Abstract
Biomechanical alterations contribute to the decreased regenerative capacity of hematopoietic stem cells (HSCs) upon aging. RhoA is a key regulator of mechanosignaling, but its role in mechanotransduction in stem cell aging remains unclear. Here we show that murine HSCs respond to increased nuclear envelope (NE) tension by inducing NE translocation of P-cPLA2, which cell-intrinsically activates RhoA. Aged HSCs experience physiologically higher intrinsic NE tension, but reducing RhoA activity lowers NE tension in aged HSCs. Feature image analysis of HSC nuclei reveals that chromatin remodeling is associated with RhoA inhibition, including restoration of youthful levels of the heterochromatin marker H3K9me2 and a decrease in chromatin accessibility and transcription at retrotransposons. Finally, we demonstrate that RhoA inhibition upregulates Klf4 expression and transcriptional activity, improving aged HSC regenerative capacity and lympho/myeloid skewing in vivo. Together, our data outline an intrinsic RhoA-dependent mechanosignaling axis, which can be pharmacologically targeted to restore aged stem cell function.
Similar content being viewed by others
Main
Aging is characterized by the decline in tissue function and is the primary risk factor for major diseases. In particular, aged hematopoietic stem cell (HSC) functional decline, critically, not only impacts on their ability to regenerate the hematopoietic system and to support lymphoid cell production over time1,2 but also directly contributes to major aging-related diseases3,4. Aging is accompanied by biomechanical changes including alterations in NE tension and in the nucleus mechanical integrity and mechanotransduction5,6. However, how to possibly target changes in nuclear mechanosignaling to explain and prevent aging of somatic stem cells is still largely underinvestigated.
In addition, epigenetic alterations are considered one of the primary hallmarks of aging7, and, despite the large amount of data demonstrating the occurrence of an epigenetic drift upon stem cell aging and disease, there is a lack of knowledge on molecular mechanisms to explain this epigenetic drift and whether it possibly associates with mechanical alterations of the nuclear and chromatin architecture.
Mechanical forces trigger multiple signaling pathways that converge in the activation of RhoA8, which is a small RhoGTPase that can cycle between an active (RhoA-GTP) and an inactive (RhoA-GDP) status. RhoA is a key regulator of mechanotransduction8 regardless of whether the activating mechanical stimulus is cell extrinsic, as occurs in cells responding to alterations of substrate stiffness9, or cell intrinsic, such as, for example, when the cell nucleus acts as a mechanosensor of genomic changes10,11,12. Thus far, in HSCs, RhoA has been shown to be important for cytokinesis13. In vivo, knocking out RhoA in bone marrow (BM) cells induces a dramatic phenotype, characterized by a multilineage hematopoietic failure due to programmed necrosis of hematopoietic progenitors, whereas HSCs retain long-term engraftment potential but fail to produce hematopoietic progenitors and lineage-defined blood cells13.
In the present study, we investigated the role of RhoA in nuclear mechanotransduction in HSCs and its involvement in preserving nuclear architecture and stem cell function upon aging. Our data reveal that RhoA activity increases upon increased NE tension in HSCs, which is intrinsically altered upon aging and can be targeted to improve in vivo function of aged blood stem cells.
Results
RhoA is necessary for HSCs to survive under increased NE tension
The nuclear membrane is able to stretch under various pathophysiological conditions involving nuclear lamina weakening, and stretching of the nucleus is thought to be a fundamental mechanism engaging nuclear mechanotransduction14,15. Therefore, we wondered if nuclear stretching might be engaging RhoA mechanosignaling pathways in HSCs. HSCs are notoriously small-sized non-adherent cells with an average cell diameter of 10 µm and a high nuclear/cytosol ratio. To induce nuclear stretching in HSCs, we took advantage of a previously established cell confinement device11,16 (Extended Data Fig. 1a). We isolated HSCs (sorted as Cd11b−, Ter119−, Cd8−, Cd5−, B220−, Gr1−, c-kit+, Sca1+, Flt3−, CD34−) from the BM of young adult (8−20 weeks old) C57Bl6 mice, and we seeded them on fibronectin-coated coverslips followed by confinement with non-adhesive coverslips functionalized with pillars of 8-µm, 5-µm and 3-µm height for 2 hours (Fig. 1a and Extended Data Fig. 1a,b). Upon confinement, HSCs were fixed and stained with DAPI and with an anti-RhoA-GTP antibody. By three-dimensional (3D) single-cell confocal microscopy, we observed that cell confinement produced a reduction of the nuclear height proportional to the applied level of confinement (Fig. 1b). The maximum nuclear diameter increased significantly already under 8-µm confinement, and it reached its maximum at almost 10 µm when HSCs were confined at 5 µm, showing a significant inverse correlation with nuclear height (Fig. 1b). We observed the formation of nucleoplasm-containing blebs, which was progressive with confinement and particularly prominent when HSCs were confined at 5 µm (Fig. 1c). This can be quantified by looking at the excess of perimeter (EOP) of the largest two-dimensional (2D) slide17 and at the major axis length, which significantly increased in HSCs under the 5-µm confinement conditions (Extended Data Fig. 1c,d). Interestingly, nuclear stretching was paralleled by a progressive alteration of the DAPI intensity profile of the nucleus and by a sharp increase in the activation levels of RhoA, which was mildly but directly correlated with the increase in nuclear diameter (Fig. 1c and Extended Data Fig. 1e). When the nuclear height was reduced to 3 µm, RhoA-GTP levels decreased dramatically. In this condition of tight confinement, HSCs displayed severe nuclear rupture, which suggests cell distress and the inability to react against such a reduction of nuclear height, as evidenced by the dramatic reduction of the nuclear volume and DAPI intensity profile, the appearance of nuclear blebs and the extreme standard deviation variability of the EOP (Fig. 1c, Extended Data Fig. 1c–e and Supplementary Videos 1–4). These data are consistent with a threshold of up to 70% compression of the nuclear height for HSC survival and consequent RhoA activation before irreversible nuclear lamina rupture, in agreement with what was described for other cells17.
a, Representative HSC gating strategy and experimental setup for the confinement experiments. b, Two representative nuclei are depicted to illustrate the strategy to measure the NHA and diameter. NHA is calculated as the sum of n measurements of the nucleus height in the y−z axis, divided by the n number of measurements. NHA and diameter are represented against the different confinement conditions used in the experiments, and a correlation plot between both is shown from three independent experiments (total: unconfined n = 25, 8 µm n = 30, 5 µm n = 16, 3 µm n = 26). Data on graphic bars show mean ± s.e.m., and they were analyzed using Mann−Whitney two-tailed tests, **P < 0.01, ***P < 0.001, ****P < 0.0001. Data on the correlation plot were analyzed by simple linear regression. Pearsonʼs r coefficient and P value are shown, ****P < 0.0001. c, Representative images of 3D immunofluorescence reconstruction from x−y and y−z axes of single HSCs under different levels of confinement of three independent experiments (total: unconfined n = 25, 8 µm n = 30, 5 µm n = 16, 3 µm n = 26). Antibody anti-RhoA-GTP was used to stain active RhoA (red). The nuclei are stained by DAPI (gray). White arrows indicate nucleoplasm-containing blebs. Scale bars, 1 µm. Videos of these confocal acquisitions are available in Supplementary Videos 1–4. Quantification of RhoA-GTP (volume of positive signal by intensity) under different confinements: unconfined, 8 µm, 5 µm and 3 µm. Data on graphic bars show mean ± s.e.m., analyzed by Mann−Whitney test, two-tailed, *P < 0.05, **P < 0.01, ***P < 0.001. Correlation plot in between the RhoA-GTP and the diameter is shown. Data on correlation were analyzed by simple linear regression. Pearsonʼs r coefficient and P value are shown, *P < 0.05. d, Representative images of 3D confocal reconstruction showing LT-HSCs from Rhoafl/fl and Rhoa−/− mice. Cells were treated in vitro with 4-hydoxy tamoxifen overnight and then confined under 5 µm for 2 hours. Cells were stained for RhoA-GTP (red) and DAPI (gray). Graph shows volume in µm3 of RhoA-GTP signal (total: Rhoafl/fl n = 20; RhoaKO n = 12). Data on graphic bars are from three independent experiments showing mean ± s.e.m., analyzed by Mann−Whitney test, two-tailed, ****P < 0.0001. e, Representative images of 3D confocal reconstruction showing RhoA-GTP signal (red) and DAPI (gray) of HSCs treated with NaB. Graph shows volume in µm3 of RhoA-GTP signal. Data on graphic bars are from three independent experiments showing mean ± s.e.m. (total: −NaB n = 31; +NaB n = 40), Mann−Whitney test, two-tailed, ****P < 0.0001. f, Representative images of 3D confocal reconstruction showing DAPI staining of nuclei from young control LT-HSCs and young LT-HSCs treated with NaB. Graph shows the maximum diameter at x−y in µm at each condition. Data on graphic bars are from three independent experiments showing mean ± s.e.m. (total: −NaB n = 22; +NaB n = 63), Mann−Whitney test, one-tailed, *P < 0.05. g, Representative images of 3D confocal reconstruction showing phospho-cPLA2 (PcPLA2) (pink) and DAPI (gray) in young control and young NaB-treated LT-HSCs. By using image analysis software (Imaris and Volocity), we compartmentalized PcPLA2 signal within the nucleus and at the NE. We assigned light pink to NE PcPLA2 and magenta to nuclear PcPLA2. NE PcPLA2 only is also shown. Right: zoomed-in image of the NE localization of PcPLA2, which was reconstructed in light pink. Graph showing the volume in µm3 of PcPLA2 signal at the NE. Videos of representative confocal acquisitions are available in Supplementary Videos 5 and 6. Data on graphic bars are from three independent experiments showing mean ± s.e.m. (total: −NaB n = 38; +NaB n = 25), Mann−Whitney, two-tailed test, ****P < 0.0001. h, Representative images of 3D confocal reconstruction showing HSCs after 16-hour incubation on hydrogels of 1-kPa or 40-kPa stiffness, stained with RhoA-GTP (red) and DAPI (gray). Graph shows volume in µm3 of RhoA-GTP signal from two independent experiments (total: after 1 kPa = 13; after 40 kPa n = 20). Data on graphic bars show mean ± s.e.m., Mann−Whitney test, two-tailed. i, Graphs showing the number of colonies and the number of cells on CFU assays of wild-type and RhoaKO HSCs after incubation on hydrogels of different stiffness (number of independent experiments per condition: wt 1 kPa n = 4; KO 1 kPa n = 8; wt 40 kPa n = 5; KO 40 kPa n = 8). Graphs show mean ± s.e.m., Mann−Whitney test, two-tailed. Scale bars, 1 µm. j,k, Graphs showing hematopoietic progenitor cells (c-Kit+) (j) and myeloid cells (Mac1+Gr1+ and Mac1+) (k) in CFU assays of wild-type and RhoaKO HSCs after incubation on hydrogels at 1 kPa or 40 kPa (number of independent experiments per condition: wt 1 kPa n = 4; KO 1 kPa n = 8; wt 40 kPa n = 5; KO 40 kPa n = 8). Graphs show mean ± s.e.m., Mann−Whitney test, two-tailed, *P < 0.05. l, Graphics depict RhoA-GTP and PcPLA2 under normal conditions and nuclear stretching, either by confinement or NaB treatment. FITC, fluorescein isothiocyanate; LT, long-term; NS, not significant; ST, short-term; LMPP, lymphoid multi-potent progenitor; wk, weeks; wt, wild-type. Panel a and l created with BioRender.
To investigate whether RhoA is necessary for the response to nuclear deformation in HSCs, we isolated HSCs from Cre-ERT2×Rhoaflox/flox mice, in which the activity of the Cre recombinase to knock out Rhoa can be induced in vitro by overnight treatment with 4-hydroxy tamoxifen (Extended Data Fig. 1f; hereafter referred to as Rhoa knockout (KO) HSCs). To note, after 12−16 hours from induction of the Cre-recombinase, levels of RhoA-GTP were clearly reduced (Extended Data Fig. 1g). Next, we confined the cells at 5 µm, according to the protocol described above, to image nuclei and quantify the impact of RhoaKO. Surprisingly, in RhoaKO HSCs, the nuclei were completely broken, and HSCs were unable to resist the 5-µm confinement (Fig. 1d).
Because our experiments were performed in vitro on sorted non-adherent HSCs, we reasoned that RhoA activation might be intrinsically induced by the mechanical tension at the NE. To investigate this hypothesis, we decided to quantify the phosphorylated form of the nuclear protein cPLA2 (P-cPLA2), which translocates to the NE due to a physical process mediated by tension at the NE11,14,15,16. We stained P-cPLA2 and quantified its NE localization, which increased when HSCs are under confinement, consistent with the increased RhoA activation (Extended Data Fig. 1h). NE P-cPLA2 catalyzes the hydrolysis of phospholipids releasing arachidonic acid11,15,16,18,19, which is a well-established activator of RhoA20,21. In agreement, we measured a clear increase in RhoA-GTP levels after treatment of young HSCs with arachidonic acid, confirming the transduction pathway (Extended Data Fig. 1I).
To further investigate whether intrinsic RhoA activation is induced by NE tension, we treated freshly sorted HSCs with sodium butyrate (NaB), a histone deacetylase inhibitor known to increase levels of histone acetylation and induce chromatin decompaction (intrinsically increasing NE tension) in different cells and also in HSCs22,23,24. RhoA activity levels were sharply upregulated in HSCs treated with 5 mM NaB, in parallel with the increase in NE tension, as shown by the increase in the nuclear diameter and P-cPLA2 translocation to the NE (Fig. 1e–g and Supplementary Videos 5 and 6)
To further support that P-cPLA2 translocation to the NE can be triggered by changes in NE tension16, we used hypotonic shock, which was previously used to increase nuclear volume11,15,25 and NE tension, and we quantified NE P-cPLA2 and RhoA activation. Consistently, the data show an increase in NE P-cPLA2 and RhoA-GTP levels in HSCs after hypotonic osmotic shock (Extended Data Fig. 1j).
