Abstract
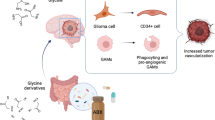
Brain tumors, including glioblastoma multiforme (GBM) and brain metastases, present a notable clinical challenge. Recent research highlights the presence of intratumor bacteria across many tumor types, yet the microbiome of brain tumors remains largely underexplored. Here we show that the microbiome of 322 brain tumors differs markedly by tumor type and location. Using multiple approaches to visualize, culture and sequence bacterial communities, we found that brain metastases harbor higher bacterial richness and diversity than GBM, with distinct microbial compositions. Moreover, metastases in posterior brain regions exhibited greater diversity than those in anterior regions. Pathway analysis revealed enrichment of bacterial metabolic pathways associated with tumor spread and metastasis in brain metastases while GBM was enriched with pathways supporting alternative phosphorus use. These findings provide valuable insights into the microbial landscape of brain tumors, highlighting tumor-type-specific and location-specific variation and suggesting potential roles for bacteria in brain tumor biology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The sequencing data generated in this study were deposited to the National Center for Biotechnology Information Sequence Read Archive under BioProject accession number PRJNA1193807. Processed data supporting the findings of this study are available within the paper and its Supplementary Information. Additional data are available from the corresponding author upon reasonable request. Source data are provided with this paper.
Code availability
No new or custom code was used in this study. Detailed methods and software packages with references are provided in the Methods.
Change history
19 January 2026
In the version of Source Data Fig. 3 initially published alongside this article, some values in the Fig. 3b table were incorrect. An updated Source Data Fig. 3 file is available in the HTML version of this article.
References
Cioffi, G. et al. Changes in survival over time for primary brain and other CNS tumors in the United States, 2004–2017. J. Neurooncol. 160, 209–219 (2022).
Js, B.-S. et al. A nomogram for individualized estimation of survival among patients with brain metastasis. Neuro Oncol. 14, 910–918 (2012).
Nejman, D. et al. The human tumor microbiome is composed of tumor type-specific intracellular bacteria. Science 368, 973–980 (2020).
Battaglia, T. W. et al. A pan-cancer analysis of the microbiome in metastatic cancer. Cell 187, 2324–2335 (2024).
Sepich-Poore, G. D. et al. The microbiome and human cancer. Science 371, eabc4552 (2021).
Yang, L., Li, A., Wang, Y. & Zhang, Y. Intratumoral microbiota: roles in cancer initiation, development and therapeutic efficacy. Signal Transduct. Target. Ther. 8, 1–24 (2023).
Zhao, J. et al. Comprehensive histological imaging of native microbiota in human glioma. J. Biophotonics 15, e202100351 (2022).
Naghavian, R. et al. Microbial peptides activate tumour-infiltrating lymphocytes in glioblastoma. Nature 617, 807–817 (2023).
Li, T. et al. Multi-omics analysis reveals the interplay between intratumoral bacteria and glioma. mSystems 10, e0045724 (2025).
Li, W. et al. Deciphering the impact of intra-tumoral bacterial infiltration on multi-omics profiles in low-grade gliomas. Front. Oncol. 15, 1582068 (2025).
Harvey, K. L., Jarocki, V. M., Charles, I. G. & Djordjevic, S. P. The diverse functional roles of elongation factor Tu (EF-Tu) in microbial pathogenesis. Front. Microbiol. 10, 2351 (2019).
Siegrist, M. S. et al. d-Amino acid chemical reporters reveal peptidoglycan dynamics of an intracellular pathogen. ACS Chem. Biol. 8, 500–505 (2013).
Fuks, G. et al. Combining 16S rRNA gene variable regions enables high-resolution microbial community profiling. Microbiome 6, 17 (2018).
Douglas, G. M. et al. PICRUSt2 for prediction of metagenome functions. Nat. Biotechnol. 38, 685–688 (2020).
Matchado, M. S. et al. On the limits of 16S rRNA gene-based metagenome prediction and functional profiling. Microb. Genom. 10, 001203 (2024).
Sun, S., Jones, R. B. & Fodor, A. A. Inference-based accuracy of metagenome prediction tools varies across sample types and functional categories. Microbiome 8, 46 (2020).
Soda, K. The mechanisms by which polyamines accelerate tumor spread. J. Exp. Clin. Cancer Res. 30, 95 (2011).
Guerra, B. et al. The mevalonate pathway, a metabolic target in cancer therapy. Front. Oncol. 11, 626971 (2021).
