Abstract
Iron-sulfur clusters are essential cofactors for the accurate cellular function of many proteins. In eukaryotic cells, the biogenesis of most iron-sulfur clusters occurs in the mitochondria and involves the action of the Cys desulfurase supercomplex, which is activated by the protein frataxin (FXN). The decrease of FXN expression and/or function results in Friedreich’s ataxia (FRDA).
In this work, several nanobodies specific to human FXN were selected via phage display, demonstrating a wide range of effects on Cys desulfurase activity and a strong interaction with FXN. Nanobody interaction stabilized wild-type and FRDA-related FXN variants in vitro. FXN-nanobody complexes were characterized by NMR, SAXS, and X-ray crystallography. Additionally, Nanobody expression was studied in human cells. The subcellular localization, direct interaction with FXN by in situ proximity ligation assay, effect on cell viability, Fe-S-dependent enzymatic activities, and oxygen consumption rates were analyzed. Significantly, nanobody expression did not alter these key metabolic variables, suggesting that the interaction with FXN did not disrupt the pathway.
As a whole, our results suggest that nanobodies can serve as binding partners for mitochondrial FXN. However, the specific effect of the nanobodies on the conformational stability of FRDA-related FXN variants in cells should be investigated.

Similar content being viewed by others
Introduction
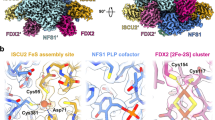
The study of iron-sulfur cluster biosynthesis at the molecular level has recently increased our understanding of some of the relationships between structure and function concerning the mitochondrial supercomplex involved in this pathway. The Fe-S clusters are at the core of many essential biological activities. Mitochondrial enzymatic functions such as NADH dehydrogenase, aconitase (ACO), succinate dehydrogenase (SDH), and complex III from the respiratory chain, lipoic acid synthetase, and processes such as nuclear DNA repair1, the cytoplasmic chemical modifications of transfer RNAs that affect base recognition between codon and anticodon2, and the cytoplasmic Fe-S cluster assembly by the cytosolic iron-sulfur cluster assembly system require the mitochondrial cysteine desulfurase supercomplex3,4. This supercomplex consists of several essential proteins, all of which are encoded by nuclear DNA (Fig. 1A, B)5. NFS1 is a pyridoxal phosphate-dependent cysteine desulfurase6,7,8 that employs Cys381 to transfer a persulfide group from the Cys substrate to the ISCU. Even though NFS1 is crucial for mitochondrial Fe-S cluster biosynthesis, it has been found in the cytosol, providing the sulfur for molybdenum cofactor biosynthesis9,10 and, more recently, it has been found at the centrosome11. ISCU is an essential scaffold protein that supports the assembly of the [2Fe-2S] cluster at its active site12,13 and is in close contact with the Cys-loop of NFS1, where the key Cys381 is located14. L-cysteine increases the binding efficiency of human ISCU2 to NFS1 nine-fold KD = 0.2 or 1.7 µM, with or without cysteine, respectively14. Additionally, NFS1 is stabilized in its active dimeric form by a small protein called ISD11 (10.5 kDa), which is essential in eukaryotes and prevents NFS1 aggregation15,16. In turn, ISD11 is stabilized by the acyl carrier protein (ACP)17,18. Furthermore, the ferredoxin system provides a natural source of electrons involving NADPH, FdxR, and FDX219,20,21,22,23.
A, B Two views of the cryoEM structure of the Cys desulfurase supercomplex (PDB ID: 6NZU) are presented. Cys desulfurase NFS1, ACP, ISD11, ISCU2, and FXN are shown in a surface representation. Only part of the supercomplex is shown in (C), the Cys381 of the NFS1 enzyme, ISCU2, and FXN are shown—additionally, the PLP cofactor, FXN Trp155, and ISCU2 assembly site. The distance between PLP and Cys381 is ∼15 Å, and between Cys381 and the ISCU2 assembly site it is ∼15 Å. ISD11 and ACP subunits were omitted for clarity. D A nanobody (VHH domain) structure scheme shows the three complementarity-determining regions: CDR1, CDR2, and CDR3. The N- and C-termini are indicated.
Another essential protein that forms the supercomplex is frataxin (FXN), the kinetic activator of NFS1, which accelerates the persulfide transfer24. FXN is imported into the mitochondrial matrix as a 210-residue precursor and processed in two steps to produce the mature FXN form (14.2 kDa), which comprises residues 81-21025,26. Current structural (Cryo-EM) and functional studies suggest that FXN simultaneously interacts with ISCU2 and with both subunits of the NFS1 dimer (Fig. 1A–C)5.
Notably, altering the function of NFS1, ISCU, or ISD11 results in rare diseases27,28,29,30. In particular, the decrease of FXN expression results in Friedreich’s ataxia (FRDA), an inherited neurodegenerative rare disease affecting 1 in 50,000 people31. In approximately 95% of FRDA patients, the transcription of the FXN gene is significantly reduced due to a trinucleotide expansion in the first intron of both alleles. Expansion ranges from 70 to more than 1000 GAA triplets in patients (700–800 repeats in most cases), while the first intron of the FXN gene in normal chromosomes contains up to 35 to 40 GAA triplets32. In the remaining 5–6% of the FRDA patients, a point mutation is found in one FXN allele combined with an expansion in the other. These mutations, in some cases, alter the stability of the protein in vitro and/or in vivo (D122Y, G130V, G137V, W173G, L198R, among others33,34,35,36), the localization and processing of the protein (R40C)37, the internal motions (L198R and CTR truncation36,38), or, as in the case of W155R or N146K, the interactions with the rest of the proteins that conform the supercomplex, dramatically reducing the activation function of FXN39.
In this context, one of our main research goals is to identify specific small binder partners that can interact with FXN in the mitochondrial environment.
We have named this strategy quaternary addition40. Recently, we selected affi_224, a synthetic FXN binder, employing the Ribosome Display strategy and archaeal Sac7D protein as the scaffold (Affitins)40. Affi_224, fused to a human mitochondrial targeting sequence, was successfully expressed in HEK-293T cell lines and interacted with FXN40.
Moreover, the presence of Affi_224 slightly increased the desulfurase activation exerted by the FXN G130V variant in vitro. Conversely, other in vitro experiments demonstrated that Affi_224 did not form a stable complex with FXN and is an inherently unstable protein that tends to form dimers. These characteristics may limit its applicability in further studies.
With this background, we explored additional scaffolds for FXN binders and chose the camelid-derived nanobody scaffold (VHH or nanobodies (NBs)). Highly specific NBs can be obtained by phage display strategy from an RNA library of camelid antibody sequences prepared after a convenient immunization of the camelid with the target protein. Camelid heavy chain antibodies are formed by two heavy chains, each one containing a variable domain (VHH), which includes three complementarity-determining regions (CDR1, CDR2, and CDR3, the latter longer than the others) (Fig. 1D). NBs are small (15 kDa) and conformationally stable proteins41 that can be expressed in E. coli. Their small size may allow for better tissue penetration42. Even though NBs usually display low immunogenicity, they can be humanized to further reduce this characteristic, making them suitable for therapeutic applications in humans43. NBs can be expressed by mammalian cells as “intrabodies” while retaining their conformational stability. An advantage of NBs is that they can be delivered by gene therapy as single genes. Treatments based on the NB for various neurodegenerative disorders and cancers are currently under development, with several in clinical trials41,44,45.
In this paper, we selected over twenty FXN-specific NBs and thoroughly investigated the interaction between FXN and four of these in both in vitro and cellular environments.
Results
Sequence analysis and structure predictions of the NB/FXN complexes
After the phage display selection, we performed an initial evaluation of NBs by assessing their interaction with recombinant FXN using a direct ELISA. FXN was bound to the plate and incubated with samples containing periplasmic proteins, which were obtained from the osmotic shock of bacteria expressing the NBs in the periplasmic compartment. Subsequently, a second selection was conducted to specifically assess the modulation of Cys desulfurase activity within the entire supercomplex (NFS1/ACP-ISD11/ISCU2/FXN)2 after the addition of the same samples containing the NBs into the enzymatic reaction. None of the samples resulted in overactivation of the desulfurase enzyme.
Intending to study a broad range of potential effects on FXN function, we selected NBs that at 1:1 NB:FXN ratio, (a) either do not inhibit or minimally inhibit the desulfurase enzyme, (b) exhibit a moderate inhibitory effect, or (c) cause high in vitro inhibition.
After the screening, 30 NBs specific for human FXN were initially sequenced. We found substantial sequence diversity in the CDR1, CDR2, and CDR3 regions (Fig. 1D and Supplementary Fig. 2A), resulting in 16 distinct sequences. Eleven NBs were chosen for protein expression in E. coli WK6 strain, purification, and in vitro studies. Four NBs (NB_4A7, NB_6B1, NB_16C10, and NB_28F6, Supplementary Fig. 1 and Supplementary Table 1) and the corresponding NB:FXN complexes were deeply studied in this work.
Alpha-Fold 346,47,48 was used to infer structures of NB:FXN complexes from their amino acid sequences. The results suggested many different types of complexes NB:FXN might be formed (Supplementary Fig. 2C). Furthermore, predictions indicated that some NBs might display an inhibitory effect because the binding region comprises a surface of FXN involved in the supercomplex architecture (e.g., an ISCU2 or NFS1 interacting surface). On the other hand, some NBs might interact with FXN surface areas that are not involved in inter-subunit interactions. In those cases, NBs may not affect the activation exerted by FXN.
NB expression in E. coli WK6
NBs were purified by NTA-Ni2+, yielding 95% pure protein (5–15 mg/L, Supplementary Fig. 1). In all cases, monomeric conformations were obtained, as judged by SEC-FPLC (see below). For NB_28F6, protein degradation was observed when the NB was stored at 4 °C. Thus, we decided to preserve the NBs at −70 °C. Apart from NB_28F6, which exhibited proteolysis, the masses of the purified NBs were in the 2 Da range of the expected masses, as deduced from the amino acid sequences, considering the predicted processing site after NB export to the periplasmic space (the processing of the signal peptide results in an N-term QVQLQ, Supplementary Fig. 2A and Supplementary Table 1).
NB effect on supercomplex activity in vitro
After NB purification, we investigated the effect of NBs addition to the enzymatic reaction on the L-Cys desulfurase catalysis (Fig. 2A and Supplementary Fig. 3). The NBs NB_4A7 and NB_16C10 exhibited a low to middle degree of inhibition, while NB_6B1 and NB_28F6 showed higher inhibition of the in vitro L-Cys-desulfurase activity. It is worth noting that the complex (NFS1/ACP-ISD11/ISCU)2 exhibits lower activity than the core (NFS1/ACP-ISD11)2. One can infer ISCU interaction with (NFS1/ACP-ISD11)2 from the observed decrease in desulfurase activity49. Higher NB concentrations of NB_6B1 and NB_28F6 produced the highest decrease in the activity of the supercomplex, like the one detected in the virtual absence of FXN, which represents a drop in the value corresponding to (NFS1/ACP-ISD11/ISCU)2. On the other hand, the inhibitory effect of higher concentrations of NB_16C10 was lower than that observed for NB_6B1 and NB_28F6, suggesting that NB_16C10 affects the conformation or the topology of the supercomplex but would not impede FXN-supercomplex interaction in these experimental conditions. Remarkably, NB_4A7 exhibited a low inhibition even at high NB concentrations, which is indicative of the formation of an activated supercomplex in the presence of the NB (Fig. 2B–E).
A The effect of the addition of NBs on the Cys-desulfurase supercomplex activity for NB_4A7, NB_6B1, NB_16C10, and NB_28F6). “NIAUF” refers to the reaction, including NFS1/ACP-ISD11/ISCU/FXN, in the absence of NB. NIA refers to a sample without ISCU2, FXN and NB, and corresponds to the activity of the core (NFS/ACP-ISD11)2, whereas NIAU corresponds to the core plus ISCU2, but without FXN. “Without Pro” corresponds to a reaction in the absence of proteins. We also studied 5:1 molar ratio. B NB_4A7, C NB_6B1, D NB_16C10, and E NB_28F6. In (A), the activity was relativized to that of NIAUF. At least two independent experiments (n = 2), including three technical replicates for each. Error bars represent the standard errors.
In vitro characterization of the NB:FXN interaction
To evaluate the in vitro NB:FXN complex formation, we first studied the interaction by size-exclusion chromatography (SEC). Purified NBs, FXN, or the mix of both proteins (injection after a 10-min incubation of FXN and NB at room temperature) were loaded in an analytical SEC system. As judged by the SEC profiles, many of the selected NBs against FXN formed stable complexes (Fig. 3A–D and Supplementary Fig. 4), and their profiles suggested a slow dissociation equilibrium. Besides this, the SEC profile corresponding to NB_15C5 was compatible with the absence of complex formation, exhibiting two peaks consistent with free FXN and free NB (Supplementary Fig. 4C).
