Abstract
Increasing evidence shows that folate can regulate neural gene function through DNA modification, while regulatory roles of folate in neural RNA modification remain largely unknown. Here we show that folate treatment significantly increased Drosophila mRNA m6A levels. RNA methylation analysis indicates that carbon metabolism pathways and neural pathways are the mainly affected pathways by folate. Folate can increase mRNA m6A modification through carbon metabolic pathway in human cells, especially in neural cells. We also show that folate treatment can significantly increase the expression of m6A-related proteins (METTL3, METTL14) and neural mRNA m6A methylation levels in mice brain. Moreover, we find that folate-producing Lactobacillus plantarum can increase host mRNA m6A modification after colonization. In conclusion, we demonstrate that folate participates in mRNA m6A modification through carbon metabolic pathway in both Drosophila and mammals, and our results suggest that folate has a significant effect on neural-related genes and pathways.
Similar content being viewed by others
Introduction
Folate is a crucial vitamin that is involved in various biochemical processes. Animals must acquire folate from their diet or gut microbes since animals cannot synthesize folate themselves1. The beneficial effect of folate supplementation during pregnancy has been known for decades, due to the role of maternal folate in brain development and function of offspring2. Importantly, increasing evidence from preclinical research has revealed the importance of maternal folate for normal neurodevelopment and brain function in the offspring. Mechanistically, folate can serve as the methyl donor to influence DNA methylation, histone methylation and protein methylation3,4,5,6. Besides, methylation modification is one of the most important epigenetic modifications crucial for brain development, folate deficiency can lead to reduced methyl donors and neural tube defects7.
All epigenetic methylation requires methyl donors, which are generated by carbon metabolic pathway8. Therefore, it can be inferred that folate is not only involved in the process of DNA methylation but also in the process of RNA methylation, which is a new layer of gene expression regulation9. Despite the critical roles of folate as a methyl donor in DNA methylation and histone methylation were recognized, the role of folate in regulating RNA methylation was far less investigated. Recently, folate was reported to affect tRNA methylation in mammalian mitochondria and regulate mRNA m5C modification in neural stem cells10,11. N6-methyladenosine (m6A) is the most prevalent mRNA modification, which is particularly abundant in the central nervous system to impact many aspects of RNA functions, including mRNA stability, RNA splicing, and translation12,13. Importantly, folate has been observed to affect RNA m6A methylation levels in hippocampal tissue as well as the learning and memory ability of rats with acute lead exposure14. However, further investigations on the role and mechanism of folate in regulating specific mRNA m6A methylation with high-throughput sequencing approaches are needed to reveal the biological functions of folate.
Besides, it has been reported that the changes in host mRNA m6A methylation levels are related to gut microbes15,16. Our previous work found that gut microbes could regulate mRNA m6A methylation levels in mice’s brain17,18,19. Currently, there is a relationship between gut microbes and brain mRNA m6A methylation, but whether and how a specific microbial metabolite regulates host brain mRNA m6A methylation are not clear. In this study, we assessed the transcriptome-wide influence of folate on RNA m6A methylation and gene expression. We found that folate significantly up-regulated the transcripts enriched in Drosophila’s carbon metabolic pathway. MeRIP-seq showed that folate supplementation regulated mRNA m6A modification of genes enriched in neural function-associated pathways. The regulatory effect of folate on Drosophila mRNA m6A methylation was confirmed by colonizing the folate-producing Lactobacillus plantarum (L. plantarum). We also verified our findings in mice, and the results showed that folate impacted mRNA m6A methylation in the brains of both male and female mice. Our study provides insights into folate-regulated transcriptome and epitranscriptome, and our results confirm the importance of folate for the functions of neural mRNAs in animals and humans.
Results
Folate supplementation regulates mRNA m6A level in Drosophila
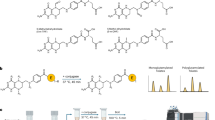
To investigate the effect of folate on mRNA m6A modification, we supplemented folate to Drosophila adult and Drosophila S2 cells. Since animals can also acquire folate from their gut microbes, we constructed germ-free Drosophila to explore the effect of folate without the interference of gut microbes (Supplementary Fig. 1a, b). For Drosophila S2 cells, we supplemented folate with cultured cells in dialyzed fetal bovine serum, which can minimize the interference from others nutrient (Fig. 1a). Subsequently, we performed a series of experiments to investigate the ability of folate in regulating mRNA m6A levels, including m6A-dot blot, m6A-immunofluorescence, western blot and MeRIP-seq and data analysis (Fig. 1a).
a Schematic diagram of this study. Folate-fed adult Drosophila and treated S2 cells were collected for several assays and MeRIP-seq sequencing. The control group (GF-Control) and folate-fed group (GF-Folate) were included in the experiments. The image is created by BioRender. b Dot blot of mRNA m6A methylation levels in Drosophila (the whole fly) treated with/o folate (n = 3). The results of dot blot were normalized to methylene blue staining. Mean ± SEM; * P < 0.05, measured with the unpaired, two-tailed Student’s t-test. c Immunofluorescence analysis showing m6A levels in S2 cells treated with/o folate (n = 7). d Western blot analysis of METTL3 in S2 cells treated with/o folate (n = 3). Mean ± SEM; *** P < 0.001, measured with the unpaired, two-tailed Student’s t-test. e RT-qPCR analysis of Mettl3 mRNA expression levels in S2 cells treated with/o folate (n = 3). Mean ± SEM; ** P < 0.01, measured with the unpaired, two-tailed Student’s t-test.
Firstly, we found that folate supplementation up-regulated mRNA m6A levels in Drosophila (Fig. 1b) and S2 cells (Fig. 1c). Western blot and qPCR analysis showed that the levels of METTL3 protein and Mettl3 mRNA increased in Drosophila S2 cells after folate supplementation (Fig. 1d, e). These results indicate that folate has a significant effect on Drosophila mRNA m6A methylation levels.
