow Progressor and Fast Progressor subgroups, which exhibited significantly different long-term survival trajectories. Similarly, in the severe asthma cohort, WEST distinguished Low Exacerbator and High Exacerbator subgroups, with high exacerbators experiencing elevated risk of severe adverse events including recurrent status asthmaticus, respiratory failure, and hypoxemia. Together, these findings indicate that WEST captures underlying dimensions of disease biology and care patterns that extend beyond codified diagnoses, providing a foundation for richer patient-state representation and more nuanced clinical stratification.
Another important feature of WEST is that it learns a latent disease representation rather than optimizing directly for a single clinical endpoint, yielding embeddings that can support multiple downstream tasks including, but not limited to, risk prediction without retraining the model. In this sense, WEST serves a complementary yet fundamentally different role from established clinical risk scores. Prognostic models such as REVEAL 2.0 in PH71, the Risk Score for Asthma Exacerbations (RSE)72, and the Asthma Exacerbation Risk (AER) score73 are supervised tools explicitly optimized to predict a single prespecified outcome (e.g., 12-month mortality in PH or 6- to 12-month exacerbation risk in asthma) using curated clinical variables. Their strength lies in calibrated, endpoint-specific prediction, but they are not designed to generalize across outcomes or reveal latent disease structure—a key limitation when working with rare diseases whose clinical subtypes may not yet be well understood.
From a statistical perspective, traditional risk scores assume a direct supervised mapping Y∣X ~ Model(β⊤X), where X denotes the high-dimensional matrix of observed EHR features, β the corresponding parameter vector, and Y a single clinical endpoint that must be explicitly observed. This formulation restricts inputs to hand- or model-selected features and depends entirely on labeled observations, leaving unlabeled data unused and capturing only a fraction of the available clinical information45. In contrast, WEST first learns a low-dimensional latent representation D = f(X) from the full EHR using weakly supervised probabilistic labels. This deep representation summarizes salient clinical information in X across a broader patient population, reducing dimensionality and improving statistical efficiency for downstream applications46. The learned representation can then support a variety of downstream tasks. For example, it can be clustered to produce discrete disease states S = Cluster(D), which capture underlying disease heterogeneity and whose differences can be evaluated using simple supervised models. Once f is learned, D and S act as latent disease descriptors that are more stable, expressive, and clinically informative than the raw feature space X, enabling the exploration of disease subtypes and progression patterns that extend beyond what traditional risk scores can reveal.
This paradigm also resonates with ideas from domain adaptation and cross-domain representation learning, which demonstrate that mapping heterogeneous datasets into a shared latent space can yield representations that generalize across settings even when their observed feature distributions differ substantially74. Likewise, recent work in cross-domain few-shot learning shows that projecting data from different domains into a common latent space—and then learning a classifier using only a small number of labeled target examples—supports effective adaptation under limited supervision75,76. Although WEST does not perform cross-institutional or cross-phenotype transfer in this study, these conceptual parallels suggest that latent representation learning may offer a promising foundation for future cross-site and cross-disease generalization efforts. Such extensions could further support the development of clinical decision support tools and facilitate discovery for rare disease phenotypes whose manifestations are not yet fully characterized.
A primary contribution of WEST is therefore its ability to learn an outcome-agnostic embedding space that supports a broad range of downstream analyses. Learned representations enable exploratory subphenotyping without predefined outcome labels—unlike traditional prognostic scores—and can be paired with supervised models to construct risk scores if desired. Accordingly, we view the subphenotyping results as hypothesis-generating rather than replacements for validated clinical risk tools, while emphasizing that WEST offers a complementary and more general framework for profiling patient state in settings where labeled data are limited, disease courses are heterogeneous, and multiple clinical outcomes are of interest.
