Abstract
Hepatocellular carcinoma (HCC) is a highly lethal malignancy, with epithelial-mesenchymal transition (EMT)-driven metastasis a key factor for poor prognosis. The C5a/C5a receptor (C5aR) pathway significantly facilitates HCC cell EMT, yet no approved anti-cancer drugs specifically target C5aR. LukS-PV, a component of Staphylococcus aureus-secreted Panton-Valentine leukocidin (PVL), specifically targets C5aR and exerts anti-tumor effects in hematological and solid tumors. However, its impact on HCC EMT and mechanisms remains unknown. Our study showed LukS-PV targets C5aR to inhibit HCC cell EMT, migration, invasion, and in vivo lung metastasis. Mechanistically, LukS-PV downregulates B-cell lymphoma 6 (BCL6), reducing histone deacetylase 6 (HDAC6) expression. Decreased HDAC6 increases heat shock protein 60 (HSPD1) acetylation, promoting its ubiquitin-mediated degradation and EMT inhibition. This study demonstrates LukS-PV targets C5aR to inhibit HCC EMT via the BCL6/HDAC6/HSPD1 axis, highlighting its potential as an HCC therapeutic agent. These findings provide valuable EMT regulatory insights and identify potential HCC therapeutic targets.
Similar content being viewed by others
Introduction
Hepatocellular carcinoma (HCC) is the sixth most prevalent malignancy and third leading cause of cancer-related mortality globally, poses a severe threat to human health1. Metastasis is a pivotal factor contributing to death of HCC patients. The epithelial-mesenchymal transition (EMT), a critical molecular event in tumor metastasis, is closely associated with poor prognosis in HCC patients2,3. Therefore, it is essential to investigate the molecular mechanisms that regulate EMT in HCC and to explore potential therapeutic approaches for inhibiting HCC metastasis.
The receptor for complement C5a, C5aR, is reportedly highly expressed in a variety of tumor types4,5. The C5a-C5aR signaling pathway can promote the growth and EMT of cancer cells, including HCC5,6. This suggests that C5aR may serve as an important molecular target for inhibiting EMT. Blocking C5a-C5aR signaling can effectively inhibit tumor growth and metastasis7,8. However, there are currently no approved anti-cancer drugs that specifically target C5aR.
PVL is a cytotoxin secreted by Staphylococcus aureus that consists of two components: LukS-PV and LukF-PV. LukS-PV can bind to C5aR on the cell membrane, then recruit LukF-PV, which results in cell perforation9,10. Our previous study found that recombinant LukS-PV has anti-tumor effects on both hematological malignancies and solid tumors by targeting C5aR, with no in vitro pore-forming activity and no observable toxic effects in vivo11,12,13.
Here, we report a critical role of LukS-PV in inhibiting EMT and metastasis of HCC by regulating acetylation modifications. Mechanistically, LukS-PV targets C5aR to inhibit BCL6 expression, thereby reducing HDAC6 levels. HDAC6 downregulation increases acetylation of its substrate, HSPD1, leading to HSPD1 ubiquitination and degradation, which ultimately inhibits EMT and metastasis in HCC. In this study, we (1) confirmed that BCL6 acts as a transcriptional activator of HDAC6 in HCC; (2) identified HSPD1 as a substrate for HDAC6-mediated deacetylation, and (3) elucidated the mechanism by which HDAC6 regulates HSPD1 acetylation to mediate its ubiquitination. This study suggests that LukS-PV may serve as a therapeutic agent for C5aR-positive HCC and that BCL6, HDAC6, and HSPD1 emerge as promising targets for HCC treatment.
Results
LukS-PV inhibits EMT and metastatic progression in HCC
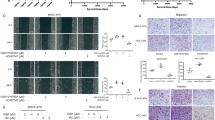
To ascertain whether LukS-PV inhibits EMT in HCC, we treated HepG2 and Huh7 cells with different concentrations of LukS-PV for 24 h. Western blot analysis showed that LukS-PV treatment significantly upregulated the epithelial marker E-cadherin and downregulated the mesenchymal markers N-cadherin and vimentin in HCC cells (Fig. 1a). Transwell assays showed that LukS-PV treatment could suppress both the migration and invasion abilities of HCC cells in a concentration-dependent manner (Fig. 1b, c). In lung metastasis models, LukS-PV treatment reduced lung metastatic nodules formation compared with the PBS group (Fig. 1d, e). Collectively, these data demonstrate that LukS-PV exerts potent anti-EMT and anti-metastatic effects in HCC.
a E-cadherin, N-cadherin, and vimentin protein expression levels were detected in HCC cells treated with different concentrations of LukS-PV for 24 h. b, c Representative images and statistical analysis of transwell migration/invasion assays in HCC cells treated with different concentrations of LukS-PV for 24 h. Scale bar, 100 μm. d Representative images of the gross lung appearance and HE staining of the lung lobes. e Statistical analysis of metastatic nodules in the lung metastasis mouse model. Scale bar, 2 mm. Transwell (mean ± SD; n = 3) and metastatic lung nodules data (mean ± SD; n = 5) were analyzed by unpaired two-tail t-test or one-way ANOVA. **, p < 0.01; ***, p < 0.001; ****, p < 0.0001; ns not significant.
LukS-PV targets C5aR to antagonize C5a and inhibit EMT in HCC
To investigate whether LukS-PV can inhibit EMT in HCC by targeting C5aR, we first verified that C5aR protein levels were higher in HCC cells than those in the human hepatic stellate cell line LX2 (Supplementary Fig. 1a). We then performed immunofluorescence experiments and found that LukS-PV co-localized with C5aR in the cell membrane of HCC cells (Supplementary Fig. 1b). Moreover, we found that C5aR knockdown attenuated LukS-PV’s inhibitory effects on HCC cell EMT, migration, and invasion, while C5aR overexpression enhanced these effects (Fig. 2a–d and Supplementary Fig. 1c, d). We further found that human-derived complement C5/C5a protein promotes HCC cell EMT, migration, and invasion, while LukS-PV inhibits this effect by antagonizing C5a (Fig. 2e–h and Supplementary Fig. 1e, f). These experiments collectively suggest that LukS-PV targets C5aR to antagonize C5a and inhibit EMT in HCC.
a Western blot analysis was used to detect the protein expression levels of E-cadherin, N-cadherin, and vimentin in HepG2 cells ± C5aR knockdown treated with or without 1 μM LukS-PV for 24 h. b Statistical analysis of transwell migration/invasion assays in HepG2 cells ± C5aR knockdown treated with or without 1 μM LukS-PV for 24 h. c Western blot analysis was used to detect the protein expression levels of E-cadherin, N-cadherin, and vimentin in Huh7 cells ± C5aR overexpression treated with or without 1 μM LukS-PV for 24 h. d Statistical analysis of transwell migration/invasion assays in Huh7 cells ± C5aR overexpression treated with or without 1 μM LukS-PV for 24 h. e, g Western blot analysis was used to detect the protein expression levels of E-cadherin, N-cadherin, and vimentin in HepG2 and Huh7 cells treated with 1 μM LukS-PV and/or 20 ng/mL C5a for 24 h. f, h Statistical analysis of transwell migration/invasion assays in HepG2 and Huh7 cells treated with 1 μM LukS-PV and/or 20 ng/mL C5a for 24 h. Transwell data (mean ± SD; n = 3) were analyzed by one-way ANOVA. *, p < 0.05; **, p < 0.01; ***, p < 0.001; ****, p < 0.0001.