Overall, the data demonstrate that intrinsic changes in the NE tension activate RhoA in HSCs. However, RhoA activity might also be triggered by the extracellular matrix (niche or extrinsic activation). To explore this possibility, we focused on tissue stiffness, a well-described mechanical stimulus inducing RhoA activity in different cells and tissues26,27. Because, in adult mammals, HSCs reside in the BM, we first measured the stiffness of this compartment, which is a semi-solid tissue with viscoelastic properties and a quite heterogeneous mechanical behavior28. To this end, we used a nanoindenter device equipped with a small and sensitive cantilever tip. We designed a matrix of several points to cover the whole area of the femoral BM tissue and obtain a map of the stiffness of the murine BM cavity (Extended Data Fig. 1k). In agreement with previous observations29, within the BM we detected areas with different levels of stiffness, ranging from 0.5 kPa to 10 kPa (on average, 4 kPa) in the inner marrow to a range of 5−50 kPa (on average, 12 kPa) at the endosteum (Extended Data Fig. 1l). Based on these results, we prepared polyacrylamide hydrogels functionalized with fibronectin to mimic in vitro the lowest (at the inner marrow; approximately 1 kPa) and the highest (endosteum; approximately 40 kPa) stiffness values that we detected in our murine BM samples. We then isolated HSCs and cultured them overnight on the functionalized hydrogels with different stiffness values. HSCs were then recovered and stained for RhoA-GTP and DAPI and used for a colony-forming unit (CFU) assay (Extended Data Fig. 1m). Interestingly, RhoA-GTP levels did not change in HSCs cultured on the stiff (40-kPa) hydrogels compared to those cultured on the 1-kPa hydrogels, contrary to what would be expected for RhoA being activated by an increased stiffness of the substrate30 (Fig. 1h). As for the CFU assay, we did not detect differences in colony number, but the number of c-Kit+ cells (hematopoietic progenitors) and the total cell number were reduced in the 40-kPa stiff hydrogels with no differences in the number of myeloid cells (Gr1+ and Mac1+ cells) (Fig. 1i–k and Extended Data Fig. 1n). Intriguingly, RhoaKO HSCs behaved as their wild-type control for all the measured parameters (Fig. 1i–k and Extended Data Fig. 1n).
Therefore, our data reveal that, in HSCs, RhoA is dispensable in the response to changes in extracellular stiffness, whereas RhoA is necessary to survive intrinsic changes in NE tension. In HSCs, RhoA activity sharply increases after NE tension raise induced by confinement, osmotic shock and chromatin decompaction after NaB treatment (Fig. 1l).
RhoA activity is increased in aged HSCs
Aging alters the biomechanical properties of tissues and cells6. Because increased tissue stiffening upon aging has been reported to induce an increase in RhoA activity levels, for example, in the hair follicle31, we wondered if RhoA activity was altered in aged hematopoietic stem and progenitor cells (HSPCs) and whether it was correlated with increased stiffness of the aged BM32. We profiled the stiffness of the aged BM by performing indentation experiments using the same nanoindenter device as reported above (Extended Data Fig. 1k). We collected several measurements alongside a matrix to cover the overall BM cavity of femurs and tibiae comparing, in parallel, young (8−20 weeks old) and aged C57Bl6 (>80 weeks old) mice. Notably, the indentation measurements showed an overall decreased BM stiffness in aged samples compared to young, resulting in a homogenously low stiffness along the transversal section of the BM (Extended Data Fig. 2a,b). To investigate whether the decreased BM stiffness was associated with changes in RhoA activity, we performed western blot and pulldown assays on HSPCs isolated from young and aged mice. We detected a significant upregulation of RhoA activity levels in aged BM cells that could be targeted and significantly reduced by treatment with a selective RhoA inhibitor (Rhosin, here referred to as Ri)33 (Extended Data Fig. 2c). Ri affects the activity of RhoA and of the two small RhoGTPases most closely related, RhoB/RhoC, by targeting the interaction with guanine nucleotide exchange factors (GEFs) at the interface33. Furthermore, we measured RhoA-GTP levels in sorted HSCs by immunofluorescence. To this end, HSCs were harvested from aged mice and incubated with or without Ri. Young HSCs were sorted alongside as control (Fig. 2a and Extended Data Fig. 2d). Aged HSCs showed a dramatic upregulation of RhoA activity, and treatment with 100 µM Ri significantly decreased levels of RhoA-GTP in aged HSCs (Fig. 2b), consistent with the western blot/pulldown results on HSPCs (Extended Data Fig. 2c). Therefore, in agreement with our conclusions based on the hydrogel experiments (Fig. 1i–k), RhoA activity appears not to be associated with the extracellular stiffness, and, whereas the aged BM stiffness decreases, RhoA activity increases in HSCs upon aging, supporting an intrinsic activation of RhoA in HSCs upon aging.
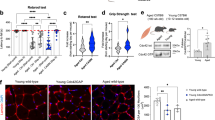
a, Representative HSC gating strategy and experimental setup for in vitro culturing experiments with and without treatment. b, Representative images of 3D confocal reconstruction of young, aged and aged+Ri HSCs stained with anti-RhoA-GTP antibody (red) and DAPI (gray). Right: RhoA-GTP volume based on intensity of the signal. Data on graphic bars are from three independent experiments showing mean ± s.e.m. Total young n = 38, aged n = 33, aged + Ri n = 32, Mann−Whitney two-tailed test, ****P < 0.0001. *P < 0.05, **P < 0.01. c, Representative images of 3D confocal reconstruction and analysis of young, aged and aged treated with AACOCF3 HSCs stained with anti-PcPLA2 showing compartmentalized PcPLA2 at the NE after segmentation. NE PcPLA2 is shown in green and DAPI in gray. The graph shows measurements of percentage volume of PcPLA2 at NE signal normalized against total volume of PcPLA2. Data on graphic bars are from three independent experiments showing mean ± s.e.m. Total young n = 32, aged n = 33, aged + AACOCF3 n = 44, Mann−Whitney two-tailed test, ***P < 0.001. d, Representative images of 3D confocal reconstruction of young, aged and aged AACOCF3-treated HSCs stained with anti-RhoA-GTP antibody (red) and DAPI (gray). The graph shows volume of RhoA-GTP signal. Data on graphic bars are from three independent experiments showing mean ± s.e.m. Total young n = 29, aged n = 28, aged + AACOCF3 n = 33, Mann−Whitney two-tailed test, *P < 0.05, **P < 0.01, ****P < 0.0001. e, Representative images of 3D reconstruction and sections at the three axes, x–y, x–z and y–z, of young, aged and aged Ri-treated LT-HSCs stained with LaminB in yellow and DAPI in gray. Arrows in white point to NE wrinkles. f, Nucleus wrinkling analysis. NE circularity of the nucleus (4πA / P2, with A the area of the nucleus and P the total perimeter of the nucleus) for aged HSCs (n = 29), aged HSCs treated with Ri (n = 32) and young HSCs (n = 30) from three independent experiments. Kruskal−Wallis test for multiple comparisons (****P < 0.0001 and ***P < 0.001). Quantification of NE excess folding parameter (1 − p / P, with p the outline of the nucleus and P the total perimeter of the nucleus) for aged HSCs (n = 29), aged HSCs treated with Ri (n = 32) and young HSCs (n = 30) from three independent experiments. Kruskal−Wallis test for multiple comparisons (****P < 0.0001 and ***P < 0.001). g, Cartoon scheme summarizing the findings. Panel a and g created with BioRender.
Therefore, we hypothesized that RhoA-GTP levels are higher in aged HSCs because their nucleus might be experiencing higher nuclear tension. Consistent with this hypothesis, we detected a significant increase of NE P-cPLA2 in aged stem cells, which is reduced by treatment with the cPLA2 inhibitor AACOCF3 (Fig. 2c and Extended Data Fig. 2e). Notably, also levels of RhoA-GTP in aged cells were sharply reduced after treatment with AACOCF3, supporting that changes in NE tension are necessary to activate RhoA in aged HSCs (Fig. 2d).
To further investigate the increased NE tension in aged stem cells, we performed additional experiments to measure the wrinkling of the NE, a structural feature of nuclear architecture that has previously been used as a measure of NE tension11,15,16,34,35. LaminB staining clearly shows increased NE circularity and reduced NE excess folding in aged HSCs compared to young (Fig. 2e,f), supporting higher NE tension in aged HSCs compared to young. Notably, RhoA inhibition sharply decreases NE circularity and restores NE excess folding in aged HSCs (Fig. 2e,f).
To corroborate further these observations, we also quantified the nuclear import of the mechanosensitive transcription factor TAZ, which is known to accumulate in the nucleus with increasing NE tension36,37. In agreement, we measured increased nuclear translocation of TAZ after 8-µm and 5-µm HSC confinement (Extended Data Fig. 2f). As also reported previously38, we quantified higher level of nuclear TAZ in aged HSCs compared to young, which is dependent on RhoA activity (Extended Data Fig. 2f). Altogether, the results support the functional connection between changes in NE tension and RhoA activity as a mechanosensitive regulator of HSC aging (Fig. 2g).
RhoA inhibition restores DAPI-intense regions in aged HSCs
To explore if changes in NE tension impact on chromatin of aged HSCs, we developed a computational approach leveraging image analysis algorithms to extract morphometric and fluorescence intensity features from 3D confocal images of HSC nuclei stained with DAPI, which is a photostable fluorescent DNA dye used in several applications to quantify DNA amount and chromatin condensation in intact nuclei39,40. First, consistent with increased NE tension, the results show that aged HSCs display a significant increase in nuclear volume, nuclear diameter, surface area, perimeter of the largest z slide and DAPI-intense region (DIR) volume, among other related features, compared to young HSCs (Fig. 3a). To note, HSC confinement (8 µm) induced a similar larger increase in the same morphometric features, supporting that these alterations of nuclear volume, size and shape in aged HSCs are compatible with increased NE tension (Fig. 3a)41.
a, Box plot displaying differences in selected feature distributions among HSCs young (n = 177 nuclei), aged (n = 164 nuclei) and young confined to 8 μm (n = 17 nuclei). Boxes depict the 25%, 50% and 75% quartiles, and the whiskers extend to the rest of the distribution. Outliers are shown as individual datapoints. Mean values are represented by a yellow dot on top of each distribution, with a yellow line aiding to visualize trends among conditions. Statistical significance was determined using the two-sided Mann−Whitney test, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. b, Line plot depicting the normalized DAPI intensity of nuclei obtained from young, aged and aged + Ri conditions as a function of 3D iso-distant intervals of 0.1 μm for up to 1.5 μm from the nuclear border. Inset: a magnified view of the intensity within the initial 0.2 μm from the nuclear border. c, Representative images of young, aged and aged + Ri nuclear intensity images are shown, with DIRs highlighted by a white contour. Box plots showing the standardized DAPI intensity of young (n = 177), aged (n = 164) and aged + Ri (n = 144) nuclei as a function of distance in 0.5-μm intervals. Boxes depict the 25%, 50% and 75% quartiles, and the whiskers extend to the rest of the distribution. Outliers are shown as individual datapoints. Statistical significance was determined using the two-sided Mann−Whitney test, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. d, Scatter plot displaying the UMAP embedding of all nuclei images, with marginal distributions for both components, colored by condition. The estimated kernel density helps to visualize high-density areas for each condition. e, UMAP embedding with background areas colored by clusters identified using the k-means clustering algorithm. Donut charts depicting the proportion of each condition per cluster are included. Marginal distributions for both UMAP components are also displayed. f, Line plots showing the change in average values for selected morphometric-related features across clusters, ordered from the cluster with highest to lowest frequency of young HSCs. Line plots depict mean values ± s.e. per cluster. g, Line plots showing the change in average values for selected intensity-related features across clusters, ordered from the cluster with lowest to highest frequency of aged + Ri HSCs. Line plots depict mean values ± s.e. per cluster. U1, UMAP1; U2, UMAP2. Panel a created with BioRender.
Next, because RhoA inhibition sharply decreases NE circularity and increases NE excess folding in aged HSCs (Fig. 2e,f), we asked whether decreasing RhoA activity might feed back to the nucleus, inducing any change in morphometric and fluorescence intensity features of aged HSC nuclei. The data revealed differences mainly in the pattern of DAPI intensity that we extracted by quantifying fluorescence intensities along 3D iso-distant intervals from the nuclear border (Extended Data Fig. 3a,b), with lower values near the nuclear border and DIRs localizing relatively far from the nuclear border toward the central part of the nucleus for aged HSCs compared to young (Fig. 3c). Most aged+Ri HSC nuclei displayed a large proportion of DIRs near the NE, like young HSCs (Fig. 3c). In addition, also DIR height, DIR major axis length, DIR distance to the border and number of DIRs were restored to a youthful level after RhoA inhibition (Extended Data Fig. 3b).
