Abstract
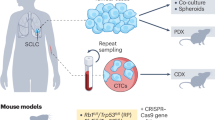
Small cell lung cancer (SCLC) is a devastating disease with limited therapeutic advancements. Although SCLC has recently been classified into four molecular subtypes, subtype-specific therapies are still lacking. Here, we established 40 patient-derived SCLC organoid lines with predominant TP53 and RB1 alterations and rare targetable genetic lesions. Transcriptome profiling divided the SCLC organoids into neuroendocrine (NE)-type SCLC and non-NE-type SCLC, with the latter characterized by YAP1 or POU2F3 expression. NE-type SCLC organoids grew independent of alveolar niche factors, whereas non-NE-type SCLC organoids relied on insulin-like growth factor (IGF)-1-driven YAP1 and AP1 activation. Therapeutic targeting of IGF-1, YAP1 and AP1 effectively suppressed the growth of non-NE-type organoids. Co-knockout of TP53 and RB1 in human alveolar cells altered their lineage toward the airway epithelium-like fate and conferred IGF-1 dependency, validating the subtype-phenotype connection. Our SCLC organoid library represents a valuable resource for developing biology-based therapies and has the potential to reshape the drug discovery landscape.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon request. Whole-exome sequencing, RNA-seq and ATAC-seq data were deposited in the DDBJ database under accession nos. JGAS000609 (study) and JGAD000738 (dataset) (https://humandbs.dbcls.jp/en/hum0357-v5). Source data are provided with this paper.
Code availability
No new code was developed in this study.
References
Dingemans, A.-M. C. et al. Small-cell lung cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 32, 839–853 (2021).
Rudin, C. M., Brambilla, E., Faivre-Finn, C. & Sage, J. Small-cell lung cancer. Nat. Rev. Dis. Primers 7, 3 (2021).
Ruano-Raviña, A., Provencio-Pulla, M. & Pérez-Ríos, M. Small cell lung cancer—a neglected disease with more data needed. JAMA Netw. Open 5, e224837 (2022).
Horn, L. et al. First-line atezolizumab plus chemotherapy in extensive-stage small-cell lung cancer. N. Engl. J. Med. 379, 2220–2229 (2018).
Paz-Ares, L. et al. Durvalumab plus platinum–etoposide versus platinum–etoposide in first-line treatment of extensive-stage small-cell lung cancer (CASPIAN): a randomised, controlled, open-label, phase 3 trial. Lancet 394, 1929–1939 (2019).
Ai, X. et al. Efficacy and safety of niraparib as maintenance treatment in patients with extensive-stage SCLC after first-line chemotherapy: a randomized, double-blind, phase 3 study. J. Thorac. Oncol. 16, 1403–1414 (2021).
Blackhall, F. et al. Efficacy and safety of rovalpituzumab tesirine compared with topotecan as second-line therapy in DLL3-High SCLC: results from the phase 3 TAHOE study. J. Thorac. Oncol. 16, 1547–1558 (2021).
Aix, S. P. et al. Combination lurbinectedin and doxorubicin versus physician’s choice of chemotherapy in patients with relapsed small-cell lung cancer (ATLANTIS): a multicentre, randomised, open-label, phase 3 trial. Lancet Respir. Med. 11, 74–86 (2023).
Peifer, M. et al. Integrative genome analyses identify key somatic driver mutations of small-cell lung cancer. Nat. Genet. 44, 1104–1110 (2012).
George, J. et al. Comprehensive genomic profiles of small cell lung cancer. Nature 524, 47–53 (2015).
Augustyn, A. et al. ASCL1 is a lineage oncogene providing therapeutic targets for high-grade neuroendocrine lung cancers. Proc. Natl Acad. Sci. USA 111, 14788–14793 (2014).
Borromeo, M. D. et al. ASCL1 and NEUROD1 reveal heterogeneity in pulmonary neuroendocrine tumors and regulate distinct genetic programs. Cell Rep. 16, 1259–1272 (2016).
