Abstract
The enteric nervous system (ENS) is involved in many gastrointestinal (GI) disorders and our understanding of how gut morphology is disrupted remains limited due to a lack of tools to investigate tissues at the organ scale. Here we present enGLOW (enteric network Gastrointestinal Lightsheet Optical Workflow), a workflow customized for high spatial-resolution investigation of the ENS in gastrointestinal samples. We demonstrate how enGLOW can extract quantitative data in cubic centimeters of intact tissue. In a single dataset, we quantify intestinal wall metrics in autofluorescence and labeled ENS-associated cells in centimeter-long segments of tissue using three-dimensional (3D) segmentation. With virtual tissue flattening, we separate neuronal plexuses in mouse and human samples, and observe variations in mucosal morphology and labeled signal distribution in genetically modified animals. As an optimized ready-to-use workflow, enGLOW expands the gut research toolbox to enable understanding of enteric network morphology at scales that encompass functional networks.
Similar content being viewed by others
Introduction
The enteric nervous system is a network of up to 600 million neurons responsible for sensing external inputs from the gut environment and the central and peripheral nervous systems1 to regulate motility2, nutrient absorption3, appetite4, immune activation5, hormone signaling6 and more. Enteric neurons7, together with other cells like enteroendocrine cells2, immune cells8, glia9,10 and interstitial cells of Cajal11, create a network whose morphology and function are spread over 8 m12 of human digestive system and whose synchronization is now recognized as a critical contributor to gastrointestinal13,14, metabolic15, neurodegenerative16,17,18,19,20 and neuropsychiatric21,22 disorders.
To understand the pathophysiology of gut diseases, microscopic imaging of histological preparations of excised samples is most common, with the only whole-gut approach consisting of the Swiss-rolling method23 to observe intestinal anatomy in centimeters of tissue. Whole-mount sample preparation can enable volumetric imaging of millimeter-scale tissues, having led to evidence linking ENS network morphology with gastrointestinal functions such as motility in animals24,25, as well as physiological responses to environmental factors such as the circadian rhythm26 and the gut microbiome27. Therapeutic strategies have also been validated using microscopic assessments of ENS morphology, spanning neuronal cell therapy in a model of enteric aganglionosis28 and myenteric plexus regeneration in modeled Chagas disease29. New approaches, such as full-field optical coherence tomography30, are advancing the field towards the exploration of ENS morphology in full-thickness mouse gut samples, but the thickness of human tissue remains a limitation for visualizing the tissue as a whole. Advances in 3D microscopy and tissue clearing have made ex vivo imaging of large intact biological samples, for example, the whole human eye31, possible. However, whole-mount preparations, relying on stretched and pinned samples to assess tissue morphology25,28,32,33 or preparation of physically separated layers24,34,35 are still common in pre-clinical and clinical gastrointestinal research with a focus on the myenteric plexus of the ENS, mostly known for coordinating motility functions, as opposed to the submucosal plexus involved in secretion and blood flow control36. In cleared human tissue, 3D confocal microscopy was applied to 1 cm × 1 cm full-thickness stretched colon resections32 and led to valuable labeling validations, in-depth characterizations of individual ganglia and demonstrated the consequence of Hirschsprung’s disease on ENS morphology. Light-sheet imaging of the cleared inflamed mouse gut highlighted its convenience and potential for having a supportive function in translational research due to its capability to investigate large intact organ segments37, but lacked the critical 3D quantitative data that can be extracted. It is also common for image processing and quantitative analyses to rely on manual segmentation32,33 thus hindering high-throughput data analyses otherwise possible, with one exception from Cairns et al.35 where 3D automated analysis to understand network connectivity and orientation was showcased.
The growing interest in understanding ENS morphology using microscopy has led to advances in image processing, segmentation and quantification methods. Software such as FIJI38 and Imaris (Bitplane) are established tools that can incorporate AI-supported segmentation methods, such as the Trainable Weka Segmentation in ImageJ35. Specific tools for the ENS, including COUNTEN39 and Gut Analysis Toolbox40, make multi-label automated pipelines available for imaged whole-mount gut samples. The advent of light-sheet fluorescence microscopy (LSFM) presents new opportunities for high-resolution volumetric characterization of large intact tissue samples in 3D. In mouse tissues, the importance of large-scale tissue clearing combined with LSFM was highlighted in a comparative study of immunofluorescence, histology and ultrastructure41. Furthermore, this approach’s value for studying the spatial distribution of sparse signals was highlighted in a model of gastrointestinal inflammation to characterize T-cell trafficking37. In non-human primate and human samples, this technique enabled the discovery of norepinephrine’s protective effect against inflammation42.
In spite of these advances, a single method combining detailed tissue processing with gut-specific labeling and imaging to enable 3D observations of intact tissues at large scales with quantitative assessments has not yet been described. To address this need, we adapted techniques well-established in other fields, including neuroscience43 and developmental biology44 (for e.g., iDISCO45), to propose the enteric network Gastrointestinal Lightsheet Optical Workflow (enGLOW). enGLOW is a gut-focused sample preparation and imaging workflow for large-scale quantification of enteric morphology, optimized here for the ENS but applicable to other structures of interest in the gut, like immune cell populations or vascular networks. We implement image processing methods taking advantage of the layered anatomy of the gut to virtually flatten tissue to reveal cellular patterns across the layers of the gut wall within 3D datasets. We demonstrate enGLOW’s diverse applicability in tissues spanning the whole gastrointestinal tract of mice and full-thickness human colon resections. This workflow uses tissue autofluorescence as an informative channel and enables the characterization of neuron, glia and interstitial cells of Cajal spatial distribution to extract quantitative information of changes in ENS morphology in the context of disease.
Results
enGLOW: enteric network Gastrointestinal Lightsheet Optical Workflow
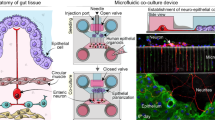
Below, we showcase several kinds of imaging data enabled by enGLOW. The enGLOW workflow itself is schematically illustrated in (Fig. 1a), and presented in detail in the “Methods” section. In addition, diagrams of the custom-machined large-sample box used to protect human samples during processing (b) and of the small animal tissue holder for imaging (c) are provided. Antibodies that were validated during the development of enGLOW can be found in Supplementary Table 1.
a Diagram describing the steps to process mouse and human tissues. Prior to enGLOW processing, antibody validation is required on cryosections (steps A–J). Sample preparation is relevant for mouse GI tissues as they are obtained fresh while human samples are received pre-fixed (steps 1–8). Both human and mouse samples then start pre-treatment which involves delipidation and background fluorescence bleaching (steps 9 to 13). Immunolabeling enables the tagging of proteins of interest with fluorescent reporters that will be imaged in lightsheet fluorescence microscopy (steps 14–19). Cleared samples (steps 20–23) are then mounted for imaging (step 24). Processing mouse samples takes 2 weeks from start to finish, while human samples take 3 weeks due to prolonged immunolabeling incubations. Actual bench time is approximately 3 h for both sample types. b Diagram with dimensions of PTFE box used to protect human tissues during enGLOW processing. c Diagram with dimensions of custom holder used to image cylindrical mouse samples. Abbreviations: RT room temperature, PBS phosphate buffered saline, PFA paraformaldehyde, DCM dichloromethane, DBE dibenzyl ether. Created in BioRender. Planchette, A. (2025) https://BioRender.com/165ygdm.
enGLOW enables anatomical observations in centimeters of intact mouse gastrointestinal tissue
The 3D nature of enGLOW data enables cellular resolution imaging of centimeter-scale volumes of intact organs, giving access to data in multiple views (Fig. 2). To achieve high-quality imaging, samples are cleared (Fig. 2a) with minimal visible distortion of the tissue. Comparing a two-dimensional (2D) view of enGLOW data of autofluorescence with a standard cryosectioned haematoxylin and eosin-labeled mouse colon confirms that the typical layers of the gut wall are distinguishable in both methods (Fig. 2a, b). In addition to gut wall anatomy, we observe neuron cell bodies labeled with HuCD in the myenteric and submucosal plexuses of the enGLOW image (Fig. 2b). With examples of autofluorescent morphology, 3D renderings (Fig. 2c) permit large-scale visualization of uninterrupted anatomy volumes, while single longitudinal slices (Fig. 2di) identify the tissue layers at high resolution along the length of the cylindrical segment (Fig. 2dii) and circumferential reslices yield views of the sample’s cross section in the circumference (Fig. 2diii). In some samples, as the one shown here (Fig. 2d), bulging in the top half of the tissue is caused by the presence of a fecal pellet during transcardiac perfusion. Other possible distortions, presenting as gaps in the submucosa region (Fig. 2e), can occur during enGLOW processing as a result of repeated tissue dehydration. If anatomical preservation of this layer is crucial, embedding mouse tissues in agarose prior to clearing (Fig. 1 step 20) could preserve this layer’s integrity as seen in human samples. Mapping maximum intensity projections of longitudinal slices reveals autofluorescent structures, such as the papilla of Vater in the small intestine (Fig. 2f-i), autofluorescence of enteric neurons in the myenteric plexus (Fig. 2f-ii) and empty blood vessels in the submucosa (Fig. 2f-iii).
a Representative photographs of 2-cm-long segments of uncleared (left) and cleared (right) mouse distal colon over a millimeter paper. b Hematoxylin and eosin (H&E) stained cryosection of mouse colon imaged in brightfield with 40X objective in a slide scanner. c Circumferential reslice of two-channel image of mouse colon acquired in mesoSPIM with 2X magnification. Tissue architecture is visible by autofluorescence (cyan) and enteric neuron cell bodies are labeled with HuCD (magenta). In (b) and (c), the identifiable layers are annotated. d 3D rendering of mouse colon in autofluorescence channel acquired with mesoSPIM 2X magnification. e Longitudinal slice (i), with close-up of identifiable layers (ii) and 3-μm-thick circumferential reslice (iii) of mouse colon. Arrow in (ii) points to detachment of submucosa from muscle that can occur during sample preparation. f Close-up of notable structures of the gastrointestinal tract, including the villi and papilla de Vater (arrow) of the small intestine (i), autofluorescence of neurons (arrows, ii) and perfused blood vessels (arrow, iii). Abbreviations: LM longitudinal muscle, CM circular muscle, SM submucosa, MM muscularis mucosae, LP lamina propria, EP epithelium.