Next, we computed dimensionality reduction analyses on our feature set to explore similarity patterns within our HSC nucleus images (Extended Data Fig. 3c,d). First, we performed feature engineering and clustering analysis on all available nucleus images to uncover potential mechanisms underlying nuclear remodeling in young, aged and aged Ri-treated HSCs. The extracted imaging features are categorized into three groups: (1) whole-nucleus-level features, (2) DIR-level features and (3) features computed from the largest 2D z slide in the x–y plane. A comprehensive summary of these features is provided in Supplementary Table 2. To address the high dimensionality and notable correlation of our feature space, we used uniform manifold approximation and projection42 (UMAP). Feature selection involved identifying statistically significant features (Mann−Whitney U-test, P < 0.05) among young and aged HSCs and among aged and aged Ri-treated HSCs (Extended Data Fig. 3c,d). These significant features were then combined and filtered to remove those exhibiting high Pearsonʼs correlation coefficient (absolute value of correlation >80%), resulting in a final set of 20 features (Extended Data Fig. 3e). The UMAP revealed that young, aged and aged+Ri HSCs exhibit overlapping yet distinct distributions, indicating underlying differences in their chromatin properties (Fig. 3d). By grouping the nucleus data points using the k-means clustering algorithm on the original feature space, we identified four distinct clusters that differ in morphometrics and intensity characteristics (Fig. 3e and Extended Data Fig. 3f,g). Analysis of individual features’ impact over the UMAP representation revealed that morphometric-related features polarize the embedding vertically, whereas intensity and DIR-related features polarize the embedding horizontally (Fig. 3e and Extended Data Fig. 3h). Subsequently, by assessing the most representative features and biological populations, we annotated the clusters as low size (LS), high size (HS), border intensity (BI) and central intensity (CI) (Fig. 3e). Feature importance for each cluster was evaluated by measuring feature statistical significance and fold change per cluster (Extended Data Fig. 3I). Cluster BI includes mostly young and aged + Ri HSCs exhibiting high intensity near the NE and lower intensity in the nuclear center, along with an increased number of small DIRs closer to the border, which tend to be less spherical compared to nuclei in other clusters (Fig. 3e–g and Extended Data Fig. 3h,i). Cluster CI predominantly consists of aged nuclei, with a reduced number of aged Ri-treated nuclei. These nuclei present with larger DIRs located farther from the NE and are notably spherical, with decreased intensity near the nuclear border (Fig. 3e–g and Extended Data Fig. 3h,i). Cluster LS mainly contains young nuclei characterized by smaller size and lower mean intensity of DIRs, with surprisingly not much accentuated intensity values near the border (Fig. 3e−g and Extended Data Fig. 3h,i). Cluster HS contains a mix of biological conditions, with nuclei characterized by larger size, DIRs positioned away from the nuclear border and a decreased aspect ratio indicating that these nuclei are wider than they are tall (Fig. 3e–g and Extended Data Fig. 3h,i). By plotting feature trajectories across clusters, morphometric features progressively decrease or increase with decreasing frequencies of young HSCs within the clusters spanning from LS to HS clusters (Fig. 3f). Interestingly, intensity features and DIR-related features progressively decrease or increase with increasing frequencies of aged+Ri HSCs from CI to BI clusters (Fig. 3g). Therefore, aged Ri-treated nuclei appear to share some morphometric similarities with aged nuclei yet exhibit intensity and DIR-related characteristics similar to those of young HSCs.
Overall, our computational approach suggests that DAPI imaging features elucidate chromatin differences in stem cells revealing a Ri-associated nuclear remodeling, which is mainly linked to changes in DAPI intensity and DIR volume, number and localization.
RhoA inhibition restores H3K9me2 at heterochromatin
Alterations in the mechanobiology of the cell have been associated with aging-dependent changes in chromatin architecture6,43, and several epigenetic alterations characterize intrinsic HSC aging44,45, among which it was previously reported a general loss of heterochromatin2,46,47. Intrigued by the observation that in aged HSCs treated with Ri some nuclear DAPI intensity and DIR-related features were significantly reverted to the level found in young HSCs (Fig. 3c and Extended Data Fig. 3b) and are associated with the nuclear remodeling induced by Ri (Fig. 3g), we focused on heterochromatin because especially DIRs are related to the most condensed portion of chromatin. Therefore, we analyzed the levels and distribution of H3K9me2, a known heterochromatin histone mark that is altered in aged HSCs2,24,48. By 3D immunofluorescence analyses, we measured a significant increase of H3K9me2 levels in aged + Ri HSCs compared to aged controls, together with a partial relocalization at the nuclear border like in young HSCs (Fig. 4a). Interestingly, treatment with UNC0646, a selective inhibitor of the histone methyltransferase G9a specific to H3K9me2 (refs. 49,50), blunts completely the effect of Ri on H3K9me2 in aged HSCs (Fig. 4a). These data suggest the requirement of G9a to increase levels of H3K9me2 in aged+Ri HSCs. Moreover, they reveal that aged HSCs are not affected by UNC0646 treatment alone, most likely because of the very low expression of G9a in these cells in basal conditions, which is partially rescued by Ri treatment (Extended Data Fig. 4a). To causally explain the role of decreased H3K9me2 in HSCs, we transduced young HSPCs (LSKs, gated as Lin−Sca-1+c-Kit+) with a retroviral vector, codifying for a histone variant in which the lysine of H3K9 is replaced by an arginine (H3R9) (Fig. 4b and Extended Data Fig. 4b). The arginine in position 9 on H3 cannot be methylated, enforcing H3K9me reduction in HSCs. We functionally validated the strategy and the H3R9 incorporation by transplanting transduced LSKs into lethally irradiated recipient mice. We measured a significant reduction of H3K9me in H3R9 myeloid progenitors compared to control vector transduced H3K9 myeloid progenitors (Fig. 4c). Young H3R9 HSCs isolated from transplanted mice present with higher RhoA-GTP levels and increased nuclear stretching compared to controls (Fig. 4d). Moreover, H3R9 HSCs show a clear premature aging phenotype upon transplantations in vivo, characterized by expansion of LSKs and granulocyte-monocyte progenitors (GMPs), reduced BM and peripheral blood regeneration, reduced B-lymphopoiesis and myeloid skewing (Fig. 4e–g). Altogether, these data demonstrate that reduced methylation of H3K9 causes nuclear stretching, increasing RhoA activation and driving aging of HSCs. Notably, RhoA inhibition restores H3K9 methylation levels in aged HSCs.
a, Left: representative images of 3D confocal reconstruction of young, aged and aged + Ri-treated HSCs stained with anti-H3K9me2 antibody (green) and DAPI (gray). Right: representative images of 3D confocal reconstruction of young, aged and aged+Ri with UNC0646 treatment. At least 10 cells were acquired per experiment per condition. Young, aged and aged + Ri data are from eight independent experiments; young UNC0646, aged UNC0646 and aged + Ri UNC0646 data are from four independent experiments. Graph shows H3K9me2 signal volume normalized against DAPI signal volume. Mann−Whitney two-tailed test, total young n = 87, aged n = 79, aged + Ri n = 81; young + UNC n = 36, aged + UNC n = 41, aged + Ri + UNC n = 41, showing mean ± s.e.m., analyzed by Mann−Whitney test, **P < 0.01, *P < 0.05, ****P < 0.0001. b, Experimental setup for retroviral transduction and transplantation of young LSK overexpressing wild-type H3K9 or mutant H3R9 with mCherry as a reporter. Transduced cells were transplanted into lethally irradiated mice. BM of transplanted mice was analyzed by flow cytometry to evaluate the regenerative capacity of transduced HSCs. mCherry+ myeloid progenitors (MPs) were sorted to measure H3K9me2 levels. mCherry+ HSCs were sorted for the analysis of RhoA activation and nuclear stretching. c, Representative images of sorted MPs transduced with H3K9 and H3R9 stained with anti-H3K9me2 antibody (magenta) and DAPI (blue). The graph shows H3K9me2 signal volume normalized against DAPI signal volume. H3K9 experimental data are from seven independent experiments and six independent experiments from H3R9, showing mean ± s.e.m., analyzed by Mann−Whitney test, two-tailed. Total H3K9 MP n = 43, H3R9 MP n = 39, n = 7 H3K9; n = 6 H3R9, showing mean ± s.e.m., ****P < 0.0001. d, Representative images of 3D confocal reconstruction of HSCs overexpressing H3K9 or H3R9 isolated from transplanted mice H3K9 experimental data from seven independent experiments, and six independent experiments from H3R9 HSCs were stained with anti-Rho-AGTP antibody (red) and DAPI (gray). Showing mean ± s.e.m., analyzed by Mann−Whitney test, two-tailed. Statistics are shown for Rho-AGTP quantification analysis, **P < 0.01, total K9 n = 56, R9 n = 61. For nuclear volume (DAPI volume), unpaired t-test (one-tailed) was used, *P < 0.05, total K9 n = 56, R9 n = 61. e, Gating strategy for the analysis of BM progenitors 12−20 weeks after transplantation. The analysis of the different populations is shown for mice transplanted with LSK transduced with either wild-type H3K9 (n = 7) or H3R9 (n = 6). Mann−Whitney statistics are shown two-tailed, **P < 0.01. f, Engraftment of mCherry−H3K9+ or mCherry−H3R9+ HSCs in mice. Representative flow cytometry charts for BM engraftment are shown. The box plots display engraftment of mCherry−H3K9+ or mCherry−H3R9+ HSCs in mice 12−20 weeks after transplantation in BM and peripheral blood (PB). Brown−Forsythe and Welch’s ANOVA tests are shown, *P < 0.05, **P < 0.01. g, Gating strategy for the analysis of B cells (B220+), T cells (Cd3+) and myeloid cells (Gr1+ and Mac1+) within the BM 12−24 weeks after transplantation. The analysis of the different populations is shown for mice transplanted with wild-type H3K9 (n = 7) transduced LSK and mutant H3R9 LSK (n = 6). Statistics are B cells, unpaired t-test, two-tailed; T cells, Mann−Whitney test, two-tailed; myeloid cells, Welch’s t-test, two-tailed. *P < 0.05, **P < 0.01. SSC-A, side scatter area. Panel b created with BioRender.
RhoA regulates chromatin accessibility at retrotransposons in aged HSCs
Previously, it was proposed that the cell nucleus can directly respond to mechanical stress by inducing chromatin remodeling, altering polymerase and transcription factor accessibility and activity51,52,53, and increased chromatin accessibility, as measured by assay for transposase-accessible chromatin using sequencing (ATAC−seq), has already been characterized as an epigenetic alteration intrinsic to HSC aging54,55. Intrigued by the possibility that RhoA inhibition might underscore a link between increased NE tension and increased chromatin accessibility in aged HSCs, we investigated further the chromatin remodeling associated with Ri treatment by performing ATAC−seq profiling of sorted young, aged and aged + Ri HSCs (Fig. 5a). Peak calling identified a total of 57,289 accessible regions consistent between samples, most of them located in introns, intergenic regions and gene promoters (Extended Data Fig. 5a,b and Supplementary Table 2). In agreement with previously published data54,55, we detected an increase in open differentially accessible regions (DARs) with aging (2,713 DARs open and 1,103 DARs closed in aged compared to young HSCs; approximately 8% false-positive rate (FPR); Fig. 5b and Supplementary Table 2). After applying the Ri treatment to aged HSCs, 743 chromatin regions opened and 355 closed (approximately 8% FPR; Fig. 5b and Supplementary Table 2). Overall, 144 DARs were detected in both comparisons (aged versus young and aged+Ri versus aged), with 85.42% of them showing accessibility levels after the Ri treatment changing in the direction of the young levels (Fig. 5c, Extended Data Fig. 5c and Supplementary Table 2). Among the regions opening in aged HSCs treated with Ri, Gene Ontology enrichment analysis revealed pathways related to cell migration, morphogenesis, adhesion and chemotaxis (Extended Data Fig. 5d and Supplementary Table 2). Interestingly, no Gene Ontologies were significantly enriched among the DARs closing in aged HSCs+Ri, and a high percentage of these closing DARs were located at retrotransposons (Fig. 5d, lower right panel), especially long terminal repeats (LTRs) and long interspersed elements (LINEs), such as ERVL-MaLR, ERVK, ERV1 and L1 families (2-fold/approximately 1 log2 fold change (log2FC) higher percentage compared to the percentage in consensus peaks; false discovery rate (FDR) = 0.0018 for LTRs and FDR = 0.014 for LINEs in a one-proportion z-test; Fig. 5d,e and Supplementary Table 2)56. Notably, in aged HSCs, we observed an opening of chromatin at retrotransposons, with a 1.65-fold (0.72 log2FC) and a 1.25-fold (0.33 log2FC) increase in the percentage of open DARs localizing in LTRs and LINEs, respectively (FDR = 3.2 × 10−13 for LTRs and FDR = 0.04998 for LINEs in a one-proportion z-test; Fig. 5d,e and Supplementary Table 2). Some of these DARs at LTRs and LINEs were overlapping with enhancers described previously57, whereas others were located in intronic and intergenic regions (Fig. 5e and Supplementary Table 2). Interestingly, retrotransposons and, in particular, LINE-1 have been suggested to directly contribute to aging of somatic cells and aging-related diseases56,58,59,60. To explore if the increase in chromatin accessibility at retrotransposons upon HSC aging is linked to increased NE tension, we performed ATAC−seq of young HSCs under 5-μm confinement and compared their chromatin accessibility profiles to those of unconfined HSCs sorted in parallel from the same mice (Fig. 5f). As a reference for DAR identification, we used the 42,632 consensus peaks identified between young and aged HSC samples (Supplementary Table 3). Data show 443 DARs opening and only 25 DARs closing in young HSCs under confinement (approximately 6.6% FPR; Fig. 5g,h, Extended Data Fig. 5e and Supplementary Table 3). Notably, whereas the DARs that are closing in confined HSCs are located at promoter transcription start site (TSS) and 5′ untranslated region (UTR) (1.67 log2FC over consensus peaks; FDR = 4.1 × 10−6 in a one-proportion z-test; Fig. 5I, lower panel), the DARs that are opening are mainly at LTRs and LINEs (0.75 and 1.18 log2FC over consensus peaks, respectively; FDR = 0.008 for LTRs and FDR = 0.003 for LINEs in a one-proportion z-test; Fig. 5I, upper panel). Overall, the results show a correlation between the increased accessibility at regulatory elements observed in aged HSCs and reverted (closed) by Ri treatment and the type of genomic regions (LTRs and LINEs) that are opened by increasing NE tension by mechanical confinement.