Horie, M., Saito, A., Ohshima, M., Suzuki, H. I. & Nagase, T. YAP and TAZ modulate cell phenotype in a subset of small cell lung cancer. Cancer Sci. 107, 1755–1766 (2016).
Huang, Y.-H. et al. POU2F3 is a master regulator of a tuft cell-like variant of small cell lung cancer. Gene Dev. 32, 915–928 (2018).
Rudin, C. M. et al. Molecular subtypes of small cell lung cancer: a synthesis of human and mouse model data. Nat. Rev. Cancer 19, 289–297 (2019).
Poirier, J. T. et al. New approaches to SCLC therapy: from the laboratory to the clinic. J. Thorac. Oncol. 15, 520–540 (2020).
Caeser, R. et al. Genomic and transcriptomic analysis of a library of small cell lung cancer patient-derived xenografts. Nat. Commun. 13, 2144 (2022).
Simpson, K. L. et al. A biobank of small cell lung cancer CDX models elucidates inter- and intratumoral phenotypic heterogeneity. Nat. Cancer https://doi.org/10.1038/s43018-020-0046-2 (2020).
Pearsall, S. M. et al. The rare YAP1 subtype of small cell lung cancer revisited in a biobank of 39 circulating tumour cell patient derived explant models (CDX): a brief report. J. Thorac. Oncol. 15, 1836–1843 (2020).
Kim, M. et al. Patient-derived lung cancer organoids as in vitro cancer models for therapeutic screening. Nat. Commun. 10, 3991 (2019).
Choi, S. Y. et al. Establishment and long-term expansion of small cell lung cancer patient-derived tumor organoids. Int. J. Mol. Sci. 22, 1349 (2021).
Wang, H.-M. et al. Using patient-derived organoids to predict locally advanced or metastatic lung cancer tumor response: a real-world study. Cell Rep. Med. 4, 100911 (2023).
Baine, M. K. et al. Small cell lung carcinoma subtypes defined by ASCL1, NEUROD1, POU2F3 and YAP1: comprehensive immunohistochemical and histopathologic characterization. J. Thorac. Oncol. 15, 1823–1835 (2020).
Qu, S. et al. Molecular subtypes of primary small cell lung cancer tumors and their associations with neuroendocrine and therapeutic markers. J. Thorac. Oncol. 17, 141–153 (2021).
Szczepanski, A. P. et al. POU2AF2/C11orf53 functions as a coactivator of POU2F3 by maintaining chromatin accessibility and enhancer activity. Sci. Adv. 8, eabq2403 (2022).
Kawasaki, K. et al. An organoid biobank of neuroendocrine neoplasms enables genotype-phenotype mapping. Cell 183, 1420–1435.e21 (2020).
Lissa, D. et al. Heterogeneity of neuroendocrine transcriptional states in metastatic small cell lung cancers and patient-derived models. Nat. Commun. 13, 2023 (2022).
Zanconato, F. et al. Genome-wide association between YAP/TAZ/TEAD and AP-1 at enhancers drives oncogenic growth. Nat. Cell Biol. 17, 1218–1227 (2015).
Ebisudani, T. et al. Genotype-phenotype mapping of a patient-derived lung cancer organoid biobank identifies NKX2-1-defined Wnt dependency in lung adenocarcinoma. Cell Rep. 42, 112212 (2023).
Cerbinskaite, A., Mukhopadhyay, A., Plummer, E. R., Curtin, N. J. & Edmondson, R. J. Defective homologous recombination in human cancers. Cancer Treat. Rev. 38, 89–100 (2012).
Rigiracciolo, D. C. et al. Correction: Rigiracciolo, D.C. et al. IGF-1/IGF-1R/FAK/YAP transduction signaling prompts growth effects in triple-negative breast cancer (TNBC) cells. Cells 9, 2619 (2020).
Ohgushi, M., Minaguchi, M. & Sasai, Y. Rho-signaling-directed YAP/TAZ activity underlies the long-term survival and expansion of human embryonic stem cells. Cell Stem Cell 17, 448–461 (2015).
