Abstract
Osteoarthritis (OA) is a degenerative joint disease that affects the cartilage and surrounding tissues. The transcription factor Kruppel-like family factor 9 (KLF9) has been identified as a regulator of tumorigenesis. However, its role in OA is still not fully understood. Herein, this study aimed to access the potential role and molecular mechanism by which KLF9 regulates OA development. KLF9 was upregulated in cartilage tissues of OA patients and medial meniscotibial ligament (MMTL)-induced OA rats, as well as in IL-1β-treated chondrocytes. Furthermore, knockdown of KLF9 inhibited OA-related cartilage injury, as evidenced by inhibiting chondrocyte extracellular matrix (ECM) degradation, increasing chondrocyte viability, and decreasing apoptosis. Conversely, overexpression of KLF9 had the opposite effect. The downstream mechanism of KLF9 was confirmed. KLF9 mediated the transcription of G protein-coupled receptor kinase 5 (GRK5) by directly targeting the GRK5 promoter. GRK5 knockdown eliminated the effects of KLF9 overexpression on chondrocyte dysfunction. It was also found that GRK5 combined with histone deacetylase 6 (HDAC6) and promoted HDAC6 phosphorylation. The use of the HDAC6 inhibitor TubastatinA also abolished the effects of GRK5 overexpression on chondrocyte ECM degradation and apoptosis. These results demonstrate that the KLF9-GRK5-HDAC6 axis plays a crucial role in promoting the progression of OA.
Similar content being viewed by others
Introduction
Osteoarthritis (OA), a chronic progressive degenerative disease of the whole joint, is a major source of pain and disability worldwide. It affects various parts of the joint, including articular cartilage, subchondral bone, ligaments, joint capsules, and synovium1. The main histopathological features of OA are progressive loss of articular cartilage, chondroosteophyte formation, and thickening of subchondral bone2. However, there are currently no effective strategies for repairing damaged cartilage, and patients with advanced OA often require joint replacement. Therefore, it is crucial to gain a better understanding of the molecular mechanisms underlying OA pathogenesis and to develop new therapeutic targets for the diagnosis and treatment of this condition.
Kruppel-like family factor 9 (KLF9) is a member of the KLF transcription factor family that is expressed in a variety of tissues, such as the brain, kidney, and lung3. Previous studies have shown that overexpression of KLF9 induces cell apoptosis in myeloma, prostate cancer, and hepatocellular carcinoma4,5,6. Notably, a recent study has identified KLF9 as a characteristic gene for OA and suggested that it may play a key role in the development of OA7. To further investigate this potential link, the current study collected articular cartilage tissues from OA patients for mRNA sequencing. The results suggested that KLF9 was highly expressed in the articular cartilage tissues of OA patients, but its explicit role in OA remains unclear.
G protein-coupled receptor kinases (GRKs) are a family of serine/threonine kinases that are known to have the ability to recognize and phosphorylate agonist-activated G protein-coupled receptors (GPCRs), resulting in their desensitization8. G protein-coupled receptor kinase 5 (GRK5) is a member of the GRK superfamily. GRK5 promotes inflammation through NF-κB signaling during myocardial infarction9. GRK5 also plays a carcinogenic role in non-small cell lung cancer and prostate cancer10,11. In addition, Sueishi et al. indicated that GRK5 inhibition attenuated cartilage degeneration by inhibiting the NF-κB signaling pathway12. Remarkably, the results of mRNA sequencing showed elevated levels of GRK5 in the articular cartilage tissues of patients with OA. Further analysis of the GRK5 promoter sequence revealed potential binding sites for KLF9, suggesting that KLF9 may regulate GRK5 transcription. This suggests that the KLF9-GRK5 axis may play an important role in OA, but this hypothesis requires further confirmation.
GRK5 is also known to be a class II histone deacetylase (HDAC) kinase, promoting the phosphorylation of HDAC513. Hitpredict analysis revealed a potential interaction between GRK5 and HDAC6. In Hela and MDA-MB-231 cells, GRK5 facilitated the phosphorylation of HDAC6Ser21, thereby enhancing its deacetylation activity14. Previous studies have revealed that HDAC6 exacerbates mitochondrial dysfunction and extracellular matrix degradation, and that the HDAC6 inhibitor Tubastatin A reduces oxidative stress in chondrocytes, thus alleviating OA15,16. However, it is currently unknown whether GRK5 exerts an available role in OA by phosphorylating HDAC6.
This study aimed to investigate the potential role and mechanism of KLF9 in the development of OA. Our research first demonstrated the accumulation of KLF9 in both human and rat articular cartilage tissues affected by OA, as well as in chondrocytes induced by IL-1β. Our data further revealed that the KLF9-GRK5-HDAC6 axis played a significant role in the pathogenesis of OA. Our findings offer new insights into the underlying mechanisms of OA and lay the groundwork for the identification of potential therapeutic targets.
Results
Differential gene expression in articular cartilage tissues of patients with OA
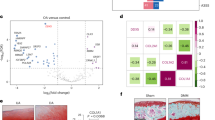
Firstly, to screen the target factors that may play a key role in OA process, six healthy and OA articular cartilage tissues of human were collected for mRNA sequencing. The volcano plot and heat map showed the expression of DEGs, of which 241 DEGs were upregulated and 265 DEGs were downregulated (Fig. 1A). To verify the functional role of the above DEGs, GO and KEGG pathway enrichment analyses were carried out. Sankey diagram of GO enrichment analysis displayed the relationship between DEGs and the pathway, and the results implied that DEGs were significantly enriched in regulation of extrinsic apoptotic signaling pathway, activation of protein kinase activity, DNA-binding transcription activator activity and negative regulation of epithelial cell proliferation (Supplementary Fig. 1A). The network diagram of GO enrichment analysis suggested the DEGs interaction of several pathways, including regulation of extrinsic apoptotic signaling pathway, negative regulation of epithelial cell proliferation, activation of protein kinase activity, DNA-binding transcription activator activity, and G protein-coupled receptor kinase activity (Supplementary Fig. 1B). In addition, KEGG enrichment analysis indicated that DEGs were involved in p53 signaling pathway, NF-kappa B signaling pathway and apoptosis (Supplementary Fig. 1C).
