Abstract
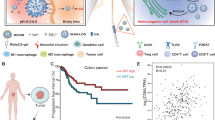
In the tumour microenvironment, accumulated lactic acid (LA) promotes tumour immune evasion by facilitating regulatory T cell (Treg) immunosuppressive function and restraining CD8+ T cell cytotoxicity, but the underlying mechanism remains elusive. Here we report that transcriptional factor MondoA-induced thioredoxin interacting protein (TXNIP) transcription is a common feature of both Treg and CD8+ T cells in response to lactic acid. In contrast to reduction in immunosuppressive capacity in MondoA-deficient Treg cells, loss of MondoA enhanced CD8+ T cell cytotoxic function in the lactic-acid-induced immunosuppressive microenvironment, by restoring glucose uptake and glycolysis. Mechanistically, lactic acid relied on sentrin/SUMO-specific protease 1 (SENP1) to stimulate the MondoA–TXNIP axis, which impaired TCR/CD28-signal-induced CD8+ T cell activation. Importantly, targeting the MondoA–TXNIP axis potentiated antitumour immunity in multiple cancer types and synergized with anti-PD-1 therapy to promote effective T cell responses in colorectal cancer. Our results demonstrate that the MondoA–TXNIP axis is a promising therapeutic target for improving cancer immunotherapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Bulk RNA-seq data that support the findings of this study have been deposited in the Gene Expression Omnibus database with the following accession codes GSE276640, GSE276642 and GSE276653. The HALLMARK_GLYCOLYSIS gene set is from the Molecular Signatures Database (MSigDB, systemic name: M5937). Source data are provided with this paper.
References
Fischer, K. et al. Inhibitory effect of tumor cell-derived lactic acid on human T cells. Blood 109, 3812–3819 (2007).
Qian, Y. et al. MCT4-dependent lactate secretion suppresses antitumor immunity in LKB1-deficient lung adenocarcinoma. Cancer Cell 41, 1363–1380 (2023).
Kumagai, S. et al. Lactic acid promotes PD-1 expression in regulatory T cells in highly glycolytic tumor microenvironments. Cancer Cell 40, 201–218 (2022).
Brand, A. et al. LDHA-associated lactic acid production blunts tumor immunosurveillance by T and NK Cells. Cell Metab. 24, 657–671 (2016).
Marrack, P. et al. Lactate regulates metabolic and pro-inflammatory circuits in control of T cell migration and effector functions. PLoS Biol. 13, e1002202 (2015).
Wang, H. et al. CD36-mediated metabolic adaptation supports regulatory T cell survival and function in tumors. Nat. Immunol. 21, 298–308 (2020).
MacIver, N. J., Michalek, R. D. & Rathmell, J. C. Metabolic regulation of T lymphocytes. Annu. Rev. Immunol. 31, 259–283 (2013).
Vaeth, M. et al. Store-operated Ca2+ entry controls clonal expansion of T Cells through metabolic reprogramming. Immunity 47, 664–679 (2017).
Bantug, G. R., Galluzzi, L., Kroemer, G. & Hess, C. The spectrum of T cell metabolism in health and disease. Nat. Rev. Immunol. 18, 19–34 (2018).
Chapman, N. M., Boothby, M. R. & Chi, H. Metabolic coordination of T cell quiescence and activation. Nat. Rev. Immunol. 20, 55–70 (2020).
Siska, P. J. et al. Mitochondrial dysregulation and glycolytic insufficiency functionally impair CD8 T cells infiltrating human renal cell carcinoma. JCI Insight 2, e93411 (2017).
Zhang, L. & Romero, P. Metabolic control of CD8+ T cell fate decisions and antitumor immunity. Trends Mol. Med. 24, 30–48 (2018).
Gerriets, V. A. et al. Metabolic programming and PDHK1 control CD4+ T cell subsets and inflammation. J. Clin. Invest. 125, 194–207 (2015).
Lim, S. A. et al. Lipid signalling enforces functional specialization of Treg cells in tumours. Nature 591, 306–311 (2021).
Gerriets, V. A. et al. Foxp3 and Toll-like receptor signaling balance Treg cell anabolic metabolism for suppression. Nat. Immunol. 17, 1459–1466 (2016).