Unfolding complexity: 3D imaging and virtual tissue flattening to separate neuronal plexuses across the GI tract
Neuronal cells form clusters known as ganglia in two plexuses, one between the muscle layers and the other in the submucosa. The orientation of the ganglia within these layers makes it challenging to observe the resulting network with 2D cross-sectional views. By applying enGLOW to five key segments of the mouse GI tract with HuCD labeling, we observe variations in the distribution of the ganglia in 3D between the esophagus, duodenum, ileum, proximal colon and distal colon (Fig. 3a).
a Two-channel mesoSPIM 2X magnification 3D renderings of 3-cm-long segments with HuCD-labeled neuronal cells (magenta) in the esophagus (E), duodenum (D), ileum (I), proximal colon (PC) and distal colon (DC). a-i Close-ups of longitudinal slices showing layers from serosa to lumen of the same samples imaged at high resolution with COLM. Cells in the myenteric plexus (MP) and submucosal plexus (SMP) are annotated with arrows. a-ii High-resolution maximum intensity projections showing the shape of enteric neuron plexuses. b XYZ coordinates of individual segmented cells extracted from an imaged distal colon plotted in 3D space (left) and unwrapped and flattened to be plotted as a 2D plot (right). c Quantitative analysis from 3D datasets of organs along the gastrointestinal tract measuring the volume of HuCD+ clusters (c-i) and the number of clusters (c-ii). n = 1 animal. d mesoSPIM 2X magnification 3D rendering of distal colon segment with labeled neuronal fibers (Tuj1, orange), inhibitory neurons (nNOS, yellow) and autofluorescence (cyan). d-i Single longitudinal slice imaged at high resolution with COLM. d-ii Maximum intensity projection of Tuj1+ neuron fibers and nNOS. e Longitudinal slice of distal colon (left) and result of virtual flattening the tissue to the serosa edge (right). f Plexus-specific segmentation of HuCD-labeled neurons using depth mapping of the first 50 image planes (equal to 160 μm) from the serosa edge. g Quantitative analysis of spatially separated neuronal plexuses, showing differences in the plexus’ distance from the serosa (g-i) and the percentage occupation of the tissue by the two plexuses (g-ii). n = 4 regions of interest in n = 1 animal. If not specified, scale bars are 100 μm. Quantitative data used for plotting can be found in tables within Supplementary Data 1. Abbreviations: MP myenteric plexus, SMP submucosal plexus. Statistics: Error bars are minimum and maximum. One-way ANOVA with Tukey multiple comparisons correction for (c-i) and (c-ii); Two-way ANOVA with Tukey multiple comparisons correction for (g-i) and (g-ii).
Close-up images of gut organs acquired in a Clarity Optimized Lightsheet Microscope (COLM) enable high-quality inspection of gut wall anatomy (see “Methods”, Figs. 3a-i and 3a-ii). Renderings of autofluorescence provide tissue architecture context for labeled structures of interest (cyan in Fig. 3a, d, e), exemplified by observing the significant variation of the muscle layers and lamina propria from the esophagus to the distal colon. For example, the esophageal muscle is thickest (Fig. 3e-i) and layered diagonally to the length of the tissue (Fig. 3e-ii), as opposed to muscles that run longitudinally and circumferentially in the other organs shown here. Single longitudinal slices (Fig. 3a-i) show tissue layering and pinpoint the location of the myenteric (right-facing arrows) and submucosal (left-facing arrows) plexuses. Maximum intensity projections yield qualitative views of enteric ganglia distribution (Fig. 3a-ii). To explore the possible processing and quantification capabilities of enGLOW data, 3D segmentation of HuCD signals were performed in Imaris (Bitplane). The XYZ coordinates of individual cells can be plotted in 3D (Fig. 3b left) or as an unwrapped 2D matrix (Fig. 3b right), enabling a variety of modeling approaches to characterize cellular distributions at large scales. Quantified volumes of HuCD-positive clusters (Fig. 3c-i) and number of HuCD-positive clusters (Fig. 3c-ii) can be analyzed across the regions of the GI tract. Cellular segmentation using enGLOW data (Supplementary Fig. 1) is also achievable using established image processing tools, such as the Gut Analysis Toolbox40 implemented with ImageJ (Supplementary Fig. 2).
Co-labeling with multiple fluorescent markers is also possible, as shown in 3D renderings (Fig. 3d) and close-ups (Figs. 3d-i and 3d-ii) of neuronal fibers (Tuj1, orange) and nitric oxide-expressing neurons (nNOS, yellow). Such labels are impactful in revealing the neuronal network at cellular resolution in large segments of tissue but present more challenges for selective segmentation and quantification, though segmenting the signal as a whole and quantifying percentage of tissue occupation is straightforward (Supplementary Fig. 3).
To distinguish myenteric and submucosal plexus morphology from a single 3D image, we take advantage of the regularity of the serosa’s edge and of the thickness of longitudinal muscle (Fig. 3e) to enable layer-specific analysis. We virtually flatten the longitudinal slices (Fig. 3e) of the tissue to the serosa which are then orthogonally resliced to observe the two neuronal plexuses separately, having originated from the same 3D image (Fig. 3f). By assigning a common heatmap of distance in micrometers, we demonstrate plexus depth variations across gut regions visually (Fig. 3f) and quantitatively (Fig. 3g-i). Throughout the GI tract, both plexuses are detected within the first 50 pixels (163 μm) from the serosa (Fig. 3g-i). Each quadrant of the sample imaged in 3D was quantified to determine whether variations in signals were present in the circumference, which in the majority of cases, was not the case. The myenteric plexus shows overall little variation in distance from the serosa along the GI tract, with a slight increase in the esophagus (40 μm) compared to approximately 20 μm in all other regions measured. The shape of myenteric ganglia changes along the GI tract, with the ganglia elongating in the circumferential axis in the proximal and distal colon compared to esophagus and small intestine. The myenteric layers in the duodenum, ileum and proximal colon also contain fewer cells per clusters while the distal colon is generally populated by larger, 10 or more cells, per cluster. The submucosal plexus shows greater variability in distance from the serosa, ranging from 40 μm to 163 μm, which could result from submucosal layer distortion during processing. Thus, visual confirmation of tissue integrity is recommended to support quantitative interpretations.
3D imaging and virtual tissue flattening to reveal cellular patterns of neurons, glia and interstitial cells of Cajal in the mouse colon
Gut function, for example motility, depends on the coordination of signals by several interacting cells including neuronal fibers, glia and interstitial cells of Cajal (ICC). With enGLOW, it is possible to make parallel observations of 3D volumes and virtually flattened 2D images to interpret signal distribution in intact organs and in specific layers of the gut wall from a single 3D image. Neuron cell bodies, labeled with HuCD (Fig. 4a), form an extensive network of ganglia that reside in the myenteric and submucosal plexus layers with sparse distribution in the longitudinal view (Fig. 4a-i) and more visible elongated ganglia structure in maximum intensity projections (Fig. 4a-ii). When flattened, the distinct spatial distribution of myenteric and submucosal ganglia and the absence of specific signal in both muscle layers is clear (Fig. 4b). Faint non-specific signal can be seen in the muscle layers matching the shape of the myenteric plexus, likely coming from imperfect flattening. These ganglia are interconnected by a net-like structure, visible when labeled with Tuj1 to target the beta III tubulin constituent of the neuronal cytoskeleton (Fig. 4c). In a longitudinal close-up of the distal colon mucosa (Fig. 4c-i), fibers are seen extending through the submucosa and weaving around colonic crypts of the lamina propria up to the layer of epithelium. At high resolution, Tuj1 labeling highlights the intricate innervation of the mucosa (Supplementary Fig. 4 and Supplementary Videos 1 and 2) which can be relevant for studying specific structures such as epithelial lining innervation, isolated lymphoid follicles or the mesentery. The dense innervation connecting ganglia in the plexuses is made of internodal strands running in longitudinal and circumferential directions (Fig. 4c-ii), which in separated layers highlights the abundance of neuronal fibers in the circular muscle as well as in the two plexuses (Fig. 4d).