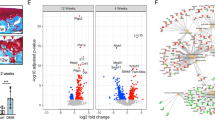
Knee articular cartilage tissues were collected from six OA patients and controls. mRNA sequencing was performed to detect gene expression in clinical samples. A The volcano plot and heat map showed the expression of DEGs. B The bar graph and heat map presented the expression of KLF family members in cartilage tissues from OA patients. C–E MicroCT, safranin O-fast green staining, OARSI score, and H&E staining were employed to determine the pathological changes of knee tissues (200×, scale bar: 100 µm; 400×, scale bar: 50 µm). $$, p < 0.01; n = 6.
The expression of KLF9 was significantly upregulated in OA human and rat articular cartilage tissues
As demonstrated by the previous mRNA sequencing results, the bar graph and heat map suggested the log2FC of KLF transcription factor family members in cartilage tissues from patients with OA (Fig. 1B). It was observed that there were 13 KLF genes. Among them, 5 genes were differentially expressed, including KLF4, KLF6, KLF9, KLF10, and KLF11. By querying the functions of these 5 genes, the function of KLF4, KLF10, and KLF11 have been reported in OA. However, the function of KLF6 and KLF9 in OA is unknown. We found that the log2FC of KLF9 is greater than that of KLF6, so KLF9 was selected as a target factor for follow-up study. To assess the expression of KLF9 on OA development in vivo, an OA rat model was established through MMTL surgery. MicroCT, safranin O-fast green staining, OARSI score, and H&E staining were then used to analyze the pathological changes in knee tissues. The histological findings revealed that the articular cartilage surface integrity was compromised and a significant loss of chondrocytes was observed in the OA group compared to the sham group (Fig. 1C–E).
Subsequently, real-time PCR and immunohistochemistry results confirmed the upregulation of KLF9 in human articular cartilage tissues affected by OA (Fig. 2A).
A The expression of KLF9 in articular cartilage tissues of OA patients and controls was detected using real-time PCR and immunohistochemical staining. B, C The expression of KLF9 in articular cartilage tissues of control and OA rats was examined by real-time PCR, western blotting, and immunohistochemical staining (400×, scale bar: 50 µm; 800×, scale bar: 25 µm). $$$$, p < 0.0001. n = 6.
Furthermore, we compared the expression levels of KLF9 in normal and OA articular cartilage using real-time PCR, western blotting, and immunohistochemistry, and found that KLF9 expression was markedly increased in the articular cartilage tissues of OA rats compared to the control group (Fig. 2B, C). Overall, these results suggested that KLF9 levels were elevated in both OA human and rat articular cartilage tissues, indicating a potential role for KLF9 in the progression of OA.
KLF9 knockdown relieved cartilage injury in OA rats
Furthermore, the in vivo effects of KLF9 knockdown were evaluated by injecting adenovirus rKLF9sh into the knee cavity of OA rats. Real-time PCR and immunohistochemical results showed successful adenovirus-mediated KLF9 knockdown 1 week after injection (Fig. 3A). MicroCT, safranin O-fast green staining, OARSI score, and H&E staining results revealed that OA rats exhibited increased cartilage surface calcification, elevated joint space stenosis, impaired articular cartilage surface integrity, and decreased chondrocytes. However, these phenotypic alterations were negated by KLF9 knockdown (Fig. 3B–D). We then investigated the effects of KLF9 on chondrocyte ECM degradation. The levels of extracellular proteolytic enzyme in cartilage degradation (anabolic marker COL2A1) and (catabolic marker MMP13) were evaluated by real-time PCR and immunohistochemistry. As indicated in Fig. 4A, KLF9 knockdown markedly increased COL2A1 levels and decreased MMP13 levels. Additionally, tissue TUNEL staining indicated that knockdown of KLF9 inhibited chondrocyte apoptosis (Fig. 4B). Following on these results, we demonstrated that KLF9 knockdown relieved cartilage injury, thus ameliorating OA in rats.
The OA model was established by MMTL method, and KLF9 knockdown adenovirus was injected into the joint cavity of rats. A The infection efficiency of KLF9 was measured using real-time PCR and immunohistochemical staining (400×, scale bar: 50 µm; 800×, scale bar: 25 µm). B–D MicroCT, safranin O-fast green staining, OARSI score, and H&E staining were used to analyze the pathological changes of knee tissues (200×, scale bar: 100 µm). $, p < 0.05; $$$, p < 0.001; $$$$, p < 0.0001. n = 6.
A The levels of COL2A1 and MMP13 were examined by real-time PCR and immunohistochemical staining (400×, scale bar: 50 µm). B Cartilage apoptosis was detected by TUNEL staining (400×, scale bar: 50 µm). $$$$, p < 0.0001. n = 6.
KLF9 aggravated IL-1β-induced chondrocyte ECM degradation and apoptosis
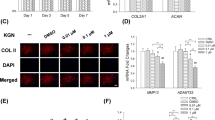
Pro-inflammatory cytokines, such as IL-1β, have been linked to the pathogenesis of OA17. Therefore, we used IL-1β to simulate OA in vitro. After 48 h of infection with adenovirus, human chondrocytes were treated with 10 ng/ml of human active IL-1β for 24 h. As shown in Supplementary Fig. 2A, chondrocytes were successfully infected with KLF9 knockdown or overexpressing adenoviruses. To assess the impact of KLF9 on chondrocyte viability, a CCK-8 assay was performed. The results displayed that IL-1β significantly reduced chondrocyte viability. However, this effect was significantly reversed by KLF9 knockdown (Fig. 5A). Data of real-time PCR and western blotting suggested that knockdown of KLF9 greatly increased COL2A1 levels and decreased MMP13 levels (Fig. 5B). Additionally, cell apoptosis analysis demonstrated that KLF9 knockdown significantly inhibited IL-1β-induced cell apoptosis, as evidenced by reduced TUNEL-positive cells, decreased Bax levels, downregulated caspase-3 and caspase-9 activities, and increased Bcl-2 levels (Fig. 5C–E). To confirm these findings, we also overexpressed KLF9. The results illustrated that KLF9 overexpression exhibited the opposite function (Fig. 5A–E), thus exacerbating IL-1β-induced chondrocyte ECM degradation and apoptosis.