Yu, R. et al. Mevalonate pathway inhibition slows breast cancer metastasis via reduced N-glycosylation abundance and branching. Cancer Res. 81, 2625–2635 (2021).
Du, X. et al. The mevalonate pathway promotes the metastasis of osteosarcoma by regulating YAP1 activity via RhoA. Genes Dis. 9, 741–752 (2022).
Gruenbacher, G. & Thurnher, M. Mevalonate metabolism governs cancer immune surveillance. Oncoimmunology 6, e1342917 (2017).
Bekkering, S. et al. Metabolic induction of trained immunity through the mevalonate pathway. Cell 172, 135–146 (2018).
Peter, S. B. & Nandhan, V. R. 31-Phosphorus magnetic resonance spectroscopy in evaluation of glioma and metastases in 3T MRI. Indian J. Radiol. Imaging 31, 873–881 (2021).
Chakrabarti, I., Cockburn, M., Cozen, W., Wang, Y.-P. & Preston-Martin, S. A population-based description of glioblastoma multiforme in Los Angeles County, 1974–1999. Cancer 104, 2798–2806 (2005).
Schroeder, T. et al. Mapping distribution of brain metastases: does the primary tumor matter? J. Neurooncol. 147, 229–235 (2020).
Nayak, L., Lee, E. Q. & Wen, P. Y. Epidemiology of brain metastases. Curr. Oncol. Rep. 14, 48–54 (2012).
Habbous, S. et al. Incidence and real-world burden of brain metastases from solid tumors and hematologic malignancies in Ontario: a population-based study. Neurooncol.Adv. 3, vdaa178 (2021).
Omuro, A. & DeAngelis, L. M. Glioblastoma and other malignant gliomas: a clinical review. JAMA 310, 1842–1850 (2013).
Schaff, L. R. & Mellinghoff, I. K. Glioblastoma and Other primary brain malignancies in adults: a review. JAMA 329, 574–587 (2023).
Dohlman, A. B. et al. The cancer microbiome atlas: a pan-cancer comparative analysis to distinguish tissue-resident microbiota from contaminants. Cell Host Microbe 29, 281–298 (2021).
Arvanitis, C. D., Ferraro, G. B. & Jain, R. K. The blood–brain barrier and blood–tumour barrier in brain tumours and metastases. Nat. Rev. Cancer 20, 26–41 (2020).
Bullman, S. et al. Analysis of Fusobacterium persistence and antibiotic response in colorectal cancer. Science 358, 1443–1448 (2017).
Fu, A. et al. Tumor-resident intracellular microbiota promotes metastatic colonization in breast cancer. Cell 185, 1356–1372 (2022).
Tokuyasu, K. T. Application of cryoultramicrotomy to immunocytochemistry. J. Microsc. 143, 139–149 (1986).
Acknowledgements
We thank the participants and their families for their contribution to this research. We also thank members of the R.S. lab for their support. R.S. is supported by the Israel Cancer Research Fund (project grant no. 22-109-PG), The Mark Foundation for Cancer Research, the Israel Science Foundation (grant no. 2927/21), the European Research Council (European Union’s Horizon 2020 research and innovation program, grant agreement no. 818086), the Fabrikant–Morse Families Research Fund for Humanity, the Knell Family Center for Microbiology, the Moross Integrated Cancer Center, the Dr. Dvora and Haim Teitelbaum Endowment Fund and the Elsie and Marvin Dekelboum Family Foundation.
Author information
Authors and Affiliations
Contributions
E.G., A.S., S.Y.-K. and R.S. conceptualized and initiated the project. A.L.-B., Y.L., A.K., A.A., T.S., A.L., I.S., O.B., Y. Zohar, L.L.P., C.R.H., P.C., S.H., J.J.M., A.S. S.Y.-K., E.S. and I.B. provided participant samples. E.G., N.G., P.C., I.G., O.G., S.H. and R.S. conducted the tissue staining and analysis. E.G., N.G., T.D. and S.LZ. performed the EM and correlative light and EM. E.G. performed the culturomics. N.G. conducted the ex vivo organ culture experiments. E.G. and Y. Zwang generated the sequencing data. E.G., L.R.-N., N.S. and R.S. performed the analysis of 16S rRNA-seq and PICRUSt2 data. Y. Zwang and I.L. helped with the data analysis. H.Z. and N.G. performed the cell line experiments. E.G. and R.S. wrote the initial manuscript. E.G., N.G., L.R.-N., Y. Zwang, N.S., A.S., S.Y.-K. and R.S. helped with discussions and manuscript revision. R.S. supervised the project.
