Abstract
Background:
Biological age derived from DNA methylation (mAge) reflects aging-related physiological changes and the risk of age-related diseases. However, the association between oral microbiome and mAge remains unclear.
Methods:
We examined associations between mAge, physical and cognitive function, and the oral microbiome in 311 community-dwelling, predominantly Chinese adults aged 85 years or older. Oral microbial composition was assessed using 16S rRNA sequencing, and mAge was estimated using four established epigenetic clocks: HorvathAge, HannumAge, PhenoAge, and GrimAge.
Results:
In this cohort (median chronological age = 88.6 years; median DNA methylation–based biological age = 81.4 years), individuals with lower mAge deviation, corresponding to a younger biological age, exhibit lower oral microbial alpha diversity based on Simpson’s index. While no taxa differences reach significance after correction for multiple testing, several short-chain fatty acid–producing genera, such as Prevotella_7 and Veillonella, show nominal associations with both mAge deviation and methylation at aging-related CpG sites, particularly in neurologically relevant genes. Higher abundance of Prevotella_7 species is associated with better cognitive performance (Mini-Mental State Examination), whereas Alloprevotella is linked to poorer cognition.
Conclusion:
Our findings highlight that, as far as we aware, previously unrecognized oral microbiome composition links to lower mAge deviation and better cognitive function among the oldest-old, suggesting a potential role of the oral microbiome in promoting healthy aging and informing future mechanistic investigations.
Plain language summary
People’s biological age, which can be estimated from changes to the structure of the molecules inherited from their parents (DNA), often differs from their actual age and reflects how well their body is aging. We studied whether the community of bacteria living in the mouth is linked to biological age and physical as well as brain function in 311 adults aged 85 years or older in Singapore. We found that people who appeared biologically younger tended to have less diverse oral bacteria within their mouths and showed distinct bacterial patterns compared with their peers. Higher amounts of some species, such as Prevotella_7, were linked to better thinking ability. These findings suggest that oral bacteria may influence healthy aging and brain health, offering potential new ways to monitor or support healthy aging through the mouth.
Similar content being viewed by others
Introduction
Despite rising life expectancy, age-related diseases remain a major burden on older adults and society1. These pathologies share basic underlying mechanisms with aging, such as epigenetic alterations2. One of the most extensively studied epigenetic alterations is DNA methylation (DNAm), which declines with age while increasing at specific Cytosine-phosphate-Guanine (CpG) islands harbouring regulatory genes, causing transcriptional noise2,3. Measuring these methylation changes in blood and other tissues, first-generation DNAm clocks have been developed to predict chronological age, while second-generation clocks have been trained on mortality risk and health indicators4,5,6,7. Both generation clocks are associated with overall disease risk, lifestyle behaviours, and other omics data8,9, with limited evidence suggesting an association between epigenetic aging and the microbiome10.
The human body harbours a complex and abundant microbiome with substantial inter-personal variability and the capacity to adapt to environmental stimuli11. The oral microbiome has been studied in relation to several systemic age-related diseases and physical and cognitive function. Lower salivary alpha diversity, which describes microbial diversity within an individual, has been associated with esophageal cancer12, lung cancer13, and frailty14,15. Similarly, beta diversity, which describes inter-individual differences in microbiome composition, has been associated with pancreatic cancer16, lung cancer13, and inconsistently with frailty14,15. Although Yang et al17. reported no significant association between oral alpha or beta diversity and cognitive function, other work shows that microbial beta diversity correlates with mental-health components related to anxiety and depression scores15, factors known to influence cognition18,19. Individual taxa abundances, on the other hand, such as those of Slackia and Prevotella species, have been positively associated with type 2 diabetes mellitus and cardiovascular disease risk, respectively20,21. Combined, these findings illustrate the oral microbiome’s association with late-life phenotypes. Given that the oral microbiome is also associated with chronological age15,22, and that certain oral taxa like Fusobacteria and Pseudomonas may alter host genome methylation patterns of genes such as NRG1 related to conserved pathways like PI3K/AKT23,24, a relationship between the oral microbiome and epigenetic age is plausible. Since the oral microbiome is malleable to lifestyle changes25, it could provide tangible interventions to optimize system health of aging individuals.
This study analyses associations between DNA methylation-based biological age, physical and cognitive function, and the oral microbiome—specifically alpha and beta diversity—in adults aged 85 years and older. Results show that individuals with a younger biological age exhibit lower oral microbial diversity based on Simpson’s index. Although no individual taxa differ significantly in abundance, short-chain fatty acid-producing genera such as Prevotella_7 and Veillonella contribute to compositional differences between groups and are linked to methylation at CpG sites related to neural function. A higher abundance of Prevotella_7 species is further associated with better cognitive performance. These findings indicate that oral microbial composition may relate to biological aging and cognitive health in the oldest-old population.