The interaction between NBs and FXN was followed by SEC for A NB_4A7, B NB_6B1, C NB_16C10, and D NB_28F6. The SEC profiles corresponding to the isolated FXN (black and gray, having three different concentrations at 2, 3, and 6 μM) or NBs (color lines). Dashed lines show the profiles corresponding to a 1:1 NB:FXN mixture. The lower elution volume for the mixture indicates the formation of a stable complex. The interaction followed by BLI. Biotin-labeled FXN variant (H177C) was titrated with each NB. Representative binding assay corresponding to E NB_4A7, F NB_6B1, G NB_16C10, and H NB_28F6 are shown. The experiment was performed in a 25 mM Tris-HC, 150 mM NaCl, pH 7.4 buffer, and 0.5 mg mL−1 BSA was added in all solutions to avoid unspecific binding of FXN and NB to the sensor and tube surfaces. NB solutions typically contain concentrations ranging from 0 to 150 nM NB. The global fitting is represented by black lines.
The binding between NBs and FXN was studied employing biolayer interferometry (BLI). To evaluate the interaction, a biotin-labeled FXN variant (H177C, Supplementary Fig. 5) was immobilized in a streptavidin sensor at 10 μg mL−1. The selected NBs exhibited high affinity with equilibrium dissociation constant KD in the nanomolar range 1–33 nM (Table S2), and association and dissociation kinetic rate coefficients ka and kd in the range 105 M−1 s−1 and 10−4 M, respectively (Fig. 3E–H and Supplementary Table 2).
Identification of the NB binding site on the FXN surface
The binding sites of this subset of NBs were studied with a resolution at the level of the amino acid residues by NMR, analyzing the chemical shift perturbations (CSP). FXN was produced in E. coli using the 15N source 15NH4Cl and purified for these experiments. The target was then titrated with each of the NBs, and 1H-15N-HSQC NMR bidimensional spectra were acquired (Fig. 4A–N and Supplementary Fig. 6–9). Then, CSP and changes in the cross-peak intensities were analyzed (Fig. 4D, H, L, P).
FXN was titrated with each NB at molar ratios of 0:1, 0.33:1, 0.66:1 and 1:1 (NB:15N-FXN), shown in blue, cyan, magenta and light green, respectively; A, B: NB_4A7, E and F: NB6B1, I and J: NB_16C10, M and N: NB_28F6 correspond to selected regions from the FXN spectra illustrating some of the residues exhibiting different patterns of CSP after incubation with the NB for each complex. C: NB_4A7, G: NB_6B1, K: NB_16C10, and O: NB_28F6, the residues exhibiting the highest CSP were mapped on the FXN structure (using a van der Waals style of representation for the residues involved and colored by element). Ribbon representation is colored as follows: alpha helix, beta strands, turns, and coils in blue, red, green, and cyan, respectively. D: NB_4A7, H: NB_6B1, L: NB_16C10, and P: NB_28F6, the calculated CSP values of amide peaks according to \({\Delta \delta }_{{{\mathrm{aV}}}}={\left[\left({\delta }_{{{{\rm{H}}}}}^{2}+\,{\delta }_{{{{\rm{N}}}}}^{2}/25\right)/2\right]}^{1/2}\). Gray bars in the right panels indicate those residues for which the CSP could not be calculated because the H-N assignments of the bound state were not determined. The complete spectra are shown in Supplementary Figs. 6–9. The experiment was performed in a 25 mM Tris-HCl, 150 mM NaCl, pH 7.4 buffer. 1H-15N HSQC experiments were performed. A list of the residues involved is provided in the Supplementary Table 3.
CSP results indicated that NBs NB_4A7, NB_6B1, and NB_16C10 bind to a similar site on the FXN surface (residues involved are shown in Supplementary Table 3). Instead, NB_28F6 presents a different binding mode (Fig. 4C, G, K, O), as indicated by the residues involved in the interaction. As judged by the discontinuous pattern of CSP observed for almost all the cross peaks involved, the results suggested, in all cases, a binding mechanism characterized by a slow exchange regime. Remarkably, some residues from the FXN core exhibited significant CSP, including aromatic side chains Phe110, Tyr123, and Phe127 and the aliphatic residue Leu113. This suggests that the interaction between NB and FXN may adjust some conformational details of the FXN structure. In the case of NB_28F6, some residues far from the suggested interaction site also exhibited high CSP (Supplementary Table 3). Among them, Tyr143, Ile145, and Trp155, the latter involved in the assembly site of the supercomplex.
Modulation of the FXN conformational stability by NB interaction
To test whether NB interactions could stabilize the FXN conformation, we conducted in vitro temperature-induced unfolding experiments. Protein denaturation was monitored by the fluorescence of the Sypro-orange probe. Briefly, when the dye interacts with the unfolded state of proteins, the quantum yield increases. For these experiments, we first used FXN G130V as a probe for highly unstable FRDA variants50,51.
Notably, the NBs were able to significantly stabilize the G130V FXN variant as judged by a significant shift (≥15 °C) in the observed Tm values (e.g., Tm values are 51.8 ± 0.3 and 70.1 ± 0.1 °C, for the G130V variant and NB_4A7: G130V_FXN complex, respectively, Fig. 5A, E). The Tm value corresponding to the complex is even higher than that observed for the wild-type FXN (Tm = 65.9 ± 0.3 °C). Similar results were obtained for the rest of the NBs (Fig. 5B–E). On the other hand, Sypro-orange dye did not exhibit a significant interaction with the NBs used in this assay at the range of temperatures tested (blue lines in Fig. 5A–D).
Experiments were carried out on a multi-well plate, and Sypro-orange fluorescence was monitored. Wild type and G130V FXN variants without NB were included as controls in black (Tm = 65.9 ± 0.3 °C) and gray (Tm = 51.8 ± 0.3 °C), respectively. A–D correspond to NB_4A7, NB_6B1, NB_16C10, and NB_28F6, respectively. Tm values for the complexes are 70.1 ± 0.1, 68.7 ± 0.1, 69.9 ± 0.1 °C, and 65.9 ± 0.1 °C for NB_4A7: G130V, NB_6B1: G130V, NB_16C10: G130V, and NB_28F6:G130V complexes, respectively). E Tm values for FXN and G130V variants and the complexes FXN G130V:NB. Stabilization of the variants followed by the thermal shift assay. Wild type (WT), D122Y, G130V, G137V, W155R, and L198R variants were incubated with NB_4A7 (F) or NB_28F6 (G). For each FXN variant, the ΔTm value was calculated as ΔTm = Tm(FXN+NB) − Tm(FXN).
For the NB_4A7 and G130V variants, control experiments demonstrated that both proteins are fully folded, as indicated by circular dichroism spectra in the Near- and Far-UV regions (Supplementary Fig. 10A, B). Moreover, an additional temperature-induced unfolding experiment monitored by circular dichroism at 222 nm (Supplementary Fig. 10C), which reflects a change in secondary structure content as the temperature increases, yielded results similar to those previously observed with Sypro-Orange. The measured Tm values were 50, 65, and 69 °C for FXN G130V, NB_4A7:FXN G130V, and NB_4A7 alone, respectively. Thus, the addition of NB_4A7 resulted in a 15 °C increase in the Tm value of the variant.
Next, we evaluated the capability of NB_4A7 to stabilize other unstable FRDA-related variants. Remarkably, the NB_4A7 interaction stabilized W155R and L198R variants (ΔTm = 11 and 17 °C, respectively). Moreover, NB_4A7 stabilized the wild-type FXN variant (ΔTm = 7 °C). In turn, the stabilization for D122Y and G137V was visibly lower (ΔTm = 3 and 4 °C, respectively) (Fig. 5F), although NB_28F6 was indeed able to stabilize these variants (ΔTm = 12.8 and 21 °C, respectively) (Fig. 5G).
Three-dimensional structure of NB:FXN complexes
To get high-resolution information on the binding mode of the NBs studied in this work, NB_4A7, NB_6B1, and NB16C10 in complex with wild-type FXN (90-210 variant) were crystallized, and their structures solved (Fig. 6). The three crystallographic complexes were solved at atomic resolutions of 1.25 Å (FXN: NB_4A7), 1.48 Å (FXN: NB_6B1), and 2.0 Å (FXN: NB_16C10). The quality of the electron density map was excellent in all three cases (Supplementary Figs. 11 and 12), allowing unambiguous modeling of both proteins and fully identifying the residues involved in protein-protein interaction. FXN residues involved in the interaction are listed in Supplementary Table 4. The structure of human FXN in our complexes presents a nearly identical structure to that reported previously (PDB ID: 1EKG)52, with RMSD values ranging from 0.21 to 0.24 Å (for superimposition of all Cα atoms). FXN presents two parallel α-helices supported by a platform provided by a five-stranded, antiparallel β-sheet. The NBs adopt the typical immunoglobulin fold, with ten β-strands forming two β-sheets connected by loops and a conserved disulfide bond between Cys22 and Cys95 (Cys95 in NB_4A7 and NB_16C10 and Cys98 in NB_6B1) (Supplementary Fig. 2A). The NB:FXN crystal structures confirmed the interaction surface and binding regions observed by NMR experiments (CSP results, Fig. 5 and Supplementary Table 3). In all cases, the two proteins in the complex are arranged in a T fashion mode with FXN orienting the acidic ridge (the loop linking α1 and β1, residues Glu114-Glu122), the loop β2-β3 (residues 135-139) and the tip of helix α1, against the central part of the NB β-sandwich. Specifically, the NBs interact with frataxin through the loop β3-β4 (residues Arg38-His46, using Supplementary Fig. 2A numbering), the β4-strand (residues Leu47-Arg50), and the loop β5-β6 (residues Asp61-Lys64); NB_4A7 and NB_16C10 also interact through Asn58 from β4 (Fig. 6A, C). Remarkably, the CDR regions of the NBs are not directly involved in binding to FXN. Some relevant electrostatic interactions are observed the NB:FXN complexes (e.g., Glu121FXN-Arg38NB, Lys135FXN-Asp61/64NB, Asp139FXN-Lys64/67NB, Phe120FXN-Arg50NB) (Fig. 6A–C and Supplementary Table 5), involving a molecular contact surface area of 581.7 Å2. Structural comparison of FXN alone and in complex with NB (Supplementary Fig. 12) reveals that major changes upon NB interaction are in Loop 1, L1, connecting α1-β1, and the stretch connecting β2–β3, L2, (Fig. 6A–C and Supplementary Fig. 12). While some variations are also observed in β5-β6 turn and the loop connecting β6 and α2, depending on the NB. Overall, a similar pattern of interactions is conserved for the interaction of FXN with NB_4A7, NB_6B1, and NB_16C10, where loops L1 and L2, as well as a punctual interaction through helix α2, are pivotal for protein-protein recognition.
A Cartoon representation of the FXN:NB_16C10 complex. Boxed areas show the most critical interactions between the NB_16C10 and FXN, box 1 colored in red and box 2 in black. Relevant residues are depicted as capped sticks and labeled. Polar interactions are represented as dotted lines. B Cartoon representation of the FXN:NB_6B1 complex. Boxed areas show the most important interactions between the NB_6B1 and Frataxin. C Cartoon representation of the FXN:NB_4A7 complex. Boxed areas show the most important interactions between the NB_4A7 and FXN.
Mutations D122Y50 and G137V35 found in two FRDA-related FXN variants occur in residues located on the interface of the NB:FXN complexes (Supplementary Fig. 13). As previously mentioned, we observed that the interaction between NB_4A7 and G137V or D122Y resulted in only a slight increase in the Tm value, consistent with a weaker interaction (Fig. 5F). Remarkably, the interaction between NB_4A7 and the D122Y or G137V variants could not be detected by BLI (Supplementary Fig. 13), suggesting that these mutations can locally alter the conformation of the FXN at the interaction site of NB_4A7. On the other hand, NB_4A7 interacted with G130V, W155R, or L198R variants (Supplementary Fig. 13).
SAXS studies on FXN:16C10 and FXN:28F6 complexes in solution
The complex FXN:NB_28F6 did not produce crystals in our hands; thus, we decided to perform small angle X-ray scattering (SAXS, Supplementary Table 6) studies to check if the interaction would follow or not the interaction pattern observed for NBs NB_16C10, NB_6B1, and NB_4A7. The complex FXN:NB_16C10 was also measured for comparison with our crystal structures. In both cases, there is a good superimposition of the experimental and theoretical scattering curves (Fig. 7A). As expected, for the FXN:NB_16C10 complex, the SAXS envelope in solution fits with the crystal structure of the complex (Fig. 7D) with the two proteins oriented in a T-fashion mode. However, the FXN:28F6 complex shows a new disposition in which the two proteins pack in a parallel fashion with the β-sheets from both proteins configured as a single long β-sheet (Fig. 7C). The model observed in the solution for the FXN:NB_28F6 complex is compatible with the predicted arrangement provided by AlfaFold3 (Supplementary Fig. 2C). As discussed below, this different arrangement for the NB_28F6 can explain the observed differences in the behavior in our cellular experiments.
