Abstract
Somatic stem cell pools comprise diverse, highly specialized subsets whose individual contribution is critical for the overall regenerative function. In the bone marrow, myeloid-biased hematopoietic stem cells (myHSCs) are indispensable for replenishment of myeloid cells and platelets during inflammatory response but, at the same time, become irreversibly damaged during inflammation and aging. Here we identify an extrinsic factor, Semaphorin 4A (Sema4A), which non-cell-autonomously confers myHSC resilience to inflammatory stress. We show that, in the absence of Sema4A, myHSC inflammatory hyper-responsiveness in young mice drives excessive myHSC expansion, myeloid bias and profound loss of regenerative function with age. Mechanistically, Sema4A is mainly produced by neutrophils, signals via a cell surface receptor, Plexin D1, and safeguards the myHSC epigenetic state. Our study shows that, by selectively protecting a distinct stem cell subset, an extrinsic factor preserves functional diversity of somatic stem cell pool throughout organismal lifespan.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data supporting the findings of the study are available from the corresponding author. All data were analyzed with standard programs and packages, as detailed in the Methods. RNA-seq data from this study are available from ArrayExpress (E-MTAB-11359 (single-cell RNA-seq of aged WT/Sema4AKO myHSCs and lyHSCs) and E-MTAB-12890 (bulk RNA-seq of WT/Sema4AKO myHSCs/lyHSCs at baseline and after acute LPS)). ATAC-seq data from this study are available from the Gene Expression Omnibus (GSE281145).
Change history
20 February 2025
A Correction to this paper has been published: https://doi.org/10.1038/s43587-025-00837-x
References
Scaramozza, A. et al. Lineage tracing reveals a subset of reserve muscle stem cells capable of clonal expansion under stress. Cell Stem Cell 24, 944–957 (2019).
Ibrayeva, A. et al. Early stem cell aging in the mature brain. Cell Stem Cell 28, 955–966 (2021).
Altshuler, A. et al. Discrete limbal epithelial stem cell populations mediate corneal homeostasis and wound healing. Cell Stem Cell 28, 1248–1261 (2021).
Farrelly, O. et al. Two-photon live imaging of single corneal stem cells reveals compartmentalized organization of the limbal niche. Cell Stem Cell 28, 1233–1247 (2021).
Hsu, Y. C., Pasolli, H. A. & Fuchs, E. Dynamics between stem cells, niche, and progeny in the hair follicle. Cell 144, 92–105 (2011).
Matatall, K. A., Shen, C. C., Challen, G. A. & King, K. Y. Type II interferon promotes differentiation of myeloid-biased hematopoietic stem cells. Stem Cells 32, 3023–3030 (2014).
Yamamoto, R., Wilkinson, A. C. & Nakauchi, H. Changing concepts in hematopoietic stem cells. Science 362, 895–896 (2018).
Yamamoto, R. et al. Clonal analysis unveils self-renewing lineage-restricted progenitors generated directly from hematopoietic stem cells. Cell 154, 1112–1126 (2013).
Florez, M. A. et al. Interferon gamma mediates hematopoietic stem cell activation and niche relocalization through BST2. Cell Rep. 33, 108530 (2020).
Beerman, I. et al. Functionally distinct hematopoietic stem cells modulate hematopoietic lineage potential during aging by a mechanism of clonal expansion. Proc. Natl Acad. Sci. USA 107, 5465–5470 (2010).
Challen, G. A., Boles, N. C., Chambers, S. M. & Goodell, M. A. Distinct hematopoietic stem cell subtypes are differentially regulated by TGF-β1. Cell Stem Cell 6, 265–278 (2010).
Sanjuan-Pla, A. et al. Platelet-biased stem cells reside at the apex of the haematopoietic stem-cell hierarchy. Nature 502, 232–236 (2013).
Carrelha, J. et al. Hierarchically related lineage-restricted fates of multipotent haematopoietic stem cells. Nature 554, 106–111 (2018).
Notta, F. et al. Distinct routes of lineage development reshape the human blood hierarchy across ontogeny. Science 351, aab2116 (2016).
Pang, W. W. et al. Human bone marrow hematopoietic stem cells are increased in frequency and myeloid-biased with age. Proc. Natl Acad. Sci. USA 108, 20012–20017 (2011).