Transcriptome-wide impact of folate on Drosophila mRNA m6A methylome
Next, we profiled the transcriptome data analysis to compare the gene expression of the germ-free control group (GF-Control) and folate-fed group (GF-Folate). Principal component analysis (PCA) showed significant separation in global gene expression in GF-Control group and GF-Folate group (Fig. 2a), the expression of thousands of genes was significantly changed (Fig. 2b). In kyoto encyclopedia of genes and genomes (KEGG) analysis, we found carbon metabolism pathway was enriched in up-regulated genes after folate treatment (Fig. 2c). RT-qPCR experiment was further performed to verify the significant increase in gene expression levels in the carbon metabolism pathway (Fig. 2d). The map with predicted enzyme names of these genes was shown in (Supplementary Fig. 1c). These results indicated that folate could promote the expression levels of genes relating to the carbon metabolism pathway in Drosophila.
a Principal component analysis (PCA) of RNA sequencing data in control and folate-fed group (n = 3). b Volcano plot of RNA sequencing data in control and folate-fed group. FDR < 0.05 and log2FC ≥ 0.5 were set as the significance thresholds. c KEGG analysis of RNA sequencing data showing functional pathways of different genes in control and folate-fed group. d RT-qPCR experiments to verify carbon metabolic pathway genes from control group and folate-fed group in S2 cells (n = 3). e Normalized frequency of m6A peaks across different regions of mRNA in GF-Control and GF-Folate group, respectively. f Representative consensus motifs and corresponding P-values of m6A peaks identified in control and folate-fed group. g Genomic annotations of m6A sites in GF-Control and GF-Folate group, respectively. h Venn diagram showing common and unique differential m6A peak in GF-Control and GF-Folate group, respectively. i GO enrichment analysis showing functional pathways of different m6A peak-containing genes in GF-Folate and GF-Control group, respectively. j Dot blot analysis of mRNA m6A methylation levels in Control-brain group and Folate-brain group, respectively (n = 3). Results were normalized to methylene blue staining. k RT-qPCR analysis of Dhfr mRNA expression levels in Control-brain group and Folate-brain group (n = 3). l RT-qPCR analysis of Mettl3 mRNA expression levels in Control-brain group and Folate-brain group (n = 3). m Western blot analysis of METTL3 in Control-brain group and Folate-brain group, respectively (n = 3). n RT-qPCR analysis to verify carbon metabolic pathway genes in Control-brain group and Folate-brain group (n = 3). Mean ± SEM; * P < 0.05, ** P < 0.01, *** P < 0.001, measured with the unpaired, two-tailed Student’s t-test.
Since we found folate regulated mRNA m6A levels of Drosophila, we profiled the landscape of mRNA m6A in GF-Control group and GF-Folate group using MeRIP-seq. We found the different peaks were mainly distributed in 3’UTR and 1st Exon (Fig. 2e). Motif analysis identified the consensus sequence DRACH (Fig. 2f), which indicates the reliability of library construction and data analysis. In addition, GF-Control group and GF-Folate group had 12939 m6A peaks and 12169 m6A peaks, respectively. Among the different m6A peaks, 975 were up-regulated and 652 were down-regulated in GF-Folate group compared to GF-Control group (Fig. 2g). Based on the different m6A peak-containing genes of GF-Folate group compared to control group (Fig. 2h), we found several neural pathways by GO enrichment analysis (Fig. 2i).
To further confirm the role of folate in neural mRNA m6A modification, we continued to perform validation experiments using Drosophila brain. The results demonstrated a substantial increase in m6A levels in Drosophila brain following folate treatment (Fig. 2j). Subsequently, we performed qPCR analysis for Dhfr, a key enzyme in folate metabolism, and Mettl3, a critical enzyme for m6A modification. The results demonstrated that the expression of both Dhfr and Mettl3 was significantly up-regulated in the Drosophila brain following folate treatment (Fig. 2k, l). We further analyzed the expression of METTL3 using Western blot (Fig. 2m). Finally, we validated the genes related to carbon metabolism pathway in Drosophila brain. Our results demonstrate that these genes were significantly up-regulated in Drosophila brain following folate treatment (Fig. 2n). The above results indicated that folate had obvious transcriptome-wide and m6A epitranscriptome-wide impact on Drosophila mRNA, especially on neural mRNAs.
The dihydrofolate reductase Dhfr mediates mRNA m6A modification in Drosophila
One-carbon metabolism intermediates, including gut microbial metabolites like cobalamin (B12), folate (B9), and riboflavin (B2), are crucial for various biosynthetic processes, including DNA methylation and histone methylation20. Folate acts as a key substrate in carbon metabolism, utilizing transporter proton-coupled folate transporter (PCFT) and replication factor c (RFC), as well as the folate receptor FRα for cellular uptake21,22,23,24. Intracellular folate is reduced to dihydrofolate (DHF) by dihydrofolate reductase (Dhfr), which is a gene involved in carbon metabolism25. DHF can be reduced to tetrahydrofolate (THF), which enters the folate cycle and is converted to 5-methyltetrahydrofolate (5-MTHF). This compound subsequently enters the methionine cycle26,27, ultimately producing S-adenosylmethionine (SAM), which is essential for methylation modifications. In our study, we found that folate up-regulated carbon metabolism pathway and mRNA m6A levels, and we also found that Dhfr gene expression was significantly up-regulated by folate (Fig. 3a), so we disturbed the expression of Dhfr in Drosophila S2 cells (Supplementary Fig. 2a, Fig. 3b). The results showed that mRNA m6A level significantly decreased after disturbing the expression of Dhfr gene (Fig. 3c).
a RT-qPCR analysis of Dhfr mRNA expression levels in S2 cells treated with/o folate (n = 3). Mean ± SEM; * P < 0.05, measured with the unpaired, two-tailed Student’s t-test. b RT-qPCR analysis of Dhfr mRNA levels after dsDhfr in S2 cells (n = 3). Mean ± SEM; *** P < 0.001, measured with the unpaired, two-tailed Student’s t-test. c Dot blot of mRNA m6A methylation levels in S2-dsGFP and S2-dsDhfr, respectively (n = 3). Results were normalized to methylene blue staining. Mean ± SEM; * P < 0.01, measured with the unpaired, two-tailed Student’s t-test. d PCA of RNA sequencing data in S2-dsGFP and S2-dsDhfr, respectively (n = 3). e Volcano plot of RNA sequencing data in S2-dsGFP and S2-dsDhfr group, respectively. FDR < 0.05 and log2FC ≥ 0.5 were set as the significance thresholds. f GO analysis of RNA sequencing data showing functional pathways of different genes in S2-dsGFP and S2-dsDhfr group, respectively. g KEGG analysis of RNA sequencing data showing functional pathways of different genes in S2-dsGFP and S2-dsDhfr group, respectively. h Normalized frequency of m6A peaks across different regions of mRNA in S2-dsGFP and S2-dsDhfr group, respectively. i Representative consensus motifs and corresponding P-values of m6A peaks identified in S2-dsGFP and S2-dsDhfr, respectively.