Several opportunities remain to extend and generalize this work. While our evaluation was retrospective and conducted within a single health system, future applications across diverse care settings and patient populations will be essential to assess robustness and broader generalizability. Because WEST learns a shared latent representation rather than task-specific features, the framework is naturally compatible with multi-site deployment and transfer learning. Although we have not yet evaluated WEST across institutions, the conceptual foundations outlined above suggest that the learned embeddings may transfer well to new settings once appropriate validation cohorts are available. Further, in its current form, WEST focuses on single-phenotype prediction. Extending the framework to multitask or cross-disease learning represents an important next step, enabling the model to leverage shared structure across related conditions and supporting cross-phenotype transfer in settings where certain diseases are too rare or poorly characterized to support standalone model training. Such extensions may be particularly powerful for accelerating discovery in rare diseases whose clinical manifestations and subtype structure are not yet fully understood. Likewise, validating both the identified phenotypes and subphenotypes in external cohorts will be critical for confirming reproducibility, characterizing cross-population stability, and assessing clinical relevance. Finally, moving from retrospective evaluation toward prospective validation and clinician-in-the-loop testing will be essential for understanding WEST’s practical utility, interpretability, and integration into real-world clinical workflows. Together, this study provides the foundational framework, empirical benchmarks, and data infrastructure necessary to support these future directions—marking a first phase toward generalizable, clinically actionable, and label-efficient transformer models for digital health.
In summary, this study provides evidence that weak supervision, when combined with transformer-based modeling, can support data-efficient learning from EHR data when high-quality labels are limited. By integrating a small set of expert-validated annotations with iteratively refined probabilistic supervision, WEST demonstrates improved diagnostic performance for PH and severe asthma cohorts at Boston Children’s Hospital. The framework also identifies patient subgroups that align with clinically meaningful sources of heterogeneity not readily captured by codified diagnoses alone. Together, these results suggest that weakly supervised transformers can move beyond traditional rule-based phenotyping by learning patient representations that reflect disease-related structure and temporal patterns in real-world EHR data.
In addition, WEST reduces reliance on extensive manual annotation while maintaining competitive performance relative to existing ML and DL baselines. By leveraging limited expert input to refine large-scale probabilistic labels, the framework facilitates more scalable and resource-efficient use of multimodal EHR data. Beyond phenotype identification, the learned representations show potential utility for downstream tasks such as subphenotyping, risk modeling, and trajectory analysis, offering a complementary alternative to single-endpoint clinical risk scores. More broadly, WEST illustrates a label-efficient modeling paradigm that may be applicable to diseases that are rare, heterogeneous, or imprecisely coded, and highlights opportunities for integrating weakly supervised representation learning into digital health research and data curation workflows.
Methods
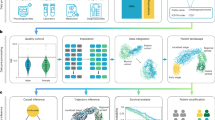
Our end-to-end WEST framework integrates representation learning with weak supervision and iterative label refinement to enable data-efficient modeling of patient state from EHR data. We first identify a high-risk patient cohort and assign initial phenotypic labels using gold- or silver-standard sources (Section “Cohort identification and labeling”). Each patient’s longitudinal clinical history is then transformed into a structured input sequence through a multi-step pre-processing pipeline that includes event aggregation, feature selection, and frequency encoding (Section “EHR sequence pre-processing”). These inputs are processed by a multi-layer transformer encoder that models dependencies among clinical concepts (Section “Transformer encoder”). We then aggregate concept-level embeddings to generate patient-level representations, apply a classification head, and iteratively refine the silver-standard labels through weak supervision (Section “Feature pooling and fine-tuning”). The framework outputs both a patient-level phenotype prediction and a low-dimensional embedding suitable for clustering and visualization. An overview of the pipeline is shown in Fig. 9.
The schematic illustrates the end-to-end workflow of WEST. (1) Cohort identification and labeling assign gold-standard (expert-validated) and silver-standard (probabilistic) labels. (2) EHR sequence pre-processing converts longitudinal structured and unstructured data into aggregated concept sequences with frequency encoding. (3) A transformer encoder models dependencies among clinical concepts. (4) Feature pooling and fine-tuning generate patient-level phenotype predictions and low-dimensional embeddings for subphenotyping. Figure created using Canva.
Cohort identification and labeling
We first define a high-risk patient cohort including individuals whose EHRs show clinical evidence suggestive of the target disease or of related conditions that confer elevated risk. For each disease-specific task, we designate a target diagnostic code or concept c*, which serves as an anchor for identifying relevant features and guiding the label refinement process.
Let i = 1, …, N index all patients in the high-risk cohort. Each patient i is assigned a label yi reflecting their phenotype status. Based on the source and reliability of the label, patients are stratified into two cohorts:
-
(1)
Gold-standard cohort: patients whose disease status has been confirmed through expert physician chart review or inclusion in a disease registry. These patients are assigned gold-standard labels, denoted \({y}_{i}^{gold}\), which serve as high-fidelity references for model training and evaluation. We allow this set to be small to ensure that the WEST pipeline is label-efficient.
