Abstract
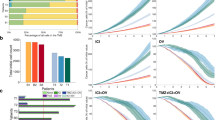
Immune checkpoint inhibition (ICI) benefits only a subset of patients with metastatic triple-negative breast cancer and determinants of response remain unclear. We assembled a longitudinal cohort of 103 female patients from the phase 2 TONIC trial, with samples spanning primary tumors, pretreatment metastases and on-treatment metastases during nivolumab therapy. We profiled 37 proteins in 270 tumors using highly multiplexed imaging and developed SpaceCat, an open-source pipeline that extracts more than 800 imaging features per sample, including cell density, diversity, spatial interactions and functional marker expression. Metastatic but not primary tumors contained features predictive of outcome. Spatial metrics such as immune diversity and T cell infiltration at tumor borders were most informative, while ratios of T cells to cancer cells and PDL1 on myeloid cells were also associated with response. Multivariate models stratified patients with the highest performance on treatment (area under the curve = 0.90). Bulk RNA-seq confirmed the predictive value of on-treatment samples. These findings highlight the value of longitudinal profiling to resolve evolving tumor microenvironment dynamics driving ICI response.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All processed imaging data from this study, including antibody staining, cell segmentation masks, and SpaceCat outputs, are publicly available online (https://www.ebi.ac.uk/biostudies/bioimages/studies/S-BIAD1288). All image analysis files are available from Zenodo (https://doi.org/10.5281/zenodo.14112852)86. The DNA-seq and RNA-seq data, as well as the response information, are available for academic use subject to the limitations of the provided informed consent. The DNA-seq and RNA-seq data from tumor biopsies of the patients with TNBC treated in the TONIC-1 trial stage 1 were deposited to the European Genome–Phenome Archive under accession number EGAS0001003535. The RNA-seq and DNA-seq data from the patients with TNBC treated in TONIC-1 trial stage 2 are not yet deposited in a public repository pending ongoing work. Both the already-deposited and the not-yet-deposited sequencing data, as well as source data supporting the findings of this study, will be made available from the corresponding author (m.kok@nki.nl) for academic use, within the limitations of the provided informed consent. Data will not be made available for commercial use. A first response to the request will be sent in <4 weeks. Data requests will be reviewed by the corresponding author and IRB of the Netherlands Cancer Institute and, after approval, applying researchers will have to sign a data transfer agreement with the Netherlands Cancer Institute. Source data are provided with this paper.
Code availability
The code to generate the figures in this paper (https://github.com/angelolab/publications/tree/main/2024-Greenwald_Nederlof_etal_TONIC), the low-level processing code (https://github.com/angelolab/toffy), the segmentation and cell assignment pipelines (https://github.com/angelolab/ark) and SpaceCat (https://github.com/angelolab/SpaceCat) are available from GitHub.
References
Larkin, J. et al. Combined nivolumab and ipilimumab or monotherapy in untreated melanoma. N. Engl. J. Med. 373, 23–34 (2015).
Hodi, F. S. et al. Improved survival with ipilimumab in patients with metastatic melanoma. N. Engl. J. Med. 363, 711–723 (2010).
Forde, P. M. et al. Neoadjuvant PD-1 blockade in resectable lung cancer. N. Engl. J. Med. 378, 1976–1986; erratum 379, 2185 (2018).
Verschoor, Y. L. et al. Neoadjuvant atezolizumab plus chemotherapy in gastric and gastroesophageal junction adenocarcinoma: the phase 2 PANDA trial. Nat. Med. 30, 519–530 (2024).
Schmid, P. et al. Event-free survival with pembrolizumab in early triple-negative breast cancer. N. Engl. J. Med. 386, 556–567 (2022).
Mittendorf, E. A. et al. Neoadjuvant atezolizumab in combination with sequential nab-paclitaxel and anthracycline-based chemotherapy versus placebo and chemotherapy in patients with early-stage triple-negative breast cancer (IMpassion031): a randomised, double-blind, phase 3 trial. Lancet 396, 1090–1100 (2020).
Emens, L. A. et al. Long-term clinical outcomes and biomarker analyses of atezolizumab therapy for patients with metastatic triple-negative breast cancer: a phase 1 study. JAMA Oncol. 5, 74–82 (2019).