a HuCD labeling of neuron cell bodies (magenta) in distal colon imaged with mesoSPIM 2X magnification. a-i Longitudinal slice view of close-up COLM acquisition of HuCD. a-ii Maximum intensity projection of gut wall COLM acquisition of HuCD. b Virtually flattened HuCD labeling in longitudinal muscle, myenteric plexus, circular muscle and submucosal plexus layers of tissue. c Tuj1 labeling of neuronal fibers (orange) in distal colon imaged with mesoSPIM 2X magnification. c-i Longitudinal slice view of close-up COLM acquisition of Tuj1. c-ii Maximum intensity projection of gut wall COLM acquisition of Tuj1. d Virtually flattened Tuj1 labeling. e S100β labeling of glia (yellow) in distal colon imaged with mesoSPIM 2X magnification. e-i Longitudinal slice view of close-up COLM acquisition of S100β. e-ii Maximum intensity projection of gut wall COLM acquisition of S100β. f Virtually flattened S100β labeling. g C-kit labeling of interstitial cells of Cajal (green) in distal colon imaged with mesoSPIM 2X magnification. g-i Longitudinal slice view of close-up COLM acquisition of c-kit. g-ii Maximum intensity projection of gut wall COLM acquisition of c-kit. h Virtually flattened c-kit labeling. Cyan channel is tissue autofluorescence in (a), (c), (e) and (g). i Percentage of area in field of view occupied by specific signal for each labeled structure. If not specified, scale bars are 100 μm. n = 3 regions of interest in n = 1 animal. Quantitative data used for plotting can be found in tables within Supplementary Data 1. Statistics: Error bars are standard error of the mean.
Enteric glia, sharing structural and functional similarities with astrocytes in the brain46, closely interact with neurons to coordinate signaling mechanisms involved in gastrointestinal reflexes and immune homeostasis in the gut9. Labeling glia by targeting the S100β protein (Fig. 4e) shows a similar pattern of distribution to both neuronal cell bodies and fibers (Fig. 4e-i). In a longitudinal slice maximum intensity projection (Fig. 4e-ii), S100β-positive strands and punctate signals are visible in the myenteric ganglia, internodal strands and tertiary plexus. Neurons are also visible by autofluorescence in the myenteric plexus (Fig. 4e-ii arrows). Glia have a widespread distribution within the layers of the gut wall, with cells and strands present in all four layers separated in the virtually flattened images (Fig. 4f).
Interstitial cells of Cajal are another important cellular component for gastrointestinal function, as they provide the pacemaker rhythm for motility and interact with neurons to modulate neurotransmitter release47. The distribution of interstitial cells of Cajal, labeled with c-kit, is distinct from the neuronal and glial patterns, as they create two overlapping grids that run parallel to the longitudinal and circular muscle (Fig. 4g). In the muscle, the myenteric ICC subtype (ICC-MY) can be seen surrounding the myenteric plexus (Fig. 4g-i) and the intramuscular subtype (ICC-IM) forms dense bundles throughout the muscle (Fig. 4g-ii). Additionally, c-kit positive signal is observed at the base of the colonic crypts in the lamina propria (Fig. 4g-i arrow) though this is known to identify a specialized colonic goblet cell48. In the virtually flattened layers, ICCs are abundant in longitudinal and circular muscle layers, as well as in the myenteric plexus region but are mostly absent in the submucosal plexus (Fig. 4h).
Quantitative analysis of flattened images can be achieved, for example by quantifying the area of tissue occupied by labeled structures (Fig. 4i). In addition to labeling and analyzing endogenous signals, enGLOW processing preserves fluorescent reporter proteins of genetic modifications as shown in the R26 LSL FSF ReaChR-mCitrine line (B6;129S-Gt(ROSA)26Sortm2.1Ksvo/J) optogenetic model (Supplementary Fig. 5).
3D imaging and virtual tissue flattening of neuronal ganglia in full-thickness human colon resections
By adapting enGLOW for processing human samples, we can image colon resections of up to 4 cm × 5 cm × 0.3 cm in 3D. As with small animal tissues, enGLOW processing involves clearing of full-thickness resections that requires embedding in agarose to maintain tissue integrity (Fig. 5a). A 2D cross-sectional reslice of autofluorescence and HuCD signals facilitates the observation of detailed layered morphology in the full-tissue-thickness (Fig. 5b). Seven major layers of the gut wall can be seen, interspersed with HuCD-labeled neurons, as well as the distinctive banding pattern of circular muscle found in the human colon (Fig. 5b double-ended arrow). In addition to the labeled cell bodies, we observe some non-specific binding in the form of small speckled signal noticeable in the submucosa and lamina propria. Human tissues thus require attention and sometimes refinement to minimize non-specific binding as seen here. As these samples are not perfused, blood vessels retain high autofluorescence that are visible specifically between the longitudinal muscle and serosa, as well as in the submucosa (Fig. 5b ‘v’). The reconstructed 3D volume (Fig. 5c) reveals approximately 14 neuronal ganglia across both myenteric and submucosal plexuses within a 1.7 cm2 area of tissue, as well as the extensive vascular network.
a Representative photographs of colon surgical resections before (left) and after (right) enGLOW processing. b 2D cross-section of autofluorescence and neurons labeled with HuCD in 3.2-mm-thick colon acquired with mesoSPIM 2X magnification, where layers from the serosa to the mucosa are visible. c 3D rendering of 1.28 cm by 1.33 cm area of healthy colon resection of the same sample as in (b) with orthogonal view in dashed box. d Top: 2D flattened projections of high-resolution images acquired with COLM 4X magnification of HuCD-labeled neurons to separate cells in the submucosal and myenteric plexuses. Bottom: Depth mapping of the layers from serosa to submucosa (first 180 image planes equal to 585 μm). e 2D cross-section of neuron cytoskeleton labeled with Tuj1 in 3.3-mm-thick colon acquired with mesoSPIM 2X magnification where layers from the muscle to the mucosa are visible. f 3D rendering of 1.28 cm by 1.33 cm area of healthy colon resection with Tuj1 labeling of neuron fibers with orthogonal view in dashed box. g-i High-resolution 2D maximum intensity projections acquired with COLM 4X magnification. g Tuj1 signal wrapping around blood vessels. h Tuj1 signal in the epithelium enabling identification of specialized neuropod cells. i Dense network of innervation throughout the mucosa.
Applying virtual flattening to the serosa of human colon tissue enables plexus-specific observations (Fig. 5d, Supplementary Video 3), with neurons of the myenteric plexus seen approximately 70 μm from the serosa (Fig. 5d purple), while neurons interspersed in smaller clusters 150 to 400 μm from the serosa make up the external submucosal (Fig. 5d orange) and internal submucosal (Fig. 5d yellow) plexuses49 in a 1.8 mm × 1.8 mm depth map close-up.
The extensive neuronal fiber network, labeled with Tuj1, is seen in all layers of the human colon, as seen in a 2D cross-sectional view (Fig. 5e). In 3D, the intricate organization of innervation makes delineating the network particularly challenging with currently available image processing methods (Fig. 5f). However, specific morphologies of how neuronal fibers spread around structures of the gut wall can be accessed by mapping intensity projections of 3D images (Fig. 5g-i). Cellular-resolution COLM images reveal the wrapping of the vascular network by neuronal fibers in the submucosa (Fig. 5g), specialized neuropod cells50 in the epithelium lining (Fig. 5h, Supplementary Video 4) and patterns of dense innervation surrounding colonic crypts (Fig. 5i).
enGLOW demonstration on tissue from genetic model of Parkinson’s disease
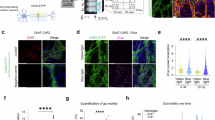
enGLOW provides rich multi-channel 3D datasets of morphology that enable investigation of morphological changes occurring in disease, as shown by implementing the workflow in colon tissues of the mitochondrial complex I genetic mouse model51 of Parkinson’s disease (MCI-Park). In this model, a homozygous knockout of the NDufs2 gene expressing a core subunit of mitochondrial complex I, specifically in dopaminergic neurons, leads to an early-onset Parkinsonian phenotype. We identify changes in tissue morphology using circumferential reslices of autofluorescent signal from four MCI-Park and four healthy animals (two of each shown in Fig. 6a). Highlighted reductions in colonic crypt length (rectangles), crypt number in the circumference, and epithelium layer thickness (circles) compared to healthy controls (Fig. 6a) were quantified and plotted (Fig. 6b).
a 2D circumferential reslices of autofluorescence channel in mouse colon from MCI-Park mice and wild type controls. Scale bars in close up boxes are 200 μm. Yellow circles and rectangles indicate the epithelium layer and the length of crypts respectively. The size is the same in all images to facilitate visual comparison. b Quantitative measurements of crypt length (b-i), crypt count (b-ii) and epithelium thickness (b-iii) obtained from images in (a). c 3D renderings of HuCD-labeled neurons in the same mice as in (a). d Quantification of segmented HuCD-labeled neurons from whole imaged volume of PD and WT samples. Metrics are count of individual cells per mm3 (d-i), count of neuron clusters per mm3 (d-ii) and volume of neuron clusters (d-iii). e Longitudinal view of two-channel images of autofluorescence and HuCD flattened to the serosa. Arrows point to aberrant HuCD signal in the mucosa, that may be an artifact that was observed in all tested MCI-Park animals. f Quantification of classified segmentation of neurons comparing the volume per mm3 of ganglia (left) and small clusters (right). g Quantification of classified segmentation of neurons comparing the count per mm3 of ganglia (left) and small clusters (right). h High-resolution single optical slice of PD1 sample with two orthogonal views of the mucosa imaged with COLM 4X magnification. i Representative close-up of high-resolution 3D rendering of Tuj1+ neuronal fibers from PD1 animal. j Quantitative analysis of volume occupied by Tuj1 signal in duodenum, jejunum and distal colon comparing PD animals and wild-type controls. Quantitative data used for plotting can be found in tables within Supplementary Data 1. Statistics: Error bars are minimum to maximum. Unpaired t-test for (b), (d), (f) and (g). One-way ANOVA with Tukey multiple comparisons correction for (j). n = 4 MCI-Park and n = 4 wild-type controls for all graphs.
