Abstract
Diffuse midline glioma (DMG) is a highly aggressive and untreatable pediatric cancer primarily arising in the pontine brainstem region, necessitating the development of representative models for treatment advance. Here we developed an FGF4-driven human brainstem organoid model, which we used to genetically engineer H3.3K27M-altered DMG. We demonstrated that brainstem pontine glial specification is critical for DMG tumorigenesis, yielding infiltrative tumors that recapitulate patient-representative intratumoral heterogeneity. Prolonged GD2 chimeric antigen receptor (CAR) T cell treatment mirrored clinical outcomes and revealed extensive transcriptional heterogeneity, from which both potent effector and dysfunctional CAR T cell populations could be identified. Furthermore, incorporation of myeloid cells generated DMG-specific microglia that reduced treatment efficacy and revealed CAR T cell functional states most vulnerable to microglia-mediated immunosuppression. Thus, we present a representative DMG model offering a months-long experimental window in vitro, which we leveraged to delineate CAR T cell functionality and microglial impact, aiding therapy development for this devastating disease.
Similar content being viewed by others
Main
Diffuse midline gliomas (DMGs) are rare and aggressive pediatric brain tumors often caused by somatic mutations in histone 3 (H3) genes, commonly a K27M substitution1, occurring at a high prevalence in the pons region of the brainstem2. Primarily, affecting children under 10 years3, they present the highest mortality rate of any cancer, with a median overall survival of only 9–15 months4,5. This detrimental prognosis necessitates a better understanding of the disease’s biology to develop effective treatments.
Single-cell analyses of H3K27M-altered DMG revealed intratumoral heterogeneity, with a spectrum of tumor cell profiles ranging from stalled stem-like oligodendrocyte progenitor cell (OPC-like) to more differentiated astrocyte (AC-like) and oligodendrocyte (OC-like) phenotypes, which closely resemble normal developmental cell types, alongside a recently identified mesenchymal-like (MES-like) state6,7,8. In addition, insights from both animal9,10,11 and human pluripotent stem-cell-derived12,13 studies suggest an early neurodevelopmental window of tumor initiation. Thus, dysregulated mechanisms during hindbrain development10,14,15, particularly involving glial progenitors in the region responsible for brainstem pons formation16, likely have a central role in driving H3K27M-altered gliomagenesis. Capturing this region-specific embryonic patterning is, therefore, crucial for accurately modeling pontine DMG. Furthermore, previous work identified a tight relationship between DMG progression and its unique environment, including neuron and synaptic signaling17,18,19, which can promote glioma growth20.
Human brain organoids have become valuable in vitro tools for investigating brain development and understanding the onset, progression and potential therapeutic targeting of nervous system disorders, including cancer21,22,23. Given the rarity and inoperable nature of DMG, which limits the availability of patients participant material2, organoids could offer a scalable model for generating DMG tumors de novo and enabling in vitro testing of emerging therapies. This includes the latest advances in immunotherapy for DMG, GD2 chimeric antigen receptor (CAR) T cells5, which showed promising, yet variable treatment outcomes between patients in a recent clinical trial24. Correlative data from this trial suggest that an expansion of the immunosuppressive myeloid compartment coincides with unfavorable treatment outcomes25. Uncovering the functional profiles of CAR T cells and their interplay with the immunosuppressive tumor microenvironment may offer key insights to improve their therapeutic outcomes in DMG.
Here, we report a human cerebral guided organoid model for the brainstem region, enriched for pontine medulla glial lineages. Genetic modeling of H3.3K27M-altered DMG in these brainstem-regionalized organoids (BrOs) replicates the infiltrative nature and transcriptomic landscape of DMG. We demonstrate the utility of this accessible human DMG organoid model (DMGO) for modeling CAR T cell functional heterogeneity during prolonged treatment (up to 1 month) and within the context of brain-resident microglia.
Results
FGF4-driven morphogen guidance patterns for brainstem identity
To generate human organoids with appropriate hindbrain brainstem identity for DMG modeling, we applied sequential morphogen guidance using a timely sequence of Wnt, dual SMAD inhibitors, retinoic acid (RA), fibroblast growth factors (FGFs) and sonic hedgehog (SHH) (Fig. 1a, Extended Data Fig. 1a,b and Supplementary Table 1). While FGF2 and FGF8 can be used in combination with RA and Wnt to pattern the midbrain26, cerebellum27 or spinal cord28 in growing organoids, we evaluated FGF4 because of its role in specifying rostral hindbrain, particularly in the pontine area29,30, as well as its involvement in the development of hindbrain-specific serotonergic neurons31. A direct comparison of replacing common FGF2 supplementation with FGF4 after 7 days of patterning demonstrated that 10 ng ml−1 FGF4 specifically promotes developing the pontine, including the prepontine to retropontine area, according to bulk sequencing data (Fig. 1b and Supplementary Table 2). Furthermore, among HOX genes important for hindbrain formation, expression of HOXB1, a marker of pontine precursor cells32, emerged already at an early stage (day 14) (Extended Data Fig. 1c) and three-dimensional (3D) imaging revealed HOXB1-expressing cells within early neurodevelopmental SOX2+ neural rosette structures (Fig. 1c). Bulk sequencing analysis from day 7 to day 84 showed that the patterning remained consistent and reproducible across and within multiple batches, as well as between human embryonic stem cell (hES cell) and induced pluripotent stem cell (iPS cell) sources (Extended Data Fig. 1d,e).
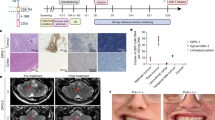
a, Schematic representation of timely morphogen stimulated patterning of hES cells and hiPS cells toward brainstem organoids and their subsequent application for DMG tumor, CAR T cell treatment and microglia-enriched tumor microenvironment modeling. b, Heat map of z score measuring relative brain region identity on the basis of VoxHunt similarity mapping for various supplemented concentrations of FGF2 or FGF4 (n = 3 experimental repeats, with n = 3 organoids pooled; total n = 9 organoids per condition). c, Immunofluorescence 3D images of a 200-µm-thick organoid slice on day 21 labeled for F-actin (white), SOX2 (yellow) and HOXB1 (red). Right, zoomed-in view of area in white inset. Scale bars, 250 µm (main image) and 25 µm (inset). A representative image of n = 2 slices from n = 2 organoids is shown. d, VoxHunt spatial correlation map of day 120 brainstem organoids with E18.5 mouse brain. The pons area is delineated in red (n = 9 organoids from three independent batches). e, Integrated UMAP representation of developing brainstem organoids from different time points, colored by cell annotation. f, Area plot following the relative distribution of cell types over time. Cell types are color-coded as in e. g, UMAP of the HNOCA34 colored for brainstem organoid presence score. A high score indicates a high likelihood that these HNOCA cells are present in the brainstem organoid dataset. Areas annotated by a dashed line indicate lineages as annotated in the HNOCA. Inset, UMAP colored by coarse regional annotation. h, Heat map showing the log2 fold compositional changes in the brainstem organoid dataset compared to the HNOCA. Positive values correspond to an increased abundance of cells from the indicated regional identity or glial lineage. NPC, neuronal precursor cell. i, Cell clusters in the HDBCA35 with gained coverage in brainstem organoids relative to the HNOCA. The horizontal line indicates the threshold used to define a cluster as gained or not. j, UMAP of the HDBCA showing, in shades of red, the HDBCA clusters gained in brainstem organoids, mostly related to oligos and glioblasts. Gray represents clusters below the threshold used to define gained. Inset, UMAP colored by coarse regional annotation. For e–j, n = 84 organoids in total with 5–24 organoids pooled per time point (details in Supplementary Table 1).
To investigate cellular composition and regional identities at higher resolution, we performed time-course single-cell RNA sequencing (scRNA-seq) across eight time points, spanning from day 5 to day 120 (Extended Data Fig. 2a and Supplementary Table 3). Following quality control (Extended Data Fig. 2b and Supplementary Table 3) and doublet filtering, we recovered 55,327 high-quality cells. Spatial similarity mapping using VoxHunt33, a tool based on Mus musculus in situ hibridization data from the Allen Brain Atlas, confirmed a hindbrain identity with a more pronounced pontine signature (Fig. 1d). We next generated an integrated uniform manifold approximation and projection (UMAP) representation of the different time points and performed cell-based annotation using reference datasets, including the recently published Human Neural Organoid Cell Atlas (HNOCA)34 and the Human Developing Brain Cell Atlas (HDBCA)35,36 (Fig. 1e and Extended Data Fig. 2c–g). Temporal analysis revealed an initial phase of high proliferation that diminished over time (Extended Data Fig. 3a), as cells transitioned from pluripotent stem cells to the neuroepithelium and radial glial cells, as well as into distinct neuronal and glial populations that emerged by days 14 and 60, respectively (Fig. 1f), reflecting the natural occurring segregation of neurogenesis and gliogenesis phases35. Projection of the organoid dataset onto the HNOCA that was regionally annotated for neuronal lineages (Fig. 1g) revealed that most neuronal precursor cells, neuroblasts and neurons originated from a heterogeneous nontelencephalic cluster with enrichment for midbrain, pons and medulla regions (Fig. 1h), which collectively form the brainstem37. These neuroblasts and early neurons expressed STMN2 and RBFOX3 (NeuN) but lacked the telencephalic marker FOXG1 (ref. 35) (Extended Data Fig. 3b). Furthermore, neurotransmitter transporter analysis revealed a predominance of excitatory (glutamatergic) and inhibitory (GABAergic) neurons, the latter known to form synapses with DMG and promote its growth17,18. Smaller proportions of cholinergic and dopaminergic neurons were also detected, consistent with their distribution in the HDBCA (Extended Data Fig. 3c,d). In addition, immunofluorescence analysis identified cells expressing tryptophan hydroxylase 2 (Extended Data Fig. 3e), a key enzyme involved in serotonergic synthesis, suggesting that, albeit undetectable at the scRNA-seq level similar to the HDBCA35, this population of neurons is present. Thus, consistent with HNOCA and adult brain data showing greater heterogeneity and intermixing among nontelencephalic neurons, including hypothalamic, brainstem and hindbrain neurons (referred to as splatter neurons) compared to cortical neurons34,38, our organoid model recapitulates this regional diversity, with an enrichment in brainstem identity compared to profiles described in most HNOCA protocols.
Pontine and medulla enrichment of glial lineages
DMG is rooted in the glial lineage8, prompting us to investigate the glial composition within our BrO model. First, we showed the presence of committed ACs (GFAP+AQP4+) and OCs (OLIG2+) at the protein level (Extended Data Fig. 3f). At the single-cell transcriptomic level, we identified glial populations spanning pre-OPCs, OPCs, committed OC precursors, glioblasts and ACs, offering a detailed representation of glial diversity and maturation states (Fig. 1e). By comparing age-matched cells of HNOCA-covered protocols, BrOs showed significant enrichment in the glial lineage, particularly glioblasts and OPCs (Fig. 1h). Additionally, we assessed glycolysis, an indicator of cell stress in brain organoids34. Consistent with models described in the HNOCA, we observed similar glycolysis levels (Extended Data Fig. 3g). However, in the glial lineage, glycolysis levels were lower (Extended Data Fig. 3h), suggesting reduced stress and a healthier metabolic state of glial cells in BrOs. Moreover, OPC (referred to as oligo in the HDBCA35) and glioblast populations demonstrated a reduced number of differentially expressed genes (DEGs) compared to HNOCA datasets (Extended Data Fig. 3i), reflecting higher transcriptional fidelity and closer alignment with primary counterparts in the HBDCA35. To date, no comprehensive region-wide analysis has been conducted on glial cells derived from organoids. However, the HBDCA revealed strong region specificity in the glial lineage during early brain development, which may be particularly relevant for H3.3K27M-altered DMG in the brainstem pontine region. Projection of the BrO datasets onto the HBDCA latent space (Extended Data Fig. 3j,k) and comparison to organoid protocols embedded in the HNCOA revealed significant coverage of glial clusters in BrOs. Notably, 13 of the 19 gained clusters exhibited pontine and medulla-specific identities (Fig. 1i,j and Extended Data Fig. 3j,k). Thus, our newly generated BrO model offers an experimental framework for studying gliogenesis within the context of pontine medulla regionality, offering relevance for DMG modeling.