Schmid, P. et al. Atezolizumab and nab-paclitaxel in advanced triple-negative breast cancer. N. Engl. J. Med. 379, 2108–2121 (2018).
Cortes, J. et al. Pembrolizumab plus chemotherapy versus placebo plus chemotherapy for previously untreated locally recurrent inoperable or metastatic triple-negative breast cancer (KEYNOTE-355): a randomised, placebo-controlled, double-blind, phase 3 clinical trial. Lancet 396, 1817–1828 (2020).
Emens, L. A. et al. Atezolizumab and nab-paclitaxel in advanced triple-negative breast cancer: biomarker evaluation of the IMpassion130 study. J. Natl Cancer Inst. 113, 1005–1016 (2021).
Loibl, S. et al. A randomised phase II study investigating durvalumab in addition to an anthracycline taxane-based neoadjuvant therapy in early triple-negative breast cancer: clinical results and biomarker analysis of GeparNuevo study. Ann. Oncol. 30, 1279–1288 (2019).
Bachelot, T. et al. Durvalumab compared to maintenance chemotherapy in metastatic breast cancer: the randomized phase II SAFIR02-BREAST IMMUNO trial. Nat. Med. 27, 250–255 (2021).
Loi, S. et al. Association between biomarkers and clinical outcomes of pembrolizumab monotherapy in patients with metastatic triple-negative breast cancer: KEYNOTE-086 exploratory analysis. JCO Precis. Oncol. 7, e2200317 (2023).
Voorwerk, L. et al. Immune induction strategies in metastatic triple-negative breast cancer to enhance the sensitivity to PD-1 blockade: the TONIC trial. Nat. Med. 25, 920–928 (2019).
Nederlof, I. et al. Neoadjuvant nivolumab or nivolumab plus ipilimumab in early-stage triple-negative breast cancer: a phase 2 adaptive trial. Nat. Med. 30, 3223–3235 (2024).
Wang, X. Q. et al. Spatial predictors of immunotherapy response in triple-negative breast cancer. Nature 621, 868–876 (2023).
Elhanani, O., Ben-Uri, R. & Keren, L. Spatial profiling technologies illuminate the tumor microenvironment. Cancer Cell 41, 404–420 (2023).
Jackson, H. W. et al. The single-cell pathology landscape of breast cancer. Nature 578, 615–620 (2020).
Keren, L. et al. A structured tumor–immune microenvironment in triple negative breast cancer revealed by multiplexed ion beam imaging. Cell 174, 1373–1387.e19 (2018).
Blomberg, O. S. et al. IL-5-producing CD4 T cells and eosinophils cooperate to enhance response to immune checkpoint blockade in breast cancer. Cancer Cell 41, 106–123.e10 (2023).
Angelo, M. et al. Multiplexed ion beam imaging of human breast tumors. Nat. Med. 20, 436–442 (2014).
Greenwald, N. F. et al. Whole-cell segmentation of tissue images with human-level performance using large-scale data annotation and deep learning. Nat. Biotechnol. 40, 555–565 (2022).
Liu, C. C. et al. Robust phenotyping of highly multiplexed tissue imaging data using pixel-level clustering. Nat. Commun. 14, 4618 (2023).
da Silva, I. P. et al. Reversal of NK-cell exhaustion in advanced melanoma by TIM-3 blockade. Cancer Immunol. Res. 2, 410–422 (2014).
Li, Y. et al. TIM-3 signaling in peripheral NK cells promotes maternal-fetal immune tolerance and alleviates pregnancy loss. Sci. Signal. 10, eaah4323 (2017).
Zhang, J. et al. Sequential actions of EOMES and T-bet promote stepwise maturation of natural killer cells. Nat. Commun. 12, 5446 (2021).
Costa, A. et al. Fibroblast heterogeneity and immunosuppressive environment in human breast cancer. Cancer Cell 33, 463–479.e10 (2018).
Wang, C. et al. Neoadjuvant camrelizumab plus nab-paclitaxel and epirubicin in early triple-negative breast cancer: a single-arm phase II trial. Nat. Commun. 14, 6654 (2023).
van der Leun, A. M., Thommen, D. S. & Schumacher, T. N. CD8+ T cell states in human cancer: insights from single-cell analysis. Nat. Rev. Cancer 20, 218–232 (2020).