-
(2)
Silver-standard cohort: patients with possible but unconfirmed diagnoses. These patients are assigned silver-standard labels, denoted \({y}_{i}^{silver}\), inferred from the EHR data. Silver-standard labels can be defined using rule-based heuristics—such as exceeding a threshold number of occurrences of c*—or derived from the probabilistic predictions of unsupervised automated phenotyping algorithms such as KOMAP36. While these criteria expand the size of the labeled dataset, silver-standard labels are inherently noisier and require iterative refinement.
The full set of training labels {yi} is drawn from both cohorts and defined as:
A central component of our framework is the iterative refinement of silver-standard labels. Unlike gold-standard labels, which remain fixed, silver-standard labels are dynamically updated during model training. After each training round, the model generates updated predictions for the silver-standard cohort, and these predicted probabilities replace the previous labels. Each round consists of training for multiple epochs—with the number of epochs treated as a tunable hyperparameter—using early stopping, followed by cross-validated evaluation to assess model performance. The silver-standard labels are then updated based on the model’s predicted probabilities, and the model is retrained using these refined labels. In this study, we performed up to three such iterative updates, stopping when the cross-validated AUC did not improve in the subsequent round to avoid overfitting to the silver-standard labels. This weakly supervised process progressively improves both label quality and model calibration, leveraging the scale and diversity of real-world EHR data to achieve more accurate phenotype classification.
EHR sequence pre-processing
We transform each patient’s raw EHR into a structured representation suitable for transformer-based learning. This pre-processing pipeline comprises three key stages: (1) sequential representation of clinical histories, (2) label-aware augmentation for gold-standard patients, and (3) construction of input embeddings via feature selection and frequency encoding.
Sequential representation of EHR data
For each patient i, the EHR is modeled as a temporal sequence of clinical events partitioned into discrete time windows. These windows reflect clinically meaningful periods such as visits, months, or hospitalization episodes. Let the patient sequence be:
where T is the number of observed time windows. Each window \({{\mathcal{V}}}_{t}\) contains a set of documented medical concepts and their associated occurrence counts:
where ctk denotes a medical concept and ntk the number of times it was recorded in window \({{\mathcal{V}}}_{t}\). The number of concepts Kt may vary across windows and patients.
Label-aware augmentation for gold-standard patients
To enhance generalization and enable effective learning from high-quality labeled examples, we apply two augmentation strategies to the gold-standard cohort: oversampling and dynamic temporal truncation. These methods address class imbalance between gold- and silver-standard cohorts and introduce variability into training.
First, we account for the limited size of the gold-standard cohort by oversampling. Each gold-standard patient is replicated r times in the training data, ensuring that high-confidence examples are adequately represented and not diluted by the larger, noisier silver cohort. This increases the frequency with which the model encounters trusted labels during training, reinforcing supervision from reliable examples. To determine r, we examine the distribution of silver-standard labels and oversample the gold-standard cases until the resulting label distribution matches the expected prevalence of the target disease within the at-risk cohort. This approach ensures that the effective proportion of gold-standard examples remains clinically realistic while preventing them from being overwhelmed by noisy silver-standard examples, thereby stabilizing training and improving model calibration.
Second, we apply temporal truncation to simulate the incompleteness and variability typical of real-world EHRs. During each training iteration, for a patient sequence \({\mathcal{P}}=\{{{\mathcal{V}}}_{1},\ldots ,{{\mathcal{V}}}_{T}\}\), we randomly sample a start and end index, tstart and tend, such that 1 ≤ tstart ≤ tend ≤ T. The truncated sequence is defined as:
This exposes the model to a variety of partial clinical trajectories—some early, some late—mimicking patients presenting at different disease stages or lacking complete documentation. Over time, this dynamic sampling increases the diversity of training examples derived from a fixed gold-standard set and improves robustness to temporal variability in real-world EHR data. By balancing dynamically truncated gold-standard examples with the larger silver-standard set and using cross-validation with early stopping to halt training before memorization occurs, these strategies collectively prevent overfitting to the augmented gold-standard cases.