To investigate the effect of Dhfr gene on gene expression profile of Drosophila S2 cells, we performed RNA sequencing. PCA showed significant separation in global gene expression in S2-dsGFP and S2-dsDhfr (Fig. 3d), and the expression of thousands of genes were significantly changed (Fig. 3e). Among the differential pathways, we found that folate-containing compound metabolic processes, NAD and NADP metabolic processes were down-regulated in S2-dsDhfr group (Fig. 3f), and in KEGG analysis, we found the one-carbon pool by the folate pathway was enriched in down-regulated genes after Dhfr interference (Fig. 3g). These results suggest that the disruption of Dhfr gene has an inhibitory effect on carbon metabolism.
We then profiled the landscape of mRNA m6A in S2-dsGFP group and S2-dsDhfr group. Peak frequency plots indicated that m6A peaks were mainly enriched in the 3′UTR and near the stop codon (Supplementary Fig. 2b), and the different peaks were mainly distributed in 3’UTR and 1st Exon (Fig. 3h). In addition, 2214 differential m6A peaks were up-regulated and 1815 were down-regulated in the S2-dsDhfr group compared to S2-dsGFP group (Supplementary Fig. 2c). Motif analysis showed that the consensus sequence changed from DRACH to DRAGH after the knockdown of Dhfr in S2 cells (Fig. 3I). Based on the down-regulated m6A peak of S2-dsDhfr group compared to S2-dsGFP group (Supplementary Fig. 2c, d), we found several neural-related pathways by GO enrichment analysis (Supplementary Fig. 2e).
Folate-producing L. plantarum can increase neural mRNA m6A methylation levels
Folate is one of the essential nutrients that can be obtained from both foods and gut microbes28. Many animals cannot synthesize folate but can rely on gut microbes to absorb folate. Lactobacilli play a key role in folate synthesis, especially the species possessing the folB, folK, folE, folP, and folQ genes (44,45). To validate the effect of folate on host mRNA m6A methylation levels, we selected L. plantarum, a bacterium capable of producing folate29, and colonized it into Drosophila.
We divided the group into control group (WT-Control) and L. plantarum colonization group (WT-LP) (Fig. 4a), followed by substantial experiments. The level of mRNA m6A methylation significantly increased in the L. plantarum colonization group (Fig. 4b). Meanwhile, we found that METTL3 protein expression was also elevated in the L. plantarum colonization group (Fig. 4c). These results suggest that L. plantarum has a significant effect on Drosophila mRNA m6A methylation levels.
a Experimental design of L. plantarum colonization in Drosophila. Adult Drosophila samples (the whole fly) with (WT-LP) or without (WT-Control) L. plantarum colonization were collected for Western assay/Dot blot assay and MeRIP-seq sequencing. The image is created by BioRender. b Dot blot of mRNA m6A methylation levels in WT-Control group and WT-LP group (n = 3). Results were normalized to SYBRTM Gold staining. Mean ± SEM; * P < 0.05, measured with the unpaired, two-tailed Student’s t-test. c Western blot analysis of METTL3 in WT-LP and WT-Control, respectively (n = 3). Mean ± SEM; * P < 0.05, measured with the unpaired, two-tailed Student’s t-test. d Venn diagram showing common and unique differential m6A peaks in WT-Control and WT-LP group, respectively. e GO enrichment analysis showing functional pathways of different m6A peak-containing genes.
We then profiled the landscape of mRNA m6A in control group and the L. plantarum colonization group (Supplementary Fig. 3a, b). Peak frequency plots indicated that m6A peaks were enriched in 3′UTR and near the stop codon (Supplementary Fig. 3c). We counted the m6A peak numbers of the control group and L. plantarum colonization group (Supplementary Fig. 3d), the different peaks were mainly distributed in 3’UTR and 1st Exon (Supplementary Fig. 3e). Motif analysis identified the consensus sequence DRACH (Supplementary Fig. 3f). Based on the different m6A peak-containing genes of L. plantarum colonization group compared to control group (Fig. 4d), we also found several neural pathways by GO enrichment analysis (Fig. 4e). These results confirmed that folate-producing L. plantarum could increase neural mRNA m6A methylation levels.
Folate and folate-producing L. plantarum up-regulated neural pathways through m6A modification
To investigate the function of folate regulating m6A in the host, we compared m6A peak-containing genes in neural pathway. After treating with folate and colonization of L. plantarum to Drosophila, the up-regulated m6A peak-containing genes in neural pathway were mainly at the mRNA 3’UTR position compared to control group. Similarly, after disrupting Dhfr gene in S2 cells, the down-regulated m6A peak-containing genes in neural pathway were also mainly at the mRNA 3’UTR position compared to control group (Fig. 5a). We also analyzed GO pathways enriched by differential m6A peak-containing genes. After intersecting the pathways of the three groups, we found 103 common pathways (Fig. 5b). The top-enriched pathways were related to neuron functions (Fig. 5c), which is consistent with the previous works reporting that folate can regulate neural gene expression.
a Normalized frequency of m6A peaks across different regions of mRNA in three groups, respectively. b Venn diagram showing the number of GO pathways among three groups. FDR < 0.05 and log2FC ≥ 1 were set as the significance thresholds. c GO enrichment analysis showing functional pathways of m6A peak-containing genes in three groups. d Venn diagram showing common and unique m6A peak-containing genes in three groups. FDR < 0.05 and log2FC ≥ 1 were set as the significance thresholds. e Representation of sema2b m6A peak from IP and Input in the 5′UTR of the sema2b transcript in three groups. Peaks were visualized using IGV.
We further intersected the genes of neuron-related pathways in each comparison, and we found three genes containing differential m6A peaks, including Sema2b, Shot, and the Tweek (Fig. 5d). We used the IGV to view the region of differential m6A peaks, and we found the changed region in Sema2b across each group (Fig. 5e). It has been reported that Sema2b is involved in the regulation of neuronal development30,31. Taken together, the profiles of MeRIP-seq data from three different sets of experiments (folate-fed Drosophila, L. plantarum-colonization Drosophila, Dhfr-interference in Drosophila S2 cells) confirmed that folate and folate-producing L. plantarum could up-regulated neural pathways through m6A modification, and Dhfr gene was shown to be involved into the regulatory process.