By segmenting HuCD signal in 3D renderings (Fig. 6c), we found a 3-fold increase in the number of individual neuronal cells compared to wild type (Fig. 6di), which was reflected in the number (Fig. 6dii) and volume (Fig. 6diii) of neuronal clusters. To clarify the source of this observation, high-resolution longitudinal slices of tissue showed unusual HuCD signal in the mucosa of all PD mice, that may be an artifact and would require further investigation (Fig.6e). 3D segmentation of HuCD signal was classified into small clusters and larger ganglia to perform quantitative analysis. The small signal clusters are in fact the source of the difference between the two groups (Fig. 6f, g). In both cluster volume and count, the ganglia do not differ in the MCI-Park mice compared to wild type, with a median volume of 10.8 mm3 (Fig. 6fi) and mean 200 ganglia per mm3 (Fig. 6gi) of tissue. Conversely, small signal clusters show an increase in median volume (12.2 mm3 vs. 7.36 mm3, Fig. 6fii) and number of segmented clusters (1088 vs. 205 per mm3, Fig. 6gii) in MCI-Park tissue compared to wild type. This difference is also correlated with the spatial distribution of HuCD signal, seen in the lamina propria of MCI-Park images of flattened longitudinal slices (Fig. 6e) and a longitudinal slice offering orthogonal views of the mucosa (Fig. 6h). The sample size of four animals makes it challenging to establish the consistency of the observed differences in HuCD signal between MCI-Park and control animals. With further experiments, it may be possible to establish whether a disrupted barrier is at the origin of these differences. Neuronal fiber labeling in MCI-Park tissue (Fig. 6i), alongside Tuj1 percentage tissue occupation in the duodenum, jejunum and distal colon (Fig. 6j), confirm that neuronal fiber remodeling is not found. As such, this dataset is not sufficient to determine whether neuronal remodeling occurs in this model of Parkinson’s disease but does highlight the diversity of analyses and the value of 3D cellular resolution investigation of potentially pathological tissues using enGLOW.
Discussion
High-resolution 3D imaging of the enteric nervous system has broad applications, ranging from establishing clearing methodologies41,52, studying cellular interactions of specialized cells in animal and human53 tissues, and unraveling pathological implications of morphology in Hirschsprung’s disease32 and colon cancer54. We present a detailed workflow optimized for cellular-resolution 3D imaging and quantification of enteric morphology at the organ scale in intact gastrointestinal mouse and human tissues, with an example of its application comparing structures in healthy versus disease in a genetic model of Parkinson’s. With enGLOW, we aim to provide a single workflow encompassing all steps necessary to achieve high-resolution 3D observation and quantification in large intact mouse and human gastrointestinal samples.
We analyze the distribution of four cellular components critical to enteric network function, demonstrating that 3D segmentation of neuronal cells, fibers, glia, and interstitial cells of Cajal provides datasets for in-depth investigation. Further processing by virtual flattening and reslicing enables the isolation of the myenteric and submucosal plexuses of the ENS, all from a single 3D dataset.
Even though enGLOW takes on average 9 days for processing centimeter-long segments of mouse gut in comparison to 1–3 days for dissected layers35 or whole-mount preparations32, it only requires 3 h of bench time and gives access to more comprehensive analyses of pathologies. Differences in structural parameters of the mucosa, such as epithelium thickness, and variations in distribution of specific labels can be observed. The method is also scalable to process full-tissue-thickness (up to 3.3 mm) human samples and provides unprecedented resolution in 3D images of excised gastrointestinal tissue at this scale. The capability of enGLOW to achieve 3D imaging in human resections creates a new opportunity for investigating network-scale morphological remodeling of the gut in human gastrointestinal diseases. However, in addition to the longer processing time of enGLOW compared to whole mount, the size of datasets resulting from enGLOW are not negligible, with 2 cm × 2 cm human samples commonly resulting in 100 gigabytes of data for three channels. Thus, the scientific advantage of accessing information at the network level needs to be well justified.
Key enGLOW adaptations enable high-quality repeatable 3D imaging by optimizing gut-specific sample preparation and mounting for imaging. For example, processing cylindrical mouse samples with a needle support inside the lumen maintains its shape and facilitates imaging using a custom-designed holder included here. However, possible detachment of the submucosa can occur as a result of dehydration and rehydration cycles during sample preparation. For human resections of up to 4 cm × 3 cm, processing inside a solvent-resistant (PTFE) box limits layer dissociation during dehydration/rehydration steps. Furthermore, mounting human resections in agarose prior to clearing facilitates high-quality 3D imaging at a 45° angle to the illumination and detection arms of the microscope.
Some limitations to the current study should be acknowledged. Results from the Parkinson’s disease cohort are insufficient to make conclusive interpretations as only four animals from each group could be compared. Due to the unknown variability of the physiological effects of this genetically-induced form of Parkinson’s disease on the gut, an extended study should be conducted to characterize this potentially pathological phenotype. In addition, we encountered potential labeling artifacts in samples collected from MCI-Park animals which also contributed to a challenging interpretation of morphological remodeling in this model and shows limitations of the proposed pipeline. Care must thus be taken to consistently validate labeling in disease samples as changes in tissue or cell permeability can lead to unusual labeling as shown here. However, evidence of perturbed epithelial barrier integrity seen in patients55, preclinical56 and primate57 models of Parkinson’s disease may explain the disruption we observe in epithelial architecture and epithelial permeability to antibody labeling. In mouse tissues, the visualizations and quantitative analyses focused on cylindrical organs of the GI tract, however processing of the stomach and the cecum was tested and is possible with enGLOW but was beyond the scope of the present study. On the other hand, enGLOW was not tested on potentially more fragile or impenetrable diseased human tissue since we only used healthy segments. While autofluorescence provides invaluable context for labeled structures, background fluorescence cannot replace label-specific cell type identification and can introduce false positives in other channels and confound quantitative analyses, especially in non-perfused human tissues. We have found a few methods to help minimize the impact of autofluorescence on false positives, including increasing the duration of the tissue bleaching step, ensuring the antibody labeling results with clean specific labeling (done by validating concentrations and number of wash steps) and using fluorescent labels in the red to far red wavelength bands. Nuclear staining, which is a common source of context around targets of interest, is possible to image with light-sheet microscopy58. However, we have found that autofluorescence in gastrointestinal tissues provides greater quality and quantity of information than would be obtained with a selective nuclear marker. Furthermore, antibody labeling can be challenging if antibodies are not able to target methanol-treated tissue and if compatible antibody species combinations are not resistant to methanol, thus making co-staining with antibodies of interest sometimes impossible. In human samples specifically, non-specific signal was observed in HuCD-labeled tissue and requires improvement. While the antibody validation phase can help address these challenges, those related to the large volume of intact tissue may only be troubleshooted in full-thickness samples. Furthermore, co-staining multiple targets can be challenging due to limitations in compatible antibody species being available to stain peripheral structures. Imaging artifacts may also occur, such as uneven signal intensity in the tissue leading to bright spots that cast a shadow through the rest of the sample in that alignment. The reliance of this method on evenly cleared samples can hinder light-sheet penetration, whereby sharp focus may be lost deeper into the tissue. In this study, only few samples for each antibody staining were shown as a means to demonstrate the varied applications of enGLOW, thus limiting observations of enteric morphology variability based on factors such as age or sex of animals. For such an understanding to be gained, more biological samples would be needed. Furthermore, we did not investigate the inter-user variability of image processing and quantitative data extraction. However, the repeatability of the proposed tissue preparation protocol was successfully tested by an inexperienced user with little prior knowledge of gastrointestinal tissue processing.
The potential impact of enGLOW comes at a time of continued discovery in the field of neurogastroenterology. Evidence of morphological remodeling of enteric tissue architecture spans a variety of mechanisms and diseases, with epithelial barrier integrity and mucosal inflammation being most prevalent across inflammatory bowel diseases59 (IBD), diabetes60,61 and neurological diseases62,63. Specialized cells such as enteroendocrine cells are an example of potentially novel targets for understanding disease in the gut64, and possibly in Parkinson’s disease65, that are accessible with enGLOW. Furthermore, enteric nervous system morphology is known to be pivotal in Hirschsprung’s disease66 and emergent observations have identified its implication in neurodegenerative diseases such as Parkinson’s67. In Hirschsprung’s, large-scale observation in intact tissue has proven critical in unraveling pathological alterations32, thus highlighting the need for large-scale cellular-resolution analysis of morphology.