LukS-PV inhibits EMT in HCC by downregulating HDAC6
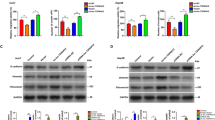
To elucidate the molecular mechanism by which LukS-PV inhibits EMT, we performed RNA-seq in HepG2 cells and found that there were significant alterations in the expression of multiple histone deacetylase (HDAC) genes following LukS-PV treatment, with HDAC6 exhibiting the most pronounced downregulation (Fig. 3a). qRT-PCR and Western blot analyses confirmed that LukS-PV treatment downregulated both HDAC6 mRNA and HDAC6 protein levels in HCC cells, while concurrently enhancing pan-acetylation levels. (Fig. 3b and Supplementary Fig. 2a, b).
a RNA sequencing was performed to identify any changes of HDACs mRNA expression levels in HepG2 cells treated with 1 µM LukS-PV for 24 h. b HDAC6 protein expression levels were detected in HepG2 and Huh7 cells treated with different concentrations of LukS-PV for 24 h. c Western blot analysis was used to examine protein expression level changes of HDAC6, E-cadherin, N-cadherin, and vimentin in Huh7 cells ± HDAC6 overexpression. d Statistical analysis of transwell migration/invasion assays in Huh7 cells ± HDAC6 overexpression. e, f Western blot analysis was used to examine protein expression level changes of HDAC6, E-cadherin, N-cadherin, and vimentin in Huh7 and HepG2 cells ± HDAC6 knockdown. g, h Statistical analysis of transwell migration/invasion assays in Huh7 and HepG2 cells ± HDAC6 knockdown. i Western blot analysis was used to examine protein expression level changes of HDAC6, E-cadherin, N-cadherin, and vimentin in Huh7 cells ± HDAC6 overexpression treated with or without 1 μM LukS-PV for 24 h. j Statistical analysis of transwell migration/invasion assays in Huh7 cells ± HDAC6 overexpression treated with or without 1 μM LukS-PV for 24 h. k, m Representative images of the gross lung appearance and HE staining of the lung lobes. l, n Statistical analysis of the metastatic nodules in the lung metastasis mouse model from the indicated experimental groups. Scale bar, 2 mm. Transwell (mean ± SD; n = 3) and metastatic lung nodules data (mean ± SD; n = 5) were analyzed by unpaired two-tail t-test or one-way ANOVA. *, p < 0.05; **, p < 0.01; ***, p < 0.001.
To further interrogate the biological function of HDAC6 in EMT of HCC, we initially observed that HDAC6 expression levels were significantly elevated in HCC cells compared with LX2 cells (Supplementary Fig. 2c). Overexpressing HDAC6 significantly promoted EMT, migration, and invasion in Huh7 cells (Fig. 3c, d and Supplementary Fig. 2d), whereas HDAC6 knockdown had the opposite effects (Fig. 3e–h and Supplementary Fig. 2e, f). Ectopic expression of HDAC6 significantly impaired the inhibitory effects of LukS-PV treatment on Huh7 cell EMT, migration, and invasion (Fig. 3i, j and Supplementary Fig. 2g). Furthermore, mice injected with HDAC6-knockdown Huh7 cells developed fewer lung metastatic nodules (Fig. 3k, l). Conversely, mice injected with HDAC6-overexpressing Huh7 cells showed enhanced lung metastasis and attenuated the inhibitory effects of LukS-PV treatment (Fig. 3m, n). These findings suggest that HDAC6 plays a crucial role in HCC cell EMT and that LukS-PV inhibits EMT in HCC by downregulating HDAC6.
LukS‑PV inhibits EMT in HCC by decreasing HDAC6 expression via BCL6 downregulation
To clarify the molecular mechanism of LukS-PV-mediated downregulation of HDAC6 expression in HCC cells, we predicted potential upstream transcription factors using five databases (UCSC, JASPAR, GTRD, GTEx, and TCGA)14,15,16,17,18. Through comprehensive analysis on the SRplot platform19, we obtained two candidates (Fig. 4a). Among them, LukS-PV treatment downregulated BCL6 expression, while exerting no significant effect on RUNX2 (Supplementary Fig. 3a). Thus, we selected BCL6 for subsequent investigations. BCL6 is reportedly highly expressed in a variety of cancer types and is associated with tumor drug resistance and metastasis20,21,22. High BCL6 expression can promote HCC metastasis and is associated with poor patient survival23. Notably, the role of BCL6 in regulating HDAC6-mediated EMT in HCC still has not been fully elucidated.
a Venn diagram showing the overlap of transcription factors among the six datasets. b, c The binding capacity of BCL6 to the HDAC6 gene promoter was validated by ChIP assays in HepG2 cells. d The binding capacity of BCL6 at the HDAC6 gene promoter was determined by ChIP-qPCR in HepG2 cells treated with or without 1 µM LukS-PV for 24 h. e Western blot analysis was used to examine the protein expression levels of BCL6, HDAC6, E-cadherin, N-cadherin, and vimentin in HCC cells ± BCL6 overexpression or knockdown. f Western blot analysis was used to examine the protein expression levels of BCL6, HDAC6, E-cadherin, N-cadherin, and vimentin in Huh7 cells ± BCL6 overexpression and/or HDAC6 knockdown. g Western blot analysis was used to examine the protein expression levels of BCL6, HDAC6, E-cadherin, N-cadherin, and vimentin in HCC cells ± BCL6 overexpression treated with or without 1 μM LukS-PV for 24 h. h, j, l Representative images of the gross lung appearance and HE staining of the lung lobes. i, k, m Statistical analysis of the metastatic nodules in the lung metastasis mouse model from the indicated experimental groups. Scale bar, 2 mm. ChIP-qPCR (mean ± SD; n = 3) and metastatic lung nodules data (mean ± SD; n = 5) were analyzed by unpaired two-tail t-test or one-way ANOVA. *, p < 0.05; **, p < 0.01; ***, p < 0.001; ****, p < 0.0001.
To determine whether BCL6 can directly regulate HDAC6 expression, we performed ChIP assays that demonstrated BCL6 binding to the HDAC6 promoter region (Fig. 4b, c). We also found that LukS-PV treatment significantly reduced its enrichment at the HDAC6 promoter region (Fig. 4d and Supplementary Fig. 3b). Moreover, overexpressing BCL6 upregulated HDAC6 expression levels and promoted EMT, migration, and invasion in HCC cells. In contrast, BCL6 knockdown had the opposite effects (Fig. 4e and Supplementary Fig. 3c, d). Furthermore, HDAC6 knockdown remarkably impaired the BCL6 overexpression-induced effects on HCC cell EMT, migration, and invasion (Fig. 4f and Supplementary Fig. 3e, f). Ectopic expression of BCL6 could attenuated the LukS-PV-mediated inhibitory effects on HCC cell EMT, migration, and invasion (Fig. 4g and Supplementary Fig. 3g, h). These findings were corroborated in vivo, BCL6 knockdown suppressed Huh7 cell lung metastasis while its overexpression induced a dramatically higher number of lung metastatic nodules and attenuated LukS-PV’s inhibitory effects (Fig. 4h–k). Notably, HDAC6 knockdown significantly attenuated the role of BCL6 in promoting Huh7 cell lung metastasis (Fig. 4l, m). Taken together, these results suggest that HDAC6 is a functional downstream target of BCL6 in HCC cells and that LukS‑PV inhibits EMT in HCC by targeting the BCL6-HDAC6 axis.
LukS-PV induces HSPD1 acetylation by inhibiting the BCL6/HDAC6 axis
To further elucidate the downstream molecular mechanisms by which HDAC6 can affect HCC metastasis, we performed IP and mass spectrometry experiments and identified 580 potential HDAC6-interacting proteins24 (Supplementary Data 1). Additionally, we performed acetylome analysis and found 134 proteins with significantly upregulated acetylation following LukS-PV treatment (Supplementary Fig. 4a and Supplementary Table 1). Furthermore, we predicted certain proteins that can interact with HDAC6 using the BioGRID25 and Pathway Commons databases26. By comprehensively analyzing these four datasets, we finally screened 19 potential candidates (Fig. 5a and Supplementary Table 2). Among them, we noted that HSPD1 has been previously reported to be associated with tumor metastasis27,28. We analyzed HSPD1 expression using the UALCAN29 and GEPIA30 websites and found elevated HSPD1 expression in HCC compared with normal tissues (Supplementary Fig. 4b), which correlated with poor patient prognosis (Supplementary Fig. 4c, d). Additionally, HSPD1 overexpression enhanced HCC cell EMT, migration, and invasion, whereas HSPD1 knockdown showed the opposite effects (Fig. 5b, c and Supplementary Fig. 4e). However, the interaction between HSPD1 and HDAC6 remains unreported in current studies.
a Venn diagram showing the overlap of proteins among the four datasets. b Statistical analysis of transwell migration/invasion assays in Huh7 cells ± HSPD1 overexpression or knockdown. c Western blot analysis was used to examine the protein expression level changes of HSPD1, E-cadherin, N-cadherin, and vimentin in Huh7 cells ± HSPD1 overexpression or knockdown. d Co-IP analysis of the interaction between HDAC6 and HSPD1 in HepG2 and Huh7 cells. e Confocal assays were used to observe the co-localization of HDAC6 and HSPD1 proteins in HepG2 and Huh7 cells treated with or without LukS-PV. Scale bar, 10 μm. f Western blot analysis was used to examine the protein expression level changes of HDAC6, HSPD1, and acetylated HSPD1 in Huh7 cells ± HDAC6 overexpression. g Western blot analysis was used to examine the protein expression level changes of HDAC6, HSPD1, and acetylated HSPD1 in HepG2 cells ± HDAC6 knockdown. h The protein expression levels of HSPD1 and acetylated HSPD1 were detected in HCC cells treated with different concentrations of LukS-PV for 24 h. i Western blot analysis was used to examine the protein expression levels of HDAC6, HSPD1, and acetylated HSPD1 in Huh7 cells ± HDAC6 overexpression treated with or without 1 μM LukS-PV for 24 h. Transwell data (mean ± SD; n = 3) were analyzed by unpaired two-tail t-test. **, p < 0.01; ***, p < 0.001.