Corresponding author
Ethics declarations
Competing interests
R.S. reports stock interest in CuResponse, Baccine and Gnomica and is a paid adviser to these companies. N.G. reports stock interest in CuResponse and is a paid advisor of CuResponse. The other authors declare no competing interests.
Peer review
Peer review information
Nature Cancer thanks Reinhild Feuerstein, Dihua Yu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Anti-EF-Tu antibody detect bacteria in mouse MC38 tumors.
106 of Escherichia coli or Salmonella typhimurium bacteria or PBS control were injected into 500mm3 MC38 tumors on the flanks of C57BL\6 mice. Twenty-four hours after bacterial injection, mice were sacrificed, and tumors were harvested and preserved as FFPE blocks. (a) Representative images of immunofluorescence staining of FFPE slices from the mouse tumors with anti-bacterial elongation factor EF-Tu antibody (see Methods for details). DAPI is shown in blue. EF-Tu in orange (Cy5). Scale bars in low-magnification images, 500 μm. Scale bars=20 μm. (b) Representative images from sequential slides stained as above without primary antibody, as a negative control. Scale bars=20 μm. For panels a,b images shown are from n = 1 mouse.
Extended Data Fig. 2 Bacteria are detected within human GBM tumors.
(a-e) Representative images of cores from tissue microarray (TMA) of human GBM (70 cores) stained by immunofluorescence for bacterial elongation factor EF-Tu and by FISH for 16S rRNA, along with their respective controls. DAPI is shown in blue. EF-Tu and 16S rRNA in orange (Cy5). Asterisks mark the region selected for higher magnification. Scale bars in low magnification images, 500 μm. Scale bars in high-magnification images, 20 μm. For panels a-e images shown are from n = 5 cores from n = 3 patients.
Extended Data Fig. 3 Bacteria are detected within human brain metastases.
(a-e) Representative images of cores from a TMA of human brain metastases (86 cores) stained by immunofluorescence for bacterial elongation factor EF-Tu and by FISH for 16S rRNA with corresponding controls. DAPI is shown in blue. EF-Tu and 16S rRNA in orange (Cy5). Asterisks mark areas selected for higher magnification. Scale bars in low-magnification images, 500 μm. Scale bars in high-magnification images, 20 μm. For panels a-e images shown are from n = 5 cores from n = 5 patients.
Extended Data Fig. 4 Intratumor bacteria reside inside cancer cells.
(a–b) Representative images of TMA cores from (a) GBM (70 cores) and (b) brain metastasis samples (86 cores) co stained for bacterial elongation factor EF-Tu (orange), and either GFAP (a) or cytokeratin (b) (red)DAPI is shown in blue. Upper left panel: low-magnification overview with marked regions shown at higher magnification below. Corresponding no primary-antibody controls are shown on the upper right panel. Scale bars in low-magnification images, 200 μm. Scale bars in high-magnification images, 10 μm. For panels a,b images shown are from n = 1 patient per group.
Extended Data Fig. 5 Bacteria in human GBM and brain metastases tumor cells are mostly found in the intracellular space.
(a-d) Transmission electron microscopy (TEM) representative images of intracellular bacteria within one human GBM tumor, detected using immunogold labeling. Sections were incubated with a primary antibody against bacterial LPS, followed by a gold-labeled secondary antibody. An enlarged view of the boxed area is shown on the right. Scale bars as indicated. (e) Control TEM representative images from adjacent sections processed without the primary anti-LPS antibody, showing the absence of gold labeling, confirming the specificity of the immunogold signal. Scale bars as indicated. The letter N marks the cell nucleus. For panels a-e images shown are from n = 1 patient.
Extended Data Fig. 6 Culturomics and D-alanine staining reveal viable intracellular bacteria in GBM tumors.
(a) Representative images of isolated bacterial colonies from fresh GBM tumors (n = 1 patient) (b) Three freshly resected GBM tumors (n = 3 patients) were incubated with fluorescently labeled D-alanine (blue) to detect viable bacteria or DMSO as control. Nuclei stained with DRAQ5 (orange). Scale bars as indicated in each frame. Each box plot represents one tumor, summarizing fluorescence intensity values from two tissue slices and 5–6 quantified regions per slice. P-values were calculated using a two-tailed Student’s t-test. Box plots show medians as central lines. Boxes represent the interquartile range (IQR), and whiskers extend to 1.5×IQR or to the minimum/maximum values if no point exceeds that range. No outliers were removed from the plots. (c) Freshly resected GBM tumor (n = 1 patient, same tumor as in Fig. 2c,d) was incubated with fluorescently labeled D-alanine (blue) ± 100 μg/ml levofloxacin to detect viable bacteria or DMSO control. Nuclei stained with DRAQ5 (orange). Scale bars=200 μm.