De novo generation of H3.3K27M-altered DMG
We next investigated whether BrOs could be exploited to model DMG tumors. Plasmids11 expressing the most common H3.3K27M-defining DMG mutation39, alongside typical accompanying and pons-specific tumor suppressor TP53 and platelet-derived growth factor A (PDGFRA) alterations3,40,41, were introduced using in situ electroporation of developing BrOs (Fig. 1a). This mutation cocktail has been shown to be time sensitive in in utero electroporation mouse models9,11,42; hence, we tested different time points of electroporation between days 11 and 28. We identified day 11 as the time point most efficiently inducing tumorigenic growth (Fig. 2a and Extended Data Fig. 4a), reinforcing the concept of a restricted early developmental time window for DMG transformation9,11. At this stage of development, we observed a dominance of radial glia and neuroepithelial stem-like cells in BrOs (Fig. 1f), aligning with earlier work identifying neural stem cells as a permissive cell state for H3.3K27M-driven neoplastic transformation9,11,12,13,14. Tracking tumor growth over 2 months showed that the resulting tumors display infiltrative growth (Fig. 2b). In contrast, the use of empty control plasmids resulted in only a few localized electroporated cells (Extended Data Fig. 4b). Whole-organoid 3D imaging at week 16 (4 months after electroporation) with tumors color-coded for invasion depth further confirmed a diffuse growth pattern characteristic of DMG (Fig. 2c). In addition, DMGOs orthotopically transplanted in immunodeficient mice were able to progress in vivo, demonstrating invasive growth (Extended Data Fig. 4c). Quantification of H3.3K27M expression, combined with dominant-negative TP53 (DNp53) and PDGFRA-D842V at the protein level, showed incorporation of all three mutations into the majority of GFP-positive cells (Fig. 2d–f). These findings illustrate DMG invasive outgrowth in our guided brain organoids dependent on combined common driver mutations typically observed.
a, Stacked bar plot quantifying electroporation efficiency (light-gray columns; day 11 versus day 14, P = 0.782) and tumor induction (dark-gray columns; day 11 versus day 14, P = 0.010) in brainstem organoids tested at various time points ranging from day 11 to day 28. NS, not significant; **P < 0.01, according to a two-tailed independent t-test. Data are shown as the mean ± s.e.m. (n = 139 BrOs in total with 23–35 BrOs per time point; details in Supplementary Table 1). b, Tumorigenic outgrowth of the same DMGO at weeks 4, 6 and 8. Representative images of n = 3 organoids. Scale bars, 500 µm. White arrowheads depict invasive and diffuse patterns. c, Representative immunofluorescence 3D image of intact DMGOs with GFP signal color-coded for z depth on a rainbow scale. The gray outline was created by masking of propidium iodide fluorescence (n = 2 DMGOs). Scale bar, 500 µm. d,e, Representative multispectral 3D images of tumor GFP (green), H3K27M (magenta) and DNp53 (yellow) (d) or tumor GFP (green) and PDGFRA (red) (e) in consecutive slices of a week 8 DMGO (n = 3 DMGOs). Scale bars, 50 µm. f, Percentage of GFP+ tumor cells expressing H3K27M, DNp53 or PDGFRA detected by multispectral 3D imaging as in d,e (n = 2 DMGOs with n = 3 ROIs imaged per DMGO). g, Methylation profile of DMGO compared to DMG or resembling tumor types. Each dot represents one patient sample. GBM, glioblastoma; EPN, ependymoma; tSNE, t-distributed stochastic neighbor embedding. For DMGO, the dot represents a pooled sample of n = 3 DMGOs. h, Integrated Force Atlas representation of DMGO tumors colored by tumor cell state (n = 14 DMGOs from four independent batches. i, Heat map representation of average transcriptomic similarity between DMGO tumor cells and in vitro and in vivo models (cell lines and PDXs) and H3K27-altered DMG, GBM and PFA1/2, H3K27M/EZHIP-mutant patient samples. The average similarity (color intensity) represents an averaged prediction score of all DMGO subsetted tumor cells mapped into a merged dataset consisting of transcriptomic model-derived and patient datasets6,7,8 (n = 14 DMGOs from four independent batches).
Representative DMG including pons-specific OPC states
To further assess the representability of our in vitro grown tumor model (DMGO), we conducted histological comparisons to patient samples sharing the same mutational profile. This showed that H3.3K27M cells (H3K27M+) display loss in H3K27 trimethylation (H3K27me3) in both patient samples and DMGOs (Extended Data Fig. 4d,e), a hallmark of H3K27-altered DMG1,39. Furthermore, our in vitro grown tumors exhibited a global methylation profile closely resembling DMG, distinguishing our tumors from glioblastoma and posterior fossa (PFA1 and PFA2) epyndomas, which present with a similar loss of H3K27M trimethylation caused by H3K27M substitution or EZHIP overexpression, respectively43 (Fig. 2g). We next conducted scRNA-seq profiling of sorted GFP+ tumor cells and, after quality control filtering, analyzed approximately 7,000 cells from 14 DMGOs (Fig. 2h, Extended Data Fig. 5a,b and Supplementary Table 4). The malignant state of these cells was further supported by the analysis of inferred copy-number variation (iCNV) from scRNA-seq data, which showed large-scale amplifications and deletions in these cells compared to healthy cells, including losses of chromosomes 10 and 13 and a gain of chromosome 19q (Extended Data Fig. 5c). Using published DMG references6,8, we first annotated cancer cell states previously described for DMG, including OPC-like, AC-like and MES-like cell states and a population of cycling cells. In line with the early developmental window of our model, we identified only few cells with a more mature OC-like phenotype. Importantly, we identified a major proportion of OPC-like tumor cells that resembled recently defined OPC-like 2 and 3 states, both described as pediatric and pons-specific pre-OPC states8 (Fig. 2h and Extended Data Fig. 5d–f). Furthermore, we found the highest similarity score between DMGOs and primary DMG patient material6, as opposed to cell lines, patient-derived xenografts (PDXs) and material from patients with glioblastoma6 or both PFA subtypes44 (Fig. 2i). Together, this highlights the ability of DMGOs to closely resemble primary DMG tumors.
Pontine glial-specific DMG tumorigenesis
We investigated the mechanisms driving tumorigenesis to identify the attributes of BrOs that appear to be critical for supporting the growth of DMG tumors. We used TrackerSeq, a PiggyBac-based genetic lineage-tracing approach (Extended Data Fig. 6a), and analyzed cancer clone dynamics at 2 months after electroporation. We retrieved 167 unique barcodes from six DMGOs and two healthy BrOs (Extended Data Fig. 6b–h and Supplementary Table 5) and detected individual clones spanning up to approximately 800 cells per barcode, indicative of cancerous transformation (Extended Data Fig. 6i,j). By comparing large versus small, traced clones (Fig. 3a,b) through DEG and METASCAPE analysis, we identified glial specification as a critical feature driving cancer clone expansion in contrast to neuronal specification enriched in smaller clones (for example, synapse organization and modulation of chemical synaptic transmission) (Fig. 3a,b). Larger clones were characterized by higher gene expression of OLIG1, a canonical OPC marker, as well as IER2, JUNB, FOS and EGR1, previously described as key markers of the OPC-like 3 pre-OPC state8. Interestingly, we also identified a higher expression of AQP1, an aquaporin previously shown to be exclusive to ACs in the human brainstem45. Furthermore, analysis of patient data7 revealed AQP1 expression in tumors located in the pons but not in those arising from the cortex or thalamic regions (Extended Data Fig. 6k). These data hint toward a glial-specific tumorigenic process that is, furthermore, dependent on pontine location. This is further illustrated by genes upregulated in large DMG tumor clones mapping back to the glial lineage of BrOs (Fig. 3c), indicating that tumorigenesis is dependent on gliogenesis. To confirm this experimentally, we performed in situ electroporation of our mutation cocktail in unguided cerebral organoids, revealing a reduction in tumor induction (Fig. 3d). In addition, the outgrowth was nondiffuse, with almost no GFP-positive tumorigenic cells carrying the H3.3K27M substitution (Fig. 3e,f). These findings demonstrate that H3.3K27M-driven tumorigenesis depends on the correct anatomical cell identity, which our BrO model recapitulates. Consensus non-negative matrix factorization (cNMF) (Fig. 3g and Extended Data Fig. 6l,m) and lineage relationship analysis identified malignant metagene programs 1 and 2 to be present in the highest number of clones (30 and 26 of 34 clones, respectively; Fig. 3h) and belonging to the OPC-like lineage, emphasizing the central role of this lineage in H3K27M DMG tumorigenesis8. More specifically, we show overlap with the pre-OPC state, OPC-like 2 (Extended Data Fig. 6m). In the context of human early gestation, regionally distinct gene signatures for the glial lineage have been suggested to underlie the strong region-specific occurrence pattern of glial-related diseases, such as DMG35. In line with this, both programs 1 and 2 specifically enrich for the hindbrain pons OC precursor lineage35, as opposed to midbrain and forebrain lineages (Fig. 3i). Collectively, these findings highlight the role of pons glial specification, captured in BrOs, in driving DMG tumorigenesis, emphasizing the need for spatial and developmental precision in modeling DMG and establishing a human-relevant experimental system for therapeutic testing.
a, Volcano plot showing top DEGs in larger and smaller clones. b, METASCAPE results showing selected GO terms from the highest-scoring summary GO terms for small and large clones. c, Presence of large (red) and small (blue) clones in the integrated UMAP representation of developing BrOs, showing a preference for gliogenesis and neurogenesis, respectively. For a–c, n = 14 DMGOs from four independent batches. d, Bar plot quantifying electroporation efficacy (light-gray columns; guided versus unguided, P = 0.342) and tumor induction (dark-gray columns; guided versus unguided, P = 0.023) for guided brainstem organoids as compared to unguided neural organoids at day 11. *P < 0.05, according to a two-tailed independent t-test. Data are shown as the mean ± s.e.m. (n = 35 organoids from three independent experiments per condition; details in Supplementary Table 1). e, Representative images of tumorigenic outgrowth (GFP, green) at weeks 4 and 8 for unguided neural organoids (n = 35 organoids from three independent batches). Scale bars, 500 µm. f, Representative multispectral 3D images of tumor GFP (green), H3K27M (magenta) and DNp53 (yellow) in unguided neural organoids (n = 2 organoids). g, UMAP of traced DMGO cells, colored by their respective highest-scoring cNMF program. h, UpSet plot displaying clonal intersection events. Only clonal families found in more than one cNMF module are depicted and filtered with at least three cells present per unique barcode. Bar plots depict the frequency of each lineage combination (top) and the number of clones that contain each program (left). Coloring of dots matches the cNMF program annotation as in g. i, Heat map presenting the mean cellular enrichment scores of cNMF programs 1 and 2 for forebrain, midbrain and hindbrain or pons oligo lineage signatures in HDBCA35. For g–i, n = 8 DMGOs from three independent batches.
Modeling CAR T cell heterogeneity and functional exhaustion
Given the relevant tumor progression observed in DMGOs, we evaluated their potential as a human in vitro platform for preclinical testing of GD2 CAR T cells (Fig. 1a), motivated by promising yet variable treatment outcomes in a recent first clinical trial in patients with H3K27M-mutant DMG24,25. By exposing untransformed BrOs to GD2 CAR T cells, we first visually inspected with brightfield imaging that the presence of GD2 CAR T cells did not affect the general health of the model (Extended Data Fig. 7a). Next, we confirmed GD2 target expression in DMGO (Extended Data Fig. 7b) and tumor cell killing through confocal imaging of cleaved caspase 3 in GFP+ tumor cells (Fig. 4a). Having established these experimental preconditions, we treated DMGOs 4 months after tumor induction by administrating CD8+ GD2 CAR T cells on days 0 and 7 and monitored T cell activation, measured by interferon-γ (IFNγ) secretion (Extended Data Fig. 7c) and tumor control (Extended Data Fig. 7d,e) over time. Similar to heterogeneous outcomes reported in individuals24,25, we observed an overall partial reduction in tumor burden (Fig. 4b) and heterogeneous response rates over time and between individual DMGOs (Extended Data Fig. 7d,e). As GD2 CAR T cell activation was evident by a robust IFNγ response for all treated DMGOs (Extended Data Fig. 7c), limited response profiles (for example, DMGO179) are unlikely to result from a lack of antigen recognition. Therapy effects could be detected after >1 month of treatment (Fig. 4c), offering advantages for modeling CAR T cell functionality in vitro in a manner that is representative of T cell states at the tumor site in vivo, including potential exhaustion profiles associated with prolonged tumor exposure. To test our model for this purpose, we sequenced over 20,000 GD2 CAR T cells retrieved from DMGOs, as well as unexposed GD2 CAR T cells (Supplementary Table 6). This revealed a substantial level of heterogeneity induced upon DMGO exposure (Fig. 4d). In GD2 CAR T cells retrieved from DMGOs, we identified nine transcriptional states (Fig. 4e) that, on the basis of a combined interrogation of curated gene signatures (Extended Data Fig. 8a), DEGs, DEG-associated Gene Ontology (GO) terms (Extended Data Fig. 8b–f), expression of canonical immune effector (Fig. 4f) and exhaustion markers (Fig. 4g) and comparison to a pan-cancer tumor-infiltrating lymphocyte (TIL) dataset including brain malignancies46 (Extended Data Fig. 8g), reflected different T cell activation, differentiation and effector states. For instance, we identified a GD2 CAR T cell population that, albeit activated (on the basis of HLA gene expression) (Extended Data Fig. 8a), does not fully differentiate toward effector function (undifferentiated; TUND) (Extended Data Fig. 8b,h), as well as an IL-2-responsive population (TIL-2) (Extended Data Fig. 8c,i), probably differentiating into effector T cells (Extended Data Fig. 8i). In addition, we observed an interferon-stimulated gene (ISG)-expressing population (TISG) (Extended Data Fig. 8a), corresponding to ISG-expressing TILs46 (Extended Data Fig. 8j) and considered as an interferon-induced activation state47,48. Other clusters included a CAR T cell population with migrating properties and interconnectivity that appeared to be influenced by its neuronal environment (Extended Data Fig. 8d), as well as proliferating (TPR) (Extended Data Fig. 8e) and metabolically stressed T cells (TMS) (Extended Data Fig. 8a,f). Importantly, we distinguished potential DMG-targeting effector T cell populations on the basis of their cytotoxic profile (Fig. 4f) and putative level of exhaustion (Fig. 4g). While one of these clusters predominantly expressed GZMK (TGZK), cytotoxic T cells (TCYT) expressed GZMB, PRF1 and IFNG (Fig. 4f). In contrast, exhausted T cells (TEX) displayed reduced IFNG and concomitant expression of immune checkpoint genes, LAG3, HAVCR2, TIGIT (ref. 49) and SELPLG50, as well as the transcriptional repressor PRDM1 associated with exhaustion51 (Fig. 4g). Weekly CAR T cell administration (days 0 and 7) did not improve TEX reduction or TCYT enrichment over a single dose (Extended Data Fig. 9a), indicating that exhaustion may set on as early as day 7.
