Abstract
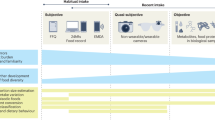
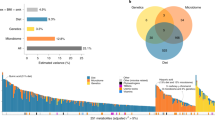
Understanding how diet shapes metabolism across diverse populations is essential to improving nutrition and health. Biomarkers reflecting diet are explored largely in European and American populations, but the food metabolome is highly complex and varies across region and culture. We assessed 1,055 plasma metabolites and 169 foods/beverages in 8,391 multi-ethnic Asian individuals and carried out diet–metabolite association analyses. Using machine learning, we developed multi-biomarker panels and composite scores for key foods, beverages and overall diet quality. Here we show these biomarker panels can be used to objectively assess dietary intakes in the Asian multi-ethnic population and can explain variances in intake prediction models better than single biomarkers. The identified diet–metabolite relationships are reproducible over time and improve prediction of clinical outcomes (insulin resistance, diabetes, body mass index, carotid intima-media thickness and hypertension), compared to self-reports. Our findings show insights into multi-ethnic diet-related metabolic variations and an opportunity to link exposure to population health outcomes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The HELIOS study individual-level datasets are not publicly available due to data privacy regulations. Researchers may apply for access through HELIOS Data Access Committee (helios_science@ntu.edu.sg). Summary-level datasets are available as source data including food–metabolite and metabolite–metabolite partial correlation coefficients, correlation coefficients between self-report against estimated intakes, ICCs from the reproducibility study, and regression coefficients relating self-reports and metabolite scores with clinical phenotypes. Source data are provided with this paper.
Code availability
Data processing, analysis and visualization were performed using R (v4.5.0) and the following packages: tidyverse (v1.3.2), readxl (v1.4.1), glue (v1.6.2), pcor (v1.1), glmnet (v4.1-7), stats (v4.2.1), irr (v0.84.1), ggplot2 (v3.4.2) and gplots (v3.1.3). Codes for analyses and visualization are available via GitHub at https://github.com/HELIOS-SG100K-LKC/Metabolic-variation-reflects-dietary-intake-in-a-multi-ethnic-Asian-population/.
References
World Health Organisation. Cardiovascular diseases (CVDs): key facts. https://www.who.int/news-room/fact-sheets/detail/cardiovascular-diseases (11 June 2021).
Barouki, R., Gluckman, P. D., Grandjean, P., Hanson, M. & Heindel, J. J. Developmental origins of non-communicable disease: Implications for research and public health. Environ. Health 11, 42 (2012).
Afshin, A. et al. Health effects of dietary risks in 195 countries, 1990-2017: a systematic analysis for the Global Burden of Disease study 2017. Lancet 393, 1958–1972 (2019).
Uusitalo, U., Pietinen, P. & Puska, P. Globalization, Diets and Noncommunicable Diseases. Dietary Transition in Developing Countries: Challenges for Chronic Disease Prevention (World Health Organisation, 2002); https://iris.who.int/bitstream/handle/10665/42609/9241590416.pdf
Xu, Y. Y. et al. The Global Burden of Disease attributable to low physical activity and its trends from 1990 to 2019: an analysis of the Global Burden of Disease study. Front. Public Health 10, 1018866 (2022).
Brennan, L., Hu, F. B. & Sun, Q. Metabolomics meets nutritional epidemiology: harnessing the potential in metabolomics data. Metabolites 11, 709 (2021).
Agudo, A. Measuring Intake of Fruit and Vegetables (World Health Organisation, 2005); https://iris.who.int/bitstream/handle/10665/43144/9241592826_eng.pdf?sequence=1
Scalbert, A. et al. The food metabolome: a window over dietary exposure. Am. J. Clin. Nutr. 99, 1286–1308 (2014).
Barabási, A.-L., Menichetti, G. & Loscalzo, J. The unmapped chemical complexity of our diet. Nat. Food 1, 33–37 (2020).
Landberg, R. et al. Future prospects for dissecting inter-individual variability in the absorption, distribution and elimination of plant bioactives of relevance for cardiometabolic endpoints. Eur. J. Nutr. 58, 21–36 (2019).
Manach, C. et al. Addressing the inter-individual variation in response to consumption of plant food bioactives: towards a better understanding of their role in healthy aging and cardiometabolic risk reduction. Mol. Nutr. Food Res. 61, 1600557 (2017).
Rothwell, J. A. et al. New biomarkers of coffee consumption identified by the non-targeted metabolomic profiling of cohort study subjects. PLoS ONE 9, e93474 (2014).