A CCK-8 assay was applied to determine cell viability. B The levels of COL2A1 and MMP13 were tested by real-time PCR and western blotting. C Chondrocyte apoptosis was detected by TUNEL staining (200×, scale bar: 100 µm; arrows represented TUNEL-positive cells). D The expression levels of Bax and Bcl-2 were detected by western blotting. E The activities of caspase-3 and caspase-9 were detected by the kits. $, p < 0.05; $$, p < 0.01; $$$, p < 0.001; $$$$, p < 0.0001. n = 3.
KLF9 mediated transcription of GRK5
Next, we probed the potential downstream molecular mechanism of KLF9. The previous results of mRNA sequencing suggested that GRK5 was elevated in the cartilage tissues of OA patients. It was also demonstrated that KLF9 and GRK5 mRNA expression in articular cartilage of clinical OA patients was positive correlated using Pearson’s correlation analysis (Fig. 6A). GSEA plot showed the expression of DEGs in the DNA-binding transcription activator activity pathway (Fig. 6B). Notably, JASPAR database prediction indicated that there were potential binding sites of KLF9 on the promoter of GRK5. To investigate whether KLF9 may targeted the promoter region of GRK5, we performed a dual luciferase reporter gene assay. It was illustrated that KLF9 obviously elevated luciferase activity, indicating a possible target between KLF9 and the GRK5 promoter (Fig. 6C). In addition, predicted graph for KLF9 binding site in the GRK5 promoter region was also displayed. The binding of KLF9 to GRK5 promoter was further confirmed by DNA pull-down and ChIP-qPCR data (Fig. 6D, E).
A Real-time PCR was used to detect the expression of GRK5 in articular cartilage of OA and control patients. In addition, correlation between KLF9 and GRK5 expression in articular cartilage of clinical OA patients was also analyzed. B GSEA plot showed the expression of DEGs in the DNA-binding transcription activator activity pathway. C Promoter fragments of different lengths of human GRK5 were inserted into pGL-3-Basic reporter plasmid and co-transfected with human KLF9 overexpression plasmid into 293T cells. Luciferase activity was detected 48 h after transfection. In addition, predicted graph for KLF9 binding site in the GRK5 promoter region was also shown. D DNA pull-down assay was used to test the binding of GRK5 promoter to KLF9 protein in human chondrocytes. E The binding of KLF9 and GRK5 promoter in human chondrocytes was also detected by ChIP-qPCR. IgG was used as the negative control. F, G The expression of GRK5 in articular cartilage tissues was detected by real-time PCR, western blotting, and immunohistochemical staining (400×, scale bar: 50 µm). n = 6. H The expression of GRK5 in human articular chondrocytes was determined by real-time PCR and western blotting. $$, p < 0.01; $$$, p < 0.001; $$$$, p < 0.0001. n = 3.
To clarify the regulation of GRK5 by KLF9, real-time PCR, western blotting, and immunohistochemistry were employed to detect the levels of GRK5 in OA rat knee cartilage tissues or IL-1β-induced chondrocytes. As expected, KLF9 overexpression led to upregulation of GRK5 levels, while KLF9 knockdown downregulated its levels (Fig. 6F–H). All of the above data suggested that KLF9 mediated transcription of GRK5 and enhanced its expression.
GRK5 knockdown abolished the promoting effects of KLF9 overexpression on chondrocyte ECM degradation and apoptosis
We then sought to confirm whether KLF9 plays a vital role in OA progression through upregulation of GRK5. Initially, chondrocytes were successfully infected with GRK5 knockdown or GRK5 overexpressing adenoviruses (Fig. 7A). CCK-8 assay results indicated that ectopic KLF9 induction markedly inhibited cell viability, while GRK5 knockdown increased it (Fig. 7B). As shown in Fig. 7C, real-time PCR results suggested that GRK5 knockdown largely upregulated COL2A1 levels and downregulated MMP13 levels, thus reversing the role of KLF9 overexpression on chondrocyte ECM degradation. In addition, TUNEL staining showed that GRK5 knockdown reduced KLF9 overexpression-induced TUNEL-positive cells (Fig. 7D). The activities of caspase-3 and caspase-9 were increased in KLF9 overexpressed cells, while decreased in GRK5 silenced cells (Fig. 7E). These results defined that GRK5 knockdown abolished the promoting effects of KLF9 overexpression on chondrocyte apoptosis. In addition, GRK5 overexpression also abrogated the inhibitory effects of KLF9 knockdown on chondrocyte ECM degradation and apoptosis (Fig. 7B–E), supporting that KLF9 participated in OA progression by modulating GRK5.
A The infection efficiency of GRK5 was tested by real-time PCR. B CCK-8 assay was employed to examine cell viability. C The levels of COL2A1 and MMP13 were tested by real-time PCR. D Chondrocyte apoptosis was tested by TUNEL staining (200×, scale bar: 100 µm; arrows represented TUNEL-positive cells). E The activities of caspase-3 and caspase-9 were detected by the kits. $, p < 0.05; $$, p < 0.01. $$$, p < 0.001; $$$$, p < 0.0001. n = 3.