a, Representative multispectral 3D imaging of GD2 CAR T cells (CD3; cyan), DMG tumor cells (GFP; green) and cleaved caspase 3 (cCasp3; red) in DMGOs (n = 2 DMGOs). Scale bar, 5 µm. b, GD2 CAR T cell treatment outcome measured as a relative change in tumor GFP intensity quantified by imaging compared to the start of treatment (100%). DMGOs were left untreated (gray line; n = 1 DMGO) or treated with mock-transduced T cells (black line; n = 2) or GD2 CAR T cells (orange line; n = 4 DMGOs); for each treatment condition, a smoothed trend line of the averaged values at different time points was plotted using the locally estimated scatter plot smoothing algorithm. Arrows indicate the time points of T cell administration. c, Representative images of the tumor GFP signal at the indicated time points for a DMGO subjected to prolonged GD2 CAR T cell treatment administrated on day 0, day 8 and day 15 (n = 1 DMGO). d, Sankey plot illustrating the shift in the relative proportions of unbiasedly identified GD2 CAR T cell clusters before and after DMGO exposure. e, UMAP visualization of annotated GD2 CAR T cell clusters. f, Cytotoxic effector molecule and cytokine gene expression across the GD2 CAR T cell clusters. g, Gene expression of selected exhaustion-associated receptors, ligands and transcription factors across the GD2 CAR T cell clusters. f,g, Dot plot representing the percentage of cells expressing selected genes. The color intensity represents the average scaled gene expression. h,i, Heat map depicting the relative expression of exhaustion markers (h) and exhaustion-associated transcription factors and functional regulators (i) in nonexposed (left) and DMGO-exposed (right) GD2 CAR T cells within the TEX cluster. TF, transcription factor. j, Super-engager signature score on a blue-to-red color scale, showing the enrichment of a previously identified T cell serial killer gene set60 atop UMAP cell embeddings of the GD2 CAR T cell dataset. The dashed outline annotates the embedding of the TCYT GD2 CAR T cell cluster. For d–j, GD2 CAR T cells retrieved from n = 4 treated DMGOs and n = 2 independent batches of unexposed GD2 CAR T cells. k, Dot chart depicting the fold enrichment in tumor killing by NCAM1+ GD2 CAR T cells over NCAM1− GD2 CAR T cells quantified as the change in tumor area detected by GFP compared to the start of treatment (n = 2 DMGOs per treatment condition). l, Percentage of cells per TCYT, TEX and THS cluster for NCAM1− GD2 CAR T cells (left; retrieved from n = 2 DMGOs) and NCAM1+ GD2 CAR T cells (right; retrieved from n = 2 DMGOs).
To confirm that exhaustion detected in our DMGO model reflects representative T cell exhaustion at the tumor site, we compared the TEX phenotype present upon DMGO exposure to preexposure GD2 CAR T cells that, although alleviated by the 4-1BB endodomain, can still display exhaustion features resulting from tonic signaling52. Indeed, a fraction of preexposure GD2 CAR T cells overlapped with our TEX cluster detected upon DMGO exposure (Extended Data Fig. 9b,c). However, separating the cells in this cluster according to experimental condition (Extended Data Fig. 9c) revealed that DMGO-exposed TEX upregulated a wide array of additional exhaustion markers (Fig. 4h), as well as known transcription factors and functional modulators of exhaustion (Fig. 4i) that include those described in cancer patients across TIL datasets (Supplementary Table 7). In addition, overlap with exhaustion markers found in the antigen-driven lymphocytic choriomeningitis virus mouse model of chronic infection53,54, as well as an in vitro model of CAR T cell dysfunction based on continuous antigen exposure55, demonstrates that the observed exhaustion profile is antigen driven (Supplementary Table 7). For in vitro model systems, this has not been achieved in the context of naturally expressed tumor antigen, only through persistent anti-CD3 and anti-CD28 antibody stimulation56 or by using repeated rounds of stimulation with antigen- pulsed57 or overexpressing58 tumor cell lines55. Thus, DMGOs model the functional heterogeneity of CAR T cells, including representative T cell functional exhaustion, an actionable axis for improving outcomes59 and, therefore, critical factor to evaluate preclinically.
NCAM1 selection of short-lived cytotoxic effector CAR T cells
Consistent with their potent cytotoxicity and lack of exhaustion, the TCYT population overlaps with the ‘killer’ gene signature of ‘super-engager’ engineered T cells that we recently identified to have profound tumor-targeting capacity and serial killing behavior in a short-term coculture assay60 (Fig. 4j). As we previously identified NCAM1 as a selection marker to enrich for this population60, we exploited this strategy (Fig. 1a) to further investigate the relevance of this CAR T cell functional profile in a prolonged treatment setting. We sorted GD2 CAR T cells on the basis of NCAM1 expression before DMGO treatment (Extended Data Fig. 9d) and compared tumor control between NCAM1+ and NCAM1− cells (Fig. 4k and Extended Data Fig. 9e). This demonstrated the initial potent antitumor activity of NCAM1+ GD2 CAR T cells, with a 1.4-fold enrichment in tumor control over NCAM1− GD2 CAR T cells on day 2. However, this enhanced potency stabilized between days 5 and 7, with NCAM1− T cells displaying more gradual antitumor activity over time, slightly outperforming NCAM1+ cells by day 7 (Fig. 4k), in line with a higher recovery of NCAM1− cells at day 14 (Extended Data Fig. 9f). To gain insight into potential transcriptomic profiles explaining these differential outcomes, we performed scRNA-seq of NCAM1− and NCAM1+ GD2 CAR T cells and mapped them back to our previously identified GD2 CAR T cell signatures (Extended Data Fig. 9g). This revealed an additional stressed GD2 CAR T cell cluster specific to NCAM1− cells (THS) (Fig. 4l and Extended Data Fig. 9h) that differed from the TMS cluster through expression of heat-shock proteins (Supplementary Table 8) and overlapped with the stress response state identified in TILs that associates with immunotherapy resistance46 (Extended Data Fig. 9i). Further aligning with the initially enhanced tumor control observed (Fig. 4k), NCAM1+ cells showed a 3.3-fold enrichment in TCYT compared to NCAM1− cells (Fig. 4l). However, in line with poor persistence of the cells (Extended Data Fig. 9f), NCAM1+ T cells were additionally enriched for TEX (Fig. 4l), explaining their reduced performance over time (Fig. 4k). Together, this identified NCAM1+ cells as a potent tumor-targeting, yet short-lived effector GD2 CAR T cell population and offers proof of concept for cell selection as a means to narrow CAR T cell functional heterogeneity before administration.
Microglia-enriched DMGO microenvironment
The upregulation of features associated with tissue residency48, including the canonical marker CD103 (ITGAE) used to identify tissue-resident T cells61,62 (Fig. 5a and Supplementary Table 7), underscores the capacity of DMGOs to model T cell performance within tissue. While this may inform strategies to enhance CAR T cell trafficking and tissue residency63, DMGOs lack the myeloid compartment, a key regulator of T cell responses. Therefore, to enhance the complexity of DMGOs, we incorporated microglia, a main component of the DMG tumor microenvironment8. We generated primitive macrophage progenitors (PMPs) from hES cells, previously shown to differentiate into mature microglia in mouse brains64 and human midbrain organoids65, as well as in coculture with neurons66. PMPs similarly integrated into BrOs and, within 7 days, adopted the ramified morphology characteristic of homeostatic microglia67 (Fig. 5b,c). Confirming functional maturation, the cells displayed typical microglia behavior, migrating to sites of myelin injection (Fig. 5d and Supplementary Video 1) and removing myelin through phagocytosis68 (Fig. 5e and Supplementary Video 2). Furthermore, 3 weeks after incorporation in BrOs, above 80% of cells expressed the microglia-specific marker P2RY12 at the protein level (Fig. 5f,g) and scRNA-seq analysis (Supplementary Table 9) demonstrated increased expression of microglia-specific transcription factors69,70 (Fig. 5h). Additionally, they resembled an adult state when referenced against microglia developmental programs identified in mice71 (Fig. 5i), further validating microglia maturation. Comparison to a myeloid cell reference dataset from DMG tumors72 confirmed microglia as opposed to macrophage identity (Fig. 5j).
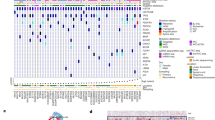
a, Gene expression of tissue-resident markers in nonexposed (top; n = 2 independent batches of unexposed GD2 CAR T cells) and DMGO-exposed (bottom; retrieved from n = 4 DMGOs) GD2 CAR T cells within the TEX cluster. Dot plot representing the percentage of cells expressing selected genes. The color intensity represents the average scaled gene expression. b, Schematic overview of PMP integration in BrOs and DMGOs and treatment with GD2 CAR T cells. c, Representative brightfield images of BrOs for mScarlet+ PMPs (white) 3 or 7 days after integration (n = 19 BrOs). Right, zoomed-in view of area in white insets. Scale bars, 500 µm (main image) and 50 µm (inset). d,e, Live 3D imaging of microglia (orange) and CFSE-labeled myelin debris (green), showing homing of microglia to sites of myelin debris injection (d) and phagocytosis of myelin debris (e) (n = 2 BrOs). Scale bars, 50 µm (d) or 10 µm (e). f, Immunofluorescence 2D images of BrO with microglia integrated for 3 weeks, labeled for DAPI (white), IBA1 (magenta) and P2RY12 (cyan) (n = 1 BrO). Scale bar, 50 µm. g, Quantification of percentage of P2RY12+ cells of total IBA1+ microglia in a BrO slice (n = 1 BrO). h, Heat map depicting the relative expression of microglia-associated transcription factors in PMPs (left) and BrO-derived microglia (right) 3 weeks after integration. i, Heat map depicting representation of microglia developmental stages71 in PMP or microglia derived from BrOs or DMGOs. j, Violin plot showing the expression level of microglia and macrophage gene signatures72 in PMP (left) and microglia derived from BrO (right). Statistical analysis was performed using a two-tailed t-test (microglia, P = 9.4 × 10−42; macrophage, P = 8.6 × 10−10). k, Violin plots showing expression levels of DMG-associated microglia states72 in microglia derived from BrO (blue) or DMGO (green). l, Dot plot showing the relative expression of selected chemokines and genes associated with immunosuppression72 in microglia derived from BrO (left) or DMGO (right). For h–l, microglia were sorted from n = 9 organoids in total (n = 5 BrOs and n = 4 DMGOs) and unexposed PMPs from n = 2 independent experiments (details in Supplementary Table 1).
Microglia incorporation in tumor-bearing DMGOs led to the acquisition of recently described DMG-associated functional phenotypes72, including an IFN-activated, phago lipid and hypoxic state (Fig. 5k). Moreover, compared to BrOs, microglia from DMGOs showed enrichment of GO terms related to antigen presentation and immune responses, such as peptide processing mediated by major histocompatibility complex class II and type I IFN response, previously described as upregulated in DMG-associated microglia and macrophages73 (Extended Data Fig. 10a). These DMG-specific microglia states were accompanied by reduced chemokine expression and upregulation of genes associated with an immunosuppressive profile, in line with patient data72 (Fig. 5l). This was confirmed by protein expression of CD163, associated with an anti-inflammatory state, and SPP1, associated with immunosuppressive lipid-laden macrophages74 (Extended Data Fig. 10b,c). Together, these findings show that, within an appropriate neuronal environment, PMPs differentiate into mature microglia that, in the presence, of DMG tumor cells adopt a DMG-specific immunosuppressive phenotype.