Mitroulis, I. et al. Modulation of myelopoiesis progenitors is an integral component of trained immunity. Cell 172, 147–161 (2018).
Bogeska, R. et al. Inflammatory exposure drives long-lived impairment of hematopoietic stem cell self-renewal activity and accelerated aging. Cell Stem Cell 29, 1273–1284 (2022).
Goodell, M. A. & Rando, T. A. Stem cells and healthy aging. Science 350, 1199–1204 (2015).
Mann, M. et al. Heterogeneous responses of hematopoietic stem cells to inflammatory stimuli are altered with age. Cell Rep. 25, 2992–3005 (2018).
Alto, L. T. & Terman, J. R. Semaphorins and their signaling mechanisms. Methods Mol. Biol. 1493, 1–25 (2017).
Kumanogoh, A. et al. Class IV semaphorin Sema4A enhances T-cell activation and interacts with Tim-2. Nature 419, 629–633 (2002).
Kumanogoh, A. et al. Nonredundant roles of Sema4A in the immune system: defective T cell priming and Th1/Th2 regulation in Sema4A-deficient mice. Immunity 22, 305–316 (2005).
Toyofuku, T. et al. Semaphorin-4A, an activator for T-cell-mediated immunity, suppresses angiogenesis via Plexin-D1. EMBO J. 26, 1373–1384 (2007).
Silberstein, L. et al. Proximity-based differential single-cell analysis of the niche to identify stem/progenitor cell regulators. Cell Stem Cell 19, 530–543 (2016).
Goncalves, K. A. et al. Angiogenin promotes hematopoietic regeneration by dichotomously regulating quiescence of stem and progenitor cells. Cell 166, 894–906 (2016).
Takamatsu, H. & Kumanogoh, A. Diverse roles for semaphorin–plexin signaling in the immune system. Trends Immunol. 33, 127–135 (2012).
Delgoffe, G. M. et al. Stability and function of regulatory T cells is maintained by a neuropilin-1–semaphorin-4a axis. Nature 501, 252–256 (2013).
Haematolymphoid Tumours. WHO Classification of Tumours 5th edn (World Health Organization, 2024).
Ho, Y. H. et al. Remodeling of bone marrow hematopoietic stem cell niches promotes myeloid cell expansion during premature or physiological aging. Cell Stem Cell 25, 407–418 (2019).
Pronk, C. J. et al. Elucidation of the phenotypic, functional, and molecular topography of a myeloerythroid progenitor cell hierarchy. Cell Stem Cell 1, 428–442 (2007).
Liang, Y., Van Zant, G. & Szilvassy, S. J. Effects of aging on the homing and engraftment of murine hematopoietic stem and progenitor cells. Blood 106, 1479–1487 (2005).
Karakaslar, E. O. et al. Transcriptional activation of Jun and Fos members of the AP-1 complex is a conserved signature of immune aging that contributes to inflammaging. Aging Cell 22, e13792 (2023).
Trizzino, M. et al. EGR1 is a gatekeeper of inflammatory enhancers in human macrophages. Sci. Adv. 7, eaaz8836 (2021).
Jia, D. et al. Cardiac resident macrophage-derived legumain improves cardiac repair by promoting clearance and degradation of apoptotic cardiomyocytes after myocardial infarction. Circulation 145, 1542–1556 (2022).
Marzio, R., Jirillo, E., Ransijn, A., Mauel, J. & Corradin, S. B. Expression and function of the early activation antigen CD69 in murine macrophages. J. Leukoc. Biol. 62, 349–355 (1997).
Konturek-Ciesla, A., Olofzon, R., Kharazi, S. & Bryder, D. Implications of stress-induced gene expression for hematopoietic stem cell aging studies. Nat. Aging 4, 177–184 (2024).
Yoshida, H. et al. The cis-regulatory atlas of the mouse immune system. Cell 176, 897–912 (2019).
King, K. Y. et al. Irgm1 protects hematopoietic stem cells by negative regulation of IFN signaling. Blood 118, 1525–1533 (2011).
Weih, F. et al. Multiorgan inflammation and hematopoietic abnormalities in mice with a targeted disruption of RelB, a member of the NF-κ B/Rel family. Cell 80, 331–340 (1995).
