Abstract
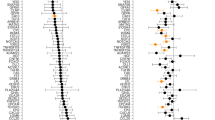
Cellular senescence increases with age and contributes to age-related declines and pathologies. We identified circulating biomarkers of senescence and related them to clinical traits in humans to facilitate future noninvasive assessment of individual senescence burden, and efficacy testing of novel senotherapeutics. Using a nanoparticle-based proteomic workflow, we profiled the senescence-associated secretory phenotype (SASP) in THP-1 monocytes and examined these proteins in 1,060 plasma samples from the Baltimore Longitudinal Study of Aging. Machine-learning models trained on THP-1 monocyte SASP associated SASP signatures with several age-related phenotypes in a test cohort, including body fat composition, blood lipids, inflammatory markers and mobility-related traits, among others. Notably, a subset of SASP-based predictions, including a high-impact SASP panel, were validated in InCHIANTI, an independent aging cohort. These results demonstrate the clinical relevance of the circulating SASP and identify potential senescence biomarkers that could inform future clinical studies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All raw MS data files and associated quantitative and statistical reports, metadata and supplementary data are available on MassIVE (dataset identifier: MSV000095315). FTP download link: ftp://massive.ucsd.edu/v08/MSV000095315/. The aggregated phenotype data have been provided as source data as well as supplementary tables. The non-aggregated clinical and SomaScan data are private because the participants did not consent to unrestricted data sharing at the time of the study conducted for BLSA. To comply with patient consent and data-sharing agreements, researchers are welcome and encouraged to request use of more detailed BLSA data for scientific projects by developing a pre-analysis plan that can be submitted for approval (https://blsa.nia.nih.gov/how-apply).
Code availability
The R scripts for the core LASSO analysis described60,61 are available at https://github.com/geroproteomics/EN_Repeat/ and https://github.com/geroproteomics/EN_Test.
References
Lee, K. A., Flores, R. R., Jang, I. H., Saathoff, A. & Robbins, P. D. Immune senescence, immunosenescence and aging. Front. Aging 3, 2673–6217 (2022).
Yousefzadeh, M. J. et al. An aged immune system drives senescence and ageing of solid organs. Nature 594, 100–105 (2021).
Ong, S.-M. et al. The pro-inflammatory phenotype of the human non-classical monocyte subset is attributed to senescence. Cell Death Dis. 9, 266 (2018).
Saul, D. et al. A new gene set identifies senescent cells and predicts senescence-associated pathways across tissues. Nat. Commun. 13, 4827 (2022).
Lin, Y. et al. Circulating monocytes expressing senescence-associated features are enriched in COVID-19 patients with severe disease. Aging Cell 22, 1474–9726 (2023).
Dey, A. K. et al. Translating senotherapeutic interventions into the clinic with emerging proteomic technologies. Biology https://doi.org/10.3390/biology12101301 (2023).
Suryadevara, V. et al. SenNet recommendations for detecting senescent cells in different tissues. Nat. Rev. Mol. Cell Biol. https://doi.org/10.1038/s41580-024-00738-8 (2024).
Basisty, N., Kale, A., Patel, S., Campisi, J. & Schilling, B. The power of proteomics to monitor senescence-associated secretory phenotypes and beyond: toward clinical applications. Expert Rev. Proteom. 17, 297–308 (2020).
Fielding, R. A. et al. Associations between biomarkers of cellular senescence and physical function in humans: observations from the lifestyle interventions for elders (LIFE) study. Geroscience 44, 2757–2770 (2022).
Tanaka, T. et al. Plasma proteomic signature of age in healthy humans. Aging Cell 17, e12799 (2018).
Tanaka, T. et al. Plasma proteomic biomarker signature of age predicts health and life span. eLife 9, e61073 (2020).
Ubaida-Mohien, C. et al. Discovery proteomics in aging human skeletal muscle finds change in spliceosome, immunity, proteostasis and mitochondria. eLife https://doi.org/10.7554/eLife.49874 (2019).
Tsitsipatis, D. et al. Proteomes of primary skin fibroblasts from healthy individuals reveal altered cell responses across the life span. Aging Cell 21, e13609 (2022).