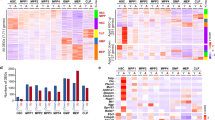
a, Representative HSC gating strategy and experimental strategy for ATAC−seq and RNA-seq of Ri-treated aged HSCs. b, Volcano plot of DARs in aged versus young and aged + Ri versus aged (moderated t-statistic, two-sided, unadjusted P < 0.005). The annotation of selected DARs is shown. Young n = 4, aged n = 5 and aged + Ri n = 5. c, Heatmap of the normalized accessibility in the 144 common DARs in aged versus young and aged+Ri versus aged. d, log2FC of the proportion of each genomic region type among the different sets of DARs over the proportions among the consensus peaks. Fisher’s exact test P value is shown. One-proportion z-test for each genomic region is shown as *FDR < 0.05, **FDR < 0.005, ***FDR < 0.0005, ****FDR < 0.00005. e, Genome tracks for the normalized ATAC−seq read distribution in several LTRs and LINEs. Location of DARs is indicated with purple lines. f, Experimental strategy for ATAC−seq of young HSCs confined in 5 μm. g, Volcano plot of DARs in young 5 μm versus young (moderated t-statistic, two-sided, unadjusted P < 0.001). The annotation of selected DARs is shown. Young n = 3 and young 5 µm n = 3. h, Heatmap of the normalized accessibility in the 468 DARs in young 5 μm versus young. i, log2FC of the proportion of each genomic region type among the different sets of DARs over the proportions among the consensus peaks. ‘nd’ indicates that no regions of that type were detected as DARs. Statistics as in d. j, Heatmap of the normalized expression of the retrotransposon subfamilies upregulated with aging (moderated t-statistic, two-sided, unadjusted P < 0.005). Legend as in c. Box plot shows the normalized expression per condition of the retrotransposon subfamilies downregulated with Ri (moderated t-statistic, two-sided, unadjusted P < 0.005). A red asterisk indicates the retrotransposon subfamilies that show significant differences in both comparisons. Young n = 3, aged n = 3 and aged+Ri n = 3. k, Radar chart showing the normalized enrichment scores (NESs) for some significantly negatively enriched GOs with Ri (permutation test, FDR < 0.05). l–o, Enrichment plot and volcano plot for the acute inflammatory response GO term (l), the interferome.org signature (m), the interferon-stimulated genes (ISG)63 (n) and the aging signature64 (o) in aged + Ri. Permutation test used for GSEA. GO, Gene Ontology. Panel a and f created with BioRender.
Next, we investigated transcriptional changes by bulk RNA sequencing (RNA-seq) analysis of young, aged and aged+Ri HSCs. We identified 38 genes upregulated and 27 downregulated in aged HSCs after treatment with Ri (approximately 5% FPR; Extended Data Fig. 5f–h and Supplementary Table 4). Consistently with the ATAC−seq data, after Ri treatment we detected a downregulation in the transcription of several retrotransposon subfamilies that are upregulated in aged HSCs, mainly LTRs but also some LINEs and DNA transposon subfamilies, such as L1 (L1ME1 and L1ME3A), ERV1 (MER110-int) and hAT-Blackjack (MER63C) (approximately 8% FPR; Fig. 5j, Extended Data Fig. 5i and Supplementary Table 4). Gene set enrichment analysis (GSEA) revealed enrichment for several Gene Ontologies related to inflammation, innate immune response activation and interferon response among the genes downregulated after Ri treatment, again consistent with a downregulation of retrotransposons56,59,61 (FDR < 0.05; Fig. 5k,l, Extended Data Fig. 5j and Supplementary Table 4). Using GSEA, we also detected a decrease in the interferome.org gene signature62 and of interferon-stimulated genes63 in aged+Ri HSCs (Fig. 5m,n and Extended Data Fig. 5k,l). Moreover, we measured a significantly negative enrichment score in aged+Ri versus aged samples for the HSC aging signature defined by Svendsen et al.64, whereas the same signature was clearly positively enriched in the aged samples compared to the young ones (Fig. 5o and Extended Data Fig. 5m).
In summary, by ATAC−seq profiling, we detected changes in chromatin accessibility in aged Ri-treated HSCs that revealed reduction of open regions at LTRs and LINE, belonging mainly to ERVL-MaLR, ERVK, ERV1 and L1 families. ATAC−seq profiling of young 5-µm-confined HSCs supports that the increased accessibility at retrotransposons observed in the aged HSCs and reverted by Ri treatment could be a consequence of increased nuclear stretching. Consistently, by bulk RNA-seq profiling of aged+Ri HSCs, we detected a reduction in the transcription of LTRs and LINEs and a downregulation of the immune response, inflammatory response, interferon response and aging gene signatures compared to aged control HSCs.
Inhibiting RhoA activity improves function of aged HSCs
Next, to further characterize the transcriptional changes in aged HSCs after Ri treatment, we focused on the few upregulated genes (Extended Data Fig. 5g,h and Supplementary Table 4), and, interestingly, we noticed three transcription factors belonging to the same family: Klf4, Klf6 and Klf2 (Fig. 6a and Supplementary Table 4). Noteworthy, the upregulation of Klfs is consistent with the opening of KLF4 motifs detected by ATAC−seq transcription factor binding motif analysis, which revealed enrichment in motifs for AP-1 and KLF4 (Fig. 6b and Supplementary Table 2). In hematopoietic cells, AP-1 can interact with chromatin remodelers to assist in the binding of other transcription factors65, and KLF4 was previously shown to be important in cell reprogramming, blood formation and mechanosensing66,67,68,69,70,71. In addition, we identified an open DAR in aged+Ri HSCs in correspondence with a recently annotated Klf4 enhancer72 (Fig. 6c), and we measured a clear increase in KLF4 protein in the nucleus of aged stem cells after Ri treatment (Extended Data Fig. 6a). Next, we analyzed the expression level of the genes targeted by a higher accessibility of KLF4 binding motifs, and most of them increased their expression after Ri (Extended Data Fig. 6b and Supplementary Table 2). Gene Ontology enrichment analysis of these genes revealed enrichment for morphogenesis, differentiation and actin polymerization (FDR < 0.05; Fig. 6d and Supplementary Table 4). Notably, actin polymerization (filamentous actin (F-actin)) has been reported to restrict nuclear stretching14,15. Consistently, whereas, in aged HSCs, F-actin is decreased compared to young, F-actin levels increase upon Ri treatment similarly to young HSCs (Fig. 6e).
a, Genome tracks for the normalized RNA-seq read distribution in Klf genes. b, Top two transcription factor binding motifs found among the DARs opening with Ri. c, Genome tracks for the normalized ATAC−seq read distribution in Klf4 gene and its enhancer in –119 kb (orange shadow). A green line indicates the DAR opening with Ri. d, Radar chart showing the minus logarithmic FDR for some significantly enriched GOs among the KLF-targeted genes (permutation test, FDR < 0.05). e, Representative images of 3D confocal reconstruction of young, aged and aged + Ri LT-HSCs stained with Phalloidin-488 (green) and DAPI (gray). The graph shows measurements of phalloidin (actin) signal volume of three independent experiments for young and two independent experiments for aged and aged Ri. Bar graph showing mean ± s.e.m., analyzed by Mann−Whitney test, two-tailed, ***P < 0.001, young n = 21, aged n = 17, aged+Ri n = 15. f, Clustered and annotated UMAP for the integrated scRNA-seq data for young (n = 3), aged (n = 3) and aged + Ri (n = 3) LSKs. CLP, common lymphoid progenitor; CMP, common myeloid progenitor; MPP, multipotent progenitor. g, Klf4 normalized expression in the UMAP for young, aged and aged+Ri. Bottom right: difference in the percentage of Klf4+ cells in aged + Ri versus aged per cluster. h, UMAP based on the transcription factor activity scores in HSCs colored by condition. i, KLF4 active cells (AUC > 0.128) in the UMAP. j, Heatmap of the activity scores for the top 30 transcription factors that are differentially active in aged + Ri versus aged (|log2FC| > 0.5 and FDR < 0.05). Color legend as in h. k, Enrichment plot for the hemogenic precursor signature77 in aged + Ri versus aged. Permutation test. AUCPct., percent.
To gain further insights into the transcriptional rewiring of aged HSCs treated with Ri, we performed single-cell RNA sequencing (scRNA-seq) of young, aged and aged Ri-treated LSK cells. We obtained a total of 60,648 cells expressing 15,049 genes. Clustering of the integrated UMAP identified 11 clusters (Fig. 6f) that were annotated as based on known marker gene expression and enrichment for several previously identified HSC and LSK signatures73,74,75 (Extended Data Fig. 6c–f and Supplementary Table 5). Compositional analysis of cell clusters showed the expected increase in the percentage of HSCs in all aged samples compared to the young ones (Extended Data Fig. 6g and Supplementary Table 5). It also revealed an increase in the percentage of MPP3 with aging and its decrease with the Ri treatment (Extended Data Fig. 6g and Supplementary Table 5), whereas there were no differences in HSC cell cycle profile after Ri treatment (Extended Data Fig. 6h and Supplementary Table 5). Interestingly, the scRNA-seq dataset showed that the increase of Klf4 expression in aged+Ri LSKs is more prominent in the HSC and MPP1 clusters (Fig. 6g). In detail, 43.58% of aged+Ri HSCs are expressing Klf4, whereas only 2.46% and 2.80% of young and aged HSCs, respectively, express this transcription factor. Differential gene expression analysis between the three conditions in the HSC cluster confirms the upregulation of several genes of the Klf family after Ri treatment (|log2FC| > 1 and FDR < 0.05; Extended Data Fig. 6i and Supplementary Table 5). To further analyze the activity of the different transcription factors and identify the network of regulated genes, we calculated transcription factor activity scores in each single HSC using SCENIC76 and generated a UMAP based on transcription factor activity scores (Fig. 6h). Interestingly, the percentage of aged+Ri HSCs with active KLF4 is 89.52%, whereas it is negligible in young and aged HSCs (1.89% and 0.69%, respectively) (Fig. 6i and Extended Data Fig. 6j). Differential analysis of the activity scores in between conditions confirms the increased activity of several KLF transcription factors as well as other transcription factors (|log2FC| > 0.5 and FDR < 0.05; Fig. 6j). Because KLF4 is known for its role in cell reprogramming and dedifferentiation, we wondered if aged+Ri HSCs show a more dedifferentiated transcriptional state. Interestingly, GSEA reveals an enrichment of the hemogenic precursor signature defined by Pereira et al.77 in the aged+Ri HSCs compared to the aged controls (Fig. 6k), supporting that Ri treatment might induce a partial transcriptional reprogramming of aged HSCs.
Because the scRNA-seq analysis indicates a partial reprogramming suggestive at the transcriptomic level of a possible functional improvement, we decided to assess the regenerative capacity of aged HSCs after Ri treatment in vivo. Previously, we correlated changes in cell epigenetic polarity of H4K16ac to function of HSCs24,78,79,80. Three-dimensional immunofluorescence staining of young, aged and aged+Ri HSCs clearly showed that Ri treatment restores H4K16ac polarity in aged stem cells (Extended Data Fig. 7a). Next, we tested in a non-competitive transplantation assay into young immunocompromised and KitW-41J mutant mice (NBSGW) the regenerative capacity of aged Ri-treated HSCs compared to young and aged control stem cells. We transplanted 200 aged HSCs treated overnight ex vivo with 100 µM Ri, alongside control recipient mice transplanted with solvent-treated young or aged HSCs (Fig. 7a). Donor mice were genetically tagged by constitutive expression in Rosa26 locus of a bright and stable red fluorescent protein (tdRFP)81, and we measured donor-derived contribution in peripheral blood by detecting RFP+ cells at several timepoints after transplantation. Notably, aged Ri-treated HSCs engrafted at the endpoint similarly to young HSCs, showing a significant increase in their peripheral blood regenerative capacity compared to aged control HSCs (Fig. 7b,c). Remarkably, Ri treatment also significantly increased the B cell lymphoid differentiation potential of aged HSCs and decreased the contribution to the myeloid lineage 18 weeks after transplantation (Fig. 7b–d). Ri treatment did not change the differentiation to the T cell compartment (Fig. 7b–d). Engraftment in BM and HSC compartments was not significantly different in Ri-treated recipients compared to either young or aged control recipients, showing a trend for both parameters to resemble young donor HSCs (Extended Data Fig. 7b,c). Overall, we conclude that inactivation of RhoA in aged HSCs functionally improves the regenerative capacity of old stem cells and their myeloid/B lymphoid skewing in vivo.
a, Experimental strategy for transplantation of young, aged and aged + Ri RFP+ HSCs into NBSGW mice. b, Percentage of engraftment in peripheral blood along the course of the transplantation at 6, 12 and 18 weeks. Four independent transplantation experiments were performed, and the initial number of recipient mice used per experiment was 3−4 per group. Young n = 9, aged n = 10, aged+Ri n = 10. Mann−Whitney test, unpaired, two-tailed, *P < 0.05, **P < 0.01, ***P < 0.001. c, Representative flow cytometry gating strategies for RFP+ cells, lymphoid cells (CD3+ and B220+) and myeloid cells (Gr1+, Gr1+Mac1+ and Mac1+) in peripheral blood at 18 weeks. d, Graphs showing the percentage of donor-derived B220+ cells, myeloid (Gr1+, Gr1+Mac1+ and Mac1+ cells) and CD3+ cells in peripheral blood at 6, 12 and 18 weeks after transplantation. Young n = 9, aged n = 10, aged+Ri n = 10. Graphs show mean ± s.e.m., Mann−Whitney test, unpaired, two-tailed, *P < 0.05, ***P < 0.001; only P < 0.05 (significant) values are shown. e, Cartoon scheme summarizing the main features rejuvenated by Ri treatment in aged HSCs. Panel a and e created with BioRender.
Discussion
The regenerative potential of HSCs declines upon aging, contributing to immunosenescence, inflammaging, clonal hematopoiesis and the increased incidence of hematopoietic disorders in older adults1,2,4. Previously, intrinsic epigenetic alterations have been associated with HSC aging, such as, for example, increased chromatin accessibility2,24,44,45,82. We also described that some of these epigenetic alterations depend on a reduction of LaminA/C expression in aged HSCs24,54, which is suggestive of a possible impairment of the mechanical properties of the nucleus. Supporting an emerging perspective that focuses on nuclear mechanoregulation9,11,37,83,84, in the present study we investigated the involvement of RhoA in nuclear mechanotrasnduction in HSCs, showing that RhoA is activated by increasing NE tension under cell confinement, by chromatin decompaction after NaB treatment, by nuclear swelling after hypotonic osmotic shock and after reduction of H3K9 methylation levels. We identify the loss of methylation of H3K9 as a likely cause not only of RhoA overactivation and increased NE tension in aged HSCs but also of many phenotypes associated with functional aging of HSCs. Interestingly, G9a activity is required for Ri restoration of H3K9me2 levels in aged HSCs. Further work is necessary to clarify mechanistically how RhoA activity downregulation affects the activity of this histone methyltransferase.