Li, J. et al. STAT1 is essential for HSC function and maintains MHCIIhi stem cells that resist myeloablation and neoplastic expansion. Blood 140, 1592–1606 (2022).
Becker-Herman, S. et al. CD74 is a regulator of hematopoietic stem cell maintenance. PLoS Biol. 19, e3001121 (2021).
Haghverdi, L., Buttner, M., Wolf, F. A., Buettner, F. & Theis, F. J. Diffusion pseudotime robustly reconstructs lineage branching. Nat. Methods 13, 845–848 (2016).
Kovtonyuk, L. V., Fritsch, K., Feng, X., Manz, M. G. & Takizawa, H. Inflamm-aging of hematopoiesis, hematopoietic stem cells, and the bone marrow microenvironment. Front. Immunol. 7, 502 (2016).
Flohr Svendsen, A. et al. A comprehensive transcriptome signature of murine hematopoietic stem cell aging. Blood 138, 439–451 (2021).
Wilson, N. K. et al. Combined single-cell functional and gene expression analysis resolves heterogeneity within stem cell populations. Cell Stem Cell 16, 712–724 (2015).
Cabezas-Wallscheid, N. et al. Identification of regulatory networks in HSCs and their immediate progeny via integrated proteome, transcriptome, and DNA methylome analysis. Cell Stem Cell 15, 507–522 (2014).
Laurenti, E. et al. CDK6 levels regulate quiescence exit in human hematopoietic stem cells. Cell Stem Cell 16, 302–313 (2015).
Fernando, R. N. et al. Cell cycle restriction by histone H2AX limits proliferation of adult neural stem cells. Proc. Natl Acad. Sci. USA 108, 5837–5842 (2011).
Qian, H. et al. Critical role of thrombopoietin in maintaining adult quiescent hematopoietic stem cells. Cell Stem Cell 1, 671–684 (2007).
Pietras, E. M. et al. Functionally distinct subsets of lineage-biased multipotent progenitors control blood production in normal and regenerative conditions. Cell Stem Cell 17, 35–46 (2015).
Adolfsson, J. et al. Identification of Flt3+ lympho-myeloid stem cells lacking erythro-megakaryocytic potential a revised road map for adult blood lineage commitment. Cell 121, 295–306 (2005).
Chen, X. et al. Bone marrow myeloid cells regulate myeloid-biased hematopoietic stem cells via a histamine-dependent feedback loop. Cell Stem Cell 21, 747–760 (2017).
Esplin, B. L. et al. Chronic exposure to a TLR ligand injures hematopoietic stem cells. J. Immunol. 186, 5367–5375 (2011).
Demel, U. M. et al. A complex proinflammatory cascade mediates the activation of HSCs upon LPS exposure in vivo. Blood Adv. 6, 3513–3528 (2022).
Walter, D. et al. Exit from dormancy provokes DNA-damage-induced attrition in haematopoietic stem cells. Nature 520, 549–552 (2015).
Komorowska, K. et al. Hepatic leukemia factor maintains quiescence of hematopoietic stem cells and protects the stem cell pool during regeneration. Cell Rep. 21, 3514–3523 (2017).
Warr, M. R. et al. FOXO3A directs a protective autophagy program in haematopoietic stem cells. Nature 494, 323–327 (2013).
Passegué, E., Wagers, A. J., Giuriato, S., Anderson, W. C. & Weissman, I. L. Global analysis of proliferation and cell cycle gene expression in the regulation of hematopoietic stem and progenitor cell fates. J. Exp. Med. 202, 1599–1611 (2005).
Cabezas-Wallscheid, N. et al. Vitamin A-retinoic acid signaling regulates hematopoietic stem cell dormancy. Cell 169, 807–823 (2017).
Yu, W. et al. Plexin-B2 mediates physiologic and pathologic functions of angiogenin. Cell 171, 849–864 (2017).
Kwok, A. J. et al. Neutrophils and emergency granulopoiesis drive immune suppression and an extreme response endotype during sepsis. Nat. Immunol. 24, 767–779 (2023).
Zhang, Y. et al. Tie2Cre-mediated inactivation of plexinD1 results in congenital heart, vascular and skeletal defects. Dev. Biol. 325, 82–93 (2009).