Shin, J.-W., Lee, E., Han, S., Choe, S.-A. & Jeon, O. H. Plasma proteomic signature of cellular senescence and markers of biological aging among postmenopausal women. Rejuvenation Research 25, 141–148 (2022).
Basisty, N. et al. A proteomic atlas of senescence-associated secretomes for aging biomarker development. PLoS Biol. 18, e3000599 (2020).
Blume, J. A.-O. X. et al. Rapid, deep and precise profiling of the plasma proteome with multi-nanoparticle protein corona. Nat. Commun. 11, 3662 (2020).
Neri, F., Basisty, N., Desprez, P.-Y., Campisi, J. & Schilling, B. Quantitative proteomic analysis of the senescence-associated secretory phenotype by data-independent acquisition. Curr Protoc 1, e32 (2021).
Starr, T., Bauler, T. J., Malik-Kale, P. & Steele-Mortimer, O. The phorbol 12-myristate-13-acetate differentiation protocol is critical to the interaction of THP-1 macrophages with Salmonella typhimurium. PLoS ONE 13, e0193601 (2018).
Knecht, S. et al. An introduction to analytical challenges, approaches, and applications in mass spectrometry-based secretomics. Mol. Cell Proteom. 22, 100636 (2023).
Ferdosi, S. et al. Engineered nanoparticles enable deep proteomics studies at scale by leveraging tunable nano-bio interactions. Proc. Natl Acad. Sci. USA 119, e2106053119 (2022).
Donovan, M. K. R. et al. Functionally distinct BMP1 isoforms show an opposite pattern of abundance in plasma from non-small cell lung cancer subjects and controls. PLoS ONE 18, e0282821 (2023).
Candia, J., Daya, G. N., Tanaka, T., Ferrucci, L. & Walker, K. A. Assessment of variability in the plasma 7k SomaScan proteomics assay. Sci. Rep. 12, 17147 (2022).
Cordon, J. et al. Identification of clinically relevant brain endothelial cell biomarkers in plasma. Stroke 54, 2853–2863 (2023).
Searle, S. D., Mitnitski, A., Gahbauer, E. A., Gill, T. M. & Rockwood, K. A standard procedure for creating a frailty index. BMC Geriatr. 8, 24 (2008).
Tanaka, T. et al. Quality specific associations of carbohydrate consumption and frailty index. Nutrients https://doi.org/10.3390/nu14235072 (2022).
Evans, D. S. et al. Proteomic analysis of the senescence-associated secretory phenotype: GDF-15, IGFBP-2, and cystatin-C are associated with multiple aging traits. J. Gerontol. A Biol. Sci. Med. Sci. https://doi.org/10.1093/gerona/glad265 (2024).
Stastna, M. & Van Eyk, J. E. Investigating the secretome: lessons about the cells that comprise the heart. Circ. Cardiovasc. Genet. 5, o8–o18 (2012).
Samsonraj, R. M., Law, S. F., Chandra, A. & Pignolo, R. J. An unbiased proteomics approach to identify the senescence-associated secretory phenotype of human bone marrow-derived mesenchymal stem cells. Bone Rep. 18, 101674 (2023).
Ozcan, S. et al. Unbiased analysis of senescence associated secretory phenotype (SASP) to identify common components following different genotoxic stresses. Aging 8, 1316–1329 (2016).
Wiley, C. D. et al. SILAC analysis reveals increased secretion of hemostasis-related factors by senescent cells. Cell Rep. 28, 3329–3337 e3325 (2019).
Herranz, N. et al. mTOR regulates MAPKAPK2 translation to control the senescence-associated secretory phenotype. Nat. Cell Biol. 17, 1205–1217 (2015).
Laberge, R. M. et al. MTOR regulates the pro-tumorigenic senescence-associated secretory phenotype by promoting IL1A translation. Nat. Cell Biol. 17, 1049–1061 (2015).
Suhre, K. et al. Nanoparticle enrichment mass-spectrometry proteomics identifies protein-altering variants for precise pQTL mapping. Nat. Commun. 15, 989 (2024).
