Abstract
Genetically encoded fluorescent biosensors became indispensable tools for biological research, enabling real-time observation of physiological processes in live cells. Recent protein engineering efforts have resulted in the generation of a large variety of fluorescent biosensors for a wide range of biologically relevant processes, from small ions to enzymatic activity and signaling pathways. However, biosensors for imaging sulfate ions, the fourth most abundant physiological anion, in mammalian cells are still lacking. Here, we report the development and characterization of a green fluorescent biosensor for sulfate named Thyone. Thyone, derived through structure-guided design from bright green fluorescent protein mNeonGreen, exhibited a large negative fluorescence response upon subsecond association with sulfate anion with an affinity of 11 mM in mammalian cells. By integrating mutagenesis analyses with molecular dynamics simulations, we elucidated the molecular mechanism of sulfate binding and revealed key amino acid residues responsible for sulfate sensitivity. High anion selectivity and sensitivity of Thyone allowed for imaging of sulfate anion transients mediated by sulfate transporter heterologously expressed in cultured mammalian cells. We believe that Thyone will find a broad application for assaying the sulfate transport in mammalian cells via anion transporters and exchangers.
Similar content being viewed by others
Introduction
Maintaining the homeostasis of physiologically important ions is a fundamental process in all living systems. Methods and techniques for monitoring intracellular ion dynamics in live cells provide invaluable insights into cell physiology and pathology. Genetically encoded fluorescence biosensors are one of the most powerful tools for visualizing physiologically relevant ions in real-time within live cells and organisms1,2. Biosensors for small ions can be categorized into three groups based on their molecular design3. The first class comprises FRET-based biosensors, which utilize a pair of fluorescent proteins (FPs) fused to the N- and C-termini of the ion-binding domain. The second class is based on the allosteric modulation of fluorescence intensity caused by the association of an ion with a binding domain, which is either fused to a circularly permutated FP or inserted into an FP. The third class is represented by ion-sensitive FPs, which contain an ion-binding motif within the protein itself. To date, numerous biosensors have been developed for a variety of small cations and anions, including calcium4,5,6,7,8, potassium9,10, proton11,12, magnesium13, zinc14, copper15, chloride16, phosphate17, etc., which enabled imaging of ion dynamics in live cells. Despite tremendous progress in optical probe engineering, genetically encoded biosensors for sulfate ion imaging in mammalian cells are still lacking18,19.
Sulfate is the fourth most abundant anion in mammals and serves as an important macronutrient, representing a major source of the essential element sulfur. The intracellular concentration of sulfate anion is primarily regulated by sulfate transporters since passive transport of hydrophilic anions through the lipid bilayer does not occur20. One class of membrane proteins that are involved in sulfate uptake in mammalian cells is represented by the SLC26 gene family, which encodes multifunctional anion exchangers that, in addition to sulfate ions, also transport other anionic substrates including chloride, bicarbonate, oxalate, iodide, and formate21. It was found that chondrodysplasias, non-syndromic hearing impairment, Pendred syndrome, and congenital and early onset Mendelian diseases are associated with mutations in human SLC26 genes22,23. For example, single point mutations and single amino acid deletions in the SLC26A2 transporter have been linked to four chondrodysplasias of varying severity depending on the level of residual sulfate transport function compared to wild-type protein24,25. Methods for quantification of sulfate influx in mammalian cells play a crucial role in studying sulfate transporters’ functions in health and disease. Radioactive sulfate uptake assays and electrophysiology techniques are the most widely used methods for the biophysical characterization of sulfate transporters22,24,25,26. However, radioactive sulfate uptake assays do not allow real-time measurements, while electrophysiological methods are laborious, indirect, and have low throughput. Sulfate-sensitive fluorescent probes can overcome the limitations of currently available methods by enabling real-time quantitative measurements of sulfate transport in live cells.
Here we report the development, characterization, and validation of a genetically encoded green fluorescent sulfate indicator, named Thyone. Thyone was derived from bright green FP mNeonGreen using structure-guided mutagenesis. The green fluorescence of Thyone, characterized by excitation/emission maxima at 509/518 nm, was quenched by sulfate ions with apparent Kd of 29 mM and tens of millisecond association kinetics in solution. By employing molecular dynamic simulations and mutagenesis analysis, we reveal that the W167I mutation plays a key role in sulfate sensitivity by enabling better access of the respective anion to the binding pocket located in the vicinity of the chromophore. Thyone preserved its high sensitivity and affinity to sulfate ion when expressed in cultured mammalian cells exhibiting 94% negative fluorescence change with Kd of 11 mM. Side-by-side comparison of Thyone with another recently reported sulfate biosensor SulfOFF-1 (ref. 18), revealed about fourfold higher sensitivity of Thyone to sulfate ions in mammalian cells. We demonstrated the utility of Thyone by imaging sulfate influx and efflux mediated by the Slc26a2 transporter expressed in cultured mammalian cells. We believe that Thyone will find application for studying sulfate transporters’ function and drug screening to identify small molecules that can rescue the sulfate uptake of mutated sulfate transporters.
Results
Development of genetically encoded fluorescent indicators for sulfate
To create an optical sulfate biosensor, we chose bright green FP mNeonGreen as a template, which was shown to have a chloride-binding cavity in close vicinity to the chromophore27. Interestingly, at acidic pH (pH≤5.5), chloride binding increases mNeonGreen fluorescence caused by chromophore deprotonation28. We suggest that remodeling the chloride-binding pocket and hydrogen-bond network near the phenolate group of the chromophore can permit interactions with the larger anions at physiologically relevant pH. Based on analysis of the mNeonGreen crystal structures, we identified four amino acid positions, namely C149/S151/W167/T183 (Fig. 1a, b), which were targeted by site-directed mutagenesis to introduce degenerate NNS codons. The generated gene library was cloned into pBAD-TorA-HisD vector, expressed in E. coli, and subjected to fluorescence-activated cell sorting (FACS) and colony screening to select the mutants with green fluorescence. The TorA sequence targeted the protein of interest expression to the E. coli periplasm, thus facilitating ion selectivity screening in intact bacteria by manipulating ion concentration29 (Fig. 1c). Ion selectivity screening was performed at neutral pH in bacterial suspension by exchanging chloride- and sulfate-containing buffers and measuring fluorescence intensity changes. The screening of several hundreds of fluorescent clones identified 14 variants that exhibited detectable fluorescence changes upon buffer exchange (Supplementary Fig. 1). The anion-sensitive mutants had various combinations of the following amino acid substitutions at the targeted positions, C149C,Y,I,T,Q,S,L, S151S,A,N,G,V, W167W,Y,I,L,F,V, and T183T,S,G, as well as at two additional positions S163S,K and F165F,L, perhaps introduced by amplification errors (Supplementary Fig. 2). Two mutants C149Y/W167Y and C149I/S151A/W167I, exhibiting the highest sensitivity to sulfate ions while preserving high brightness, were further assessed for sensitivity to fluoride (F-), chloride (Cl-), bromide (Br-), iodide (I-), nitrate (NO3-), thiocyanate (SCN-), and sulfate (SO42-) anions in comparison to the parental protein mNeonGreen. Fluorescence of both mutants was quenched by F- < NO3- < SCN- < SO42−, however, the addition of Cl- and Br- anions increased their fluorescence by ~30-60% and ~17-74%, respectively (Fig. 1d). Iodide had only a minor influence on fluorescence intensity. For comparison, mNeonGreen showed less than 10% fluorescence change to all ions except for SCN- and SO42−, which quenched its fluorescence by 14% and 27%, respectively, under screening conditions (Fig. 1d). Due to the higher sensitivity to sulfate ions, the C149I/S151A/W167I mutant was selected for further evaluation as a fluorescent sulfate indicator. We named the selected variant Thyone, after the mortal mother of Dionysus from Greek mythology, since the pronunciation resembles Greek word for sulfur.
a The selected mutation sites in the mNeonGreen protein (PDB ID: 5LTR). b The immediate chromophore environment in mNeonGreen showing amino acids selected for mutagenesis and the associated H-bonds to chloride ion and chromophore. The chromophore is displayed in yellow stick, and residues are displayed as a green stick with chloride ion in magenta. Labels of residues showed as one letter code, and H-bonds showed as dashed yellow lines with distances in Å. c Schematic depiction of the Thyone development workflow. The selected amino acid positions in mNeonGreen were randomized using site-directed mutagenesis, yielding a mutant library, which was expressed in the periplasm of E. coli bacteria. Generated mutants were selected using FACS based on green fluorescence brightness. After growing colonies on Petri dishes, protein expression and extraction were carried out, and ΔF/F was determined upon the Cl-/SO42- anion exchange (created in BioRender.com). d Selectivity of mNeonGreen, mNG-C149Y/W167Y, and Thyone to anions as measured by the change of fluorescence intensity under the addition of 180 mM anions. The potassium salt is used for each anion tested including F-, Cl-, I-, Br-, NO3-, SCN-, SO42- at pH 7.4. Data are collected at the emission from 510 to 600 nm after the exaction at 480 nm.