Savas, P. et al. Single-cell profiling of breast cancer T cells reveals a tissue-resident memory subset associated with improved prognosis. Nat. Med. 24, 986–993 (2018).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Mariathasan, S. et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544–548 (2018).
Ali, H. R. et al. Imaging mass cytometry and multiplatform genomics define the phenogenomic landscape of breast cancer. Nat. Cancer 1, 163–175 (2020).
Tibshirani, R. Regression shrinkage and selection via the Lasso. J. R. Stat. Soc. B 58, 267–288 (1996).
McNamara, K. L. et al. Spatial proteomic characterization of HER2-positive breast tumors through neoadjuvant therapy predicts response. Nat. Cancer 2, 400–413 (2021).
Chen, L. et al. CD38-mediated immunosuppression as a mechanism of tumor cell escape from PD-1/PD-L1 blockade. Cancer Discov. 8, 1156–1175 (2018).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Liu, B. et al. Temporal single-cell tracing reveals clonal revival and expansion of precursor exhausted T cells during anti-PD-1 therapy in lung cancer. Nat. Cancer 3, 108–121 (2022).
Wu, T. D. et al. Peripheral T cell expansion predicts tumour infiltration and clinical response. Nature 579, 274–278 (2020).
Oliveira, G. & Wu, C. J. Dynamics and specificities of T cells in cancer immunotherapy. Nat. Rev. Cancer 23, 295–316 (2023).
Luoma, A. M. et al. Tissue-resident memory and circulating T cells are early responders to pre-surgical cancer immunotherapy. Cell 185, 2918–2935.e29 (2022).
Yost, K. E. et al. Clonal replacement of tumor-specific T cells following PD-1 blockade. Nat. Med. 25, 1251–1259 (2019).
Shiao, S. L. et al. Single-cell and spatial profiling identify three response trajectories to pembrolizumab and radiation therapy in triple negative breast cancer. Cancer Cell 42, 70–84.e8 (2024).
Denkert, C. et al. Tumour-infiltrating lymphocytes and prognosis in different subtypes of breast cancer: a pooled analysis of 3771 patients treated with neoadjuvant therapy. Lancet Oncol. 19, 40–50 (2018).
Geurts, V. C. M. et al. Tumor-infiltrating lymphocytes in patients with stage I triple-negative breast cancer untreated with chemotherapy. JAMA Oncol. 10, 1077–1086 (2024).
Loi, S. et al. Tumor-infiltrating lymphocytes and prognosis: a pooled individual patient analysis of early-stage triple-negative breast cancers. J. Clin. Oncol. 37, 559–569 (2019).
Bianchini, G., De Angelis, C., Licata, L. & Gianni, L. Treatment landscape of triple-negative breast cancer—expanded options, evolving needs. Nat. Rev. Clin. Oncol. 19, 91–113 (2022).
Bidard, F.-C. et al. Switch to fulvestrant and palbociclib versus no switch in advanced breast cancer with rising ESR1 mutation during aromatase inhibitor and palbociclib therapy (PADA-1): a randomised, open-label, multicentre, phase 3 trial. Lancet Oncol. 23, 1367–1377 (2022).
Bassez, A. et al. A single-cell map of intratumoral changes during anti-PD1 treatment of patients with breast cancer. Nat. Med. 27, 820–832 (2021).
Ribeiro, J. M. et al. 156TiP Impact of neoadjuvant immunotherapy in early stage breast cancer before standard therapy (BIS-Program). ESMO Open 8, 101495 (2023).
Simon, R. Optimal two-stage designs for phase II clinical trials. Control. Clin. Trials 10, 1–10 (1989).
Huang, M. et al. Association of pathologic complete response with long-term survival outcomes in triple-negative breast cancer: a meta-analysis. Cancer Res. 80, 5427–5434 (2020).
Cortazar, P. et al. Pathological complete response and long-term clinical benefit in breast cancer: the CTNeoBC pooled analysis. Lancet 384, 164–172 (2014).
Schwartz, L. H. et al. RECIST 1.1-Update and clarification: from the RECIST committee. Eur. J. Cancer 62, 132–137 (2016).