Feature engineering and embedding construction
To prepare each sequence \({\mathcal{P}}\) or its truncated version \({\mathcal{P}}{\prime}\) as input to the transformer, we construct a structured representation through several pre-processing steps. Let \({\mathcal{C}}=\{({c}_{1},{n}_{1}),({c}_{2},{n}_{2}),...,({c}_{K},{n}_{K})\}\) denote the set of unique concepts and their cumulative counts across a patient’s selected time period, whether from \({\mathcal{P}}\) or \({\mathcal{P}}{\prime}\). Each concept \({c}_{k}\in {\mathcal{C}}\) is mapped to a vector representation ek using a pre-trained embedding model (PEM) for clinical concepts such as SapBERT77, CODER78, MUGS60, or ONCE36:
Since the transformer model operates in a hidden space of dimension dmodel, we project each embedding into this space via a learnable linear transformation:
where \({{\bf{W}}}^{proj}\in {{\mathbb{R}}}^{{d}_{model}\times {d}_{input}}\) and \({{\bf{b}}}^{\,{\rm{p}}{\rm{r}}{\rm{o}}{\rm{j}}}\in {{\rm{{\mathbb{R}}}}}^{{d}_{{\rm{m}}{\rm{o}}{\rm{d}}{\rm{e}}{\rm{l}}}}\) are learnable parameters.
Given the potentially large number of unique concepts in \({\mathcal{C}}\), we perform feature selection to retain only those most relevant to the target condition. This serves two purposes: (1) reducing noise from unrelated concepts, and (2) lowering computational burden, since transformer attention scales quadratically with the number of input tokens79. To identify relevant features, we compute the cosine similarity between the embedding of each concept and that of the target concept c*, representing the disease condition of interest:
where ek and e* are the respective embeddings. The top K* concepts with the highest similarity scores are retained:
The target c* is always included to ensure phenotype-specific information is preserved. Each nk denotes the total count of concept ck across all relevant time windows.
At this stage, we have constructed an aggregated set \({{\mathcal{C}}}^{* }\) comprising unique clinical concepts and their corresponding cumulative frequencies, which summarize a patient’s longitudinal medical history. To encode concept frequency—serving as a proxy for clinical significance, capturing aspects such as chronicity or ongoing management—we introduce a frequency-based embedding mechanism. Each patient-specific cumulative count nki for concept ck is projected into the model’s embedding space through a two-layer feedforward network with a Swish-Gated Linear Unit (SwiGLU) activation function, a gated variant of the linear unit shown to improve expressivity and training stability in transformer feedforward layers80:
with learnable parameters:
Unlike traditional positional encodings used in NLP, this representation is grounded in concept frequency rather than token order, offering a tailored signal for clinical models sensitive to the recurrence and persistence of medical events. The final representation of each selected concept is obtained by summing its embedding and patient-specific frequency encoding:
Here, zki is the input token for concept ck for patient i to the transformer. If concept ck is not observed for patient i, we set zki = 0. This formulation allows the model to simultaneously capture semantic similarity across medical concepts and their implicit clinical significance based on frequency. The final patient sequence is:
Transformer encoder
Our model builds on a multi-layer transformer encoder but adapts it for the challenges of weakly supervised phenotyping. The encoder serves two purposes simultaneously: (1) patient-level classification, where the model predicts the probability that a patient has the target condition, and (2) representation learning, where it generates low-dimensional embeddings useful for clustering and visualization.
Each patient sequence Zi is processed through stacked transformer encoder layers. Within each layer, multi-head self-attention models dependencies among medical concepts, enabling the network to focus on the parts of the record most informative for the target disease. Standard architectural elements—including residual connections, layer normalization, and feedforward networks with nonlinear activations—are incorporated to ensure stable training. Full mathematical details are provided in Section S1 of the Supplementary Materials, which describe the multi-layer transformer architecture employed by WEST. The derivations clarify the inner workings of the transformer encoder, including its attention mechanism, projection layers, and feedforward components.
Feature pooling and fine-tuning
After passing through multiple transformer layers, the sequence of contextualized embeddings is aggregated into a fixed-length patient representation using mean pooling:
This approach allows the model to capture contributions from all medical concepts while accommodating sequences of varying lengths. The pooled patient representation xi is passed through a classification head—a linear layer followed by a sigmoid activation—to produce a probability score:
where Wclass and bclass are learnable parameters. The sigmoid function σ( ⋅ ) maps the logit to a probability in the range (0, 1). Model training employs binary cross-entropy (BCE) loss, which provides a well-calibrated probabilistic objective for binary classification and naturally accommodates the soft, probabilistic silver-standard labels used in our weakly supervised setting. After each training round, the best-performing model on the validation set is used to update silver-standard labels using its predicted probabilities:
This iterative label refinement allows the model to incorporate its own predictions, progressively improving phenotype classification over training cycles.