Characterization of Thyone in vitro
To characterize the spectral and biochemical properties in vitro, we expressed Thyone in E. coli and purified it using the standard metal affinity chromatography method. Absorption and fluorescence spectra of the purified protein solutions were measured in sulfate-free buffer and the sulfate buffer containing 100 mM SO42− at neutral pH. The absorption spectrum of apo-Thyone had two major peaks with maxima at 509 nm and 404 nm, corresponding to deprotonated and protonated forms of the GFP-like chromophore, respectively (Fig. 2a, Table 1). The introduced mutations reduced both the extinction coefficient and quantum yield (QY) of the fluorescent form, thus the molecular brightness of apo-Thyone was ~7-fold lower than that of the parental mNeonGreen (Table 1). Administration of sulfate shifted equilibrium to the protonated form of the chromophore, which correspondingly resulted in quenching green fluorescence by >5.5-fold when excited at 470 nm (Fig. 2a, b). Sulfate binding also reduced quantum yield and fluorescence lifetime of Thyone though less significantly than the extinction coefficient at 509 nm (Table 1). Thyone retained high sensitivity to sulfate with -ΔF/F of ~90% in the pH range from 6.0 to 7.4, which was substantially broadened and shifted to physiologically relevant pH compared to its precursor mNeonGreen (Supplementary Fig. 3). Titration of sulfate ions at neutral pH and constant ionic strength did not affect the fluorescence emission profile and maximum (Fig. 2d). The corresponding normalized ΔF/F curve was fitted well to a single binding site model with affinity (apparent dissociation constant, Kd) of 33 ± 3 mM for sulfate (Fig. 2e). According to these results, the estimated dynamic range of SO42− detection in solution was from ~5 mM to ~100 mM. To characterize sulfate binding kinetics, fluorescence quenching was measured at [SO42−] = 100 mM using stopped-flow fluorimetry. The bi-exponential association curve was characterized by tens of millisecond fast component demonstrating 50% fluorescence quenching in about 20 ms (Fig. 2f). The slow component accounted for less than 25% of maximal fluorescence change and occurred on the second time scale. Overall, these results demonstrated that sulfate binding quenched Thyone fluorescence on a subsecond time scale by stabilizing the protonated state of the chromophore as indicated by a shift in the pKa from 7.2 to 8.6 and decreasing QY of the deprotonated form of the chromophore.
a Absorption spectra of Thyone in the presence (100 mM; red line) and absence (black line) of sulfate ions. Data are normalized to the absorbance peak at 509 nm in the sulfate-free condition. b Fluorescence emission spectra of Thyone in the presence (100 mM; red line) and absence (black line) of sulfate ions excited at 470 nm. Data are normalized to the absorbance peak at 518 nm in the sulfate-free condition. c Fluorescence intensity of Thyone in SO42−-bound (100 mM; red line) and SO42−-free (0 mM; black line) states and fluorescence changes as a function of pH (n = 3 independent experiments; data points, means; error bars, SD). d The fluorescence spectra of Thyone at different sulfate concentrations at pH = 7.4. Spectral data are normalized to the fluorescence intensity peak at 518 nm for sulfate-free conditions. e Plot of normalized -∆F/F against different sulfate concentrations fitted with DoseResp Model for data shown in panel (d) (n = 2 independent experiments; squares, data points; dashes, mean; red line, fitting curve). f Average fluorescence trace for association kinetics of Thyone at [SO42−] = 100 mM measured by stopped-flow fluorimetry (n = 14 independent experiments; solid line, mean; shaded area, SD).
To assess selectivity, we characterized the affinity of Thyone to different anions compared to its precursor mNeonGreen. The mNG-C149Y/W167Y variant was used as an additional reference to compare the influence of the introduced mutation on anion sensitivity and affinity. We measured fluorescence changes upon titration of the F-, Cl-, Br-, I-, NO3-, HCO3-, SCN-, and SO42− anions in the range from 0 to 180 mM at pH 7.4 and calculated affinities for each anion by fitting to a single binding site model (Supplementary Fig. 4–6). Fluorescence changes induced by the selected anions were consistent with the results obtained in the screening experiments (Fig. 1d and Fig. 3a–c). Thyone showed comparable affinities to Br-, SCN-, and SO42− anions, which were in the range from 22 to 37 mM (Fig. 3d, Supplementary Fig. 6). Importantly, Thyone exhibited negligible fluorescence changes in response to the physiological concentrations of Cl- and HCO3-(Fig. 3d), the most abundant physiological anions, whose concentrations can reach up to 60 mM30 and 33 mM31,32, respectively, in mammalian cells. However, it should be noted that the KHCO3 solution used for Thyone titration was not buffered to the neutral pH due to its high buffer capacity. Therefore, we could not rule out the influence of pH during HCO3- titration as the pH of Thyone solution with HCO3- might be increasing with increased HCO3- concentration (see Methods section for details). In turn, nitrate induced almost twice smaller fluorescence changes (about −50% of ΔF/F) compared to SCN- and SO42−.The sulfate affinity of Thyone was 2.6- and 1.5-fold higher than that of mNeonGreen and mNG-C149Y/W167Y, respectively (Fig. 3d). Overall, Thyone was characterized by higher sensitivity and affinity to sulfate ions compared to mNeonGreen and the mNG-C149Y/W167Y variant while exhibiting high ion selectivity to SO42− vs Cl- and HCO3- thus representing a promising optical tool for sulfate detection and imaging under physiological conditions.
a–c The fluorescence changes of mNeonGreen, mNG-C149Y/W167Y, and Thyone upon titration against selected anions at pH 7.4 (n = 2 independent experiments each; data points, means; error bars, SD). d Kd values of mNeonGreen, mNG-C149Y/W167Y, and Thyone to respective anions calculated based on the data shown in (a–c) (n = 2 two independent experiments each; bar graphs, mean).
Mechanism of sulfate binding
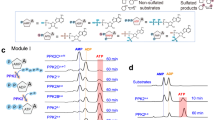
To gain more insight into the molecular mechanism of sulfate binding, we investigated the individual and synergistic contributions of each introduced mutation into sulfate affinity and sensitivity of Thyone. We generated all six possible combinations of single and double mutants and performed their titration with sulfate anion in the range from 0 mM to 180 mM in solution using the parental mNeonGreen protein and another sulfate biosensor SulfOFF-1, also derived from mNeonGreen18, as references. All generated mutants exhibited negative fluorescence responses upon sulfate titration (Fig. 4a). The variants containing the W167I mutation, including Thyone, appeared to be more sensitive to sulfate compared to all other tested proteins, demonstrating more than 83% fluorescence quenching at 180 mM (Fig. 4b). In turn, ΔF/F for SulfOFF-1 was only 63%, which was comparable to the mNeonGreen double mutant C149I/S151A (Fig. 4b). Correspondingly, the sulfate affinity of the W167I-containing mutants was in the range of 40–45 mM, which was at least ~1.7-fold lower than that of other mutants (Fig. 4c). The chloride sensitivity of the generated mutants remained very low (less than 10% of ΔF/F), which was comparable to that of SulfOFF-1 and Thyone (Supplementary Fig. 7). According to these results, W167I is the key mutation that enables sulfate sensitivity and high affinity of Thyone. Furthermore, Thyone outperformed SulfOFF-1 in terms of sulfate sensitivity and affinity.
a The fluorescence changes of mNeonGreen, mNeonGreen mutants, SulfOFF-1, and Thyone to sulfate upon titration against selected anions at pH 7.4 (n = 6 independent experiments for mNG, mNG-C149I, mNG-S151A, mNG-W167I, mNG-C149I/S151A, Thyone each and n = 3 independent experiments for mNG-C149I/W167I, mNG-S151A/W167I, SulfOFF-1; dots, means; error bars, standard deviations). b Fluorescence changes of mNeonGreen, mNeonGreen mutants, SulfOFF-1, and Thyone to sulfate at 180 mM and pH7.4 (n = 6 independent experiments for mNG, mNG-C149I, mNG-S151A, mNG-W167I, mNG-C149I/S151A, Thyone each and n = 3 independent experiments for mNG-C149I/W167I, mNG-S151A/W167I, SulfOFF each; dots, data points; bars, means; error bars, standard deviations). c The corresponding Kd to sulfate of mNeonGreen, mNeonGreen mutants, SulfOFF-1, and Thyone to sulfate calculated based on the data shown in (a) (n = 6 independent experiments for mNG-C149I, mNG-S151A, mNG-W167I, mNG-C149I/S151A, Thyone each and n = 3 independent experiments for mNG, mNG-C149I/W167I, mNG-S151A/W167I, SulfOFF-1 each; dots, data points; bars, means; error bars, standard deviations). d The predicted entry pathways of Thyone through constant pH MD simulations. Pathway 1, outlined in orange: Sulfate initially binds to surface residues K166, K152, and R150. When the sulfate is moved between the proximal β sheets, after 20 ns the sulfate is bound inside close to the original pocket. Pathway 2, outlined in blue: Sulfate initially binds between β sheets 7 and 10 to K152, K153 and R150. When the sulfate is moved between the proximal beta sheets, after 20 ns the sulfate was bound with K153, V203 and H72 partially inside the β barrel. The final bound position of sulfate is shown in the central panel. Hydrogen bonds are shown in cyan dashed lines, with residues shown in gray stick, sulfate shown in colored stick, and the chromophore shown in yellow stick. e Structural changes upon binding of sulfate to Thyone and W167I systems, with the original position of the chromophore shown in yellow stick, its position with sulfate bound shown in gray stick, binding residues shown in green stick, sulfate residue shown in colored stick, and hydrogen bonds shown in dashed cyan. (1) Representative structure of Thyone in transparent green cartoon, with mutated positions are highlighted in pink (I167) and purple (S151 and C149). (2) Zoom of labeled binding pocket of Thyone. (3) The change in chromophore position upon binding of sulfate to Thyone. (4) The beta barrel structure of W167I where mutated position is highlighted in pink (I167). (5) Zoom of labeled binding pocket of W167I. (6) The change in chromophore position upon binding of sulfate to W167I.