Seymour, L. et al. iRECIST: guidelines for response criteria for use in trials testing immunotherapeutics. Lancet Oncol. 18, e143–e152 (2017).
Risom, T. et al. Transition to invasive breast cancer is associated with progressive changes in the structure and composition of tumor stroma. Cell 185, 299–310.e18 (2022).
McCaffrey, E. F. et al. The immunoregulatory landscape of human tuberculosis granulomas. Nat. Immunol. 23, 318–329 (2022).
Liu, C. C. et al. Reproducible, high-dimensional imaging in archival human tissue by multiplexed ion beam imaging by time-of-flight (MIBI-TOF). Lab. Invest. 102, 762–770 (2022).
Bosse, M. et al. MIBI and IHC solutions. protocols.io https://doi.org/10.17504/protocols.io.bhmej43e (2021).
Bosse, M. IHC staining V.1. protocols.io https://doi.org/10.17504/protocols.io.bf6ajrae (2021).
Bosse, M. et al. MIBI staining V.5. protocols.io https://doi.org/10.17504/protocols.io.dm6gprk2dvzp/v5 (2022).
Bosse, M. et al. Staining sequenza. protocols.io https://doi.org/10.17504/protocols.io.bmc6k2ze (2021).
Camacho, C. et al. Antibody lyophilization. protocols.io https://doi.org/10.17504/protocols.io.bhmgj43w (2021).
Greenwald, N. et al. Greenwald, Nederlof et al extended methods V.2. protocols.io https://doi.org/10.17504/protocols.io.e6nvw44k7lmk/v2 (2025).
Medina-Martínez, J. S. et al. Isabl Platform, a digital biobank for processing multimodal patient data. BMC Bioinformatics 21, 549 (2020).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Okonechnikov, K., Conesa, A. & García-Alcalde, F. Qualimap 2: advanced multi-sample quality control for high-throughput sequencing data. Bioinformatics 32, 292–294 (2016).
Benjamin, D. et al. Calling somatic SNVs and indels with Mutect2. Preprint at bioRxiv https://doi.org/10.1101/861054 (2019).
Kim, S. et al. Strelka2: fast and accurate calling of germline and somatic variants. Nat. Methods 15, 591–594 (2018).
Rosenthal, R., McGranahan, N., Herrero, J., Taylor, B. S. & Swanton, C. DeconstructSigs: delineating mutational processes in single tumors distinguishes DNA repair deficiencies and patterns of carcinoma evolution. Genome Biol. 17, 31 (2016).
Shen, R. & Seshan, V. E. FACETS: allele-specific copy number and clonal heterogeneity analysis tool for high-throughput DNA sequencing. Nucleic Acids Res. 44, e131 (2016).
Ali, H. R. et al. Genome-driven integrated classification of breast cancer validated in over 7,500 samples. Genome Biol. 15, 431 (2014).
Szolek, A. et al. OptiType: precision HLA typing from next-generation sequencing data. Bioinformatics 30, 3310–3316 (2014).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Richman, L. P., Vonderheide, R. H. & Rech, A. J. Neoantigen dissimilarity to the self-proteome predicts immunogenicity and response to immune checkpoint blockade. Cell Syst. 9, 375–382 (2019).
Ayers, M. et al. IFN-γ-related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Invest. 127, 2930–2940 (2017).
Bagaev, A. et al. Conserved pan-cancer microenvironment subtypes predict response to immunotherapy. Cancer Cell 39, 845–865.e7 (2021).
Shah, S. P. et al. The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature 486, 395–399 (2012).
Lehmann, B. D. et al. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J. Clin. Invest. 121, 2750–2767 (2011).
van den Eynden, J., Jiménez-Sánchez, A., Miller, M. L. & Larsson, E. Lack of detectable neoantigen depletion signals in the untreated cancer genome. Nat. Genet. 51, 1741–1748 (2019).
Fang, Z. & Peltz, G. An automated multi-modal graph-based pipeline for mouse genetic discovery. Bioinformatics 38, 3385–3394 (2022).
Tietscher, S. et al. A comprehensive single-cell map of T cell exhaustion-associated immune environments in human breast cancer. Nat. Commun. 14, 98 (2023).