Microglia impact GD2 CAR T cell therapy responses
Correlative clinical data suggest that expansion of the immunosuppressive myeloid compartment may be associated with poor GD2 CAR T cell outcomes25 and myeloid cells, including microglia, are also implicated in CAR T cell-induced toxicity75. To address this experimentally, we performed GD2 CAR T cell treatment in DMGOs with integrated microglia. Confocal imaging showed interactions between GD2 CAR T cells and microglia within tumors (Fig. 6a). Moreover, we observed increased cytokine secretion in the presence of both GD2 CAR T cells and integrated microglia (Fig. 6b). This included upregulated chemokines related to myeloid cell chemotaxis, including MCP3 and CXCL1, as well as myeloid-cell-associated growth factors, such as macrophage colony-stimulating factor (M-CSF). We also observed elevated interleukin (IL)-1α and IL-6 levels, key proinflammatory cytokines linked to CAR T cell (neuro)toxicity and clinically targeted to manage adverse effect24, suggesting that microglia-integrated DMGOs may serve as a relevant model to study CAR T cell–microglia interactions underlying treatment-related toxicity. Furthermore, analysis of DMGO-induced CAR T cell transcriptional heterogeneity in the presence of microglia (Extended Data Fig. 10d and Fig. 6c), showed an increased proportion of the exhausted cluster and identified a microglia-affected GD2 CAR T cell population (TMA), aligning with undifferentiated TILs, as well as naive and tissue-resident memory cells, from the pan-cancer TIL atlas46 (Extended Data Fig. 10e). To validate the low effector profile of the TMA population, we compared curated gene signatures from the same resource46 across GD2 CAR T cell clusters (Fig. 6d and Extended Data Fig. 10f). TMA cells showed reduced activation and effector signatures, including cytotoxicity, but higher senescence-related genes. Thus, the presence of microglia shifts the transcriptional profile of GD2 CAR T cells toward reduced effector function. In line with this, when we monitored CAR T cell-mediated tumor control, it was reduced in integrated-microglia DMGOs (Fig. 6e). Together, these findings demonstrated that microglia integrated in DMGOs yield a suitable representative phenotype for probing CAR T cell function and toxicity. This establishes a direct role for microglia in shaping CAR T cell functional states and impairing tumor control.
a, Representative immunofluorescence 3D images of a 200-µm-thick slice containing microglia, 1 week after start of GD2 CAR T cell treatment. Cells are labeled for DAPI (white), GFP+ DMG tumor cells (green), IBA1+ microglia (orange), CD3+ T cells (cyan) and cCasp3 (red). n = 2 DMGOs. Bottom, zoomed-in view of area in white inset (n = 2 DMGOs). Scale bar, 10 µm. b, Heat map depicting the fold change in concentration of selected cytokines, chemokines and growth factors of DMGOs containing microglia normalized against no microglia on day 3, day 7 and day 14 after GD2 CAR T cell addition (n = 6 DMGOs). TNF, tumor necrosis factor. c, Percentage of cells within GD2 CAR T cell clusters, including a new microglia-affected cluster, for nonexposed GD2 CAR T cells (left), GD2 CAR T cells retrieved from DMGO without (middle) or with integrated microglia (right). d, Heat map highlighting the average scaled expression of curated TIL gene signatures46 in the TMA GD2 CAR T cell cluster. e, Reduction in tumor area (normalized z score per DMGO relative to time point 0) after the addition of GD2 CAR T cells in DMGO without (blue) or with (orange) integrated microglia. The arrow indicates the time point of GD2 CAR T cell administration. Data are shown as the mean ± s.e.m. Statistical analysis at each time point was performed using a linear mixed-effects model, accounting for experimental and organoid variation. Reported P values are two-sided and were adjusted for multiple comparisons using the false discovery rate (Benjamini–Hochberg) (t3, P = 0.04389; t7, P = 0.04389; t10, P = 0.08409; t14, P = 0.0578). For b–e, n = 6 DMGOs with microglia and n = 6 DMGOs without microglia from four independent batches.
Discussion
Here, we showed that morphogen-guided patterning with FGF4 and RA produces BrOs, characterized across region-specific neuronal and glial lineages through benchmarking against recent single-cell organoid and brain atlases34,35,36. BrOs enable spatial and temporal modeling of the developing brainstem, including pons-specific features, the regional origin and niche for H3.3K27M-altered DMG. We demonstrated an interplay between the H3.3K27M substitution and pontine glial fate in driving DMG tumorigenesis, resulting in an experimentally accessible organoid model that recapitulates features of DMG tumors. While DMG patient-derived organoids are emerging as platforms for drug testing60,76,77, our model offers a complementary approach to generate in vitro tumors for this rare and fatal disease, for which tissue samples are limited. Moreover, the use of iPS cells as a cell source establishes the potential for individualized modeling, supporting personalized drug evaluation and direct comparison to clinical outcomes. However, because the model is derived from hES cells and iPS cells, it is limited in recapitulating postnatal tissue, as reported for other neural organoids34,78. Furthermore, the current model does not capture complex intraregional interactions, which are particularly relevant in the pons—a key relay between the forebrain and motor or sensory pathways. Assembloid methodologies79,80, such as linking cortical organoids with BrOs and DMGOs, could improve neuronal health and lineage diversity while enabling modeling of DMG invasion and progression across brain regions in response to neural secretion18,81 and activity17,19,20,82. Such neuronal interplay should be further characterized in DMGOs in the future to validate the model for these critical processes, for instance, through live calcium imaging of synaptic signaling or retrograde labeling of GABAergic neurons using viral tracers17.
Despite current limitations, the model provides a therapeutically relevant platform to interrogate CAR T cell function in DMG tissue-specific context. Prolonged treatment of DMGOs reflects the variable outcomes seen in individuals24,25,83, revealing CAR T cell heterogeneity and functional exhaustion. From this heterogeneity, we identified the most potent yet short-lived CAR T cell population and validated a means to enrich for these cells, offering a potential approach to optimize therapy composition84. This model could also be leveraged to uncover how CAR T cells modulate cancer cell states and reveal mechanisms of acquired resistance that may be cotargeted to improve clinical efficacy.
Given the recognition that the tumor microenvironment can significantly impact treatment response, we integrated microglia, a key immune component in DMG70. In line with representative tumor cell states observed in DMGOs, microglia differentiated into DMG-specific and immunosuppressive profiles72. This enabled an experimental interrogation of microglia impact on CAR T cell functionality, revealing increased exhaustion and a TMA population, marked by stalled differentiation and low effector function. This shift toward dysfunction correlated with reduced tumor control, offering a framework to counteract microglia-induced resistance and enhance antitumor efficacy. While T cell exhaustion might be addressed through immune checkpoint inhibition, strategies targeting the newly identified TMA population require further investigation. Our imaging data revealing direct microglia–CAR T cell interactions, suggest that dissecting the underlying signaling involved could be a critical starting point. Furthermore, similar approaches could be used to assess microglial influence on tumor phenotypes, as prior studies linked MES tumor states to tumor-associated macrophages8.
Altogether, we generated a human brainstem organoid model for DMG with applications toward understanding CAR T cell functionality in the context of the tumor microenvironment that could aid further therapy development for this detrimental disease.
Methods
Ethics
All murine experiments were conducted in compliance with the Animal Welfare Body of the Princess Máxima Center for Pediatric Oncology based on local and international regulations under CCD license AVD39900 202216507. For the use of DMG samples, patients and/or their parents or guardians provided written informed consent according to national laws and in agreement with the declaration of Helsinki (2013). This study was approved by the Institutional Review Board (IVB) and registered under national registry number 2020.142.
Stem cell culture
Brain organoids were generated from three different cell lines encompassing H9 (WA09, WiCell) and H1 (WA01, WiCell) hES cells (both derived from human blastocysts85) and C7-a iPS cells (RUID 06C52463, derived from CD4+ T cells). The iPS cell line C7-a was obtained from Rutgers University Cell and DNA Repository and contains a Cre-inducible H3.3K27M reading frame in the endogenous H3F3A locus13. Cell lines were cultured in mTeSR Plus medium (StemCell Technologies, 100-0276) and incubated at 37 °C with 5% CO2. The cells were grown on Matrigel-coated (Corning, 354277) six-well plates and passaged when 70–80% confluent by nonenzymatic detachment of colonies using Gentle Cell Dissociation Reagent (GCDR) (StemCell Technologies, 100-0485). All cell cultures were routinely tested for the presence of Mycoplasma species.
Embryoid body (EB) formation
Cells were washed with 1× Dubecco’s PBS (Gibco, 14190144), detached with GCDR and spun down at 300g for 5 min, before resuspending in base medium (1:1 advanced DMEM/F-12 medium (Gibco, 12634010) and Neurobasal medium (Gibco, 10888022), supplemented with 1× GlutaMax (Gibco, 35050061)) and counting. A total of 70,000 cells per ml were added to day 0 medium (base medium, 10 µM Y-27632 (ROCKi, AbMole BioScience, M1817) and 4 ng ml−1 FGF2 (PeproTech, 100-18 C)). For EB formation, 7,000 cells in 100 µl of medium were seeded per well of an ultralow-attachment treated U-bottom 96-well plate (Nexcelom, ULA96U020; PHC Europe, MS-9096UZ) and incubated at 37 °C with 5% CO2. From day 2 to day 21, patterning medium (base medium, 1× N2 (Gibco, 17502048) and 1 mg ml−1 heparin solution (StemCell Technologies, 07980)) was used.
Organoid patterning
In week 1, week 1 medium (patterning medium, 50 ng ml−1 FGF2, 1 µM dorsomorphin (DM; StemCell Technologies, 72102), 10 µM SB431542 (SB43; StemCell Technologies, 72232) and 3 µM CHIR99021 (CHIR; StemCell Technologies, 72052)) was used. On day 2, 100 µl of week 1 medium was added per well. On day 5, 100 µl of medium per well was replaced with fresh week 1 medium. In the second week, week 2 medium (patterning medium, 1 µM DM, 10 µM SB43, 3 µM CHIR, 10 ng ml−1 FGF4 (StemCell Technologies, Cat.#78103.1), 10 µM all-trans RA (StemCell Technologies, 72262) and 1 µM purmorphamine (PMA; StemCell Technologies, 72202) was used. On day 7, 190 µl of medium was replaced with fresh week 2 medium and, on day 9, 100 µl of medium was replaced. On day 11, EBs were embedded in 12 µl Matrigel droplets and five droplets were transferred to each well of a 12-well suspension plate (Greiner Bio-One,665102) with 1 ml of week 2 medium and incubated at 37 °C with 5% CO2. In week 3, week 3 medium (patterning medium, 10 ng ml−1 FGF4, 10 µM RA and 1 µM PMA) was used. Until day 21, the medium was refreshed every 2 days with week 3 medium. On day 17, the plates were placed on an orbital shaker inside a 5% CO2 incubator at 37 °C. From day 21 onward, the medium was refreshed every 2–3 days with maturation medium (1:1 advanced DMEM/F-12 medium and neurobasal medium, supplemented with 1× GlutaMax, 0.5× N2, 0.5× B27 without vitamin A (Gibco, 12587010) and 1× penicillin–streptomycin (Pen–Strep, Gibco, 15140122).
In situ electroporation
PCAGPbase, PBCAG_DNp53_IRES_luciferase, PBCAG_PDGFRA-D842V_IRES_eGFP and PBCAG_H3K27M_eGFP plasmids were used to induce tumor growth in hES cell-derived organoids. In iPS cell-derived organoids, the H3K27M-expressing plasmid was replaced with 1.00 µg µl−1 SSi-Cre. PCAGPbase and PB_Venus were used as a control. All plasmids were kindly provided by the T. N. Phoenix laboratory11. For genetic lineage tracing, 1.50 µg µl−1 TrackerSeq86 was added to the plasmid mix.
On day 11, unless stated otherwise, organoids were injected with a mixture of plasmid DNA (1.50 µg µl−1 per plasmid) and 0.1% (w/v) FastaGreen (Merck, F7252-5G) using a FemtoJet 4i (Eppendorf, 5252000013) with the following parameters: injection pressure, 15 hPa; compensation pressure, 5 hPa. Subsequently, electroporation was performed using a NEPA21 Super Electroporator (Nepagene) and CUY650P1 (Nepagene) tweezers with the following parameters: voltage, 50 V; pulse length, 10 ms; pulse interval, 50 ms; number of pulses, four; decay rate, 10%. Transfer pulse parameters were as follows: voltage, 20 V; pulse length, 50 ms; pulse interval, 50 ms; number of pulses, five; decay rate, 40%. Using the impedance (kΩ) measurement of the NEPA21 Super Electroporator, voltage was automatically readjusted to optimize cell perforation and viability per individual organoid. Electroporation was performed by applying a shock twice in orthogonal direction and organoids were incubated at 37 °C with 5% CO2 for at least 2 h to recover.
DMGO orthotopic transplantation
No statistical methods were used to predetermine sample sizes but they are close to those previously published87. NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ (NSG) mice (The Jackson Laboratory, 005557) were housed at 45–65% humidity and 20.5–23.5 °C with a 12-h light–dark cycle, in specific-opportunist-pathogen-free conditions using individually ventilated cages and sterile food and water ad libitum. Ten 3–4-week-old male and female NSG mice were anesthetized using isoflurane/O2 inhalation and transferred to a stereotaxic frame. Eye ointment was applied and 0.05 mg kg−1 buprenorphine was injected subcutaneously. After removing hair from the surgical site, a 1-cm incision was made in the skin to expose the skull and 3 mg kg−1 lidocaine was applied topically. Under a stereo microscope, a Dremel was used to drill a circular groove of 5 mm in the skull above the right cerebral cortex. Cortex buffer was applied before the dura mater and 2 mm3 of brain tissue was removed to accommodate the DMGO transplant. DMGOs were preselected on the basis of GFP signal 1–2 weeks after electroporation and, if too big in size, cut in half before transplantation. After placing the DMGO, the brain was covered with a neuropatch, the skull was closed with dental cement and the wound was closed using skin glue. After surgery, 0.06 mg ml−1 carprofen was provided in the drinking water for 3–5 days and mice were monitored 2–3 times per week for signs of weight loss, lack of grooming and/or reduced mobility.