Hyperparameter tuning
We performed hyperparameter tuning using a two-fold cross-validation procedure to robustly select model configurations. For each hyperparameter setting, the model was trained on one fold and evaluated on the other. A random search strategy was employed to explore the following hyperparameter space: batch size ∈ {64, 128, 256}, learning rate ∈ {5e−4, 1e−3, 2e−3}, hidden dimension ∈ {32, 64, 128}, number of transformer layers ∈ {2, 3, 4}, dropout rate ∈ {0.3, 0.7}, and number of training epochs ∈ {30, 50}. AUC served as the primary selection metric, and the chosen hyperparameters for each fold were subsequently used to train the final models.
Implementation
We implement WEST in Python 3.12.11 using PyTorch for model development and scikit-learn for evaluation. The WEST pipeline automates cohort pre-processing, hyperparameter optimization, cross-validation-based model selection, model evaluation, and iterative silver-label refinement across training rounds. Models are trained on a single NVIDIA GPU (48–80 GB VRAM) using early stopping based on validation performance. Each training run requires approximately 2–4 h per fold, and a complete two-round pipeline, including hyperparameter search, evaluation, and label updates, completes within 16–20 h. The implementation supports both interactive execution and parallelized SLURM submission, with independent GPU jobs per cross-validation fold to maximize utilization.
Data availability
The EHR data used in this study were obtained from Boston Children's Hospital and contain protected health information that cannot be shared publicly due to patient privacy regulations and institutional data use agreements. Access to these data is therefore restricted and cannot be distributed outside the institution. Derived, de-identified summary results supporting the findings of this study are available from the corresponding author upon reasonable request and subject to institutional approval.
Code availability
The complete codebase, data-processing utilities, and training scripts implementing the WEST framework are openly available at https://github.com/kfgreco/WEST.
References
Health, T. L. G. The landscape for rare diseases in 2024. Lancet Glob. Health 12, e341 (2024).
Wang, C. M. et al. Operational description of rare diseases: a reference to improve the recognition and visibility of rare diseases. Orphanet J. Rare Dis. 19, 334 (2024).
Marwaha, S., Knowles, J. W. & Ashley, E. A. A guide for the diagnosis of rare and undiagnosed disease: beyond the exome. Genome Med. 14, 23 (2022).
Boulanger, V., Schlemmer, M., Rossov, S., Seebald, A. & Gavin, P. Establishing patient registries for rare diseases: rationale and challenges. Pharm. Med. 34, 185–190 (2020).
Mak, C. M. et al. Computer-assisted patient identification tool in inborn errors of metabolism–potential for rare disease patient registry and big data analysis. Clin. Chim. Acta 561, 119811 (2024).
Rubinstein, Y. R. et al. The case for open science: rare diseases. JAMIA Open 3, 472–486 (2020).
Bauskis, A., Strange, C., Molster, C. & Fisher, C. The diagnostic odyssey: insights from parents of children living with an undiagnosed condition. Orphanet J. Rare Dis. 17, 233 (2022).
Stoller, J. K. The challenge of rare diseases. Chest 153, 1309–1314 (2018).
Sreih, A. G. et al. Diagnostic delays in vasculitis and factors associated with time to diagnosis. Orphanet J. Rare Dis. 16, 1–8 (2021).
Gunne, E. et al. A retrospective review of the contribution of rare diseases to paediatric mortality in ireland. Orphanet J. Rare Dis. 15, 1–8 (2020).
Mazzucato, M. et al. Estimating mortality in rare diseases using a population-based registry, 2002 through 2019. Orphanet J. Rare Dis. 18, 362 (2023).
eClinicalMedicine. Raising the voice for rare diseases: under the spotlight for equity. EClinicalMedicine 57, 101941 (2023).
Gherasim, A., Dao, A. & Bernstein, J. A. Confounders of severe asthma: diagnoses to consider when asthma symptoms persist despite optimal therapy. World Allergy Organ. J. 11, 1–11 (2018).
Kavanagh, J., Jackson, D. J. & Kent, B. D. Over-and under-diagnosis in asthma. Breathe 15, e20–e27 (2019).
Ruopp, N. F. & Cockrill, B. A. Diagnosis and treatment of pulmonary arterial hypertension: a review. JAMA 327, 1379–1391 (2022).