Folate increases brain mRNA m6A methylation levels in mice
After the observed effect of folate on mRNA m6A modification in Drosophila, we continued to explore the effect of folate on mRNA m6A modification levels in mammals. Mice were fed with folate to examine m6A changes in multiple tissues of male and female mice (brain, spleen, liver, kidney, heart, thymus, and testis). We found that METTL3 and METTL4 protein expression levels were significantly elevated in the brain of female and male mice after folate feeding (Fig. 6a, b), but not in other tissues (Supplementary Fig. 4).
a Western blot result of METTL3/METTL14 in brain from the four groups (Male-Control vs Male-Folate, Female-Control vs Female-Folate, n = 3 for each group). b Quantification of western blot analysis (n = 3). Mean ± SEM; ** P < 0.01, *** P < 0.001, measured with the unpaired, two-tailed Student’s t-test. c Normalized frequency of m6A peaks across different regions of mRNA in four groups, respectively. d Number of m6A peaks identifies in different groups. e Representative consensus motifs and corresponding P-values of m6A peaks identified in four groups. f Venn diagram showing common and unique GO pathways using mRNAs with up-regulated m6A peaks. g GO enrichment analysis showing functional pathways of different m6A peak-containing genes in four groups. h Venn diagram showing the common and unique differential neuron-related GO pathway in four groups. i Representation of Sema5b m6A peak from IP and Input in 5′UTR of the Sema5b transcript in four groups. Peaks were visualized using IGV.
By MeRIP-seq data analysis, peak frequency plots showed that m6A peaks were enriched in 3’UTR for both female and male mice (Fig. 6c, and Supplementary Fig. 5a). We found obviously up-regulated peaks in brain after feeding folate both in male and female mice, indicating up-regulate m6A levels by folate (Fig. 6d). The consensus sequence we identified in mice were DRACH, which was consistent with mammalian system (Fig. 6e).
Next, we analyzed GO pathways enriched for the genes with differential mRNA m6A peaks in male and female. We found 2584 up peaks and 1361 up peaks in female and male, with 879 peaks shared between females and males (Fig. 6f). In common pathways, we found that the top-enriched pathways were related to neuron functions (Fig. 6g, Supplementary Table S1). After folate treatment, up-regulated m6A peaks were also enriched in neural pathways both in male and female mice (Supplementary Fig. 5b–f). We identified 63 shared neuron-related pathways (Fig. 6h). Among these genes, Sema5b is the homolog of Sema2b gene in Drosophila (above-mentioned). IGV results showed that differential m6A levels are located in 5’UTR of Sema5b gene (Fig. 6i), which are consistent with the results from Drosophila.
Dhfr-mediated mRNA m6A methylation through folate in mammalian cells
It is well known that folate-producing bacteria mainly dwell in the gut, and mammals absorb most of the folate through the gut. Moreover, it has been reported that folate is involved in neural development2,32. Our above results indicated that the dihydrofolate reductase gene Dhfr might be involved in the regulatory process. Therefore, we used human colorectal carcinoma HCT116 cells and human glioblastoma U251 cells to validate the role of Dhfr in regulating m6A modification through folate. Dot blot assay showed that the overall m6A modification of mRNA was significantly elevated after folate treatment in two tested cell lines (Fig. 7a, b), and we found that protein expression level of METTL3 was significantly elevated after folate treatment (Fig. 7c, d). RT-qPCR experiments also revealed that mRNA expression level of Mettl3 was significantly elevated after folate treatment in two cell lines (Fig. 7e, f). To demonstrate the role of METTL3 in the upregulation of m6A modification by folate treatment, shRNA was transfected into two cell lines (HCT116 and U251) to knock down the expression of Mettl3. After successfully knocking down Mettl3 in cells, we treated the cells with folic acid and measured the levels of m6A modification using dot blot assay. The results showed that the expression of METTL3 was significantly knocked down (Supplementary Fig. 6a, b), and the m6A level in dot blot assays did not change (Supplementary Fig. 6b, d). These results indicated the regulatory role of folate on m6A is METTL3-depedent.
a Dot blot of mRNA m6A methylation levels in U251 cells treated with/o folate (n = 3). Results are normalized to methylene blue staining. b Dot blot of mRNA m6A methylation levels in HCT116 cells treated with/o folate (n = 3). Results are normalized by methylene blue staining. c Western blot analysis of METTL3 in U251 cells treated with/o folate (n = 3). d Western blot analysis of METTL3 in HCT116 cells treated with/o folate (n = 3). e RT-qPCR analysis of Mettl3 mRNA expression levels in U251 cells treated with/o folate (n = 3). f RT-qPCR analysis of Mettl3 mRNA expression levels in HCT116 cells treated with/o folate (n = 3). g RT-qPCR analysis of Dhfr mRNA expression levels in U251 cells treated with/o folate (n = 3). h RT-qPCR analysis of Dhfr mRNA expression levels in HCT116 cells treated with/o folate (n = 3). i RT-qPCR analysis of Dhfr mRNA expression levels in U251 cells from siNC, siDhfr and siDhfr + 5-MTHF group (n = 3). j Western blot analysis of METTL3 in U251 cells from siNC, siDhfr and siDhfr + 5-MTHF group (n = 3). k RT-qPCR analysis of Mettl3 mRNA expression levels in U251 cells from siDhfr and siDhfr + 5-MTHF group (n = 3). l Dot blot of mRNA m6A methylation levels in U251 cells from siNC, siDhfr and siDhfr + 5-MTHF group (n = 3). Results are normalized by methylene blue. Mean ± SEM; * P < 0.05, ** P < 0.01, *** P < 0.001, measured with the unpaired, two-tailed Student’s t-test.
Subsequently, we explored whether folate could increase mammalian m6A modification through carbon metabolic pathway which was enriched in Drosophila experiments. We first examined the mRNA expression level of Dhfr in these cells, there was a significant increase of Dhfr expression after folate treatment in two cell lines (Fig. 7g, h). Secondly, we successfully disturbed the expression of Dhfr in U251 cells (Fig. 7i), and set up control group (siNC), Dhfr gene knockdown group (siDhfr), 5-MTHF rescue group (siDhfr + 5-MTHF). As 5-MTHF is downstream of Dhfr in the pathway, we added it to U251 cells after interfering Dhfr and 5-MTHF did not increase the expression of Dhfr as expected (Fig. 7i). Western blot analysis demonstrated that METTL3 protein expression was significantly reduced after the knockdown of Dhfr, but METTL3 could be recovered after supplementation with 5-MTHF (Fig. 7j). RT-qPCR demonstrated that supplementation with 5-MTHF after Dhfr knockdown also led to a significant increase in mRNA expression in Mettl3 compared to siDhfr group (Fig. 7k). Finally, we used dot blot to detect the m6A modification level in each group, the results showed that m6A modification in siDhfr group was significantly decreased compared to siNC group; the m6A modification was significantly restored when 5-MTHF was supplemented to the cells after siDhfr (Fig. 7l). From all these results, we conclude that folate can increase mammalian m6A modifications, especially in the neural cells, via carbon metabolic pathway through Dhfr gene.