Doering, T. A. et al. Network analysis reveals centrally connected genes and pathways involved in CD8+ T cell exhaustion versus memory. Immunity 37, 1130–1144 (2012).
Xu, L. et al. A comprehensive single-cell breast tumor atlas defines epithelial and immune heterogeneity and interactions predicting anti-PD-1 therapy response. Cell Rep. Med. 5, 101511 (2024).
Newman, A. M. et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 37, 773–782 (2019).
Greenwald, N. et al. Temporal and spatial composition of the tumor microenvironment predicts response to immune checkpoint inhibition. Zenodo https://doi.org/10.5281/zenodo.14112852 (2025).
Acknowledgements
We thank the patients and their families for taking part in the TONIC trials. TONIC trial costs were supported by Bristol Myers Squibb. We thank all supporting clinical trial staff, particularly nurse specialists and the Departments of Medical Oncology, Surgery, Radiology and Pathology of the participating centers. We thank the Netherlands Cancer Institute Core Facility of Molecular Pathology and Biobanking for their support in processing of samples. Support with title page creation and format was provided by AuthorArranger, a tool developed at the National Cancer Institute (NCI). Figures and graphics were created with BioRender.com. We thank T. Murty for constant support. N.F.G. was supported by NCI CA246880, NCI CA264307 and the Stanford Graduate Fellowship. L.M. was supported by the Stanford School of Medicine Dean’s Fellowship. Collection and processing of samples was made possible by the Dutch Cancer Foundation (grant 2016-10510). Research in the laboratory of M.K. is funded by the Netherlands Organization for Scientific Research and Victoria’s Secret Global Fund for Women’s Cancers Rising Innovator Research Grant, in partnership with Pelotonia and the American Association for Cancer Research. M.A. was supported by National Institutes of Health grants 5U54CA20997105, 5DP5OD01982205, 1R01CA24063801A1, 5R01AG06827902, 5UH3CA24663303, 5R01CA22952904, 1U24CA22430901, 5R01AG05791504, 5R01AG05628705, 5U24CA22430903, 3U54HL165445-03S1, 5R01AG05628705 and 5R01AG05791505, the Department of Defense (W81XWH2110143) and other funding from the Wellcome Trust, the Bill and Melinda Gates Foundation, the Cancer Research Institute, the Parker Institute for Cancer Immunotherapy and the Breast Cancer Research Foundation. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
N.F.G., I.N., C.C., M.A. and M.K. conceptualized the project. I.N. collected the longitudinal samples and clinical data. M.d.G. collaborated on the clinical data collection. L.V. and V.G. coordinated the TONIC trial procedures. M.d.M. processed the FFPE tissues for IHC, isolated the DNA and RNA from tissue biopsies and constructed the TMAs. M.J.v.d.V. and H.M.H. annotated the FFPE tissue for TMA construction and provided a pathology review of each tissue slide. M.J.v.d.V. funded the longitudinal sampling and logistics of the project. T.N.S. and S.C.B. provided guidance. N.F.G., C.C.F., Z.K., Y.B., E.M., H.P., T.R., A.D., F.J.H. and M.B. generated the MIBI data. N.F.G., C.S., D.Y.D., A. Kong, I.N., S.R.V., E.M., C.C.L., J.S.R., A. Kagel, B.G.S. and R.T. processed and analyzed the MIBI data. S.P., K.E.H., A. Khan, N.F.G., D.Y.D., C.C.L., C.Y.Y., L.M., C.S.-V. and Z.M. processed and analyzed the DNA-seq and RNA-seq data. N.F.G., I.N., C.S., D.Y.D., S.P., A. Kong and S.R.V. generated the figures. N.F.G., I.N., C.C., M.K. and M.A. wrote the paper. C.C., M.K. and M.A. supervised the project. All authors provided feedback on the paper.