Ding, J. B., Oh, W. J., Sabatini, B. L. & Gu, C. Semaphorin 3E–Plexin-D1 signaling controls pathway-specific synapse formation in the striatum. Nat. Neurosci. 15, 215–223 (2011).
Kim, M. H. et al. A late-lineage murine neutrophil precursor population exhibits dynamic changes during demand-adapted granulopoiesis. Sci. Rep. 7, 39804 (2017).
Passegue, E., Wagner, E. F. & Weissman, I. L. JunB deficiency leads to a myeloproliferative disorder arising from hematopoietic stem cells. Cell 119, 431–443 (2004).
Abram, C. L., Roberge, G. L., Hu, Y. & Lowell, C. A. Comparative analysis of the efficiency and specificity of myeloid-Cre deleting strains using ROSA-EYFP reporter mice. J. Immunol. Methods 408, 89–100 (2014).
Dykstra, B. et al. Long-term propagation of distinct hematopoietic differentiation programs in vivo. Cell Stem Cell 1, 218–229 (2007).
Meng, Y. et al. Epigenetic programming defines haematopoietic stem cell fate restriction. Nat. Cell Biol. 25, 812–822 (2023).
Lee, J. W. et al. RUNX3 regulates cell cycle-dependent chromatin dynamics by functioning as a pioneer factor of the restriction-point. Nat. Commun. 10, 1897 (2019).
Pinho, S. et al. Lineage-biased hematopoietic stem cells are regulated by distinct niches. Dev. Cell 44, 634–641 (2018).
Morita, Y., Ema, H. & Nakauchi, H. Heterogeneity and hierarchy within the most primitive hematopoietic stem cell compartment. J. Exp. Med. 207, 1173–1182 (2010).
Luchsinger, L. L., de Almeida, M. J., Corrigan, D. J., Mumau, M. & Snoeck, H. W. Mitofusin 2 maintains haematopoietic stem cells with extensive lymphoid potential. Nature 529, 528–531 (2016).
Song, Z. et al. An NAD+-dependent metabolic checkpoint regulates hematopoietic stem cell activation and aging. Nat. Aging 4, 1384–1393 (2024).
Kwak, H. J. et al. Myeloid cell-derived reactive oxygen species externally regulate the proliferation of myeloid progenitors in emergency granulopoiesis. Immunity 42, 159–171 (2015).
Zhu, H. et al. Reactive oxygen species-producing myeloid cells act as a bone marrow niche for sterile inflammation-induced reactive granulopoiesis. J. Immunol. 198, 2854–2864 (2017).
Wang, L. et al. Expression of Semaphorin 4A and its potential role in rheumatoid arthritis. Arthritis Res. Ther. 17, 227 (2015).
Young, K. et al. Decline in IGF1 in the bone marrow microenvironment initiates hematopoietic stem cell aging. Cell Stem Cell 28, 1473–1482 (2021).
Arthur, L. et al. Cellular and plasma proteomic determinants of COVID-19 and non-COVID-19 pulmonary diseases relative to healthy aging. Nat. Aging 1, 535–549 (2021).
Choi, Y. I. et al. PlexinD1 glycoprotein controls migration of positively selected thymocytes into the medulla. Immunity 29, 888–898 (2008).
Dai, X. et al. A novel role for myeloid cell-specific neuropilin 1 in mitigating sepsis. FASEB J. 31, 2881–2892 (2017).
Zeng, A. G. X. et al. Identification of a human hematopoietic stem cell subset that retains memory of inflammatory stress. Preprint at bioRxiv https://doi.org/10.1101/2023.09.11.557271 (2023).
Xia, J. et al. Semaphorin-plexin signaling controls mitotic spindle orientation during epithelial morphogenesis and repair. Dev. Cell 33, 299–313 (2015).
Galvin, A. et al. Cell cycle analysis of hematopoietic stem and progenitor cells by multicolor flow cytometry. Curr. Protoc. Cytom. 87, e50 (2019).
Feng, C. G., Weksberg, D. C., Taylor, G. A., Sher, A. & Goodell, M. A. The p47 GTPase Lrg-47 (Irgm1) links host defense and hematopoietic stem cell proliferation. Cell Stem Cell 2, 83–89 (2008).