Taken together, our study demonstrated that folate can regulate neural mRNA m6A epitranscriptome in Drosophila and mammals through one carbon metabolism pathway. Mechanistically, folate can increase the expression of m6A-related proteins (METTL3, METTL14). Folate-producing L. plantarum can also affect host mRNA m6A modification in a similar pattern (Fig. 8).
Proposed working model in this study. The image is created by BioRender.
Discussion
Our previous studies found that gut microbes have a great influence on mRNA m6A methylation levels17,18,19, but the mechanisms by which gut microbes affect m6A modification are unclear. We speculate that gut microbes can affect host mRNA m6A modification through microbial metabolites. Besides, folate plays an important role in neural development and is associated with neurodegenerative diseases32,33, but its specific mechanism of action still needs to be thoroughly investigated. In this study, we demonstrated that the microbial metabolite folate and folate-producing L. plantarum can increase m6A modifications in both Drosophila and mammals. Mechanistically, we demonstrated that folate can significantly increase the expression of m6A-related proteins (METTL3, METTL4) and the dihydrofolate reductase gene Dhfr is involved in the regulatory process. L. plantarum has all the genes to synthesize folate34, it also has relatively high survival rate in the gastrointestinal35. Therefore, we selected L. plantarum as the folate-producing bacteria in our validation experiments.
Our results revealed that m6A methylation levels changed in Drosophila after L. plantarum colonization in wild-type Drosophila containing gut microbes. The effect of L. plantarum on m6A methylation levels may require interaction with specific strains in the gut. L. plantarum has been considered as a beneficial bacterium with applications in neurological diseases since it can reduce the production of trimethylamines TMA, alleviate cognitive deficits, and protect against neurological complications from brain infections36. In our study, MeRIP-seq analysis showed that L. plantarum-affected genes were indeed enriched in neural pathways, it is speculated that L. plantarum may also increase m6A modification of other pathways through different metabolites.
Previous studies have shown that folate can activate the metabolic regulator AMPKα and may promote oligodendrocyte survival and differentiation by promoting DNA synthesis37. Folate treatment also alleviates neuropathic pain and enhances recovery after spinal cord injury by methylating the promoter of matrix metallopeptidase 238. In our study, we found that folate regulates m6A modification, and the pathways enriched in MeRIP-seq were more obvious than those in RNA-seq. The ability of m6A modification to affect mRNA translation has been reported in many studies5,39,40,41. Together, folate may affect mRNA translation processes or other processes of neural genes by regulating m6A modifications via carbon metabolism.
Moreover, we found that folate can up-regulate the m6A modification of neural pathway genes in both male and female mice, with more pronounced impact in females. Gender differences in the expression of genes related to DNA methylation have also been reported in the hypothalamus, in which females also have higher expression levels than males42. Another study also found significant gender differences in DNA methylation sites as age changed43. The effect of folate on DNA methylation is indeed a different phenomenon in females and males, whether and how the effect of folate on mRNA m6A modification is gender-specific need to be explored in further.
Methods
Folate supplementation and L. plantarum colonization in Drosophila
The conventional Drosophila melanogaster strain w1118 was reared on yeast-glucose medium (1 L water, 100 g yeast, 100 g glucose, 1.2% agar, 0.1% potassium sorbate) at 25 °C, 70% relative humidity, and a photoperiod of 12 L: 12D. The collected Drosophila eggs were coated in the sterilized medium, and L. plantarum was added evenly dropwise to the coated Drosophila egg medium. The culture tubes were sealed with antibacterial sealing film and placed in the incubator.
Folate supplementation in mice
Three-week-old C57BL/6 J SPF mice (six mice for male and female, respectively) were purchased, and each gender of mice was randomly divided into two groups with three mice in each group. All mice had free access to a sterilized diet and water, and the corn bedding was regularly changed. Each mouse was fed with folate via gavage at a volume of 0.1 mL/day and a concentration of 500 μg/kg/day for 14 days. After 14 days, mice organs (including the brain, heart, liver, kidney, spleen, thymus, and testis) were dissected and collected. Mice organ samples were immediately frozen in liquid nitrogen and stored at −80 °C until use. The protocol for an animal experiment in this study was reviewed and approved by the Institutional Animal Care and Use Committee (IACUC) of South China Normal University (protocol code SCNU-SLS-2022-009). We have complied with all relevant ethical regulations for animal use.
Folate or 5-methyltetrahydrofolate supplementation in cultured cells
Drosophila S2 cells were maintained in the Wang lab and were cultured at 28 °C in SIM-SF (Gibco) insect cell culture medium with 10% fetal calf serum, and the cells were sub-cultured every three days. U251 cells were provided by Prof. Xiaodong Ma at South China Normal University and maintained in the Wang lab. HCT116 cells were provided by Prof. Hongsheng Wang at Sun Yat-sen University and maintained in the Wang lab. U251 cells and HCT116 were maintained in DMEM (Gibco) with 10% FBS at 37 °C in 5% CO2. At the confluence of 60 ~ 70%, 180 μL of folate or 5-methyltetrahydrofolate was taken from the stock (9 mg/mL) and added to the cell culture medium (2.5 mL). After 48 h, cells were suspended and centrifuged at 1500 rpm for 5 min at 4 °C to collect cell pellets. The harvested cells were used for subsequent protein or RNA extraction experiments.
Culture and colonization of L. plantarum
The strain L. plantarum (ATCC 8014) was purchased from the Guangdong Institute of Microbiology in China. For bacterial recovery, the MRS liquid medium was prepared in advance and sterilized at 121 °C for 15 min. The bacteria stock was taken out from −80 °C and slowly dissolved at 4 °C, MRS medium was then added to bacteria solution and cultured at 37 °C for 24 ~ 48 h. The bacteria passage was performed at a ratio of 5% for static culture when the OD600 value of the bacteria solution reached 1.