Methods
Study Design
The SG9026,27,28,29 is a cohort drawn from two longitudinal population health studies, the Singapore Chinese Health Study (SCHS) and the Singapore Longitudinal Ageing Study (SLAS), to describe the oldest-old demographic of community-living individuals aged 85 years and above with no diagnosis of dementia in Singapore. Out of the 1158 participants in the SG90 cohort, this study investigated a subset of 311 participants for whom both salivary oral microbiome and blood epigenetic data have been collected. Ethical approval for the SG90 study was secured from the Institutional Review Board (IRB) at the National University of Singapore (SCHS-SG90 approval number H-17-027, SLAS-SG90 approval number LB-15-081, Bio-sample collection DSRB approval number 2019/00439). All participants provided their informed written consent prior to participation. The current study has IRB approval for the use and analyses of primary and secondary data derived from the SG90 study. Comprehensive details of the approved data collection and analyses are described in a previous publication29.
Data collection
Participant characteristics
Demographic and lifestyle information were obtained as described elsewhere26,28. In brief, age, sex, ethnicity, education level, and smoking habits were gathered from participants through a self-reporting survey with a trained interviewer. Body weight was assessed to the nearest 0.1 kilogram (kg) using a Soehnle Exacta Comfort digital weighing scale (Model S63315 PSD). Height was measured to the nearest centimetre (cm) by means of a rigid, self-retracting metallic tape measure. Body mass index (BMI) was calculated using the formula weight/height² (kg/m²). Cognitive function was assessed using the Mini-Mental State Examination (MMSE), adapted for application in Singapore30. Physical performance was evaluated using the Short Physical Performance Battery (SPPB), which includes a repeated chair stand test, a balance test, and a 6-meter fast-paced walking test, with a total score ranging from 0 to 12, with 12 indicating the highest lower limb functionality31. Finally, physical function was examined using the Barthel Basic Activities of Daily Living (BADL) scale32 and the Lawton Instrumental Activities of Daily Living (IADL) scale33, with higher scores representing higher independence in both metrics.
Salivary oral microbiome
Saliva samples were unstimulated and collected in 50 mL polypropylene tubes without stabilizing reagents; dentate status was not recorded. Participants were instructed to refrain from eating, drinking, smoking, chewing gum, brushing teeth, or using mouthwash on the morning of collection. Samples were transported on ice packs and stored at −80 °C within 4 h, after processing into 1 mL aliquots. A 150 μL fresh aliquot was used for DNA extraction on the same day using the MasterPure™ DNA Purification Kit (cat. no. MC85200, Lucigen, LGC Biosearch Technologies, WI, USA).
Polymerase chain reactions (PCR) were set up with Thermo Scientific Phusion™ Hot Start Flex 2X Master Mix (cat. no. M0536), 341 F/805 R primers targeting the V3-V4 region of 16S rDNA, generating amplicons of approximately 465 bp. The thermal cycling conditions included 35 cycles with an initial denaturation at 98 °C for 30 s, followed by 10 s at 98 °C, 30 s at 54 °C, and 45 s at 72 °C, concluding with a final extension at 72 °C for 10 min. PCR products were isolated using 2% agarose gel and subsequently purified with AMPure XP beads (cat. no. A63880, Beckman Coulter Genomics, Danvers, MA, USA) before they were quantified by Qubit assay (cat. no. Q32851, Invitrogen, USA). Qualified libraries were selected based on concentrations > 2 nM as determined by Agilent 2100 Bioanalyzer (Agilent Technologies, USA) and KAPA Library Quantification Kits (cat. no. KK4824, Kapa Biosciences, Woburn, MA, USA). Sequencing was done using NovaSeq 6000 SP paired-end reads (2 × 250 bp, 500 cycles). The resulting paired-ends then had barcode and primers truncated (cutadapt v1.9, ‘-g R1 -G R2 -n 1 -O 17 -m 100’), were merged with overlap using FLASH (v1.2.8, ‘-m 10 -M 100 -x 0.25 -t 1 -z’), and subjected to quality control (fqtrim, ‘-P 33 -w 100 -q 20 -l 100 -m 5 -p 1 -V -o trim.fastq.gz’). Chimeric sequences were filtered using VSEARCH (v2.3.4, default parameters). Finally, dereplication was done with DADA2. The resulting amplicon sequence variants (ASVs) were then assigned using the SILVA database (release 132, confidence level > 0.7)34. Each library had between 9623 and 72412 reads, with a median of 49173 reads.