Christodoulou, C. et al. Live-animal imaging of native haematopoietic stem and progenitor cells. Nature 578, 278–283 (2020).
Ema, H. et al. Adult mouse hematopoietic stem cells: purification and single-cell assays. Nat. Protoc. 1, 2979–2987 (2006).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Cunningham, F. et al. Ensembl 2022. Nucleic Acids Res. 50, D988–D995 (2022).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Reimand, J. et al. Pathway enrichment analysis and visualization of omics data using g:Profiler, GSEA, Cytoscape and EnrichmentMap. Nat. Protoc. 14, 482–517 (2019).
Fang, Z., Liu, X. & Peltz, G. GSEApy: a comprehensive package for performing gene set enrichment analysis in Python. Bioinformatics 39, btac757 (2023).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291 (2019).
McInnes, L., Healy, J. & Melville, J. UMAP: uniform manifold approximation and projection for dimension reduction. Preprint at arXiv https://doi.org/10.48550/arXiv.1802.03426 (2020).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Lun, A. T., Bach, K. & Marioni, J. C. Pooling across cells to normalize single-cell RNA sequencing data with many zero counts. Genome Biol. 17, 75 (2016).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. https://doi.org/10.1038/nbt.4314 (2018).
Nestorowa, S. et al. A single-cell resolution map of mouse hematopoietic stem and progenitor cell differentiation. Blood 128, e20–e31 (2016).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. https://doi.org/10.14806/ej.17.1.200 (2011).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Acknowledgements
The authors would like to acknowledge the Core Facilities of Fred Hutchinson Cancer Center; E. Regiarto, M. Baum and K. Krum for technical help; and A. Wilkinson and S. Mckinney-Freeman for helpful discussions and critical reading of the manuscript. This study was supported by National Institutes of Health grant RO1 HL148189, the New Development Fund from the Fred Hutchinson Cancer Center and Leukemia and Lymphoma Society Translational Research Program award to L.S. E.M.P. is a Scholar of the Leukemia and Lymphoma Society.
Author information
Authors and Affiliations
Contributions
L.S. conceived the study. D.T., S.G., S.Z., S. Radtke, H.-P.K., S. Rodriguez, N.C., A.P., A.G., E.M.P. and D.T.S. characterized mutant mouse strains. N.S., C.B., D.G. and J.A.S. performed intravital microscopy experiments. E.M., E.I.C., N.K.W., S.J.K., B.G., P.K., D.J., A.T. and A.S. generated and analyzed bulk and single-cell RNA-seq data. L.Z., I.B., B.P. and D.H.J. generated and analyzed ATAC-seq data. A.K. provided the Sema4AKO mice; T.W. provided the Sema4A-floxed mice; and C.N. provided the vWF-Tomato mice. M.M., C.M. and F.L. performed the bone histology analysis. J.-G.C. and S.Z.J. provided human HSC gene expression data. L.S. wrote the manuscript, with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
L.S., S. Radtke and H.P.-K. are listed inventors on patent application 18/717,971 relating to this work. All other authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks Yu Hou and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The absence of Sema4A leads to excessive myeloid expansion and premature hematopoietic aging-like phenotype.
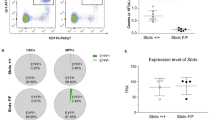
A. Representative images of H&E staining of femurs from aged WT/Sema4AKO mice (n = 4 mice per group). B. Bone marrow cellularity of aged WT/Sema4AKO mice (n = 4 mice per group). C. Gating strategy for flow cytometric analysis of primitive bone marrow subsets (WT aged mice are shown). LT-HSC denotes long-term HSC. D. Gating strategy for flow cytometric identification of myHSC and lyHSC based on intensity of CD150 expression. E. Absolute numbers of myHSC and lyHSC in young and aged WT mice (n = 6 mice for young group and n = 4 mice for aged group) F. Representative plots of myHSC quantification in aged WT/Sema4KO mice. G. Absolute number of myHSC and lyHSC in individual aged WT/Sema4AKO mice. P values are shown. Statistical significance was assessed by two-tailed t-test. Mean +/- (SEM) are shown.
Extended Data Fig. 2 The absence of Sema4A leads to functional attrition of myHSC during aging.