GRK5 bound to HDAC6 and promoted its phosphorylation
In accordance with evidence that GRK5, as a class II histone deacetylase (HDAC) kinase, promotes the phosphorylation of HDAC518,19, we also shed light on the downstream mechanism of GRK5. Markedly, hitpredict analysis revealed a potential binding between GRK5 and HDAC6 (https://www.hitpredict.org/interaction.php?Value=714595). A previous study has shown that HDAC6 exacerbates mitochondrial disorder and extracellular matrix degradation15. Based on this, we hypothesized that GRK5 may heighten OA progression by promoting the phosphorylation of HDAC6. To this end, we focused on the regulatory mechanism of these two proteins. Through immunofluorescence analysis, we noticed that IL-1β enhanced the colocalization of GRK5 and HDAC6 in chondrocytes (Fig. 8A). Co-IP results further confirmed the interaction between GRK5 and HDAC6 in IL-1β-treated chondrocytes (Fig. 8B). To determine the specific regions of GRK5 that bind to HDAC6, we constructed GRK5 overexpression plasmids containing different domains (with a flag label) and co-transfected them with human HDAC6 overexpression plasmids (with a his label) in HEK-293T cells. Co-IP results showed that all regions of GRK5 were able to bind to HDAC6 (Fig. 8C). Next, we infected IL-1β-treated chondrocytes with adenovirus carrying either wild type GRK5 or a kinase-inactive mutant (GRK5 K215R), and confirmed successful infection (Fig. 8D). Furthermore, we demonstrated that wild type GRK5 promoted HDAC6 phosphorylation, while the GRK5 K215R mutant and GRK5 knockdown inhibited its phosphorylation (Fig. 8E). We explored the effects of GRK5 on chondrocyte ECM degradation and apoptosis. The results showed that wild type GRK5 promoted the chondrocyte ECM degradation and apoptosis, which was abolished by GRK5 K215R (Fig. 8F, G). These findings suggested that GRK5 bound to HDAC6 and promoted its phosphorylation. Finally, we examined the effects of KLF9 on HDAC6 phosphorylation in vivo and in vitro, and the results showed that KLF9 knockdown inhibited the phosphorylation of HDAC6, while KLF9 overexpression showed the opposite function (Fig. 8H).
A The colocalization of GRK5 and HDAC6 in human chondrocytes was detected by immunofluorescence double staining (400×, scale bar: 50 µm). B Co-IP was used to determine the binding of GRK5 to HDAC6 in human chondrocytes. C The human GRK5 overexpressing plasmid with different domains (with a flag tag) was constructed and co-transfected with the human HDAC6 overexpressing plasmid (with a his tag) to 293T cells. After transfection for 48 h, the binding of GRK5 and HDAC6 was detected in Co-IP. D human chondrocytes were infected with GRK5 overexpressing plasmid (with flag tag) and kinase-inactive K215R GRK5 overexpressing plasmid (with flag tag), and western blotting was used to detect flag levels. E The levels of total HDAC6 and phosphorylated HDAC6 in chondrocytes were detected by western blotting. F The levels of COL2A1 and MMP13 were detected by real-time PCR. G Chondrocyte apoptosis was tested by TUNEL staining (200×, scale bar: 100 µm; arrows represented TUNEL-positive cells). H The levels of total HDAC6 and phosphorylated HDAC6 were determined in articular cartilage tissues (n = 6) and in chondrocytes (n = 3) by co-IP and western blotting. $, p < 0.05; $$, p < 0.01. $$$, p < 0.001; $$$$, p < 0.0001. n = 3.
To validate the role of GRK5 in OA progression through regulation of HDAC6, we conducted additional experiments using a specific HDAC6 inhibitor. Interestingly, the HDAC6 inhibitor Tubastatin A reversed the effects of GRK5 overexpression on chondrocyte ECM degradation and apoptosis, as evidenced by increasing cell viability and decreasing chondrocyte ECM degradation, TUNEL-positive cells, as well as caspase-3 and caspase-9 activities (Fig. 9A–D), indicating that the GRK5-HDAC6 axis was involved in OA progression.
A CCK-8 assay was used to test cell viability. B The levels of COL2A1 and MMP13 were tested by real-time PCR. C Chondrocyte apoptosis was determined by TUNEL staining (200×, scale bar: 100 µm; arrows represented TUNEL-positive cells). D The activities of caspase-3 and caspase-9 were examined by the kits. $, p < 0.05; $$, p < 0.01; $$$, p < 0.001; $$$$, p < 0.0001. n = 3.
Discussions
OA is a chronic age-related degenerative disease. Chondrocyte apoptosis is an important index in promoting the progression of OA20. In this study, we found that KLF9 as a pathogenic factor of OA, was highly expressed in in cartilage tissues of both OA rats and patients, as well as IL-1β-induced chondrocytes. Through in vivo and in vitro assays, we demonstrated that knockdown of KLF9 effectively reduced chondrocyte ECM degradation and apoptosis, while overexpression of KLF9 had the opposite effect. Further investigation revealed that KLF9 mediated the transcription of GRK5, and that overexpression of GRK5 counteracted the effects of KLF9 knockdown on chondrocyte ECM degradation and apoptosis. Subsequently, the downstream mechanism of GRK5 was uncovered, which confirmed that GRK5 bound to HDAC6 and promoted its phosphorylation. Strikingly, the HDAC6 inhibitor Tubastatin A eliminated the effects of GRK5 overexpression on chondrocyte ECM degradation and apoptosis. These observations suggested that the KLF9-GRK5-HDAC6 axis may be a novel mechanism for OA progression.
As a pivotal transcription factor, KLF9 is highly expressed in multiple tissues, including the brain and lungs. KLF9 was also found to be involved in cell proliferation, differentiation, and neurodevelopment21. In addition, KLF9 has been identified a regulator of tumorigenesis in various types of tumors, including colorectal, breast, and ovarian cancers22,23,24. Growing studies indicated that KLF9 promoted cell apoptosis in prostate and pancreatic cancer25,26. Of note, one study suggested that KLF9 may be a characteristic gene for OA and could potentially be involved in its development7. Despite the growing number of studies on KLF9, the exact mechanism by which it contributes to OA remains unclear. In this study, we observed elevated levels of KLF9 in OA human and rat articular cartilage tissues, as well as in IL-1β-induced chondrocytes. Furthermore, our in vivo and in vitro experiments demonstrated, for the first time, that knocking down KLF9 reduced cartilage damage and inhibited chondrocyte ECM degradation and apoptosis. Conversely, overexpression of KLF9 showed the opposite effect, exacerbating the development of OA.