Ebisudani, T. et al. Direct derivation of human alveolospheres for SARS-CoV-2 infection modeling and drug screening. Cell Rep. 35, 109218 (2021).
Liu, Q. et al. Proteogenomic characterization of small cell lung cancer identifies biological insights and subtype-specific therapeutic strategies. Cell 187, 184–203.e28 (2024).
Soria, J.-C. et al. Osimertinib in untreated EGFR-mutated advanced non–small-cell lung cancer. N. Engl. J. Med. 378, 113–125 (2017).
Ito, T. et al. Loss of YAP1 defines neuroendocrine differentiation of lung tumors. Cancer Sci. 107, 1527–1538 (2016).
Wu, Q. et al. YAP drives fate conversion and chemoresistance of small cell lung cancer. Sci. Adv. 7, eabg1850 (2021).
Chen, P. et al. YAP1 expression is associated with survival and immunosuppression in small cell lung cancer. Cell Death Dis. 14, 636 (2023).
Chiappori, A. A. et al. A randomized phase II study of linsitinib (OSI‐906) versus topotecan in patients with relapsed small‐cell lung cancer. Oncol. 21, 1163–1164 (2016).
Sutherland, K. D. et al. Cell of origin of small cell lung cancer: inactivation of Trp53 and Rb1 in distinct cell types of adult mouse lung. Cancer Cell 19, 754–764 (2011).
Cui, M. et al. PTEN is a potent suppressor of small cell lung cancer. Mol. Cancer Res. 12, 654–659 (2014).
Park, J. W. et al. Reprogramming normal human epithelial tissues to a common, lethal neuroendocrine cancer lineage. Science 362, 91–95 (2018).
Gao, D. et al. Organoid cultures derived from patients with advanced prostate cancer. Cell 159, 176–187 (2014).
Tamagawa, H. et al. Wnt-deficient and hypoxic environment orchestrates squamous reprogramming of human pancreatic ductal adenocarcinoma. Nat. Cell Biol. 26, 1759–1772 (2024).
Fujii, M. et al. Human intestinal organoids maintain self-renewal capacity and cellular diversity in niche-inspired culture condition. Cell Stem Cell 23, 787–793.e6 (2018).
Togasaki, K. et al. Wnt signaling shapes the histologic variation in diffuse gastric cancer. Gastroenterology 160, 823–830 (2021).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Fujii, M. et al. A colorectal tumor organoid library demonstrates progressive loss of niche factor requirements during tumorigenesis. Cell Stem Cell 18, 827–838 (2016).
Islam, S. M. A. et al. Uncovering novel mutational signatures by de novo extraction with SigProfilerExtractor. Cell Genom. 2, 100179 (2022).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinform. 12, 323 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Wilkerson, M. D. & Hayes, D. N. ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics 26, 1572–1573 (2010).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9, R137 (2008).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Zerbino, D. R., Johnson, N., Juettemann, T., Wilder, S. P. & Flicek, P. WiggleTools: parallel processing of large collections of genome-wide datasets for visualization and statistical analysis. Bioinformatics 30, 1008–1009 (2014).
Manabe, T. et al. IGF2 autocrine-mediated IGF1R activation is a clinically relevant mechanism of osimertinib resistance in lung cancer. Mol. Cancer Res. 18, 549–559 (2020).
Ohta, Y. et al. Cell–matrix interface regulates dormancy in human colon cancer stem cells. Nature 608, 784–794 (2022).
Acknowledgements
We thank C. Yonekawa and M. Shibuya for technical assistance. This work was supported by the Japan Agency for Medical Research and Development (grant nos. 19cm0106206h0004, 20gm1210001, 21cm0106576h0002 and 23ck0106722h0002), by the JSPS KAKENHI (grant nos. JP22K08290, JP21J21306 and JP22KJ2670), by Japan Science and Technology Agency (JST) Moonshot R&D (grant no. JPMJMS2022), by JST FOREST (grant no. JPMJFR2215) and Doctoral Student Grant-in-Aid Program by the Ushioda Memorial Fund. T.F., A.M. and T. Shinozaki were supported by the Japan Society for the Promotion of Science Research Fellowships for Young Scientists. This work was supported by JST ERATO JPMJER2303. This work was also supported by the Takahashi Industrial and Economic Research Foundation and the Masabumi Mori Endowed Research Laboratory for Conquering Lung Cancer.