Western Blot analysis
Protein samples were lysed with RIPA lysate, centrifuged, the supernatant was collected. Protein concentration was determined by a BCA assay kit and protein samples were heated at 105 °C for 5 min. After SDS-PAGE, proteins were transferred to PVDF membranes at 100 V for 1.5 h. Membranes were blocked in 5% skim milk for 1.5 h at room temperature and incubated with primary antibodies including anti-METTL3 antibody (15073-1-AP, Proteintech), anti-METTL14 antibody (HPA038002, Merck), β-actin (30101ES60, Yeasen) at 4 °C overnight. The membranes were washed twice with TBST, followed by incubation with Horseradish Peroxidase Labeled goat anti-rabbit IgG (A0208, Beyotime Biotechnology) or Horseradish Peroxidase Labeled goat anti-mouse IgG (A0216, Beyotime Biotechnology) for 1.5 h at room temperature. Finally, the membranes were soaked in ECL luminescent solution and put into a chemiluminescence image analysis system to take the pictures. The intensities of protein bands were quantified using the ImageJ software and GraphPadPrism9.
Dot Blot analysis of m6A level in poly(A)-RNA
Total RNAs from tissues or cells were extracted using the Trizol method according to the manufacturer’s instructions. Poly(A)-RNA was purified from total RNA using Dynabeads™ mRNA Purification Kit (#61006, Invitrogen). An equal volume of poly(A)-RNA sample was heated at 95 °C for 3 min by heat shock and immediately cooled on ice. The denatured RNAs were dropped onto Amersham Hybond-XL (RPN303S, GE Healthcare) and air dry. After sampling, the nitrocellulose membrane was placed in a UV crosslinker and irradiated at 0.12 Joule for 5 min. The membrane was stained with SYBR™ Gold (S11494, Invitrogen) or 0.02% methylene blue solution. The membrane was photographed with a gel image analyzer, and ImageJ software was used to calculate the gray value of mRNA spots. The membrane was washed twice with Washing Buffer (1X PBS + 0.02% Tween-20) and blocked with 5% skim milk for 1 h at room temperature. The membrane was incubated with m6A antibody (E1610S, NEB) at 4 °C overnight followed by incubation with Horseradish Peroxidase Labeled goat anti-rabbit IgG (A0208, Beyotime Biotechnology) for 1 h at room temperature. Finally, the membrane was incubated with ECL luminescent solution and put into a chemiluminescence image analysis system to take the pictures. The intensities of protein bands were quantified using the ImageJ software and GraphPadPrism9.
Immunofluorescence
Drosophila S2 cells were grown in 12-well plates pretreated with the slides, and the cell culture medium was removed after folate treatment. Cells were fixed with 4% w/v formaldehyde (P0099, Beyotime), permeabilized with 0.5% v/v Triton X-100 (DH351-4, Dingguo Biotechnology Co., Ltd.) in phosphate-buffered saline (C0221A, Beyotime) for 30 min. The slides were blocked for 30 min at room temperature in a blocking buffer with 5% w/v goat serum (C0265, Beyotime). After blocking, the cells were incubated with rabbit anti-m6A antibody (E1610S, NEB) at 4 °C overnight and then incubated with a goat anti-rabbit IgG (H + L) Cross-Adsorbed Secondary Antibody, Alexa Fluor™ 488 (#A-11008, Invitrogen) for 1 h. The slides were mounted with nail polish and stained with DAPI Staining Solution (#ab228549, Abcam). A FLUOVIEW FV3000 confocal laser scanning microscope (Olympus, Shinjuku, Japan) was used to capture immunofluorescence images of the samples.
RT-qPCR analysis
Total RNAs were extracted with Trizol and NanoDrop 2000 spectrophotometer (ThermoFisher) was used to quantify the concentration of total RNA at a wavelength of 260 nm. RNA reverse transcription reactions were performed using the Hifair® III First Strand cDNA Synthesis Kit (#11139ES60, Yeasen). Three biological replicates were used for quantitative reverse transcription PCR analysis using HieffSYBR Mixture (#11201ES08, Yeasen). In S2 cells, the relative mRNA level of gene expression was measured with Rp49/Gapdh as an internal control and analyzed by the 2-∆∆Ct method. In U251 and HCT116 cells, the relative mRNA level of gene expression was measured with Gapdh as an internal control. Primer sequences are shown in Supplementary Table S2.
RNAi using dsRNA in S2 cells
The T7 RiboMAX™ Express RNAi System (#P1700, Promega, WI, USA) was used to synthesize dsRNA molecules following the manufacturer’s protocol. The dsRNA primers for dsDhfr were designed on the NCBI website. We also prepared dsRNA targeting green fluorescent protein (GFP) as the control for RNAi experiments. The pre-cultured S2 cells were transferred to a 6-well plate for adherent culture. RNA interference could be started when the cells grew to about 80%. A total of 20 μg dsRNA targeting dsDhfr and 10 μL FuGENE® HD Transfection Reagent (#E2311, Promega) were added to 100 μL serum-supplemented medium containing cultured S2 cells. After 48 h of incubation, cells were collected for RNA and protein extraction for subsequent experiments. The dsRNA sequences are shown in Supplementary Table S2.
RNAi using siRNA in human cells
3 × 105 HCT116/U251 cells were cultured in a 6 cm dish containing 4 mL of DMEM, and 24 h later the cells were transfected with Lipofectamine 2000 (Invitrogen) and 50 nM siRNA. After 7 h, the transfection mixture was removed, and the cells were replaced with normal medium or treated with reagents. 48 h later, the cells were harvested for protein or RNA for subsequent experiments. The siRNA sequences are shown in Supplementary Table S2.
RNAi using shRNA in human cells
3×105 HCT116/U251 cells were cultured in a 6 cm dish containing 4 mL of DMEM, and 24 h later the cells were transfected with Lipofectamine 2000 (Invitrogen) and 100 nM shRNA. After 7 h, the transfection mixture was removed, and the cells were replaced with normal medium or treated with reagents. 48 h later, the cells were harvested for protein or RNA for subsequent experiments. The shRNA sequences are shown in Supplementary Table S2.
Construction of RNA-seq libraries and m6A MeRIP-seq libraries
About 1.5 µg of purified poly(A)-RNA was fragmented into ~100 nt fragments using RNA fragmentation buffer (AM8740, Thermo Scientific). One-tenth of fragmented RNAs were used for the construction of RNA-seq libraries (Input) and the left RNAs were used for the construction of m6A MeRIP-seq libraries (IP). m6A IP was conducted using an EpiMark® N6-Methyladenosine Enrichment Kit (#E1610S, NEB) following the product manual. The resultant m6A-enriched and untreated poly(A)-RNA fragments were used to construct strand-specific IP and Input libraries, respectively. The VAHTS® Universal V6 RNA-seq Library Prep Kit for Illumina (NR604-02, Vazyme Biotech. Co., Ltd.) and VAHTS® RNA Adapters set3 for Illumina (N809, Vazyme Biotech. Co., Ltd.) provided the necessary reagents for first-strand and second-strand cDNA synthesis, adapter ligation, and library amplification. RNA-seq and MeRIP-seq of the prepared libraries were performed in Berry Genomics on the NovaSeq 6000 platform (Illumina, CA, USA) to obtain paired-end reads of 150 bp. Library quality was assessed on an Agilent Bioanalyzer 4200 TapeStation before loading onto the sequencer, ~8 G of raw reads was obtained for each library.