Gibbons, H. et al. Demonstration of the utility of biomarkers for dietary intake assessment; proline betaine as an example. Mol. Nutr. Food Res. 61, 1700037 (2017).
Vázquez-Manjarrez, N. et al. Discovery and validation of banana intake biomarkers using untargeted metabolomics in human intervention and cross-sectional studies. J. Nutr. 149, 1701–1713 (2019).
Li, C. et al. Development and validation of a metabolite score for red meat intake: an observational cohort study and randomized controlled dietary intervention. Am. J. Clin. Nutr. 116, 511–522 (2022).
Rothwell, J. A. et al. A metabolomic study of biomarkers of habitual coffee intake in four European countries. Mol. Nutr. Food Res. 63, 1900659 (2019).
Prevention, Pesticides and Toxic Substances (7508C) (United States Environmental Protection Agency, 1999).
Sri Harsha, P. S. C. et al. Biomarkers of legume intake in human intervention and observational studies: a systematic review. Genes Nutr. 13, 25 (2018).
Perera, T. et al. Identification and monitoring of metabolite markers of dry bean consumption in parallel human and mouse studies. Mol. Nutr. Food Res. 59, 795–806 (2015).
Roca-Saavedra, P. et al. Phytanic acid consumption and human health, risks, benefits and future trends: a review. Food Chem. 221, 237–247 (2017).
Wang, Y., Hodge, R. A., Stevens, V. L., Hartman, T. J. & McCullough, M. L. Identification and reproducibility of plasma metabolomic biomarkers of habitual food intake in a US diet validation study. Metabolites 10, 382 (2020).
Cuparencu, C. et al. Biomarkers of meat and seafood intake: an extensive literature review. Genes Nutr. 14, 35 (2019).
Cuparencu, C. et al. The anserine to carnosine ratio: an excellent discriminator between white and red meats consumed by free-living overweight participants of the PREVIEW study. Eur. J. Nutr. 60, 179–192 (2021).
Hanhineva, K. et al. Nontargeted metabolite profiling discriminates diet-specific biomarkers for consumption of whole grains, fatty fish, and bilberries in a randomized controlled trial. J. Nutr. 145, 7–17 (2015).
Rafiq, T. et al. Nutritional metabolomics and the classification of dietary biomarker candidates: a critical review. Adv. Nutr. 12, 2333–2357 (2021).
McCullough, M. L. et al. Metabolomic markers of healthy dietary patterns in US postmenopausal women. Am. J. Clin. Nutr. 109, 1439–1451 (2019).
Low, D. Y. et al. Diet-related metabolites associated with cognitive decline revealed by untargeted metabolomics in a prospective cohort. Mol. Nutr. Food Res. 63, 1900177 (2019).
Cheah, I. K., Tang, R. M. Y., Yew, T. S. Z., Lim, K. H. C. & Halliwell, B. Administration of pure ergothioneine to healthy human subjects: uptake, metabolism, and effects on biomarkers of oxidative damage and inflammation. Antioxid. Redox Signal. 26, 193–206 (2017).
Weigand-Heller, A. J., Kris-Etherton, P. M. & Beelman, R. B. The bioavailability of ergothioneine from mushrooms (Agaricus bisporus) and the acute effects on antioxidant capacity and biomarkers of inflammation. Prev. Med. 54, S75–S78 (2012).
Kubec, R., Svobodová, M. & Velíšek, J. Gas-chromatographic determination of S-methylcysteine sulfoxide in cruciferous vegetables. Eur. Food Res. Technol. 213, 386–388 (2001).
Pujos-Guillot, E. et al. Mass spectrometry-based metabolomics for the discovery of biomarkers of fruit and vegetable intake: citrus fruit as a case study. J. Proteome Res. 12, 1645–1659 (2013).
Lloyd, A. J., Beckmann, M., Favé, G., Mathers, J. C. & Draper, J. Proline betaine and its biotransformation products in fasting urine samples are potential biomarkers of habitual citrus fruit consumption. Br. J. Nutr. 106, 812–824 (2011).
Playdon, M. C. et al. Comparing metabolite profiles of habitual diet in serum and urine. Am. J. Clin. Nutr. 104, 776–789 (2016).
Guertin, K. A. et al. Metabolomics in nutritional epidemiology: identifying metabolites associated with diet and quantifying their potential to uncover diet-disease relations in populations. Am. J. Clin. Nutr. 100, 208–217 (2014).
Irwig, M. S., El-Sohemy, A., Baylin, A., Rifai, N. & Campos, H. Frequent intake of tropical fruits that are rich in beta-cryptoxanthin is associated with higher plasma beta-cryptoxanthin concentrations in Costa Rican adolescents. J. Nutr. 132, 3161–3167 (2002).