A, B Fit of the experimental scattering curve (green and pink dots) and theoretical scattering (green and pink lines) computed for the model of FXN:NB_28F6 and FXN:NB_16C10. C Plot showing the normalized pair-distance distribution function P(r) for the complexes reflecting the distance distribution. D The upper figure represents the superimposition on the ab initio-determined SAXS envelope (pale gray) for FXN:NB_28F6 complex with the AlphaFold3 predicted model. The bottom figure represents the superimposition on the ab initio-determined SAXS envelope (pale gray) for FXN:NB_16C10 with its crystal structure-based model.
Docking of the NB:FXN complex onto the supercomplex structure
Whether the selected NBs might block the interaction of FXN with the rest of the supercomplex depends on the specific structure of the NB:FXN complex and how it fits into the supercomplex structure throughout the catalytic cycle. To gain structural information on that hypothesis, we combined crystallographic NB:FXN complexes (this work) with the previous structural data for the supercomplex (NFS1/ACP-ISD11/ISCU/FXN)2 provided by cryo-electron microscopy (PDB ID: 6NZU5). The different NB:FXN structures (Fig. 8A) were docked onto the supercomplex by the superimposition of the FXN subunit. In the type I NB-FXN binding (NBs NB_4A7, NB_6B1, and NB_16C10, Fig. 8B), only slight clashes between the NB and the rest of the subunits of the supercomplex were predicted. The predicted contacts that the NB may establish through its β3-β4 connector with one of the NFS1 subunits (Fig. 8B) could lead to the inhibitory effects observed in vitro, promoting FXN dissociation or directly blocking FXN binding to the supercomplex. In the case of NB_16C10, residues that could be involved in the clashes are Pro41NB-Asn245NFS1, Gly42NB-Arg271NFS1, Lys43NB-Glu120NFS1, Arg45NB-Arg273NFS1, and His46NB-Arg273NFS1 (using the numbering shown in Supplementary Fig. 2A for the NB and the numbering used in PDB ID: 6NZU for NFS1). Whether the plasticity of the protein chains might adequately accommodate this stretch could depend on the amino acid sequence of the NB β3-β4 connector (Fig. 8B), thus determining the rigidity/flexibility behavior and, ultimately, the inhibitory power of these NBs that bind using the same site. Given that NB_4A7 and NB_16C10 have an extra Gly in that stretch, we expect more flexibility than in the case of NB_6B1, in which an Asn residue occupies the place of the Gly44. Nevertheless, the edition by reengineering β3-β4 connectors will be a key for obtaining better FXN binders. Since the supercomplex exhibited higher enzymatic activity with NB_4A7 compared to NB_6B1 or NB_16C10, we considered NB_4A7 as a viable candidate for evaluating the interaction with the supercomplex in vitro.
A The X-ray structure of complex FXN:NB_16C10 was superimposed to the cryo-EM structure of the supercomplex (PDB ID: 6NZU). Gray surfaces represent NFS1subunits and ISCU2. B Residues from NB_16C10 involved in a possible clash with NFS1 subunit are shown. C SEC-FPLC profile of the supercomplex (NFS1/ACP-ISD11/ISCU2/FXN)2 in the presence of NB_4A7 (blue) or in the absence of the NB_4A7 (light blue). The SEC profile corresponding to the subcomplex (NFS1/ACP-ISD11/ISCU2)2 in the presence of the NB_4A7 (dark green, NIAU/NB) or the absence (light green, NIAU). FXN alone or NB_4A7 alone were also loaded (black and orange, respectively). Buffer or buffer plus PLP profiles are shown as controls (dashed black and gray lines, respectively). D Interaction between biotin-FXN and NB_4A7 (orange dashed line), the subcomplex (NFS1/ACP-ISD11/ISCU2)2 (NIAU, orange dotted line), or biotin-FXN with NB_4A7 and subcomplex (NFS1/ACP-ISD11/ISCU2)2 (NIAU + NB_4A7, orange full line) followed by Interferometry. E Interaction between biotin-FXN and NB_28F6 (magenta dashed line), the subcomplex (NFS1/ACP-ISD11/ISCU2)2 (NIAU, magenta dotted line), or biotin-FXN with NB_28F6 and (NFS1/ACP-ISD11/ISCU2) 2 (NIAU + NB_28F6, magenta full line) by Interferometry. Buffer was 25 mM Tris-HCl, 150 mM NaCl, 1 mM DTT, 1 mM Cys, 10 μM PLP, pH 8.0. (NFS1/ACP-ISD) concentration was 1 μM, and ISCU2 was 3 μM.
Remarkably, when the supercomplex formed by NFS1, ACP-ISD11, ISCU2, and FXN subunits was loaded in SEC-FPLC system, in the presence of NB_4A7, the elution of the complete assembly shifted to a lower elution time (from 37.65 to 37.28 min, black arrow 2 to 3, absence or presence of NB_4A7) (Fig. 8C). By contrast, when FXN was absent, the presence of NB_4A7 did not modify the elution time of the subcomplex (NFS1/ACP-ISD11/ISCU) (Fig. 8C, elution time 38.4 min, black arrow 1 dark and light green, with or without NB_4A7), a fact indicative of the NB specificity for FXN. More importantly, these results suggest that NB_4A7 can bind FXN in the context of the supercomplex. Additionally, it was evident that a fraction of the NB bound to the free FXN shifted its elution time to lower values, indicating the stabilization of the complex NB:FXN (elution time shifted from 44.4 to 43.3 min in the absence or presence of NB_4A7, respectively). It is worthy of note that under these experimental conditions, the interaction of FXN with NFS1 is not so strong; in fact, other research groups reported a dissociation constant for wild-type FXN of KD ~ 3 ± 1 μM5. Thus, we expected a high fraction of unbound FXN (∼55% unbound, ∼45% bound53). Moreover, considering the dilution due to the SEC column volume, the unbound FXN fraction may be higher (a half dilution would result in ∼72% unbound and ∼28% bound).
We also conducted an interferometry experiment to investigate whether the supercomplex and NB_4A7 can bind simultaneously to FXN (Fig. 8D). In this experiment, streptavidin sensors were sensitized using the biotin-labeled FXN H177C variant, (Supplementary Fig. 5) and, after that, sensors were incubated with solution containing both NB and the subcomplex (NFS1/ACP-ISD11/ISCU2)2 in the presence of a 3:1 excess of ISCU2: (NFS1/ACP-ISD11/ISCU2)2, L-Cys and PLP to increase the affinity between ISCU2 and the core complex (NFS1/ACP-ISD11)214. Our results showed that the signal corresponding to the complete mix of proteins is even higher than the one observed for the NB_4A7 alone, suggesting that the interaction of FXN with the NB_4A7 does not impede supercomplex formation. Additionally, the KD measured for FXN: (NFS1/ACP-ISD11/ISCU2)2 was ∼0.6 μM, a value in the range of the dissociation constant previously measured under similar conditions (0.2 μM)14.
On the other hand, when a similar experiment was carried out using NB_28F6 (Fig. 8E), which binds to the helix 1, the signal observed for the subcomplex (NFS1/ACP-ISD11/ISCU2)2 and NIAU + NB_28F6 was similar to that observed for NB_28F6, suggesting that this NB inhibits the binding of the subcomplex to FXN, in agreement with structural results (SAXS and NMR).
Expression of NBs in the mitochondria of human cell lines
To evaluate the effects of NB:FXN interaction on the cellular metabolism, we first studied whether the NBs could be successfully imported to the mitochondrial matrix when expressed in human cell lines. We used transfected HeLa Kyoto cells to evaluate this by immunofluorescence. We prepared vectors that included the NB sequence in frame with the citrate synthase mitochondrial transit sequence (MTS) for mitochondrial matrix localization. The cells were co-transfected with the Discosoma red fluorescent protein (dsRed Mito), which also included an MTS (cytochrome c oxidase subunit VIII). The transfected cells were analyzed after 48 h post-transfection.
As expected, when NBs were detected using an anti-VHH polyclonal serum, which recognizes the NBs invariable regions (followed by an anti-rabbit secondary antibody labeled with Alexa 488), the NB signal co-localized with that obtained with dsRed-Mito, strongly suggesting a mitochondrial localization of the NBs. This behavior was observed for the four nanobodies studied in this work (Supplementary Fig. 14A).
Importantly, the expression of NBs NB_4A7, NB_6B1 and NB_16C10 did not alter the viability of the cells (Supplementary Fig. 14B). The absence of significant toxicity after NB transfection and the lack of increased cytosolic immunofluorescence related to NBs suggest that NBs did not remain in the cytosol and do not prevent FXN from reaching the mitochondria. However, more experiments will be done, especially with unstable FXN variants or those that may remain longer in the cytosol. Therefore, we demonstrated that the NBs are localized in the mitochondria and that their expression did not significantly affect the cell viability. In turn, although the expression of NB_28F6 was very low, it was still possible to evaluate its subcellular localization, and the analysis suggests that it is localized in the mitochondria. On the other hand, the viability analysis of cells transfected with NB_28F6 should be examined cautiously due to the lower expression levels and the possibility that the effects may not be comparable to those of the other NBs.
NB expression and interaction between FXN and NB in HeLa Kyoto and HEK-293T cells. Characterization of Fe-S clusters related to mitochondrial bioenergetics
The interaction between FXN and NB in the cellular environment was investigated (Fig. 9). The results clearly showed that co-expressing NB_4A7 (using the citrate synthase MTS) and FXN (in its 210-residue precursor form) led to the colocalization of both proteins in a similar subcellular compartment (Fig. 9H), supporting previous findings that indicate mitochondrial localization of the NB (Supplementary Fig. 14A).
A–H Immunofluorescence to assess NB_4A7 and FXN in the cellular environment using HeLa Kyoto cells. (A–D) Transfection with an empty plasmid; neither FXN nor NB_4A7 was transiently overexpressed (FXN is endogenous). E–H Both FXN precursor and NB_4A7 were transiently overexpressed from pCDNA3.1 vectors. Detection of FXN and NB_4A7 was performed using anti-FXN (host species: mouse, ABCAM ab110328) and anti-HA MAB (host species: rabbit, Cell Signaling HA-Tag (C29F4)). Labeled secondary antibodies detected FXN at 488 nm (secondary Antibody, Alexa Fluor™ 488) and NB at 543 nm (secondary Antibody, Alexa Fluor™ 594). The merged fluorescence signals are shown (yellow, D and H). Nuclei were counterstained with Hoechst (blue). I, J Proximity Ligation Assay (PLA) in HeLa Kyoto cells. Dots indicate the proximity of FXN to NB_4A7. Cells were fixed, washed, permeabilized, and incubated with primary antibodies, anti-FXN (mouse) and anti-HA (rabbit). PLA reactions were carried out, and cells were mounted using in situ mounting medium with DAPI. Two independent experiments were performed, and the presence or absence of red dots was analyzed in at least five different fields. Fluorescence microscopy images were then acquired. In (I), an empty vector was used, while (J), NB_4A7, and FXN precursors were transiently overexpressed using pCDNA3.1 vectors. Immunofluorescence was performed 48 h after transfection. K Co-immunoprecipitation of NB:FXN. HEK-293T cells were transfected with either an empty vector or a vector encoding the NB_16C10 sequence for protein expression. IP: His indicates the lanes where co-immunoprecipitation was performed using an anti-His tag antibody. FXN was detected with an anti-FXN monoclonal antibody, while NB was detected using the anti-His antibody. GAPDH was used to assess the protein content in the input of the co-IP experiments. The symbols + and – indicate the presence or absence of the respective vectors for cell transfection (empty or encoding NB_16C10). The scale bar is the same for all panels and corresponds, in all cases, to 30.0 μm.
Next, we employed the in situ proximity ligation assay (PLA, Fig. 9I, J and Supplementary Fig. 15) to determine whether the interaction between FXN and NB occurs within a cellular environment, particularly in HeLa Kyoto cells. Both FXN precursor and NB_4A7 were overexpressed. Most importantly, the PLA-positive signal (Fig. 9J) demonstrated that FXN and NB are within 40 nm of each other, indicating direct contact.
The interaction between NBs and FXN in a cellular environment was further studied through co-immunoprecipitation. Given that N_16C10 has a middle inhibitory effect on desulfurase activity compared to NB_4A7 and NB_6B1 and given that it exhibits the lowest KD for FXN among the solved complexes (Supplementary Table 2), we evaluated this NB as a strong candidate for examining possible interactions in a more complex environment. The NB candidate, NB_16C10, was expressed in HEK-293T cells, and its interaction with endogenous FXN was examined. Notably, endogenous FXN was co-immunoprecipitated with NB_16C10 using an anti-His-tag antibody, indicating that a stable NB:FXN complex formed in the cells (Fig. 9).
Additionally, samples corresponding to the co-immunoprecipitation were investigated by mass spectrometry. The analysis of the peptides corresponding to the recovered proteins indicated a strong interaction between the NB_16C10 and FXN (Supplementary Table 7). The identification of peptides corresponding to citrate synthase suggests that the NB precursor containing the MTS of this mitochondrial enzyme was immunoprecipitated. However, since the immunoprecipitation was performed using an anti-His antibody, this does not necessarily indicate an interaction between the NB precursor and FXN in the cellular environment. Future research should focus on whether the NB and FXN can interact in the cytosol or, conversely, whether the precursor of the NB might interact with FXN in the mitochondrial matrix.