Galiè, N. et al. 2015 ESC/ERS guidelines for the diagnosis and treatment of pulmonary hypertension: the Joint Task Force for the Diagnosis and Treatment of Pulmonary Hypertension of the European Society of Cardiology (ESC) and the European Respiratory Society (ERS): endorsed by: Association for European Paediatric and Congenital Cardiology (AEPC), International Society for Heart and Lung Transplantation (ISHLT). Eur. Heart J. 37, 67–119 (2016).
Humbert, M. et al. 2022 ESC/ERS guidelines for the diagnosis and treatment of pulmonary hypertension: developed by the Task Force for the Diagnosis and Treatment of Pulmonary Hypertension of the European Society of Cardiology (ESC) and The European Respiratory Society (ERS). Endorsed by the International Society for Heart and Lung Transplantation (ISHLT) and the European Reference Network on Rare Respiratory Diseases (ERN-LUNG). Eur. Heart J. 43, 3618–3731 (2022).
Brown, L. M. et al. Delay in recognition of pulmonary arterial hypertension: factors identified from the reveal registry. Chest 140, 19–26 (2011).
Chung, K. F. Diagnosis and management of severe asthma. In Seminars in Respiratory and Critical Care Medicine (ed. O’Byrne, P. M.), Vol. 39, 091–099 (Thieme Medical Publishers, 2018).
Sheikh, A. et al. Difficult-to-treat and severe asthma in adolescents and adult patients: diagnosis and management. Global Initiative for Asthma (GINA) (2018).
Levy, M. L. et al. Why asthma still kills: the National Review of Asthma Deaths (NRAD) confidential enquiry report. Royal College of Physicians, (London, 2014).
D’Agnolo, H. M. et al. Creating an effective clinical registry for rare diseases. United European Gastroenterol. J. 4, 333–338 (2016).
Gliklich, R., Dreyer, N. & Leavy, M. Registries for Evaluating Patient Outcomes: A User’s Guide 3rd edn (Agency for Healthcare Research and Quality (US), 2014).
Hageman, I. C., van Rooij, I. A., de Blaauw, I., Trajanovska, M. & King, S. K. A systematic overview of rare disease patient registries: challenges in design, quality management, and maintenance. Orphanet J. Rare Dis. 18, 106 (2023).
Garcelon, N., Burgun, A., Salomon, R. & Neuraz, A. Electronic health records for the diagnosis of rare diseases. Kidney Int. 97, 676–686 (2020).
Ahmad, S. G. et al. IoT based smart wearable belt for tracking fetal kicks and movements in expectant mothers. IEEE Sensors J. 25, 27322–27333 (2025).
Hassan, A. et al. Enhanced model for gestational diabetes mellitus prediction using a fusion technique of multiple algorithms with explainability. Int. J. Comput. Intell. Syst. 18, 1–33 (2025).
Hassan, A., Nawaz, S., Tahira, S. & Ahmed, A. Preterm birth prediction using an explainable machine learning approach. Artif. Intell. Appl. 0, 1–14 (2025).
Hassan, A. & Ahmed, A. Predicting Parkinson’s disease progression: a non-invasive method leveraging voice inputs. Comput. Sci. 8, 66–82 (2023).
Walsh, S. L., Calandriello, L., Silva, M. & Sverzellati, N. Deep learning for classifying fibrotic lung disease on high-resolution computed tomography: a case-cohort study. Lancet Respir. Med. 6, 837–845 (2018).
Huang, P. et al. Deep machine learning predicts cancer risk in follow-up lung screening. Lancet Digit. Health 1, e353–e362 (2019).
Kaplan, A. et al. Artificial intelligence/machine learning in respiratory medicine and potential role in asthma and copd diagnosis. J. Allergy Clin. Immunol. Pract. 9, 2255–2261 (2021).
Geva, A. et al. A computable phenotype improves cohort ascertainment in a pediatric pulmonary hypertension registry. J. Pediatr. 188, 224–231 (2017).
Shivade, C. et al. A review of approaches to identifying patient phenotype cohorts using electronic health records. J. Am. Med. Inform. Assoc. 21, 221–230 (2014).
Alzoubi, H. et al. A review of automatic phenotyping approaches using electronic health records. Electronics 8, 1235 (2019).
Xiong, X. et al. Knowledge-driven online multimodal automated phenotyping system. medRxiv https://doi.org/10.1101/2023.09.29.23296239 (2023).
Yu, S. et al. Enabling phenotypic big data with phenorm. J. Am. Med. Inform. Assoc. 25, 54–60 (2018).