To gain atomic-level insights into the mechanisms of sulfate binding, we performed constant pH molecular dynamics (CpHMD) simulations for Thyone and mNeonGreen-W167I proteins. For both systems, we carried out free diffusion and sulfate-bound simulations to determine how sulfate enters the protein barrel and the stability of sulfate binding within the previously identified chloride-binding pocket. To further elucidate the binding pathways of Thyone, 2 short (20 ns) separate surface bound simulations were carried out as well, since the free diffusion replicates indicated two topologically different entry points from the surface of the protein to the binding pocket. (Fig. 4d). The free diffusion simulations indicated that for both systems sulfate primarily binds to Arg150, Lys152, Lys153 and Lys166 on the surface of the β-barrel. The introduced isoleucine at the 167th position in both systems removes the bulky nonpolar side chain of tryptophan, leading to increases in hydrogen bonding of the adjacent β-strands (between Ser168 and Asp147) and a decrease of distance between them, in the order mNeonGreen (mNG) < mNG-W167I < Thyone (Supplementary Tables 1–4). This contraction leads to an opening between β-strands 7 and 10, further leading to specific entry points we observe in the simulations (Fig. 4d, Supplementary Fig. 8). Our results suggest that, consistent with the experimental data, the W167I mutation is the key mutation for promoting sulfate diffusion into the binding pocket.
For the single W167I mutant, we captured the free diffusion of a sulfate ion into the β-barrel in two of the three replicate trajectories. We observed the sulfate in W167I was also initially bound to the protein surface via the sidechain of Arg150, similar to previously reported results for mNG and SulfOFF-1 proteins18. Then, the surface-bound sulfate was guided into the β-barrel by Lys153 and Ser163 located in the vicinity of the anion-binding pocket (Supplementary Fig. 8). Once bound, sulfate was stable in the chloride binding pocket and was found to have hydrogen bonds with His72, Arg205, Ser151, Ser163, Thr183, Tyr185, and Arg98 with high percentage occupancy (Supplementary Fig. 9, Supplementary Table 1, 2). Additionally, upon sulfate binding, we observed a rotation in the chromophore conformation similar to that of mNG as reported by Ong et al.18 (Supplementary Movie 1). The final conformation adopted by the chromophore of W167I was very similar to that of mNG (Supplementary Fig. 9). Normal mode PCA analysis revealed that, upon sulfate binding, the highest contributing vibrational mode of the protein has a drastically lowered mobility and flexibility compared to the apo protein motion, implying stable sulfate binding (Supplementary Fig. 10).
Compared to mNG-W167I, Thyone has two more nonpolar residues introduced to the inside of the β-barrel (Ala151 and Ile149). These mutations created two additional charged areas on the surface of the β-barrel where sulfate binds favorably: Pathway 1 with Lys166, Lys152, and Arg150 sidechains, and Pathway 2 with Lys152, Lys153, and Arg150 sidechains (Fig. 4d). Experimentally, Thyone has a bimodal binding with fast and slow components (Fig. 2f), and we attributed this to the differences between our simulations of the original binding pathway in the single mutation and the new Pathway 1 and Pathway 2. Pathway 1 is faster than Pathway 2, though both require overcoming a small energetic barrier from outside the barrel to diffuse in. However, once the sulfate passes these charged regions, the sulfate rapidly proceeds to the original binding pocket, in less than 20 ns in Pathway 1 and more than 20 ns in Pathway 2 (Fig. 4d). The S151A mutation has caused the other binding pocket residues to have increased hydrogen bonding with the bound sulfate, especially Arg98, Thr183, and Ser163, which gain ~20% more hydrogen bonding percentage occupancy compared to that of W167I (Supplementary Tables 1–4). We observed a similar change in chromophore conformation, indicating that the actual sensing mechanism is likely similar between SulfOFF-1, mNG-W167I, and Thyone, though the final conformation differs slightly (Fig. 4e). The sulfate-bound simulation confirms that sulfate was stable in the chloride binding pocket and makes hydrogen bonds with Arg98, Thr183, Ser163, Tyr185, Lys153, and His72 (Supplementary Tables 1–4). We observe a similar effect on the vibrational modes of the protein as in W167I, although the decrease in mobility was more exaggerated. Taken together, our MD results agreed well with the experimental data for sulfate sensitivity, kinetics and importance of specific point mutations for sulfate binding while revealing specific molecular interactions occurring during sulfate association and binding.
Characterization of Thyone in mammalian cells
The high sulfate sensitivity of Thyone in solution encouraged us to validate its performance as a sulfate biosensor in mammalian cells. While we were finalizing this study, Ong et al. reported SulfOFF-1, another green fluorescent sulfate biosensor, which was also derived from mNeonGreen18. According to the literature data, the SulfOFF-1 biosensor exhibited high affinity and selectivity to sulfate ions in solution18 (Table 1). However, it was not characterized in mammalian cells. Therefore, we sought to compare the sulfate sensitivity of SulfOFF-1 and Thyone in mammalian cells using the parental protein mNeonGreen as an additional control. Upon expression in CHO-K1 cells, all tested FPs produced easily detectable green fluorescence, which was evenly distributed throughout the cytoplasm and nuclei of cells without any signs of aggregation or mislocalization (Fig. 5a). Sulfate ionophore-treated CHO-K1 cells expressing Thyone demonstrated large (ΔF/F of -56%) and reversible fluorescence response to sulfate-containing buffer (Fig. 5b, c; here and below for live cell imaging we used a bath perfusion system for buffer exchange at a flow rate of 1 ml/min, which may result in a slow fluorescence response of the biosensors compared to in vitro measurements reported in Fig. 2f due gradual build-up of the ion concentrations). Under identical conditions, SulfOFF-1 produced only a moderate fluorescence response of less than 12% (Fig. 5b, c). Perfusion of sulfate-containing buffer without ionophore did not result in any fluorescence changes in Thyone-expressing cells (Supplementary Fig. 11). We also observed the dependence of fluorescence changes amplitude on ionophore concentration. Fluorescence responses, matching in amplitude those recorded with the highest tested concentration of ionophore, were obtained when SulfOFF-1 and Thyone were expressed on the extracellular side of the plasma membrane using the pDisplay vector (Fig. 5d,e,f). Importantly, despite a very minor fluorescence response to chloride in solution (Fig. 3c), Thyone did not respond to chloride anion in HEK cells (Fig. 5e). In turn, mNeonGreen expressed on the extracellular membrane did not respond to sulfate or chloride ions (Fig. 5d–f). Despite variations in the baseline brightness of cells due to different expression levels of the indicators, the amplitudes of fluorescence changes were consistent with average standard deviation not exceeding 20% of the relative amplitude values (Fig. 5b, e; see also Supplementary Fig. 12 for single-cell single trial traces). Large fluorescence response and high ion selectivity to sulfate make Thyone a more suitable probe for imaging sulfate dynamics in live mammalian cells compared to SulfOFF-1.