Author information
Authors and Affiliations
Contributions
T.F., J.H., K.F., H.Y. and Toshiro Sato conceived and designed the study. K.T., M. Oda. and M.F. performed bioinformatics analysis. T.F., J.H., T.E., A.M., K.S., T. Shinozaki., M. Okada., A.S., H. Takaoka., F.I., L.S. and M.M performed in vitro experiments. T.F., J.H. and F.I. performed in vivo experiments. K.E., S.T. and Y.K. performed immunostaining. K.E., Y.O. and Y.K. performed statistical analysis for immunostaining experiments. K.O., Takashi Sato, I.K., K.A., T.H., H.A., S.I., H. Terai and K.S. provided clinical samples and advice from clinical and biological viewpoints. T.F., K.T., J.H., H.Y. and Toshiro Sato wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Toshiro Sato is an inventor on several patents related to organoid culture. All other authors declare no competing interests.
Peer review
Peer review information
Nature Cancer thanks Chong Chen, Hongbin Ji, Martin Sos and Anish Thomas for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Histological recapitulation of SCLC in organoids and xenografts.
(a) Representative H&E, Synaptophysin, CD56 and CHGA staining of the parental clinical sample and SCLC organoids (n = 10 clinical sample and organoid pairs). Scale bar: 100 mm. Five independent organoids were evaluated with similar results. (b) Representative H&E, Synaptophysin, CD56 and CHGA staining of the parental clinical sample (left), organoid (middle) and organoid xenograft (right) of SCLC organoids (n = 6 clinical sample, organoid and xenograft pairs). Scale bar: 100 mm. Five independent organoids were evaluated with similar results.
Extended Data Fig. 2 Genetic alterations and fusion genes in SCLC organoids.
(a) Comparison of copy-number alterations between clinical sample and organoid pairs (10 pairs). The copy-number state is shown as the copy-number ratio between the sample genome and the diploid genome. (b) The DENND2A-BRAF fusion gene detected in SCLC03 organoid line. Sanger sequencing result is shown (bottom). (c) The UBE4B-PIK3CD fusion gene detected in SCLC05. Sanger sequencing result is shown (bottom). (d) The effect of a pan-Raf inhibitor (LY3009120, 0.1 μM) on SCLC organoids with DENN2A-BRAF fusion (SCLC03) or those without (SCLC12 and SCLC19) (n = 3 lines). The organoids were plated as single cells and treated with or without LY3009120 for 21 days. The values indicate the organoid area relative to the mean control area. Scale bar: 500 μm. This experiment was repeated independently twice with similar results. (e) Total (t) and phospho (p)-ERK immunoblotting of SCLC03 and SCLC12 following LY3009120 treatment (n = 2 lines). The organoids were treated with 0, 0.1 or 1 μM LY3009120 for 4 hours. The values indicate the intensity ratio between phospho- and total ERK compared to the control. (f) AKT inhibitor treatment (AZD5363, 0.1 μM) on SCLC organoids with (SCLC05) or without (SCLC12 and SCLC19) UBE4B-PIK3CD fusion (n = 3 lines). The organoids were plated as single cells and treated with AZD5363 for 21 days. The values indicate the organoid area relative to the mean control area. Scale bar: 500 μm. This experiment was repeated independently twice with similar results. (g) Total (t) and phospho (p)-AKT immunoblotting of SCLC05 and SCLC12 following AZD5363 treatment (n = 2 lines). Cell lysates were collected after 4-hour treatment with 0, 0.1 or 1 μM AZD5363. The number indicates relative staining intensity p/t AKT compared to the control.