To determine whether HSPD1 is a substrate of HDAC6, we performed co-IP assays to confirm the physical association between HSPD1 and HDAC6 (Fig. 5d). Immunofluorescence staining revealed the colocalization of HDAC6 and HSPD1 in the cytoplasm, and LukS-PV treatment led to a significant reduction in the fluorescence intensity of both proteins in the cytoplasmic compartment (Fig. 5e). Additionally, we found that HDAC6 overexpression reduced HSPD1 acetylation while increasing its protein levels, whereas HDAC6 knockdown showed the opposite results (Fig. 5f, g). LukS-PV treatment similarly elevated HSPD1 acetylation and decreased its expression (Fig. 5h), with these effects being reversed by HDAC6 or BCL6 overexpression and C5aR knockdown (Fig. 5i and Supplementary Fig. 4f, g). Moreover, HDAC6 knockdown remarkably impaired the BCL6 overexpression-induced effects on HSPD1 (Supplementary Fig. 4h). Taken together, these data suggest that HSPD1 is a substrate of HDAC6 and that LukS-PV treatment induces HSPD1 acetylation and decreases HSPD1 expression by inhibiting the BCL6/HDAC6 axis.
LukS-PV inhibits EMT in HCC by upregulating HSPD1 acetylation levels and promoting its ubiquitination degradation
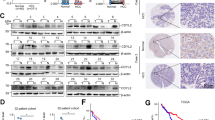
Previous work has indicated that hyperacetylation of HSPD1 leads to its degradation via the ubiquitin-proteasome system31. We used CHX chase experiments and showed that HDAC6 knockdown and LukS-PV treatment both reduced HSPD1 stability, while HDAC6 overexpression stabilized it (Fig. 6a, b and Supplementary Fig. 5a). Furthermore, the inhibitory effect of LukS-PV or HDAC6 knockdown on HSPD1 protein levels was reversed by MG132 (Fig. 6c, d). These data suggest that LukS-PV- and HDAC6-mediated HSPD1 level changes may occur via the ubiquitin-proteasome pathway.
a HepG2 cells ± HDAC6 knockdown were treated with 10 μM CHX for different times, then HSP60 protein expression was analyzed using Western blots. b Huh7 cells were treated with or without 1 μM LukS-PV in the presence of 10 μM CHX for different times, then HSP60 protein expression was analyzed using Western blots. c HepG2 cells ± HDAC6 knockdown treated with or without 20 μM MG132 for 5 h, then HSPD1 protein expression was analyzed using Western blots. d HepG2 and Huh7 cells were treated with 1 μM LukS-PV for 24 h and/or 20 μM MG132 for 5 h, then HSP60 protein expression was analyzed using Western blots. e Huh7 cells were treated with or without 1 μM LukS-PV for 24 h and 20 μM MG132 for 5 h, then the HSPD1 acetylation and ubiquitination levels were detected by IP and Western blot assays. f HepG2 cells ± HDAC6 knockdown were treated with 20 μM MG132 for 5 h, then the HSPD1 acetylation and ubiquitination levels were detected by IP and Western blot assays. g Huh7 cells ± HDAC6 overexpression were treated with 20 μM MG132 for 5 h, then the HSPD1 acetylation and ubiquitination levels were detected by IP and Western blot assays. h Huh7 cells ± HDAC6 overexpression were treated with or without 1 μM LukS-PV for 24 h and 20 μM MG132 for 5 h, then the HSPD1 acetylation and ubiquitination levels were detected by IP and Western blot assays. i Huh7 cells infected with WT HSPD1 or the indicated HSPD1 mutants were treated with 20 μM MG132 for 5 h, then the HSPD1 acetylation and ubiquitination levels were detected by IP and Western blot assays. j Huh7 cells infected with WT HSPD1 or the indicated HSPD1 mutants were treated with or without 1 μM LukS-PV for 24 h and 20 μM MG132 for 5 h, then the HSPD1 acetylation and ubiquitination levels were detected by IP and Western blot assays.
Further experiments showed that both LukS-PV treatment and HDAC6 knockdown could upregulate the acetylation and ubiquitination levels of HSPD1 (Fig. 6e, f), whereas HDAC6 overexpression showed the opposite results (Fig. 6g). The effect of LukS-PV on HSPD1 was attenuated by the overexpression of HDAC6 and BCL6, as well as by C5aR knockdown (Fig. 6h and Supplementary Fig. 5b, c). Additionally, the effect of BCL6 overexpression on HSPD1 was abrogated by HDAC6 knockdown (Supplementary Fig. 5d), thereby validating that LukS-PV induces HSPD1 acetylation and ubiquitination by targeting C5aR to inhibit the BCL6/HDAC6 axis.
Through further acetylomics analysis, we found that LukS-PV treatment predominantly affected two acetylation sites of HSPD1, K236 and K249, both of which are highly conserved among species (Supplementary Fig. 5e). We then mutated these two lysine (K) residues to glutamine (Q) and arginine (R), which mimicked the acetylation and deacetylation levels of the protein. Notably, HDAC6 knockdown failed to raise the acetylation level of 2KR mutants (Supplementary Fig. 5f). The KQ mutants had dramatically increased HSPD1 ubiquitination, while the KR mutants reduced it (Fig. 6i). Furthermore, LukS-PV treatment significantly increased ubiquitination levels of WT HSPD1 but not of 2KQ or 2KR mutants (Fig. 6j). Thus, the above results indicate that LukS-PV promotes HSPD1 ubiquitination via K236/K249 acetylation by downregulating HDAC6.
We next investigated the influence of HSPD1 acetylation on HCC cell EMT, migration, and invasion. Functional assays revealed that deacetylation mutants of HSPD1 enhanced HCC cell EMT, migration and invasion, with 2KR showing the strongest effects (Fig. 7a–c and Supplementary Fig. 6a–c). We also found that the 2KR mutant weakened the inhibitory effect of LukS-PV treatment on these phenotypes more than HSPD1-WT (Fig. 7d–f and Supplementary Fig. 6d–f). In vivo, HSPD1 knockdown reduced the number of lung metastatic nodules (Fig. 7g). Overexpressing HSPD1 promoted HCC metastasis, whereas HSPD1-2KR significantly attenuated LukS-PV’s anti-metastatic effects (Fig. 7h). Consistent with the in vitro findings, immunohistochemical analysis showed that, LukS-PV decreased the expression of BCL6, HDAC6, HSPD1, and N-cadherin while increasing E-cadherin expression, and that mice in the HSPD1-2KR group exhibited stronger resistance to LukS-PV treatment (Supplementary Fig. 6g). Taken together, these results suggest that hypoacetylation of the K236 and K249 residues of HSPD1 is critical for its pro-metastatic role in HCC, and LukS-PV treatment inhibits EMT in HCC by upregulating HSPD1 acetylation levels.
a Western blot analysis was used to determine the protein expression level changes of E-cadherin, N-cadherin, vimentin, HSPD1, and acetylated HSPD1 in Huh7 cells infected with WT HSPD1 or the indicated HSPD1 mutants. b, c Statistical analysis of transwell migration/invasion assays in Huh7 and HepG2 cells infected with WT HSPD1 or the indicated HSPD1 mutants. d Western blot analysis was used to examine the protein expression levels of E-cadherin, N-cadherin, vimentin, HSPD1, and acetylated HSPD1 in Huh7 cells ± HSPD1-WT or HSPD1-2KR treated with or without 1 μM LukS-PV for 24 h. e, f Statistical analysis of transwell migration/invasion assays in Huh7 cells and HepG2 cells ± HSPD1-WT or HSPD1-2KR treated with or without 1 μM LukS-PV for 24 h. g, h Representative images of the gross lung appearance and HE staining of the lung lobes, and statistical analysis of the metastatic nodules in the lung metastasis mouse model from the indicated experimental groups. Scale bar, 2 mm. Transwell (mean ± SD; n = 3) and metastatic lung nodules data (mean ± SD; n = 5) were analyzed by unpaired two-tail t-test or one-way ANOVA. *, p < 0.05; **, p < 0.01; ***, p < 0.001; ****, p < 0.0001.