a Representative fluorescence images of live CHO cells expressing untagged SulfOFF-1 (top) and Thyone (bottom) in sulfate-free and sulfate-containing extracellular solution containing sulfate ionophore (n = 61, 112 cells for SulfOFF-1 and Thyone, respectively, from 1 transfection each). Scale bar, 10 μm. b Average single-trial traces of SulfOFF-1 (top) and Thyone (bottom) fluorescence upon adding and washing out sulfate-containing buffer (n = 61, 112 cells for SulfOFF-1 and Thyone, respectively, from 1 transfection each). Solid line, mean; shaded area, s.d. c Quantification of SulfOFF-1 and Thyone fluorescence response for the experiments shown in panel (b) (n = 61, 112 cells from 1 transfection). Box plots with notches are used throughout this paper: narrow part of notch, median; top and bottom of the notch, 95% confidence interval for the median; top and bottom horizontal lines, 25% and 75% percentiles for the data; whiskers extend 1.5 × the interquartile range from the 25th and 75th percentiles; horizontal line, mean; outliers, dots. One-way ANOVA with 95% confidence intervals (alpha = 5%, one-sided) was performed to calculate reported p value. d Representative fluorescence images of live HEK cells expressing mNeonGreen (top), SulfOFF-1 (middle), and Thyone (bottom) on the extracellular surface of the plasma membrane in sulfate-free and sulfate-containing extracellular solution (n = 58, 47, 46 cells for mNeonGreen, SulfOFF-1, and Thyone, respectively, from 2 to 3 transfections each). Scale bars, 10 μm. e Averaged single-trial optical traces of mNeonGreen (top), SulfOFF-1 (middle), and Thyone (bottom) fluorescence upon adding and washing out sulfate- and chloride-containing buffers (n = 58, 47, 46 cells for mNeonGreen, SulfOFF-1, and Thyone, respectively, from 2 to 3 transfection each). Solid line, mean; shaded area, s.d. f Quantification of fluorescence response for mNeonGreen, SulfOFF-1, and Thyone to sulfate anion for the experiments shown in panel (e) (n = 58, 47, 46 cells for mNeonGreen, SulfOFF-1, and Thyone, respectively, from 2 to 3 transfection each). One-way ANOVA with 95% confidence intervals (alpha = 5%, one-sided) was performed to calculate reported p value.
To further characterize sulfate sensitivity, we expressed Thyone in HEK293 cells using pDisplay system, which enabled a faster and more complete ion exchange compared to ionophore-treated cells and performed titration from [SO42−] = 0 to 100 mM (Fig. 6a). Imaging positive cells revealed uniform membrane localization of Thyone with only a minor fraction of intracellular fluorescence (Fig. 6b). Thyone demonstrated a robust response to 5 mM sulfate buffer, reaching almost complete quenching of fluorescence, about -98% of ΔF/F, at 100 mM sulfate buffer (Fig. 6c). Fluorescence quenching was reversible, completely restoring baseline fluorescence in sulfate-free buffer. The fitting fluorescence response curve determined affinity to sulfate at about 11 mM (Fig. 6d, e; however, it should be that due to the distinct environment, the affinity of the sensor on the extracellular membrane may be different from that in the cytoplasm). Furthermore, we observed only a minor decrease in Thyone baseline fluorescence by ~10–20% due to photobleaching during long-term live cell imaging for over 90 min (traces presented in Figs. 5, 6, 7 were corrected for photobleaching; see Methods for details). Altogether, the results demonstrated that Thyone can serve as a reliable fluorescent biosensor for sulfate ions in mammalian cells, enabling long-term recordings.
a Linear map of the expression cassette for expression of Thyone at the extracellular surface of the plasma membrane (top) and schematics of the automated perfusion system used for buffer exchange experiments (created in BioRender.com). b Representative fluorescence images of Thyone-expressing cells in SO42−-free or SO42−-containing buffers (n = 72 cells from 5 independent transfections). Scale bar, 20 μm. c Representative single-trial fluorescence trace of Thyone performed with a series of SO42− concentrations at pH 7.4. d Quantification of fluorescence response to sulfate in cells expressing Thyone for the experiments shown in panel (c) (n = 44 cells from 5 independent transfections). Squares, mean, error bars, SD. e Plot of normalized -∆F/F against different sulfate concentrations fitted using nonlinear fitting (Hill I) for the data shown in panel (d) (n = 44 cells from 5 independent transfections; squares, data points; error bar, SD; red line, fitting curve).
a Representative fluorescence images of CHO cells expression Thyone (top), Thyone/Slc26a2 (middle), and Thyone/Slc26a2 supplemented with DIDS (bottom) in sulfate-free and sulfate-containing extracellular solution (n = 21, 96, 59 cells, respectively, from 2 independent transfections each). Scale bar, 10 μm. b Averaged single-trial traces of Thyone fluorescence in cells expressing only Thyone (top), Thyone/Slc26a2 (middle), and Thyone/Slc26a2 supplemented with DIDS (bottom) upon perfusion of sulfate- and chloride-containing buffers for the cells marked with white dashed lines in panel (a) (n = 21, 96, 59 cells, respectively, from 2 independent transfections each). Solid line, mean; shaded area, s.d. c Quantification of fluorescence response for Thyone, Thyone/Slc26a2, and Thyone/Slc26a2 supplemented with DIDS to sulfate anion for the experiments shown in panel (b) (n = 21, 96, 59 cells, respectively, from 2 independent transfections each). One-way ANOVA with 95% confidence intervals (alpha = 5%, one-sided) was performed to calculate reported p value. d Representative fluorescence images of live HEK cells co-expression Thyone, pHTomato, and Slc26a2 in sulfate-free (top) and sulfate-containing (bottom) extracellular solution (n = 58 cells from 1 transfection). e Averaged single trial traces of pHTomato (top) and Thyone (bottom) fluorescence upon perfusion of sulfate- and chloride-containing buffers (n = 58 cells from 1 transfection). Solid line, mean; shaded area, s.d.
Probing sulfate transport in mammalian cells
The biochemical properties of Thyone provide a unique ability to optically record sulfate ion dynamics in live mammalian cells in real-time, which was not previously realized with any other optical probe for sulfate ions reported to date. To test the applicability of Thyone, we decided to image sulfate ion transients mediated by the Slc26a2 protein, which serves as SO42−/2OH- or SO42−/2Cl- exchanger in cartilage and several epithelia22. The Slc26a2-mediated sulfate fluxes were assayed in cultured mammalian cells by recording green fluorescence of Thyone in real-time under fluorescence microscope. Exposing Slc26a2-expressing cells bathed in Cl-/SO42−-free solution to 2 mM SO42- resulted in a significant reduction of green fluorescence (Fig. 7a, b). Fluorescence drop occurred almost at a steady rate reaching about 28% in ~11 min (Fig. 7c). Fluorescence could be rapidly recovered to the initial level upon replacing sulfate buffer with chloride buffer (Fig. 7b). At the same time, CHO-K1 cells expressing only Thyone did not show any changes in fluorescence under the identical treatment (Fig. 7a, b). Furthermore, the fluorescence intensity decrease was only 5% when Slc26a2-expressing cells were treated with diisothiocyanostilbene-2,2’-disulfonic acid (DIDS), a known inhibitor of anion transporters33. As sulfate influx can be associated with changes in intracellular pH depending on extracellular pH22, it was essential to confirm that the reduction of Thyone fluorescence was not due to intracellular pH changes. To monitor intracellular pH we co-expressed red fluorescent pH sensor, pHTomato34, with Thyone and Slc26a2 and conducted buffer exchange experiments while monitoring green and red fluorescence (Fig. 7d). Consistently with earlier experiments, Thyone demonstrated a reversible fluorescence drop upon SO42−/Cl--free → SO42− → Cl- buffer exchange. However, the red fluorescence of pHTomato remained constant throughout all buffer exchange steps (Fig. 7e). These results indicated that the drop in Thyone fluorescence was not associated with changes in intracellular pH but rather occurred solely due to sulfate influx/efflux.
Discussion
We presented the development, characterization, and application of the first practical genetically encoded green fluorescent sulfate biosensor, Thyone. Our protein engineering effort was inspired by the recent discovery that a bright green FP mNeonGreen exhibits intrinsic sensitivity to some anions at acidic pH, with chloride being the most prominent28. We suggested that remodeling chromophore-protein interactions might shift anion sensitivity to a more physiologically relevant pH range and fine-tune anion specificity. To explore this possibility, we employed structure-guide mutagenesis complemented by high-throughput screening to select mNeonGreen mutants with sensitivity to sulfate ions at neutral pH. Since bacterial culture media contains sodium chloride, which is a crucial component for cell growth (our attempts to remove or reduce its concentration in culture media resulted in poor bacteria growth or even cell death), it was essential to remove traces of chloride ions before anion sensitivity assessment to avoid any screening artifacts. Periplasmic expression, previously utilized for screening other genetically encoded biosensors for calcium6,29, facilitated chloride ion removal from bacteria after protein expression and subsequent buffer exchange in a high-throughput fashion sufficient for screening hundreds of variants in one round. Our screening approach identified the mutant with a sizeable negative fluorescence response characterized by a millimolar affinity to sulfate anion at neutral pH. We suggested that introducing hydrophobic aliphatic amino acids in the close vicinity of the phenolate group of the chromophore shifted chromophore pKa to neutral pH values and perhaps enlarged the anion binding cavity inside the β-barrel, supported by the distances computed from MD simulations (Fig. 1b, Supplementary Table 5). Based on the spectroscopic characterization, a fluorescence decrease upon association with sulfate ions mainly occurs via static quenching (Fig. 2a). The reduction of fluorescence lifetime and associated drop in quantum yield indicate the contribution of the dynamic quenching mechanism, although to a smaller extent than static quenching (Table 1). We proposed that the molecular mechanism of fluorescence quenching in Thyone is similar to that previously reported for phiYFP35. According to the proposed mechanism, oxyanions such as nitrate impede excited state conversion by distorting the antiparallel conformation between the chromophore Y66 phenolate and Y203 phenolic rings, which results in fluorescence quenching35. Therefore, we propose that optimization of the anion binding cavity in mNeonGreen resulted in increased sulfate sensitivity at neutral pH.