Remarkably, no proteins other than FXN, NB, and those recovered when cell samples were transfected with the empty vector were consistently retrieved, suggesting that the method used for cellular lysis may perturb the stability of other complexes involving protein-protein interactions of lower affinities than that of NB:FXN (KD for FXN in the nanomolar range, Supplementary Table 2).
The effect of NB expression on FXN expression was studied. The densitometric analysis of Western blot membranes from three independent experiments showed similar levels of FXN among the cells transfected with an empty vector (Supplementary Fig. 16). Besides this, the expression of NB_4A7, NB_6B1, and NB_16C10 in HEK-293T cells was substantial (Supplementary Fig. 16B). However, the expression of NB_28F6 was significantly lower (Supplementary Fig. 16B) than the others (~90% lower, as evidenced by Western blotting).
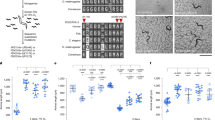
Transfected HEK-293T cells were evaluated to identify possible adverse effects of the NB expression on Fe-S cluster-dependent enzymatic activities from ACO and SDH. The expression of NBs showed only slight modulations of ACO and SDH activities (Fig. 10A, B). Only the expression of the NB_16C10 led to a slight decrease of the SDH enzyme in HEK-293T cells (Fig. 10B). More experiments will be conducted to evaluate whether this is a direct effect on the enzyme through the alteration of Fe-S cluster assembly activity, or if, alternatively, the modulation of SDH is a consequence of more complex processes involving transcription, translation, or even post-translational modifications.
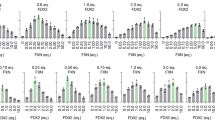
A Aconitase and B succinate dehydrogenase activities (ACO and SDH, respectively) were quantified 48 h after transfection. OCR profiles are shown for cells transfect with C NB_4A7, D NB_6B1, E NB_16C10, and F NB_28F6. G The OCR parameters. After basal O2 consumption recording, oligomycin (an ATP synthase inhibitor) is added, and the coupling to the ATP synthesis is measured. Then, the addition of FCCP (a protonophore that uncouples oxygen consumption from ATP synthesis) allows us to measure the maximum respiratory capacity to use OXPHOS. This also enables the measurement of the spare respiratory capacity (SRC) as the difference between maximal and basal OCR. Finally, rotenone and antimycin A (Complex I and Complex III inhibitors, respectively) are added to block the electron transport chain. For panels A and B, three independent experiments were analyzed (n = 3), each with two technical replicates. For C–F, two independent experiments were analyzed (n = 2), including three technical replicates for each. Statistical analysis was performed using one-way ANOVA, followed by Dunnett´s test. Differences were considered significant when P ≤ 0.05. Error bars represent the standard errors.
To gain a more global picture of the effect of the FXN_ NB expression in these cells, we measured the oxygen consumption rate (OCR) 48 h after transfection (Fig. 10C–F), which is indicative of oxidative phosphorylation (OXPHOS) efficiency. Our results suggested that the OCR is not altered when NB_6B1 or 16C10 are expressed. These NBs exhibited similar extents in the increase of the OCR after p-trifluoromethoxyphenylhydrazone (FCCP) treatment, revealing a similar maximal respiratory capacity (Fig. 10G), as non-transfected cells. On the other hand, NB_4A7 exhibited a slight decrease in the basal respiratory capacity, possibly compatible with the slight decrease of SDH (Complex II, not significant) activity. However, the observed metabolic changes seem subtle, given the considerable mitochondrial protein expression of NB_4A7, 6B1, and 16C10.
In turn, when the HEK-293T cells were transfected with a vector encoding NB_28F6, although the expression of this NB was significantly lower than that of the other NBs (Supplementary Fig. 16B), a decrease in basal and maximal respiration was observed (Fig. 10F), indicating a reduction of mitochondrial metabolism.
Discussion
In this work, we characterized new molecular tools to explore the interaction of FXN with foreign binder partners. With this aim, twenty NBs specific to FXN were selected by phage display. We found that NBs can increase FXN conformational stability in vitro. It is therefore plausible to hypothesize that NB interaction may potentially rescue unstable FXN variants linked to FRDA. However, this hypothesis remains to be tested, as our study did not assess the effect of NB interaction on the conformational stability of FRDA-related FXN variants within a cellular context. Binding affinities were in the nanomolar range of 1–33 nM, suggesting a strong interaction, a key feature for complex formation in the cellular environment. SEC, NMR, and BLI results also suggested a slow dissociation equilibrium. In addition, in vitro L-Cys-desulfurase activity could be modulated by the NBs; while 1:1 molar ratio does not affect L-Cys desulfurase activity for some NBs (NB_4A7, 5A8, 29F7, and 83G4, Fig. 2 and Supplementary Fig. 3), an increase to 1:5 ratio was inhibitory. As at 1:1 ratio, ∼83% of the FXN is in complex with NB_4A7 (KD = 33.0 nM), whereas ∼17% is free (calculated using the ligand binding simulation tool53), the inhibition observed at 5:1 indicates a more complex interaction of the NB with the rest of the proteins of the system, suggesting unspecific binding under these conditions. The relevance of this fact will be further studied. On the other hand, binding of NB_4A7 to FXN did not inhibit the interaction of FXN with the supercomplex (Fig. 8), in agreement with the preservation of ∼85% and ∼70% of Cys desulfurase activity observed in vitro for 1:1 and 1:5 ratios, respectively (Fig. 2).
Different experimental techniques (NMR, X-ray crystallography, and SAXS) enabled us to characterize the interaction between the FXN and the NBs. Two distinct types of binding, which we have designated as Type I binding (as we find for NBs NB_16C10, NB_6B1, and NB_4A7), and Type II binding (as found for NB_28F6) were found (Figs. 4, 6, and 7). The complexes with a Type I binding have two key polar interactions with the FXN residues of the loop L1 of FXN; specifically, residues Glu121, Pro117, and Thr119, involving a molecular contact surface area of 581.7 Å2. Residue Glu121 plays a crucial role in the protein interaction of Type I by forming a salt bridge interaction with residue Arg38 and engaging in polar interactions with Ser62 of the NBs. Pro117 and Thr119 also play an important role by establishing H-bonds with residues His46 and Leu47 of β5 from the NBs. Even though Asn52 from CDR2 forms an H-bond with backbone Gly138 from FXN, and Val98, Pro99, and Pro100 from CDR3 are at Van der Waals distance of FXN, in Type I interaction mode, the recognition is not made using the three segments CDR1, CDR2, and CDR3. However, this is common for NBs that use a far vaster diversity of structural stretch combinations to bind antigens54.
To study the effect of NBs in vivo, human cell lines were transfected with vectors encoding four different NBs. Three presented a considerable expression level; instead, NB_28F6 exhibited significantly lower expression. The four transfected NBs were found in the mitochondria of HeLa Kyoto cells. Remarkably, the expression of the NBs in HEK-293T and HeLa Kyoto cells did not alter FXN expression or cell viability. However, it will be important to assess the impact of NBs on the viability of other cell types and patient-derived cells.
Considering the degree of modulation of L-Cys desulfurase in vitro, the higher expression level of NBs in the cells, and the plausible broader spectrum of functions that FXN might exert inside the cells, including direct interactions with iron, aconitase55,56, superoxide dismutase57 or Complex I from the respiratory chain58, we reasoned that the expression of the NBs could affect the Fe-S cluster-dependent enzymatic activities and energetics. However, the expression of NB_6B1 and NB_4A7 did not have effects on Fe–S–dependent ACO and SDH activities, and neither did the expression of NB_16C10 have effects on ACO, although it did exhibit a slight inhibition on SDH (Fig. 10). This last fact could be compatible with the absence of in-cell inhibition of mitochondrial L-Cys desulfurase NFS1 supercomplex in HEK-293T cells overexpressing NBs. Moreover, in a more global picture of the mitochondrial metabolism, we demonstrated that the expression of NB_4A7, NB_6B1 and NB_16C10 in HEK-293T did not alter mitochondrial respiration 48 h after transfection, suggesting that the mitochondrial global metabolism is not significantly perturbed when these NBs are expressed, imported into the mitochondria and eventually interacting with FXN (as judged by co-immunoprecipitation and in situ proximity ligation assay). However, the effect of nanobody expression on a wide range of Fe-S cluster-dependent enzymes should be assessed to understand its impact on cellular FXN function. Nanobody expression might alter specific enzymatic activities, which do not necessarily affect mitochondrial energetics.
Remarkably, NB_28F6 yielded a similar profile of ACO and SDH activities. However, under the same conditions, NB_28F6 showed very poor expression levels (∼5 times lower on average, Supplementary Fig. 16). Thus, we could not evaluate its effect on metabolism with confidence. The observed decrease of the OCR (even with a considerably lower protein expression level than the other NBs) suggests that NB_28F6 expression affects mitochondrial OCR through FXN intervention. In particular, the binding surface of FXN involved in the interaction with NB_28F6 could explain the inhibitory effects observed in vitro and in vivo (Figs. 2 and 10). As evidenced by our NMR results, it involves a larger portion of helix α1 in FXN, the acidic ridge of FXN and NB_28F6 binding resulted in the inhibition of the FXN-supercomplex interaction (Fig. 8). Unfortunately, the crystal structure of the FXN: NB_28F6 complex could not be obtained. We believe that some heterogeneity in the sample, arising from partial proteolysis of this NB (as indicated by mass spectrometry results), might impede the crystallization process. These dissimilar results between the in vitro and in-cell effects can be understood considering that the in vitro measured activity involves only a partial reaction, the first steps of Cys desulfurase reaction, employing DTT as a non-physiological reducing agent. In contrast, in the cell, the formation of the Fe-S cluster necessary to sustain Fe-S dependent enzymes involves several steps: the transfer of persulfide to ISCU2, the delivery of electrons (reduction) by FDX2, the assembly of the iron-sulfur cluster on ISCU2, and the transfer of the cluster to target subunits and enzymes.
The dynamic behavior of the L-Cys NFS1 desulfurase supercomplex was further characterized last year by including the electron donor for cluster assembly Ferredoxin 2 (FDX2)21. Recently, it was determined that this protein has a multifaceted binding mode (two steps involving the C-terminal partially folded stretch of FDX2) to the supercomplex59. Moreover, it was demonstrated that FDX2 binds to the same supercomplex region as FXN. This indicates that precise exchange dynamics are essential for accurate function. Changes in FXN or FDX2 affinity by the supercomplex could significantly impact the kinetics of the catalytic cycle. The interaction of FXN with the NB, making subtle clashes of the NB and NFS1 structures, might have unexpected effects on function, suggesting that the reaction studied in vitro, the L-Cys desulfurization in the presence of DTT, may show a different profile compared to the complete reaction occurring in the cell, involving the Fe-S cluster formation, which depends on FDX2/FXN dynamics.
Regarding the interaction of NBs with FXN variants, NB_4A7 was able to interact with and stabilize some of the FRDA variants assayed (G130V, W155R, L198R), though not all. Specifically, variants D122Y and G137V, which carry mutations located at the interaction surface, were not significantly stabilized in vitro by this NB.
Notably, G130V, like many other FRDA-related FXN variants, exhibits less Cys desulfurase activation capability compared to the wild-type FXN. Specifically, this variant exhibits an in vitro activation of 50–60% compared to the control60. Additionally, the FXN G130V concentration in vivo is low51,61; it may be degraded, presumably due to its low conformational stability50. Furthermore, altered mitochondrial import51 and MTS processing62 were described for this variant.
The analysis of in vitro experiments suggested that NB_4A7 does not increase the activation capability of G130V (Supplementary Fig. 3B). Considering also the observation that NBs can endow G130V with conformational stability, these results indicate that G130V may have a locally altered conformation in both states, free or in complex with the NB; thus, when a Val residue occupies position 130, the side chain is at van der Waals distance from residues Ser385 and Glu387, near of Cys381, which is involved in the transfer of persulfide from the PLP catalytic center to the ISCU2 assembly site. Thus, Val130 could directly alter the activation capability.
Currently, our laboratory is investigating the potential of producing NBs in a Trojan version for cell penetration and delivery63. These NBs can be easily produced using a bacterial expression system, making their production cost-effective and efficient. It is worth mentioning that the premature folding of the Trojan NB precursor may cause the accumulation of NB in the cytosol or the stalling of NB at the protein import apparatus, leading to proteotoxic stress64. Thus, NB proteostasis should be studied to prevent such stress and other effects of mitochondrial NB import. Additionally, although the immunogenicity of NBs is reduced65, in vivo experiments will require a deeper understanding of the interaction between these specific NBs and the immune system.
In this work, we successfully localized NBs in the mitochondrial matrix, which exhibit a high affinity for FXN and stabilize a series of FRDA-related variants in vitro. Moreover, the analysis of the in situ Proximity Ligation Assay results demonstrated the direct interaction between NB and FXN in the cellular environment. These findings open the door to a general strategy based on intervening in mitochondrial biochemistry using specific nanobodies. These new tools may help in adding specific players to the macromolecular environment closest to the supercomplex.