Burri, B., La Frano, M. R. & Zhu, C. Absorption, metabolism, and functions of β-cryptoxanthin. Nutr. Rev. 74, 69–82 (2016).
Hanhineva, K. et al. Qualitative characterization of benzoxazinoid derivatives in whole grain rye and wheat by LC-MS metabolite profiling. J. Agric. Food Chem. 59, 921–927 (2011).
Meng, L. et al. Occurrence, temporal variation (2010–2018), distribution, and source appointment of per- and polyfluoroalkyl substances (PFAS) in mollusks from the Bohai Sea, China. ACS EST Water 2, 195–205 (2022).
Pallister, T. et al. Characterizing blood metabolomics profiles associated with self-reported food intakes in female twins. PLoS ONE 11, e0158568 (2016).
Gorgani, L., Mohammadi, M., Najafpour, G. D. & Nikzad, M. Piperine—the bioactive compound of black pepper: from isolation to medicinal formulations. Compr. Rev. Food Sci. Food Saf. 16, 124–140 (2017).
Praticò, G., Gao, Q., Manach, C. & Dragsted, L. O. Biomarkers of food intake for Allium vegetables. Genes Nutr. 13, 34 (2018).
Chietra, W. Forms, Analysis and Stability of Selenium and Vitamin E in Flour and Asian Noodles. PhD thesis, RMIT Univ. (2015).
Schmidt, C. et al. Lactoyl leucine and isoleucine are bioavailable alternatives for canonical amino acids in cell culture media. Biotechnol. Bioeng. 118, 3395–3408 (2021).
Ulaszewska, M. et al. Food intake biomarkers for apple, pear, and stone fruit. Genes Nutr. 13, 29 (2018).
Fung, T. T. et al. Diet-quality scores and plasma concentrations of markers of inflammation and endothelial dysfunction. Am. J. Clin. Nutr. 82, 163–173 (2005).
Chiuve, S. E. et al. Alternative dietary indices both strongly predict risk of chronic disease. J. Nutr. 142, 1009–1018 (2012).
Appel, L. J. et al. A clinical trial of the effects of dietary patterns on blood pressure. DASH Collaborative Research Group. N. Engl. J. Med. 336, 1117–1124 (1997).
Bromage, S. et al. Development and validation of a novel food-based global diet quality score (GDQS). J. Nutr. 151, 75s–92s (2021).
Satija, A. et al. Healthful and unhealthful plant-based diets and the risk of coronary heart disease in U.S. adults. J. Am. Coll. Cardiol. 70, 411–422 (2017).
Sadhu, N. et al. Metabolome-wide association identifies ferredoxin-1 (FDX1) as a determinant of cholesterol metabolism and cardiovascular risk in Asian populations. Nat. Cardiovasc. Res. 4, 567–583 (2025).
Nieuwenhuijse, H. in Encyclopedia of Dairy Sciences 3rd edn (eds Paul L. H. M. et al.) 549–554 (Academic Press, 2022).
Pantzaris, T. P. & Ahmad, M. J. Properties and Utilization of Palm Kernel Oil. Palm Oil Developments 35 (Malaysian Palm Oil Board, 2001).
Dos Santos, N. J. A. et al. Effect of dietary palm kernel oil on the quality, fatty acid profile, and sensorial attributes of young bull meat. Foods https://doi.org/10.3390/foods11040609 (2022).
Pan, A. et al. Red meat consumption and risk of type 2 diabetes: 3 cohorts of US adults and an updated meta-analysis. Am. J. Clin. Nutr. 94, 1088–1096 (2011).
Liu, G. et al. Cooking methods for red meats and risk of type 2 diabetes: a prospective study of U.S. women. Diabetes Care 40, 1041–1049 (2017).
Field, B. C., Gordillo, R. & Scherer, P. E. The role of ceramides in diabetes and cardiovascular disease regulation of ceramides by adipokines. Front. Endocrinol. 11, 569250 (2020).
Rebholz, C. M., Lichtenstein, A. H., Zheng, Z., Appel, L. J. & Coresh, J. Serum untargeted metabolomic profile of the Dietary Approaches to Stop Hypertension (DASH) dietary pattern. Am. J. Clin. Nutr. 108, 243–255 (2018).
Gadgil, M. D. et al. Metabolomic profile of the Healthy Eating Index-2015 in the multiethnic study of atherosclerosis. J. Nutr. https://doi.org/10.1016/j.tjnut.2023.05.030 (2023).