If mice reached the study (21 days) or humane endpoint according to the monitoring of clinical symptoms, they were put under deep anesthesia by intraperitoneal injection of 75 mg kg−1 ketamine and 1 mg kg−1 medetomidine. Transcardiac perfusion was performed with PBS and 4% paraformaldehyde and, after resection, brains were cut into 300-μm sections using a vibratome. Staining, clearing and imaging were performed as described below. Antibodies used are listed in Supplementary Table 10.
Multispectral large-scale single-cell resolution 3D (mLSR-3D) imaging
Organoids were fixed in 4% paraformaldehyde (Sigma-Aldrich, 441244) for 30 min at 4 °C, washed three times in PBST (1:1,000 Tween-20 in 1× PBS) for 15 min at 4 °C, embedded in 4% low-melting-point agarose (Invitrogen, 16520-050) and sliced into 100–250-µm sections using a Leica VT 1200 S vibratome. mLSR-3D was performed on the sliced organoids as described previously88. All combinations of primary and secondary antibodies used are listed in Supplementary Table 10. The slices were imaged using a Zeiss LSM 880 confocal microscope with a ×25 (numerical aperture (NA): 0.8) objective and Leica Stellaris with ×20 (NA: 0.75) and ×40 (NA: 1.3) objectives. Alternatively, intact organoids were fixed and cleared using the organic solvent-based vDISCO method89 and imaged using a Leica SP8 microscope with a ×16 (NA: 0.6) BABB-compatible objective.
2D formalin-fixed paraffin-embedded (FFPE) tissue imaging
Routine histopathology procedures were followed to obtain FFPE tissue for World Health Organization standardized tumor classification. Patient and organoid material was sliced into 3-µm sections before hematoxylin and eosin and subsequent stainings. Immunohistochemical staining was performed on the Leica BOND RX fully automated research stainer using the bond polymer refine detection kit (Leica, DS9800). Stained tissue sections were analyzed by an experienced neuropathologist.
DNA methylation profiling
The DNA methylation profile of a pooled DMGO sample consisting of three independent replicates was compared to cases of DMG, glioblastoma and PFA ependymoma obtained from published datasets90,91. Data were loaded in R (version 4.3.1), probe filtering was performed using package ChAMP92 and each array platform was processed separately (HumanMethylation450 or EPIC) using the ‘minfi’ method93 and filtering out probes located on single-nucleotide polymorphisms or sex chromosomes or with detection P value > 0.01. Raw β values were merged using function combineArrays and normalized with method BMIQ94. In total, 10,000 probes with the highest s.d. were selected and the Pearson correlation between samples was calculated, weighted by the inverse of variance. This correlation matrix was used to compute a distance matrix, which served as the input for the Rtsne function from the Rtsne package.
Cyclic immunofluorescence imaging
Organoid FFPE sections were deparaffinized in xylene (three times, 3 min each) followed by rehydration in a graded alcohol series for 1 min each (100% twice, 95% twice and 70% once). Sections were washed in deionized water (two times, 1 min each) and put in target retrieval solution, pH 9 (Agilent Dako). Antigen retrieval was performed for 40 min at 95 °C. Sections were allowed to cool to room temperature and washed for 5 min in deionized water followed by storage in PBS until further use. Cyclical immunofluorescence imaging was performed as previously described95. All combinations of primary and secondary antibodies used are listed in Supplementary Table 10. Imaging was performed on a Leica DMi8 Thunder imaging system with an HC PL APO ×20 (NA: 0.80) objective. Images of each cycle were aligned based on DAPI signal using a previously developed tool (https://github.com/Dream3DLab/CycFluoCoreg). The resulting composite images were imported into QuPath (version 0.4.4)96, where nuclei were segmented using a cell expansion of 2.5 μm. An object classifier using RandomTrees was trained for each marker on two separate images. These object classifiers were combined into a composite classifier that was applied to all images. The resulting dataset containing the count of classified cells in each image was exported to R for quantification and visualization.
Transcriptomics and genetic lineage-tracing processing and analysis for BrO and DMGO
The detailed step-by-step approaches were deposited to the protocols.io repository (https://doi.org/10.17504/protocols.io.6qpvrwq13lmk/v1)97.
GD2 CAR T cell expansion and selection
CD8 GD2 CAR T cells (14G2a GD2-4-1BBz CAR) and donor-matched mock-transduced CD8 T cells were produced as previously described98. CAR T cells and mock-transduced T cells were expanded using a rapid expansion protocol99 and cryopreserved after 14 days. T cells were thawed and rested in RPMI-1640 medium, supplemented with GlutaMax, 10% FBS (Thermo Fisher, 10500064), 1% Pen–Strep, 50 U per ml IL-2 (Miltenyi, 130-097-743), 2,000 U per ml IL-7 (Miltenyi, 130-095-367) and 50 U per ml IL-15 (Miltenyi, 130-095-760) for 3 days at 37 °C with 5% CO2. For selection of GD2 CAR T cells based on NCAM1 expression, a similar expansion protocol was used but without the addition of IL-15 and Daudi cells. After resting, cells were washed and stained for 30 min at 4 °C in flow cytometry (FC) buffer (1× PBS with 2% FBS) with live–dead fixable near-IR dead cell stain (1:1,000; Thermo Fisher), CD3–APC (1:80; BD BioLegend, clone SK7) and NCAM1–HiLyte-488 (1:200; QVQ, FSH-10B10). CD3+NCAM1− or CD3+NCAM1+ GD2 CAR T cells underwent fluorescence-activated cell sorting on a BD FACSAria II and were immediately used for experiments.
GD2 CAR T cell treatment
Firstly, 4 months after tumor induction, DMGOs were untreated or treated with 500,000 CD8+ GD2 CAR T cells or mock-transduced CD8+ T cells per DMGO added on days 0 and 7. For prolonged treatment, GD2 CAR T cells were administrated on days 0, 8 and 15. Tumor size was monitored by imaging using a Leica Thunder DMi8 microscope with a ×10 objective. After THUNDER software-mediated computational clearing of the imaging data, tumor size for each time point was quantified using Fiji. Background signal, defined as GFP-negative areas within the organoid, was subtracted. The organoid surface was set as the region of interest (ROI) and mean gray values of the GFP channel for the ROI were calculated. Supernatant of the cocultures was collected and IFNγ concentration was measured with ELISA (R&D Systems, DY285B). Untransformed BrOs were similarly treated and organoid appearance was monitored by brightfield imaging.
DMGO GD2 expression analysis
Firstly, 60 days after electroporation, a DMGO sample was dissected for the tumor region, mechanically dissociated and cultured for two additional weeks to expand tumor cells. Cells were retrieved from the culture plate using StemPro Accutase (Gibco, A1110501) and passed through a 70-µm Flowmi cell strainer (Merck, BAH136800070) to create a single-cell suspension. Dissociated cells were centrifuged at 500g for 5 min at 4 °C and resuspended and washed in FC buffer. Cells were either left unstained or stained with live–dead fixable near-IR dead cell stain (1:1,000; Thermo Fisher) and GD2–PE (1:200; clone 14.G2a, BD Biosciences, 562100) for 30 min at 4 °C. Cells were washed twice in FC buffer, acquired on a Sony SH800s (Sony Biotechnology) and analyzed using FlowJo software (version 10.9.0).
scRNA-seq of GD2 CAR T cells
DMGOs treated with GD2 CAR T cells were dissociated and washed twice with PBS +/+ (Mg2+/Ca2+, 3% FBS). Cell suspensions were filtered using a 70-µm Flowmi cell strainer and stained in FC buffer with CD3–APC (1:80; BD Biosciences, clone SK7) and live–dead fixable near-IR dead cell stain (1:1000; Thermo Fisher) for 30 min at 4 °C. CD3+ T cells were sorted on a CytoFLEX SRT benchtop cell sorter (Beckman Coulter) and immediately processed for scRNA-seq.
Preprocessing and analysis of GD2 CAR T cell scRNA-seq datasets
Doublets were identified and removed using the scDblFinder package100, with default settings. Low-quality cells (>15% mitochondrial content, <200 or >6,500 genes or >35,000 reads) were removed. The Seurat (version 4)101 workflow was used to normalize and scale reads and 3,000 highly variable genes were determined. Cell-cycle confounding effects were eliminated by the removal of cell-cycle-related genes from the variable features of the dataset. Principal component analysis was performed using the ‘RunPCA’ function. The first 30 principal components were used for nonlinear dimensionality reduction applying the UMAP102 method with the ‘RunUMAP’ function of the Seurat package. Clustering analysis was performed using the Seurat package ‘FindNeighbors’ and ‘FindClusters’ functions. To identify subpopulations, marker genes for each cluster were determined through the ‘FindAllMarkers’ function. Markers (adjusted P value < 0.05) were examined to profile genes associated with known CD8 T cell subsets, as well as project previously published signatures. In addition, DEGs were used as input for GO enrichment analysis using the GO resource (https://geneontology.org). To integrate three batches of NCAM1-selected cells, we applied the Seurat-based canonical correlation analysis integration method. As integration features, we used 1,000 variable features from each dataset, along with DEGs between conditions to account for biological variability. To assign cell populations to the clusters identified in Fig. 4e, we estimated the proportion of cells from these clusters for every cluster of the integrated dataset. Newly emerging populations were defined on the basis of differentially expressed markers and their origin, categorized by whether they originated from exposed, unexposed, NCAM1+ or NCAM1− populations. To determine the identity of the TMA cluster, we mapped our GD2 CAR T cell subsets to the CD8+ TIL resource dataset46 using Seurat’s FindTransferAnchors(). The GD2 CAR T cell identities were then transferred to this dataset with TransferData(), retaining only high-confidence predictions (score > 0.5). These transferred identities were used to calculate the proportion of TMA GD2 CAR T cells within each CD8+ TIL subset.
T cell signature projection
To evaluate the expression of established T cell signatures, we used a gene signature specific to serial killer engineered T cells that we previously obtained60. Using the VISION R package103, we computed and visualized the overall enrichment of the identified gene set atop UMAP cell embeddings of our dataset. In addition, we projected our GD2 CAR T cell signatures onto a pan-cancer CD8 TIL atlas46 through T Cell Map (https://singlecell.mdanderson.org/TCM/) using the VISION package. For each GD2 CAR T cell subset, markers obtained through DEG analysis were filtered using an adjusted P value < 0.00001.
PMP generation and integration
PMPs were generated using an adjusted protocol104. Briefly, 70–80% confluent H1 stem cells were detached with GCDR. For EB formation, 7,000 cells were plated per well of an ultralow-attachment treated U-bottom 96-well plate (Greiner Bio-One, 650970) in mTeSR+ (StemCell Technologies, 100-0276) medium, containing 50 μM ROCK inhibitor (Y-27632; AbMole, M1817), 50 ng ml−1 bone morphogenetic protein 4 (StemCell Technologies, 78211), 50 ng ml−1 VEGF (PeproTech, 100-20-100ug) and 20 ng ml−1 SCF (Miltenyi Biotec, 130-093-991). On day 2, fresh medium was added and, on day 4, EBs were transferred to a six-well plate with X-VIVO 15 medium (Lonza, BE02-060F), containing 1× GlutaMax (Gibco), 1× Pen–Strep (Gibco), 100 ng ml−1 M-CSF (PeproTech, 300-25-50ug) and 25 ng ml−1 IL-3 (PeproTech, AF-200-03-10ug). The medium was refreshed once a week. After ~3 weeks, the release of PMPs in the supernatant was observed. PMPs were collected and 100,000–200,000 cells were added per brain organoid in maturation medium (1:1 advanced DMEM/F-12 (Gibco) and Neurobasal (Thermo Fisher) medium, 1× GlutaMax, 0.5× N2 (Gibco), 0.5× B27 without vitamin A (Gibco) and 1× Pen–Strep). Organoids were kept on a microtiter orbital shaker inside a 37 °C 5% CO2 incubator for 1–3 weeks for PMP integration and differentiation. For CAR T cell treatment experiments, organoids were sectioned into 200-µm-thick slices using a vibratome, transferred to a 24-well suspension plate in 750 µl of maturation medium and incubated at 5% CO2 and 37 °C for 3 days. A total of 50,000–200,000 PMPs were added per slice in 750 µl of maturation medium in a 24-well suspension plate for 7 days before adding 200,000 CD8+ GD2 CAR T cells in 750 µl of maturation medium. Tumor size during treatment was monitored by imaging using a Leica DMIL LED FLUO microscope with a ×10 objective.
SORT-seq of microglia and GD2 CAR T cells
BrOs and DMGOs containing microglia and optionally treated with GD2 CAR T cells were dissociated 21 days after initial microglia incorporation as described above for the preparation of scRNA-seq and TrackerSeq libraries. Dissociated cells, control PMPs and unexposed GD2 CAR T cells were washed and stained in FC buffer with CD3–BV421 (1:100; BD Biosciences, clone SK7) and live–dead fixable near-IR dead cell stain (1:1,000; Thermo Fisher) for 30 min at 4 °C. CD3+ T cells and mScarlet+ microglia and PMPs were single-cell-sorted into 386-well plates containing well-specific barcoded primers (Single Cell Discoveries) on a Sony SH800s (Sony Biotechnology). Plates containing sorted cells were immediately snap-frozen on dry ice and processed for SORT-seq by Single Cell Discoveries. Cells were heat-lysed at 65 °C followed by complementary (cDNA) synthesis. All the barcoded material from one plate was pooled into one library and amplified using in vitro transcription. Library preparation was performed following the CEL-Seq2 protocol105 to prepare a cDNA library for sequencing using TruSeq small RNA primers (Illumina). The DNA library was paired-end sequenced on an Illumina Nextseq 500, with high output, using a 1× 75-bp Illumina kit (read 1: 26 cycles, index read: 6 cycles, read 2: 60 cycles).