Corresponding authors
Ethics declarations
Competing interests
N.F.G. is an advisor for Cellformatica. T.N.S. is advisor for Allogene Therapeutics, Asher Bio, Merus, Neogene Therapeutics and Scenic Biotech, is a stockholder in Allogene Therapeutics, Asher Bio, Cell Control, Celsius, Merus and Scenic Biotech and is venture partner at Third Rock Ventures, all outside of the current work. M.K. reports research funding to the institute from BMS, Roche and AstraZeneca/MedImmune and an advisory role/speakers fee (all compensated to the institute) for Alderaan, BMS, Domain Therapeutics, Medscape, Roche, MSD and Daiichi Sankyo, outside the submitted work. M.A. is a named inventor on patent US20150287578A1, which covers the mass spectrometry approach used by MIBI to detect elemental reporters in tissue using secondary-ion mass spectrometry. M.A. is a board member and shareholder in IonPath, which develops and manufactures the commercial MIBI platform. The other authors declare no competing interests.
Peer review
Peer review information
Nature Cancer thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 ROI selection and sample summary.
a Workflow for selecting cores to be included in the study. Hematoxylin and eosin (H&E) slides were digitized in slidescore.com, annotated by a dedicated breast cancer pathologist, and demarcated with areas of interest. Each identified region was punched with a 1.5 mm core and placed onto a tissue microarray (TMA). b Illustrative examples of the H&E image of the entire core, the cropped inset of the portion of the core selected for MIBI analysis, and the corresponding MIBI image generated from the selected region. c Histogram showing the number of fields-of-view (x-axis) acquired on MIBI across all timepoints from all patients (y-axis) d Upset plot showing the overlap of distinct timepoints of MIBI data across the patients in the cohort. e Venn diagram showing the overlap between DNA, RNA, and MIBI data for baseline samples (N = 93 patients) f Venn diagram showing the overlap between RNA and MIBI data for pre-nivo samples (N = 77 patients) g Same as f) for on-nivo (N = 66 patients).
Extended Data Fig. 2 Cell clustering scheme and cell type prevalence.
a Diagram illustrating the three levels of cell clustering (broad, intermediate, detailed), and the relationship between clusters in each level b Number of cells of each cell type in the broad clusters c Number of cells of each cell type in the intermediate clusters d Number of cells of each cell type in the detailed clusterings. b, c, d are all based on all primary and metastatic tumors in the sample set. e Mean count of cell types (broad clusters) for each timepoint f Mean count of cell types (intermediate clusters) for each timepoint g Average proportion of each intermediate cancer subpopulation in metastatic samples (N = 101 patients) h Average proportion of each intermediate structural subpopulation in metastatic samples (N = 101 patients) i Average proportion of each intermediate immune subpopulation in metastatic samples (N = 101 patients) j PD1 expression in Wang et al. Box plot: Lower bound is 1st quartile, center is median, upper bound is 3rd quartile, whiskers extend to 1.5*IQR beyond bound.
Extended Data Fig. 3 Feature extraction pipeline schematics.
a Cartoon illustrating how compartment masks are defined b Table illustrating how functional marker thresholds are used to binarize cells into positive or negative for each marker; these assignments are then used to generate positivity proportion statistics per image c Schematic showcasing how diversity scores are calculated d Cartoon showing how the radius around a cell is used for neighborhood diversity e Cartoon showing how cell distances are used to compute the linear distance feature f Cartoon showing how the mixing score is calculated, along with examples of high and low mixing g Schematic showing how the fiber segmentation pipeline is used to generate features h Schematic showing how the extracellular matrix pipeline is used to generate features.
Extended Data Fig. 4 Quantification of features across tumor compartments.
a Heatmap showing features that are consistent across compartments. Each row represents a different feature, and each column is a distinct compartment. Features are normalized to the whole image value b Same as above, but for features that are enriched in specific compartment(s) c Histogram showing threshold used to identify varying vs. non-varying compartment features d Distribution of the CD8 T / CD4 T ratio, a feature that changed across compartments (n = 895 features) e Distribution of Smooth Muscle density, a feature that changed across compartments (n = 1084 features) f Distribution of Ki67 positivity in Cancer 1, a feature that did not change across compartments g Distribution of PD-L1 positivity in CD68 macrophages, a feature that did not change across compartments (n = 956 features) h Heatmap showing expression of markers across distinct tile-based clusters (n = 507 features) i Proportion of TIM3+ Structural cells across ECM neighborhoods (n = 227 FOVs) j Proportion of TIM3 + CD68 Macrophages cells across ECM neighborhoods (n = 200 FOVs) k Proportion of PDL1 + CD68 Macrophages cells across ECM neighborhoods (n = 200 FOVs) Box plot: Lower bound is 1st quartile, center is median, upper bound is 3rd quartile, whiskers extend to 1.5*IQR beyond bound.