We further clarified the detailed molecular mechanism of KLF9 in regulating cartilage injury. GO enrichment analysis indicated that DEGs were involved in the regulation of G protein-coupled receptor kinase activity. In addition, the results of mRNA sequencing showed that GRK5, a member of the GRK superfamily, was elevated in the cartilage tissues of OA patients. Accumulating evidence suggested that GRK5 promoted inflammation via the NF-κB pathway in myocardial infarction and arthritis9,27. GRK5 also acted as a key contributor to the growth and metastasis of non-small cell lung and prostate cancer cells10,11. Of note, Sueishi et al. suggested that GRK5 silencing suppressed cartilage degeneration by inhibiting the NF-κB signaling pathway12. In this study, we observed a positive correlation between GRK5 and KLF9 expression in the cartilage tissues of OA patients. According to the JASPAR database, there are potential binding sites for KLF9 on the promoter of GRK5. We then demonstrated that KLF9 directly targeted the promoter region of GRK5 and upregulated its expression. Furthermore, overexpression of GRK5 reversed the effects of KLF9 knockdown on chondrocyte ECM degradation and apoptosis, suggesting that KLF9 may regulate OA progression by affecting GRK5 levels.
Although we have identified a potential mechanism by which KLF9 regulates the OA process through GRK5, the specific role of GRK5 in modulating OA-related chondrocyte injury remains to be fully understood. Growing studies introduced that GRK5 was an HDAC kinase that promoted the phosphorylation of HDAC513,28,29. With hitpredict prediction, we found a potential combination between GRK5 and HDAC6. A previous study proved that GRK5 facilitated the phosphorylation of HDAC6Ser21, thus increasing its deacetylation activity14. Remarkably, HDAC6 fostered mitochondrial disorder and extracellular matrix degradation in chondrocytes, and that the HDAC6 inhibitor Tubastatin A repressed oxidative stress15,16. Herein, we proposed that GRK5 elicited OA progression by promoting the phosphorylation of HDAC6. Our results were consistent with those of previous studies demonstrating that GRK5 bound to HDAC6 and enhanced HDAC6 phosphorylation, while a GRK5 K215R mutation and GRK5 knockdown inhibited HDAC6 phosphorylation. Furthermore, we found that the HDAC6 inhibitor Tubastatin A counteracted the effects of GRK5 overexpression on chondrocyte ECM degradation and apoptosis, suggesting that GRK5 may facilitate OA progression through the phosphorylation of HDAC6.
In conclusion, this study ascertains for the first time that KLF9 is upregulated in both human and rat articular cartilage tissues, as well as IL-1β-induced chondrocytes. Furthermore, the KLF9-GRK5-HDAC6 axis has been identified as a contributor to OA development by promoting chondrocyte ECM degradation and apoptosis. These findings offer a new perspective on the pathological mechanism of OA.
Experimental procedures
Human cartilage samples collection and mRNA sequencing
Six healthy cartilage samples (4 females, aged 15, 16, 18, and 19; 2 males, aged 15 and 15) and six articular cartilage samples (5 females, aged 56, 64, 69, 70, and 77; 1 male, aged 72) from OA patients were obtained. Part of the control group was from knee cartilage fragments caused by knee trauma, and the remaining part was from the lateral femoral condyle cartilage fragments obtained after total knee arthroplasty in patients with valgus knee deformity. The human articular cartilage tissue collection and human cartilage tissue-related experiments were approved by Ethical Committee of the Shengjing Hospital of China Medical University and following the guidelines of the Declaration of Helsinki. The articular cartilage tissues collected from OA patients and controls were used for mRNA sequencing.
Total RNA was used as input material for RNA sample preparation. mRNA was purified from total RNA using poly-T oligo-attached magnetic beads. The first strand cDNA was synthesized with M-MuLV reverse transcriptase (RNase H), followed by the second strand cDNA synthesized with DNA polymerase I and RNase H. PCR was performed using Phusion high-fidelity DNA polymerase. Finally, PCR products were purified using the AMPure XP system, and library quality was assessed on the Agilent 5400 system (USA) and quantified by real-time PCR. The qualified libraries were sequenced on Illumina platforms with PE150 strategy in Novogene (Beijing, China). The original fluorescence image files obtained from Illumina platform were transformed to short reads (Raw data) by base calling and Raw data were recorded in FASTQ format. For the bioinformatics analysis, differentially expressed gene (DEG, |log2FC| > 1 and p < 0.05) analysis in articular cartilage tissues of OA patients and controls was performed using DESeq2 software (version 1.24.0). GO and KEGG enrichment analyses of DEGs were also conducted by cluster-Profiler software (version 4.6.2).
Animal OA model and adenovirus infection
All experiments were performed in accordance with Institutional Animal Care and Use Committee (IACUC) guidelines and approved by the Shengjing Hospital of China Medical University. Male Wistar rats (12 weeks old) were randomly divided into 4 groups (Con, OA, OA+shNC, and OA+rKLF9sh) and kept in a room with a 12-h/12-h dark/light cycle and constant temperature. One week later, the OA model was established. A small incision (1 cm) was made on the medial parapatellar area of the right knee joint, followed by incision of the medial capsule. Subsequently, the medial meniscotibial ligament (MMTL) was transected. Finally, the joint capsule and skin were sutured separately. After 7 days of operation, the corresponding adenovirus (15 µl, 4 × 1010/ml, 10 days later, one additional virus injection) was injected into the joint cavity of rats in OA+shNC and OA+rKLF9sh groups. At 4 weeks after the operation, the rats were sacrificed, and knee cartilage tissues were collected for follow-up experiments30.
For adenovirus infection, shRNA targeting KLF9 (AGAGCTTGTTGGACCTGAA
CATTCAAGAGATGTTCAGGTCCAACAAGCTCTTTTTT) or shRNA targeting GRK5 (GGACCATAGACAGAGATTACTTTCAAGAGAAGTAATCTCTGTCTATG
GTCCTTTTT) were inserted into pShuttle-CMV. KLF9 CDS and GRK5 CDS were also inserted into pShuttle-CMV.