Huang, T. et al. Protein coronas on functionalized nanoparticles enable quantitative and precise large-scale deep plasma proteomics. Preprint at bioRxiv https://doi.org/10.1101/2023.08.28.555225 (2023).
Coppe, J. P., Desprez, P. Y., Krtolica, A. & Campisi, J. The senescence-associated secretory phenotype: the dark side of tumor suppression. Annu. Rev. Pathol. 5, 99–118 (2010).
Ishaq, A., Tchkonia, T., Kirkland, J. L., Siervo, M. & Saretzki, G. Palmitate induces DNA damage and senescence in human adipocytes in vitro that can be alleviated by oleic acid but not inorganic nitrate. Exp. Gerontol. 163, 111798 (2022).
Xue, W. J. et al. High glucose and palmitic acid induces neuronal senescence by NRSF/REST elevation and the subsequent mTOR-related autophagy suppression. Mol. Brain 15, 61 (2022).
Maeda, M., Hayashi, T., Mizuno, N., Hattori, Y. & Kuzuya, M. Intermittent high glucose implements stress-induced senescence in human vascular endothelial cells: role of superoxide production by NADPH oxidase. PLoS ONE 10, e0123169 (2015).
Minamino, T. et al. A crucial role for adipose tissue p53 in the regulation of insulin resistance. Nat. Med. 15, 1082–1087 (2009).
Schafer, M. J. et al. Exercise prevents diet-induced cellular senescence in adipose tissue. Diabetes 65, 1606–1615 (2016).
Xu, M. et al. Senolytics improve physical function and increase lifespan in old age. Nat. Med. 24, 1246–1256 (2018).
Aversa, Z. et al. Calorie restriction reduces biomarkers of cellular senescence in humans. Aging Cell 23, e14038 (2024).
St Sauver, J. L. et al. Biomarkers of cellular senescence and risk of death in humans. Aging Cell 22, e14006 (2023).
Korbecki, J., Olbromski, M. & Dziegiel, P. CCL18 in the progression of cancer. Int. J. Mol. Sci. https://doi.org/10.3390/ijms21217955 (2020).
Singh, A. et al. CCL18 aggravates atherosclerosis by inducing CCR6-dependent T-cell influx and polarization. Front. Immunol. 15, 1327051 (2024).
Tchkonia, T., Zhu, Y., Deursen, J. V., Campisi, J. & Kirkland, J. L. Cellular senescence and the senescent secretory phenotype: therapeutic opportunities. J. Clin. Invest. 123, 966–972 (2013).
Tsuchiya, S. et al. Establishment and characterization of a human acute monocytic leukemia cell line (THP-1). Int. J. Cancer 26, 171–176 (1980).
Tsuchiya, S. et al. Induction of maturation in cultured human monocytic leukemia cells by a phorbol diester1. Cancer Res. 42, 1530–1536 (1982).
Pham, T. V., Henneman, A. A. & Jimenez, C. R. iq: an R package to estimate relative protein abundances from ion quantification in DIA-MS-based proteomics. Bioinformatics 36, 2611–2613 (2020).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Shock, N. W. Normal Human Aging: the Baltimore Longitudinal Study of Aging (US Department of Health and Human Services, Public Health Service, National Institutes of Health, National Institute on Aging, Gerontology Research Center, 1984).
Ferrucci, L. The Baltimore Longitudinal Study of Aging (BLSA): a 50-year-long journey and plans for the future. J. Gerontol. A Biol. Sci. Med. Sci. 63, 1416–1419 (2008).
Schrack, J. A. et al. Assessing the “physical cliff”: detailed quantification of age-related differences in daily patterns of physical activity. J. Gerontol. A Biol. Sci. Med. Sci. 69, 973–979 (2014).
Ferrucci, L. et al. Subsystems contributing to the decline in ability to walk: bridging the gap between epidemiology and geriatric practice in the InCHIANTI study. J. Am. Geriatr. Soc. 48, 1618–1625 (2000).
Candia, J. A.-O. et al. Assessment of variability in the SOMAscan assay. Sci. Rep. 7, 2045–2322 (2017).