While we were finalizing this study, Ong et al. reported another turn-off sulfate biosensor called SulfOFF-1, which was also derived from mNeonGreen by rational mutagenesis of the two β-bulge residues18. Based on the reported properties measured in solution, SulfOFF-1 was characterized by higher anion selectivity, molecular brightness, and sulfate affinity compared to Thyone (Table 1). On the other hand, Thyone demonstrated about ~1.5-fold larger fluorescence response to sulfate, which was coupled with tens of millisecond sulfate association kinetics. Fast dynamics of the Thyone biosensor could be beneficial for characterizing rapid transients of sulfate with minimal artifacts. Despite certain advantages of SulfOFF-1 in solution, such as fivefold higher molecular brightness and significantly higher sulfate affinity, in our hands, SulfOFF-1 had limited performance in mammalian cells mainly due to a narrow fluorescence dynamic range, which was only 20% compared to about 95% for Thyone. The more reliable response of Thyone makes it biosensor of choice for real-time monitoring of sulfate transients in mammalian cells, which we capitalized on to probe sulfate transport mediated by the Slc26a2 transporter in CHO cells. However, it should be noted that both SulfOFF-1 and Thyone are intensiometric biosensors, which cannot readily report the absolute concentration of sulfate ions. From this perspective, FRET-based biosensors for sulfate, such as FLIP-SP, might represent a better option for ratiometric imaging of sulfate concentration19. The FLIP-SP biosensor was used in bacteria, yeasts, and plants, although this biosensor requires further thorough characterization in mammalian cells. Alternatively, the mNeonGreen-derived sulfate biosensors can be engineered into ratiometric biosensors by fusing with spectrally complementary FPs, the strategy previously successfully applied for ratiometric chloride biosensors36,37. Therefore, we propose that the mNeonGreen-derived sulfate biosensors represent a promising platform for developing optimized intensiometric and ratiometric biosensors. For example, as SulfOFF-1 and Thyone do not share common mutations relative to mNeonGreen, one potential approach could be to combine mutations from SulfOFF-1 and Thyone to achieve higher anion selectivity couple with large fluorescence response and rapid association kinetics. Furthermore, the simple molecular design of Thyone, which is a single-FP-based biosensor, should allow for a straightforward combination with other green biosensors using a spatial multiplexing approach38.
The MD results confirm several aspects of Thyone’s unique efficiency. The sensing mechanism and sulfate binding pocket residues are similar between all current and previously tested systems, including SulfOFF-1. Additionally, we confirm that the W167I mutation is key for overcoming the original kinetic barrier and that it alone has similar results to the original simulations for SulfOFF-1. For Thyone itself, we showed that the increase in nonpolar residues within the barrel allows for multiple entry pathways and a particularly fast entry mode via Pathway 1. This qualitatively matches the mix of fast/slow binding kinetics observed experimentally. We also demonstrate consistency with the original hypothesis that the increase in nonpolar residues increases the anionic volume by showing a larger distance between the nonpolar residues and the chromophore for Thyone vs. W167I and the parental mNeonGreen.
To demonstrate the applicability of Thyone in mammalian cells, we visualized sulfate transients in mammalian cells mediated by the Slc26a2 anion exchanger. Our results were in complete agreement with previously reported properties of Slc26a2 anion exchanger characterized in Xenopus oocytes using ion-selective electrodes and whole-cell patch clamp recordings22. We observed sulfate influxes upon incubation of Thyone/Slc26a2-expression cells in chloride-free sulfate buffer (Fig. 7). Based on the amplitude of fluorescence change, intracellular sulfate concentration could reach about 5 mM. Rapid efflux of sulfate ions was triggered by exchanging sulfate for chloride in extracellular buffer, restoring basal concentration of sulfate. Since we did not detect intracellular pH changes during buffer exchange, Slc26a2 performed as SO42−/2Cl− exchanger. As expected, the inhibitor of anion transporters DIDS efficiently blocked sulfate influx (Fig. 7). These results demonstrated the potential applicability of Thyone for studying anion transport by the Slc26 exchangers and may promote establishing drug screening platforms similar to that based on genetically encoded chloride biosensors for cystic fibrosis drug development39,40. However, it should be noted that the sulfate concentration in mammals under physiological conditions is typically lower than 2 mM. For example, sulfate concentration in human serum is in the range from 0.29 to 0.41 mM41, and the intracellular concentration in human bronchial epithelial cells can reach ~1.9 mM in chloride-free medium while under physiological conditions, it is at 0.33 mM42. Therefore, the dynamic range of Thyone, which spans from ~2 mM to ~100 mM (Fig. 5e), might be slightly above the typical sulfate concentrations under physiological conditions. Nevertheless, we believe that Thyone may represent a safer and more straightforward alternative to well-established radioactive sulfate uptake assays and electrophysiology techniques for monitoring sulfate homeostasis in live cells. Moreover, the mechanistic insight into sulfate sensitivity revealed by the MD study and mutagenesis analysis can guide further optimization of Thyone to enhance its sensitivity and selectivity for sulfate anion. For example, one strategy to improve the sulfate affinity of Thyone is to enhance access of the sulfate ions to the binding site by modifying the β-bulge gate as was done in the SulfOFF-1 biosensor18.
Materials and methods
Plasmid construction and site-directed mutagenesis
The mNeonGreen gene was synthesized de novo by Epoch Life Science Inc. based on the sequence reported in the original publication43 and subcloned into pBAD-TorA-HisD and pBAD-HisD vectors. The site-directed library of mNeonGreen was synthesized de novo as gBlocks (Epoch Life Science Inc., China) with degenerated NNS codons at the following amino acid positions 149, 151, 167, and 183. The gBlock was amplified with corresponding primers using PCR and subcloned into the pBAD-TorA-HisD vector. The site-directed mutagenesis of Thyone was performed using overlap extension PCR followed by subcloning into pBAD-HisD vector. The SulfOFF-1 gene was generated by site-directed mutagenesis using mNeonGreen as a template. The SLC26A2 gene in pCMV-SPORT6 vector was purchased from Miaoling Biotech Co., Ltd (China). The NPY-pHTomato plasmid was obtained from Addgene (plasmid #83501). For expression in mammalian cells, the genes of mNeonGreen, SulfOFF-1, and Thyone were cloned into the pDisplay vector (pDisplay-pHuji; Addgene plasmid #61556) and pN1 vector (Invitrogen) following the reagent’s protocol (Hieff Clone Plus One-step cloning kit, YEASEN Biotech), respectively.
PrimeStar Max master mix (Clontech, USA) was used for high-fidelity PCR amplification. Restriction endonucleases were purchased from New England BioLabs (USA) and used according to the manufacturer’s protocols. Ligations were performed using T4 DNA ligase (New England BioLabs, USA) or NovoRec® plus one-step Cloning kit (Novoprotein, China). Small-scale isolation of plasmid DNA was performed with Mini-Prep kits (Qiagen, Germany); large-scale DNA plasmid purification was done with GenElute HP Endotoxin-Free Plasmid Maxiprep Kits (Sigma-Aldrich, USA). Purified plasmids and bacterial colonies were sequenced by Sanger method (Youkang Biotechnology, Hangzhou, China).
The site-directed gene library in the expression vector was electroporated into TOP10 E. coli host cells (Biomed, China) and expressed in LB medium supplemented with ampicillin (0.1 mg/ml) and L-arabinose (0.02% w/v; Sangon Biotech Co., Ltd., China). The bacterial library was sorted using BD FACSMelody™ Cell Sorter (BD Biosciences, USA) using 488 nm excitation and 525/50 nm filter and plated on LB/agar Amp+/Ara+ medium. Green fluorescent colonies were selected under Olympus SZX16 stereomicroscope (Japan) equipped with SPECTRA III light engine (Lumencor, USA) and a color CCD camera (BGIMAGING, China) and grown in duplicated 24 deep-well RB blocks (Invitrogen, CS15124) with 4 ml LB Amp+ for plasmid extraction and LB Amp+/Ara+ for protein expression (overnight at 37 °C and 200 rpm). Overnight culture expressing proteins was palleted and washed three times with chloride and sulfate free buffer, then imaged under stereomicroscope before and after addition of sulfate-containing buffer consisting of 100 mM solution of sodium sulfate or chloride-containing buffer composed of 100 mM solution of sodium chloride.