Histological and immunochemical analysis
The paraffin blocks were cut into 5 µm sections and stained with hematoxylin for 5 min (Solarbio, Beijing, China) and eosin (H&E) for 3 min. For safranin O-fast green staining, the paraffin-embedded sections were stained with fast green staining solution for 2 min and safranin staining solution for 2 min. Immunohistochemical staining was performed according to a standard protocol. Briefly, the paraffin-embedded sections were deparaffinized and rehydrated, and the endogenous peroxidase was blocked by 3% hydrogen peroxide. Afterwards, the sections were incubated with 1% BSA (Sangon Biotech, Shanghai, China) for 15 min at room temperature, followed by incubation with primary antibody (KLF9, 1:100, Bioss, Changzhou, China or GRK5, 1:50, A3899, ABclonal) overnight at 4 °C. Next day, HRP-conjugated secondary antibody was employed to detect the primary antibody for 60 min. Finally, a DAB substrate (Solarbio) was used for coloration. Images were taken using a microscope (Olympus Inc., Tokyo, Japan).
Cell culture and treatment
Human articular chondrocytes obtained from iCell Bioscience Inc (Shanghai, China) were cultured in the special culture medium for chondrocytes (iCell) at 37 °C with 5% CO2. Chondrocytes were infected with KLF9 overexpression or KLF9 knockdown adenovirus (MOI = 100) for 48 h and treated with human active IL-1β (10 ng/ml, MCE, USA) for 24 h. In addition, 48 h after infection with KLF9-overexpressing adenovirus, chondrocytes were treated with 10 ng/ml IL-1β and 5 µM HDAC6 inhibitor Tubastatin A (Aladdin, Shanghai, China) for 24 h.
Cell viability assay
Cell viability was tested using a CCK-8 (KeyGEN Biotech, Nanjing, China) according to the manufacturer’s instructions. In brief, cells were plated in 96-well plates at a density of 4 × 103 cells/well. CCK-8 solution (10 µl) was added to each well and the plate was incubated at 37 °C for 2 h. Subsequently, absorption levels were detected at a wave-length of 450 nm using a microplate reader (BIOTEK, USA).
TUNEL staining
DNA damage was detected by TUNEL staining. The chondrocytes or cartilage sections were deparaffinized, rehydrated, and treated with 0.1% Triton X at 37 °C for 8 min. TUNEL reaction was conducted at 37 °C for 60 min using the in-situ cell death assay kit (Roche, Switzerland) according to the manufacturer’s instructions. The nuclei were stained with DAPI (Aladdin, Shanghai, China). The staining was captured under a fluorescence microscope (Olympus Inc.).
Detection of caspase-3 and caspase-9 activities
The activities of Caspase-3 (Beyotime, Shanghai, China) and Caspase-9 (Beyotime) were determined according to the instructions of the respective assay kits.
Real-time PCR
Total RNA was extracted from cartilage tissues and chondrocytes using TRIpure (Biotek, Beijing China). One microgram of total RNA was applied to synthesize cDNA using All-in-One First-Strand SuperMix reverse transcriptase (Magen Biotechnology, Guangzhou, China). Thereafter, cDNA was amplified with real-time PCR system (BIONEER, Korea) using SYBR green kit (Solarbio). The relative mRNA expression was normalized against β-actin using the 2-ΔΔCt method. The primer sequences were as follow:
homo GRK5 F, 5′-TTGACCGCTTTCTCCAGT-3′;
homo GRK5 R, 5′-GCTTGCAGGCATACATTTT-3′;
homo KLF9 F, 5′-TGGACCTGAACAAGTACCGA-3′;
homo KLF9 R, 5′-CCAGGATGGAGGAGGGA-3′;
homo β-actin F, 5′-GGCACCCAGCACAATGAA-3′;
homo β-actin R, 5′-TAGAAGCATTTGCGGTGG-3′;
rat GRK5 F, 5′-AGACTTACACCGTGAGAACA-3′;
rat GRK5 R, 5′-GCCAACAGTGCCTACCC-3′;
rat KLF9 F, 5′-TTGGACCTGAACAAATACCG-3′;
rat KLF9 R, 5′-CCAGAGTGGAGGAGGGAGA-3′;
rat COL2A1 F, 5′-CCCCGAGGTGACAAAG-3′;
rat COL2A1 R, 5′-AGGGATTCCATTAGAGCC-3′;
rat MMP13 F, 5′-GCCAGAACTTCCCAACCA-3′;
rat MMP13 R, 5′-ACCCTCCATAATGTCATACCC-3′;
rat β-actin F, 5′-GGAGATTACTGCCCTGGCTCCTAGC-3′;
rat β-actin R, 5′-GGCCGGACTCATCGTACTCCTGCTT-3′.
Western blotting
The cells were harvested and lysed in a RIPA lysate containing 1 mM PMSF on ice for 5 min. A BCA protein quantity kit purchased from Beyotime was used to determine the protein concentration. An equal amount of supernatant (20 µg per lane) was dissolved by 10% SDS-PAGE gel and transferred to the PVDF membranes (Abcam, Cambridge, UK) at 400 mA current. Then blocked membranes were incubated with primary antibodies (KLF9, 1:1000, bs-3644R, Bioss; Bax, 1:1000, A0207, ABclonal, Wuhan, China; Bcl-2, 1:1000, A21592, ABclonal; GRK5, 1:1000, A3899, ABclonal; HDAC6, 1:1000, AF6485, Affinity, Changzhou, China and p-HDAC6, 1:1000, AF3485, Affinity) overnight at 4 °C. Next, the membranes were incubated with HRP- conjugated secondary antibodies (1:5000) at room temperature for 45 min. Finally, ECL reagent (Beyotime) was used to detect immunoreactive bands. The density analysis was performed using Gel-Pro-Analyzer software.
Dual luciferase reporter gene assay
The GRK5 promoter fragments with different lengths were cloned into the pGL-3-basic firefly luciferase reporter vector and co-transfected into HEK-293T cells with the human KLF9 overexpression plasmid. Renilla was used as a control plasmid. After 48 h of transfection, the luciferase activity was detected by the dual luciferase reporter gene detection kit (KeyGEN Biotech).