Moreover, the field of neurogastroenterology has seen recent advances in functional recording of gastrointestinal activity, as seen by new tools for pre-clinical in-vivo assessment of gut electrophysiology68,69. By accessing large volumes of tissue, enGLOW can enable understanding of function in the context of gastrointestinal tissue morphology through structural assessments of networks of involved cells.
In the context of gut status implication in a wide range of diseases, our method addresses the need for high-quality assessments of gut tissues in large segments of anatomically intact samples. The added image processing applications, specifically the capability to separate layers of the gut wall from intact samples and from a single dataset, significantly increase the value of enGLOW. Morphological remodeling of the gut at scales that encompass functional networks can be explored, with access to thousands of cells across variable functional regions in a single large dataset. enGLOW has the potential to impact pre-clinical and clinical research in the context of Hirschsprung’s disease but also IBD, diabetes, autism and Parkinson’s disease. Also, the possibility of parallel processing of the gut from the same animal whose brain is being processed with other relevant techniques, will allow enGLOW to advance gut–brain axis research and offer new insights into related diseases.
Methods
Gastrointestinal tissue collection
Gastrointestinal tract samples were excised from wild-type adult C57BL/6 mice and transgenic MCI Parkinson model mice51 (Ndufs2tm1.1Job Slc6a3tm1.1(cre)Bkmn/SurmJ genetically-modified line, Jackson Laboratories strain #036313). Gastrointestinal tissues from wild type and MCI-Park animals were collected separately but by the same experimenter and underwent the same procedure. The MCI-Park tissues were perfusion-fixed by the laboratory of Prof. Courtine (NeuroRestore, EPFL) and dissected by the same experimenter as in the wild-type animals. All animals were raised under specific pathogen-free conditions and handled in compliance with Swiss Veterinary Law guidelines. Wild type and transgenic animals were bred and raised in the same animal facility with corresponding diet, housing conditions and light cycles. All procedures were approved by the Veterinary Office of the Canton of Geneva (GE241D). Animals whose tissues were collected for this manuscript were aged 10–24 weeks of age and were wild-type. We did not select for sex.
All human samples were collected at Nantes University Hospital, France. The study protocol was approved by the French Ethics Committee for Research on Humans (DC-2008-402). Patients undergoing surgery gave their approval to take part in the study. Healthy segments of tissue distant from resected colorectal cancer were provided for this study. All ethical regulations relevant to human research participants were followed.
Overview of enGLOW procedures
enGLOW is adapted from the iDISCO+ protocol45 to optimally process mouse and human gastrointestinal tissues, in terms of sample handling, incubation lengths, antibody validation, sample mounting and subsequent imaging. A detailed step-by-step schematic of the enGLOW protocol can be found in Fig. 1. The workflow takes two weeks for mouse samples and three weeks for human resections, due to prolonged incubations to accommodate for the larger human samples. Actual bench time is approximately 3 h for both pipelines.
Antibody labeling validation on sections
Validation of antibodies on methanol-treated sections was systematically necessary to find primary antibodies that effectively labeled structures in the peripheral nervous system, listed in Supplementary Table 1. To do this, 12–25-μm-thick cryosections of either mouse or human tissue mounted on slides are incubated with 100% methanol for 3 h, with methanol refreshes every 30 min. This is followed by 1 h in blocking buffer at room temperature (RT, 6% donkey serum ThermoFisher 31873, 10% DMSO Sigma 472301 in PBS 0.2% Triton-X100 Merck 108603), overnight incubation at RT with primary antibody, 3× 10-min washes in 0.2% PBS-Tween 20 (Sigma P9416) at RT, 1 h secondary antibody at RT, 3× 10-min washes 0.2% PBS-Tween 20 at RT, DAPI (1:1000) counterstaining for 10 min at RT, a final 10-min wash in 0.2% PBS-Tween 20 at RT and mounting with cover slips. Slides were left to rest for minimum 12 h before imaging in either a slide scanner or a confocal microscope.
enGLOW mouse sample collection and preparation
One of the key enGLOW optimizations focuses on protecting tissue layers and maintaining shape for imaging of intact organ segments. Animals are deeply anesthetized by intraperitoneal injection of sodium pentobarbital (150 mg/kg). Blood is removed by transcardiac perfusion of 10 mL phosphate buffered saline (PBS), then tissues are fixed by perfusing 10 mL of 4% paraformaldehyde (PFA, EMS 15710) diluted in PBS. To access the gastrointestinal organs, the abdomen is opened from pelvis to rib cage. Cylindrical organs are identified (includes esophagus, duodenum, jejunum, ileum, proximal and distal colon) and cut in segments of up to 3 cm in length. To empty the lumen, if content is visibly present, angled forceps are used to gently press a segment from the outside and slide to push out any content. To track oral versus aboral physiological orientation, the oral end is marked by a 2 mm longitudinal snip into the tissue. In cases where cylindrical morphology is a hindrance for downstream analysis, for example if flattened visualizations of the myenteric plexus are of greater interest, enGLOW is compatible with gut tissues cut lengthwise and pinned flat during post-fixation (as is done for whole-mount processing33). For non-cylindrical organs (i.e., stomach and cecum), a 1–2 cm portion of tissue is excised and cleaned the same way but physiological orientation is lost. The mesentery, found on the outside of the gut wall, often creates bright artifacts in the images and complicates virtual flattening so it is removed by gentle pinching and peeling with forceps. To minimize shrinkage and stop collapsing of cylindrical samples, segments are post fixed in 4% PFA at 4 °C overnight with a steel needle or glass capillary (0.6 mm diameter or 23 G) inserted into the lumen. It is possible to pause the workflow at this step and store samples at 4 °C before continuing, however, in this case the needle or capillary shall be removed.
enGLOW human sample collection and preparation
Macroscopically normal human ileocolonic resections (proximal margin) were immediately transported to the laboratory and placed into ice-cold oxygenated Kreb’s solution containing (mM; 117 NaCl, 4.7 KCl, 1.2 MgCl2, 1.2 NaH2PO4, 25.0 NaHCO3, 2.5 CaCL2 and 11 glucose). Samples were pinned flat and stretched (mucosa up) in a dissection dish, fixed with 4% paraformaldehyde in PBS for 3 h at room temperature, or overnight at 4 °C. Tissues can be stored in 0.1% NaN3 in PBS at 4 °C for several years. Larger human resections can be cut with surgical scissors to the desired size for each experiment. As their edges are sensitive to tearing between circular muscle and submucosa, it is recommended to cut the tissue immediately before starting the next step.
enGLOW pre-treatment
This step consists of removing lipids and bleaching background fluorescence. Due to delipidation fragilizing the integrity of the submucosa in human resections, samples are caged in a custom PTFE box with holes to allow the flow of solutions during pre-treatment, which is then placed in a beaker and covered with parafilm for incubations. The size of the box depends on the largest expected size of human resections to be processed that is still compatible with the imaging set-up. Our box was made by cutting a PTFE block to 30 mm in width, 40 mm in length and 10 mm in height, followed by coring the block to an open box with walls 2 mm thick. Holes were drilled with a 5 mm drill bit at 3 mm intervals in all three walls. The lid was made from a 3 mm thin sheet of PTFE. A 2.4 mm diameter hole was drilled in each of the four corners of the lid and box to enable screwing the box shut. If PTFE is not accessible, a material that is resistant to dichloromethane and dibenzyl ether can be used.
In a fume hood, samples are dehydrated in consecutive 20%, 40%, 60%, 80%, and 100% methanol (Sigma 32213) incubations diluted in distilled water at room temperature, for 1 h each (mouse) or 1.5 h each (human). Samples are cooled in a second 100% methanol incubation for 1 h (mouse) or 1.5 h (human) at 4 °C. Lipids are removed in 66% dichloromethane (DCM, Sigma 34856) diluted in 100% methanol overnight at 4 °C. Two 100% methanol washes remove remnants of DCM for 1 h each at 4 °C. Samples are bleached in 5% H2O2 (Reactolab 99230) diluted in 100% methanol overnight at 4 °C. This is followed by rehydration to an aqueous solution with a reverse serial dilution of methanol and water (80%, 60%, 40%, 20%, and PBS) for 1 h each (mouse) or 1.5 h each (human). If no immunolabeling is desired or the samples have genetic expression of labels, this rehydration step is skipped and samples can go directly to delipidation step (21) in the clearing stage. All incubations are under agitation. It is possible to pause the workflow at this step and store samples at 4 °C in PBS before continuing.