RNA-seq data analysis
Adapters and raw reads containing low-quality bases were first removed by applying trim_galore (version 0.6.6) to all RNA-seq raw data including the MeRIP-seq input libraries. The resulting reads of at least 35 bp were mapped to the reference genome (mm39, dm6-FLYBASE). The obtained comparison results were removed and sorted using samtools (version 1.3.1) for PCR duplicates. The featureCounts program44 in the Subread package (version 2.0.1) at SourceForge was used to count the reads that mapped to genes. The genes that showed an average read count of less than 10 in any group or lacked read count in any replicate were primarily filtered. The R package DESeq2 (version 1.36.0)45 was used for the identification of differentially expressed genes (DEGs) by setting a false discovery rate-corrected P value < 0.05 and log2FC > 0.5 as the thresholds for significance. R package DESeq2 was also used for read count normalization across samples by size factors and gene expression was measured based on normalized read counts.
MeRIP-seq data analysis
We first processed the raw data of MeRIP-seq IP libraries with the same upstream pipeline employed for RNA-seq. The mapping results of both MeRIP-seq IP and input libraries were used for calling m6A peaks. Differential m6A peaks of the single-factor comparisons between groups were analyzed using the R package exomePeak2 (version 1.8.1), with Poisson’s generalized linear model (GLM) used as a quantitative method to call the m6A peaks of individual groups as well as the m6A peaks of single-factor comparative differences between groups. To reduce false positives, the called m6A peaks from each biological replicate were considered significant and retained for subsequent analyzes according to the following thresholds: peak width < 1500 bp, log2FC > 1, FDR < 0.05, Input.RPM > 0.1, IP.RPM > 0.1. Similarly, the called differential m6A peaks between groups were considered significant under the following thresholds: peak width < 1500 bp, |diff.log2FC | >1, FDR < 0.05, Input.Control.RPM > 0.1, IP.Control.RPM > 0.1, Input.Treated.RPM > 0.1, IP.Treated.RPM > 0.1. HOMER (version 3.12) was used for de novo motif searching around peak summit-centered (mouse, peak 300 bp region; Drosophila, peak 250 bp) regions. The R packages ChIPseeker (version 1.32.0)46 and Guitar (version 2.12.0)47 were used for annotation and representation of the distribution characteristics of the significant/differential m6A peaks. The bamCoverage script in deeptools (version 3.5.1) was used to perform read-coverage trajectory calculations on the results of mapping IP libraries and Input libraries to the genome. Integrative Genomics Viewer (IGV) software (version 2.14.0)48 was used to display read coverage tracks of target genes.
Statistics and reproducibility
Biological replicates (n) are defined as experiments performed on independent biological sources. Repeated experiments for all statistics are labeled in the corresponding figure legend. Key experiments were independently repeated at least three times to ensure reliability and reproducibility. All data were expressed as the mean ± SEM and representative data were shown. A two-tailed unpaired Student’s t-test was employed to analyze the experimental data. The level of significance was set at P < 0.05; *P < 0.05; **P < 0.01; ***P < 0.001, ns: non-significant. Data and graphics were processed using GraphPad Prism 8.0.2 (GraphPad Software, La Jolla, CA, USA) and Adobe Illustrator 2024 (Adobe, San Jose, CA, USA).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
References
Rossi, M., Amaretti, A. & Raimondi, S. Folate production by probiotic bacteria. Nutrients 3, 118–134 (2011).
Prevention of neural tube defects: results of the Medical Research Council Vitamin Study. MRC Vitamin Study Research Group. Lancet 338, 131–137 (1991).
Barua, S. et al. Single-base resolution of mouse offspring brain methylome reveals epigenome modifications caused by gestational folic acid. Epigenet. Chromatin. 7, 3 (2014).
Burton, M. A., Antoun, E., Penailillo, R. S., Burdge, G. C. & Lillycrop, K. A. Folic acid induces intake-related changes in the mammary tissue transcriptome of C57BL/6 mice. Nutrients 12, 2821 (2020).
Zheng, Y. & Cantley, L. C. Toward a better understanding of folate metabolism in health and disease. J. Exp. Med. 216, 253–266 (2019).
Crider, K. S., Yang, T. P., Berry, R. J. & Bailey, L. B. Folate and DNA methylation: a review of molecular mechanisms and the evidence for folate’s role. Adv. Nutr. 3, 21–38 (2012).
Cao, R., Xie, J. & Zhang, L. Abnormal methylation caused by folic acid deficiency in neural tube defects. Open Life Sci. 17, 1679–1688 (2022).
Mosca, P., Leheup, B. & Dreumont, N. Nutrigenomics and RNA methylation: role of micronutrients. Biochimie 164, 53–59 (2019).
Song, J. & Yi, C. Chemical modifications to RNA: a new layer of gene expression regulation. ACS Chem. Biol. 12, 316–325 (2017).
Morscher, R. J. et al. Mitochondrial translation requires folate-dependent tRNA methylation. Nature 554, 128–132 (2018).
Xu, X. et al. Folate regulates RNA m(5)C modification and translation in neural stem cells. BMC Biol. 20, 261 (2022).
Frye, M., Harada, B. T., Behm, M. & He, C. RNA modifications modulate gene expression during development. Science 361, 1346–1349 (2018).
Shafik, A. M. et al. N6-methyladenosine dynamics in neurodevelopment and aging, and its potential role in Alzheimer’s disease. Genome Biol. 22, 17 (2021).
Li, N. et al. The effects of folic acid on RNA m6A methylation in hippocampus as well as learning and memory ability of rats with acute lead exposure. J. Funct. Foods 76, 104276 (2021).
Jabs, S. et al. Impact of the gut microbiota on the m(6)A epitranscriptome of mouse cecum and liver. Nat. Commun. 11, 1344 (2020).
Chen, S. et al. Fusobacterium nucleatum reduces METTL3-mediated m(6)A modification and contributes to colorectal cancer metastasis. Nat. Commun. 13, 1248 (2022).