Extended Data Fig. 5 Evolution of the TME through treatment.
a Summary of features that changed between baseline and on-nivo samples across all patients b Ratio of Cancer cells to T cells in stroma border at baseline and on-nivo timepoint across all patients c Proportion of all CD45RO+ cells at baseline and on-nivo timepoint across all patients d Diversity of cells surrounding Mac_Other cells at baseline and on-nivo timepoint across all patients e Summary of features that changed between baseline and on-nivo samples across paired samples(3.5) f Ratio of Cancer cells to T cells in stroma border at baseline and on-nivo timepoint across paired samples g Ratio of Cancer cells to unclassified cells in stroma border at baseline and on-nivo timepoint across paired samples h Ratio of NK cells to T cells at baseline and on-nivo timepoint across paired samples For panels b-d and f-h, N = 101 patients.
Extended Data Fig. 6 Relationship between MIBI- and RNA-based features.
a Distribution of ssGSEA normalized enrichment scores (NES) across MSigDB signatures related to cytokine signaling (N = 210 samples) b Volcano plot showing correlation coefficient (x-axis) and p-value from an unpaired two-sided t-test (y-axis) for pairs RNA- and MIBI-based features c Correlation between HLA1+ cells (MIBI) and MHCI (RNA). Scatter points represent individual data values, the line indicates the linear regression fit, and the shaded area shows the 95% confidence interval of the fit. d Correlation between CD45RO + CD4 + T cells at the border (MIBI) with a TGFb gene score (RNA) e Correlation between HLA1+ Cancer cells (MIBI) and interferon signaling (RNA) f Correlation between diversity surrounding CD8 T cells in the cancer core (MIBI) with a signature for T cell exhaustion (RNA) g Enrichment of MIBI-RNA feature pairs in top correlated features based on the compartment the MIBI feature is defined in Box plot: Lower bound is 1st quartile, center is median, upper bound is 3rd quartile, whiskers extend to 1.5*IQR beyond bound.
Extended Data Fig. 7 Outcomes associations.
a Enumeration of the number of top MIBI features which are ratios compared to the number which are densities b Volcano plot of RNA-seq features associated with response c Top RNA features associated with response, organized by biological category d Comparison of number of DNA and RNA features associated with response e Number of RNA-seq features associated with response by timepoint f The number of times a given MIBI feature was repeated across distinct timepoints among the top 100 features g Same as Fig. 4b, but with all labels included h Types of features associated with response from baseline metastatic timepoint i Top 50 features associated with response from baseline timepoint, colored by compartment and sorted by feature type j Cancer/Immune mixing score stratified by response status across timepoints: primary (N = 51, 7 Yes / 44 No), baseline (N = 50, 10 Yes / 40 No), pre nivo (N = 39, 7 Yes / 32 No), on nivo (N = 37, 7 Yes / 30 No). N=number of patients. k Proportion of Ki67+ Cancer 1 cells stratified by response status across timepoints l Proportion of GLUT1+ Cancer 1 cells stratified by response status across timepoints Box plot: Lower bound is 1st quartile, center is median, upper bound is 3rd quartile, whiskers extend to 1.5*IQR beyond bound. For j and k: primary (N = 59, 9 Yes / 50 No), baseline (N = 74, 12 Yes / 62 No), pre nivo (N = 67, 11 Yes / 56 No), on nivo (N = 57, 8 Yes / 49 No). N=number of patients.