enGLOW immunolabeling
For tissue immunolabeling, the needle is removed from the lumen of mouse tissues and the human samples remain in the PTFE box. All incubations are done at 37 °C on a shaker. All samples are incubated in permeabilization buffer (0.2% Triton-X100 Merck #108603, 20% DMSO Sigma 472301 and 0.3 M Glycine Sigma G7126-500G in PBS) for 1 day followed by blocking buffer (6% donkey serum ThermoFisher 31873, 10% DMSO Sigma 472301 in PBS 0.2% Triton-X100 Merck 108603) for 1 day. Samples are first labeled with primary antibody (1:500 most common dilution upon validation) in blocking buffer for 1 day (mouse) or 2 days with a solution refresh after 1 day (human). The larger human samples benefit from a solution refresh after 1 day, particularly in cases where the antigen is highly abundant in the tissue (for example Tuj1). All samples are washed minimum 5 times for 30 min each in 2% PBS-Tween 20 (Sigma P9416). Human samples can benefit from more wash cycles, going up to 10 washes with refreshes every 30 min. Secondary antibody conjugation (1:1000 most common dilution upon validation) is done in blocking buffer for 1 day (mouse) or 2 days with a solution refresh after 1 day (human). Samples are washed again a minimum of 5 times for 30 min each in 2% PBS-Tween 20. It is possible to pause the workflow at this step and store samples at 4 °C in PBS before continuing.
enGLOW clearing
To enable the optimal quality with light-sheet fluorescence microscopy, samples are made transparent by clearing. During clearing, the needle is once again inserted into the lumen of the mouse samples to retain a straight cylindrical morphology. To maintain the shape of human samples and simplify mounting into the microscope, human samples are embedded in 2% agarose (Thermo 17850) and a block is cut out according to the size of the imaging cuvette. To prepare for clearing, samples are dehydrated in consecutive 20%, 40%, 60%, 80%, and 100% methanol (Sigma 32213) incubations diluted in distilled water at room temperature for 1 h each (mouse) or 1.5 h each (human). At this point, the needle is removed from cylindrical mouse samples. Residual lipids are removed in 66% dichloromethane (DCM, Sigma 34856) diluted in 100% methanol for 3 h. The samples are washed in 100% DCM twice for 15 min each. The final clearing step is done without shaking in 100% dibenzyl ether (DBE, Sigma 108014) for at least 1 h (mouse) or 1.5 h (human) and until the sample is visually transparent. During the DCM and DBE incubation, containers must be filled to the top to prevent oxidation of the sample. All clearing steps are performed in a fume hood at room temperature and cleared samples are kept in 100% DBE protected from light in opaque tubes until imaging stored in a ventilated cabinet.
enGLOW 3D imaging with light-sheet microscopy
To facilitate repeatable mounting of cleared mouse samples, we designed a holder compatible with imaging in the lightsheet microscopes. This holder is made of machined stainless steel rail with two vertical prongs (40 mm tall, 4.6 mm wide and 2 mm deep) attached to a square base (2 mm tall, 10 mm wide and 10 mm deep) with holes (3 mm diameter) to allow fluid flow. Two needle points with magnetic heads (7.5 mm long, 0.5 mm thick) can slide inside of the rail allowing height adjustment to stabilize mouse samples.
In lightsheet microscopy, samples are illuminated with excitation light and the emitted light either from endogenous autofluorescent constituents or from exogenous labels is collected at 90° angle from the excitation axis (see Fig. 1 step 24). Tissue immobilization and orientation are crucial components of ensuring repeatable high-quality image acquisitions. For mouse tissue, samples are pinned in the custom holder to position the oral end of the sample upwards ensuring its longitudinal axis is orthogonal to both excitation and emission planes. The holder is placed in a quartz imaging cuvette that is filled to the top with 100% DBE to avoid cleared sample oxidation. For human tissues, it is recommended to position agarose-embedded samples in the DBE-filled cuvette at a 45° angle to the excitation and emission paths of the microscope, as shown in Fig. 1 step 24. The imaging cuvettes are then placed in a larger cuvette in the lightsheet microscope filled with Cargille oil (Cargille labs #16484) and positioned in place with a magnetic holder. Tissue autofluorescence is excited at 488 nm (detection with a bandpass filter 530/43), with two other channels (long pass filter with cutoff at 561 nm and long pass filter with cutoff at 663 nm) being available for fluorescent targets. The 663 nm channel is preferred for less abundant targets due to the inherently lower background tissue fluorescence and lower scattering.
Both our lightsheet microscopes generate 16-bit TIFF files that undergo pre-processing (stitching and channel merging with TeraStitcher70) prior to segmentation and quantitative analysis.
MesoSPIM microscope for large-scale imaging
The mesoSPIM71 is an open-source lightsheet microscopy platform developed for high-throughput, high-resolution imaging of large, optically cleared biological specimens (cm size). The system utilizes a long working distance objective (Olympus MVPLAPO), which is compatible with a range of refractive indices (RI 1.33–1.56), making it versatile across different tissue clearing protocols like CLARITY, iDISCO+, and CUBIC. Illumination is achieved via galvo-scanned laser light sheets and electronic tunable lens,
while detection is handled by sCMOS cameras optimized for large-format, low-noise acquisition. It also includes a motorized XYZ stage for volumetric tiling and stitching of large datasets. Control is provided through open-source Python-based software, offering automated image acquisition, laser control, and real-time feedback. The mesoSPIM architecture is described in detail in the following paper71. Images from mouse samples are typically acquired with the 2X zoom of the Olympus MVX-10 objective with a voxel size of 3.26 μm × 3.26 μm × 3 μm (x, y, z) and a field of view of 7 × 7 mm2. The zoom factor selected for human samples is based on the size of the sample and the desired spatial resolution and typically ranges between the 2X and 1X zoom.
Clarity optimized light-sheet microscopy for high spatial resolution imaging
The CLARITY-optimized light-sheet microscope, or COLM, was developed in the Deisseroth laboratory by Raju Tomer72 specifically to meet the demands of high-resolution imaging of CLARITY-cleared tissues. The detection axis is orthogonal to the illumination, allowing for optical sectioning with reduced phototoxicity and high speed. Detection is achieved using high-performance, long working distance multi-immersion objectives such as the Olympus 16X NA 0.6 and WD 8 mm, which are specially designed to work in refractive index-matched media around 1.33 to 1.56, the typical range for CLARITY-cleared tissues. COLM’s sample chamber is custom-fabricated with index-matched optical windows (quartz) to minimize chromatic and spherical aberrations and to provide a stable imaging environment. Multicolor imaging is supported through laser lines at 488, 561, and 647 nm, with appropriate filter wheels allowing for rapid channel switching. The system is described in detail in the following paper72. To observe specific cell types in mouse tissues, we use the 10X/0.6 CLARITY WD 8 mm objective with a voxel size of 0.59 μm × 0.59 μm × 3 μm (x, y, z) and field of view of 1.208 × 1.208 mm2. Since biological constituents are significantly larger in human tissue, we use the 4X/0.28 (d) WD 29 mm objective with a voxel size of 1.44 μm × 1.44 μm × 5 μm (x, y, z) and a field of view of 2.94 mm × 2.94 mm2.
Spectral unmixing in mesoSPIM and COLM
The imaging systems (mesoSPIM and COLM) use high-quality objectives specifically selected to minimize chromatic aberration across a broad range of wavelengths, especially in high-refractive-index (RI) media such as those used in cleared tissues (e.g., dibenzyl ether, RI ~ 1.56). This “built-in” correction of axial chromatic aberration ensures that, at acquisition, the various fluorophores are imaged in focus without post-hoc z-registration. While residual chromatic shifts can occur, empirical studies and specifications for these objectives indicate that such shifts are generally within the range of 2–3 µm in plane, and do not exceed 4–5 µm axially, even in high-resolution configurations. Importantly, the biological structures we examine—such as enteric neuronal cell bodies, ganglia, glial networks, and interstitial cells of Cajal—are typically spaced tens to hundreds of microns apart, and are distributed across distinct anatomical layers (e.g., myenteric vs. submucosal plexus). Therefore, the expected extent of chromatic aberration is significantly smaller than the inter-structure distances.
Virtual flattening of images of GI tract to the serosa for virtual separation of layers
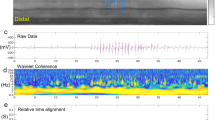
To separate layers of the gut wall in 3D images of intact tissues, we developed a virtual flattening algorithm in Python that identifies the serosa (outermost layer of gut wall) and performs numerical 3D alignment. Serosa segmentation is sequentially performed on the autofluorescence channel of a longitudinal image stack. First, the background is attenuated using a rolling ball of radius 25 pixels73. A mask is then created by thresholding the autofluorescence signal. The serosa location is found as the first positive pixel in the mask for each row in the images. Positions are smoothed using a lowpass Butterworth (order 6, cutoff 5%) filter. Finally, each pixel row is shifted according to the found serosa position so that they are in the same horizontal position. Labeled fluorescence channels are then corrected using the same coordinates since all channels are acquired consecutively without changing tissue orientation. From the flattened longitudinal slice stack, an orthogonal reslice from the serosa side of the image is computed with ImageJ to create a new stack of the flattened tissue in plane. The python code has been made available at the following link: https://zenodo.org/records/15736375.