Materials and methods
DNA sequencing, protein concentration, and mass spectrometry analysis
Identity was verified by DNA sequencing using the Macrogen facility. Absorption spectra were obtained with a JASCO V-730 BIO spectrophotometer (Tokyo, Japan). Protein concentration was determined using the extinction coefficient obtained from the amino acid sequence using the ProtParam tool from ExPASy. Protein purity was estimated by SDS-PAGE analysis, and the analysis of the intact masses was performed by mass spectrometry at the National Laboratory of Research and Services in Peptides and Proteins (FCEN-UBA, Argentina) using an LCQ DUO ESI ion trap (Thermo Finnigan) or QExactive Orbitrap (Thermo Scientific) spectrometers.
HPLC analysis of protein samples
Reverse phase HPLC analysis was performed using a JASCO system equipped with an autoinjector, UV detector, and a thermostatic oven at 25 °C. Gradients from 0 to 100% acetonitrile were made, and 0.05% TFA (v/v) were added to the solvents and samples. The columns used were analytical C18 (Higgins Analytical, Inc., U.S.A.), and the flow was 1.0 mL min−1. Proteins were monitored at 220 nm. Before injection, all samples were centrifuged.
Expression and purification of the human supercomplex core (NFS1/ACP-ISD11)2
The DNA sequences of the human mature form of the human cysteine desulfurase NFS1 enzyme (NFS1Δ55), the ISD11, and the human mitochondrial ACP (the mature form) were optimized for protein expression in E. coli BL21 (DE3) by BIO BASIC Inc (Markham ON, Canada). NFS1Δ55 and ISD11 were cloned in a pETDuet-1 plasmid, whereas ACP was cloned in a pACYCDuet-1 for co-expression. The NFS1 amino acid sequence included the RGSHHHHHH tag in the N-terminal for purification and antibody recognition. Protein expression was induced by 1 mM IPTG when the bacterial culture reached OD600nm = 1.0. Co-expression was carried out overnight at 20 °C (at 250 rpm). The purification of the complex (NFS1/ACP-ISD11)2 was performed from the soluble fraction of E. coli BL21 DE3 cultures using a Ni2+-NTA-agarose column (first step). The protein was eluted with 20 mM Tris-HCl, 300 mM NaCl, 500 mM imidazole, pH 8.0. After that, 2 mM DTT was added and the protein was dialyzed overnight against the same buffer, without imidazole, at 4 °C. Afterwards, it was frozen at −70 °C. This protocol minimized aggregation and oxidation of the protein. Before each experiment, (NFS1/ACP-ISD11)2 was thawed in ice and centrifuged. Protein concentration was determined spectroscopically using an absorption coefficient ε280nm = 53750 M–1 cm–1.
Expression and purification of ISCU2
The ISCU2 DNA sequence corresponding to the mature form was optimized for E. coli overexpression by Explora Biotech (Rome, Italy) and subcloned in a pE22b plasmid, with a C-terminal His6 tag. Protein induction was carried out by adding 1 mM IPTG final concentration (3 h, 37 °C, and 250 rpm). ISCU2 was purified using a Ni2+-NTA-agarose column equilibrated with 20 mM Tris-HCl, 300 mM NaCl, pH 7.5. The elution was performed with 20 mM Tris-HCl, 300 mM NaCl, 500 mM imidazole, pH 7.5. The collected elution fractions were analyzed by SDS-PAGE and pooled. After that, an extensive dialysis step was performed (4 °C, in a 20 mM Tris-HCl, 300 mM NaCl, pH 7.5, 1 mM DTT). Protein purity was >95%, as evaluated in SDS-PAGE. No aggregation was observed, as inferred from size exclusion chromatography (SEC) analysis. The protein was stored at –70 °C until use. Protein concentration was determined spectroscopically using an absorption coefficient ε280nm = 11,460 M–1 cm–1 (1 mg/mL protein solution represents Abs280nm = 0.70). The zinc content in ISCU samples was evaluated by atomic absorption spectroscopy at the Departamento de Bromatología, Facultad de Farmacia y Bioquímica, Universidad de Buenos Aires. Zinc concentration in ISCU2 preparations was 0.14: 1 (zinc: ISCU2, molar ratio).
Expression and purification of FXN
Human FXN (residues 90–210) was overexpressed and purified as previously described for the wild-type protein36. Briefly, bacterial cultures (E. coli BL21 (DE3), 2–3 L Terrific Broth, pH 7.2) were grown at 37 °C and 280 rpm. Protein expression was induced at DO = 0.8–1.0 with 1.0 mM IPTG. After induction (3.5 h), bacteria were centrifuged (6000 rpm), and the pellet was stored at –20 °C until cell disruption by sonication (in an ice-water bath). The soluble fraction was separated by centrifugation (10,000 rpm, 30 min). The soluble fraction was incubated with 10 mM EDTA and loaded onto an ion exchange chromatography (DEAE DE52 matrix). The protein was eluted with a 300 mL linear gradient from 0.0 to 1.0 M NaCl (the buffer was 20 mM Tris-HCl, 1 mM EDTA, pH 7.0). Fractions with FXN (identified by SDS-PAGE) were loaded onto a Sephadex G–100 column (SEC, 93 cm × 62.7 cm, equilibrated with 20 mM Tris-HCl, 100 mM NaCl, 1.0 mM EDTA, pH 7.0). FXN concentration was determined spectroscopically using an absorption coefficient ε 280nm = 26,930 M–1 cm–1 (1 mg/mL protein solution represents Abs280nm = 2.00). Purity was >98% as evaluated in SDS-PAGE.
NFS1 cysteine desulfurase activity
For enzymatic desulfurization of L-Cys to L-Ala and sulfide by the (NFS1/ACP-ISD11/ISCU/FXN)2 supercomplex, concentrations of proteins, substrate and the reducing agent DTT were set according to a previous paper by Tsai and Barondeau66. Reactions contained 1.0 μM NFS1/ACP-ISD11, 3.0 μM ISCU2, and 1.0 μM FXN, and samples were supplemented with 10 μM PLP, 2.0 mM DTT and 1.0 μM FeSO4 (final concentrations). In all cases, the reaction buffer was 50 mM Tris-HCl and 200 mM NaCl, pH 8.0, and reactions were started by the addition of 1.0 mM L-Cys (or variable L-Cys concentrations, as we described below). Samples were incubated at room temperature (25 °C) for 30 min. For the reactions including NBs, each NB was preincubated with FXN (10 min), and then both proteins were added to the reaction mix.
The methylene blue method
Sulfide was determined by the methylene blue method66,67. For this analysis, H2S production was stopped by adding 50 μL of 20 mM N,N-dimethyl p-phenylenediamine in 7.2 M HCl and 50 μL of 30 mM FeCl3 (prepared in 1.2 M HCl). Under these conditions, the production of methylene blue took 20 min. After that, samples were centrifuged for 5 min at 12,000 x g and the supernatant was separated. Absorbance at 670 nm was measured.
Llama immunization and library construction
A llama located at INTA’s Camelids Experimental Unit was immunized intramuscularly with 150 μg per dose of human FXN 90-210 recombinant protein on days 0, 14, 28, and 50. Complete Freund’s adjuvant was used for the first dose, and incomplete Freund’s adjuvant for the following boosts. Antibody responses were monitored by ELISA on serum samples taken before each immunization. Llama inoculation and sample collection were conducted by trained staff, and the study was approved by the Animal Care and Use Committee of INTA (CICUAE) under protocol No. FR6.2-3/2020. Four days after the last boost, 150 mL of anticoagulated blood was used to isolate lymphocytes by Ficoll Paque Plus (GE Life Sciences, 17-1440-02) filled Leucosep tubes (GBO, 227290), total RNA was extracted (RNAeasy Midi, Qiagen 75144) and cDNA was prepared with oligo(dT) primers (First Strand cDNA Synthesis Kit, Roche 04379012001). VH and VHH genes were amplified with CALL001 (5’-GTCCTGGCTGCTCTTCTACAAGG-3’) and CALL002 (5’-GGTACGTGCTGTTGAACTGTTCC-3’) primers. A PCR amplicon of 0.7 kb was purified from gel (Wizard® SV Gel and PCR Clean-Up System, Promega A9282) and then used as a template in a nested PCR to specifically amplify the VHH fragments using VHH-BACK-SAPI (5’-CTTGGCTCTTCTGTGCAGCTGCAGGAGTCTGGRGGAGG-3’) and VHH-FORWARD-SAPI (5’-TGATGCTCTTCCGCTGAGGAGACGGTGACCTGGGT-3’) primers. A Golden Gate assembly was performed to clone the VHH sequences between the two SapI sites of the phagemid vector pMECS-GG, following a protocol previously described68. Electro-competent E. coli TG1 cells (Lucigen 60502-1) were transformed with the purified ligation mixture and plated on a selective agar medium. A library of 1.8 × 109 individual transformants was obtained69.
Selection of FXN specific NBs
To produce recombinant specific NBs, 1 mL of bacteria from the stock library was grown in 2 × TY until OD600nm = 0.6; afterward, VCS M13 helper phage (Stratagene, 200251) was used to infect exponentially growing bacteria. The resulting NB phage display library was panned three times on microtiter plates (Maxisorp Nunc) coated overnight at 4 °C with 10 μg of each recombinant protein in 100 μL of PBS, and 100 μL of PBS for the negative controls. The next day, wells were washed with PBST (PBS + 0.05% Tween 20) and blocked with 2% skim milk in PBST. Approximately 1 × 1012 phage particles were preincubated with 10 μL of blocking solution in 100 μL of PBS for 30 min at room temperature by head-over-head rotation, then added on positive and negative wells and incubated for 2 h on a vibrating platform (500 rpm.). During the first panning round, wells were washed 10 times with PBST, while wells were washed 20 and 25 times for the second and third rounds, respectively; 5 min incubation on a vibrating platform (300 rpm) was performed every five washes. Specific phage particles were eluted with 0.25 mg mL−1 trypsin solution (Sigma-Aldrich, T1426) for 30 min followed by neutralization with 4 mg mL−1 AEBSF solution (Carl Roth, 2931.3). A second elution step was performed by adding exponentially growing E. coli TG1 cells to positive and negative wells, which were then incubated for 30 min at 37 °C. Trypsin-eluted phage particles were amplified by infection of exponentially growing E. coli TG1 cells and later superinfected with VCS M13 helper phage. Phage particles obtained after both elution strategies were purified using PEG 6000/NaCl precipitation and used for the next selection round. To obtain specific binders, individual TG1 colonies were screened by ELISA using periplasmic extract. For this, 95 colonies from the positive wells (different panning rounds and elution strategies) and 1 colony from the negative well were inoculated in 1 mL of 2 × TY medium containing 100 μg mL−1 ampicillin and 0.1% glucose in a deep well plate. NB expression was induced after bacteria incubation for 3 h at 37 °C and 200 rpm. with 1 mM IPTG. After a 4 h induction, bacterial cultures were centrifuged, the pellets were frozen and thawed twice to disrupt cells and resuspended in 120 μL of PBS. On the other hand, the periplasmic extract (see below, NB purification) was used to determine the specificity of NB binding and to study their functional activity by the methylene blue method.
Screening for antigen binders
For screening experiments, the study of NB binding to FXN was conducted by ELISA. For this, microtiter plates (Maxisorp, Nunc) were coated overnight at 4 °C with 200 ng/well of recombinant FXN or an irrelevant protein as negative controls, diluted in PBS. After three washes with PBST, wells were blocked with 3% skim milk in PBST, and 50 μL of the periplasmic extract was added to each well and incubated at room temperature for 2 h. After washing with PBST to remove the excess of NB, specific binding was detected with horseradish peroxidase (HRP)-linked anti-HA antibody (Abcam, ab1190) diluted 1:1500. Finally, 50 μL of TMB substrate (3,3’, 5,5’ tetramethylbenzidine, BD 555214) was added. Absorbance at 450 nm was measured using an ELISA reader (TECAN).
Nanobody production and purification
A DNA sequence corresponding to the NB, preceded by a signal peptide for export to the periplasm (MKYLLPTAAAGLLLLAAQPAMA), C-terminal hemagglutinin (YPYDVPDY), and His (HHHHHH) tags, was cloned in an expression vector (pMECS), and E. coli WK6 was transfected with these constructs. Different expression protocols were carried out (temperature was modified from 20 to 37 °C, and expression was performed for 4 h or overnight). Some differences concerning yield and purity were observed for specific NBs. However, we chose to follow the same protocol for all the NBs. Protein induction was carried out by adding 1.0 mM IPTG final concentration (4 h, 37 °C, and 190 rpm). Periplasm fluid was recovered by osmotic shock. The pellet from 2 L of WK6 cell culture was resuspended in 30 mL of TES buffer (100 mM Tris-HCl pH 8.0, 1 mM EDTA, and 20% sucrose) pre-chilled in an ice-water bath. The resuspended cells were incubated for 60 min. After that, 90 mL of pre-chilled deionized water was added, and cells were incubated overnight. Both incubations were carried out on an ice-water bath in a rocking shaker (3D motion). The suspension was centrifuged, and the supernatant was loaded onto a Ni2+-NTA-agarose column equilibrated with 20 mM Tris-HCl, 300 mM NaCl, pH 7.5. The elution of the NBs was performed with 20 mM Tris-HCl, 300 mM NaCl, 500 mM imidazole, and pH 7.5. The collected fractions were analyzed by SDS-PAGE and pooled. After that, an extensive dialysis step was performed (4 °C, in a 20 mM Tris-HCl, 300 mM NaCl, pH 7.5). Protein purity was >95%, as evaluated in SDS-PAGE. No aggregation was observed, as inferred from SEC analysis. Pure NB was stored at −70 °C until use. When osmotic shock was performed, protein preparations of higher purity were obtained compared with lysis by sonication.