LukS-PV targets C5aR to inhibit EMT in HCC via the BCL6/HDAC6/HSPD1 axis
From the above findings, we aimed to determine the expression levels of C5aR, BCL6, HDAC6, and HSPD1, as well as the acetylation levels of HSPD1, in 23 matched pairs of fresh primary HCC tissues and their corresponding adjacent non-tumor tissues. C5aR, BCL6, and HSPD1 expression levels were significantly higher in HCC tissues than in the adjacent non-tumor tissues. High HDAC6 expression levels were also found in 78.26% (18/23) of the HCC tissues examined, while HSPD1 acetylation levels were lower in the HCC tissues (Fig. 8a, b). Collectively, our findings demonstrate that in HCC cells, BCL6 binds to the HDAC6 promoter to promote HDAC6 expression, and subsequently HDAC6 interacts with HSPD1 in the cytoplasm and mediates its deacetylation at residues K236 and K249, which in turn prevents HSPD1 from being degraded via the ubiquitin-proteasome pathway. The stable expression of HSPD1 ultimately drives EMT and metastasis in HCC. In contrast, LukS-PV can target C5aR to suppress BCL6 expression, leading to downregulation of HDAC6 and subsequent upregulation of HSPD1 acetylation at residues K236 and K249. Hyperacetylated HSPD1 is degraded through the ubiquitin-proteasome system, ultimately impairing EMT and metastasis in HCC (Fig. 8c).
a C5aR, BCL6, HDAC6, HSPD1, and acetylated HSPD1 levels were determined in 23 pairs of fresh HCC tissues and their corresponding adjacent non-tumor tissues. b Quantification data of (a) were expressed as mean ± SD (n = 23). c Schematic diagram of LukS-PV inhibits HCC metastasis in this research (By Figdraw). Statistical analysis was conducted using paired t-test. **, p < 0.01; ***, p < 0.001; ****, p < 0.0001; ns not significant.
Discussion
The complement C5a receptor, C5aR, is aberrantly expressed in human cancer cells32. Furthermore, C5a has been shown to enhance the motility and invasion of human cancer cells that express C5aR both in vivo and in vitro32. Blocking C5a-C5aR signaling can effectively inhibit tumor growth and metastasis7,8. Our previous studies have consistently demonstrated the antitumor activity of LukS-PV against HCC cells with high C5aR expression. Specifically, we confirmed that LukS-PV inhibits HCC cells proliferation both in vitro and in vivo13,33,34, and suppresses HCC cells migration in vitro by downregulating HDAC6 and targeting the TNNC1/PI3K/AKT axis24,35. In this study, our results further demonstrates that LukS-PV targets C5aR to inhibit EMT and metastasis of HCC cells via the BCL6/HDAC6/HSPD1 axis. Our study elucidates a key epigenetic mechanism by which LukS-PV inhibits EMT in HCC by specifically targeting C5aR. Importantly, we demonstrate that LukS-PV antagonizes C5a-mediated metastatic effects in HCC cells, thereby highlighting its potential as a targeted biotherapeutic agent against C5aR-positive HCC.
Acetylation is an important epigenetic modification that regulates the function and activity of proteins. Recent studies have demonstrated that acetylation is crucially involved in the development and progression of tumors36,37. HDACs regulate histone acetylation homeostasis and ensure gene expression and protein stability, thereby affecting a variety of intracellular biological processes, and HDACs have also been associated with tumor development38. HDAC6 belongs to the HDAC class II family and is a potential therapeutic target in certain cancers, as it is closely associated with oncogenic transformation and tumor metastasis39,40,41. The HDAC6 inhibitors has been used in clinical studies for cancer treatment42,43,44. Our previous study demonstrated that LukS-PV downregulates HDAC6 to inhibit HCC cell migration in vitro24. However, the precise molecular mechanisms remained incompletely characterized. Here, we identified BCL6 as the upstream transcription factor of HDAC6 and HSPD1 as its downstream substrate. We elucidated the mechanistic by which LukS-PV inhibits EMT in HCC through regulating HDAC6, supporting the notion that HDAC6 inhibition as a viable therapeutic strategy for HCC treatment.
The proto-oncogene BCL6 is widely expressed in human cells and is involved in the regulation of humoral immunity and lymphoma survival21,45,46. BCL6 is a key gene contributing to the development of B-cell lymphomas47, and plays an important role in many solid tumors, making it an important therapeutic target. BCL6 has been reported to exhibit dual roles in promoting and repressing transcription23,48,49,50. For example, in HCC, BCL6 interacts with the histone variant H2A.Z and promotes the transcriptional expression of downstream genes, which promotes HCC cells proliferation and metastasis23. However, BCL6 expression levels were found to be significantly reduced in gastric cancer, BCL6 could inhibit gastric cancer proliferation and metastasis by transcriptional repression of frizzled class receptor 7 (FZD7)50. In this study, we identified BCL6 as a key transcription factor that can regulate HDAC6 expression, confirming that BCL6 could promote HCC metastasis by regulating HDAC6 expression. Given that LukS-PV inhibits the BCL6/HDAC6 axis to suppress HCC progression, we propose potential mechanisms for BCL6 downregulation: binding to C5aR to suppress C5a-C5aR-mediated pathways (e.g., NF-κB, PI3K/AKT) regulating BCL6 transcription or protein stability51,52; promoting BCL6 ubiquitination-proteasomal degradation via specific E3 ligases or deubiquitinases; and repressing BCL6 transcription by altering histone acetylation at its promoter through downregulating certain HDACs. Further studies are warranted to validate these hypotheses.
HDAC6 can affects tumor progression by regulating the acetylation levels of several substrates. Deng et al. reported that HDAC6 can destabilize A-kinase anchoring protein 12 (AKAP12) by regulating its deacetylation, thereby promoting colon cancer metastasis53. Wen et al. reported that HDAC6 deacetylates signal transducer and activator of transcription 1 (STAT1) and promotes NF-κB p65 and STAT1 entry into the nucleus to regulate the transcriptional expression of PD-L1, thereby facilitating tumor cell immune evasion54. In this study, we found that HDAC6 could promote HCC cells metastasis by deacetylating HSPD1, which is a highly conserved molecular chaperone protein and important HSP family member31. Notably, HSPD1 is primarily localized to mitochondria and can also be detected in the cytoplasm, cell membrane, and extracellular space, with its extra-mitochondrial localization significantly enhanced in tumor cells55,56,57. This subcellular distribution pattern is crucial for the interaction between HDAC6 and HSPD1 in the cytoplasm. Moreover, HSPD1 is closely associated with cancer progression, as it can inhibit tumor cell apoptosis and promote tumor growth and metastasis28,56,58,59. Specifically, HSPD1 promotes EMT and metastasis through multiple mechanisms across various cancers: it facilitates β-catenin nuclear translocation to drive pro-metastatic transcription28, stabilizes vascular endothelial growth factor A (VEGFA) to enhance angiogenesis60, and activates AKT/mTOR and MAPK signaling to boost cell migration61,62. Several studies have shown that HSPD1 inhibitors and modulators are promising new anti-cancer drugs55,63. However, HSPD1 expression patterns and functions in HCC are poorly understood. Our data identify HDAC6 as a deacetylase of HSPD1 and describe a key post-translational regulatory mechanism for HSPD1, in which acetylation of the evolutionarily conserved K236/K249 residues triggers ubiquitin-dependent proteasomal degradation of HSPD1.