Autofluorescence morphology quantification
Parameters that were measured in autofluorescence include crypt length, crypt number and epithelium thickness. Automated segmentation in the autofluorescence channel is challenged by the lack of morphological specificity of the fluorescence signal. For this reason, we relied on manual extraction of quantitative parameters. Longitudinal slices were used to measure distances in ImageJ, manually drawing from the edge of the crypt on the lumen side to the apex of the crypt facing inwards. The thickness of the epithelium was measured using the same approach. To quantify the number of crypts in the circumference, circumferential reslices were used and crypts were manually annotated and quantified in ImageJ.
Labeled morphology segmentation and quantification
Data from multi-channel lightsheet acquisitions are rendered in 3D using Imaris software (Bitplane v9.8.2), where signals are segmented and quantified. The surfaces segmentation tool is used to isolate and quantify signals ranging from punctate cell structures to fibrous networks in the whole sample or in defined regions of interest. The process for segmentation of either individual cells or cell clusters is shown in supplementary Fig. 1.
XYZ coordinate plotting
To plot spatial distribution of single cells in 2D or 3D as shown in Fig. 3b, individual cells are segmented using the Surfaces tool in Imaris. In the data saving tab, the ‘Position’ option is selected and saved as a csv file. The header row of the resulting table should read “X ; Y ; Z ; ID” and all other columns should be removed. The 3D graph is plotted using matplotlib in Python, while the 2D plot is achieved by unwrapping the coordinates along the length of the sample using MATLAB (Version 24.2.0.2863752 (R2024b) Update 5), both codes have been made available at the following link: https://zenodo.org/records/15736375.
Histopathology
Gut samples are collected from perfused animals, post-fixed overnight in 4% PFA (EMS #15710) in PBS and cryoprotected in 30% sucrose (Sigma S9378) at 4 °C for 24 h. Cylindrical samples are frozen in isopentane (Reactolab 524391) at −55 °C cooled in a SnapFrost® 2 (Excilone, France). Sections of 20 μm thick are cut on a Leica CM3050S cryostat (Leica Microsystems GmbH, Germany) and mounted on microslides SuperFrost® Plus Menzel Epredia and stained with hematoxylin and eosin (H&E) in a Leica ST5020-CV5030 Stainer Integrated Workstation. Slides are imaged using an Olympus VS120 slide scanner.
Statistics and reproducibility
The data presented in this work are used to demonstrate the various image processing and data analysis applications made possible by the use of enGLOW. For this reason, the majority of figures are made up of single biological replicates (Figs. 2, 3 and 4). Where statistics are shown (Figs. 3c, g and 4i), the purpose is to show the range and distribution of values extracted from each biological sample. When comparing no more than two groups, unpaired t-tests were used. For comparison of more than two groups, one- and two-way ANOVAs were used with Tukey correction for multiple comparisons. Exact p-values are shown when significant. In Fig. 6, each group contains 4 biological replicates (MCI-Park PD and wild-type controls). Statistical analysis and interpretation of results obtained in these groups are limited due to the sample size and unknown expected variability of the biological phenomenon captured with the images. The methods described are highly reproducible once experimental conditions, such as the quality of labeling, are tested and established.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All images of mouse samples have been made available in the NeurO-GI Zenodo repository at the following link: https://zenodo.org/records/1489199574. Images of human samples are available from the corresponding author on reasonable request due to the large size of the datasets. Numerical data underlying all graphs in the article can be found in the Supplementary Data 1 file.
Code availability
Image processing codes, including 1-channel and 2-channel flattening, 3D xyz coordinate plotting and 2D xyz unwrapping, are available in the form of python notebooks in the NeurO-GI Zenodo repository at the following link: https://zenodo.org/records/1573637574.
References
Kang, Y.-N., Fung, C. & Vanden Berghe, P. Gut innervation and enteric nervous system development: a spatial, temporal and molecular tour de force. Development 148, dev182543 (2021).
Spencer, N. J. & Hu, H. Enteric nervous system: sensory transduction, neural circuits and gastrointestinal motility. Nat. Rev. Gastroenterol. Hepatol. 17, 338–351 (2020).
Mourad, F. H. & Saadé, N. E. Neural regulation of intestinal nutrient absorption. Prog. Neurobiol. 95, 149–162 (2011).
Näslund, E. & Hellström, P. M. Appetite signaling: from gut peptides and enteric nerves to brain. Physiol. Behav. 92, 256–262 (2007).
Yoo, B. B. & Mazmanian, S. K. The enteric network interactions between the immune and nervous systems of the gut. Immunity 46, 910–926 (2017).
Ye, L. & Liddle, R. A. Gastrointestinal hormones and the gut connectome. Curr. Opin. Endocrinol. Diab. Obes. 24, 9–14 (2017).
Furness, J. B. In Reference Module in Biomedical Sciences (Elsevier, 2015).
Wang, H., Foong, J. P., Harris, N. L. & Bornstein, J. C. Enteric neuroimmune interactions coordinate intestinal responses in health and disease. Mucosal Immunol. 15, 27–39 (2022).
Seguella, L. & Gulbransen, B. D. Enteric glial biology, intercellular signalling and roles in gastrointestinal disease. Nat. Rev. Gastroenterol. Hepatol. 18, 571–587 (2021).
Boesmans, W. et al. Imaging neuron-glia interactions in the enteric nervous system. Front. Cell. Neurosci. 7, 1–9 (2013).
Al-Shboul, O. The importance of interstitial cells of Cajal in the gastrointestinal tract. Saudi J. Gastroenterol. 19, 3–15 (2013).
Hounnou, G., Destrieux, C., Desmé, J., Bertrand, P. & Velut, S. Anatomical study of the length of the human intestine. Surg. Radiol. Anat. 24, 290–294 (2002).
Holland, A. M., Bon-Frauches, A. C., Keszthelyi, D., Melotte, V. & Boesmans, W. The enteric nervous system in gastrointestinal disease etiology. Cell. Mol. Life Sci. 78, 4713–4733 (2021).
Bonaz, B. L. & Bernstein, C. N. Brain-gut interactions in inflammatory bowel disease. Gastroenterology 144, 36–49 (2013).
Crudele, L., Gadaleta, R. M., Cariello, M. & Moschetta, A. Gut microbiota in the pathogenesis and therapeutic approaches of diabetes. eBioMedicine 97, 104821 (2023).
Mulak, A. & Bonaz, B. Brain-gut-microbiota axis in Parkinson’s disease. World J. Gastroenterol. 21, 10609–10620 (2015).
Chalazonitis, A. & Rao, M. Enteric nervous system manifestations of neurodegenerative disease. Brain Res. 1693, 207–213 (2018).
Cammann, D. et al. Genetic correlations between Alzheimer’s disease and gut microbiome genera. Sci. Rep. 13, 5258 (2023).
Kowalski, K. & Mulak, A. Brain-gut-microbiota axis in Alzheimer’s disease. J. Neurogastroenterol. Motil. 25, 48–60 (2019).
Marizzoni, M. et al. A peripheral signature of Alzheimer’s disease featuring microbiota-gut-brain axis markers. Alzheimers Res. Ther. 15, 101 (2023).
Anand, N., Gorantla, V. R. & Chidambaram, S. B. The role of gut dysbiosis in the pathophysiology of neuropsychiatric disorders. Cells 12, 54 (2022).
Petra, A. I. et al. Gut-microbiota-brain axis and its effect on neuropsychiatric disorders with suspected immune dysregulation. Clin. Ther. 37, 984–995 (2015).
Le Naour, J. et al. Improved Swiss-rolling method for histological analyses of colon tissue. MethodsX 9, 101630 (2022).
Li, Z. et al. Regional complexity in enteric neuron wiring reflects diversity of motility patterns in the mouse large intestine. eLife 8, e42914 (2019).
Nestor-Kalinoski, A. et al. Unique neural circuit connectivity of mouse proximal, middle, and distal colon defines regional colonic motor patterns. Cell. Mol. Gastroenterol. Hepatol. 13, 309–337.e3 (2022).
Zhang, Y. et al. Circadian rhythm disruption modulates enteric neural precursor cells. differentiation leading to gastrointestinal motility dysfunction via the NR1D1/NF-κB axis. J. Transl. Med. 22, 975 (2024).
Vicentini, F. A. et al. Intestinal microbiota shapes gut physiology and regulates enteric neurons and glia. Microbiome 9, 210 (2021).
Bhave, S. et al. Enteric neuronal cell therapy reverses architectural changes in a novel diphtheria toxin-mediated model of colonic aganglionosis. Sci. Rep. 9, 18756 (2019).
Khan, A. A. et al. Enteric nervous system regeneration and functional cure of experimental digestive Chagas disease with trypanocidal chemotherapy. Nat. Commun. 15, 4400 (2024).
Durand, T. et al. Visualizing enteric nervous system activity through dye-free dynamic full-field optical coherence tomography. Commun. Biol. 6, 236 (2023).
Darche, M. et al. Light sheet fluorescence microscopy of cleared human eyes. Commun. Biol. 6, 1025 (2023).
Graham, K. D. et al. Robust, 3-dimensional visualization of human colon enteric nervous system without tissue sectioning. Gastroenterology 158, 2221–2235.e5 (2020).