Hydrodynamic behavior of the NB:FXN protein complexes
SEC-FPLC was performed using a Superose-6 column (GE Healthcare). Protein concentration was 6–10 μM, a volume of 100 μL was typically injected, and the running buffer was 20 mM Tris-HCl, 100 mM NaCl, at pH 7.4. The experiment was conducted at room temperature (∼25 °C) at a 0.5 mL/min flow rate. A JASCO HPLC instrument was used. It had an automatic injector, a quaternary pump, and a UV–Vis UV-2075 (elution was monitored at 280 nm).
Thermal shift assay
Temperature-induced denaturation of NB:FXN complexes using the FRDA-related G130V FXN variant was monitored by the change in the Sypro Orange dye fluorescence using protein at a 5.0 μM concentration in 50 mM sodium phosphate buffer, pH 7.4. Samples of FXN G130V alone, wild-type FXN alone or NB alone, and samples without any protein were also included as controls. The dye was used at 2× (as suggested by Thermo Fisher Scientific). The temperature slope was 1 °C min−1 (from 20 to 90 °C). Excitation and emission ranges were 470–500 and 540–700 nm, respectively. The fluorescence signal was quenched in the aqueous environment but became unquenched when the probe was bound to the apolar residues upon unfolding. Experiments by triplicate were carried out in a Step One Real-Time-PCR instrument (Applied Biosystems, CA, U.S.A.).
NB titration by NMR and the NB interaction sites on the FXN surface
15N labeled FXN was prepared as before60. Samples for NMR experiments contained 0.1 mM 15N-labeled protein in a buffer supplemented with 5% D2O. NMR experiments were performed at 22 °C in a Bruker 600 MHz Avance III spectrometer equipped with a TXI probe. The NMR data were processed with NMRPipe70 and analyzed using NMRViewJ71. 1H–15N HSQC experiments were performed to follow the interaction by CSPs. The 15N-labeled FXN was titrated with each NB at molar ratios of 0:1, 0:33, 0.66, and 1:1 (NB:15N-FXN). The experiment was performed in a 25 mM Tris-HCl, 150 mM NaCl, pH 7.4 buffer. CSP values of amide peaks were calculated using the equation \({\Delta \delta }_{{{\mathrm{aV}}}}={\left[\left({\delta }_{{{\mathrm{HN}}}}^{2}+\,{\delta }_{{{{\rm{N}}}}}^{2}/25\right)/2\right]}^{1/2}\), where ΔδAV, ΔδHN, and Δδ15N represent the average chemical shift value, proton chemical shift, and nitrogen chemical shift changes, respectively72.
Biolayer interferometry experiments
The experiments were carried out using a BLItz instrument (Sartorius). For this experiment, a recombinant FXN H177C variant (>98% pure) was labeled with a bifunctional biotin-maleimide probe of long arm Cat# SP-1501-12 (Vector). Variant FXN H177C-biotin was purified by G25 (separated from the free probe), and extensive dialysis was performed and stored at −70 °C. The sensors were hydrated in TBS buffer (25 mM Tris-HCl, 150 mM NaCl, pH 7.4) for these experiments and supplemented with 0.5 mg mL−1 BSA (TBS-BSA) for 10 min. Subsequently, FXN-biotin binding to the streptavidin sensor surface was carried out. Each experiment consisted of a baseline (100 s, 250 μL), an association (300 s, 4 μL), and a dissociation (300 s, 250 μL). After each experiment, the sensor was regenerated by glycine 10 mM pH 1.0 (5 s, 300 μL) and TBS-BSA buffer washing.
Cell culture, treatments, and materials
HEK-293T cells (kindly provided by Dr. Ibanez, INQUIMAE, UBA) and Hela Kyoto cells (kindly provided by Dr. Matias Blaustein, iB3, UBA) were grown in high glucose (4.5 g L−1 glucose) Dulbecco’s modified Eagle’s medium (DMEM, Thermo Fisher Scientific) supplemented with 10% fetal bovine serum (FBS, Natocor), penicillin/streptomycin (100 units mL−1 and 100 µg mL−1, respectively, Thermo Fisher Scientific) and 110 mg L−1 of sodium pyruvate (Thermo Fisher Scientific) in a 37 °C humidified incubator containing 5% CO2. Polyethylenimine (PEI, PolyAR, UBA) was used for transfection. Briefly, cells were plated (2 × 106 HEK-293T cells per 100 mm plate, 5 × 104 HeLa Kyoto cells per well in 12 well plate) and grown for 24 h before transfection. pCMV_MTS_NB_4A7, pCMV_MTS_NB_6B1, pCMV_MTS_NB_16C10, and pCMV_MTS_NB_28F6 vectors encoding each NB (optimized for mammalian cells expression) preceded by the mitochondrial transit signal (MTS) from the citrate synthase enzyme for the mitochondrial matrix localization were transfected according to the PEI manufacturer’s instructions. For the immunofluorescence assays, p-ds-Red2-mito_vector (kindly provided by Dr. Roxana Gorojod, IQUIBICEN, UBA) was co-transfected with the MTS_NB in the same conditions. After transfection, cells were grown for 48 h.
SDS-PAGE and Western blotting assay
Protein lysates were prepared using RIPA low salt buffer (20 mM Tris-HCl, 150 mM NaCl, 0,1% Tween 20, 2 mM EDTA, pH 7.40) and complete protease inhibitor solution (Thermo Fisher). Protein concentration was determined using a Bradford reagent (Thermo Fisher). Protein samples (either purified or complete lysates) were boiled in a sample buffer (4% SDS, 20% glycerol, 120 mM Tris-HCl, pH 6.8, 0.002% bromophenol blue, 200 mM 2-mercaptoethanol) and subjected to 16% SDS-PAGE. Electrophoresis was carried out at room temperature for 20 min at 90 V and 1.5 h at 150 V. Proteins were stained with Coomassie Brilliant Blue G-250. When Western blotting analyses were performed, proteins were transferred to either a PVDF or a nitrocellulose membrane (0.2 μm, Thermo Fisher and BioRad) for 1 h at 100 V. Membranes were blocked for 1 h at room temperature with 5% skimmed milk in 0.05% Tween TBS buffer. Blocked membranes were incubated overnight at 4 °C with either an anti-human FXN mAb (abcam, ab 110328), an anti-His6 mAb (MA-125, Thermo Fisher), or an anti-GAPDH mAb (SC-47724). Afterward, HRP-conjugated anti-mouse was incubated for 1 h at room temperature and visualized by enhanced chemiluminescence (Clarity Substrate, Biorad) using Amersham Imager 680.
Aconitase and succinate dehydrogenase activity after NB transfection
Briefly, HEK-293T cells were plated (2 × 106 cells per 100 mm plate) and grown for 24 h before transfection (using PEI) with pCMV_MTS_NB_4A7, pCMV_MTS_NB_6B1, pCMV_MTS_NB_16C10, and pCMV_MTS_NB_28F6, according to the PEI manufacturer’s instructions (an empty vector control was included). After 48 h, protein lysates were prepared using either PBS buffer or aconitase reaction buffer and complete protease inhibitor (Thermo Fisher). Protein concentration was determined using a Bradford reagent (Thermo Fisher). SDH activity was measured using the SDH Activity Assay Kit (Colorimetric) (Abcam; ab228560). Aconitase activity was measured using the Aconitase Enzyme Kit (Abcam, ab109712). All assays were performed following the manufacturer’s instructions.
Mitochondrial oxygen consumption rate measurements after NB transfection
The mitochondrial OCR and ECAR were monitored in vivo in real-time using a Seahorse XFp analyzer (XFp, Agilent). For this, HEK-293T cells were plated (2 × 105 cells in a 6-well plate) and grown for 24 h before transfection (using PEI) with pCMV_MTS_NB_4A7, pCMV_MTS_NB_6B1, pCMV_MTS_NB_16C10, and pCMV_MTS_NB_28F6, according to the PEI manufacturer’s instructions (an empty vector was included as a control condition). After 24 h of transfection, 2.5 × 104 cells/well were plated on pre-coated (with PEI, Sigma) Seahorse XFp Cell Culture Miniplates (Agilent) and maintained in culture conditions for 24 h, when bioenergetics was assessed. Seahorse assays were performed on confluent plates (criterion: cells covering 99.90 ± 0.05% of the plate surface, quantified over a grid of 25 × 25 µm). This criterion was confirmed by quantification of DAPI-positive nuclei at the end of the Seahorse assays. On the day of the assay, the culture medium was aspirated and replaced by XF Base Medium (Agilent) supplemented with 25 mM D-glucose, 1 mM sodium pyruvate, and 2 mM l-glutamine, pH 7.4. Cells were incubated with this medium for 1 h at 37 °C in a non-CO2 incubator, and then the microplate was loaded into the Seahorse XFp Analyzer (Agilent) following the manufacturer’s instructions. Cells were titrated with 0.125–2.0 µM carbonyl cyanide FCCP to render the maximum oxygen consumption rate (OCR), and these concentrations were used for experiments. The OCR was determined at the beginning of the assay (basal OCR) and after the sequential addition of 1.0 µM oligomycin (Oligo), 1.0 µM FCCP, and 0.5 µM rotenone plus antimycin A (Rot/AA). Three basal rates and three response rates (after adding a compound) were measured, and the average of these rates was used for data analysis. Respiratory parameters were obtained as follows: basal respiration is the baseline OCR. Respiration driving proton leak is the OCR after the addition of 1 µM Oligo. Respiration driving ATP synthesis is the Basal respiration minus Respiration driving proton leak. Maximum respiration is the OCR after the addition of 1 µM FCCP. Spare respiratory capacity (SRC) is the maximum respiration minus the basal respiration.
Values were expressed as a percentage of the OCR corresponding to the last baseline rate (100%). Data were normalized to the cell number at the end of the assay.
Co-immunoprecipitation assay
HEK-293T cell lysates (500 μg) were incubated overnight at 4 °C under agitation with 4 μg of a 6×-His tag antibody (MA 1-135, Thermo Fisher). Binding to protein G magnetic beads (Biorad) was performed for 1 h at 4 °C (previously, beads were washed with the same RIPA buffer). Three washes were performed after binding and then eluted with 25 μL of a 1× sample buffer (for SDS-PAGE) containing 8% 2-mercaptoethanol. After that, the co-immunoprecipitated proteins were analyzed by Western blotting.
LC-MS analysis
For mass spectrometry analysis, the SDS-PAGE fragments were treated according to LaBaer and coworkers73. A volume of 4 μL of each sample was injected. Peptide separations were performed on a nano HPLC Ultimate3000 (Thermo Scientific) using a nano column EASY-Spray ES901 (15 cm × 50 μm ID, PepMap RSLC C18). The mobile phase flow rate was 300 nL/min using 0.1% formic acid in water (solvent A) and 0.1% formic acid and 100% acetonitrile (solvent B). The gradient profile was set as follows: 4-30% solvent B for 64 min, 30-80% solvent B for 7 min and 80% solvent B for 1 min. MS analysis was performed using a Q-Exactive HF mass spectrometer (Thermo Scientific). 1.9 kV of liquid junction voltage and 250 °C of capillary temperature were used for ionization. The entire scan method employed a m/z 375–2000 mass selection, an Orbitrap resolution of 120,000 (at m/z 200), a target automatic gain control (AGC) value of 1e6, and a maximum injection time of 100 msec. After the survey scan, the 15 most intense precursor ions were selected for MS/MS fragmentation. Fragmentation was performed with a normalized collision energy of 28 eV, and MS/MS scans were acquired with a dynamic first mass, AGC target was 5e5, resolution of 30000 (at m/z 200), isolation window of 1.4 m/z units, and maximum IT was 55 ms. Charge state screening was enabled to reject unassigned, singly charged, and equal or more than six protonated ions. A dynamic exclusion time of 25 s was used to discriminate against previously selected ions.
Mass spectrometry data analysis
Using standardized workflows, MS data were analyzed with Proteome Discoverer (versión 2.4.1.15). Mass spectra *.raw files were searched against a database from Homo sapiens (UP000005640) and NB-16c10 sequence. Precursor and fragment mass tolerance were set to 10 ppm and 0.02 Da, respectively, allowing two missed cleavages. The following modifications were set: −Max. Equal modifications per peptide: 3. Max Dynamic Modifications per peptide: 4. The dynamic modifications included in the analysis were (i) oxidation (+15.995 Da), (ii) N-terminal modification, acetylation (+42.011 Da), (iii) N-terminal modification, Met-loss (−131.040 Da), (iv) N-terminal modification Met-loss plus acetylation (−89.030 Da); the static modification included was the carbamidomethylation (+57.021 Da).