Differently from other cell types30,31, we show that, in HSCs, RhoA is not involved in transducing changes in extracellular stiffness, because RhoA is dispensable in the response to changes induced by culturing HSCs on hydrogel with high stiffness. However, because of the complexity of the mechanical BM microenvironment, we cannot exclude that RhoA might be activated by other extracellular signaling when HSCs are within their niche in vivo.
Furthermore, our results reveal that NE tension is physiologically increased upon HSC aging and that NE tension is necessary to activate RhoA in aged HSCs. RhoA activity in aged HSCs can be targeted by a specific small-molecule inhibitor33,85, which restores H3K9me2 levels and recovers the phenotypes of young HSC nuclei, such as DAPI intensity, DIR volume, number and localization. This is further supported by our machine learning approach, which demonstrates that DAPI imaging of morphometric and intensity features can be used to explain differences between HSCs and to inform on the age and fitness of the stem cells. To note, DAPI intensity and nuclear morphology features may vary with cell cycle state, whereas, in our experimental setting, HSCs were mainly in G0/early G1 with no differences across samples.
Notably, by ATAC−seq and RNA-seq, we detected a downregulation in chromatin accessibility and transcription at LTRs and LINE and a decrease in inflammation, immune response, interferon-responsive genes and aging signatures in aged Ri-treated HSCs (Extended Data Table 1). Moreover, after treatment of aged HSCs with Ri, we measured an increased transcription of Klf4, an opening of KLF4 binding motifs and an increased activity of KLF4 transcription factor (Extended Data Table 1). Genes targeted by KLF4 are enriched for pathways related to actin polymerization, and the increased levels of F-actin are likely acting to restrict nuclear stretching in aged Ri-treated HSCs (Extended Data Table 1). Moreover, scRNA-seq profiling revealed that KLF4 also targets genes enriched for a signature of hemogenic precursors, compatible with a partial reprogramming of aged HSCs, supporting the functional improvement in the regenerative capacity and myeloid/lymphoid skewing of aged+Ri stem cells in transplantation assays (Fig. 7e and Extended Data Table 1).
Thus far, several reports associated the aging process of different cell types with an epigenetic drift involving loss of heterochromatin and H3K9me and a disregulation of normally silenced and closed retrotransposons56,58,59,86,87. Here we report that an intrinsic nuclear mechanosignaling pathway dependent on RhoA can be pharmacologically targeted to revert these drifted epigenetic features, improving function of aged somatic stem cells (Fig. 7e). In the present study, the function of young, aged and aged+Ri HSCs was tested in a non-competitive transplantation assay, isolating sorted stem cells to be transplanted after in vitro treatment in young immunocompromised and KitW-41J mutant (NBSGW) recipient mice. Therefore, recipient mice were not challenged by preconditioning regimens that would disrupt the BM architecture and the cellular composition of the niche, likely exposing donor HSCs to a profoundly distinct mechanical environment88. However, in these settings, we did not perform secondary transplants with donor-derived HSCs, and further work will be required to evaluate the effect of RhoA inhibition on HSC function longer term. To note, whereas dominant negative RhoA was previously reported to increase HSC engraftment in mice89, overactivation of RhoA has been associated with functional impairment of human HSCs, further supporting possible translations of our findings90. Altogether, our data shed light on how intrinsic nuclear mechanotransduction controls aging-related epigenetic drift of somatic stem cells and uncover a potential target for improving tissue homeostasis over time.
Methods
Ethical compliance for mouse experiments
All mouse experiments were performed in compliance with ethical regulations according to the Spanish Law for Animal Protection and Welfare Code and were previously approved in project AR18008/10399 by the Biomedical Research Institute of Bellvitge Ethical Committee for Animal Experimentation (CEEA-IDIBELL) as well as by the Generalitat of Catalunya.
Reagents
A list of reagents, chemicals, commercial kits and antibodies is provided as Supplementary Table 6.
Mice
Young and aged C57BL/6 mice were obtained from the internal divisional stock (derived from mice obtained from The Jackson Laboratory (JAX)). Young and aged acRFP C57BL/6 mice were obtained from the internal divisional stock (originally kindly donated by Hans Jörg Fehling, Ulm University, and previously described81). The NBSGW mice were obtained from the internal divisional stock (derived from mice obtained from JAX, 026622) and were maintained as homozygotes. Rhoaflox mice were described previously13 and crossed with Cre-ERT2 mice (JAX, 008463). All mice were housed in the animal barrier facility under pathogen-free conditions at IDIBELL. Young mice discussed in this Article are 10−20 weeks old, and aged mice are at least 80 weeks old. Young mice were randomized for sex, and an equal number of male and female mice were used across samples. Aged mice were all females. Mice were not randomized for age, and operators were not blinded because aged mice require special care and attention. For the transplantation study, NBSGW mice were randomized for sex, and an equal number of male and female mice were used across samples. Mice that failed to recover from blood sampling and mice that died due to laboratory errors were excluded. Mice that needed to be euthanized because they were scored as ‘weak and about to die’ according to our approved animal license protocol for evaluating mouse health status remained part of the dataset. Allocation to control or treated group was done randomly (4−5 mice in each experimental group in four different experimental batches). All animals were maintained according to the recommendations of the European Convention for the Protection of Vertebrate Animals used for Experimental and other Scientific Purposes (ETS 123). Animals were housed in groups of up to four animals per cage in Makrolon Type II (long) cages with bedding and paper nesting material. The animals had access to food (ssniff, V1124-3) and water ad libitum. Animals were kept at a day/night rhythm of 12 hours/12 hours throughout the entire experiment.
Immunofluorescence staining and confocal images acquisition
Freshly sorted HSCs were seeded on fibronectin-coated glass coverslips. For polarity staining, HSCs were incubated for 12–16 hours in HBSS + 10% FBS + 1% penicillin−streptomycin and, when indicated, treated with 100 µM Ri33, 5 mM NaB, 20 µM cPLA2 inhibitor (AACOCF316), hypotonic medium, UNC0646 and arachidonic acid or left untreated. After incubation at 37 °C, 5% CO2, 3% O2 in growth factor-free medium, cells were fixed with BD Cytofix fixation buffer (BD Biosciences). After fixation, cells were gently washed with PBS, permeabilized with 0.2% Triton X-100 (Sigma-Aldrich) in PBS for 20 minutes and blocked with 10% donkey serum (Sigma-Aldrich) for 30 minutes. Primary and secondary antibody incubations were performed for 1 hour at room temperature. Cells were stained with a DAPI dilution 1:500 in PBS of DAPI 1 µg µl−1 (Thermo Fisher Scientific) for 10 minutes at room temperature and washed twice with PBS. Coverslips were mounted with ProLong Gold Antifade reagent without DAPI (Invitrogen, Molecular Probes). Samples were imaged with an Axio Observer Z1 microscope (Zeiss) equipped with a ×63 PH3 objective and an LSM 880 confocal microscope (Zeiss) equipped with a ×63 objective. z-stacks were obtained by automatically scanning along the z axis of the cell with a confocal microscope and acquiring a picture of the in-focus plane every 0.2−0.4 μm.
Immunofluorescence image analysis and rendering
Samples for immunofluorescence quantification analysis were rigorously sorted, stained and imaged in parallel within the same experiment to minimize any possible technical variability. Image acquisition at the confocal was carried out consistently in between experiments regarding laser power, z-stack size and gain master. Antibody specificities were validated using a knockout model in the case of RhoA-GTP or using a control sample only with the secondary antibody in the staining protocol for the rest of the stainings. Quantification of protein signals was done using Volocity software 7.0 (Quorum Technologies) using the ‘volume by intensity’ tool, which sets a threshold for the positive signal against the negative. Positive signal threshold was set for each experiment by using RhoAKO for RhoA-GTP and with the secondary antibody for the rest of the stainings. Morphometric measurements were done using Volocity software 6.5 or by our computational vision approach. Quantifications of DAPI volume were done with the ‘volume’ tool and nuclear height average (NHA), and diameter was done using the tool ‘line’. Immunofluorescence image 3D reconstruction and rendering were performed using Imaris 9.5.0 (Oxford Instruments) using the ‘surface’ tool, keeping the threshold constant for the signal in between experiments and conditions. As for polarity scoring, the localization of each single stained protein was considered polarized when a clear asymmetric distribution was visible by drawing a line across the middle of the cell. A total of 50−100 HSCs were singularly analyzed per sample. Data are plotted as percentage of the total number of cells scored per sample.
3D nuclear DAPI image preprocessing
HSC nucleus microscopy images were exported as Carl Zeiss CZI files for downstream analysis and processed using the Python programming language. Images acquired with a different microscope than the ones specified above, and images belonging to experiments in which more than 30 days passed between nucleus staining and image data acquisition were excluded from these analyses. As a first quality control, images displaying total pixel intensities higher than 4 × 108 were labeled as overexposed to DAPI and discarded. Similarly, images with increased Gaussian noise (estimated noise s.d. >8) were labeled as noisy and also excluded from further analyses. The remaining images were then corrected using Chambolle’s total variation denoising method91. Because image acquisition measurements were dynamically adjusted to the size of the nucleus, each 3D image was resized to achieve a uniform resolution of 10 pixels per micrometer in all dimensions through isotropic interpolation that accounted for variations in the number of z-stacks obtained. For each image, a nucleus binary mask was extracted using the Otsu segmentation method92 after smoothing with a Gaussian filter and allowing for hysteresis to preserve the nuclear border with higher confidence. Potential holes in the binary mask due to low-intensity areas within the nucleus were filled and marked as part of the segmentation. Both the intensity image and the respective nucleus mask were centered in the container array grid by trimming the background and padding the image borders symmetrically. Marginal intensities outside the nucleus mask were removed. DAPI intake within the nucleus is affected by both biological and technical factors. To mitigate batch effects related to technical variation in fluorescence signal, we standardized the pixel intensity distribution within each nucleus mask to yield 0 mean and 1 s.d. using z-score normalization. This approach removes technical artifacts and ensures that downstream analyses reflect biological rather than acquisition-related variability. We validated our nuclear segmentation pipeline by manually annotating 426 slices from nine nuclei (three per condition) using Napari, excluding the top and bottom 20% of z-slices. Automated Otsu-based segmentations showed high agreement with manual masks (Dice: 0.96 ± 0.02; Precision: 0.99 ± 0.01; Recall: 0.93 ± 0.03). Full details and representative overlays are provided in the supplementary methods.
Intensity by distance to nuclear border analyses
Intensity by distance analyses were performed at iso-distance intervals of 0.1 µm from the boundary of the nucleus segmentation, using a distance-transformed mask. A distance-transformed mask is a mask in which each pixel value represents the shortest geodesic distance to the nearest mask boundary. The average intensity value of all pixels within a 3D band with a thickness of 0.15 µm was reported at each measurement interval. Young, aged and aged+Ri conditions were measured up to a distance of 1.6 µm from the nuclear border. Box plots of DAPI intensity by discrete distance ranges were computed, taking into account the average intensity of all pixels within the specified distance boundaries for each interval.
Multivariate feature analyses
Most morphometric and intensity features were measured with the scikit-image Python library93 regionprops_table() function at the nuclear mask level, at the DIR level and on the largest 2D slide in the x−y plane from each 3D image, comprising a total of 39 features (Supplementary Table 2). The width, length and height of each nucleus were obtained from its bounding box. Height deviation was calculated as the average standard deviation of height in the x dimension for each y−z slide. The aspect ratio was defined as the ratio of height to length. The surface area was calculated using the Marching Cubes algorithm after smoothing the nucleus mask with a Gaussian filter. The intensity ratio was computed as the ratio of average intensity within the 1−1.5-µm distance interval to the 0−0.5-µm distance interval from the nuclear border. The EOP was computed as the proportion of the difference of the nuclear perimeter compared to the perimeter of an ellipse with the same major and minor axis length as the nucleus mask. Detailed information about each of these features is shown in Supplementary Table 2.
DIRs were defined based on intensity, not spatial pattern. Specifically, DIRs correspond to the top 20% of standardized DAPI intensity values across all 3D young nuclei, yielding a fixed threshold of 0.6 after z-score normalization. This approach identifies DIRs as high-intensity subregions within each nucleus. More information about the definition of DIRs, including visualizations for various nuclei and conditions, is included in the supplementary methods.
DIRs were segmented using the watershed method on the thresholder images. The intensity standardization of the images allowed for the choice of an absolute thresholding value for all nuclei. We set this parameter to the quantile 80% of the intensity distribution for all young nuclei. Individual DIRs were labeled and segmented with watershed by identifying the intensity peaks as the local maxima in the Euclidean distance transform of the thresholder images. From the total of segmented DIRs, we filtered out those displaying a voxel volume smaller than 0.2 µm. Morphometric and intensity features were measured on the resulting DIR segmentation mask in the same manner as with the nuclear mask. DIR distance to border was computed as the average of the distances for each voxel within each DIR to the nuclear mask border. After the calculation of these measurements, a second quality control was conducted to eliminate images that produced artifacts in the nuclear and DIR segmentation. The images were filtered out if they produced nuclear masks with a voxel volume smaller than 40 µm3, an EOP larger than 0.25 or a DIR volume larger than 5 µm3, which mostly belonged to confined nuclei that were damaged during the experimental setup and failed nuclear mask segmentations. We proceeded with 177 young nuclei, 164 aged nuclei and 144 aged+Ri nuclei.
Feature selection was conducted to reduce dimensionality while preserving biological interpretability. We started from an initial set of 39 features derived from 3D nuclear masks, DIRs and 2D projections (Supplementary Table 1). These included 17 whole-nucleus features, 15 DIR-level features and seven from the largest 2D z-slice.
Feature selection was conducted in two steps. First, we identified features that significantly differed between biological conditions (young versus aged and aged versus aged+Ri) using a two-sided Mann−Whitney U-test (P < 0.05). This yielded 23 features distinguishing young from aged and 18 distinguishing aged from aged+Ri. The union of these (n = 29) was retained.