Mazzoni, M. et al. Quantitative analysis of enteric neurons containing choline. acetyltransferase and nitric oxide synthase immunoreactivities in the submucosal and myenteric plexuses of the porcine colon. Cell Tissue Res. 383, 645–654 (2021).
Greene, J. G., Noorian, A. R. & Srinivasan, S. Delayed gastric emptying and enteric nervous system dysfunction in the rotenone model of Parkinson’s disease. Exp. Neurol. 218, 154–161 (2009).
Cairns, B. R., Jevans, B., Chanpong, A., Moulding, D. & McCann, C. J. Automated computational analysis reveals structural changes in the enteric nervous system of nNOS deficient mice. Sci. Rep. 11, 17189 (2021).
Fleming, M. A., Ehsan, L., Moore, S. R. & Levin, D. E. The enteric nervous system and its emerging role as a therapeutic target. Gastroenterol. Res. Pract. 2020, 1–13 (2020).
Zundler, S. et al. Three-dimensional cross-sectional light-sheet microscopy imaging of the inflamed mouse gut. Gastroenterology 153, 898–900 (2017).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Kobayashi, Y. et al. COUNTEN, an AI-driven tool for rapid and objective structural analyses of the enteric nervous system. eneuro 8, ENEURO.0092-21.2021 (2021).
Sorensen, L. et al. Gut Analysis Toolbox—automating quantitative analysis of enteric neurons. J. Cell Sci. 137, jcs261950 (2024).
Neckel, P. H., Mattheus, U., Hirt, B., Just, L. & Mack, A. F. Large-scale tissue clearing (PACT): technical evaluation and new perspectives in immunofluorescence, histology, and ultrastructure. Sci. Rep. 6, (2016).
Sun, Y. et al. Sarm1-mediated neurodegeneration within the enteric nervous system protects against local inflammation of the colon. Protein Cell 12, 621–638 (2021).
Flinn, H., Cruz-Pineda, L., Montier, L., Horner, P. J. & Villapol, S. iDISCO tissue clearing whole-brain and light sheet microscopy for high-throughput imaging in a mouse model of traumatic brain injury. Methods Mol. Biol. https://doi.org/10.1007/978-1-0716-3662-6_39 (2024).
Vieites-Prado, A. & Renier, N. Tissue clearing and 3D imaging in developmental biology. Development 148, dev199369 (2021).
Renier, N. et al. iDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging. Cell 159, 896–910 (2014).
Grundmann, D. et al. Enteric Glia: S100, GFAP, and beyond. Anat. Rec. 302, 1333–1344 (2019).
Klein, S. et al. Interstitial cells of Cajal integrate excitatory and inhibitory neurotransmission with intestinal slow-wave activity. Nat. Commun. 4, 1630 (2013).
Rothenberg, M. E. et al. Identification of a cKit+ colonic crypt base secretory cell that supports Lgr5+ stem cells in mice. Gastroenterology 142, 1195–1205.e6 (2012).
Wedel, T. et al. Organization of the enteric nervous system in the human colon demonstrated by wholemount immunohistochemistry with special reference to the submucous plexus. Ann. Anat. 181, 327–337 (1999).
Kaelberer, M. M., Rupprecht, L. E., Liu, W. W., Weng, P. & Bohórquez, D. V. Neuropod cells: the emerging biology of gut-brain sensory transduction. Annu. Rev. Neurosci. 43, 337–353 (2020).
González-Rodríguez, P. et al. Disruption of mitochondrial complex I induces progressive Parkinsonism. Nature 599, 650–656 (2021).
Liu, Y. A. et al. Optical clearing improves the imaging depth and signal-to-noise ratio for digital analysis and three-dimensional projection of the human enteric nervous system. Neurogastroenterol. Motil. 23, e446–57 (2011).
Liu, Y. A. et al. 3-D imaging, illustration, and quantitation of enteric glial network in transparent human colon mucosa. Neurogastroenterol. Motil. 25, e324–38 (2013).
Valès, S. et al. Tumor cells hijack enteric glia to activate colon cancer stem cells and stimulate tumorigenesis. eBioMedicine 49, 172–188 (2019).
Clairembault, T. et al. Structural alterations of the intestinal epithelial barrier in Parkinson’s disease. Acta Neuropathol. Commun. 3, 12 (2015).
Pellegrini, C. et al. Enteric α-synuclein impairs intestinal epithelial barrier through caspase-1- inflammasome signaling in Parkinson’s disease before brain pathology. NPJ Park. Dis. 8, 9 (2022).
Coletto, E. et al. Dysregulation of epithelial ion transport and neurochemical changes in the colon of a Parkinsonian primate. NPJ Park. Dis. 7, 9 (2021).
Lu, C.-H. et al. Lightsheet localization microscopy enables fast, large-scale, and three-dimensional super-resolution imaging. Commun. Biol. 2, 177 (2019).
Quansah, E. et al. Intestinal epithelial barrier integrity investigated by label-free techniques in ulcerative colitis patients. Sci. Rep. 13, 2681 (2023).
Damms-Machado, A. et al. Gut permeability is related to body weight, fatty liver disease, and insulin resistance in obese individuals undergoing weight reduction. Am. J. Clin. Nutr. 105, 127–135 (2017).
Sorini, C. et al. Loss of gut barrier integrity triggers activation of islet-reactive T cells and autoimmune diabetes. Proc. Natl. Acad. Sci. USA 116, 15140–15149 (2019).
Li, F. et al. Leaky gut plays a critical role in the pathophysiology of autism in mice by activating the lipopolysaccharide-mediated toll-like receptor 4–myeloid differentiation factor 88–nuclear factor kappa B signaling pathway. Neurosci. Bull. 39, 911–928 (2023).
Munoz-Pinto, M. F. et al. Gut-first Parkinson’s disease is encoded by gut dysbiome. Mol. Neurodegener. 19, 78 (2024).
Nwako, J. G. & McCauley, H. A. Enteroendocrine cells regulate intestinal homeostasis and epithelial function. Mol. Cell. Endocrinol. 593, 112339 (2024).
Rodrigues, P. V. et al. Transcellular propagation of fibrillar α-synuclein from enteroendocrine to neuronal cells requires cell-to-cell contact and is Rab35-dependent. Sci. Rep. 12, 4168 (2022).
Eisenberg, J. D. et al. Three-dimensional imaging of the enteric nervous system in human pediatric colon reveals new features of Hirschsprung’s disease. Gastroenterology 167, 547–559 (2024).
Baumuratov, A. S. et al. Enteric neurons from Parkinson’s disease patients display ex vivo aberrations in mitochondrial structure. Sci. Rep. 6, 33117 (2016).
Srinivasan, S. S. et al. Luminal electrophysiological neuroprofiling system for gastrointestinal neuromuscular diseases. Device 2, 100400 (2024).
Xue, J. et al. Scalable balloon catheter assisted contact enhancement of 3D electrode array for colon electrophysiological recording. Sens. Actuators B Chem. 424, 136955 (2025).
Bria, A. & Iannello, G. TeraStitcher—a tool for fast automatic 3D-stitching of teravoxel-sized microscopy images. BMC Bioinformatics 13, 316 (2012).
Voigt, F. F. et al. The mesoSPIM initiative: open-source light-sheet microscopes for imaging cleared tissue. Nat. Methods 16, 1105–1108 (2019).
Tomer, R., Ye, L., Hsueh, B. & Deisseroth, K. Advanced CLARITY for rapid and high-resolution imaging of intact tissues. Nat. Protoc. 9, 1682–1697 (2014).
Sternberg, C. C. U. Biomedical image processing. Computer 16, 22–34 (1983).
Planchette, A., Sobolewski, A. & Scholler, J. enGLOW Image Processing Codes. https://doi.org/10.5281/zenodo.15736375 (2025).
Acknowledgements
We thank the laboratory of Prof. Courtine (NeuroRestore, EPFL), especially Dr. Antoine Collomb-Clerc and Dr. Quentin Barraud, for providing fixed samples of gastrointestinal tracts from MCI-Park model animals. We are grateful to the Advanced Lightsheet Imaging Center (ALICe) at the Wyss Center for Bio and Neuroengineering, Geneva. We thank Samantha Barale for her contributions to validating the applicability of the pipeline. This project has received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation program (grant agreement No. 853378).
Author information
Authors and Affiliations
Contributions
A.P. and M.J.G. formulated the initial idea for the work presented in this paper. A.P., A.S., and C.B. collected mouse samples. M.N., T.D., and P.N. provided human samples. I.G. developed custom imaging holder for mouse samples. K.W. developed custom human sample processing tools. A.P. and I.G. imaged the samples. Y.C. and S.P. assisted in imaging of samples. J.S. developed virtual flattening code and assisted A.P. with flattening image processing. A.P. processed all images and formatted figures. A.P. drafted the initial version of the paper. A.P., A.S., Y.C., T.D., P.N., G.B., M.N., S.P., and M.J.G. interpreted, critically reviewed and revised the manuscript for intellectual content.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Philippa Seika and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Joao Valente. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Planchette, A., Gantar, I., Scholler, J. et al. enGLOW 3D microscopy of the enteric nervous system in cleared human and mouse gut. Commun Biol 9, 357 (2026). https://doi.org/10.1038/s42003-026-09643-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-026-09643-6