Immunofluorescence assays
For fluorescent reporter assays, cells were plated either into 12-well plates with precision cover glasses or 8-well (Nunc® Lab-Tek® II) coverglass imaging plates. After 48 h transfection, cells were fixed with 4% paraformaldehyde in PBS for 10 min at room temperature, washed 3 times with PBS, and permeabilized for 5 min with 0.2% Triton X-100 in the same buffer. Blocking was performed for 1 h in 1% BSA solution. After incubation for 1 h or overnight (4 ºC) with the primary antibody and then the secondary antibody (Thermo Fisher) in blocking solution for 1 h, cells were extensively washed with PBS, and nuclei were stained with Hoescht dye. Images were captured on an Olympus FV-1000 fluorescence microscope with a 60× oil immersion objective and a Coolsnap HQ2 CCD camera (Photometrics). Alternatively, images were captured with a Zeiss LSM980 instrument and analyzed with Zen System software.
Proximity ligation assay (PLA)
PLA was performed using a Duolink® In Situ Red Starter Kit Mouse/Rabbit (DUO92101, Sigma-Aldrich). HeLa Kyoto cells were fixed with 4% PFA in PBS for 10 min, then washed three times with PBS for 5 min each, followed by permeabilization with 0.2% (v/v) Triton X-100 in PBS for 5 min. After three additional PBS washes for 5 min each, cells were blocked with 1% (w/v) BSA in PBS at room temperature for 1 h. Next, the cells were incubated with primary antibodies: an anti-FXN (mouse) and an anti-HA (rabbit), as previously described for the immunofluorescence experiments. The remaining steps followed the manufacturer’s instructions and those described by Doni and co-workers58. Briefly, cells were washed and incubated for 60 min at 37 °C in a preheated, humidified chamber with a dilution of oligonucleotide-conjugated secondary antibodies (anti-rabbit PLUS® DUO82002, Sigma-Aldrich, and anti-mouse MINUS® DUO82004, Sigma-Aldrich). After washing, the cells were incubated with DNA Ligase for 30 min at 37 °C. Subsequently, they were washed and kept in the dark for 100 min at 37 °C with a solution containing DNA polymerase and the red fluorescently labeled oligonucleotides. Cells were washed again and mounted using Duolink in situ mounting medium with DAPI (DUO82040, Sigma-Aldrich). Finally, fluorescence microscopy images were captured with a Zeiss LSM980 instrument and analyzed with Zen System software.
Assembly of the NB:FXN complexes
For structural purposes, FXN:NB_16C10, FXN:NB_4A7 and FXN:NB_6B1 complexes were assembled by incubation and then copurified by using the His-tag present in the nanobodies. Briefly, the complexes were copurified by HisTrap FF column using an imidazole gradient from 20 mM to 500 mM in 50 mM Tris-HCl, pH 7.5, 300 mM NaCl, and 1 mM DTT. The complexes were eluted at 130 mM of imidazole. The eluted fractions containing the desired complexes were identified using SDS-PAGE, and concentrations were determined by NanoDrop. All the proteins were collected and dialyzed for imidazole removal using a Spectra/PorTM 1RC dialysis membrane (of 6–8 kDa MWCO). The resulting protein solutions were concentrated on an Amicon Ultra-4 Centrifugal Filter (MWCO 10 kDa; Millipore Sigma, Burlington, MA). The final concentration for FXN:NB_16C10, FXN:NB_4A7, and FXN:NB_6B1 was 7.5 mg mL−1, 3.5 mg mL−1, and 11 mg mL−1, respectively.
NB:FXN crystallization experiments
Initial crystallization screening using the copurified complexes was performed at 298 K using the sitting-drop vapor-diffusion method with an available collection of commercial conditions. The drops consisted of 0.2 μL of protein solution (7.5 mg mL−1, 3.5 mg mL−1 or 11 mg mL−1 of protein complex in a buffer containing 30 mM Tris-HCl pH 7.5, 300 mM NaCl, 1 mM DTT) and 0.2 μL well solution, and were equilibrated against 60 μL of well solution. The extensive screening rendered crystals in several conditions and their corresponding diffraction quality was checked on the beamline BL13-XALOC at the ALBA Synchrotron (Barcelona, Spain). Best diffracting crystals were subsequently scaled up and optimized, yielding crystals grown in 15% PEG 10 K, 0.1 M ammonium acetate and 0.1 M Bis-Tris pH 5.5 (FXN:NB_16C10); 0.2 M Na thiocyanate pH 6.9, 20% PEG 3350 (FXN:_NB4A7); and 0.1 M HEPES pH 7.5, 0.8 M potassium sodium tartrate (FXN:NB_6B1).
X-ray data collection and processing
All data were collected from frozen crystals at 100 K with the PILATUS 6 M detector at beamline XALOC (ALBA Synchrotron, Barcelona, Spain). Data processing and scaling were accomplished using XDS74, POINTLESS, and AIMLESS75 as implemented in autoPROC76. Statistics for the crystallographic data and structure solution are summarized in Table 1.
Crystal structure determination, model building, and refinement
The NB:FXN structures were solved by the molecular-replacement method, as implemented in the program PHASER77,78 using ab initio models given by AlphaFold47,48 as search models. Then, its initial models were subjected to iterative cycles of model building and refinement with Coot79 and REFMAC80, respectively. Final refinement cycles were performed with PHENIX81, yielding the refinement and data collection statistics summarized in Table 1. Figures were generated using PyMOL82 and ChimeraX83. The final refinement parameters are summarized in Table 1.
Small angle X-ray scattering (SAXS)
SAXS experiments were performed at the beamline B21 of the Diamond Light Source84. Samples of 45 μL of all the complexes at different concentrations were loaded onto an SRT-C SEC-300 (Sepax) column equilibrated in buffer (20 mM Tris-HCl pH 8.0 and 150 mM NaCl) and connected to an Agilent 1200 HPLC system at 18 °C. The continuously eluting samples were exposed for 3 s in 10 s acquisition blocks using an X-ray wavelength of 1 Å and a sample-to-detector (Eiger 4 M) distance of 3.7 m. The data covered a momentum transfer range of 0.0032 < q < 0.34 Å−1. The frames recorded immediately before the sample elution were subtracted from the protein scattering profiles. The Scåtter software package (www.bioisis.net) was used to analyze data, buffer-subtraction, scaling, merging, and checking possible radiation damage of the samples. The Rg value was calculated with the Guinier approximation, assuming that at very small angles q < 1.3/Rg. The particle distance distribution, Dmax, was calculated from the scattering pattern with GNOM, and shape estimation was carried out with DAMMIF/DAMMIN; all these programs are included in the ATSAS package85,86. The protein molecular mass was estimated with GNOM. Interactively generated PDB-based homology models were made using the program COOT by manually adjusting the X-ray structures obtained in this work into the envelope given by SAXS until a good correlation between the real-space scattering profile calculated for the homology model matched the experimental scattering data. This was computed with the program FoXS87.
Statistics and reproducibility
For comparisons among multiple groups, one-way ANOVA was used, followed by a Dunnett’s test for comparisons against a single control group or a Bonferroni’s test for all pairwise comparisons. The mean ± SEM (Standard error of the mean) was usually presented. Analysis was performed using GraphPad Prism 10.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Atomic coordinates and structure factors are available in the Protein Data Bank (PDB) under the accession codes 9HO6, 9HO4, and 9HO5. All supporting data for this study are included in the article and its Supplementary Information. The uncropped Western blot images are provided in Supplementary Figs. 17 and 18. The original data can be accessed in the Excel file named “Supplementary_data_MFP_2025.xlsx”. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE88 partner repository with the dataset identifier PXD061538. Additional information is available from the corresponding authors upon reasonable request.
References
Fan, X. et al. Iron-regulated assembly of the cytosolic iron-sulfur cluster biogenesis machinery. J. Biol. Chem. 298, 102094 (2022).
Pandey, A. K., Pain, J., Dancis, A. & Pain, D. Mitochondria export iron-sulfur and sulfur intermediates to the cytoplasm for iron-sulfur cluster assembly and tRNA thiolation in yeast. J. Biol. Chem. 294, 9489–9502 (2019).
Paul, V. D. & Lill, R. Biogenesis of cytosolic and nuclear iron-sulfur proteins and their role in genome stability. Biochim. Biophys. Acta 1853, 1528–1539 (2015).
Lill, R., Srinivasan, V. & Muhlenhoff, U. The role of mitochondria in cytosolic-nuclear iron-sulfur protein biogenesis and in cellular iron regulation. Curr. Opin. Microbiol. 22, 111–119 (2014).
Fox, N. G. et al. Structure of the human frataxin-bound iron-sulfur cluster assembly complex provides insight into its activation mechanism. Nat. Commun. 10, 2210 (2019).
Biederbick, A. et al. Role of human mitochondrial Nfs1 in cytosolic iron-sulfur protein biogenesis and iron regulation. Mol. Cell Biol. 26, 5675–5687 (2006).
Fosset, C. et al. RNA silencing of mitochondrial m-Nfs1 reduces Fe-S enzyme activity both in mitochondria and cytosol of mammalian cells. J. Biol. Chem. 281, 25398–25406 (2006).
Black, K. A. & Dos Santos, P. C. Shared-intermediates in the biosynthesis of thio-cofactors: Mechanism and functions of cysteine desulfurases and sulfur acceptors. Biochim. Biophys. Acta 1853, 1470–1480 (2015).
Marelja, Z. et al. The L-cysteine desulfurase NFS1 is localized in the cytosol where it provides the sulfur for molybdenum cofactor biosynthesis in humans. PLoS ONE 8, e60869 (2013).
Marelja, Z., Stocklein, W., Nimtz, M. & Leimkuhler, S. A novel role for human Nfs1 in the cytoplasm: Nfs1 acts as a sulfur donor for MOCS3, a protein involved in molybdenum cofactor biosynthesis. J. Biol. Chem. 283, 25178–25185 (2008).
Neukranz, Y. et al. Analysis of the cellular roles of MOCS3 identifies a MOCS3-independent localization of NFS1 at the tips of the centrosome. Biochemistry 58, 1786–1798 (2019).
Tong, W. H. & Rouault, T. A. Functions of mitochondrial ISCU and cytosolic ISCU in mammalian iron-sulfur cluster biogenesis and iron homeostasis. Cell Metab. 3, 199–210 (2006).
Parent, A. et al. Mammalian frataxin directly enhances sulfur transfer of NFS1 persulfide to both ISCU and free thiols. Nat. Commun. 6, 5686 (2015).
Boniecki, M. T., Freibert, S. A., Muhlenhoff, U., Lill, R. & Cygler, M. Structure and functional dynamics of the mitochondrial Fe/S cluster synthesis complex. Nat. Commun. 8, 1287 (2017).
Richards, T. A. & van der Giezen, M. Evolution of the Isd11-IscS complex reveals a single alpha-proteobacterial endosymbiosis for all eukaryotes. Mol. Biol. Evol. 23, 1341–1344 (2006).
Wiedemann, N. et al. Essential role of Isd11 in mitochondrial iron-sulfur cluster synthesis on Isu scaffold proteins. EMBO J. 25, 184–195 (2006).
Herrera, M. G. et al. Structure of the Human ACP-ISD11 Heterodimer. Biochemistry 58, 4596–4609 (2019).
Herrera, M. G., Pignataro, M. F., Noguera, M. E., Cruz, K. M. & Santos, J. Rescuing the rescuer: on the protein complex between the human mitochondrial Acyl carrier protein and ISD11. ACS Chem. Biol. 13, 1455–1462 (2018).
Cai, K., Frederick, R. O., Tonelli, M. & Markley, J. L. Interactions of iron-bound frataxin with ISCU and ferredoxin on the cysteine desulfurase complex leading to Fe-S cluster assembly. J. Inorg. Biochem. 183, 107–116 (2018).
Cai, K., Tonelli, M., Frederick, R. O. & Markley, J. L. Human mitochondrial ferredoxin 1 (FDX1) and ferredoxin 2 (FDX2) both bind cysteine desulfurase and donate electrons for iron-sulfur cluster biosynthesis. Biochemistry 56, 487–499 (2017).
Gervason, S. et al. Physiologically relevant reconstitution of iron-sulfur cluster biosynthesis uncovers persulfide-processing functions of ferredoxin-2 and frataxin. Nat. Commun. 10, 3566 (2019).
Sheftel, A. D. et al. Humans possess two mitochondrial ferredoxins, Fdx1 and Fdx2, with distinct roles in steroidogenesis, heme, and Fe/S cluster biosynthesis. Proc. Natl. Acad. Sci. USA 107, 11775–11780 (2010).
Webert, H. et al. Functional reconstitution of mitochondrial Fe/S cluster synthesis on Isu1 reveals the involvement of ferredoxin. Nat. Commun. 5, 5013 (2014).