Extended Data Fig. 7 Matched metastasis samples from the same patient show lower Jaccard beta diversity with minimal change in effect size.
(a-c) Box plots showing Jaccard beta diversity comparisons between matched brain metastasis samples- multiple samples from the same patient and non-matched brain metastases samples from different patients. Only patients with more than one brain metastasis sample are included in the matched group. Analyses are presented for combined breast and lung cancer brain metastatic samples (matched: n = 45 pairwise comparisons from n = 37 patients, non-matched: n = 7,458 pairwise comparisons from n = 93 patients) (a), breast cancer brain metastases only (BCBrM; matched: n = 34 pairwise comparisons from n = 26 patients, non-matched: n = 2,522 pairwise comparisons from n = 42 patients) (b), and lung cancer brain metastases only (LCBrM; matched: n = 11 pairwise comparisons from n = 11 patients, non-matched: n = 1,264 pairwise comparisons from n = 51 patients) (c). Data points represent individual pairwise Jaccard index values. P-values were calculated using a two-tailed Student’s t-test. Box plots show medians as central lines. Boxes represent the interquartile range (IQR), and whiskers extend to 1.5×IQR or to the minimum/maximum values if no point exceeds that range. No outliers were removed from the plots.
Extended Data Fig. 8 Distinct microbial signatures in GBM compared to brain metastases.
(a) Permanova results corresponding to Fig. 4a, showing tumor type-specific explained variance (R²) in the ordination space. Significance was assessed using a two-sided Permanova test with 999 permutations, followed by pairwise post hoc comparisons. P-values were adjusted for multiple comparisons using Bonferroni correction. Reported values represent explained variance (R²) and adjusted P-values for tumor-type–specific sample groups. (b-c) Principal coordinate analysis (PCoA) based on Jaccard dissimilarity indexes, comparing bacterial species profiles in GBM, breast cancer brain metastases (BCBrM), and lung cancer brain metastases (LCBrM) from (b) samples collected at Rambam and Galilee medical centers together (c) samples from all medical centers. All bacteria that passed filtering criteria in any of the tumor types were included in the analysis. (d-e) Volcano plots demonstrating the differential distribution of bacterial taxa (order, class and phylum level) between (d) GBM and breast cancer brain metastases, (e) GBM and lung cancer brain metastases. A two-sided Wilcoxon test was used to calculate the P values. Dot size reflects the FDR value, with larger dots indicating lower FDR values. Significant taxa are color-coded according to the associated tumor type, meeting thresholds of effect size >5%, P-value < 0.05, and FDR-corrected Q value < 0.1. (f) Box plots showing Bray-Curtis beta diversity comparisons between matched and unmatched pairs of primary lung tumors and brain metastasis. Matched pairs represent samples from the same patient (n = 5 patients); For each patient, two samples from each condition (primary and metastasis) were included when available, resulting in multiple pairwise comparisons per patient. Unmatched comparisons include pairs from different patients. The two-sided Wilcoxon test was used to calculate the P-values. Box plots show medians as central lines. Boxes represent the interquartile range (IQR), and whiskers extend to 1.5×IQR or to the minimum/maximum values if no point exceeds that range. No outliers were removed from the plots. (g) PCoA based on Bray-Curtis dissimilarity indexes, comparing bacterial species profiles in matched pairs of primary lung tumors and brain metastasis (LCBrM) from five patients (n = 5 patients). For panels a-e: n = 151 patients for GBM, n = 79 tumor samples from n = 45 patients for BCBrM, n = 71 tumor samples from n = 56 patients for LCBrM.
Extended Data Fig. 9 Location-based variation of bacterial richness, diversity, and prevalence in brain metastases.