Myelin phagocytosis assay
CSFE-labeled myelin debris, kindly provided by the L. Akkari lab74, was injected into PMP-integrated organoid slices using a glass needle and a FemtoJet 4i. The slices were immediately imaged on a Leica STELLARIS microscope at 37 °C and 5% CO2 overnight with a time interval of 5–10 min.
Luminex analysis
Protein was detected in the culture supernatant by Luminex. Acquisition of data was performed using a FLEXMAP 3D system (Bio-Rad) with xPONENT 4.3u1 software (Luminex). Data analysis was performed using Bio-Plex Manager 6.2 (Bio-Rad). All assays were performed at the ISO9001:2008-certified MultiPlex Core Facility of the University Medical Center Utrecht.
Statistics and reproducibility
Statistics on bulk sequencing data were computed using built-in functions of R (‘stats’, version 4.3.1) through a one-way analysis of variance with a post hoc Tukey honestly significant difference test. Statistics on electroporation efficiency and tumor induction were calculated using two-tailed independent t-tests (function: t.test). All statistical tests were performed with a confidence interval of at least 95% (α = 0.05). Data distribution was assumed to be normal but this was not formally tested, unless otherwise indicated, and no statistical method was used to predetermine sample size. To evaluate tumor response to GD2 CAR T cells in the presence or absence of microglia, the normalized tumor area at each time point was analyzed using a linear mixed-effects model, accounting for fixed and random effects related to batch and organoid variation. Multiple models were tested and the best-fitting model was selected. P values were adjusted for multiple comparisons using the false discovery rate (Benjamini–Hochberg). The investigators were not blinded to allocation during experiments and outcome assessment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Raw sequencing and methylation data that support the findings of this study were deposited to the European Genome–Phenome Archive under accession codes E-MTAB-15147 and E-MTAB-15559, respectively. Processed sequencing data were deposited to Zenodo (https://doi.org/10.5281/zenodo.16992353)106. Sequencing metadata are provided in Supplementary Tables 2–4, 6 and 9. Source data are provided with this paper.
Code availability
All used R and Python scripts for analysis are available from GitHub (https://github.com/Dream3DLab/DMGO_analysis). Pipelines for analyzing TrackerSeq data can also be found on GitHub (https://github.com/anna-alemany/TrackerSeq_BROs).
References
Louis, D. N. et al. The 2021 WHO classification of tumors of the central nervous system: a summary. Neuro Oncol. 23, 1231–1251 (2021).
Findlay, I. J. et al. Pharmaco-proteogenomic profiling of pediatric diffuse midline glioma to inform future treatment strategies. Oncogene 41, 461–475 (2022).
Schwartzentruber, J. et al. Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 482, 226–231 (2012).
Vuong, H. G., Ngo, T. N. M., Le, H. T. & Dunn, I. F. The prognostic significance of HIST1H3B/C and H3F3A K27M mutations in diffuse midline gliomas is influenced by patient age. J. Neurooncol. 158, 405–412 (2022).
Thomas, B. C. et al. CAR T cell therapies for diffuse midline glioma. Trends Cancer 9, 791–804 (2023).
Filbin, M. G. et al. Developmental and oncogenic programs in H3K27M gliomas dissected by single-cell RNA-seq. Science 360, 331–335 (2018).
Jessa, S. et al. K27M in canonical and noncanonical H3 variants occurs in distinct oligodendroglial cell lineages in brain midline gliomas. Nat. Genet. 54, 1865–1880 (2022).
Liu, I. et al. The landscape of tumor cell states and spatial organization in H3-K27M mutant diffuse midline glioma across age and location. Nat. Genet. 54, 1881–1894 (2022).
Pathania, M. et al. H3.3K27M cooperates with Trp53 loss and PDGFRA gain in mouse embryonic neural progenitor cells to induce invasive high-grade gliomas. Cancer Cell 32, 684–700 (2017).
Larson, J. D. et al. Histone H3.3 K27M accelerates spontaneous brainstem glioma and drives restricted changes in bivalent gene expression. Cancer Cell 35, 140–155 (2019).
Patel, S. K. et al. Generation of diffuse intrinsic pontine glioma mouse models by brainstem targeted in utero electroporation. Neuro Oncol. 22, 381–392 (2019).
Funato, K., Major, T., Lewis, P. W., Allis, C. D. & Tabar, V. Use of human embryonic stem cells to model pediatric gliomas with H3.3K27M histone mutation. Science 346, 1529–1533 (2014).
Haag, D. et al. H3.3-K27M drives neural stem cell-specific gliomagenesis in a human iPSC-derived model. Cancer Cell 39, 407–422 (2021).
Bressan, R. B. et al. Regional identity of human neural stem cells determines oncogenic responses to histone H3.3 mutants. Cell Stem Cell 28, 877–893 (2021).
Pun, M. et al. Common molecular features of H3K27M DMGs and PFA ependymomas map to hindbrain developmental pathways. Acta Neuropathol. Commun. 11, 25 (2023).
Jessa, S. et al. Stalled developmental programs at the root of pediatric brain tumors. Nat. Genet. 51, 1702–1713 (2019).
Barron, T. et al. GABAergic neuron-to-glioma synapses in diffuse midline gliomas. Nature 639, 1060–1068 (2025).
Taylor, K. R. et al. Glioma synapses recruit mechanisms of adaptive plasticity. Nature 623, 366–374 (2023).
Venkatesh, H. S. et al. Neuronal activity promotes glioma growth through neuroligin-3 secretion. Cell 161, 803–816 (2015).
Venkataramani, V. et al. Glutamatergic synaptic input to glioma cells drives brain tumour progression. Nature 573, 532–538 (2019).
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013).
Bian, S. et al. Genetically engineered cerebral organoids model brain tumor formation. Nat. Methods 15, 631–639 (2018).
Hendriks, D. et al. Human fetal brain self-organizes into long-term expanding organoids. Cell 187, 712–732 (2024).
Monje, M. et al. Intravenous and intracranial GD2-CAR T cells for H3K27M+ diffuse midline gliomas. Nature 637, 708–715 (2025).
Majzner, R. G. et al. GD2-CAR T cell therapy for H3K27M-mutated diffuse midline gliomas. Nature 603, 934–941 (2022).
Qian, X. et al. Brain-region-specific organoids using mini-bioreactors for modeling ZIKV exposure. Cell 165, 1238–1254 (2016).
Muguruma, K., Nishiyama, A., Kawakami, H., Hashimoto, K. & Sasai, Y. Self-organization of polarized cerebellar tissue in 3D culture of human pluripotent stem cells. Cell Rep. 10, 537–550 (2015).
Andersen, J. et al. Generation of functional human 3D cortico-motor assembloids. Cell 183, 1913–1929 (2020).
Ye, W., Shimamura, K., Rubenstein, J. L. R., Hynes, M. A. & Rosenthal, A. FGF and Shh signals control dopaminergic and serotonergic cell fate in the anterior neural plate. Cell 93, 755–766 (1998).
Wurst, W. & Bally-Cuif, L. Neural plate patterning: Upstream and downstream of the isthmic organizer. Nat. Rev. Neurosci. 2, 99–108 (2001).
Lu, J. et al. Generation of serotonin neurons from human pluripotent stem cells. Nat. Biotechnol. 34, 89–94 (2016).
Philippidou, P. & Dasen, J. S. Hox genes: choreographers in neural development, architects of circuit organization. Neuron 80, 12–34 (2013).
Fleck, J. S. et al. Resolving organoid brain region identities by mapping single-cell genomic data to reference atlases. Cell Stem Cell 28, 1148–1159.e8 (2021).
He, Z. et al. An integrated transcriptomic cell atlas of human neural organoids. Nature 635, 690–698 (2024).
Braun, E. et al. Comprehensive cell atlas of the first-trimester developing human brain. Science 382, eadf1226 (2023).
Albiach, A. M. et al. Futile wound healing drives mesenchymal-like cell phenotypes in human glioblastoma. Preprint at bioRxiv https://doi.org/10.1101/2023.09.01.555882 (2023).
The Federative International Programme for Anatomical Terminology. Terminologia Anatomica 2nd edn (FIPAT, 2019).
Siletti, K. et al. Transcriptomic diversity of cell types across the adult human brain. Science 382, eadd7046 (2023).
Harutyunyan, A. S. et al. H3K27M induces defective chromatin spread of PRC2-mediated repressive H3K27me2/me3 and is essential for glioma tumorigenesis. Nat. Commun. 10, 1262 (2019).
Mackay, A. et al. Integrated molecular meta-analysis of 1,000 pediatric high-grade and diffuse intrinsic pontine glioma. Cancer Cell 32, 520–537(2017).
Khuong-Quang, D.-A. et al. K27M mutation in histone H3.3 defines clinically and biologically distinct subgroups of pediatric diffuse intrinsic pontine gliomas. Acta Neuropathol. 124, 439–447 (2012).
Siddaway, R. & Hawkins, C. Modeling DIPG in the mouse brainstem. Neuro Oncol. 22, 307–308 (2020).
Mariet, C. et al. Posterior fossa ependymoma H3 K27-mutant: an integrated radiological and histomolecular tumor analysis. Acta Neuropathol. Commun. 10, 137 (2022).
Gillen, A. E. et al. Single-cell RNA sequencing of childhood ependymoma reveals neoplastic cell subpopulations that impact molecular classification and etiology. Cell Rep. 32, 108023 (2020).
Gao, J. et al. Cellular localization of aquaporin-1 in the human and mouse trigeminal systems. PLoS ONE 7, e46379 (2012).
Chu, Y. et al. Pan-cancer T cell atlas links a cellular stress response state to immunotherapy resistance. Nat. Med. 29, 1550–1562 (2023).
Zheng, L. et al. Pan-cancer single-cell landscape of tumor-infiltrating T cells. Science 374, abe6474 (2021).
Szabo, P. A. et al. Single-cell transcriptomics of human T cells reveals tissue and activation signatures in health and disease. Nat. Med. 10, 4706 (2019).
Anderson, A. C., Joller, N. & Kuchroo, V. K. Lag-3, Tim-3, and TIGIT: co-inhibitory receptors with specialized functions in immune regulation. Immunity 44, 989–1004 (2016).
Tinoco, R. et al. PSGL-1 is an immune checkpoint regulator that promotes T cell exhaustion. Immunity 44, 1190–1203 (2016).
Shin, H. et al. A role for the transcriptional repressor Blimp-1 in CD8+ T cell exhaustion during chronic viral infection. Immunity 31, 309–320 (2009).
Long, A. H. et al. 4-1BB costimulation ameliorates T cell exhaustion induced by tonic signaling of chimeric antigen receptors. Nat. Med. 21, 581–590 (2015).
Beltra, J.-C. et al. Developmental relationships of four exhausted CD8+ T cell subsets reveals underlying transcriptional and epigenetic landscape control mechanisms. Immunity 52, 825–841 (2020).
Khan, O. et al. TOX transcriptionally and epigenetically programs CD8+ T cell exhaustion. Nature 571, 211–218 (2019).
Good, C. R. et al. An NK-like CAR T cell transition in CAR T cell dysfunction. Cell 184, 6081–6100 (2021).
Jenkins, E., Whitehead, T., Fellermeyer, M., Davis, S. J. & Sharma, S. The current state and future of T-cell exhaustion research. Oxf. Open Immunol. 4, iqad006 (2023).
Trefny, M. P. et al. Deletion of SNX9 alleviates CD8 T cell exhaustion for effective cellular cancer immunotherapy. Nat. Commun. 14, 86 (2023).
Vardhana, S. A. et al. Impaired mitochondrial oxidative phosphorylation limits the self-renewal of T cells exposed to persistent antigen. Nat. Immunol. 21, 1022–1033 (2020).
Chow, A., Perica, K., Klebanoff, C. A. & Wolchok, J. D. Clinical implications of T cell exhaustion for cancer immunotherapy. Nat. Rev. Clin. Oncol. 19, 775–790 (2022).
Dekkers, J. F. et al. Uncovering the mode of action of engineered T cells in patient cancer organoids. Nat. Biotechnol. 41, 60–69 (2023).
Ganesan, A.-P. et al. Tissue-resident memory features are linked to the magnitude of cytotoxic T cell responses in human lung cancer. Nat. Immunol. 18, 940–950 (2017).
Schenkel, J. M. & Masopust, D. Tissue-resident memory T cells. Immunity 41, 886–897 (2014).
Jung, I.-Y. et al. Tissue-resident memory CAR T cells with stem-like characteristics display enhanced efficacy against solid and liquid tumors. Cell Rep. Med. 4, 101053 (2023).
Xu, R. et al. Human iPSC-derived mature microglia retain their identity and functionally integrate in the chimeric mouse brain. Nat. Commun. 11, 1577 (2020).
Sabate-Soler, S. et al. Microglia integration into human midbrain organoids leads to increased neuronal maturation and functionality. Glia 70, 1267–1288 (2022).