Our study has several limitations. Firstly, the precise mechanism by which LukS-PV downregulates BCL6 requires further investigated. Secondly, we have not identified the key E3 ubiquitin ligase that regulates HSPD1 degradation. Finally, our animal experiments only used a lung metastasis model. Generating a patient-derived tumor xenografts model would further validate the in vivo anti-tumor effects of LukS-PV observed here. We will explore these in future studies.
In conclusion, our findings demonstrate that LukS-PV exerts anti-tumor activity on C5aR-positive HCC cells via the BCL6/HDAC6/HSPD1 axis. We show that C5aR, BCL6, HDAC6 and HSPD1 promote EMT in HCC, suggesting that these molecules could serve as therapeutic targets for HCC. Moreover, our study reveals a key mechanism for regulating EMT in HCC and also lays the foundation for the development of potential targeted therapies against HCC.
Methods
Cell culture
The LX2, HEK293T, HepG2, SNU449, Hep3B and Huh7 cell lines were purchased from Shanghai Cell Bank of Chinese Academy of Sciences (Shanghai, China). Cells were cultured at 37 °C in 5% CO2 with RPMI-1640 (Gibco, Grand Island, NY, USA) or DMEM (Gibco) supplemented with 10% fetal bovine serum (FBS, Lonsera, Ciudad de la Costa, Uruguay) and 1% penicillin/streptomycin (Biosharp, Hefei, China). All cells were cultured in an incubator at 37 °C containing 5% CO2. Cells were identified by short tandem repeat profiling and tested for Mycoplasma contamination every 2 months using the Mycoplasma Detection Kit (Biosharp, #BL1470A). All reagents used in this study are listed in Supplementary Table 3.
Production and purification of recombinant LukS-PV and LukS-PV-GFP
The LukS-PV sequence was amplified from PVL-positive Staphylococcus aureus isolates by PCR. Recombinant LukS-PV was generated as described previously64. Purification of recombinant LukS-PV was performed using the His-Bind Purification Kit (Millipore, #70239, Bedford, MA, USA) following the manufacturer’s protocol. To generate the recombinant LukS-PV-GFP, the LukS-PV sequence was digested with BamHI (Promega, #R6021) and XhoI (Promega, #R6161), then ligated into the pMD-18T vector (Takara, Tokyo, Japan) to produce pMD-18T-LukS-PV plasmids. After double enzyme digestion, the plasmids were connected to the pet28a-N6H-GFP vector (provided by Prof. Jian-Ye Zang, University of Science and Technology of China). DH5α bacteria were transformed with this construct, then the clones were verified by DNA sequencing. The identified plasmids were converted into BL21 bacteria. The method for purifying LukS-PV-GFP was the same as used for purifying recombinant LukS-PV. All the primers used are listed in Supplementary Table 4.
Cell transfection and infection
siRNA-C5aR and C5aR overexpression plasmid were purchased from General Biotechnology (Chuzhou, China). HCC cells were transfected with siRNA or plasmid using Lipofectamine 2000 (Invitrogen, #11668019, Carlsbad, CA, USA) following the manufacturer’s protocol. Transfection efficiency was assessed by Western blot analysis (see below). To generate lentiviral expression constructs, HDAC6 cDNA was amplified and cloned into the pCDH-CMV-MCS-EF1-Puro vector. To generate viruses for stable expression or knockdown of target genes, HEK293T cells were co-transfected with pCDH-HDAC6 or pLKO.1-shRNA and lentivirus package plasmids (psPAX2 and pMD2.G), respectively. The cell supernatant containing the viral particles was collected, filtered through a 0.45-mm filter, and used to infect HepG2 or Huh7 cells. Stable pools of infected cells were selected with 2.5 μg/mL puromycin. The established HCC cells were maintained in a growth medium containing 1 μg/mL puromycin. The lentivirus for BCL6 overexpression was purchased from GenePharma Biotechnology (Shanghai, China). All HSPD1 mutants and wild-type HSPD1 constructs were purchased from Tsingke Biotech (Shanghai, China). The lentivirus infection and stable HCC cells screening methods were in accordance with the manufacturer’s protocol. All the oligonucleotides used are listed in Supplementary Table 4.
Western blot analysis
After the indicated treatments, cells were lysed in RIPA buffer (Beyotime Biotechnology, Shanghai, China) containing a protease inhibitor cocktail (Beyotime) on ice for 30 min. The total protein concentration of each sample was quantified using a BCA protein assay Kit (Beyotime, #P0012). The protein samples were boiled in 5× sodium dodecylsulfate (SDS) loading buffer for 15 min. The samples were then separated using 7.5% or 10% SDS-PAGE electrophoresis, transferred to nitrocellulose membranes (Millipore, #HATF00010). The membranes were blocked in 5% skim milk and incubated with the primary antibody overnight at 4 °C. Then the membranes were washed in TBST and incubated with HRP-conjugated secondary antibodies for 1.5 h at room temperature. The protein bands were detected with an enhanced chemiluminescence kit (Epizyme, #SQ201, Shanghai, China). The antibodies used are listed in Supplementary Table 5.
RNA extraction and quantitative real-time PCR
Total RNA was extracted from HCC cells using TRIzol reagent (Invitrogen, USA). cDNA was synthesized with the HiScript II Q RT SuperMix (Vazyme, #R223-01, Nanjing, China), and qRT-PCR was performed using SYBR Green Master Mix (Vazyme, #Q711-02), all in accordance with the respective manufacturers’ protocols. The relative mRNA expression levels of target genes were normalized to GAPDH using the 2−ΔΔCt method. All experiments were conducted in triplicate. The qRT-PCR primers used are listed in Supplementary Table 4.
Transwell migration and invasion assay
For migration assays, 1 × 105 cells in 200 μL serum-free medium were seeded into the upper chamber (Corning, #353097, NY, USA), while 700 μL medium containing 10% FBS was added to the lower chamber. For invasion assays, the chamber inserts were pretreated with Matrigel (Corning) diluted in pre-cooled serum-free medium and incubated at 37 °C for at least 4–5 h for gelling. Subsequently, 1 × 105 cells in 200 μL serum-free medium were seeded into the Matrigel-pretreated upper chamber, and 700 μL 10% FBS-containing medium was added to the lower chamber. After a 24 h incubation, any cells in the lower chamber were fixed in 4% paraformaldehyde (BIOMIKY, Shanghai, China) and stained with 0.1% crystal violet solution (Biosharp).
Immunofluorescence (IF) staining
Cells were pre-plated into a glass bottom cell culture dish (Biosharp, #BS-20-GJM). The cells were fixed in 4% paraformaldehyde for 15 min, then permeabilized with 0.2% Triton X-100 (Sigma-Aldrich, #T9284, St. Louis, MO, USA) for 5 min at room temperature. Next, the cells were blocked with 3% BSA (BioFroxx, Einhausen, Germany) at room temperature for 1 h, then incubated with primary antibodies in blocking buffer overnight at 4 °C. Next day, the cells were incubated with or without secondary antibodies (Alexa Fluor-594) at room temperature for 1 h, depending on the primary antibodies being directly conjugated. Cell nuclei were stained with DAPI (Biosharp). Images were visualized with a Zeiss LSM800 confocal microscope and analyzed using software.
RNA sequencing (RNA-seq)
Total RNA was isolated from HepG2 cells treated with PBS or 1 μM LukS-PV for 24 h using the RNeasy Mini Kit (QIAGEN, #74106, Germany) following the manufacturer’s protocol. Paired-end libraries were synthesized with the TruSeq RNA sample preparation kit (Illumina, #RS-122-2001, USA). In brief, the poly(A)-containing mRNA molecules were purified using oligo (dT) attached to magnetic beads. Library construction and sequencing were performed at Shanghai Sinomics Corporation (Shanghai, China).