Uhl, F. M. et al. Metabolic reprogramming of donor T cells enhances graft-versus-leukemia effects in mice and humans. Sci. Transl. Med. 12, eabb8969 (2020).
Watson, M. J. et al. Metabolic support of tumour-infiltrating regulatory T cells by lactic acid. Nature 591, 645–651 (2021).
Stoltzman, C. A. et al. Glucose sensing by MondoA:Mlx complexes: a role for hexokinases and direct regulation of thioredoxin-interacting protein expression. Proc. Natl Acad. Sci. USA 105, 6912–6917 (2008).
Ahn, B. et al. MondoA coordinately regulates skeletal myocyte lipid homeostasis and insulin signaling. J. Clin. Invest. 126, 3567–3579 (2016).
Wu, N. et al. AMPK-dependent degradation of TXNIP upon energy stress leads to enhanced glucose uptake via GLUT1. Mol. Cell 49, 1167–1175 (2013).
Sullivan, W. J. et al. Extracellular matrix remodeling regulates glucose metabolism through TXNIP destabilization. Cell 175, 117–132 (2018).
Nakayama, Y., Mukai, N., Kreitzer, G., Patwari, P. & Yoshioka, J. Interaction of ARRDC4 with GLUT1 mediates metabolic stress in the ischemic heart. Circ. Res. 131, 510–527 (2022).
Ahn, B. et al. MondoA drives muscle lipid accumulation and insulin resistance. JCI Insight 5, e129119 (2019).
Lu, Y. et al. MondoA–thioredoxin-interacting protein axis maintains regulatory T-cell identity and function in colorectal cancer microenvironment. Gastroenterology 161, 575–591(2021).
Zhu, Y. et al. MEK1-dependent MondoA phosphorylation regulates glucose uptake in response to ketone bodies in colorectal cancer cells. Cancer Sci. 114, 961–975 (2023).
Angelin, A. et al. Foxp3 reprograms T cell metabolism to function in low-glucose, high-lactate environments. Cell Metab. 25, 1282–1293 (2017).
Zhou, J. et al. Lactate supports Treg function and immune balance via MGAT1 effects on N-glycosylation in the mitochondria. J. Clin. Invest. 134, e175897 (2024).
Feng, Q. et al. Lactate increases stemness of CD8+ T cells to augment anti-tumor immunity. Nat. Commun. 13, 4981 (2022).
Kaymak, I. et al. Carbon source availability drives nutrient utilization in CD8+ T cells. Cell Metab. 34, 1298–1311(2022).
Chen, D. et al. Pan-cancer analysis implicates novel insights of lactate metabolism into immunotherapy response prediction and survival prognostication. J. Exp. Clin. Cancer Res. 43, 125 (2024).
Zhang, B. et al. Dynamic glycolytic reprogramming effects on dendritic cells in pancreatic ductal adenocarcinoma. J. Exp. Clin. Cancer Res. 43, 271 (2024).
Cao, J. et al. Effects of altered glycolysis levels on CD8+ T cell activation and function. Cell Death Dis. 14, 407 (2023).
Liu, W. et al. Lactate regulates cell cycle by remodelling the anaphase promoting complex. Nature 616, 790–797 (2023).
Pagès, F. et al. Effector memory T cells, early metastasis, and survival in colorectal cancer. N. Engl. J. Med. 353, 2654–2666 (2005).
Han, J., Khatwani, N., Searles, T. G., Turk, M. J. & Angeles, C. V. Memory CD8+ T cell responses to cancer. Semin. Immunol. 49, 101435 (2020).
Zhang, L. et al. Single-cell analyses inform mechanisms of myeloid-targeted therapies in colon cancer. Cell 181, 442–459 (2020).
Buck, M. D., Sowell, R. T., Kaech, S. M. & Pearce, E. L. Metabolic instruction of immunity. Cell 169, 570–586 (2017).
Li, X. et al. Navigating metabolic pathways to enhance antitumour immunity and immunotherapy. Nat. Rev. Clin. Oncol. 16, 425–441 (2019).
Wang, F. et al. ZFP91 disturbs metabolic fitness and antitumor activity of tumor-infiltrating T cells. J. Clin. Invest. 131, e144318 (2021).
Wherry, E. J. & Kurachi, M. Molecular and cellular insights into T cell exhaustion. Nat. Rev. Immunol. 15, 486–499 (2015).