(a, b) Box plots showing bacterial richness (a) and Shannon diversity index (b) per independent brain metastasis sample from breast cancer brain metastases (BCBrM; frontal n = 19 tumor samples from n = 13 patients, posterior n = 27 tumor samples from n = 16 patients) and lung cancer brain metastases (LCBrM; frontal n = 16 tumor samples from n = 16 patients, posterior n = 17 tumor samples from n = 14 patients). Each “n” corresponds to a distinct patient-derived metastasis specimen (biological replicate). (c, d) Box plots showing bacterial richness (c) and Shannon diversity index (d) in brain metastases from the frontal (n = 35 tumor samples from n = 29 patients), temporal (n = 11 tumor samples from n = 9 patients), parietal (n = 5 tumor samples from n = 3 patients), occipital (n = 16 tumor samples from n = 12 patients), and cerebellum (n = 28 tumor samples from n = 19 patients) brain regions. (e, f) Box plots comparing bacterial richness (e) and Shannon diversity index (f) between brain metastases located in the frontal (n = 35 tumor samples from n = 29 patients), midbrain (n = 16 tumor samples from n = 12 patients), and posterior (n = 44 tumor samples from n = 30 patients) regions. (g) Paired line plot comparing bacterial species richness between matched anterior vs posterior brain metastases from the same patient. Each line connects two independent metastasis samples from the same patient (n = 3 patients). (h, i) Volcano plots showing the differential prevalence of bacterial taxa between frontal (n = 35 tumor samples from n = 29 patients) and cerebellum (n = 28 tumor samples from n = 19 patients) (h), and between (i) frontal (n = 35 tumor samples from n = 29 patients) and occipital (n = 16 tumor samples from n = 12 patients) (i) brain regions in all brain metastases samples. Dot size reflects the FDR value, with larger dots indicating lower FDR values. Significant taxa are color-coded by tumor location, meeting thresholds of effect size >5%, P-value < 0.05, and FDR-corrected Q-value < 0.05. BrM, All brain metastases samples. P-values were calculated using a two-tailed Student’s t-test for (a–f), and a one-sided binomial test for (h,i). (a-f) Box plots show medians as central lines. Boxes represent the interquartile range (IQR), and whiskers extend to 1.5×IQR or to the minimum/maximum values if no point exceeds that range. No outliers were removed from the plots. All “n” values indicate the number of independent biological replicates (patient-derived brain metastasis samples) analyzed. No technical replicates were used for statistical testing. Control groups are defined by anatomical location (different brain regions).
Extended Data Fig. 10 Prevalence of selected genera across brain regions in GBM and brain metastases.
(a-d) Bar plots showing the prevalence of chosen genera from Fig. 6d in occipital and cerebellum tumors compared to frontal tumors in breast cancer and lung cancer brain metastases. P-values were calculated using two-sided Fisher’s exact test. (e) Box plot showing the Shannon diversity index per sample in GBM tumors compared to all brain metastases in anterior (GBM: n = 94 patients, BrM: n = 51 tumor samples from n = 38 patients) and occipital (GBM: n = 7 patients, BrM: n = 16 tumor samples from n = 12 patients) brain regions. P-value was calculated using a two-tailed Student’s t-test. (f) Scatter plot showing the effects of bacterial pre-conditioned media on the response of two human glioma cell lines (T98G and U251) to TMZ treatment. Each point represents one of 30 bacterial species from genera that were identified to be prevalent in GBM tumors (n = 30 bacterial species). The x- and y-axes represent the rescue score (rScore) as explained in the methods section, where positive values represent PCM-mediated resistance and negative values represent an additive or synergistic effect between the PCM and TMZ. Colored dots represent different bacterial genera. A linear regression line is shown to illustrate the relationship between responses across cell lines. BrM, All brain metastases samples. BCBrM, Breast cancer brain metastases samples. LCBrM, Lung cancer brain metastases samples. for panels a-d: frontal: n = 35 tumor samples from n = 29 patients, occipital: n = 16 tumor samples from n = 12 patients, cerebellum: n = 28 tumor samples from n = 19 patients.
Supplementary information
Supplementary Video 1 (download MP4 )
Confocal Z-stack reconstruction of representative images shown in Fig. 1d,e. The video demonstrates of bacterial EF-Tu staining (white) with (DAPI positive, blue) Cells, supporting the presence of bacteria within the host cell cytoplasm.
Supplementary Video 2 (download MP4 )
Confocal Z-stack reconstruction of representative images shown in Fig. 1d,e. The video demonstrates co-localization of bacterial EF-Tu staining (white) with (DAPI positive, blue) Cells, supporting the presence of bacteria within the host cell cytoplasm.
Supplementary Tables 1–7 (download XLSX )
Supplementary Tables 1–7.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gigi, E., Gavert, N., Raijman-Nagar, L. et al. Characterization of the tumor microbiome of brain metastases and glioblastoma reveals tumor-type-specific and location-specific microbial signatures. Nat Cancer 6, 1761–1776 (2025). https://doi.org/10.1038/s43018-025-01073-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43018-025-01073-3