Mass spectrometry-based label-free quantitative acetylated proteomics
Label-free quantitative acetylated proteomics was conducted by PTM Biolabs (Hangzhou, China). Lysis buffer (8 M urea, 1% protease inhibitor, 3 μM TSA, 50 mM NAM) was added to the samples, the lysates were sonicated and centrifuged, then the supernatant were collected. For digestion, the protein solution was initially treated with 5 mM dithiothreitol at 56 °C for 30 min and alkylated with 11 mM iodoacetamide at room temperature in darkness for 15 min. The protein sample was then diluted by adding 100 mM TEAB to urea concentration less than 2 M. Finally, trypsin was added at a 1:50 trypsin-to-protein mass ratio for the first digestion overnight and 1:100 trypsin-to-protein mass ratio for a second 4 h-digestion. To enrich the acetylated modified peptides, the tryptic peptides were dissolved in IP buffer solution (100 mM NaCl, 1 mM EDTA, 50 mM Tris-HCl, 0.5% NP-40, pH 8.0), and the supernatant was transferred to a pre-washed acetylation resin (PTM Bio, #PTM104) and incubate overnight at 4 °C with gentle shaking. The resin was then washed four times with IP buffer solution and twice with deionized water. Finally, the resin-bound peptides were eluted a total of three times with 0.1% trifluoroacetic acid eluent, and the eluent was collected and lyophilized under vacuum. For liquid chromatography-tandem mass spectrometry (LC-MS/MS) analysis, the resulting peptides were desalted using C18 ZipTips (Millipore) according to the manufacturer’s instructions.
Chromatin immunoprecipitation (ChIP) assay
ChIP DNA was purified using the SimpleChIP enzymatic chromatin IP kit (Cell Signaling Technology, #9003, Danvers, MA, USA) following the manufacturer’s protocol. HepG2 cells were cross-linked with 1% formaldehyde, nuclei were prepared immediately, chromatin was digested with micrococcal nuclease. The supernatant was collected by sonication and centrifugation, and incubated overnight at 4 °C with an anti-BCL6 antibody or relevant non-specific IgG, Protein G magnetic beads were then added and incubated for 2 h. ChIP DNA was purified, then quantified by PCR and qPCR. The gene-specific primer sequences used for ChIP-PCR and ChIP-qPCR are listed in Supplementary Table 4.
Immunopurification (IP) assay and mass spectrometry
Cells were lysed in IP lysis buffer containing protease inhibitor cocktail. Lysates were incubated with antibodies overnight at 4 °C. Next day, Protein A/G PLUS-Agarose beads (Santa Cruz, CA, USA) were added to the lysates and incubated for 4 h. The beads were washed three times with lysis buffer, mixed with an equal volume of 2× SDS loading buffer, and boiled at 100 °C for 5 min, then analyzed by Western blot analysis. Huh7 cells stably overexpressing HDAC6 were lysed for IP and mass spectrometry analysis to identify proteins interacting with HDAC6, as previously described24.
Acetylation assay
For HSPD1 acetylation assays, the treated cells were lysed in IP lysis buffer containing a protease inhibitor cocktail. Lysates were incubated overnight at 4 °C with anti-HSPD1 antibody or anti-acetylated-lysine antibody. Next day, Protein A/G PLUS-agarose beads were added to the lysates and incubated for 4 h. The beads were washed three times with lysis buffer, mixed with an equal volume of 2× SDS loading buffer, and boiled at 100 °C for 5 min, then analyzed by Western blot analysis.
Cycloheximide (CHX) chase and ubiquitination assay
Cells were incubated with 10 μM CHX (Beyotime) and collected at different time points. HSPD1 protein expression was determined by Western blot analysis. For HSPD1 ubiquitination assays, the treated cells were incubated with 20 μM MG132 (Beyotime) for 5 h before harvesting. The cells were then lysed in IP lysis buffer containing a protease inhibitor cocktail. Lysates were incubated with an anti-HSPD1 antibody overnight at 4 °C. Next day, Protein A/G PLUS-agarose beads were added to the lysates and incubated for 4 h. The beads were washed three times with lysis buffer, mixed with an equal volume of 2× SDS loading buffer, and boiled at 100 °C for 5 min, then analyzed by Western blot analysis.
In vivo metastasis assay
All animal experiments were approved by the Ethics Committee of the University of Science and Technology of China (2024-N(A)-173). We have complied with all relevant ethical regulations for animal use. BALB/c nude mice (male, 4 weeks old) were purchased from Beijing Vital River Laboratory Animal Technology Co., Ltd (Beijing, China). All mice were housed 5 per cage, and all experiments were performed in the specific pathogen-free (SPF) animal facility of the First Affiliated Hospital of University of Science and Technology of China. Cages were randomly positioned to minimize environmental confounders, and experimental procedures were conducted in random sequence to reduce operational bias. For the in vivo metastasis assay, mice were randomly divided into treatment groups (n = 5 per group). The sample size was determined based on our laboratory’s prior experience with HCC in vivo models, where it consistently detects biologically significant effects. Huh7 cells (8 × 106 cells/200 μL) were injected into BALB/c nude mice (n = 5) via the tail vein. Weekly continuous tail vein injections of LukS-PV (300 μg/kg body weight per mouse) or PBS were initiated two weeks after cell inoculation. Eight weeks after the initial cell injection, the mice were euthanized using cervical dislocation. Lungs were removed, fixed in 4% paraformaldehyde, embedded in paraffin, and sectioned for subsequent hematoxylin and eosin (HE) staining and immunohistochemistry (IHC) analysis.
For HE staining, sections were directly stained with hematoxylin and eosin, and metastatic foci were counted. For IHC: after deparaffinization, endogenous peroxidase activity was blocked with 3% hydrogen peroxide for 10 min, followed by antigen retrieval via microwave heating in 10 mmol/L sodium citrate buffer (pH 6.0). Sections were blocked with 0.1% Triton X-100-containing 5% BSA buffer at room temperature, then incubated with primary antibodies at 4 °C overnight. After PBS washing, sections were incubated with secondary antibodies for 30 min at room temperature, developed with DAB substrate, and counterstained with hematoxylin.
To evaluate the regulatory effect of LukS-PV on the in vivo metastasis of HCC cells, a lung metastasis model was established by tail vein injection of HCC cells into immunodeficient nude mice. This approach avoids immune rejection of human-derived Huh7 cells and has been widely used in studies to assess the in vivo metastatic capacity of cancer cells. Additionally, an in vivo experimental system based on multiple stably infected cell lines was constructed, which enables the effective evaluation of the specific roles of BCL6, HDAC6, and HSPD1 in HCC cell metastasis in vivo. Notably, this model directly mimics the metastatic process of human HCC cells in vivo, significantly enhancing the relevance of the in vivo metastasis assay to human biology. Therefore, the selection of this animal model is highly consistent with the scientific objectives of this study. The results obtained from this model not only provide reliable in vivo experimental evidence for BCL6, HDAC6, and HSPD1 as potential therapeutic targets for human HCC but also offer critical preliminary data supporting the development of LukS-PV as a candidate anti-HCC metastasis drug. Additionally, the experimental system established and the molecular mechanisms validated in this study lay a solid foundation for subsequent investigations into the regulatory roles of the BCL6/HDAC6/HSPD1 axis in the complex tumor microenvironment, as well as for further evaluations of the clinical translational potential of LukS-PV.
For the in vivo metastasis assay, mice were enrolled if no immediate adverse reactions were observed at the tail vein injection site. Animals with injection failure or death within 7 days post-injection were excluded, with no additional exclusions made in this study. A total of 135 mice were used, and each individual mouse was considered an independent experimental unit. The in vivo metastasis assay included one primary outcome measure: the number of metastatic nodules in the lungs, quantified by microscopic examination of tissue sections. This indicator directly reflects the severity of lung metastasis of HCC cells, with a higher number of nodules indicating a more significant metastatic burden.
To avoid subjective bias affecting the judgment of outcome indicators during data analysis, four investigators participated in the animal experiments with clearly defined responsibilities: Investigators Pengsheng Ding and Bing Lu jointly took charge of the core operational procedures. They strictly followed the preset protocol to administer Huh7 cells, PBS, or LukS-PV to the respective groups, and co-conducted mouse euthanasia, dissection, and lung tissue collection at the end of the experiment. Given the need for precise matching of treatments to groups during administration and clear association of lung tissues with their corresponding groups post-dissection, both researchers had full access to the group allocation information. Prior to data analysis, all lung tissue samples were anonymously coded by Investigator Xuexue Xu. Subsequently, Investigator Shanshan Zhang counted lung metastatic nodules and conducted data analysis based on the coded samples, remaining unaware of the corresponding group assignments throughout the process. Only after all statistical analyses were completed did Xuexue Xu decrypt the codes to link samples to their actual groups, thereby effectively eliminating potential subjective bias during analysis.