To remove redundancy, we applied pairwise Pearsonʼs correlation filtering, discarding features with |r| > 0.8. In cases of redundancy, the retained feature was selected based on domain relevance. The resulting non-redundant feature set (n = 20) included nine whole-nucleus features, eight DIR features and 3D from 2D projections. These were used as input for UMAP embedding and clustering (Extended Data Fig. 3e). The final set shows low overall inter-feature correlation (mean |r| = 0.22) and captures the morphometric and intensity-based variation across conditions. These features were standardized and used to train the UMAP42 model on young, aged and aged+Ri-treated nuclei. Clustering was conducted in the original multidimensional parameter space using the k-means algorithm94 with k = 4 and depicted in the UMAP embedding. This value of k yielded the best silhouette scores for all clusters (Extended Data Fig. 3f). The resulting clusters revealed biologically relevant groups combining images from the young, aged and aged+Ri conditions. Volcano plots showing feature importance for each of the identified clusters were generated by calculating the statistical significance (Mann−Whitney U-test) and the log2FC of the average of each feature between the clusters and the average of the remaining clusters. Clustering quality and stability were evaluated using multiple validation metrics and subsampling analysis, as described in detail in the supplementary methods.
Confinement assay
The confinement device used in this protocol was adapted to a single-well plate from a previously described method11,95. In brief, the confinement device was made by a magnetic container, two metallic rings, a polymeric ring and a closing ring (Extended Data Fig. 1a). The compression is mediated by a pillar coverslip, a polymeric piston and the magnetic lid of the device that exerts the confinement pressure. In total, 3,000 HSCs were seeded in a volume of 40 µl of HBSS + 10% FBS + 1% penicillin−streptomycin onto a 35-mm glass coverslip. The coverslip was previously functionalized with fibronectin: 40 µl of fibronectin (50 µg ml−1) was applied upon the surface of the coverslip for 2 hours at room temperature, blocked with the same volume of BSA 2% for 1 hour at 37 °C and washed with PBS Ca2+/Mg2+. The coverslip with cells was mounted in the confinement device and compressed with pillars of 3 µm, 5 µm and 8 µm for 2 hours at 37 °C in a hypoxic incubator (5% CO2, 3% O2). Cells were then fixed directly in the confinement device with 1 ml of paraformaldehyde 4.21% (BD Cytofix, Thermo Fisher Scientific, 15817828) at 4 °C for 15 minutes. The sample on the coverslip was then removed from the confiner and stocked at 4 °C in PBS for a maximum of 2 weeks before staining.
Nuclear wrinkling analysis
To analyze nuclear wrinkling in LaminB-stained HSCs, we developed an automatic image analysis pipeline in MATLAB (MathWorks, R2024b). Fluorescence images of LaminB were acquired in an LSM 880 confocal microscope (Zeiss), as mentioned above in ‘Immunofluorescence staining and confocal images acquisition’. For each dataset, the equatorial maximum cross-section section was identified from z-slices, and three slices were selected above and below, covering approximately 50% of the nucleus surface with optimal resolution of surface wrinkles in the lateral x−y dimension. Selected image slices were then subjected to binning (factor n = 3) using a bilinear interpolation to reduce noise and facilitate subsequent processing steps, effectively preserving key structural information. A threshold intensity of 25 counts was applied to create a binary mask and to segment the regions of interest representing the nuclear envelope. A Gaussian blur with a sigma of 3 pixels was applied to the binary mask to smooth the segmented regions and facilitate the subsequent skeletonization of NE features based on a morphological thinning algorithm. The skeletonization process enabled us to extract the structural features of the segmented Lamin signal, highlighting the wrinkles of the NE. To remove structural artifacts outside the nuclear region of interest, the skeletonized image was filled to generate an inner region (representing the inside of the nuclear wrinkles) and an outer region (around the nucleus periphery) by applying an inverted mask to the filled skeleton. This allowed us to separate the NE from the surrounding structures and detect the points representing nuclear invaginations. The coordinates of the nuclear surface and inner invaginations represent the total nuclear periphery and were extracted for further quantitative metrics of nuclear wrinkles. We derived the nuclear circularity (C = 4πA / P2) from the total perimeter P and area A of nuclear cross-sections from multiple z-slices for each cell. The circularity provides a quantitative measure of NE deformations, with values close to 1 indicating a more circular shape and values <1 indicating a highly deformed and wrinkled nuclear surface. In addition, we derived the excess folding parameter (E = 1 − p / P) as the ratio of the boundary outline of the nucleus p to the total perimeter P of nuclear cross-sections, with values close to 0 resembling a circular shape and values closer to 1 indicating increased nuclear wrinkling. Statistical data analysis was performed in GraphPad Prism (version 10.2.3.) using a Kruskal−Wallis test for multiple comparisons.
Statistics and reproducibility
Statistical data analysis derived from protein quantification from both image data and western blot and transplantation analysis were performed using GraphPad Prism version 10.2.3. For the statistics all over the dataset, the procedure was the following: outliers were removed, and data distribution was tested for normality before choosing the appropriate statistical test. The statistical test applied is indicated in the corresponding figure legend. An exception was done in the case of H3K9me2 immunofluorescence signal quantification in Fig. 4a, in which we did not remove outliers to preserve the high biological variability for this marker in HSCs, in agreement with previous reports24,48. As for experiments involving mice, sample size was determined according to the standard deviation of the measured parameter. We used StatMate and the statistical guide of Prism to help with sample size calculation. No method of randomization was used to assign mice to experimental groups, but groups were balanced by sex, except for aged mice that were all females. For in vitro experiments, no statistical methods were used to predetermine sample size, but our sample sizes are similar to those reported in previous publications11,24,53,96. Outliers were excluded as based on the ROUT method, aggressiveness Q = 1% (recommended by Prism). In Fig. 7c,d, data were excluded when engraftment did not reach 0.1% of peripheral blood, when the mouse died before the endpoint and in case the value was marked as an outlier by Prism.
Owing to space constraints, additional information on methods is provided in the supplementary information file.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The source data underlying Figs. 1, 2, 4, 6e and 7 and Extended Data Figs. 1, 2, 4, 6a and 7 are provided as a Source Data file. The DAPI-stained raw and preprocessed microscopy images used for the analysis shown in Fig. 3 and Extended Data Fig. 3 are provided in the following repository: https://doi.org/10.34810/data697 (ref. 97). Raw sequencing data are deposited at the Gene Expression Omnibus (accession number GSE233989). Dilutions and catalog numbers of all commercial antibodies are provided in Supplementary Table 6. All other data supporting the findings of this study are available from the corresponding author upon reasonable request.
Code availability
The source code for the DAPI-stained microscopy image analyses shown in Fig. 3 and Extended Data Fig. 3 is available at https://github.com/biomedical-data-science/hsc_rhoa. The code and main results of the ATAC−seq, RNA-seq and scRNA-seq data analysis underlining Figs. 5 and 6, Extended Data Figs. 5 and 6 and Supplementary Tables 2–5 are deposited in the following repository: https://doi.org/10.34810/data697. The code for the measurement of nuclear wrinkling will be made available upon reasonable request.
References
Brunet, A., Goodell, M. A. & Rando, T. A. Ageing and rejuvenation of tissue stem cells and their niches. Nat. Rev. Mol. Cell Biol. 24, 45–62 (2022).
Mejia-Ramirez, E. & Florian, M. C. Understanding intrinsic hematopoietic stem cell aging. Haematologica 105, 22–37 (2020).
Matteini, F., Montserrat-Vazquez, S. & Florian, M. C. Rejuvenating aged stem cells: therapeutic strategies to extend health and lifespan. FEBS Lett. 598, 2776–2787 (2024).
Andersson, R., Mejia-Ramirez, E. & Florian, M. C. Haematopoietic ageing in health and lifespan. Nat. Cell Biol. 27, 1398–1410 (2025).
Starodubtseva, M. N. Mechanical properties of cells and ageing. Ageing Res. Rev. 10, 16–25 (2011).
Phillip, J. M., Aifuwa, I., Walston, J. & Wirtz, D. The mechanobiology of aging. Annu. Rev. Biomed. Eng. 17, 113–141 (2015).
López-Otín, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. Hallmarks of aging: an expanding universe. Cell 186, 243–278 (2023).
Lessey, E. C., Guilluy, C. & Burridge, K. From mechanical force to RhoA activation. Biochemistry 51, 7420–7432 (2012).
Burridge, K., Monaghan-Benson, E. & Graham, D. M. Mechanotransduction: from the cell surface to the nucleus via RhoA. Philos. Trans. R. Soc. B 374, 20180229 (2019).
Goldyn, A. M., Rioja, B. A., Spatz, J. P., Ballestrem, C. & Kemkemer, R. Force-induced cell polarisation is linked to RhoA-driven microtubule-independent focal-adhesion sliding. J. Cell Sci. 122, 3644–3651 (2009).
Venturini, V. et al. The nucleus measures shape changes for cellular proprioception to control dynamic cell behavior. Science 370, eaba2644 (2020).
Mistriotis, P. et al. Confinement hinders motility by inducing RhoA-mediated nuclear influx, volume expansion, and blebbing. J. Cell Biol. 218, 4093–4111 (2019).
Zhou, X. et al. RhoA GTPase controls cytokinesis and programmed necrosis of hematopoietic progenitors. J. Exp. Med. 210, 2371–2385 (2013).
Enyedi, B. & Niethammer, P. Nuclear membrane stretch and its role in mechanotransduction. Nucleus 8, 156–161 (2017).
Enyedi, B., Jelcic, M. & Niethammer, P. The cell nucleus serves as a mechanotransducer of tissue damage-induced inflammation. Cell 165, 1160–1170 (2016).
Lomakin, A. J. et al. The nucleus acts as a ruler tailoring cell responses to spatial constraints. Science 370, eaba2894 (2020).
Le Berre, M., Aubertin, J. & Piel, M. Fine control of nuclear confinement identifies a threshold deformation leading to lamina rupture and induction of specific genes. Integr. Biol. 4, 1406 (2012).
Nielsen, L. K., Risbo, J., Callisen, T. H. & Bjørnholm, T. Lag-burst kinetics in phospholipase A2 hydrolysis of DPPC bilayers visualized by atomic force microscopy. Biochim. Biophys. Acta 1420, 266–271 (1999).
Leidy, C. et al. Membrane restructuring by phospholipase A2 is regulated by the presence of lipid domains. Biophys. J. 101, 90–99 (2011).
Garcia, M. C., Williams, J., Johnson, K., Olden, K. & Roberts, J. D. Arachidonic acid stimulates formation of a novel complex containing nucleolin and RhoA. FEBS Lett. 585, 618–622 (2011).
Garcia, M. C. et al. Arachidonic acid stimulates cell adhesion through a novel p38 MAPK-RhoA signaling pathway that involves heat shock protein 27. J. Biol. Chem. 284, 20936–20945 (2009).
Chen, T. et al. Histone acetylation is involved in hsp70 gene transcription regulation in Drosophila melanogaster. Arch. Biochem. Biophys. 408, 171–176 (2002).
Kulka, L. A. M., Fangmann, P.-V., Panfilova, D. & Olzscha, H. Impact of HDAC inhibitors on protein quality control systems: consequences for precision medicine in malignant disease. Front. Cell Dev. Biol. 8, 425 (2020).
Grigoryan, A. et al. LaminA/C regulates epigenetic and chromatin architecture changes upon aging of hematopoietic stem cells. Genome Biol. 19, 189 (2018).
Roffay, C. et al. Passive coupling of membrane tension and cell volume during active response of cells to osmosis. Proc. Natl Acad. Sci. USA 118, e2103228118 (2021).
Piccolo, S., Dupont, S. & Cordenonsi, M. The biology of YAP/TAZ: Hippo signaling and beyond. Physiol. Rev. 94, 1287–1312 (2014).
Dupont, S. et al. Role of YAP/TAZ in mechanotransduction. Nature 474, 179–183 (2011).
Chen, X. et al. Mechanical heterogeneity in the bone microenvironment as characterized by atomic force microscopy. Biophys. J. 119, 502–513 (2020).
Zhang, P. et al. The physical microenvironment of hematopoietic stem cells and its emerging roles in engineering applications. Stem Cell Res. Ther. 10, 327 (2019).
Ingallina, E. et al. Mechanical cues control mutant p53 stability through a mevalonate−RhoA axis. Nat. Cell Biol. 20, 28–35 (2018).
Koester, J. et al. Niche stiffening compromises hair follicle stem cell potential during ageing by reducing bivalent promoter accessibility. Nat. Cell Biol. 23, 771–781 (2021).
Zhang, X. et al. Harnessing matrix stiffness to engineer a bone marrow niche for hematopoietic stem cell rejuvenation. Cell Stem Cell 30, 378–395 (2023).
Shang, X. et al. Rational design of small molecule inhibitors targeting RhoA subfamily Rho GTPases. Chem. Biol. 19, 699–710 (2012).
Al Jord, A. et al. Cytoplasmic forces functionally reorganize nuclear condensates in oocytes. Nat. Commun. 13, 5070 (2022).
Jackson, J. A. et al. Scaling behaviour and control of nuclear wrinkling. Nat. Phys. 19, 1927–1935 (2023).
Totaro, A., Panciera, T. & Piccolo, S. YAP/TAZ upstream signals and downstream responses. Nat. Cell Biol. 20, 888–899 (2018).
Kalukula, Y., Stephens, A. D., Lammerding, J. & Gabriele, S. Mechanics and functional consequences of nuclear deformations. Nat. Rev. Mol. Cell Biol. 23, 583–602 (2022).
Kim, K. M. et al. Taz protects hematopoietic stem cells from an aging-dependent decrease in PU.1 activity. Nat. Commun. 13, 5187 (2022).
Mascetti, G., Carrara, S. & Vergani, L. Relationship between chromatin compactness and dye uptake for in situ chromatin stained with DAPI. Cytometry 44, 113–119 (2001).
Linhoff, M. W., Garg, S. K. & Mandel, G. A high-resolution imaging approach to investigate chromatin architecture in complex tissues. Cell 163, 246–255 (2015).