Protein purification and spectroscopic measurements
For in vitro characterization, selected proteins were expressed in TOP10 (Tsingke, China) in LB medium supplemented with 0.2% w/v L-arabinose overnight at 37°C followed by culturing at 16 °C for 18 h. For data in Fig. 1, the harvested bacteria cells were subjected to 3 cycles of freezing and thawing of bacterial cell pellet at −80 °C followed by protein extraction with B-PER™ (ThermoFisher, USA). For data in Figs. 2 and 3, the harvested bacteria cells were sonicated in the chloride/sulfate-free buffer containing 38 mM sodium gluconate, 100 mM potassium gluconate, 20 mM HEPES at pH 7.4 followed by ultracentrifugation at 18,000 × g for 30 min at 4 °C. The liquid supernatant was incubated with nickel nitrilotriacetic acid (Ni-NTA) agarose resin (YEASEN Biotech, China) for 20 min at 4 °C and then transferred to affinity chromatography column (Beyotime, China). Recombinant proteins were eluted with the buffer containing 400 mM imidazole and dialyzed overnight at 4 °C with the chloride/sulfate-free buffer. The protein solutions were centrifuged at 12,000 rpm for 20 min to discard the potential precipitated protein and collected the fluorescent supernatant.
The absorption spectra were measured by UV-Vis-NIR Spectrophotometer (Shimadzu, Japan) or VarioSkanTM Lux multimode microplate reader (ThermoFischer Scientific, USA) with the emission ranging from 250 to 600 nm. The fluorescence spectra were measured by FS5 Spectrofluorometer (Edinburgh Instruments, UK) with the exaction at 470 nm and the emission ranging from 480 to 700 nm. Extinction coefficients were measured using alkaline denaturation methods, which is well established for FPs44. Briefly, FP solution in PBS was vigorously mixed with 2 M NaOH in a 1:1 ratio, and the absorption spectrum was recorded within 5 min and compared to that in PBS. The absolute fluorescence quantum yield (QY) was directly measured using an FLS1000 spectrometer equipped with an integrating sphere accessory (Edinburgh Instruments, UK) according to the manufacturer’s protocol.
For the pH titration, fluorescence data was collected by Varioskan LUX Plate reader (ThermoFisher Scientific, USA) at the emission ranging from 500 nm to 600 nm with the exaction at 480 nm. The pH buffers were prepared by dissolving potassium gluconate for sulfate-free buffers or potassium sulfate for sulfate-containing buffers in pH buffers ranging from 4 to 10 (Cat#280-4.00 to Cat#280-10.00; Hydrion Buffer Chemvelopes, USA) and subsequent measurements of the final pH using pH meter (Mettler Toledo, China). The protein solution of Thyone and mNeonGreen were added to the corresponding pH buffers at 23-25oC in the absence and presence of 100 mM SO42−
For ion selectivity measurement, protein solutions were titrated with buffers containing potassium fluoride (Aladdin, China), potassium iodide (Macklin, China), potassium bromide (Innochem, China), potassium nitrate (Sinopharm Chemical Reagent, China), potassium thiocyanate (Aladdin, China), potassium bicarbonate (Aladdin, China), and potassium sulfate (Sigma-Aldrich, Germany). Fluorescence was recorded at an increasing ionic concentration and strength by adding the corresponding salt solution (solution of KHCO3 was not pH adjusted to pH=7.4 due to high buffer capacity) from 0 to 180 mM to the protein solution in the sulfate-free buffer (sulfate-free buffer containing 38 mM sodium gluconate, 100 mM potassium gluconate, 20 mM HEPES at pH 7.4). The integrated fluorescence was calculated to determine the Kd using a data analysis function named DoseResp in OriginPro 2019b.
For stopped-flow kinetic measurements, data were collected by a Stopped Flow Spectrometer (SX 20, Applied Photophysics Ltd., UK). Fluorescence traces with excitation at 460 nm with 1 nm bandwidth, and the emission wavelength at 520/10 nm were recorded in 5 sequential repeats set up automatically. The protein solution was diluted with sulfate-free buffer (38 mM sodium gluconate, 20 mM HEPES at pH 7.4) and mixed with an equal volume of 200 mM SO42− buffer (38 mM sodium gluconate, 200 mM potassium sulfate, 20 mM HEPES at pH 7.4) at 24°C. The fluorescence change was recorded for 500 ms, and a total of 1000 time points was collected.
Characterization in mammalian cells
HEK293T (ATCC® ACS-4500™) and CHO-K1 cells (ATCC® CCL-61™) were grown on 12-mm round glass coverslips coated with Matrigel (Millipore, USA) in Dulbecco’s modified Eagle’s medium (Servicebio, China) containing 10% fetal bovine serum (YEASENN Biotech, China) at 37 °C incubators (ESCO Scientific, Singapore). Transient transfection was performed at cell density around 40-50% with 0.5 μg plasmid DNA by liposomal transfection method according to reagent’s protocol (YEASENN Biotech, China). For co-expressing sulfate transporter with Thyone, the ratio of pN1-Thyone and Slc26a2 was 1:3. For co-expressing sulfate transporter and pHTomato, the ratio of pN1-Thyone, pHTomato, and Slc26a2 was 1:1:3. The imaging was performed using Nikon Ti2-E widefield microscope equipped with Spectra III Light Engine (LumenCore, USA), the ORCA-Flash 4.0v3 sCMOS camera (Hamamatsu, USA), and 20x/0.75 objective lens controlled by NIS Elements software using YFP (excitation 515/12 nm, emission 550/40 nm) and RFP (excitation 555/28 nm, 594/40 nm) channel.
The high sulfate buffer refers to the solution containing 100 mM magnesium sulfate, 38 mM sodium gluconate, 20 mM HEPES at pH 7.4. The corresponding concentrations of sulfate buffers were prepared by mixing the high sulfate buffer and the sulfate-free buffer (38 mM sodium gluconate, 100 mM potassium gluconate, 20 mM HEPES at pH 7.4). Sulfate ionophore I, 3-[Bis(3-phenylthioureidomethyl)] benzene (Santa Cruz, USA) was dissolved in ethanol and diluted to SO42−-free buffer at the final concentration of 10 mM, which was determined experimentally in this study (Supplementary Fig. 11). The Thyone sensor calibration in the cytoplasm of mammalian cells was not conducted since long-term (>30 min) imaging of cells treated with sulfate ionophore I was not feasible due to the gradual deterioration of cell conditions with every buffer exchange (perhaps due to plasma membrane disruption). The imaging of cells at different concentrations of sulfate buffers was carried out using the customized perfusion system introduced in our previous work10, which provides consecutive environment changes in the imaging chamber.
For experiments in CHO-K1 co-expressing Slc26a2 or in HEK293T co-expressing pHTomato and Slc26a2 shown in Fig. 7, the modified HEPES-buffered ND96 solution contained 96 mM sodium chloride, 2 mM potassium chloride, 1.8 mM calcium chloride, 1 mM magnesium chloride and 5 mM HEPES at pH 7.5 in liter-size. Although the ND96 solution is commonly used for Xenopus oocyte experiments22, we did not observe any detrimental effect of the used buffers on cultured mammalian cells (no cell shrinking or swelling, as well as no observed cell death). Therefore, we concluded that these buffers might also be appropriate for mammalian cell culture imaging. Cl--free buffer was prepared by replacing chloride with gluconate, which included as follows: 96 mM sodium gluconate, 2 mM potassium gluconate and 5 mM HEPES at pH 7.5 in liter-size. For testing sulfate influx, magnesium sulfate was added to Cl--free buffer to a final concentration of 2 mM. A stock solution of 100 mM diisothiocyanostilbene-2,2’-disulfonic acid (DIDS) (Aladdin, China) was dissolved in DMSO and prepared freshly to a final concentration of 50 μM in Cl--free buffer containing sulfate.
Computational methods
We carried out constant pH MD simulations for both Thyone and W167I systems for two setups: free diffusion, where sulfate ions are free to associate with and enter into the protein system, and bound systems, where a sulfate is placed at the bound position of chloride in the initial crystal structure and the sulfate binding position of SulfOFF-1 as in previous work18. After determining potential entry points from the free diffusion simulations, we performed additional short simulations where the sulfate was pushed into the barrel slightly to determine if the entry points could lead to the sulfate entering into the binding pocket. These simulations were run in triplicate, for a total of 16 trajectories and a total simulation time of 5840 ns.
In order to prepare the mutated systems, we used the published crystal structure of mNeonGreen at pH=4.5 (PDB ID – 5LTP)27 and replaced the mutated positions with relevant residues (Thyone: W167I, C149I, and S151A, single mutation: W167I) using the Dunbrack rotamer library within the ChimeraX molecular visualization program45. Then, we used MolProbity to add hydrogens and optimize hydrogen bonding networks46. For each system, we added 7 sulfates and 7 sodium ions to neutralize the overall charge of the system. After that, we solvated the systems using the TIP3P water model with a water pad of 12 Å from the surface of the protein47. To generate the topology and input coordinates we used the tleap program in AMBER20 (ref. 48). We used the ff14SB force field for the protein, and XFP chromophores for the chromophore49,50. Custom parameters were generated for the sulfate ions where we carried out RESP fitting for the partial charges, and used geometric parameters from the literature51,52. We generated the cpin files and new topology files with new intrinsic radii using the cpinutil.py module within AMBER20, with the Generalized Born implicit solvation model settings. Considering the pH of the solution (pH=7.0) we chose to titrate the residues Asp, Glu and His of all the systems. Specifics for the individual types of simulations are detailed below.