Extended Data Fig. 3 TF subtyping SCLC organoids.
(a) (Top): Representative ASCL1, NEUROD1, POU2F3 and YAP1 staining of the parental clinical sample and organoid pairs (n = 15 lines). Five independent organoids were evaluated with similar results. Scale bar: 100 μm. (Bottom): Consistency of immunostaining between parental tissue and organoid pairs (n = 19 lines). Positive staining was defined as more than 20 % of cancer cells express corresponding transcription factors in the nucleus. The red square indicates positive/negative staining pattern is matched between the parental tissue and organoid pairs. (b) Representative YAP1 staining of SCLC17 organoid (n = 1 line). Scale bar: 100 mm. Five independent organoids were evaluated with similar results. (c) Representative YAP1 staining in the xenografts of ASCL1-subtype (SCLC19), NEUROD1-subtype (SCLC05), POU2F3-subtype (SCLC14 and SCLC17) and YAP1-subtype (SCLC11, SCLC15 and SCLC23) organoids (n = 7 xenograft tumors). Scale bar: 100 mm. Three mice were evaluated with similar results. (d) Frequency of SCLC subtypes by immunohistochemistry for clinical samples. (Left): A pie chart indicating the frequency of the indicated subtypes in the organoid successful cohort (n = 19). (Right): A pie chart indicating the frequency of the indicated subtypes in the organoid unsuccessful cohort (n = 20). (e) Comparison of gene expressions between indicating culture conditions (ASCL1-subtype (SCLC03 and SCLC07), NEUROD1-subtype (SCLC13), POU2F3-subtype (SCLC30) and YAP1-subtype (SCLC11)) (n = 5 lines). Four transcription factors and neuroendocrine marker genes are shown.
Extended Data Fig. 4 Molecular characterization of 4-TF subtypes.
(a) Consensus clustering of SCLC organoids using the transcriptome data (n = 40 lines). Clustering result using 2 clusters is shown. (b) Hierarchical clustering of SCLC organoids using genes differentially expressed between YAP1-subtype and other SCLCs (Top 20 genes) (n = 40 lines). (c) Gene ontology analysis of genes upregulated in ASCL1 and NEUROD1-subtype and YAP1 and POU2F3-subtype (n = 40 lines). Top 10 pathways with the lowest FDR are listed. (d) Enrichment of neuron- and airway-associated genes in patient SCLC samples27 (n = 59 samples for ASCL1 and NEUROD1 subtypes. n = 29 samples for YAP1 and POU2F3 subtypes). Each dot shows one SCLC clinical sample. Box plots represent the median (center line), upper and lower quartiles (box limits) and 1.5× interquartile range (whiskers). Statistics, two-side Wilcoxon rank sum test. Each dot represents one lines. (e) Motif enrichment in ASCL1-subtype (n = 7 lines), NEUROD1-subtype (n = 2 lines) and POU2F3-subtype (n = 4 lines) organoids. Merged peaks were used for each subtype. The motifs were sorted by the adjusted p value in motif enrichment analysis. Statistics information is available in the Methods section. (f) Lists of airway-specific and alveolar-specific motifs (top 10 motifs) (n = 3 airway organoid lines and n = 3 alveolar organoid lines). Statistics information is available in the Methods section.
Extended Data Fig. 5 Niche factor dependency of SCLC organoids.