Long, J. T. & Lammerding, J. Nuclear deformation lets cells gauge their physical confinement. Dev. Cell 56, 156–158 (2021).
McInnes, L., Healy, J., Saul, N. & Großberger, L. UMAP: uniform manifold approximation and projection. J. Open Source Softw. 3, 861 (2018).
Goldman, R. D. et al. Accumulation of mutant lamin A causes progressive changes in nuclear architecture in Hutchinson–Gilford progeria syndrome. Proc. Natl Acad. Sci. USA 101, 8963–8968 (2004).
Chambers, S. M. et al. Aging hematopoietic stem cells decline in function and exhibit epigenetic dysregulation. PLoS Biol. 5, e201 (2007).
Sun, D., et al. Epigenomic profiling of young and aged HSCs reveals concerted changes during aging that reinforce self-renewal. Cell Stem Cell 14, 673–688 (2014).
Pal, S. & Tyler, J. K. Epigenetics and aging. Sci. Adv. 2, e1600584 (2016).
Tsurumi, A. & Li, W. X. Global heterochromatin loss: a unifying theory of aging? Epigenetics 7, 680–688 (2012).
Djeghloul, D. et al. Age-associated decrease of the histone methyltransferase SUV39H1 in HSC perturbs heterochromatin and B lymphoid differentiation. Stem Cell Rep. 6, 970–984 (2016).
Liu, F. et al. Optimization of cellular activity of G9a inhibitors 7-aminoalkoxy-quinazolines. J. Med. Chem. 54, 6139–6150 (2011).
Padeken, J., Methot, S. P. & Gasser, S. M. Establishment of H3K9-methylated heterochromatin and its functions in tissue differentiation and maintenance. Nat. Rev. Mol. Cell Biol. 23, 623–640 (2022).
Kirby, T. J. & Lammerding, J. Emerging views of the nucleus as a cellular mechanosensor. Nat. Cell Biol. 20, 373–381 (2018).
Zhang, D., Zhang, R., Song, X., Yan, K. C. & Liang, H. Uniaxial cyclic stretching promotes chromatin accessibility of gene loci associated with mesenchymal stem cells morphogenesis and osteogenesis. Front. Cell Dev. Biol. 9, 664545 (2021).
Nava, M. M. et al. Heterochromatin-driven nuclear softening protects the genome against mechanical stress-induced damage. Cell 181, 800–817 (2020).
Grigoryan, A. et al. Attrition of X chromosome inactivation in aged hematopoietic stem cells. Stem Cell Rep. 16, 708–716 (2021).
Itokawa, N. et al. Epigenetic traits inscribed in chromatin accessibility in aged hematopoietic stem cells. Nat. Commun. 13, 2691 (2022).
Gorbunova, V. et al. The role of retrotransposable elements in aging and age-associated diseases. Nature 596, 43–53 (2021).
Lara-Astiaso, D. et al. Immunogenetics. Chromatin state dynamics during blood formation. Science 345, 943–949 (2014).
De Cecco, M. et al. L1 drives IFN in senescent cells and promotes age-associated inflammation. Nature 566, 73–78 (2019).
Della Valle, F. et al. LINE-1 RNA causes heterochromatin erosion and is a target for amelioration of senescent phenotypes in progeroid syndromes. Sci. Transl. Med. 14, eabl6057 (2022).
Hidaoui, D. et al. Targeting heterochromatin eliminates chronic myelomonocytic leukemia malignant stem cells through reactivation of retroelements and immune pathways. Commun. Biol. 7, 1555 (2024).
Clapes, T. et al. Chemotherapy-induced transposable elements activate MDA5 to enhance haematopoietic regeneration. Nat. Cell Biol. 23, 704–717 (2021).
Rusinova, I. et al. INTERFEROME v2.0: an updated database of annotated interferon-regulated genes. Nucleic Acids Res. 41, D1040–D1046 (2013).
Bouman, B. J. et al. Single-cell time series analysis reveals the dynamics of HSPC response to inflammation. Life Sci. Alliance 7, e202302309 (2024).
Flohr Svendsen, A. et al. A comprehensive transcriptome signature of murine hematopoietic stem cell aging. Blood 138, 439–451 (2021).
Edginton-White, B. et al. A genome-wide relay of signalling-responsive enhancers drives hematopoietic specification. Nat. Commun. 14, 267 (2023).
Di Giammartino, D. C. et al. KLF4 is involved in the organization and regulation of pluripotency-associated three-dimensional enhancer networks. Nat. Cell Biol. 21, 1179–1190 (2019).
Park, C. S., Shen, Y., Lewis, A. & Lacorazza, H. D. Role of the reprogramming factor KLF4 in blood formation. J. Leukoc. Biol. 99, 673–685 (2016).
Xie, L. et al. A dynamic interplay of enhancer elements regulates Klf4 expression in naïve pluripotency. Genes Dev. 31, 1795–1808 (2017).
Roisman, A. et al. Loss of KLF6 recapitulates molecular and functional changes associated with aging in human hematopoietic stem and progenitor cells. Blood 134, 447 (2019).
Adelman, E. R. et al. Aging human hematopoietic stem cells manifest profound epigenetic reprogramming of enhancers that may predispose to leukemia. Cancer Discov. 9, 1080–1101 (2019).
McGinn, J. et al. A biomechanical switch regulates the transition towards homeostasis in oesophageal epithelium. Nat. Cell Biol. 23, 511–525 (2021).
Mas, G. et al. In vivo temporal resolution of acute promyelocytic leukemia progression reveals a role of Klf4 in suppressing early leukemic transformation. Genes Dev. 36, 451–467 (2022).
Wilson, N. K. et al. Combined single-cell functional and gene expression analysis resolves heterogeneity within stem cell populations. Cell Stem Cell 16, 712–724 (2015).
Rodriguez-Fraticelli, A. E. et al. Single-cell lineage tracing unveils a role for TCF15 in haematopoiesis. Nature 583, 585–589 (2020).
Cabezas-Wallscheid, N. et al. Vitamin A-retinoic acid signaling regulates hematopoietic stem cell dormancy. Cell 169, 807–823 (2017).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Pereira, C.-F. et al. Hematopoietic reprogramming in vitro informs in vivo identification of hemogenic precursors to definitive hematopoietic stem cells. Dev. Cell 36, 525–539 (2016).
Florian, M. C. et al. Cdc42 activity regulates hematopoietic stem cell aging and rejuvenation. Cell Stem Cell 10, 520–530 (2012).
Florian, M. C. et al. Aging alters the epigenetic asymmetry of HSC division. PLoS Biol. 16, e2003389 (2018).
Montserrat-Vazquez, S. et al. Transplanting rejuvenated blood stem cells extends lifespan of aged immunocompromised mice. npj Regen. Med. 7, 78 (2022).
Luche, H., Weber, O., Nageswara Rao, T., Blum, C. & Fehling, H. J. Faithful activation of an extra-bright red fluorescent protein in ‘knock-in’ Cre-reporter mice ideally suited for lineage tracing studies. Eur. J. Immunol. 37, 43–53 (2007).
Chambers, S. M. & Goodell, M. A. Hematopoietic stem cell aging: wrinkles in stem cell potential. Stem Cell Rev. 3, 201–211 (2007).
Maurer, M. & Lammerding, J. The driving force: nuclear mechanotransduction in cellular function, fate, and disease. Annu. Rev. Biomed. Eng. 21, 443–468 (2019).
Xia, Y., Pfeifer, C. R., Cho, S., Discher, D. E. & Irianto, J. Nuclear mechanosensing. Emerg. Top. Life Sci. 2, 713–725 (2018).
Hegde, S. et al. Inhibition of RHOA activity preserves the survival and hemostasis function of long-term cold-stored platelets. Blood 144, 1732–1746 (2024).
De Cecco, M. et al. Transposable elements become active and mobile in the genomes of aging mammalian somatic tissues. Aging 5, 867–883 (2013).
Wang, Y. et al. L1 drives HSC aging and affects prognosis of chronic myelomonocytic leukemia. Signal Transduct. Target. Ther. 5, 205 (2020).
Horton, P. D., Dumbali, S. & Wenzel, P. L. Mechanoregulation in hematopoiesis and hematologic disorders. Curr. Stem Cell Rep. 6, 86–95 (2020).
Ghiaur, G. et al. Inhibition of RhoA GTPase activity enhances hematopoietic stem and progenitor cell proliferation and engraftment. Blood 108, 2087–2094 (2006).
Jaganathan, B. G., Anjos-Afonso, F., Kumar, A. & Bonnet, D. Active RHOA favors retention of human hematopoietic stem/progenitor cells in their niche. J. Biomed. Sci. 20, 66 (2013).
Chambolle, A. An algorithm for total variation minimization and applications. J. Math. Imaging Vis. 20, 89–97 (2004).
Otsu, N. A threshold selection method from gray-level histograms. In IEEE Transactions on Systems, Man, and Cybernetics 62–66 (IEEE, 1979); https://doi.org/10.1109/TSMC.1979.4310076
Walt, S. vander et al. scikit-image: image processing in Python. PeerJ 2, e453 (2014).
Hartigan, J. A. & Wong, M. A. Algorithm AS 136: a k-means clustering algorithm. J. R. Stat. Soc. Ser. C 28, 100–108 (1979).
Le Berre, M., Zlotek-Zlotkiewicz, E., Bonazzi, D., Lautenschlaeger, F. & Piel, M. Methods for two-dimensional cell confinement. Methods Cell. Biol. 121, 213–229 (2014).
Sacma, M. et al. Haematopoietic stem cells in perisinusoidal niches are protected from ageing. Nat. Cell Biol. 21, 1309–1320 (2019).
Mejia-Ramirez, E. V. A. et al. Replication data for ‘Targeting RhoA activity rejuvenates aged hematopoietic stem cells’. Repositori de Dades de Recerca https://doi.org/10.34810/data697 (2025).
Acknowledgements
We thank support from L. Traveset Martinez, IDIBELL Innovation Unit. We acknowledge support from M. M. Gaudes, head of Technical Facilities at IDIBELL, together with J. A. Vaquero (IDIBELL FACS and Flow Cytometry SCT), A. Ventura (IDIBELL Mouse Facility SCT) and J. Repulles and S. Mendizuri (IDIBELL Bioimaging SCT). We also thank E. Castaño, B. Barroso and B. Torrejon (CCiT-UB, Bellvitge). We thank G. Lunazzi and the National Center for Genomic Analysis (CNAG, Barcelona) for support with scRNA-seq experiments. We thank the CERCA Program/Generalitat de Catalunya for institutional support. We thank C. Lazaro (LCAM laboratory, ICO-HUB) for supporting sequencing experiments. We acknowledge the following funding sources: European Research Council grant 101002453 (M.C.F.); Spanish Ministry of Science, Innovation and University grants RYC2018-025979-I (M.C.F.) and PGC2018-102049-B-I00 (M.C.F.) and PID2021-123922NB-I00 (M.C.F.); Department of Research and University of the Generalitat of Catalunya and AGAUR (expedient 2021 SGR 00888) (M.C.F.). P.I.P received an INPhINIT Incoming fellowship from ‘la Caixa’ Foundation (ID 100010434) with code 2022-4167. P.P. and P.I.P. acknowledge support from the grant CEX2023-0001290-S funded by MCIN/AEI/ 10.13039/501100011033, and support from the Generalitat de Catalunya through the CERCA Program. P.P acknowledges her AI4S fellowship within the “Generación D” initiative by Red.es, Ministerio para la Transformación Digital y de la Función Pública, for talent attraction (C005/24-ED CV1), funded by NextGenerationEU through PRTR. V.R. acknowledges financial support from the Ministerio de Ciencia e Innovación through the Plan Nacional (PID2020-117011GB-I00) and funding from the European Union’s Horizon EIC-ESMEA Pathfinder program under grant agreement number 101046620.
Author information
Authors and Affiliations
Contributions
Conceptualization: E.M.-R., M.C.F., P.I.P. and B.W. Methodology: S.M.-V., E.M.-R., M.C.F., P.I.P., F.P., F.A., J.C.R., B.W., F.M., D.M., L.R. and S.W. Investigation: S.M.-V., E.M.-R., M.C.F., P.I.P. and B.W. Visualization: S.M.-V., E.M.-R., P.I.P. and L.M. Funding acquisition: P.P., V.R. and M.C.F. Project administration: M.C.F. Supervision: Y.Z., V.R., A.R., P.P. and M.C.F. Writing—original draft: E.M.-R., S.M.-V., M.C.F. and P.I.P. Writing—review and editing: A.R., V.R., Y.Z., P.P. and M.C.F.
Corresponding authors
Ethics declarations
Competing interests
The findings presented in this study are covered under patent application number EP25382180.5, filed on 27 February 2025 (F.M.C. and Y.Z.). Y.Z. and F.M.C. declare no additional financial or non-financial competing interests. The remaining authors declare no financial or non-financial competing interests.
Peer review
Peer review information
Nature Aging thanks Pamela Wenzel, Haitham Shaban and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Increasing nuclear envelope tension but not changes in substrate stiffness activates RhoA in hematopoietic stem cells.