Haenseler, W. et al. A highly efficient human pluripotent stem cell microglia model displays a neuronal-co-culture-specific expression profile and inflammatory response. Stem Cell Rep. 8, 1727–1742 (2017).
Sabogal-Guaqueta, A. M. et al. Brain organoid models for studying the function of iPSC-derived microglia in neurodegeneration and brain tumours. Neurobiol. Dis. 203, 106742 (2024).
Hughes, A. N. & Appel, B. Microglia phagocytose myelin sheaths to modify developmental myelination. Nat. Neurosci. 23, 1055–1066 (2020).
Gosselin, D. et al. An environment-dependent transcriptional network specifies human microglia identity. Science 356, eaal3222 (2017).
Park, D. S. et al. iPS-cell-derived microglia promote brain organoid maturation via cholesterol transfer. Nature 623, 397–405 (2023).
Matcovitch-Natan, O. et al. Microglia development follows a stepwise program to regulate brain homeostasis. Science 353, aad8670 (2016).
Andrade, A. F. et al. Immune landscape of oncohistone-mutant gliomas reveals diverse myeloid populations and tumor-promoting function. Nat. Commun. 15, 7769 (2024).
Lin, G. L. et al. Non-inflammatory tumor microenvironment of diffuse intrinsic pontine glioma. Acta Neuropathol. Commun. 6, 51 (2018).
Kloosterman, D. J. et al. Macrophage-mediated myelin recycling fuels brain cancer malignancy. Cell 187, 5336–5356 (2024).
Vinnakota, J. M. et al. Targeting TGFβ-activated kinase-1 activation in microglia reduces CAR T immune effector cell-associated neurotoxicity syndrome. Nat. Cancer 5, 1227–1249 (2024).
Sarnow, K. et al. Neuroimmune-competent human brain organoid model of diffuse midline glioma. Neuro Oncol. 27, 369–382 (2025).
Deligne, C. et al. Establishing a living biobank of pediatric high-grade glioma and ependymoma suitable for cancer pharmacology. Neuro Oncol. 27, 1325–1340 (2025).
Gordon, A. et al. Long-term maturation of human cortical organoids matches key early postnatal transitions. Nat. Neurosci. 24, 331–342 (2021).
Birey, F. et al. Assembly of functionally integrated human forebrain spheroids. Nature 545, 54–59 (2017).
Kim, J. et al. Human assembloid model of the ascending neural sensory pathway. Nature 642, 143–153 (2025).
Qin, E. Y. et al. Neural precursor-derived pleiotrophin mediates subventricular zone invasion by glioma. Cell 170, 845–859 (2017).
Venkatesh, H. S. et al. Electrical and synaptic integration of glioma into neural circuits. Nature 573, 539–545 (2019).
Vitanza, N. A. et al. Intracerebroventricular B7-H3-targeting CAR T cells for diffuse intrinsic pontine glioma: a phase 1 trial. Nat. Med. 31, 861–868 (2025).
Albelda, S. M. CAR T cell therapy for patients with solid tumours: key lessons to learn and unlearn. Nat. Rev. Clin. Oncol. 21, 47–66 (2024).
Thomson, J. A. et al. Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147 (1998).
Bandler, R. C. et al. Single-cell delineation of lineage and genetic identity in the mouse brain. Nature 601, 404–409 (2022).
Ishahak, M. et al. Genetically engineered brain organoids recapitulate spatial and developmental states of glioblastoma progression. Adv. Sci. (Weinh.) 12, e2410110 (2025).
Ineveld, R. L. V. et al. Multispectral confocal 3D imaging of intact healthy and tumor tissue using mLSR-3D. Nat. Protoc. 17, 3028–3055 (2022).
Cai, R. et al. Whole-mouse clearing and imaging at the cellular level with vDISCO. Nat. Protoc. 18, 1197–1242 (2023).
Auffret, L. et al. A new subtype of diffuse midline glioma, H3 K27 and BRAF/FGFR1 co-altered: a clinico-radiological and histomolecular characterisation. Acta Neuropathol. 147, 2 (2024).
Capper, D. et al. DNA methylation-based classification of central nervous system tumours. Nature 555, 469–474 (2018).
Tian, Y. et al. ChAMP: updated methylation analysis pipeline for Illumina BeadChips. Bioinformatics 33, 3982–3984 (2017).
Aryee, M. J. et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 30, 1363–1369 (2014).
Teschendorff, A. E. et al. A beta-mixture quantile normalization method for correcting probe design bias in Illumina Infinium 450 k DNA methylation data. Bioinformatics 29, 189–196 (2012).
Watson, S. S. et al. Microenvironmental reorganization in brain tumors following radiotherapy and recurrence revealed by hyperplexed immunofluorescence imaging. Nat. Commun. 15, 3226 (2024).
Bankhead, P. et al. QuPath: open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Ariese, H. C. R. et al. Comprehensive transcriptomic analysis of brainstem-regionalized organoids and associated diffuse midline glioma organoids. protocols.io https://doi.org/10.17504/protocols.io.6qpvrwq13lmk/v1 (2025).
Andersch, L. et al. CD171- and GD2-specific CAR-T cells potently target retinoblastoma cells in preclinical in vitro testing. BMC Cancer 19, 895 (2019).
Marcu-Malina, V. et al. Redirecting αβT cells against cancer cells by transfer of a broadly tumor-reactive γδT-cell receptor. Blood 118, 50–59 (2011).
Germain, P.-L., Lun, A., Meixide, C. G., Macnair, W. & Robinson, M. D. Doublet identification in single-cell sequencing data using scDblFinder. F1000Res. 10, 979 (2021).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
McInnes, L., Healy, J. & Melville, J. UMAP: uniform manifold approximation and projection for dimension reduction. Preprint at https://doi.org/10.48550/arXiv.1802.03426 (2018).
DeTomaso, D. et al. Functional interpretation of single cell similarity maps. Nat. Commun. 10, 4376 (2019).
Gutbier, S. et al. Large-scale production of human iPSC-derived macrophages for drug screening. Int. J. Mol. Sci. 21, 4808 (2020).
Hashimshony, T. et al. CEL-Seq2: sensitive highly-multiplexed single-cell RNA-Seq. Genome Biol. 17, 77 (2016).
Ariese, H. C. R. & Alieva, M. Transcriptomic profiling of BrO, DMGO and GD2 CAR T cells: processed datasets. Zenodo https://doi.org/10.5281/zenodo.15356013 (2025).
Acknowledgements
All imaging was performed at the Princess Máxima Imaging Center. We thank the Princess Máxima Center Single-Cell Genomics Facility, the Leiden Genome Technology Center and Single Cell Discoveries for performing scRNA-sequencing, R. Moeniralam and E. de Boed from the Princess Máxima Center Pathology Diagnostic Laboratory for performing immunohistochemical staining and the FC facilities at the Princess Máxima Center and Laboratory of Translational Immunology and M. Nicolasen for cell sorting. We thank J. Lammers for providing gene expression data from patient material, Z. Odé for her collaboration on establishing cerebral organoids, H. R. Johnson and A. Zomer for assistance with in vivo experiments and J. Bunt for sharing his knowledge on neural development. This work was financially supported by the Princess Máxima Center for Pediatric Oncology, Oncode Institute, the Children Cancer Free (KiKa) Foundation (grant no. 473) and the Charlie Teo Foundation (Research Rebel-Alegra's Army grant). A.C.R was supported by an European Research Council starting grant (2018 project, no. 804412) and N.B. was supported by a research grant from Stichting Proefdiervrij.
Author information
Authors and Affiliations
Contributions
N.B. and A.C.R conceptualized the work with critical input from H.C.R.A., A.K.L.W., N.D., C.R.M., H.G.S., C.H., M.A. and A.A. and wrote the manuscript with support from E.J.W. H.C.R.A., C.H. and N.B. grew pontine organoids and performed DMG tumor induction. H.C.R.A., C.R.M., N.B., S.d.B. and M.A. analyzed bulk and scRNA-seq datasets with critical input from H.G.S. and F.K. N.D., H.C.R.A. and N.B. performed and analyzed lineage-tracing and cNMF experiments supervised by A.A. and A.C.R. C.M. provided TrackerSeq and critical input for analysis of lineage-tracing experiments. N.B., H.C.R.A. and M.B.R. performed multispectral 3D imaging using specific technology provided by A.E. T.J.M.v.d.B. performed cyclic immunofluorescence imaging, R.V.U.C. performed live-cell imaging, D.J.K. performed FC and S.P. and E.M. performed DNA methylation profiling. A.K.L.W. and C.H. integrated microglia and performed all GD2 CAR T cell experiments with help from A.M.C. for T cell expansion and L.C.D.E.G. for cell sorting. A.K. provided GD2 CAR T cells and J.K., Z.S. and S.N. provided protocols for T cell rapid expansion. E.J.W., A.K.L.W., F.K., E.v.V. and M.A. analyzed DMGO CAR T cell treatment data, including scRNA-seq analysis of GD2 CAR T cells and microglia. M.E.G.K. contributed critical knowledge on DMG histopathological features and performed immunohistochemical staining on DMGOs. M.R. and M.K. provided healthy brain organoids as a refence dataset for iCNV analysis and knowledge related to brain organoid generation. S.d.B. offered critical computational support and infrastructure for data analysis.
Corresponding author
Ethics declarations
Competing interests
A.C.R. and N.B. are listed as inventors on a pending patent related to the novel BrO model (P382144NL). A.K.L.W., E.J.W., M.A. and A.C.R. are listed as inventors on a pending patent related to the development of marker-based T cell selection (P102253NL). The other authors declare no competing interests.
Peer review
Peer review information
Nature Cancer thanks Harry Hill and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Brainstem organoid specification and reproducibility.
a, Overview of patterning approach and corresponding representative brightfield images of brainstem organoids over time. n = 3 BrOs for early time points, n = 15 BrOs for timepoint 60 days, scale bar of single organoids and multiple organoids is 500 µm and 2 mm, respectively. b, Schematic representation of a human foetal brain in gestational week (GW) 5 with indicated morphogens influencing the differentiation of hindbrain rhombomeres (r) and their HOX gene code respectively. c, Heatmap showing relative bulk RNA expression of homeobox (HOX) genes at week 2 and 3 depicted as log2 fold change normalized to week 1. n = 3 pooled BrOs per batch. d, Boxplot representation of Spearman’s rank coefficient between different organoid batches from week 2 to 12. Box plots show the median (center line, 50th percentile), the box spans the interquartile range (25th–75th percentiles), whiskers extend to the most extreme observations within 1.5×IQR of the quartiles (defining the plotted minima and maxima), points beyond this range are not displayed. n = 3 organoids pooled per batch from n = 3 independent batches per timepoint. e, PCA of organoids at different timepoints from week 1 to 12 (grey scale) and derived from hESCs (circles) or iPSCs (square). n = 3 organoids pooled per batch from n = 3 independent batches for weeks 2-12; n = 3 organoids pooled from n = 2 independent batches for week 1 and n = 3 organoids pooled from n = 1 batch for iPSC-derived.
Extended Data Fig. 2 Quality control and integration of brainstem organoids time course scRNA-seq data.
a, Unintegrated UMAP representation of developing brainstem organoids, colored by collection day. Bargraph (upper left) depicts the contribution of each day to the final dataset. b, Violin plots showing the number of genes, number of counts, percentage of mitochondrial genes and percentage of ribosomal genes after filtering on a sample-by-sample basis. Box plots inside violin plots show the median (center line, 50th percentile), the box spans the interquartile range (25th–75th percentiles), whiskers extend to the most extreme observations within 1.5×IQR of the quartiles (defining the plotted minima and maxima). c, scIB integration benchmarking of different assessed integration methods, showing metrics for preservation of biological variation and batch correction. d, scPoli latent embedding of brainstem organoid cells, colored by timepoint (top) and snapseed annotation (bottom). e, scPoli integrated UMAP of brainstem organoids, colored by timepoint. f, scPoli integrated UMAP representation showing three age bins, early (day 5 – day 20), mid (day 30 – day 60) and late (day 90 – day 120). g, Matrixplot of scPoli (HNOCA34) or scANVI (HDBCA35) mediated label transfer annotation compared to the final annotation. For panels a-g, n = 84 organoids in total with 5-24 organoids pooled per timepoint, see Supplementary Table S1 for details.
Extended Data Fig. 3 scRNA-seq and 3D imaging characterization of brainstem organoids.
a, Areaplots showing relative ratios of the cell cycle phases over the different timepoints. b, Dotplot showing the expression of selected markers for each cell type annotation, colored by their respective cell class. FOXG1 is represented in bold, indicating absence of expression. c, Annotation of neurotransmitter transporters (NTT) for cells assigned as neuroblast or neuron. d, Comparison of NTT found in the brainstem organoid dataset and the HDBCA35 pons. For panels a-d, n = 84 organoids in total with 5-24 organoids pooled per timepoint, see Supplementary Table S1 for details. e, Representative 3D confocal image of a 200 µm thick organoid slice at day 110 labelled for TUBB3 (orange) and TPH2 (blue). n = 2 BrOs. White insert indicates zoom area displayed on the right. Overview image and zoom scale bars 250 µm and 25 µm, respectively. f, Representative optical section of immunofluorescent 3D imaging of a 200 µm thick organoid slice at week 16 labelled for neurofilament (NF, white), GFAP (red-to-white gradient) and AQP4 (green) (left) or for DAPI (white) and OLIG2 (red) (right). n = 2 BrOs, scale bars = 50 µm. g, Glycolysis scores over the HNOCA34 and BrO datasets. Dashed line represents the mean score over all HNOCA datasets. h, Glycolysis scores for the OPC (left) and Glioblast (right) lineages in the HNOCA and brainstem organoids (BrOs), statistically analyzed using a permutation test. i, DEG analysis comparing Oligo (left) and Glioblast (right) from the HNOCA and brainstem organoids (BrOs) to the HDBCA counterparts. j, UMAP of the HDBCA, colored by the brainstem organoid presence scores. k, Proportional brainstem organoid presence scores for the HDBCA, showing the ratio of cells being represented in brainstem organoids (BrOs) per annotated cell class. For panels g-k, n = 84 organoids in total with 5-24 organoids pooled per timepoint, see Supplementary Table S1 for details.