Banda, J. M., Halpern, Y., Sontag, D. & Shah, N. H. Electronic phenotyping with aphrodite and the Observational Health Sciences and Informatics (OHDSI) data network. AMIA Summits Transl. Sci. Proc. 2017, 48 (2017).
Nogues, I.-E. et al. Weakly semi-supervised phenotyping using electronic health records. J. Biomed. Inform. 134, 104175 (2022).
Yang, S., Varghese, P., Stephenson, E., Tu, K. & Gronsbell, J. Machine learning approaches for electronic health records phenotyping: a methodical review. J. Am. Med. Inform. Assoc. 30, 367–381 (2023).
Banda, J. M., Seneviratne, M., Hernandez-Boussard, T. & Shah, N. H. Advances in electronic phenotyping: from rule-based definitions to machine learning models. Annu. Rev. Biomed. Data Sci. 1, 53–68 (2018).
Callahan, T. J. et al. Characterizing patient representations for computational phenotyping. In AMIA Annual Symposium Proceedings, Vol. 2022, 319-328 (American Medical Informatics Association, Washington, DC USA, 2023).
Choi, E. et al. Multi-layer representation learning for medical concepts. In Proc. 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, 1495–1504 (Association for Computing Machinery, New York, NY USA, 2016).
Weng, W.-H. & Szolovits, P. Representation learning for electronic health records. Preprint at arXiv https://doi.org/10.48550/arXiv.1909.09248 (2019).
Rajkomar, A. et al. Scalable and accurate deep learning with electronic health records. NPJ Digital Med. 1, 18 (2018).
Bengio, Y., Courville, A. & Vincent, P. Representation learning: a review and new perspectives. IEEE Trans. Pattern Anal. Mach. Intell. 35, 1798–1828 (2013).
Fridgeirsson, E. A., Sontag, D. & Rijnbeek, P. Attention-based neural networks for clinical prediction modelling on electronic health records. BMC Med. Res. Methodol. 23, 285 (2023).
Banerjee, J. et al. Machine learning in rare disease. Nat. Methods 20, 803–814 (2023).
Schaefer, J., Lehne, M., Schepers, J., Prasser, F. & Thun, S. The use of machine learning in rare diseases: a scoping review. Orphanet J. Rare Dis. 15, 1–10 (2020).
Yang, J., Triendl, H., Soltan, A. A., Prakash, M. & Clifton, D. A. Addressing label noise for electronic health records: insights from computer vision for tabular data. BMC Med. Inform. Decis. Mak. 24, 183 (2024).
Wu, H., Yamal, J. M., Yaseen, A. & Maroufy, V. Statistics and Machine Learning Methods for EHR Data: From Data Extraction to Data Analytics (CRC Press, 2020).
Mitani, A. A. & Haneuse, S. Small data challenges of studying rare diseases. JAMA Netw. Open 3, e201965–e201965 (2020).
Vaswani, A. et al. Attention is all you need. Adv. Neural Inf. Process. Syst. 30 (2017).
Yang, Z., Mitra, A., Liu, W., Berlowitz, D. & Yu, H. Transformehr: transformer-based encoder-decoder generative model to enhance prediction of disease outcomes using electronic health records. Nat. Commun. 14, 7857 (2023).
Li, Y. et al. Behrt: transformer for electronic health records. Sci. Rep. 10, 7155 (2020).
Rasmy, L., Xiang, Y., Xie, Z., Tao, C. & Zhi, D. Med-bert: pretrained contextualized embeddings on large-scale structured electronic health records for disease prediction. NPJ Digital Med. 4, 86 (2021).
Hirszowicz, O. & Aran, D. ICU bloodstream infection prediction: a transformer-based approach for EHR analysis. In International Conference on Artificial Intelligence in Medicine, 279–292 (Springer, 2024).
Kraljevic, Z. et al. Foresight—a generative pretrained transformer for modelling of patient timelines using electronic health records: a retrospective modelling study. Lancet Digital Health 6, e281–e290 (2024).
Yu, S., Cai, T. & Cai, T. NILE: fast natural language processing for electronic health records. Preprint at arXiv https://doi.org/10.48550/arXiv.1311.6063 (2013).
Li, M. et al. Multisource representation learning for pediatric knowledge extraction from electronic health records. NPJ Digital Med. 7, 319 (2024).