Wang, X. et al. Transcriptome-wide reprogramming of N(6)-methyladenosine modification by the mouse microbiome. Cell Res. 29, 167–170 (2019).
Yang, M. et al. Antibiotic-induced gut microbiota dysbiosis modulates host transcriptome and m(6)A epitranscriptome via bile acid metabolism. Adv. Sci.11, e2307981 (2024).
Xu, Z. et al. Microbiome-induced reprogramming in post-transcriptional landscape using nanopore direct RNA sequencing. Cell Rep. 43, 114798 (2024).
Friso, S., Udali, S., De Santis, D. & Choi, S. W. One-carbon metabolism and epigenetics. Mol. Asp. Med. 54, 28–36 (2017).
Matherly, L. H., Hou, Z. & Deng, Y. Human reduced folate carrier: translation of basic biology to cancer etiology and therapy. Cancer Metastasis Rev. 26, 111–128 (2007).
Elnakat, H. Ratnam M. distribution, functionality and gene regulation of folate receptor isoforms: implications in targeted therapy. Adv. Drug Deliv. Rev. 56, 1067–1084 (2004).
Zhao, R., Diop-Bove, N., Visentin, M. & Goldman, I. D. Mechanisms of membrane transport of folates into cells and across epithelia. Annu Rev. Nutr. 31, 177–201 (2011).
Matherly, L. H. & Goldman, D. I. Membrane transport of folates. Vitam. Horm. 66, 403–456 (2003).
Kostic, A. D., Xavier, R. J. & Gevers, D. The microbiome in inflammatory bowel disease: current status and the future ahead. Gastroenterology 146, 1489–1499 (2014).
Moran, R. G., Colman, P. D., Rosowsky, A., Forsch, R. A. & Chan, K. K. Structural features of 4-amino antifolates required for substrate activity with mammalian folylpolyglutamate synthetase. Mol. Pharm. 27, 156–166 (1985).
Menezo, Y., Elder, K., Clement, A. & Clement, P. Folic acid, folinic acid, 5 methyl tetrahydrofolate supplementation for mutations that affect epigenesis through the folate and one-carbon cycles. Biomolecules 12, 197 (2022).
Donnelly, J. G. Folic acid. Crit. Rev. Clin. Lab Sci. 38, 183–223 (2001).
Tamene, A. et al. Lactobacillus plantarum P2R3FA Isolated from traditional cereal-based fermented food increase folate status in deficient rats. Nutrients 11, 2819 (2019).
Orr, B. O., Fetter, R. D. & Davis, G. W. Retrograde semaphorin-plexin signalling drives homeostatic synaptic plasticity. Nature 550, 109–113 (2017).
Wu, Z. et al. A combinatorial semaphorin code instructs the initial steps of sensory circuit assembly in the Drosophila CNS. Neuron 70, 281–298 (2011).
van Gool, J. D., Hirche, H., Lax, H. & De Schaepdrijver, L. Folic acid and primary prevention of neural tube defects: a review. Reprod. Toxicol. 80, 73–84 (2018).
Shulpekova, Y. et al. The concept of folic acid in health and disease. Molecules 26, 3731 (2021).
Gilbert, J. A. et al. Microbiome-wide association studies link dynamic microbial consortia to disease. Nature 535, 94–103 (2016).
van den Nieuwboer, M., van Hemert, S., Claassen, E. & de Vos, W. M. Lactobacillus plantarum WCFS1 and its host interaction: a dozen years after the genome. Micro Biotechnol. 9, 452–465 (2016).
Wu, Y. et al. Lactobacillus plantarum-derived postbiotics prevent Salmonella-induced neurological dysfunctions by modulating gut-brain axis in mice. Front. Nutr. 9, 946096 (2022).
Weng, Q. et al. Folate metabolism regulates oligodendrocyte survival and differentiation by modulating AMPkalpha activity. Sci. Rep. 7, 1705 (2017).
Miranpuri, G. S. et al. Folic acid modulates matrix metalloproteinase-2 expression, alleviates neuropathic pain, and improves functional recovery in spinal cord-injured rats. Ann. Neurosci. 24, 74–81 (2017).
Choi, S. W., Friso, S., Keyes, M. K. & Mason, J. B. Folate supplementation increases genomic DNA methylation in the liver of elder rats. Br. J. Nutr. 93, 31–35 (2005).
Huang, H. et al. Recognition of RNA N(6)-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat. Cell Biol. 20, 285–295 (2018).
Wang, X. et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117–120 (2014).
Cisternas, C. D., Cortes, L. R., Bruggeman, E. C., Yao, B. & Forger, N. G. Developmental changes and sex differences in DNA methylation and demethylation in hypothalamic regions of the mouse brain. Epigenetics 15, 72–84 (2020).
Yusipov, I. et al. Age-related DNA methylation changes are sex-specific: a comprehensive assessment. Aging 12, 24057–24080 (2020).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Yu, G., Wang, L. G. & He, Q. Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Cui, X. et al. Guitar: an R/bioconductor package for gene annotation guided transcriptomic analysis of RNA-related genomic features. Biomed. Res. Int. 2016, 8367534 (2016).
Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative genomics viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform 14, 178–192 (2013).
Acknowledgements
We thank all members from the Wang lab for technical assistance and insightful discussion. This work was supported by the National Natural Science Foundation of China (32070615), Guangdong Provincial Natural Science Foundation (2022A1515010569), Guangzhou Science and Technology Project (2024A04J6265), and Guangdong Province Universities and Colleges Pearl River Scholar Funded Scheme for X.W. This work was also partially supported by Science and Technology Planning Project of Guangdong Province, China (2023B1212060050).
Author information
Authors and Affiliations
Contributions
Conceptualization: X.W. Methodology: A.L., Z.H. and Z.X. Investigation: A.L., Z.H., Z.X., S.L., Y.J., R.L. Y.L. and M.Y. Visualization: A.L., Z.X., M.Y. and J.H. Supervision: X.W. Writing: A.L., X.W. and C.H.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Arne Klungland and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Kaliya Georgieva. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, A., Huo, Z., Xu, Z. et al. Transcriptome-wide regulation of folate on neural mRNA m6A methylome via carbon metabolism in Drosophila and mammals. Commun Biol 8, 1151 (2025). https://doi.org/10.1038/s42003-025-08623-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-08623-6
This article is cited by
-
The role of N6-methyladenosine associated genes in ischemic stroke risk: interplay with environmental factors
Journal of Thrombosis and Thrombolysis (2025)