Human tissue analysis
Patient selection strictly followed predefined inclusion and exclusion criteria: the inclusion criterion was patients with pathologically confirmed primary HCC, while the exclusion criteria comprised incomplete clinical data, postoperative complications, missing tissue samples, prior receipt of interventional therapy, chemotherapy, or radiotherapy, and positive serological results for HBV, tuberculosis, syphilis, or HIV (or patients in the acute phase of any infectious disease). Primary HCC tissues and the corresponding adjacent nontumor tissues were obtained from 23 patients at the First Affiliated Hospital of University of Science and Technology of China from March 2023 to April 2024. None of the patients received neoadjuvant therapy before partial hepatectomy. Western blot analysis was used to evaluate relevant protein expression levels in the HCC tissues. This study was approved by the Ethics Committee of the First Affiliated Hospital of University of Science and Technology of China (2024KY-401). All ethical regulations relevant to human research participants were followed. This study adheres to the Declaration of Helsinki.
Statistics and reproducibility
Data are presented as mean ± SD. GraphPad Prism software 9.5 was used to conduct data analyses and calculate p-values. Data from two groups were analyzed using Student’s t test, while data from multiple groups were analyzed using one-way ANOVA. Before parametric analyses, the homogeneity of variances was evaluated. As the homogeneity assumption was not met, an appropriate ANOVA approach accommodating unequal variances was used for group comparisons. Differences were considered statistically significant when p < 0.05. All experiments were independently repeated at least three times to ensure consistency of results. To uphold the reproducibility of experimental results, all assays were designed with sufficient sample sizes to ensure robust statistical power. The number of replicates for each experiment is indicated in the relevant figure legends and corresponding sections of the manuscript: for in vitro studies, replicates refer to independent experiments carried out on distinct days using separate cell cultures, while in vivo replicates are determined by the number of animals assigned to each experimental group.
Ethics approval and consent to participate
All animal care and experimental procedures were approved by the Ethics Committee of the University of Science and Technology of China (2024-N(A)-173). Authorization for the use of primary HCC tissues and their corresponding adjacent nontumor tissues in this study was obtained from the Ethics Committee of the First Affiliated Hospital of the University of Science and Technology of China (2024KY-401). Informed consent was obtained from all human participants. All procedures involving human tissues were conducted in accordance with institutional and national ethical standards.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data supporting the findings of the paper are present in the main text and the Supplementary information. Source data can be found in Supplementary Data 2. Uncropped blot or gel images are available in Supplementary Fig. 7, attached as Supporting Information to this manuscript. The acetylome data have been deposited in the ProteomeXchange Consortium via the PRIDE65 partner repository with the dataset identifier PXD057287. The RNA-seq data are available in the NCBI Sequence Read Archive (SRA) under BioProject PRJNA1404140. All other data are available from the corresponding author on reasonable request.
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Giannelli, G., Koudelkova, P., Dituri, F. & Mikulits, W. Role of epithelial to mesenchymal transition in hepatocellular carcinoma. J. Hepatol. 65, 798–808 (2016).
Shibue, T. & Weinberg, R. A. EMT, CSCs, and drug resistance: the mechanistic link and clinical implications. Nat. Rev. Clin. Oncol. 14, 611–629 (2017).
Chen, J. et al. Complement C5a/C5aR pathway potentiates the pathogenesis of gastric cancer by down-regulating p21 expression. Cancer Lett. 412, 30–36 (2018).
Hu, W. H. et al. C5a receptor enhances hepatocellular carcinoma cell invasiveness via activating ERK1/2-mediated epithelial-mesenchymal transition. Exp. Mol. Pathol. 100, 101–108 (2016).
Piao, C. et al. Complement 5a stimulates macrophage polarization and contributes to tumor metastases of colon cancer. Exp. Cell Res. 366, 127–138 (2018).
Markiewski, M. M. et al. Modulation of the antitumor immune response by complement. Nat. Immunol. 9, 1225–1235 (2008).
Luan, X. et al. Blockade of C5a receptor unleashes tumor-associated macrophage antitumor response and enhances CXCL9-dependent CD8(+) T cell activity. Mol. Ther. J. Am. Soc. Gene Ther. 32, 469–489 (2024).
Spaan, A. N. et al. The staphylococcal toxin Panton-Valentine leukocidin targets human C5a receptors. Cell Host Microbe 13, 584–594 (2013).
Jayasinghe, L. & Bayley, H. The leukocidin pore: evidence for an octamer with four LukF subunits and four LukS subunits alternating around a central axis. Protein Sci. 14, 2550–2561 (2005).
Shan, W. et al. LukS-PV, a component of Panton-Valentine leukocidin, exerts potent activity against acute myeloid leukemia in vitro and in vivo. Int. J. Biochem. Cell Biol. 61, 20–28 (2015).
Sun, X. X. et al. LukS-PV-regulated microRNA-125a-3p promotes THP-1 macrophages differentiation and apoptosis by down-regulating NF1 and Bcl-2. Cell. Physiol. Biochem. 44, 1093–1105 (2017).
Shi, L. et al. Toxin protein LukS-PV targeting complement receptor C5aR1 inhibits cell proliferation in hepatocellular carcinoma via the HDAC7-Wnt/β-catenin axis. J. Biol. Chem. 301, 108148 (2025).
Raney, B. J. et al. The UCSC genome browser database: 2024 update. Nucleic Acids Res. 52, D1082–d1088 (2024).
Kolmykov, S. et al. GTRD: an integrated view of transcription regulation. Nucleic Acids Res. 49, D104–d111 (2021).
Lonsdale, J. et al. The genotype-tissue expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Weinstein, J. N. et al. The Cancer Genome Atlas pan-cancer analysis project. Nat. Genet. 45, 1113–1120 (2013).
Rauluseviciute, I. et al. JASPAR 2024: 20th anniversary of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 52, D174–d182 (2024).
Tang, D. et al. SRplot: a free online platform for data visualization and graphing. PLoS ONE 18, e0294236 (2023).
Walker, S. R. et al. The transcriptional modulator BCL6 as a molecular target for breast cancer therapy. Oncogene 34, 1073–1082 (2015).
Cardenas, M. G. et al. The expanding role of the BCL6 oncoprotein as a cancer therapeutic target. Clin. Cancer Res. 23, 885–893 (2017).
Guo, J. et al. BCL6 confers KRAS-mutant non-small-cell lung cancer resistance to BET inhibitors. J. Clin. Investig. 131, e133090 (2021).
Yuan, Y., Cao, W., Zhou, H., Qian, H. & Wang, H. H2A.Z acetylation by lincZNF337-AS1 via KAT5 implicated in the transcriptional misregulation in cancer signaling pathway in hepatocellular carcinoma. Cell Death Dis. 12, 609 (2021).
Xu, X., Ding, P., Shi, L., Wu, G. & Ma, X. LukS-PV inhibits hepatocellular carcinoma cells migration by downregulating HDAC6 expression. BMC Cancer 22, 630 (2022).
Oughtred, R. et al. The BioGRID database: a comprehensive biomedical resource of curated protein, genetic, and chemical interactions. Protein Sci. 30, 187–200 (2021).
Rodchenkov, I. et al. Pathway commons 2019 update: integration, analysis and exploration of pathway data. Nucleic Acids Res. 48, D489–d497 (2020).
Forouzanfar, F., Barreto, G., Majeed, M. & Sahebkar, A. Modulatory effects of curcumin on heat shock proteins in cancer: a promising therapeutic approach. Biofactors 45, 631–640 (2019).
Tsai, Y. P. et al. Interaction between HSP60 and beta-catenin promotes metastasis. Carcinogenesis 30, 1049–1057 (2009).
Chandrashekar, D. S. et al. UALCAN: an update to the integrated cancer data analysis platform. Neoplasia 25, 18–27 (2022).
Tang, Z. et al. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 45, W98–w102 (2017).
Caruso Bavisotto, C. et al. Hsp60 post-translational modifications: functional and pathological consequences. Front. Mol. Biosci. 7, 95 (2020).
Nitta, H. et al. Enhancement of human cancer cell motility and invasiveness by anaphylatoxin C5a via aberrantly expressed C5a receptor (CD88). Clin. Cancer Res. 19, 2004–2013 (2013).
Wang, Z. et al. LukS-PV inhibits hepatocellular carcinoma progression by downregulating HDAC2 expression. Mol. Ther. Oncolytics 17, 547–561 (2020).