Extended Data Fig. 8 Timepoint-specific features.
a Box plots showing the distribution of the B / NK ratio in the cancer core feature in responders and non-responders, across the four distinct timepoints in the cohort, primary (N = 21, 4 Yes / 17 No), baseline (N = 15, 5 Yes / 10 No), pre nivo (N = 13, 2 Yes / 11 No), on nivo (N = 15, 4 Yes / 11 No). b Same as a), with CD68 Macrophage density as the feature, primary (N = 59, 9 Yes / 50 No), baseline (N = 79, 12 Yes / 67 No), pre nivo (N = 70, 13 Yes / 57 No), on nivo (N = 62, 12 Yes / 50 No) c Same as a), with Vim+ in CD4 T cells as the feature, primary (N = 53, 7 Yes / 46 No), baseline (N = 58, 11 Yes / 47 No), pre nivo (N = 52, 9 Yes / 43 No), on nivo (N = 52, 12 Yes / 40 No) d Same as a), with Structural cell density in the stroma border as the feature, primary (N = 59, 9 Yes / 50 No), baseline (N = 76, 12 Yes / 64 No), pre nivo (N = 66, 12 Yes / 54 No), on nivo (N = 58, 8 Yes / 50 No) e Same as a), with Other / Cancer cell ratio in the stroma core as the feature, primary (N = 59, 9 Yes / 50 No), baseline (N = 78, 12 Yes / 66 No), pre nivo (N = 69, 13 Yes / 56 No), on nivo (N = 62, 12 Yes / 50 No) f Same as a), with Nk / Other cell ratio in the stroma border as the feature, primary (N = 56, 7 Yes / 49 No), baseline (N = 72, 12 Yes / 60 No), pre nivo (N = 65, 12 Yes / 53 No), on nivo (N = 53, 7 Yes / 46 No) Box plot: Lower bound is 1st quartile, center is median, upper bound is 3rd quartile, whiskers extend to 1.5*IQR beyond bound. N=number of patients for all.
Extended Data Fig. 9 Multivariate modeling approach and benchmarking.
a Diagram illustrating the cross validation approach used for model training b Histogram showing the number of times the top features were selected across the 10 different iterations of model training c Comparison of the importance score of the top features identified by the model across distinct timepoints, stratified by MIBI vs. RNA d Overlap across timepoints of top features identified by the MIBI models e Overlap across timepoints of top features identified by the RNA models f AUROC evaluated for the on-nivo MIBI data using separate train, val, test split without cross validation. g Same as above, for AUPRC h Accuracy of multivariate model with decreasing number of retained features i Relative accuracy of combined MIBI + RNA model compared to best performing unimodal model Box plot: Lower bound is 1st quartile, center is median, upper bound is 3rd quartile, whiskers extend to 1.5*IQR beyond bound.
Extended Data Fig. 10 Comparison with previously identified predictive features.
a Overlap in pre-treatment predictive features originally identified in Wang et al. b Overlap in pre-treatment predictive features after running SpaceCat on the data from Wang et al. c Proportion of Ki67+ Cancer cells stratified by response across timepoints in Wang et al., baseline (N = 109, 55 pCR / 54 RD), on treatment (N = 67, 28 pCR / 39 RD) d Proportion of Ki67 + CD8 T cells stratified by response across timepoints in Wang et al., baseline (N = 113, 56 pCR / 57 RD), on treatment (N = 97, 48 pCR / 49 RD) e Density of Cancer cells stratified by response across timepoints in Wang et al., baseline (N = 108, 54 pCR / 54 RD), on treatment (N = 93, 47 pCR / 46 RD) f Comparison of SpaceCat features, original features, and combined features to predict outcome in Wang et al. data g Comparison of SpaceCat features, Wang et al. features, and combined features to predict outcome in TONIC data h Overlap in on-treatment predictive features originally identified in Wang et al. (4.2) i Overlap in on-treatment predictive features after running SpaceCat on the data from Wang et al. j Distribution of top features across timepoints when using only features defined in Wang et al. k Change in top 100 predictive features when including cell subsets defined in Wang et al. l Change in multivariate model specific cell clusters that are predictive, as well as multivariate results Box plot: Lower bound is 1st quartile, center is median, upper bound is 3rd quartile, whiskers extend to 1.5*IQR beyond bound. N=number of patients.
Supplementary information
Supplementary Table (download XLSX )
Supplementary Tables 1–11.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Greenwald, N.F., Nederlof, I., Sowers, C. et al. Temporal and spatial composition of the tumor microenvironment predicts response to immune checkpoint inhibition in metastatic TNBC. Nat Cancer 7, 435–450 (2026). https://doi.org/10.1038/s43018-026-01114-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43018-026-01114-5