(a) Dependency of SCLC organoids on Wnt/R-spondin or Noggin. BMP4 was used in addition to Noggin withdrawal to test the resistance to BMP activation. Images of SCLC organoids cultures for 21 days from single cells with the indicated conditions are shown (n = 6 lines). The values show the organoid area relative to the control area. Scale bar: 500 μm. Three independent experiments were evaluated with similar results. (b) Summary of Wnt/R-spondin and Noggin dependencies in SCLC organoids (n = 30 lines). (c) Dependency of SCLC organoids on IGF-1, EGF or FGF-2 signaling. Images of SCLC organoids cultured for 21 days from single cells in the indicated conditions are shown (n = 7 lines). The values show the organoid area relative to the control area. Scale bar: 500 μm. Three independent experiments were evaluated with similar results. (d) The expression of IGF1R and EGFR in the RNA-seq data of normal airway (gray, n = 3 lines), normal alveolar (brown, n = 3 lines), ASCL1 and NEUROD1-subtype SCLC (blue; ASCL1-subtype, n = 20 lines, purple; NEUROD1-subtype, n = 7 lines), POU2F3 and YAP1-subtype SCLC (orange; YAP1-subtype, n = 9 lines, green; POU2F3-subtype, n = 4 lines) organoids. Each dot shows one organoid line. Box plots represent the median (center line), upper and lower quartiles (box limits) and 1.5× interquartile range (whiskers). NS, not significant, Statistics, two-tailed Welch’s t-test.
Extended Data Fig. 6 EGFR and IGF1R expressions in SCLC organoids.
(a) Flow cytometry analysis of cell-surface EGFR in ASCL1-subtype (n = 3 lines), NEUROD1-subtype (n = 1 line), POU2F3-subtype (n = 1 line), YAP1-subtype (n = 3 lines), normal alveolar (n = 2 lines) and normal airway (n = 2 lines) organoids. EGFR positive cells are gated based on the threshold of the top 5% fluorescence intensity of the isotype control. (b) Flow cytometry analysis of cell-surface IGF1R in ASCL1-subtype (n = 3 lines), NEUROD1-subtype (n = 3 lines), POU2F3-subtype (n = 2 lines), YAP1-subtype (n = 4 lines), normal alveolar (n = 2 lines) and normal airway (n = 2 lines) organoids.
Extended Data Fig. 7 IGF1R and downstream activation in YAP1-subtype SCLC organoids.
(a) Immunoblotting of the indicated proteins in SCLC organoids (n = 8 lines). The proteins were extracted 4 hours after IGF-1 stimulation (middle lane in each line) and IGF-1 stimulation and 1 μM linsitinib (right lane in each line). (b) The expression of AP1 genes FOS, FOSB, FOSL2, JUN, JUNB and JUND in ASCL1 and NEUROD1-subtype SCLC (blue; ASCL1-subtype, n = 20 lines, purple; NEUROD1-subtype, n = 7 lines), POU2F3 and YAP1-subtype SCLC (orange; YAP1-subtype, n = 9 lines, green; POU2F3-subtype, n = 4 lines) organoids. Each dot shows one SCLC organoid line. Box plots represent the median (center line), upper and lower quartiles (box limits) and 1.5× interquartile range (whiskers). NS; not significant, One-way ANOVA test, followed by Turkey’s post-hoc test. p = 0.8743 for FOS, p = 0.4049 for FOSB, p = 0.4288 for JUN and p = 0.5917 for JUNB (NE-type versus non-NE-type). (c) The expression of FOSL1 mRNA in SCLC11 and SCLC15 (YAP1-subtype) organoids treated with or without the AKT inhibitor AZD5363 (0.1 mM, 4 hours) analyzed by quantitative RT-PCR (n = 2 lines with 3 technical replicates for each line). Data are shown as mean ± SD. Statistics, two-tailed Welch’s t-test. (d) FOSL1 staining in YAP1-subtype (SCLC10 and SCLC23) and ASCL1-subtype (SCLC12 and SCLC19) clinical samples (n = 4 clinical samples). Scale bar, 100 mm. (e) The effect of AP1 inhibition on the growth of SCLC organoids. The organoids were seeded as single cells and treated with or without the AP1 inhibitor SR11302 (1 mM) for 21 days (n = 3 lines). Organoid images on day 21 (left) and area quantification (right). Each dot shows one well, and data are shown as mean ± SD. Data are from three technical replicates for each line.
Extended Data Fig. 8 YAP1/TAZ knockdown in SCLC organoids.