To perform free diffusion simulations, the 7 sulfate ions were placed in different random positions for each replicate. After an initial minimization followed by heating to 300 °K and solvent equilibration in the NPT ensemble (4 ns total), we ran production simulations of Thyone and W167I for 400 ns in triplicate in the NVT ensemble. This is the same procedure with the conditions described under free diffusion simulation in our previous work18. In brief, we used a Langevin thermostat with a friction constant of 5 ps-1, smooth particle mesh Ewald for the Coulombic interactions, a cutoff of 8 Å for the other nonbonded interactions, a salt concentration of 0.1 M, and a constant pH value of 7.0.
To perform sulfate-bound simulations, 1 of the 7 sulfate ions was placed in the original chloride binding pocket in the mNeonGreen (PDB 5LTP) crystal structure. Then, we repeated the same procedure as discussed under free diffusion simulations.
When running surface bound simulations, we chose snapshots from the free diffusion simulation trajectories where 1 of the 7 sulfates was bound to the surface near a potential entry point. We then shifted that sulfate 1–2 Å inward between the closest two beta sheets to the entry point. Then, we followed the same initial equilibration procedure under free diffusion simulations, and ran short 20 ns production simulations.
Data analysis
Data were analyzed using the following software: NIS-Elements Advance Research software (Version 5.21.00 and 5.30.00), Excel (Microsoft Office 365, USA), OriginPro (2019b, OriginLab), Pro-Data SX Software, and Fiji ImageJ 2.9.01/1.53t. For fluorescence signal intensity analysis, regions of interest (ROI), including cells and a neighboring cell-free region (background), were selected either manually or automatically by NIS-Elements Advance Research software. Fluorescence images in Fig. 5a, b, Fig. 5b, and Fig. 7a, due to low fluorescence level under high sulfate concentration were displayed after the noise reduction by NIS-Elements Denoise.ai software. Mean intensity was extracted for each ROI in YFP/RFP channels, with the corresponding background being subtracted. Cells that detached or exhibited significant morphological changes during long-term imaging were excluded from the analysis. Dead cells were also excluded from analysis. The mean values of ratios (F/F0) indicating the normalized intracellular brightness of the fluorescent proteins were reported in all experiments. For calibration in cultured cells, the imaging datasets were corrected for movement before analysis using ImageJ plugin Stabilizer (http://www.cs.cmu.edu/~kangli/code/Image_Stabilizer.html). Fluorescence traces in Figs. 5, 6, and 7 were corrected for photobleaching by fitting baseline fluorescence with exponential function in Origin 2019b software. For computational data analysis, we carried out root mean square deviation (RMSD), root mean square fluctuations (RMSF), hydrogen bonding analysis, and correlation analysis using the cpptraj module in AMBER20 (ref. 53). For the visualization of hydrogen bonding networks of sulfate we used PyMol. To create the movies of free diffusion in W167I and Thyone, and to visualize the normal mode data with NMWiz, we used VMD54.
Statistics and reproducibility
All statistical analyses were performed using OriginPro software. The results are presented as means ± standard deviation (SD). We did not perform statistical tests to determine the sample size. The sample size in each experiment was indicated in the figure legends. All attempts to reproduce the results were successful.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The numerical source data for the graphs and plots for all main text figures and Supplementary Figs. 3-6 are available in Supplementary Data 1. All essential data collected for this study is available from the FigShare repository (https://doi.org/10.6084/m9.figshare.27143766) (ref. 55). The computational inputs used to run minimization, heating, equilibration, and simulations of systems for MD have been deposited to Zenodo, and are available from https://doi.org/10.5281/zenodo.10416670 (ref. 56). In addition, the database includes topology files, initial coordinates of all the systems, the last frames of the sulfate-bound simulations of both Thyone and W167I, and the force field parameters created for sulfate (Supplementary Table 6, 7). A short simulation video of sulfate diffusion in W167I has been deposited as well. All other data used in this study are available upon reasonable request from the corresponding authors. All plasmids and their annotated full-length nucleotide sequences are available from WeKwikGene with the following barcodes: pBAD-HisD-Thyone (https://wekwikgene.wllsb.edu.cn/plasmids/0000363); pBAD-HisD-mNeonGreen (https://wekwikgene.wllsb.edu.cn/plasmids/0000364); pN1-mNeonGreen (https://wekwikgene.wllsb.edu.cn/plasmids/0000365); pDisplay-Thyone (https://wekwikgene.wllsb.edu.cn/plasmids/0000366); pN1-Thyone (https://wekwikgene.wllsb.edu.cn/plasmids/0000367); pDisplay-sulfOFF-1 (https://wekwikgene.wllsb.edu.cn/plasmids/0000370); pCMV-SPORT6-slc26a2 (https://wekwikgene.wllsb.edu.cn/plasmids/0000371); pN1-sulfOFF-1 (https://wekwikgene.wllsb.edu.cn/plasmids/0000372); pBluescriptR-slc26a2 (https://wekwikgene.wllsb.edu.cn/plasmids/0000373). The sequences for Thyone is deposited in the GenBank databases (accession number ON981382).
References
Torres-Ocampo, A. P. & Palmer, A. E. Genetically encoded fluorescent sensors for metals in biology. Curr. Opin. Chem. Biol. 74, 102284 (2023).
Piatkevich, K. D., Murdock, M. H. & Subach, F. V. Advances in engineering and application of optogenetic indicators for neuroscience. Appl. Sci. 9, 562 (2019).
Baek, K., Ji, K., Peng, W., Liyanaarachchi, S. M. & Dodani, S. C. The design and evolution of fluorescent protein-based sensors for monoatomic ions in biology. Protein Eng. Des. Sel. 34, 1–9 (2021).
Chen, T.-W. W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Subach, O. M. et al. Novel genetically encoded bright positive calcium indicator NCaMP7 based on the mneongreen fluorescent protein. Int. J. Mol. Sci. 21, 1644 (2020).
Barykina, N. V. et al. Green fluorescent genetically encoded calcium indicator based on calmodulin/M13-peptide from fungi. PLoS One 12, e0183757 (2017).
Subach, O. M. et al. YTnC2, an improved genetically encoded green calcium indicator based on toadfish troponin C. FEBS Open Bio 13, 2047–2060 (2023).
Subach, O. M. O. M., Barykina, N. V. N. V., Anokhin, K. V. K. V., Piatkevich, K. D. K. D. & Subach, F. V. F. V. Near-infrared genetically encoded positive calcium indicator based on GAF-FP bacterial phytochrome. Int. J. Mol. Sci. 20, 3488 (2019).
Shen, Y. et al. Genetically encoded fluorescent indicators for imaging intracellular potassium ion concentration. Commun. Biol. 2, 1–10 (2019).
Torres Cabán, C. C. et al. Tuning the sensitivity of genetically encoded fluorescent potassium indicators through structure-guided and genome mining strategies. ACS Sens. 7, 1336–1346 (2022).
Bizzarri, R. et al. Development of a novel GFP-based ratiometric excitation and emission pH indicator for intracellular studies. Biophys. J. 90, 3300–3314 (2006).
Shen, Y., Rosendale, M., Campbell, R. E. & Perrais, D. pHuji, a pH-sensitive red fluorescent protein for imaging of exo- and endocytosis. J. Cell Biol. 207, 419–432 (2014).
Pérez, V. et al. MagIC, a genetically encoded fluorescent indicator for monitoring cellular Mg2+ using a non-Förster resonance energy transfer ratiometric imaging approach. 20, 101203 https://doi.org/10.1117/1.JBO.20.10.101203 (2015).
Minckley, T. F. et al. Sub-nanomolar sensitive GZnP3 reveals TRPML1-mediated neuronal Zn2+ signals. Nat. Commun 10, 1–14 (2019).
Liang, G.-T. et al. Enhanced small green fluorescent proteins as a multisensing platform for biosensor development. Front. Bioeng. Biotechnol. 10, 1039317 (2022).
Grimley, J. S. et al. Visualization of synaptic inhibition with an optogenetic sensor developed by cell-free protein engineering automation. J. Neurosci. 33, 16297–16309 (2013).
Gu, H. et al. A novel analytical method for in vivo phosphate tracking. FEBS Lett. 580, 5885–5893 (2006).
Ong, W. S. Y. et al. Rational Design of the β-Bulge Gate in a Green Fluorescent Protein Accelerates the Kinetics of Sulfate Sensing**. Angew. Chemie Int. Ed. e202302304 https://doi.org/10.1002/ANIE.202302304 (2023).
Fatima, U. et al. A non-invasive tool for real-time measurement of sulfate in living cells. Int. J. Mol. Sci. 21, 2572 (2020).