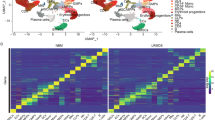
A. Gating strategy for quantification of overall reconstitution by donor-derived HSC and analysis of lineage composition of donor-derived graft. B. Myeloid/lymphoid ratio of peripheral blood donor-derived cells (CD45.2) in WT mice (CD45.1) which were competitively transplanted with aged WT myHSC or aged WT lyHSC (CD45.2) (n = 5 animals per group). C. Gating strategy for quantification of lineage contribution by donor-derived cells. D. Distribution of pairwise Spearman’s correlation distances between aged WT and Sema4AKO myHSC (left) and lyHSC (right) (n = 642 cells across 2 biological replicates and 4 technical replicates). In box plot, centre line shows median, box limits indicate upper and lower quartiles, whiskers extend to minimum and maximum values E. Volcano plots showing DEGs in myHSC from aged WT and Sema4AKO mice. The x and y axes indicate the expression fold change (FC) (log2) and the false discovery rate (FDR) ( − log10) for each gene versus controls, respectively. Legends highlight upregulated (red) or downregulated (blue) transcripts, as well as genes not passing cutoff criteria for FC (black) and FDR (gray). Selected representative genes are shown (n = 2 biological replicates per genotype). F. Distribution of diffusion pseudotime values of myHSC and lyHSC from aged WT mice (n = 642 cells across 2 biological replicates and 4 technical replicates). In the inner box plots of the violinplots, the white point shows median value, box limits indicate upper and lower quartiles, whiskers extend to minimum and maximum values. P values are shown. Statistical significance was assessed by two-tailed t-test (*P < 0.05, **P < 0.01), except for the diffusion pseudotime analysis where two-tailed Wilcoxon rank sum test was used. Mean +/- SEM are shown.
Extended Data Fig. 3 The absence of Sema4A in young animals leads to selective myHSC hyperactivation at the steady state.
A, B. MyHSC cell cycle analysis (n = 6 mice per group) (A) and myHSC short-term (4-day) BrdU incorporation (n = 5 mice per group) (B) in WT/Sema4AKO mice. C, D. LyHSC cell cycle analysis (n = 6 mice per group) (C) and lyHSC short-term (4-day) BrdU incorporation (n = 4 mice per group) (D) in WT/Sema4AKO mice. E. Myeloid/lymphoid ratio of peripheral blood donor-derived cells in WT mice (CD45.1) which were competitively transplanted with WT myHSC or LyHSC (CD45.2). F, G. Frequency (F) and mean fluorescent intensity of Flk2 expression (G) of MPP4 in the bone marrow of WT mice (CD45.1) after competitive transplantation of WT/Sema4AKO myHSC (myHSC recipients) (CD45.2) and WT/Sema4AKO lyHSC (lyHSC recipients) (CD45.2). Quantification and representative plots are shown (n = 5 for myHSC recipient groups, n = 4 for WT lyHSC recipient group and n = 3 for Sema4AKO lyHSC recipient group). P values are shown. Statistical significance was assessed by two-tailed t-test. Mean +/- SEM are shown.
Extended Data Fig. 4 Sema4AKO myHSC are hypersensitive to acute innate immune activation.
A. Absolute number of WT myHSC/lyHSC at baseline and post-LPS injection at indicated time points (n = 5 mice for baseline group, n = 6 mice for 72 hours group, n = 7 mice for 120 hours group, n = 4 mice for 168 hours group). B. Representative flow cytometry plots of myHSC at baseline and 72 hours after LPS injection. C. Absolute number of WT MPPs at baseline and post-LPS injection at indicated time points (n = 5 mice for baseline group, n = 6 mice for 72 hours group, n = 7 mice for 120 hours group, n = 4 mice for 168 hours group). D. Frequency of myHSC and MPPs in WT/Sema4AKO mice 72 hours post-LPS injection (n = 4 mice per group). P values are shown. *P < 0.05, **P < 0.01. Statistical significance was assessed by two-tailed t-test. Mean +/- SEM are shown.
Extended Data Fig. 5 PlxnD1fl/fl is a functional receptor for Sema4A on myHSC.