Tay, C., Tanaka, A. & Sakaguchi, S. Tumor-infiltrating regulatory T cells as targets of cancer immunotherapy. Cancer Cell 41, 450–465 (2023).
Le, D. T. et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357, 409–413 (2017).
Chalabi, M. et al. Neoadjuvant immunotherapy leads to pathological responses in MMR-proficient and MMR-deficient early-stage colon cancers. Nat. Med. 26, 566–576 (2020).
Chen, Y. et al. Spatiotemporal single-cell analysis decodes cellular dynamics underlying different responses to immunotherapy in colorectal cancer. Cancer Cell 42, 1268–1285 (2024).
Liu, X. et al. Inhibition of PCSK9 potentiates immune checkpoint therapy for cancer. Nature 588, 693–698 (2020).
Kaech, S. M. & Cui, W. Transcriptional control of effector and memory CD8+ T cell differentiation. Nat. Rev. Immunol. 12, 749–761 (2012).
Henning, A. N., Roychoudhuri, R. & Restifo, N. P. Epigenetic control of CD8+ T cell differentiation. Nat. Rev. Immunol. 18, 340–356 (2018).
Chang, C.-H. et al. Posttranscriptional control of T cell effector function by aerobic glycolysis. Cell 153, 1239–1251 (2013).
Chang, C.-H. et al. Metabolic competition in the tumor microenvironment is a driver of cancer progression. Cell 162, 1229–1241 (2015).
Ho, P.-C. et al. Phosphoenolpyruvate Is a metabolic checkpoint of anti-tumor T cell responses. Cell 162, 1217–1228 (2015).
Scharping, N. E. et al. The tumor microenvironment represses T cell mitochondrial biogenesis to drive intratumoral T cell metabolic insufficiency and dysfunction. Immunity 45, 374–388 (2016).
Yu, Y. R. et al. Disturbed mitochondrial dynamics in CD8+ TILs reinforce T cell exhaustion. Nat. Immunol. 21, 1540–1551 (2020).
Dahl, J. A. & Collas, P. A rapid micro chromatin immunoprecipitation assay (microChIP). Nat. Protoc. 3, 1032–1045 (2008).
Galmozzi, A. et al. PGRMC2 is an intracellular haem chaperone critical for adipocyte function. Nature 576, 138–142 (2019).
Liu, Q. et al. Non-oxidative pentose phosphate pathway controls regulatory T cell function by integrating metabolism and epigenetics. Nat. Metab. 4, 559–574 (2022).
Acknowledgements
We thank members of the laboratory of X.T. and B.L. for their suggestions. This work was supported by the National Key Research and Development Program of China (grant no. 2019YFA0906100 to B.L. and X.T.) and the National Natural Science Foundation of China (grant nos. 82330080, 82425042 and 32371361 to X.T., grant nos. 82003012, 82273235 to Y.L. and grant no. 82173002 to L.W.). We appreciate the support from Core Facility of Basic Medical Sciences, Shanghai Jiao Tong University School of Medicine. We are grateful for the support from the Innovative research team of high-level local universities in Shanghai (grant no. SHSMU-ZDCX20210902 to R.J. and X.T. and grant no. SHSMU-ZDCX20210601 to B.L. and Y.L.) and Shanghai Frontiers Science Center of Cellular Homeostasis and Human Diseases. We thank F. Zhu for valuable discussions and scRNA-seq data analysis. The Extended Data Fig. 7g was created with BioRender.com.
Author information
Authors and Affiliations
Contributions
N.X., R.J., B.L., Y.L. and X.T. designed the research; N.X., Y.L., Y.Z. and Y.H. performed experiments; Q.L., Y.L., S.S., W.H., J.H., Y.W. and S.Q. contributed to mouse and cellular experiments; L.T. and Z.C. contributed to metabolic analysis; L.W., P.Z., H.L., and J.C. contributed to scientific discussion; Y.Z. and M.L. performed and interpreted bioinformatic analyses; P.C., P.D. and W.Z. provided patient samples; N.X. and Y.Z. performed data analysis and interpretation; N.X., Y.Z., Y.L. and X.T. wrote the manuscript. All authors have read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
B.L. is a co-founder of Biotheus and chairman of its scientific advisory board. X.T. has a patent related to the MondoA–TXNIP therapy in cancers. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Ilaria Elia, Guideng Li and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Alfredo Giménez-Cassina in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Lactic acid upregulates TXNIP and ARRDC4 levels in CD8+ T cells and Treg cells, related to Fig. 1.