Nie, Z. et al. LukS-PV inhibits the proliferation of hepatocellular carcinoma cells by maintaining FOXO3 stability via the PI3K/AKT signaling pathway. Cell. Signal. 95, 110357 (2022).
Ma, F. et al. LukS-PV inhibits hepatocellular carcinoma cells migration via the TNNC1/PI3K/AKT axis. OncoTargets Ther. 13, 10221–10230 (2020).
Sun, L., Zhang, H. & Gao, P. Metabolic reprogramming and epigenetic modifications on the path to cancer. Protein Cell 13, 877–919 (2022).
Guertin, D. A. & Wellen, K. E. Acetyl-CoA metabolism in cancer. Nat. Rev. Cancer 23, 156–172 (2023).
Manzotti, G., Ciarrocchi, A. & Sancisi, V. Inhibition of BET proteins and histone deacetylase (HDACs): crossing roads in cancer therapy. Cancers 11, 304 (2019).
Lee, Y. S. et al. The cytoplasmic deacetylase HDAC6 is required for efficient oncogenic tumorigenesis. Cancer Res. 68, 7561–7569 (2008).
Aldana-Masangkay, G. I. & Sakamoto, K. M. The role of HDAC6 in cancer. J. Biomed. Biotechnol. 2011, 875824 (2011).
Yang, P. H., Zhang, L., Zhang, Y. J., Zhang, J. & Xu, W. F. HDAC6: physiological function and its selective inhibitors for cancer treatment. Drug Discov. Ther. 7, 233–242 (2013).
Yussuf Khamis, M. et al. Overcome the tumor immunotherapy resistance by combination of the HDAC6 inhibitors with antitumor immunomodulatory agents. Bioorg. Chem. 109, 104754 (2021).
Biersack, B., Nitzsche, B. & Hopfner, M. Immunomodulatory properties of HDAC6 inhibitors in cancer diseases: new chances for sophisticated drug design and treatment optimization. Semin. Cell Dev. Biol. 154, 286–294 (2024).
Gu, Z. et al. Advances in dual-targeting inhibitors of HDAC6 for cancer treatment. Eur. J. Med. Chem. 275, 116571 (2024).
Cattoretti, G. et al. BCL-6 protein is expressed in germinal-center B cells. Blood 86, 45–53 (1995).
Bunting, K. L. & Melnick, A. M. New effector functions and regulatory mechanisms of BCL6 in normal and malignant lymphocytes. Curr. Opin. Immunol. 25, 339–346 (2013).
Basso, K. & Dalla-Favera, R. BCL6: master regulator of the germinal center reaction and key oncogene in B cell lymphomagenesis. Adv. Immunol. 105, 193–210 (2010).
Dent, A. L., Vasanwala, F. H. & Toney, L. M. Regulation of gene expression by the proto-oncogene BCL-6. Crit. Rev. Oncol. Hematol. 41, 1–9 (2002).
Zeng, G., Xu, Y., Li, Z. & Deng, G. BCL6 promotes transcription of GPR61 to suppress IL-1β-induced osteoarthritis progression in C28/I2 cells. Microbiol. Immunol. 69, 148–156 (2025).
Guo, S. et al. The malignancy suppression and ferroptosis facilitation of BCL6 in gastric cancer mediated by FZD7 repression are strengthened by RNF180/RhoC pathway. Cell Biosci. 13, 73 (2023).
Fang, J., Wang, J., Zhao, X., Yang, Y. & Xiao, Y. KLHDC8A knockdown in normal ovarian epithelial cells promoted the polarization of pro-tumoral macrophages via the C5a/C5aR/p65 NFκB signaling pathway. Cell. Immunol. 409-410, 104913 (2025).
Ajona, D., Ortiz-Espinosa, S. & Pio, R. Complement anaphylatoxins C3a and C5a: emerging roles in cancer progression and treatment. Semin. Cell Dev. Biol. 85, 153–163 (2019).
Deng, Y. et al. HDAC6-dependent deacetylation of AKAP12 dictates its ubiquitination and promotes colon cancer metastasis. Cancer Lett. 549, 215911 (2022).
Wen, Y. et al. HDAC6 inhibitor ACY-1215 enhances STAT1 acetylation to block PD-L1 for colorectal cancer immunotherapy. Cancer Immunol. Immunother. 73, 7 (2024).
Cappello, F. et al. Hsp60 chaperonopathies and chaperonotherapy: targets and agents. Expert Opin. Ther. Targets 18, 185–208 (2014).
Cappello, F., Conway de Macario, E., Marasa, L., Zummo, G. & Macario, A. J. Hsp60 expression, new locations, functions and perspectives for cancer diagnosis and therapy. Cancer Biol. Ther. 7, 801–809 (2008).
O’Malley, J., Kumar, R., Inigo, J., Yadava, N. & Chandra, D. Mitochondrial stress response and cancer. Trends Cancer 6, 688–701 (2020).
Ghosh, J. C., Dohi, T., Kang, B. H. & Altieri, D. C. Hsp60 regulation of tumor cell apoptosis. J. Biol. Chem. 283, 5188–5194 (2008).
Ghosh, J. C., Siegelin, M. D., Dohi, T. & Altieri, D. C. Heat shock protein 60 regulation of the mitochondrial permeability transition pore in tumor cells. Cancer Res. 70, 8988–8993 (2010).
Zheng, D. et al. LncRNA LINC01503 promotes angiogenesis in colorectal cancer by regulating VEGFA expression via miR-342-3p and HSP60 binding. J. Biomed. Res. 39, 286–304 (2024).
Mandal, J. P. et al. PKCδ mediates mitochondrial ROS generation and oxidation of HSP60 to relieve RKIP inhibition on MAPK pathway for HCC progression. Free Radic. Biol. Med. 163, 69–87 (2021).
Zhang, Y. et al. HSPD1 supports osteosarcoma progression through stabilizing ATP5A1 and thus activation of AKT/mTOR signaling. Int. J. Biol. Sci. 20, 5162–5190 (2024).
Meng, Q., Li, B. X. & Xiao, X. Toward developing chemical modulators of Hsp60 as potential therapeutics. Front. Mol. Biosci. 5, 35 (2018).
Ma, X., Chang, W., Zhang, C., Zhou, X. & Yu, F. Staphylococcal Panton-Valentine leukocidin induces pro-inflammatory cytokine production and nuclear factor-kappa B activation in neutrophils. PLoS ONE 7, e34970 (2012).
Perez-Riverol, Y. et al. The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res. 50, D543–d552 (2022).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (81972001 to X.L.M., 82203436 to S.S.Z., 82102478 to X.F.W.), Young Research Fund of Wannan Medical College (WK2022F18 to Y.Y.W.), the Natural Science Foundation of Anhui Province (2208085QH250 to S.S.Z.) and the University Natural Science Research Project of Anhui Province (2023AH053403 to Z.C.N.). The Schematic diagram was drawn by Figdraw. We thank J. Iacona, Ph.D., from Liwen Bianji (Edanz) (www.liwenbianji.cn) for editing the English text of a draft of this manuscript.
Author information
Authors and Affiliations
Contributions
Conceptualization, P.S.D., X.L.M., and S.S.Z.; methodology, P.S.D. and L.S.; validation, P.S.D., L.S., X.X.X., G.L., Y.Y.W., Z.C.N., X.F.W., W.J.C., and Y.Y.D.; formal analysis, P.S.D., X.X.X., B.L., and S.S.Z.; investigation, P.S.D., X.X.X., and B.L.; resources, X.L.M.; data curation, P.S.D., L.S., X.X.X., G.L., and S.S.Z.; writing–original draft, P.S.D.; writing– review and editing, P.S.D., L.S., X.L.M., and S.S.Z.; visualization, P.S.D., G.L., and S.S.Z.; supervision, W.J.C., Y.Y.D., X.L.M., and S.S.Z.; project administration, X.L.M. and S.S.Z.; funding acquisition, Y.Y.W., X.F.W., Z.C.N., X.L.M., and S.S.Z. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Joo-Yong Lee and Linchong Sun for their contribution to the peer review of this work. Primary handling editors: Katharine Irvine and Mengtan Xing.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ding, P., Shi, L., Xu, X. et al. LukS-PV targeting C5aR inhibits EMT in hepatocellular carcinoma via the BCL6/HDAC6/HSPD1 axis. Commun Biol 9, 314 (2026). https://doi.org/10.1038/s42003-026-09640-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-026-09640-9