A. Expression of PlxnB2 in myHSC and lyHSC in WT mice (n = 3 mice per group). B. Frequency of PlxnB2-expressing myHSC and lyHSC in WT mice (n = 3 mice per group). C, D. Expression of PlxnD1 mRNA in WT myHSC (n = 4 mice per group) (C) and percentage of Nrp1-expressing WT myHSC (D) 72 hours post-LPS injection, as assessed by RNA-Seq and flow cytometry, respectively (n = 4 mice for PBS group and n = 5 mice for LPS group). E. Frequency of PlxnD1+ CD34+ hematopoietic stem/progenitor cells in patients with sepsis, as assessed by scRNA Seq in Kwok et al61. Displayed is a cumulative analysis of “non-zero” PlxnD1 expression values in single HSPC obtained from 26 patients with sepsis and 6 healthy volunteers. F, G. PlxnD1 excision validation in Lin-Kit+Sca1+ cells by genomic DNA PCR (n = 2 mice per group) (F) and Q-PCR (n = 7 mice for PlxnD1fl/fl Cre(-) group and n = 3 mice for PlxnD1fl/fl Cre(+) group) (G). H-L. Absolute number of myHSC and lyHSC (n = 10 mice for PlxnD1fl/fl Cre(-) group and n = 6 mice for PlxnD1fl/fl Cre(+) group) (H), other primitive hematopoietic cells (n = 10 mice for PlxnD1fl/fl Cre(-) group and n = 6 mice for PlxnD1fl/fl Cre(+) group) (I), mature cells (n = 7 mice for PlxnD1fl/fl Cre(-) group and n = 4 mice for PlxnD1fl/fl Cre(+) group) (J), myHSC cell cycle analysis (n = 7 mice for PlxnD1fl/fl Cre(-) group and n = 4 mice for PlxnD1fl/fl Cre(+) group) (K), lyHSC cell cycle analysis (n = 7 mice for PlxnD1fl/fl Cre(-) group and n = 4 mice for PlxnD1fl/fl Cre(+) group) (L) in PlxnD1fl/fl Mx1Cre(+) and PlxnD1fl/fl Cre(-) mice at baseline (n = 5 mice for PlxnD1fl/fl Cre(-) group and n = 3 mice for PlxnD1fl/fl Cre(+) group). M. Frequency of LT-HSC in the bone marrow of WT mice (CD45.1) after competitive transplantation of PlxnD1fl/fl Mx1Cre(+) and PlxnD1fl/fl Cre(-) myHSC (myHSC recipients) (CD45.2) and PlxnD1fl/fl Mx1Cre(+) and PlxnD1fl/fl Cre(-) lyHSC (lyHSC recipients) (CD45.2) (n = 5 mice for myHSC receipient groups, n = 4 mice for PlxnD1fl/fl Cre(-) lyHSC recipient group and n = 5 mice for PlxnD1fl/fl Cre(+) lyHSC recipient group). N, O. Frequency (N) and mean fluorescent intensity of Flk2 expression (O) of MPP4 in the bone marrow of WT mice (CD45.1) after competitive transplantation of PlxnD1fl/fl Mx1Cre(+) and PlxnD1fl/fl Cre(-) myHSC (myHSC recipients) and PlxnD1fl/fl Mx1Cre(+) and PlxnD1fl/fl Cre(-) lyHSC (lyHSC recipients), quantification and representative plots are shown for MPP4 frequency (n = 5 mice for myHSC receipient groups, n = 4 mice for PlxnD1fl/fl Cre(-) lyHSC recipient group and n = 5 mice for PlxnD1fl/fl Cre(+) lyHSC recipient group). P. Frequency of myHSC and MPPs in PlxnD1fl/fl Mx1Cre(+) and PlxnD1fl/fl Cre(-) mice 72 hours post-LPS injection (n = 3 mice for PlxnD1fl/fl Cre(-) group and n = 4 mice for PlxnD1fl/fl Cre(+) group). P values are shown. Statistical significance was assessed by two-tailed t-test. Mean +/- SEM are shown.
Extended Data Fig. 6 Neutrophils serve as a physiologically important source of Sema4A.