Duggan, M. R. et al. Proteome-wide analysis identifies plasma immune regulators of amyloid-β progression. Brain Behav. Immun. 120, 604–619 (2024).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Olinger, B. EN_Repeat. Zenodo https://doi.org/10.5281/zenodo.15042404 (2025).
Olinger, B. EN_Test. Zenodo https://doi.org/10.5281/zenodo.15042410 (2025).
Acknowledgements
This work was supported by the National Institute on Aging (NIA) Intramural Research Program, NIH. N.B. was supported by a SenNet NIH Common Fund Grant (NIA U54 AG079779, PI Elisseeff) and a Hevolution GRO grant (HF-GRO-23-1199068-44). We gratefully acknowledge L. Brick and NIDA/NIA Visual Media for assistance in figure preparation and G. Howard for editing of the manuscript.
Author information
Authors and Affiliations
Contributions
R.B., D.T., A.R., T.N., M.S. and L.C. performed the experiments. B.O. performed the computational analysis. A.D. was involved in MS data analysis. R.B., B.O. and N.B. prepared the manuscript. T.T., G.D., Z.P. and J.C. helped in performing clinical associations. E.S., K.W. and L.F. provided the clinical data. M.G. and N.B. provided the facilities and guidance for the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks Sundeep Khosla, Yannick van Sleen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Optimization of IR-induced senescence in THP-1 monocytes.
a, Line plot indicating the cell number after exposure to different doses of IR indicates inhibition of cell proliferation up to 7 days after exposure to different doses of IR. b–e, Bar plots showing increased expression of known senescence markers over time after exposure to different doses of IR. f, Bar plots showing elevated SPiDER β-gal confirming induction of senescence in IR-treated THP-1 cells. Statistical analysis was performed using two-tailed Students’ t-test (**P < 0.01). g, Representative fluorescence microscopy images from Edu incorporation assay after exposure to different doses of IR indicating inhibition of cell proliferation on Day 7 after exposure to different doses of IR. Data are represented as mean +/− standard deviation for n number of replicates in all graphs.
Extended Data Fig. 2 Validation of Cell Identity after IR Treatment.
a, Classical SA Beta-gal staining performed 7 days after IR treatment showed increased number of beta galactosidase positive cells as indicated by the formation of blue color in senescent cells as compared to the proliferating controls. b, Brightfield microscopy images to assess any change in morphology after senescence induction shows conservation of suspension cell characteristic of monocytes 7 days after IR treatment. Flow cytometry analysis performed to confirm cell identity after IR treatment shows no increase the monocyte differentiation markers c, CD14 and d, CD11b indicating conservation of monocyte cell lineage after IR treatment.
Extended Data Fig. 3 NanoParticle Analysis.
a, Peptide expression level is compared between quantification methods of 3 proliferating and 3 senescent monocyte samples, using NanoParticles or Neat (control). Only human peptides present in at least 4 of 6 samples, showing differential expression (Pval < 0.05, t-tests) in neat samples were included (n = 428). b, Protein expression level is compared between quantification methods of 3 proliferating and 3 senescent monocyte samples, using NanoParticles or Neat (control). Only human proteins present in at least 4 of 6 samples, and showing differential expression (Pval < 0.05, t-tests) in neat samples were included (n = 132) For all figures, human mapped peptides and proteins were used. c, CV (SD / Mean log10 intensities) was calculated for all proteins present in at least 2 of 3 Senescent samples and 2 of 3 Proliferating samples in Both Neat and NP MaxRep rollup (N = 919). d, The number of identified peptides in any sample in either NP or Neat treatments, classified as either human, bovine, or mixed. e, The distribution of the peptide log10 intensities sorted by species, quantified using either the standard (Neat) or Proteograph (NP) workflows. f, The sum of the log10 intensities sorted by species, quantified using either the standard (Neat) or Proteograph (NP) workflows.