Hannun, Y. A. & Obeid, L. M. Sphingolipids and their metabolism in physiology and disease. Nat. Rev. Mol. Cell Biol. 19, 175–191 (2018).
Wu, G. Amino acids: metabolism, functions, and nutrition. Amino Acids 37, 1–17 (2009).
Brennan, L. & de Roos, B. Role of metabolomics in the delivery of precision nutrition. Redox Biol. 65, 102808 (2023).
Eichelmann, F. et al. Lipidome changes due to improved dietary fat quality inform cardiometabolic risk reduction and precision nutrition. Nat. Med. https://doi.org/10.1038/s41591-024-03124-1 (2024).
Wang, X. et al. The Health for Life in Singapore (HELIOS) study: delivering precision medicine research for Asian populations. Preprint at medRxiv https://doi.org/10.1101/2024.05.14.24307259 (2024).
Association, A. D. Classification and diagnosis of diabetes: standards of medical care in diabetes—2021. Diabetes Care 44, S15–S33 (2020).
HEARTS D: Diagnosis and Management of Type 2 Diabetes (World Health Organization, 2020); https://www.who.int/publications/i/item/who-ucn-ncd-20.1
Unger, T. et al. 2020 International Society of Hypertension Global Hypertension Practice Guidelines. Hypertension 75, 1334–1357 (2020).
Whitton, C. et al. Relative validity and reproducibility of a food frequency questionnaire for assessing dietary intakes in a multi-ethnic asian population using 24-h dietary recalls and biomarkers. Nutrients https://doi.org/10.3390/nu9101059 (2017).
Whitton, C., Rebello, S. A., Lee, J., Tai, E. S. & van Dam, R. M. A healthy Asian a posteriori dietary pattern correlates with a priori dietary patterns and is associated with cardiovascular disease risk factors in a multiethnic Asian population. J. Nutr. 148, 616–623 (2018).
Fung, T. T. et al. Adherence to a DASH-style diet and risk of coronary heart disease and stroke in women. Arch. Intern. Med. 168, 713–720 (2008).
Disease Burden (Ministry of Health Singapore, 2023); https://www.moh.gov.sg/others/resources-and-statistics/disease-burden
Ford, L. et al. Precision of a clinical metabolomics profiling platform for use in the identification of inborn errors of metabolism. J. Appl. Lab. Med. 5, 342–356 (2020).
Evans, A. M., DeHaven, C. D., Barrett, T., Mitchell, M. & Milgram, E. Integrated, nontargeted ultrahigh performance liquid chromatography/electrospray ionization tandem mass spectrometry platform for the identification and relative quantification of the small-molecule complement of biological systems. Anal. Chem. 81, 6656–6667 (2009).
Evans, A. M. et al. High resolution mass spectrometry improves data quantity and quality as compared to unit mass resolution mass spectrometry in high-throughput profiling metabolomics. Metabolomics https://doi.org/10.4172/2153-0769.1000132 (2014).
Sumner, L. W. et al. Proposed minimum reporting standards for chemical analysis. Metabolomics 3, 211–221 (2007).
Zou, H. & Hastie, T. Regularization and variable selection via the elastic net. J. R. Stat. Soc. B 67, 301–320 (2005).
Mbatchou, J. et al. Computationally efficient whole-genome regression for quantitative and binary traits. Nat. Genet. 53, 1097–1103 (2021).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Sollis, E. et al. The NHGRI-EBI GWAS Catalog: knowledgebase and deposition resource. Nucleic Acids Res. https://doi.org/10.1093/nar/gkac1010 (2022).
Szklarczyk, D. et al. STRING v11: protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–D613 (2019).
Acknowledgements
We thank the participants of the HELIOS study and the HELIOS operation team for recruitment, organization and data/sample collection. This study (NTU IRB: 2016-11-030) is supported by Singapore Ministry of Health’s (MOH) National Medical Research Council (NMRC) under its OF-LCG funding scheme (MOH-000271) and CS-IRG funding scheme (MOH-001708), NMRC through National Cohorts Office (P2022-01-03) and National Research Foundation, Singapore, through NMRC and the Precision Health Research, Singapore (PRECISE) under the National Precision Medicine programme (NMRC/PRECISE/2020) and intramural funding from Nanyang Technological University, Lee Kong Chian School of Medicine and National Healthcare Group.