Chen, T. & Guestrin, C. Xgboost: a scalable tree boosting system. In Proc. 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, 785–794 (Association for Computing Machinery, New York, NY USA, 2016).
Maaten, L. vd & Hinton, G. Visualizing data using t-sne. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Ramos, J. et al. Using tf-idf to determine word relevance in document queries. In Proc. First Instructional Conference on Machine Learning, Vol. 242, 29–48 (Citeseer, 2003).
Van der Maaten, L. & Hinton, G. Visualizing data using t-sne. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Amorim, L. D. & Cai, J. Modelling recurrent events: a tutorial for analysis in epidemiology. Int. J. Epidemiol. 44, 324–333 (2015).
Ratner, A. et al. Snorkel: rapid training data creation with weak supervision. VLDB J. 29, 709–730 (2020).
Sohn, K. et al. Fixmatch: simplifying semi-supervised learning with consistency and confidence. Adv. Neural Inf. Process. Syst. 33, 596–608 (2020).
Ren, Z., Wang, S. & Zhang, Y. Weakly supervised machine learning. CAAI Trans. Intell. Technol. 8, 549–580 (2023).
Liao, K. P. et al. High-throughput multimodal automated phenotyping (MAP) with application to phewas. J. Am. Med. Inform. Assoc. 26, 1255–1262 (2019).
Ferté, T. et al. Automatic phenotyping of electronical health record: Phevis algorithm. J. Biomed. Inform. 117, 103746 (2021).
Benza, R. L. et al. Predicting survival in patients with pulmonary arterial hypertension: the reveal risk score calculator 2.0 and comparison with ESC/ERS-based risk assessment strategies. Chest 156, 323–337 (2019).
Bateman, E. D. et al. Development and validation of a novel risk score for asthma exacerbations: the risk score for exacerbations. J. Allergy Clin. Immunol. 135, 1457–1464 (2015).
Hatoun, J., Correa, E. T., MacGinnitie, A. J., Gaffin, J. M. & Vernacchio, L. Development and validation of the asthma exacerbation risk score using claims data. Acad. Pediatr. 22, 47–54 (2022).
Teshima, T., Sato, I. & Sugiyama, M. Few-shot domain adaptation by causal mechanism transfer. In International Conference on Machine Learning, 9458–9469 (PMLR, 2020).
Guan, J., Zhang, M. & Lu, Z. Large-scale cross-domain few-shot learning. In Proc. Asian Conference on Computer Vision (Springer, 2020).
Jing, T., Xia, H., Hamm, J. & Ding, Z. Marginalized augmented few-shot domain adaptation. IEEE Trans. Neural Netw. Learn. Syst. 35, 12459–12469 (2023).
Liu, F., Shareghi, E., Meng, Z., Basaldella, M. & Collier, N. Self-alignment pretraining for biomedical entity representations. Preprint at arXiv https://doi.org/10.48550/arXiv.2010.11784 (2020).
Yuan, Z. et al. Coder: knowledge-infused cross-lingual medical term embedding for term normalization. J. Biomed. Inform. 126, 103983 (2022).
Wu, Z. et al. Token statistics transformer: linear-time attention via variational rate reduction. Preprint at arXiv https://doi.org/10.48550/arXiv.2412.17810 (2024).
Shazeer, N. Glu variants improve transformer. Preprint at arXiv https://doi.org/10.48550/arXiv.2002.05202 (2020).
Acknowledgements
This work was supported in-part by the National Institutes of Health under awards R01LM013614 (NLM), R01HL170151 (NHLBI), and U01TR002623 (NCATS). We also acknowledge support from the PrecisionLink Biobank for Health Discovery at Boston Children’s Hospital.
Author information
Authors and Affiliations
Contributions
K.F.G., Z.Y., and T.C. conceptualized and designed the study. K.F.G. and Z.Y. implemented and benchmarked the WEST framework, and conducted comprehensive analyses of model performance and results. K.F.G. and S.M.S. retrieved and processed the electronic health record data. M.L., H.T., and A.G. contributed to algorithm design and data interpretation. K.D.M. and B.A.R. provided domain expertise, contributed to study design and interpretation of findings, and co-supervised the research together with T.C. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Greco, K.F., Yang, Z., Li, M. et al. A weakly supervised transformer for rare disease diagnosis and subphenotyping from EHRs with pulmonary case studies. npj Digit. Med. 9, 211 (2026). https://doi.org/10.1038/s41746-026-02406-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-026-02406-x