(a) Whole-mount immunofluorescence YAP1 staining of SCLC organoids treated with the FAK inhibitor defactinib (0.1 mM, 4 hours) (n = 1 line). YAP1 (green) and nuclei (blue). The percentage of nuclei with YAP1 positivity is shown on the top-left. Scale bar: 100 mm. (b) The constructs of Dox-inducible shYAP/TAZ cassettes. (c) Sorting of cells with an efficient co-transduction of shYAP1 and shTAZ (n = 3 lines). Doxycycline-inducible shYAP1 and shTAZ vectors include TagBFP and mRFP cassettes, respectively. (d) YAP1 and TAZ (WWTR1) knockdown efficiencies in YAP1-subtype (SCLC11 and SCLC15) and ASCL1-subtype (SCLC12) SCLC organoids analyzed by quantitative RT-PCR (n = 3 lines with three technical replicates for each line). Knockdown was induced with a 48-hour treatment with 500 ng/ml doxycycline (DOX). The expression of CYR61 (CCN1) was also analyzed as a YAP1 and TAZ target gene. Data are shown as mean ± SD. (e) The effect of YAP1/TAZ knockdown on the growth of SCLC organoids (n = 3 lines). The organoids were plated as single cells and treated with or without 500 ng/ml doxycycline for 21 days. Bright-field images were overlaid with BFP (YAP1) and RFP (TAZ) fluorescence. The values indicate organoid area relative to the untreated control area. Scale bar: 500 mm. Three independent organoids were evaluated with similar results.
Extended Data Fig. 9 Therapeutic effects of IGF-1 inhibition on YAP1-subtype SCLC organoids.
(a) Dose-response curves of SCLC cell lines representing indicated subtypes treated with linsitinib (n = 8 cell lines (ASCL1-subtype (n = 2 cell lines), NEUROD1-subtype (n = 1 cell line), POU2F3-subtype (n = 2 cell line), YAP1-subtype (n = 3 cell lines)). Three technical replicates for each line. (b) The effect of AKT, FAK of IGF1R blockade on the growth of SCLC organoids (n = 4 lines). The organoids were plated as single cells and were treated with or without AZD5363 (0.1 mM), defactinib (0.1 mM) and linsitinib (0.1 mM) for 21 days. Number indicates the relative organoid area compared to the complete medium (left). Representative images (left) and area measurement (right). Each dot indicates one well, and data are shown as mean ± SD. Scale bar: 500 mm. NS; not significant. Statistics, one-way ANOVA, followed by Tukey’s post-hoc test. Data are from three technical replicates for each line.
Extended Data Fig. 10 TP53/RB1 knockout in human normal airway organoids.
(a) The growth of human normal lung airway organoids (n = 1 line) (left) and TR-KO lung airway organoids (n = 1 line) (right) cultured with the indicated conditions. Scale bar: 500 mm. Three independent experiments were evaluated with similar results. (b) Representative ASCL1, NEUROD1, POU2F3 and YAP1 staining in the TR-KO lung airway organoids (n = 1 line). Scale bar, 100 mm. Three independent organoids were evaluated with similar results. (c) Representative H&E, SYP, CHGA and CD56 staining of normal (n = 1 line) and TR-KO airway organoids (n = 1 line). Scale bar, 100 mm. Three independent organoids were evaluated with similar results.
Supplementary information
Supplementary Table (download XLSX )
Clinical Information of SCLC organoids.
Source data
Source Data Figs. 2–7 and Source Data Extended Data Figs. 2–5 and 7–10 (download XLSX )
Statistical Source Data.
Source Data Figs. 4 and 5 and Source Data Extended Data Figs. 2 and 7 (download PDF )
Uncropped blot images.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fukushima, T., Togasaki, K., Hamamoto, J. et al. An organoid library unveils subtype-specific IGF-1 dependency via a YAP–AP1 axis in human small cell lung cancer. Nat Cancer 6, 874–891 (2025). https://doi.org/10.1038/s43018-025-00945-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43018-025-00945-y
This article is cited by
-
Advancing small cell lung cancer metastasis research: innovations in preclinical mouse models
Cancer and Metastasis Reviews (2025)