(a) The table of all differentially expressed genes of RNA-seq in both CD8+ T cells and Treg cells between the control and lactic acid-treated groups. (b, c) Volcano plot of CD8+ T cells (b) and Treg cells (c) treated with or without 10 mM lactic acid. (d, e) QPCR (d, n = 3 independent samples) and western blotting (e) analysis of human CD8+ T cells activated by anti-CD3/CD28 antibodies and treated with vehicle control, 10 mM lactic acid or 10 mM sodium lactate. (f-h) Flow cytometric and quantitative analysis of TNF (f), IFN-γ, GZMB (g) and IL-2 (h) expression of activated mouse CD8+ T cells treated with vehicle control, 10 mM lactic acid or 10 mM sodium lactate. (i) QPCR analysis of function-associated gene expression in human CD8+ T cells treated with or without 10 mM lactic acid (n = 3 independent samples). (j) QPCR analysis of function-associated gene expression in human iTreg cells treated with or without 10 mM lactic acid (n = 3 independent samples). (k) Flow cytometric and quantitative analysis of Treg suppressive capacity using in vitro suppression assays. Treg cells and CTV-labelled mouse CD8+ T cells were co-cultured for 48 h and treated with or without 5 mM lactic acid for 24 h (control: n = 4 cultures, lactic acid: n = 3 cultures). The data were representative of three independent experiments and presented as mean ± s.e.m. Statistical significance was calculated by the unpaired twotailed Student’s ttest (d and f-h) and two-way ANOVA followed by Sidak’s multiple comparisons test (i-k). ns, not significant.
Extended Data Fig. 2 The MondoA-TXNIP axis plays a critical role in lactic acid-reduced CD8+ T cell cytokine production.
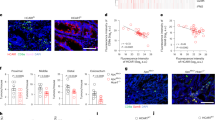
(a-d) ChIP analysis of MondoA occupancy at the TXNIP and ARRDC4 promoters in activated human CD8+ T cells (a, b) and Treg cells (c, d) (n = 3 cultures). (e) Immunofluorescent analysis of subcellular localization of MondoA in human CD8+ T cells treated with or without 10 mM lactic acid. Representative merged images of MondoA (red) and DAPI (blue) were shown. Scale bars, 7.5 μm. (f) Quantification of the colocalization of DAPI and MondoA in Extended Data Fig. 2e using Pearson’s correlation coefficients (n = 7 independent samples). (g, h) QPCR (g, n = 3 mice) and western blotting (h) analysis of mRNA and protein levels of MondoA and TXNIP in T cells of MondoAfl/fl and MondoAfl/flCd4-Cre mice. The data were representative of three independent experiments and presented as mean ± s.e.m. Statistical significance was calculated by the unpaired twotailed Student’s ttest (a-d, f) and two-way ANOVA followed by Sidak’s multiple comparisons test (g).
Extended Data Fig. 3 MondoA ablation restores glycolysis and oxidative phosphorylation disturbed by lactic acid in CD8+ T cells, related to Fig. 4.
(a, b) GSEA of glycolysis (a) and oxidative phosphorylation (b) between WT and MondoA-deficient CD8+ T cells. (c) Western blot analysis of GLUT1 of WT and MondoA-deficient CD8+ T cells. (d) 2-NBDG uptake by flow cytometric analysis and quantitative results of the MFI of 2-NBDG in WT and MondoA-deficient CD8+ T cells. (e, f) Glycolytic rate (e) and glycolytic capacity (f) of CD8+ T cells isolated from MondoAfl/fl and MondoAfl/flCd4-Cre mice. (g-j) Analysis of glucose consumption (g), lactic acid production (h), maximal respiration (i) and spare respiratory capacity (j) in CD8+ T cells isolated from MondoAfl/fl or MondoAfl/flCd4-Cre mice and stimulated with anti-CD3/CD28 antibodies for 36 h. (k, l) GSEA of glycolysis (k) and oxidative phosphorylation (l) of CD8+ T cells treated with or without 10 mM lactic acid for 24 h. (m) Quantification of mitochondrial numbers of WT and MondoA-deficient CD8+ T cells with or without 10 mM lactic acid treatment (WT-Ctrl n = 23 independent samples, WT-LA n = 22 independent samples, KO-Ctrl n = 29 independent samples, KO-LA n = 25 independent samples). (n, o) Quantification of number (n) (WT-Ctrl n = 19 independent samples, WT-LA n = 26 independent samples, KO-Ctrl n = 34 independent samples, KO-LA n = 26 independent samples) and maximal width (o) (n = 30 independent samples) of mitochondrial cristae in WT and MondoA-deficient CD8+ T cells treated with or without 10 mM lactic acid for 24 h. The data were representative of three independent experiments and presented as mean ± s.e.m. Statistical analyses were determined by unpaired twotailed Student’s ttest.