ChIP-qPCR assay
ChIP detection kit (Beyotime) was used for ChIP assay. The cells were treated with 1% formaldehyde at 37 °C for 10 min to crosslink the protein and DNA. After cell lysis, ultrasound treatment was performed to cut genomic DNA. The samples after ultrasonic treatment were treated with ChIP Dilution containing 1 mM PMSF. 20 µl sample was used as input. The remaining sample was incubated with 70 µl Protein A + G Agarose/Salmon Sperm DNA at 4 °C for 30 min. Then DNA-protein complexes were incubated with 5 M NaCl to de-crosslink. After that, the DNA fragments were extracted with phenol chloroform for real-time PCR. The primers were as follows:
ChIP-qPCR, F, 5′-AGAGTCGGCTTCTGAACGG-3′;
ChIP-qPCR, R, 5′-CCTGGAAATGGGCTGTCTT-3′.
DNA pull-down
According to the length/mass ratio of 2 µg/1000 bp, biotin-labeled target DNA probes with corresponding mass were added into DNA beads buffer to 100 µl and added into Streptavidin magnetic beads. Next, nucleoprotein was extracted, and protein concentration was measured with the BCA protein assay kit. After nucleic acid removal and protein sample washing, the pull-down experiment was carried out. The pre-washed protein samples were mixed with 500 µl binding buffer, 5 µl poly (dI·dC), 5 µl protease inhibitor, 5 µl DTT, 9 µl EDTA and 4.5 µl EGTA. Then mixture was added to the probe-magnetic bead complex and incubated at 4 °C for 1 h. Finally, magnetic beads were incubated with 60 µl protein elution buffer and 0.6 µl DTT at 37 °C for 2 h. The magnetic beads were collected, and 15 µl protein sample was used for western blotting.
Co-IP assay
Cells were lysed in a RIPA lysate containing 1 mM PMSF and proteins were isolated. Total protein concentration was measured with a BCA protein quantity kit. As indicated by the immunoprecipitation test kit (Pierce, USA), the AminoLink conjugated resin was washed with 200 µl 1 × crosslinking buffer. IP antibody was added to the crosslinked buffer and ultrapure water with a volume of 200 µl. The above mixture was added to the centrifugal column with sodium cyanoborohydride solution and incubated at room temperature for 120 min. After centrifugation and resin washing, the lysate was added to the resin that had cured the corresponding antibody and incubated at room temperature for 2 h. Next day, after centrifugation, the column was placed in a new collection tube, then 200 µl IP lysate was added and centrifuged. Finally, the samples were employed to western blotting. The antibodies were as follows: GRK5 (1:1000, A3899, ABclonal), HDAC6 (1:1000, AF6485, Affinity) and anti-phosphoserine (1:3000, ab9332, Abcam).
Immunofluorescence double staining
The chondrocytes were fixed in 4% formaldehyde for 15 min and treated in 0.1% Triton X-100 (Beyotime) at 37 °C for 30 min. Subsequently, the samples were blocked with 1% BSA for 15 min, followed by incubation with primary antibodies (GRK5, 1:50, A3899, Abclonal and HDAC6, 1:400, 67250-1-Ig, Proteintech, Wuhan, China) overnight at 4 °C. The samples were then incubated with Cy3-conjugated secondary antibodies (1:200) at room temperature for 60 min. The nuclei were counterstained with DAPI. Finally, the samples were observed by a fluorescence microscopy (Olympus Inc).
Statistics and reproducibility
Statistical analysis was performed using GraphPad Prism 8.0 software (USA). The data are the means of 3 or 6 independent experiments mentioned in the figure legends. Data (Mean ± SD) between multiple groups were analyzed by one-way or two-way ANOVA with a post-hoc test (Tukey). Data between two groups were compared by the paired t-test. Correlation between KLF9 and GRK5 expression was analyzed with Person’s test. p < 0.05 was considered to be significantly different.
Data availability
The raw data for mRNA sequencing have been deposited at NCBI (GSE285234). All the data are available in the article and Supplementary files. Source data are provided in this paper in Supplementary Data 1. Uncropped western blots are found in Supplementary Data 2. Any remaining information can be obtained from the corresponding author upon reasonable request.
References
Loeser, R. F., Goldring, S. R., Scanzello, C. R. & Goldring, M. B. Osteoarthritis: a disease of the joint as an organ. Arthritis Rheumatol. 64, 1697–1707 (2012).
Wu, Y. et al. Exosomes rewire the cartilage microenvironment in osteoarthritis: from intercellular communication to therapeutic strategies. Int. J. Oral. Sci. 14, 40 (2022).
Ying, M. et al. Kruppel-like factor-9 (KLF9) inhibits glioblastoma stemness through global transcription repression and integrin alpha6 inhibition. J. Biol. Chem. 289, 32742–32756 (2014).
Mannava, S. et al. KLF9 is a novel transcriptional regulator of bortezomib- and LBH589-induced apoptosis in multiple myeloma cells. Blood 119, 1450–1458 (2012).
Shen, P. et al. KLF9, a transcription factor induced in flutamide-caused cell apoptosis, inhibits AKT activation and suppresses tumor growth of prostate cancer cells. Prostate 74, 946–958 (2014).
Sun, J. et al. Transcription factor KLF9 suppresses the growth of hepatocellular carcinoma cells in vivo and positively regulates p53 expression. Cancer Lett. 355, 25–33 (2014).
Zhang, J. et al. KLF9 and EPYC acting as feature genes for osteoarthritis and their association with immune infiltration. J. Orthop. Surg. Res. 17, 365 (2022).
Pitcher, J. A., Freedman, N. J. & Lefkowitz, R. J. G protein-coupled receptor kinases. Annu. Rev. Biochem. 67, 653–692 (1998).
Nagasaka, A. et al. GRK5-mediated inflammation and fibrosis exert cardioprotective effects during the acute phase of myocardial infarction. FEBS Open Bio 13, 380–391 (2023).