Author information
Authors and Affiliations
Contributions
J.C.C., J.D.B., P.E., E.R., J.L., E.S.L. and J.N. acquired funding for the HELIOS study. J.C.C., K.E.W., P.A.S., R.S. and G.A.M. acquired funding for metabolomics analysis. J.C.C. and D.Y.L. conceptualized the study. D.Y.L., T.H.M., N.S., P.R.J., R.D., K.E.W., P.A.S., R.S. and G.A.M. generated the data. D.Y.L., T.H.M., N.S. and P.R.J. analysed the data. J.C.C. and D.Y.L. wrote the initial manuscript draft. M.L., T.H.M., N.S., P.R.J. and P.A.S. provided critical feedback to the manuscript draft. All authors reviewed and contributed to revision of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
K.E.W. and G.A.M. are employees of Metabolon. The other authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Rui Wang-Sattler and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Yanina-Yasmin Pesch, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Indices of diet quality components within the HELIOS study population.
a) Sankey illustration of diet quality components. b) Radar plots showing distribution of individual components across diet quality indices. c) Density plots of diet quality scores across age and ethnicity. MUFA, monounsaturated fatty acids; PUFA, polyunsaturated fatty acids; SFA, saturated fatty acids; bev, beverage; proc, processed; HF, high fat; LF, low fat.
Extended Data Fig. 2 Model prediction performance of dietary intakes and diet quality.
a) Percent (%) error observed between self-reported and estimated intakes (log-transformed) of foods and beverages in respective test datasets (n = 2,336-2,513; refer to Supplementary Table 9 for exact sample sizes of test datasets). b) Density distribution of prediction error % of foods and beverages in respective test datasets with dotted line indicating median error and text insert indicating median prediction accuracy (formula: 1 - error %).
Extended Data Fig. 3 Summary of coffee and tea intakes.
a) Percent (%) of variations of total daily coffee and tea intakes recorded via FFQ in the study population. b) Proportion of kilocalories across the top 5 most consumed variations of coffee or tea.
Extended Data Fig. 4 Associations between environmental contaminants and clinical phenotypes.
Effect sizes of environmental contaminants (that comprise various dietary biomarker panels), adjusted for age, sex, ethnicity and batch. Estimates were obtained from independent linear regression (for HOMA-IR, FMI, cIMT) or logistic regression (for T2D, hypertension), based on the following sample sizes: T2D (n = 1,040 T2D, n = 7,351 non-T2D), hypertension (n = 2,411 with hypertension, n = 5,980 without hypertension). Association strength represents log-transformed P values. Circle border represents significant associations (P < 0.05). HOMA-IR, homeostatic model assessment for insulin resistance; T2D, type 2 diabetes; FMI, fat mass index; cIMT, carotid intima-media thickness.
Supplementary information
Supplementary Tables 1–10 (download XLSX )
Supplementary Table 1: List of 1,055 metabolites in the study population. Supplementary Table 2: Partial correlation coefficients of FFQ food/beverage items and metabolites (r ≥ 0.15 and FDR P < 0.05), adjusted for age, sex, ethnicity and batch. Supplementary Table 3: Effect sizes of metabolites (within the multi-biomarker panels) from associations with key foods and beverages, in regression models adjusted for age, sex, batch and ethnicity or genetic ancestry. Supplementary Table 4: Summary table of metabolite and genetic correlations for annotation of unknowns in the dietary biomarker panels. Supplementary Table 5: Summary of metabolic pathways enriched in dietary quality indices. Supplementary Table 6: Effect sizes of nutrients (% kcal) from associations with dietary quality indices. Supplementary Table 7: Effect sizes of dietary intakes (derived from self-report or metabolite score) from associations with clinical phenotypes. Supplementary Table 8: Effect sizes of diet quality (derived from self-report or metabolite score) from associations with clinical phenotypes. Supplementary Table 9: Sample sizes for training and test datasets of each food and beverage dataset. Supplementary Table 10. Effect sizes of independent metabolite–SNP associations above genome-wide significance (P < 5 × 10−8) and genes/loci in the top associations.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Table 1 (download XLSX )
Statistical source data.
Source Data Extended Data Table 2 (download XLSX )
Statistical source data.
Source Data Extended Data Table 3 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Low, D.Y., Mina, T.H., Sadhu, N. et al. Metabolic variation reflects dietary exposure in a multi-ethnic Asian population. Nat Metab 7, 1939–1954 (2025). https://doi.org/10.1038/s42255-025-01359-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42255-025-01359-x
This article is cited by
-
Adversity, adiposity, nutrition and metabolic well-being in multi-ethnic Asia
Nature Metabolism (2026)
-
The Health for Life in Singapore (HELIOS) Study: delivering precision medicine research for Asian populations
Nature Communications (2025)