Extended Data Fig. 4 Deletion of MondoA shows minimal effect in T cell homeostasis, related to Fig. 5.
(a) Flow cytometric gating strategy of TCRβ+CD8+ T cells, TCRβ+CD4+ T cells and TCRβ+CD4+FOXP3+ Treg cells in the spleen of mice. (b-d) Flow cytometric analysis of proportions of CD4+ T cells (b), CD8+ T cells (c) and Treg cells (d) in the spleen, mesenteric and peripheral lymph nodes (mLNs and pLNs) from 8-week-old WT and MondoAfl/flCd4-Cre mice. (e, f) Flow cytometric analysis of the percentage of CD4− CD8− (DN), CD4+ CD8+ (DP), CD4+ CD8− (SP-CD4) and CD4− CD8+ (SP-CD8) in the thymus of 8-week-old WT and MondoAfl/flCd4-Cre mice. (g, h) Expression of CD62L and CD44 in CD4+ T cells (g) and CD8+ T cells (h) in the spleen, mLNs and pLNs of 8-week-old WT and MondoAfl/flCd4-Cre mice. (i, j) Flow cytometric analysis of the percentage of Ki67-expressing CD4+ T cells (i) and CD8+ T cells (j) in the spleen, mLNs and pLNs from 8-week-old WT and MondoAfl/flCd4-Cre mice. (k) Flow cytometric analysis of the percentage of IFN-γ and TNF producing CD8+ T cells in the spleen, mLNs and pLNs from 8-week-old WT and MondoAfl/flCd4-Cre mice. The data were representative of three independent experiments and presented as mean ± s.e.m. Statistical analyses were performed using two-way ANOVA followed by Sidak’s multiple comparisons test (b-k). ns, not significant.
Extended Data Fig. 5 MondoA ablation potentiates tumor-infiltrating CD8+ T cell antitumor immunity in vivo, related to Fig. 5.
(a, b) Flow cytometric (a) and quantitative (b) analysis of the percentage of CD4+ T cells and CD8+ T cells from the dLNs of MondoAfl/fl and MondoAfl/flCd4-Cre mice injected subcutaneously with MC38 colorectal cancer cells (n = 6 mice). (c-f) Flow cytometric (c, d) and quantitative (e, f) analysis of IFN-γ and TNF-α production by CD4+ and CD8+ T cells in the dLNs of MondoAfl/fl (n = 6 mice) and MondoAfl/flCd4-Cre (n = 8 mice) mice injected subcutaneously with MC38 colorectal cancer cells. (g, h) Flow cytometric (g) and quantitative (h) analysis of IFN-γ-producing and TNF-producing CD45.2+ T cells in dLNs of CD45.1 mice injected subcutaneously with MC38-OVA colorectal cancer cells and adoptively transferred with MondoAfl/flCd4-Cre OT-I and MondoAfl/fl OT-I CD8+ T cells (n = 4 mice). The data were representative of three independent experiments and presented as mean ± s.e.m. Statistical significance was calculated by the two-way ANOVA followed by Sidak’s multiple comparisons test. ns, not significant.
Extended Data Fig. 6 SBI-477 reinforces CD8+ T cell antitumor immunity, related to Fig. 6.