Extended Data Fig. 4 LSPs Show Age-Independent Predictive Potential.
a, Pearson linear models were constructed using covariates only (age, sex, race, and eGFR), LSPs only, or LSPs and covariates. Correlation coefficients for all models are shown by trait. b, This analysis is repeated but including fat percent as a covariate. Pearson linear models were constructed using covariates only (age, sex, race, and eGFR, and fat percent), LSPs only, or LSPs and covariates. Correlation coefficients for all models are shown by trait.
Extended Data Fig. 5 Senescence Signatures Predict Frailty.
a, LASSO modeling was used with age, sex, race, and eGFR as covariates in one model, and with CRP and IL-6 added as two additional covariates in a second model, for feature selection of proteins implicated in either BMI or walking pace. Selected features were used for trait prediction (90% train, 10% test). b, Senescence signatures were selected for a 44-component frailty index. Ten rounds of cross-validated (90% train, 10% test) trait prediction using a linear model with LSPs only was performed. Total sample sizes per trait are indicated in Table 1. c, Proteins chosen via machine learning and positively associated with frailty are shown by their association with the frailty index using linear modeling, with covariates age, sex, race, and eGFR.
Extended Data Fig. 6 LASSO-Selected Proteins by Cohort.
a, 220 monocyte SASP were detected in both the BLSA (7k SomaScan) and Inchianti (1.3k SomaScan). LASSO modeling was used for feature selection in both Inchianti and BLSA. The number of LSPs selected via LASSO for each train in the BLSA are shown, including 220 SASP and 2 covariates (age and sex). b, LSPs selected in InCHIANTI, including 220 SASP and 2 covariates (age and sex). c, Linear models were constructed using only proteins selected in both studies for each trait. Spearman’s correlation of predicted values of linear models trained on the BLSA and observed values in InCHIANTI. d, Spearman’s correlation of predicted values of linear models trained on InCHIANTI and observed values in the BLSA.
Extended Data Fig. 7 Permutation Tests and LASSO Optimized Lambda Values.
a, Principal-Component Analysis was used to condense the high-impact panel into a composite senescence burden score in the BLSA. Principal Component 1 was used to represent an eigengene for the high-impact panel. With the InCHIANTI cohort ranked from low to moderate to high senescence burden, linear trait trends reveal that positive traits HDL and Walking Pace show a negative trend, while negative traits BMI and CRP show a positive trend. b, Permutation tests comparing the predictive potential is shown for LSPs compared with randomly selected proteins. Linear models for each trait were created either using LSPs or randomly selected proteins of the same size. Models were trained on 80% of the data and used to predict the clinical traits for the remaining 20%. Randomly selected proteins models were trained and tested 100,000 times per trait and compared with the accuracy of the LSP-only model. Red dotted lines show where the Spearman’s correlation of the LSP-only model lies in relation to the bell curve for the randomly selected protein models.
Supplementary information
Supplementary Table 1 (download XLSX )
Replicate data from quantification of proteins from MS-based secretomics of senescent and nonsenescent monocytes.
Supplementary Table 2 (download XLSX )
log2 fold changes, P values, FDR of senescence/proliferating proteins in monocytes; age associations (Rho), P values and FDR for circulating plasma proteins in BLSA.
Supplementary Table 3 (download XLSX )
Phenotype table for InCHIANTI study.
Supplementary Table 4 (download XLSX )
ANOVA testing of nested linear models comparing covariate-only model (age, sex, race, eGFR) with ENSPs + covariates model.
Supplementary Table 5 (download XLSX )
Permutation testing comparing ENSP-only linear model predictive accuracy compared to 100,000 iterations of randomly selected proteins.
Supplementary Table 6 (download XLSX )
qPCR primers.
Source data
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Rights and permissions
About this article
Cite this article
Olinger, B., Banarjee, R., Dey, A. et al. The secretome of senescent monocytes predicts age-related clinical outcomes in humans. Nat Aging 5, 1266–1279 (2025). https://doi.org/10.1038/s43587-025-00877-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43587-025-00877-3
This article is cited by
-
Blood proteomics for quantitative biomarkers of cellular therapies
Biomarker Research (2025)
-
Collaboration networks and knowledge structure in cancer and cellular senescence research (2000–2025): a bibliometric perspective
Discover Oncology (2025)