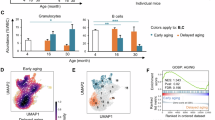
A, B. Sema4A mRNA (A) and protein expression (B) at baseline, 24 hours after LPS injection and upon aging (n = 3 mice per group per condition). C. Relative contribution of distinct cellular subsets to Sema4A production in the bone marrow 24 hours after LPS injection and upon aging (n = 4 mice per group). D. Representative calviarial intra-vital microscopy images of WT myHSC transplanted into lethally irradiated WT/Sema4AKO recipients. Red – progeny of transplanted myHSC, green – bone, blue – collagen. Scale bar – 10 microne. E. Relative contribution of distinct cellular subsets to Sema4A production in the bone marrow 24 hours after 950 cGy irradiation, as estimated by flow cytometry with Sema4A antibody (n = 3 mice per group). Bar graph and representative flow cytometry plots are shown. F. Gating strategy for assessing Sema4A expression in Ly6Ghigh vs Ly6Glow neutrophils. G. Quantification and representative histogram of Sema4A deletion from neutrophils in Sema4Afl/fl Mrp8-Cre(+) mice (n = 5 mice per group). Sema4A antibody-stained neutrophils from Sema4AKO mouse were used as a negative control. H-J. Absolute number of primitive hematopoietic cells (H), myHSC and lyHSC (I) and mature cells (J) at baseline in Sema4Afl/fl Mrp8-Cre(+) and Sema4Afl/fl Mrp8-Cre(-) mice (n = 4 mice per group). K. Frequency of myHSC and MPPs in Sema4Afl/fl Mrp8-Cre(+) and Sema4Afl/fl Mrp8-Cre(-) mice 72 hours after LPS injection (n = 6 mice for Sema4Afl/fl Mrp8-Cre(-) group and n = 5 mice for Sema4Afl/fl Mrp8-Cre(+) group). L. Cell cycle analysis of myHSC from Sema4Afl/fl Mrp8-Cre(+) and Sema4Afl/fl Mrp8-Cre(-) mice 72 hours after LPS injection (n = 6 mice for Sema4Afl/fl Mrp8-Cre(-) group and n = 5 mice for Sema4Afl/fl Mrp8-Cre(+) group). M, N. Overall percentage of donor-derived peripheral blood cells (M) and lineage contribution by donor-derived cells (N) in WT mice (CD45.1) which were competitively transplanted with myHSC from low-dose LPS-treated Sema4Afl/fl Mrp8-Cre(+) and Sema4Afl/fl Mrp8-Cre(-) (CD45.2) mice (n = 5 mice per group). O. Frequency of Sema4A+ cells in peripheral blood lymphoid subsets from patients with sepsis and healthy volunteers (HV), as assessed by single cell RNA-Seq in Kwok et al.61. Displayed is a cumulative analysis of “non-zero” PlxnD1 expression values in single HSPC obtained from 26 patients with sepsis and 6 healthy volunteers. The plots show the median (middle line), interquartile range (box) and minimum to maximum values (whiskers) throughout. P. Sema4A mean fluorescent intensity of Sema4A+Ly6Ghigh neutrophils in the bone marrow of young and aged WT mice (n = 5 mice for young WT group and n = 4 mice for aged WT group). P values are shown. Statistical significance was assessed by two-tailed t-test. Mean +/- SEM are shown.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes 1–3, Figs. 1–3 and figure legends.
Supplementary Table 1 (download PDF )
List of antibodies.
Supplementary Table 2 (download PDF )
List of oligonucleotides.
Source data
Source Data Fig. 1 (download XLSX )
Statistical Source Data.
Source Data Fig. 2 (download XLSX )
Statistical Source Data.
Source Data Fig. 3 (download XLSX )
Statistical Source Data.
Source Data Fig. 4 (download XLSX )
Statistical Source Data.
Source Data Fig. 5 (download XLSX )
Statistical Source Data.
Source Data Fig. 6 (download XLSX )
Statistical Source Data.
Source Data Fig. 7 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical Source Data.
Source Data Extended Data Fig. 5 (download JPG )
Uncropped gel.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical Source Data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Toghani, D., Gupte, S., Zeng, S. et al. Niche-derived Semaphorin 4A safeguards functional identity of myeloid-biased hematopoietic stem cells. Nat Aging 5, 558–575 (2025). https://doi.org/10.1038/s43587-024-00798-7
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43587-024-00798-7
This article is cited by
-
The ageing immune system as a driver of systemic ageing
Nature Reviews Immunology (2026)