Markovich, D. Physiological roles and regulation of mammalian sulfate transporters. Physiol. Rev. 81, 1499–1533 (2001).
Alper, S. L. & Sharma, A. K. The SLC26 gene family of anion transporters and channels. Mol. Asp. Med. 34, 494–515 (2013).
Ohana, E., Shcheynikov, N., Park, M. & Muallem, S. Solute carrier family 26 member a2 (Slc26a2) protein functions as an electroneutral SO 42-/OH -/Cl - exchanger regulated by extracellular Cl. J. Biol. Chem. 287, 5122–5132 (2012).
Dawson, P. & Markovich, D. Pathogenetics of the Human SLC26 Transporters. Curr. Med. Chem. 12, 385–396 (2012).
Karniski, L. P. Functional expression and cellular distribution of diastrophic dysplasia sulfate transporter (DTDST) gene mutations in HEK cells. Hum. Mol. Genet. 13, 2165–2171 (2004).
Karniski, L. P. Mutations in the diastrophic dysplasia sulfate transporter (DTDST) gene: Correlation between sulfate transport activity and chondrodysplasia phenotype. Hum. Mol. Genet. 10, 1485–1490 (2001).
Girard, J. P., Baekkevold, E. S., Feliu, J., Brandtzaeg, P. & Amalric, F. Molecular cloning and functional analysis of SUT-1, a sulfate transporter from human high endothelial venulles. Proc. Natl Acad. Sci. Usa. 96, 12772–12777 (1999).
Clavel, D. et al. Structural analysis of the bright monomeric yellow-green fluorescent protein mNeonGreen obtained by directed evolution. urn:issn:2059-7983 72, 1298–1307 (2016).
Tutol, J. N., Kam, H. C. & Dodani, S. C. Identification of mNeonGreen as a pH-Dependent, Turn-On Fluorescent Protein Sensor for Chloride. ChemBioChem 20, cbic.201900147 (2019).
Barykina, N. V. et al. A new design for a green calcium indicator with a smaller size and a reduced number of calcium-binding sites. Sci. Rep. 6, 1–15 (2016).
Lüscher, B. P., Vachel, L., Ohana, E. & Muallem, S. Cl- as a bona fide signaling ion. Am. J. Physiol. - Cell Physiol. 318, C125–C136 (2020).
Li, Y., Zhou, X. & Sun, S. X. Hydrogen, bicarbonate, and their associated exchangers in cell volume regulation. Front. Cell Dev. Biol. 9, 683686 (2021).
Jones, R. T., Faas, G. C. & Mody, I. Intracellular bicarbonate regulates action potential generation via KCNQ channel modulation. J. Neurosci. 34, 4409–4417 (2014).
Malekova, L. et al. Inhibitory effect of DIDS, NPPB, and phloretin on intracellular chloride channels. Pflug. Arch. Eur. J. Physiol. 455, 349–357 (2007).
Li, Y. & Tsien, R. W. pHTomato, a red, genetically encoded indicator that enables multiplex interrogation of synaptic activity. Nat. Neurosci. 2012 157 15, 1047–1053 (2012).
Chen, C. et al. Excitation ratiometric chloride sensing in a standalone yellow fluorescent protein is powered by the interplay between proton transfer and conformational reorganization. Chem. Sci. 12, 11382–11393 (2021).
Arosio, D. et al. Simultaneous intracellular chloride and pH measurements using a GFP-based sensor. Nat. Methods 7, 516–518 (2010).
Sato, S. S. et al. Simultaneous two-photon imaging of intracellular chloride concentration and pH in mouse pyramidal neurons in vivo. Proc. Natl Acad. Sci. Usa. 114, E8770–E8779 (2017).
Linghu, C. et al. Spatial multiplexing of fluorescent reporters for imaging signaling network dynamics. Cell 183, 1682–1698.e24 (2020).
Sui, J. et al. Optimization of a yellow fluorescent protein-based iodide influx high-throughput screening assay for cystic fibrosis transmembrane conductance regulator (CFTR) modulators. 656–669 https://doi.org/10.1089/adt.2010.0312 (2010).
Merkert, S. et al. High-throughput screening for modulators of CFTR activity based on genetically engineered cystic fibrosis disease-specific iPSCs. Stem Cell Rep. https://doi.org/10.1016/j.stemcr.2019.04.014 (2019).
Cole, D. E. C. & Evrovski, J. Quantitation of sulfate and thiosulfate in clinical samples by ion chromatography. J. Chromatogr. A 789, 221–232 (1997).
Mohapatra, N. K. et al. Sulfate concentrations and transport in human bronchial epithelial cells. Am. J. Physiol. - Cell Physiol. 264, C1231–C1237 (1993).
Shaner, N. C. et al. A bright monomeric green fluorescent protein derived from Branchiostoma lanceolatum. Nat. Methods 10, 407–409 (2013).
Subach, O. M. et al. cNTnC and fYTnC2, genetically encoded green calcium indicators based on troponin C from fast animals. Int. J. Mol. Sci. 23, 14614 (2022).
Pettersen, E. F. et al. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. urn:issn:0907-4449 66, 12–21 (2009).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Case: Amber18 (University of San Francisco) - Google Scholar. https://scholar.google.com/scholar_lookup?title=Amber18&author=DA+Case&author=IY+Ben-Shalom&author=SR+Brozell&author=DS+Cerutti&author=TE+Cheatham&publication_year=2018&.
Breyfogle, K. L., Blood, D. L., Rosnik, A. M. & Krueger, B. P. Molecular dynamics force field parameters for the EGFP chromophore and some of its analogues. J. Phys. Chem. B 127, 5772–5788 (2023).
Maier, J. A. et al. ff14SB: Improving the accuracy of protein side chain and backbone parameters from ff99SB. J. Chem. Theory Comput. 11, 3696–3713 (2015).
Bayly, C. I., Cieplak, P., Cornell, W. D. & Kollman, P. A. A well-behaved electrostatic potential based method using charge restraints for deriving atomic charges: The RESP model. J. Phys. Chem. 97, 10269–10280 (1993).
Huige, C. J. M. & Altona, C. Force field parameters for sulfates and sulfamates based on ab initio calculations: Extensions of AMBER and CHARMm fields. J. Comput. Chem. 16, 56–79 (1995).
Roe, D. R. & Cheatham, T. E. PTRAJ and CPPTRAJ: Software for processing and analysis of molecular dynamics trajectory data. J. Chem. Theory Comput. 9, 3084–3095 (2013).
Humphrey, W., Dalke, A. & Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Piatkevich, K. D., Yang, L. & Lai, C. Genetically encoded green fluorescent sensor for probing sulfate transport activity of solute carrier family 26 member a2 (Slc26a2) Protein. FigShare https://figshare.com/articles/dataset/_b_Genetically_Encoded_Green_Fluorescent_Sensor_for_Probing_Sulfate_Transport_Activity_of_Solute_Carrier_Family_26_Member_a2_Slc26a2_Protein_b_/27143766 (2024).
Pathiranage, V. & Walker, A. R. Genetically encoded green fluorescent sensor for probing sulfate transport activity of solute carrier family 26 member a2 (Slc26a2) Protein. Zenodo https://doi.org/10.5281/ZENODO.10416670 (2024).
Acknowledgements
We thank Stavrini Papadaki from Westlake Laboratory for verifying all plasmid sequences and depositing them to WeKwikGene. We thank the Flow Cytometry Core Facility at Westlake University for their help with cell sorting. We also thank Zhong Chen and Xingyu Lu from Westlake University for their assistance with spectroscopic measurements. We thank the WSU Supercomputing Grid for computational support. This work was supported by start-up funding from the Foundation of Westlake University, National Natural Science Foundation of China grant 32050410298 and 32171093, 2020 BBRF Young Investigator Grant 28961 to K.D.P. and by the Ministry of Science and Higher Education of the Russian Federation for the development of the Kurchatov Center for Genome Research 075-15-2019-1659 within the state assignment of NRC "Kurchatov Institute" and by the Russian Science Foundation project 21-74-20135 (protein purification) to F.V.S. The authors thankfully acknowledge support from the National Science Foundation−NSF through the grant NSF CHE2338804 to A.R.W.
Author information
Authors and Affiliations
Contributions
C.L. developed Thyone and performed its characterization. C.L., with help from F.V.S., characterized Thyone in vitro. R.W. prepared random libraries and performed preliminary screening. C.L. and L.Y. performed calibration experiments in cell culture. V.P. and A.R.W. performed MD simulations. C.L. and K.D.P. analyzed the data and, together with V.P. and A.R.W., wrote the manuscript. K.D.P. initiated the project, made high-level designs and plans, and oversaw all aspects.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Manuel Breuer. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lai, C., Yang, L., Pathiranage, V. et al. Genetically encoded green fluorescent sensor for probing sulfate transport activity of solute carrier family 26 member a2 (Slc26a2) protein. Commun Biol 7, 1375 (2024). https://doi.org/10.1038/s42003-024-07020-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-024-07020-9