Fox, N. G., Das, D., Chakrabarti, M., Lindahl, P. A. & Barondeau, D. P. Frataxin accelerates [2Fe-2S] cluster formation on the human Fe-S assembly complex. Biochemistry 54, 3880–3889 (2015).
Koutnikova, H., Campuzano, V. & Koenig, M. Maturation of wild-type and mutated frataxin by the mitochondrial processing peptidase. Hum. Mol. Genet. 7, 1485–1489 (1998).
Schmucker, S., Argentini, M., Carelle-Calmels, N., Martelli, A. & Puccio, H. The in vivo mitochondrial two-step maturation of human frataxin. Hum. Mol. Genet. 17, 3521–3531 (2008).
Nordin, A., Larsson, E. & Holmberg, M. The defective splicing caused by the ISCU intron mutation in patients with myopathy with lactic acidosis is repressed by PTBP1 but can be derepressed by IGF2BP1. Hum. Mutat. 33, 467–470 (2012).
Saha, P. P. et al. The presence of multiple cellular defects associated with a novel G50E iron-sulfur cluster scaffold protein (ISCU) mutation leads to development of mitochondrial myopathy. J. Biol. Chem. 289, 10359–10377 (2014).
Farhan, S. M. et al. Exome sequencing identifies NFS1 deficiency in a novel Fe-S cluster disease, infantile mitochondrial complex II/III deficiency. Mol. Genet. Genom. Med. 2, 73–80 (2014).
Saha, P. P., Srivastava, S., Kumar, S. K. P., Sinha, D. & D’Silva, P. Mapping key residues of ISD11 critical for NFS1-ISD11 subcomplex stability: implications in the development of mitochondrial disorder, COXPD19. J. Biol. Chem. 290, 25876–25890 (2015).
Pandolfo, M. & Pastore, A. The pathogenesis of Friedreich ataxia and the structure and function of frataxin. J. Neurol. 256, 9–17 (2009).
Pandolfo, M. Friedreich ataxia: detection of GAA repeat expansions and frataxin point mutations. Methods Mol. Med. 126, 197–216 (2006).
Bellanda, M. et al. Exploring iron-binding to human frataxin and to selected Friedreich ataxia mutants by means of NMR and EPR spectroscopies. Biochim. Biophys. Acta Proteins Proteom. 1867, 140254 (2019).
Espeche, L. D. et al. Conformational stability, dynamics and function of human frataxin: Tryptophan side chain interplay. Arch. Biochem. Biophys. 715, 109086 (2022).
Faggianelli, N. et al. Analyzing the effects of a G137V mutation in the FXN gene. Front. Mol. Neurosci. 8, 66 (2015).
Faraj, S. E., Roman, E. A., Aran, M., Gallo, M. & Santos, J. The alteration of the C-terminal region of human frataxin distorts its structural dynamics and function. FEBS J. 281, 3397–3419 (2014).
Van Driest, S. L., Gakh, O., Ommen, S. R., Isaya, G. & Ackerman, M. J. Molecular and functional characterization of a human frataxin mutation found in hypertrophic cardiomyopathy. Mol. Genet. Metab. 85, 280–285 (2005).
Roman, E. A. et al. Protein stability and dynamics modulation: the case of human frataxin. PLoS ONE 7, e45743 (2012).
Castro, I. H. et al. Frataxin Structure and Function. Subcell. Biochem. 93, 393–438 (2019).
Pignataro, M. F. et al. Selection of synthetic proteins to modulate the human frataxin function. Biotechnol. Bioeng. 120, 409–425 (2023).
Muyldermans, S. A guide to: generation and design of nanobodies. FEBS J. 288, 2084–2102 (2021).
Bao, G., Tang, M., Zhao, J. & Zhu, X. Nanobody: a promising toolkit for molecular imaging and disease therapy. EJNMMI Res. 11, 6 (2021).
Sang, Z., Xiang, Y., Bahar, I. & Shi, Y. Llamanade: an open-source computational pipeline for robust nanobody humanization. Structure 30, 418–429.e413 (2022).
Messer, A. & Butler, D. C. Optimizing intracellular antibodies (intrabodies/nanobodies) to treat neurodegenerative disorders. Neurobiol. Dis. 134, 104619 (2020).
Yang, E. Y. & Shah, K. Nanobodies: next generation of cancer diagnostics and therapeutics. Front. Oncol. 10, 1182 (2020).
AlQuraishi, M. AlphaFold at CASP13. Bioinformatics 35, 4862–4865 (2019).
Jumper, J. et al. Applying and improving AlphaFold at CASP14. Proteins 89, 1711–1721 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Sewell, K. E. et al. Direct cysteine desulfurase activity determination by NMR and the study of the functional role of key structural elements of human NFS1. ACS Chem. Biol. 18, 1534–1547 (2023).
Correia, A. R., Pastore, C., Adinolfi, S., Pastore, A. & Gomes, C. M. Dynamics, stability and iron-binding activity of frataxin clinical mutants. FEBS J. 275, 3680–3690 (2008).
Lazaropoulos, M. et al. Frataxin levels in peripheral tissue in Friedreich ataxia. Ann. Clin. Transl. Neurol. 2, 831–842 (2015).
Dhe-Paganon, S., Shigeta, R., Chi, Y. I., Ristow, M. & Shoelson, S. E. Crystal structure of human frataxin. J. Biol. Chem. 275, 30753–30756 (2000).
Paakkonen, J., Janis, J. & Rouvinen, J. Calculation and visualization of binding equilibria in protein studies. ACS Omega 7, 10789–10795 (2022).
Mitchell, L. S. & Colwell, L. J. Analysis of nanobody paratopes reveals greater diversity than classical antibodies. Protein Eng. Des. Sel. 31, 267–275 (2018).
Bulteau, A. L. et al. Frataxin acts as an iron chaperone protein to modulate mitochondrial aconitase activity. Science 305, 242–245 (2004).
Mansilla, S. et al. Redox sensitive human mitochondrial aconitase and its interaction with frataxin: in vitro and in silico studies confirm that it takes two to tango. Free Radic. Biol. Med. 197, 71–84 (2023).
Doni, D. et al. A combined spectroscopic and in silico approach to evaluate the interaction of human frataxin with mitochondrial superoxide dismutase. Biomedicines 9. https://doi.org/10.3390/biomedicines9121763 (2021).
Doni, D. et al. Human frataxin, the Friedreich ataxia deficient protein, interacts with mitochondrial respiratory chain. Cell Death Dis. 14, 805 (2023).
Steinhilper, R. et al. Two-stage binding of mitochondrial ferredoxin-2 to the core iron-sulfur cluster assembly complex. Nat. Commun. 15, 10559 (2024).
Castro, I. H. et al. Relationship between activity and stability: design and characterization of stable variants of human frataxin. Arch. Biochem. Biophys. 691, 108491 (2020).
Doni, D. et al. The displacement of frataxin from the mitochondrial cristae correlates with abnormal respiratory supercomplexes formation and bioenergetic defects in cells of Friedreich ataxia patients. FASEB J. 35, e21362 (2021).
Clark, E., Butler, J. S., Isaacs, C. J., Napierala, M. & Lynch, D. R. Selected missense mutations impair frataxin processing in Friedreich ataxia. Ann. Clin. Transl. Neurol. 4, 575–584 (2017).
Kardani, K., Milani, A., Shabani, H. S. & Bolhassani, A. Cell penetrating peptides: the potent multi-cargo intracellular carriers. Expert Opin. Drug Deliv. 16, 1227–1258 (2019).
Borgert, L., Becker, T. & den Brave, F. Conserved quality control mechanisms of mitochondrial protein import. J. Inherit. Metab. Dis. 47, 903–916 (2024).
Ackaert, C. et al. Immunogenicity risk profile of nanobodies. Front. Immunol. 12, 632687 (2021).
Tsai, C.-L. & Barondeau, D. P. Human frataxin is an allosteric switch that activates the Fe−S cluster biosynthetic complex. Biochemistry 49, 9132–9139 (2010).
Moest, R. R. Hydrogen sulfide determination by the methylene blue method. Anal. Chem. 47, 1204–2205 (1975).
Romao, E. et al. Construction of high-quality camel immune antibody libraries. Methods Mol. Biol. 1701, 169–187 (2018).
Pavan, M. F. et al. SARS-CoV-2 specific nanobodies neutralize different variants of concern and reduce virus load in the brain of h-ACE2 transgenic mice. Viruses 16. https://doi.org/10.3390/v16020185 (2024).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Johnson, B. A. Using NMRView to visualize and analyze the NMR spectra of macromolecules. Methods Mol. Biol. 278, 313–352 (2004).
Xu, J., Sarma, A. V. S., Wei, Y., Beamer, L. J. & Van Doren, S. R. Multiple ligand-bound states of a phosphohexomutase revealed by principal component analysis of NMR peak shifts. Sci. Rep. 7, 5343 (2017).
Link, A. J. & Labaer, J. In-gel trypsin digest of gel-fractionated proteins. Cold Spring Harb. Protoc. 2009, https://doi.org/10.1101/pdb.prot5110 (2009).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Crystallogr. D Biol. Crystallogr. 69, 1204–1214 (2013).
Vonrhein, C. et al. Data processing and analysis with the autoPROC toolbox. Acta Crystallogr. D Biol. Crystallogr. 67, 293–302 (2011).
McCoy, A. J. Solving structures of protein complexes by molecular replacement with Phaser. Acta Crystallogr. D Biol. Crystallogr. 63, 32–41 (2007).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P. & Debreczeni, J. E. The use of molecular graphics in structure-based drug design. Methods Mol. Biol. 841, 143–159 (2012).
Winn, M. D., Murshudov, G. N. & Papiz, M. Z. Macromolecular TLS refinement in REFMAC at moderate resolutions. Methods Enzymol. 374, 300–321 (2003).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Schrödinger, LLC. The PyMOL Molecular Graphics System, Version 1.8 (2015).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Cowieson, N. P. et al. Beamline B21: high-throughput small-angle X-ray scattering at Diamond Light Source. J. Synchrotron Radiat. 27, 1438–1446 (2020).
Franke, D. et al. ATSAS 2.8: a comprehensive data analysis suite for small-angle scattering from macromolecular solutions. J. Appl. Crystallogr. 50, 1212–1225 (2017).
Petoukhov, M. V. et al. New developments in the ATSAS program package for small-angle scattering data analysis. J. Appl. Crystallogr. 45, 342–350 (2012).
Schneidman-Duhovny, D., Hammel, M., Tainer, J. A. & Sali, A. FoXS, FoXSDock and MultiFoXS: single-state and multi-state structural modeling of proteins and their complexes based on SAXS profiles. Nucleic Acids Res. 44, W424–W429 (2016).
Perez-Riverol, Y. et al. The PRIDE database at 20 years: 2025 update. Nucleic Acids Res. 53, D543–D553 (2025).
Acknowledgements
This work was supported by Universidad de Buenos Aires UBACyT20020190100338BA, CONICET, and Friedreich´s Ataxia Research Alliance (FARA, grant 2023-2025). The work in Spain was funded by the Spanish Ministry of Science and Innovation by grants PID2020-115331GB-100 and PID2023-153118OB-100 to J.A.H., and PID2022-136750NB-I00 to R.M. We gratefully acknowledge the staff of the XALOC beamline at the ALBA Synchrotron in Barcelona for their valuable assistance with data collection. We would also like to thank Lic. Alejo Cantoia, Dr. Gernán Rosano, and Dr. Eduardo Ceccarelli from the UEM-IBR (Mass spectrometry unit of the Instituto de Biología Molecular y Celular de Rosario, Argentina) for technical assistance and data processing of LC-MS experiments.
Author information
Authors and Affiliations
Contributions
M.F.P. and J.S. performed the experiments, analyzed the results and structures, and wrote the paper. N.B.F. performed co-immunoprecipitations, Western blots, and analysis of the results. A.E.G., R.M., and J.A.H.D. performed the crystallization, determined the X-ray structure, analyzed the structures, and wrote the paper. I.G.M. performed SAXS experiments and analysis. J.G., A.E.G., and N.A.R. expressed and purified some NB candidates. M.N. purified FXN and the NFS1/ACP-ISD11 and ISCU proteins. A.V. performed the acquisition and processing of fluorescence microscopy images. H.G.G. helped to measure ACO and SDH enzymatic activities. M.A. acquired 2D NMR spectra. L.I.I., M.P.P., G.V.P., and M.B. made the library of nanobodies and the selection by phage display.
Corresponding authors
Ethics declarations
Competing interests
M.F.P., J.S., J.G., and L.I.I. hold a related National patent application: “NANOANTICUERPOS CONTRA FRATAXINA (FXN) O FRAGMENTOS DE LA MISMA Y MÉTODOS AFINES”. Application number: P20230103084 (Argentina). Application date: 15/11/2023. All other authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary handling editors: Laura Rodriguez Perez. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pignataro, M.F., Fernández, N.B., Garay-Alvarez, A. et al. Nanobodies as tools for studying human frataxin biology. Commun Biol 9, 181 (2026). https://doi.org/10.1038/s42003-025-09458-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-09458-x