Extended Data Fig. 4 Tumorigenic growth and patient-representability of DMGO.
a, b, Representative image of GFP expression (a; tumor-inducing mix) or control (b; PiggyBac backbone including CAG-mVenus) as a measure of tumor outgrowth at week 6. n = 3 DMGOs and 3 controls, scale bars = 1 mm. c, Representative 3D confocal image of a 300 µm brain section of a DMGO-transplanted NSG mouse developing tumor outgrow labelled for CD31 (red), tumor-GFP (green) and DAPI (grey). n = 2 mice, scale bar = 500 µm. d, Representative routine histopathological characterization of a DMG patient tumor sample harbouring H3.3K27M, TP53 and PDGFRA mutations. Top panels; staining for H3K27M and H3K27me3. Bottom panels; Haematoxylin and eosin (HE), glial fibrillary acidic protein (GFAP) and neurofilament (NF). n = 7 patients. e, Representative routine histopathological characterization of a DMGO at day 120. Top panels; staining for H3K27M and H3K27me3. Bottom panels; Haematoxylin and eosin (HE), glial fibrillary acidic protein (GFAP) and neurofilament (NF). n = 5 DMGOs.
Extended Data Fig. 5 Quality control, processing and analysis of DMGO scRNA-seq data.
a, Violin plots showing the number of genes, number of counts, percentage mitochondrial genes and percentage of ribosomal genes after filtering on a sample-by-sample basis. Box plots inside violin plots show the median (center line, 50th percentile), the box spans the interquartile range (25th–75th percentiles), whiskers extend to the most extreme observations within 1.5×IQR of the quartiles (defining the plotted minima and maxima). b, Unintegrated UMAP representation of DMGO cells, colored by sequencing batch. c, iCNV profile of DMGO cells showing chromosomal aberrations compared to healthy cells from the BrO time course data shown in Fig. 1e. Orange depicts tumor and green represents healthy cells. d, Dotplot showing marker gene expression for different tumor states, color-coded for their respective annotation. e, Matrixplot with the obtained tumor state annotations compared to reference datasets7,8. f, Mapping scores of DMGO cells in Liu et al.8, showing high presence of the annotated cell states, except for cycling cells. For panels a-f, n = 14 DMGOs from 4 independent batches.
Extended Data Fig. 6 Barcode representation after quality control and filtering and cNMF program annotation.
a, Schematic representation of the applied approach for simultaneously recovering transcriptomic information, HTO hashtags and lineage barcodes from DMGOs on a single cell level. A nested PCR strategy was applied for the TrackerSeq barcode. b–f, Quality control assessment and filters used to select barcodes from experimental replicate 1 (top) and experimental replicate 2 (bottom). Histogram depicting the total number of UMI counts prior to any filtering (b). Histogram displaying read counts and the cut-off (red dashed line) set as a minimum total read count of log102 and log103 for experimental replicate 1 and 2, respectively (c). Scatter plot depicting read counts plotted against UMI counts and the applied threshold for minimum total read counts indicated (red dashed line) (d). Histogram depicting the mean oversequence per barcode and thresholding applied (dashed blue line) (e). Scatter plot depicting read counts plotted against max mean oversequence showing both thresholds applied (f). g, UMAP embedding of lineage-traced DMGOs and BrO controls. Cells are colored according to unique lineage barcodes. Cells without barcodes are presented in gray. h, Representation of each DMGO and BrO control in the final UMAP embedding. i, Bargraph depicting the total number of cells for each clonal barcode after applying the filtering of >3 cells per clonal family. j, Pie charts of the relative size of each recovered clonal family among all barcoded cells per sample used for the large versus small clone comparison. Percentage is depicted for clonal families that are equal to, or above 20%, which are defined as large clones. For panel b-j, n = 14 DMGOs and n = 2 BrOs, see Supplementary Table S1 for details. k, Dotplot showing normalized expression of AQP1 and AQP4 in DMG cells7. In contrast to AQP1, AQP4 - a canonical AC-like marker - was present in DMG tumors across all locations. l. Module scores of gene programs as derived from cNMF projected onto the UMAP of lineage-traced cells. m, Heatmap of Jaccard index scores, indicating the overlap of the cNMF derived programs to tumor state annotations. For panels b-j, l, m, n = 14 DMGOs and n = 2 BrOs, see Supplementary Table S1 for details. For panel l and m, n = 14 DMGOs and n = 2 BrOs, see Supplementary Table S1 for details.
Extended Data Fig. 7 DMGO GD2 expression and CAR T cell mediated tumor control.
a, Representative brightfield images at day 14 of untransformed brainstem organoids (BrOs) left untreated (n = 8 BrOs) or exposed to GD2 CAR T cells (n = 8 BrOs) at day 0 and 7. b, DMGO tumor cell GD2 expression (orange) analyzed by flow cytometry compared to an unstained control (black). n = 1 DMGO. c, d, IFNγ levels measured in the culture supernatant (c) and GD2 CAR T cell treatment outcome measured as tumor GFP intensity relative to the start of treatment (day 0, 100%) with a smoothed line trend plotted between the values at different timepoints for each DMGO using the LOESS algorithm (d). DMGOs were either left untreated (gray line, n = 1 DMGO), treated with mock transduced T cells (black lines, n = 2 DMGOs), or GD2 CAR T cells (orange lines, n = 4 DMGOs). Arrows indicate the timepoints of T cell administration. e, Images of tumor GFP signal on day 0, 7, 10 and 14 for an untreated DMGO (n = 1 DMGO) and DMGOs treated with mock transduced T cells (n = 2 DMGOs), or GD2 GAR T cells (n = 4 DMGOs). GD2 CAR T cells and mock transduced T cells were administrated at day 0 and 7. For panels c-e, DMGOxxx indicates individual DMGO sample ID for reference across data figures.
Extended Data Fig. 8 Key marker genes, GO terms and reference data projection of GD2 CAR T cell clusters.
a, Dot plot showing key marker gene expression (selected from the top 20 DEGs) across the GD2 CAR T cell clusters. Dot size is proportional to the percentage of cells expressing a gene and color intensity to the average scaled gene expression. Grid colors highlight genes that are closely related in function; HLA genes (green), metabolic stress-related genes (red) and ISGs (blue). b–f, Selected significant GO terms associated with the DEGs of the TUND (b), TIL-2 (c), TMI (d), TPR (e) and TMS (f) GD2 CAR T cell clusters. g, UMAP visualization of the CD8+ TIL clusters from the pan-cancer atlas46 used as a reference dataset. Annotated clusters are highlighted because of their overlap with, or use in defining the GD2 CAR T cell clusters. h–j, Marker gene signatures (DEG analysis adjusted p-value < 0.00001) of the TUND (h), TIL-2 (i) and TISG (j) GD2 CAR T cell clusters projected onto the CD8+ TIL dataset from g. For panels a-j, GD2 CAR T cells retrieved from n = 4 treated DMGOs.
Extended Data Fig. 9 Non-exposed, DMGO-exposed and NCAM1 subset characterization of GD2 CAR T cells.
a, Percentage of cells within GD2 CAR T cell clusters, including those identified in Fig. 4e, a cluster enriched in non-exposed cells, and the NCAM1− specific THS cluster. Data is shown separately for GD2 CAR T cells retrieved from DMGO at day 14 after administration weekly (at day 0 and 7, left; n = 4 DMGOs) or once (at day 0, right; n = 2 DMGOs). b, UMAP representation of the integrated GD2 CAR T cell dataset, illustrating the distribution of non-exposed GD2 CAR T cells. c, UMAP embedding of DMGO-exposed (pink) and non-exposed (green) GD2 CAR T cells within the TEX cluster. For panel b and c, GD2 CAR T cells retrieved from n = 4 DMGOs and n = 2 independent batches of unexposed GD2 CAR T cells. d, Applied gating strategy (top panels) and obtained purity (bottom panels) of sorted NCAM1− and NCAM1+ GD2 CAR T cells. e, Representative images of tumor control measured by GFP imaging in DMGOs treated with sorted NCAM1− (top) or NCAM1+ (bottom) GD2 CAR T cells. f, Number of NCAM1− and NCAM1+ GD2 CAR T cells retrieved from each DMGO sample after two weeks of treatment. g, Proportion of cells within GD2 CAR T cell clusters, including those identified in Fig. 4e, a cluster enriched in non-exposed cells, and the NCAM1− specific THS cluster. Data is shown separately for NCAM1− GD2 CAR T cells (left) and NCAM1+ GD2 CAR T cells (right). h, Selected significant GO terms associated with the DEGs of the THS NCAM1− specific GD2 CAR T cell cluster. i, Upregulated marker gene signature (DEG analysis adjusted p-value < 0.05; avg_log2FC > 0) of the THS GD2 CAR T cell cluster projected onto the pan-cancer CD8+ TIL dataset46. For panels d-i, n = 2 DMGOs for NCAM1− and n = 2 DMGOs for NCAM1+ cells.
Extended Data Fig. 10 Microglia characterization in brainstem organoids and DMGOs and transcriptomic phenotype of microglia-affected T cells (TMA).
a, Selected significant GO terms associated with DEGs in microglia derived from BrOs (left; n = 5 BrOs) or DMGOs (right; n = 4 DMGOs). b, Representative Immunofluorescent 2D images of DMGOs with microglia, 3 weeks after integration, labeled for DAPI (white), IBA1 (green), SPP1 (magenta) and CD163 (blue). n = 3 DMGOs, scale bar = 50 µm. c, Quantification of the percentage of CD163+ (left) or SPP1+ cells (right) in IBA1+ microglia in DMGOs. n = 3 DMGOs. d, Gating strategy used to sort mScarlet+ PMP/microglia and CD3+ GD2 CAR T cells for scRNA-seq. e, Bargraph depicting the proportion of TMA GD2 CAR T cells matched to identified pan-cancer CD8 TIL subsets46. f, Heatmap depicting the average scaled expression of curated gene signatures from the CD8 TIL reference dataset46 across the GD2 CAR T cell clusters. For panels d-f, n = 6 DMGOs with microglia and n = 6 DMGOs without microglia.
Supplementary information
Supplementary Tables (download XLSX )
Supplementary Tables 1–10.
Supplementary Video 1 (download MP4 )
Microglia home to sites of myelin debris injection. Live 3D imaging of microglia (orange) and CFSE-labelled myelin debris (green). Areas of myelin debris injection are outlined by white dashed lines. Comparison of microglia homing at 0 and 14.5 h provided at the end.
Supplementary Video 2 (download MOV )
Microglia phagocytose myelin debris. Live 3D imaging of microglia (orange) and CFSE-labelled myelin debris (green). White arrows indicate myelin debris that is phagocytosed by microglia outlined by white dashed lines.
Source data
Source Data Fig. 1 (download XLSX )
Source data and mean gains for each HDBCA cluster in the BrO model.
Source Data Fig. 2 (download XLSX )
Source data and electroporation efficiency data, including statistical analysis and number of tumor cells positive for H3K27M, p53 and PDGFRA.
Source Data Fig. 3 (download XLSX )
Source data and electroporation efficiency data, including statistical analysis, DEGs and selected GO terms for large versus small clones, gene lists derived from cNMF and oligo lineage signatures derived from Braun et al.39.
Source Data Fig. 4 (download XLSX )
Source data, tumor GFP intensity and fold enrichment in tumor control for NCAM1+ over NCAM1− GD2 CAR T cells.
Source Data Fig. 5 (download XLSX )
Source data.
Source Data Fig. 6 (download XLSX )
Source data.
Source Data Extended Data Fig. 3 (download XLSX )
Source data and DEGs from single-cell sequencing of glioblasts and OPCs from BrO, HNOCA and HDBCA datasets.
Source Data Extended Data Fig. 6 (download XLSX )
Source data and gene lists derived from cNMF.
Source Data Extended Data Fig. 8 (download XLSX )
Source data and DEGs from the GD2 CAR T clusters.
Source Data Extended Data Fig. 9 (download XLSX )
Source data and DEGs from the GD2 CAR T clusters.
Source Data Extended Data Fig. 10 (download XLSX )
Source data and DEGs in microglia derived from DMGO compared to BrO.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bessler, N., Wezenaar, A.K.L., Ariese, H.C.R. et al. De novo H3.3K27M-altered diffuse midline glioma in human brainstem organoids to dissect GD2 CAR T cell function. Nat Cancer 7, 316–333 (2026). https://doi.org/10.1038/s43018-025-01084-0
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43018-025-01084-0