(a-c) Flow cytometric (a) and quantitative (b, c) analysis of IFN-γ and TNF of activated mouse CD8+ T cells treated with the vehicle control or SBI-477. (d) CD8+ effector T cells isolated from OT-I mice were treated with or without SBI-477, incubated with MC38-OVA cells for 24 h, and LDH in culture was measured to assess cytotoxicity. (e) QPCR analysis of Txnip and Arrdc4 levels in WT and MondoA-deficient activated CD8+ T cells treated with or without SBI-477. (f, g) QPCR analysis of MondoA and Txnip mRNA levels in tumors (f) and skeletal muscles (g) from MC38 tumor-bearing mice treated with or without SBI-477 (n = 3 mice). (h, i) Flow cytometric (h) and quantitative (i) analysis of 2-NBDG levels in tumor-infiltrating CD8+ T cells from MC38 tumor-bearing mice treated with or without SBI-477 (control: n = 7 mice, SBI-477: n = 6 mice). (j, l) Flow cytometric analysis of the percentage of IFN-γ and TNF production by CD8+ T cells in the dLNs (j) or spleen (l) from MC38 tumor-bearing mice treated with or without SBI-477. (k, m) Quantification of the percentage of T cells, CD4+ and CD8+ T cells as well as IFN-γ and TNF production by CD4+ and CD8+ T cells in the dLNs (k) or spleen (m) from MC38 tumor-bearing mice treated with or without SBI-477 (n = 3 mice). (n) Scheme for treating KPC1199 tumor-bearing mice with or without SBI-477 and representative tumor images. (o, p) Tumor growth (o) and tumor weight (p) of KPC1199 tumor-bearing mice treated with or without SBI-477 (n = 7 mice). (q, r) Flow cytometric (q) and quantitative analysis (r) of GZMB levels in tumor-infiltrating CD8+ T cells from KPC1199 tumor-bearing mice treated with or without SBI-477 (n = 7 mice). (s) CCK-8 assays for assessing the effect of SBI-477 on MC38 cell proliferation (n = 6 independent samples). The data are representative of three independent experiments and are presented as mean ± s.e.m. Statistical significance was calculated by two-way ANOVA followed by Sidak’s multiple comparisons test (f, g, k, m and o) and by Tukey’s multiple comparisons (e) or unpaired twotailed Student’s ttest (b, c, d, i, p and r). ns, not significant.
Extended Data Fig. 7 MondoA inhibition improves cancer immunotherapy in anti-PD-1-resistant tumors, related to Fig. 7.
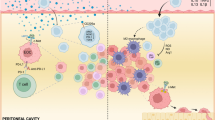
(a-c) Levels of TXNIP mRNA in tumor-infiltrating CD8+ T cells and Treg cells from MSS and MSI colorectal cancer patients in dataset GSE236581. (d-f) Levels of TXNIP mRNA in tumor-infiltrating CD8+ T cells and Treg cells from colorectal cancer patients responsive or non-responsive to the anti-PD-1 antibody in dataset GSE236581. (g) The MondoA-TXNIP axis plays a critical role in lactic acid-induced immunosuppressive tumor microenvironment by regulating the balance between CD8+ T cells and Treg cells. Lactic acid increases TXNIP and ARRDC4 expression by promoting MondoA transcriptional activity in CD8+ T cells and Treg cells and impairs the balance between CD8+ T cells and Treg cells. In the high lactic acid microenvironment, targeting the MondoA-TXNIP axis promotes GLUT1 expression and glycolysis, enhances CD8+ T cell cytotoxicity and attenuates Treg suppressive function, therefore potentiating antitumor immune responses. This figure was created in BioRender. Statistical analyses were determined by two-tailed Wilcoxon rank-sum test (a-f).
Supplementary information
Supplementary Information (download PDF )
Supplementary Table 1.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 1 (download PDF )
Unprocessed western blots.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download PDF )
Unprocessed western blots.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1/Table 1 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 1/Table 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2/Table 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2/Table 2 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 3/Table 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3/Table 3 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 4/Table 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5/Table 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6/Table 6 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, N., Zhu, Y., Han, Y. et al. Targeting MondoA–TXNIP restores antitumour immunity in lactic-acid-induced immunosuppressive microenvironment. Nat Metab 7, 1889–1904 (2025). https://doi.org/10.1038/s42255-025-01347-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42255-025-01347-1
This article is cited by
-
SENP1 drives glycolysis and cisplatin resistance in gastric cancer via desumoylating ENO1
Journal of Experimental & Clinical Cancer Research (2025)
-
Dual gains from targeting MondoA in T cells
Nature Metabolism (2025)