Jiang, L. P. et al. GRK5 functions as an oncogenic factor in non-small-cell lung cancer. Cell Death Dis. 9, 295 (2018).
Chakraborty, P. K. et al. G protein-coupled receptor kinase GRK5 phosphorylates moesin and regulates metastasis in prostate cancer. Cancer Res. 74, 3489–3500 (2014).
Sueishi, T. et al. GRK5 inhibition attenuates cartilage degradation via decreased NF-kappaB signaling. Arthritis Rheumatol. 72, 620–631 (2020).
Martini, J. S. et al. Uncovering G protein-coupled receptor kinase-5 as a histone deacetylase kinase in the nucleus of cardiomyocytes. Proc. Natl. Acad. Sci. USA 105, 12457–12462 (2008).
Lagman, J. et al. G protein-coupled receptor kinase 5 modifies cancer cell resistance to paclitaxel. Mol. Cell. Biochem. 461, 103–118 (2019).
Zheng, Y. et al. Inhibition of histone deacetylase 6 by Tubastatin A attenuates the progress of osteoarthritis via improving mitochondrial function. Am. J. Pathol. 190, 2376–2386 (2020).
Shen, Z. et al. Inhibition of HDAC6 by Tubastatin A reduces chondrocyte oxidative stress in chondrocytes and ameliorates mouse osteoarthritis by activating autophagy. Aging 13, 9820–9837 (2021).
Fei, J., Liang, B., Jiang, C., Ni, H. & Wang, L. Luteolin inhibits IL-1beta-induced inflammation in rat chondrocytes and attenuates osteoarthritis progression in a rat model. Biomed. Pharmacother. 109, 1586–1592 (2019).
Xiang, H. et al. GRK5 promoted renal fibrosis via HDAC5/Smad3 signaling pathway. FASEB J. 38, e23422 (2024).
Sasaki, S. et al. Type 2 diabetes susceptibility gene GRK5 regulates physiological pancreatic beta-cell proliferation via phosphorylation of HDAC5. iScience 26, 107311 (2023).
Charlier, E. et al. Insights on molecular mechanisms of chondrocytes death in osteoarthritis. Int. J. Mol. Sci. 17, https://doi.org/10.3390/ijms17122146 (2016).
Yan, X. et al. Progesterone receptor inhibits the proliferation and invasion of endometrial cancer cells by up regulating Kruppel-like factor 9. Transl. Cancer Res. 9, 2220–2230 (2020).
Brown, A. R. et al. Kruppel-like factor 9 (KLF9) prevents colorectal cancer through inhibition of interferon-related signaling. Carcinogenesis 36, 946–955 (2015).
Ybanez, W. S. & Bagamasbad, P. D. Kruppel-like factor 9 (KLF9) links hormone dysregulation and circadian disruption to breast cancer pathogenesis. Cancer Cell Int. 23, 33 (2023).
Wang, K., Liu, S., Dou, Z., Zhang, S. & Yang, X. Loss of Kruppel-like factor 9 facilitates stemness in ovarian cancer ascites-derived multicellular spheroids via Notch1/slug signaling. Cancer Sci. 112, 4220–4233 (2021).
Shen, P. et al. KLF9 suppresses cell growth and induces apoptosis via the AR pathway in androgen-dependent prostate cancer cells. Biochem. Biophys. Rep. 28, 101151 (2021).
Zhong, Z. et al. Expression of KLF9 in pancreatic cancer and its effects on the invasion, migration, apoptosis, cell cycle distribution, and proliferation of pancreatic cancer cell lines. Oncol. Rep. 40, 3852–3860 (2018).
Toya, M. et al. G protein-coupled receptor kinase 5 deletion suppresses synovial inflammation in a murine model of collagen antibody-induced arthritis. Sci. Rep. 11, 10481 (2021).
Zhang, Y. et al. Nuclear effects of G-protein receptor kinase 5 on histone deacetylase 5-regulated gene transcription in heart failure. Circ. Heart Fail. 4, 659–668 (2011).
Gold, J. I., Gao, E., Shang, X., Premont, R. T. & Koch, W. J. Determining the absolute requirement of G protein-coupled receptor kinase 5 for pathological cardiac hypertrophy: short communication. Circ. Res. 111, 1048–1053 (2012).
Iijima, H. et al. Subchondral plate porosity colocalizes with the point of mechanical load during ambulation in a rat knee model of post-traumatic osteoarthritis. Osteoarthr. Cartil. 24, 354–363 (2016).
Acknowledgements
The work was supported by grants from the National Natural Science Foundation of China (Grant No. 82172479).
Author information
Authors and Affiliations
Contributions
L.H.B. designed the project; X.N.Z. performed the experiments and wrote the draft of the manuscript; P. J., H.Q.T., and Y.F.W. analyzed data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
The human articular cartilage tissue collection and human cartilage tissue-related experiments were approved by Ethical Committee of the Shengjing Hospital of China Medical University and following the guidelines of the Declaration of Helsinki. All experiments were performed in accordance with Institutional Animal Care and Use Committee (IACUC) guidelines and approved by the Shengjing Hospital of China Medical University.
Peer review
Peer review information
Communications Biology thanks Shu Hu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Joao Valente.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhou, X., Jiang, P., Tan, H. et al. KLF9-GRK5-HDAC6 axis aggravates osteoarthritis pathogenesis by promoting chondrocyte extracellular matrix degradation and apoptosis. Commun Biol 8, 23 (2025). https://doi.org/10.1038/s42003-025-07460-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-07460-x
This article is cited by
-
Integrated analysis identifies conserved transcription factors TCF12, E2F1, and TEAD4 with diagnostic and therapeutic value in osteoarthritis subchondral bone
Genes & Genomics (2026)
-
KLF9-Mediated Transcriptional Promotion of HMGB2 Accelerates Cardiomyocyte Apoptosis, Inflammation, and Ferroptosis in Myocardial Ischemia/Reperfusion Injury
Cardiovascular Toxicology (2025)