A. Confinement device scheme explaining the mounting strategy and how the cells are confined. B. Gating strategy for the isolation of LT-HSC from young BM cells from a representative experiment. C. Stripplot showing the increase of the excess of perimeter (EOP) upon confinement measured in 2D. D. Stripplot showing an increase in major nuclear axis length upon confinement at different heights. E, DAPI intensity profiles of representative confocal images from HSC under different levels of confinement as a function of the distance to the nuclear border. F, RhoA locus genetic maps of RhoaWT and Rhoafl mice and schematic result of in vitro Cre tamoxifen-induced deletion of Rhoa exon3. Black arrowhead stands for the flox sequence location at the Rhoa locus. PCR results are shown for the Rhoa floxed (RAp1&RAp2) and RhoaKO (RAp1&RAp3) pair of primers performed on BM cells genomic DNA from Cre+/− Rhoawt/wt and Cre+/−Rhoafl/fl mice after in vitro tamoxifen treatment and control. Sizes of expected PCR products are shown in each case. G, Representative confocal images, and their respective 3D reconstruction in LT-HSC after o/n treatment with tamoxifen for induction of RhoaKO. Cells were stained against RhoAGTP (red) and DAPI (gray). Graph shows the volume for RhoAGTP signal in WT and induced RhoaKO without confinement. Mann-Whitney, unpaired two-tailed is shown *p < 0.05. H. Representative image of NE PcPLA2 (green) in 5 µm confined HSC. Graph comparing levels of PcPLA2 at the NE in unconfined and 5 µm confined cells. Mann-Whitney, unpaired two-tailed is shown ****p < 0.0001, n = 3. I, Representative image of RhoA-GTP (red) in young HSC cultured with or without AA. DAPI is shown in grey. The graphs compare levels of RhoA-GTP in young HSCs ± AA. Mann-Whitney, unpaired two-tailed is shown *p < 0.05. n = 3. J, Representative image of NE PcPLA2 (green) and RhoA-GTP (red) in young HSC cultured in isotonic and 0.75X hypotonic conditions. DAPI is shown in grey. The graphs compare levels of NE PcPLA2 and RhoA-GTP in isotonic and hypotonic samples. Mann-Whitney, unpaired two-tailed is shown **p < 0.01, *p < 0.05. n = 3. K, Schematic representation of the Chiaro NanoIndenter and design of bone measurement experiment. L, Graph showing the stiffness of the endosteum and inner marrow within femurs from young mice. A stiffness variation graph of a representative single femur is shown. Mann-Whitney, unpaired two-tailed ****p < 0.0001. M, Schematic representation of the experimental design for the 2D culturing of LT-HSC on different stiffness hydrogels and later analysis by immunofluorescence (IF) and methocult culturing. Created in BioRender. Mejía-Ramírez, E. (2025) https://BioRender.com/0w6komx. N, Graph representing the number of cells obtained after 10 days on methocult from RhoAwt and RhoaKO LT-HSC cultured in different stiffness hydrogels. Mann-Whitney, unpaired two-tailed.
Extended Data Fig. 2 NE tension and nuclear translocation of TAZ are increased in aged HSCs.
A, Superposed stiffness variation graphs showing the measurements of representative femurs on a young and an aged mouse. B, Stiffness of the BM of femurs of young and aged mice at the endosteum and inner marrow. n = 3, with a total of 69-80 measurements per condition, Mann-Whitney, two-tailed test *p < 0.05; ****p < 0.0001. C, Representative western blot of RhoAGTP pulled down from BM cells from young, aged and aged treated in vitro with Ri. Antibody against RhoA was used to detect pulled down RhoAGTP and total RhoA in whole protein extracts. Western blot of paired protein extraction samples against RhoA total and actin is shown as reference. n = 3, Mann-Whitney test, unpaired one-tailed *p < 0.05. D, Gating strategy for the isolation of LT-HSC from young and aged BM cells from representative experiments. E. Representative images of 3D confocal reconstruction and analysis of young, aged and aged treated with AACOCF3 HSCs stained with anti-PcPLA2. These images show total PcPLA2 (green) and the NE PcPLA2 signal (red) which overlaps completely with the total signal and appears in yellow (merged of red and green). Yellow arrows point to representative accumulation of NE PcPLA2 which can be more or less visible also depending on 3D rotation of the cells. Graphs are showing total PcPLA2 vol in µm3, percentage of NE PcPLA2 volume relative to total PcPLA2 volume and relative to DAPI volume. n = 3 Mann-Whitney, two-tailed test *p < 0.05; **p < 0.01; ***p < 0.001 ****p < 0.0001. F, Representative images of 3D confocal reconstruction stained with anti-TAZ and its analysis of HSC from young, aged and aged treated with Ri along confined young cells at 8 µm and 5 µm. n = 3, Mann-Whitney, two-tailed test *p < 0.05; **p < 0.01; ***p < 0.001 ****p < 0.0001.
Extended Data Fig. 3 Identification of image-derived morphometric and DAPI intensity features.
A. Schematic representation of the analyses of DAPI intensity by distance to the nuclear border. Created in BioRender. Mejía-Ramírez, E. (2025) https://BioRender.com/htlxngv. B. Stripplot displaying differences in morphometric and DIRs-related feature distributions among young (n = 177), aged (n = 164), and aged + Ri (n = 144) nuclei. Mean values are represented by a yellow dot on top of each distribution, with a yellow line aiding to visualize trends among conditions. Statistical significance was determined using the Mann-Whitney two-sided test. C. Heatmap depicting the correlation matrix for the 11 significant features identified among young and aged nuclei, after discarding highly correlated features. Marginal dendrograms exhibit clusters of feature similarity. The accompanying UMAP embedding, fitted with the specified features, is shown on the right. D. Heatmap depicting the correlation matrix for the 11 significant features identified among aged and aged Ri-treated nuclei, after discarding highly correlated features. Marginal dendrograms exhibit clusters of feature similarity. The accompanying UMAP embedding, fitted with the specified features, is shown on the right. E. Heatmap depicting the correlation matrix for the 20 features combined from the previous pairwise comparisons (young and aged, aged and aged Ri-treated) after discarding highly correlated features. F. Silhouette plot showing the silhouette scores for K-Means clustering (K = 4) for each nucleus in our dataset, with scores colored by cluster assignment. A red dashed vertical line indicates the overall silhouette score. G. Schematic representation of the multidimensional analysis pipeline. Features are extracted from whole DAPI intense microscopy images and used to construct a lower dimensional representation. Feature selection involved identifying statistically significant features among young and aged, and among aged and aged Ri-treated nuclei, followed by correlation-based filtering. H. Scatterplots illustrating the UMAP embedding constructed using nuclei from young, aged, and aged Ri-treated conditions, with color indicating variations in selected morphometric and intensity-related features. I. Volcano plots showing the most significant features among clusters. The x-axis represents the log2 feature fold change of the indicated cluster against the remaining clusters, while the y-axis indicates the significance of this change. Dot color intensity corresponds to significance, and the red horizontal line marks a significance threshold at a p-value of 0.005.
Extended Data Fig. 4 Reduced expression of G9a in aged HSCs.
A, Representative images of 3D confocal reconstruction of young, aged and aged+Ri treated HSCs stained with G9a antibody (red) and DAPI (grey). Mann-Whitney two-tailed, **p < 0.01, *p < 0.05. B. Flow cytometric analysis of transduced LSK pre-transplantation, representing 3 independent batches of transplantation. Mann-Whitney test, unpaired two-tailed, p = 0.700.
Extended Data Fig. 5 Transcriptional and chromatin accessibility changes in aged HSCs.
A, Proportions of the consensus peaks annotations in a donut chart. B, PCA plot for the ATAC-seq normalized read counts in the consensus peaks for the Ri-treatment experiment. C, Venn diagram for the number of common DARs in aged vs young and aged+Ri vs aged. D, Radar chart showing the minus logarithmic FDR for the significantly enriched GOs among the genes annotated to the DARs opening with Ri (permutation test, FDR < 0.1). E, PCA plot for the ATAC-seq normalized read counts in the aged vs young DARs for the young confined HSCs experiment. F, PCA plot for the RNA-seq normalized read counts in genes. Legend as in B. G, Volcano plots of differentially expressed (DE) genes in aged vs young and aged+Ri vs aged (moderated t-statistic, two-sided, unadjusted p < 0.001). Selected genes are shown. H, Venn diagram for the number of common DE genes in aged vs young and aged+Ri vs aged. I, Venn diagram for the number of common DE retrotranspon subfamilies in aged vs young and aged+Ri vs aged. Legend as in H. J–M, Enrichment plot and volcano plot for the acute inflammatory response GO term (J), the Interferome.org signature (K), the interferon-stimulated genes (L) and the aging signature (M) in aged+Ri vs aged. Permutation test. NES = normalized enrichment score. FDR = false discovery rate. Number of genes used for the analysis is also indicated.
Extended Data Fig. 6 Inhibiting RhoA activity induces Klf4 expression without altering cell cycle in aged HSCs.
A, Representative images of 3D confocal reconstruction of young, aged and aged+Ri treated HSCs stained with KLF4 antibody (red) and DAPI (grey). Mann-Whitney two-tailed, *p < 0.05. Experimental mice per arm n = 2. B, t-statistic for the KLF4-targeted genes in the aged+Ri vs aged comparison in ATAC-seq and RNA-seq. The number of genes with positive and negative RNA-seq t-statistic is shown. Top upregulated genes are shown. C, Main marker genes used for cluster annotation. D, Cell scores for the HSC signature (MolO) defined by Wilson et al. E, F, Cell assignments for the low/high-output signature by Rodriguez-Fraticelli et al. (E) and the dormant/active signature by Cabezas-Wallscheid et al. (F). G, Percentage of cells in each cluster per sample. scCODA’s statistically credible changes are shown. *Effects Final Parameter (EFP) < 0.5; **EFP < 1; ***EFP < 1.5. H, Percentage of HSCs in each cell cycle phase per sample. scCODA’s effects were all non-statistically credible. I, Log2FC against the average normalized expression for the DE analysis in aged vs young and aged+Ri vs aged in the HSC cluster (moderated t-statistic, two-sided, |log2FC| > 1 and FDR < 0.05). Selected genes are shown. J, CTCF and HOXA10 active cells (AUC > 0.007 and 0.01, respectively) in the UMAP.
Extended Data Fig. 7 Inhibiting RhoA activity improves epigenetic polarity of aged HSCs.
A, Representative images of 3D confocal reconstruction of young, aged and aged+Ri HSCs stained with anti-H4K16ac antibody (green), RhoAGTP (red), and DAPI (gray) as reference. Dotted line indicates symmetry axes. The graph shows quantification of polar cell number for H4K16ac. N = 3-5. 50 cells were counted per sample, per experiment, one-way Anova, *p < 0.05; **p < 0.01. B, Percentage of engraftment in bone marrow at 20wk post-transplantation, Young n = 9; aged n = 7; aged+Ri n = 9. Mann-Whitney test, unpaired two-tailed. C, Percentage of donor-derived HSC, ST-HSCs and LMPPs 20wk after transplantation, Young n = 9; aged n = 6; aged+Ri n = 9. Mann-Whitney test, unpaired two-tailed.
Supplementary information
Supplementary Information (download PDF )
Supplementary methods and Supplementary Figs. 1−7
Supplementary Table 1 (download PDF )
Morphometric and intensity features extracted from DAPI microscope images used for the multivariate feature analysis.
Supplementary Table 2 (download XLSX )
Data for the ATAC−seq experiment on young, aged and aged+Ri HSCs. Information on the samples, the consensus peaks, the DARs, the Fig. 5d statistics, the Gene Ontology enrichment, the transcription factor motif analysis and the KLF4-targeted genes.
Supplementary Table 3 (download XLSX )
Data for the ATAC−seq experiment on young and young 5-μm confined HSCs. Information on the samples, the consensus peaks, the DARs and the Fig. 5i statistics.
Supplementary Table 4 (download XLSX )
Data for the RNA-seq experiment on young, aged and aged+Ri HSCs. Information on the samples, the differentially expressed genes, the differentially expressed retrotransposons, the GSEA and the Gene Ontology enrichment of upregulated KLF4-targeted genes.
Supplementary Table 5 (download XLSX )
Data for the scRNA-seq experiment on young, aged and aged+Ri LSKs. Information on the samples, the cluster markers, the compositional analysis, the differentially expressed genes and the differentially active regulons.
Supplementary Table 6 (download XLSX )
Reagents table used in this study.
Supplementary Video 1 (download MP4 )
Representative 3D immunofluorescence reconstructions of an unconfined HSC. The nucleus is stained by DAPI (gray). RhoA-GTP was stained with an RhoA-GTP antibody, and is shown in red. Scale bars =1 µm.
Supplementary Video 2 (download MP4 )
Representative 3D immunofluorescence reconstructions of 8µm-confined HSC. The nucleus is stained by DAPI (gray). RhoA-GTP was stained with an RhoA-GTP antibody, and is shown in red. Scale bars =1 µm.
Supplementary Video 3 (download MP4 )
Representative 3D immunofluorescence reconstructions of 5µm-confined HSC. The nucleus is stained by DAPI (gray). RhoA-GTP was stained with an RhoA-GTP antibody, and is shown in red. Scale bars =1 µm.
Supplementary Video 4 (download MP4 )
Representative 3D immunofluorescence reconstructions of 3µm-confined HSC. The nucleus is stained by DAPI (gray). RhoA-GTP was stained with an RhoA-GTP antibody and is shown in red. Scale bars =1 µm.
Supplementary Video 5 (download MP4 )
Representative 3D confocal reconstructions of HSCs showing phospho-cPLA2 (PcPLA2) (light pink) and DAPI (gray) in young control HSCs. By using image analysis software Imaris we have compartmentalized PcPLA2 signal within the nucleus and at the nuclear envelope (NE). We have assigned light pink to NE PcPLA2 and magenta to nuclear PcPLA2 after compartmentalization. Surfaces have been created for better visualization. Scale bar =1 µm.
Supplementary Video 6 (download MP4 )
Representative 3D confocal reconstructions of HSCs showing phospho-cPLA2 (PcPLA2) (light pink) and DAPI (gray) in young NaB treated HSCs. By using image analysis software Imaris we have compartmentalized PcPLA2 signal within the nucleus and at the nuclear envelope (NE). We have assigned light pink to NE PcPLA2 and magenta to nuclear PcPLA2 after compartmentalization. Surfaces have been created for better visualization. Scale bar =1 µm.
Source data
Source Data Fig. 1 (download PDF )
Source Data of the blot in Extended Data Fig. 2c with unprocessed total protein and stained blot.
Source Data Fig. 2 (download XLSX )
Source Data of all graph figures + extended data figures.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mejía-Ramírez, E., Picazo, P.I., Walter, B. et al. Targeting RhoA nuclear mechanoactivity rejuvenates aged hematopoietic stem cells. Nat Aging 6, 68–87 (2026). https://doi.org/10.1038/s43587-025-01014-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43587-025-01014-w
This article is cited by
-
Squishing, squeezing and stretching age hematopoietic stem cells
Nature Aging (2026)