Epigenetic clock measures
Fasted peripheral venous blood samples were collected to isolate peripheral blood mononuclear cells (PBMCs) using CPT™ Mononuclear Cell Preparation Tubes (cat. no. 362753, BD Vacutainer, NJ, USA). DNA was extracted from PBMCs using the QIAamp DNA Blood Mini Kit (cat. no. 51104, Qiagen, Hilden, Germany), with quality evaluated using the Agilent TapeStation (Agilent, CA) and quantification performed using the Qubit fluorometer (Thermo Fisher Scientific, MA). Approximately 1 µg of DNA per sample was bisulfite-treated using the Zymo EZ DNA Methylation Kit (cat. no. D5001, Zymo, CA) for methylation analysis. Genome-wide DNA methylation was measured using the Illumina Infinium MethylationEPIC v1.0 BeadChip (cat. no. WG-317-1001, Illumina, CA), covering over 850,000 CpG sites across the genome.
The Horvath DNAm Age (PCHorvath)7, Hannum DNAm Age (PCHannum)6, DNAm PhenoAge (PCPhenoAge)5, and DNAm GrimAge (PCGrimAge)4 were computed using a principal-component (PC) based implementation to maximize reliability35. Normalization was performed according to the most effective data processing pipelines previously found for each specific PC epigenetic clock36. The median epigenetic age (median mAge) for each individual, representing the central tendency of the values obtained from all four clocks, was calculated and then subtracted by their chronological age to derive the mAge deviation. Negative mAge deviation values indicate that an individual is epigenetically younger than their chronological age, while positive values suggest the opposite. Previous studies have linked all four clocks to different diseases and risk factors, with each clock showing unique associations in both the type and magnitude of its effects8. This variability underscores the value of combining the clocks to capture a broader spectrum of aging-related contributions. For example, mAge deviation evaluated using GrimAge was linked with substance use, while Horvath’s clock was uniquely associated with total cholesterol, and PhenoAge was able to capture hip-to-waist ratio9. The decision to use the median of these four clocks was taken to align with the ABLE intervention study protocol37.
Data processing and curation
Five participants had blood and saliva samples collected on different dates between one and 28 days apart. One participant had missing biological data collection dates. Given that both the microbiome and blood epigenetic age display changes even on a daily timescale11,38, all six samples were ruled out from the dataset, leaving 311 individuals. Ten individuals’ BMI values, one participant’s smoking status, one participant’s education level, one SPPB score, five MMSE scores, and four IADL scores were not reported. Given the small proportion of absent data and a skewed distribution of the BMI, SPPB, MMSE, and IADL variables, their missing values were imputed using median imputation. For the same reason, smoking status and education level were imputed using mode imputation. Since blood and saliva were collected on different dates from the demographic and participant characteristics, an interaction test was first conducted to assess the significance of this time difference at p < 0.05. Among participants with complete date information (n = 224), the median lag between biosample collection and (A) demographic/MMSE/IADL/BADL assessments, (B) smoking-status questionnaire, and (C) SPPB / BMI testing was 336, 335, and 189 days, respectively (Supplementary Fig. S1). Because the date of demographic and physical testing data collection was missing for 87 individuals, a three-step approach was subsequently employed. First, among participants with complete dates (n = 224), baseline ordinary least-squares (OLS) models (statsmodels v0.14.2) were fitted using confounders and microbiome principal components as inputs and mAge deviation as the outcome. Model performance was then compared to OLS models that included either only the main time effect or both the main time effect and interaction terms, using nested ANOVA. For the MMSE, SPPB, IADL, and BADL measures, these variables were used as the outcome in place of mAge deviation. In the second step, the missingness of date information was confirmed to be significantly correlated with sex and ethnicity, prompting multiple imputation by chained equations (MICE) with 10 iterations each to generate five distinct datasets (miceforest v5.6.2). Baseline, main-effect, and interaction-effect models were refitted in each of the five imputed datasets and compared via nested ANOVA. Finally, min- and max-bounding imputation was performed as a sensitivity analysis, followed by further model-fit comparisons for each outcome. Results indicated a significant time and time-interaction effect for BADL and SPPB determined by sensitivity analysis. Following significant associations of BADL and SPPB with either alpha or beta diversity, tests have therefore been adjusted to account for interaction effects. Although BMI, IADL, and MMSE occasionally showed significant main or interaction time coefficients under certain sensitivity conditions, effect sizes were minimal, and ANOVA did not support improved model fit upon adding these terms; they were therefore omitted from later analyses. For further details, please refer to Supplementary Data 1 (complete-case, non-imputed analyses), Supplementary Data 2 (sensitivity analyses using minimum- and maximum-bound imputations), and Supplementary Data 3 (multiple-imputation analyses pooled across five datasets), each containing baseline, main-effect, and interaction models with corresponding ANOVA comparisons and README documentation.
To compare participants with higher and lower mAge deviation, the cohort was divided into two groups based on the median of the mAge deviation variable. This approach preserved statistical power for differential abundance analyses and increased the likelihood of achieving a good fit in the machine learning classifier by reducing noise and simplifying the training process, particularly given the limited sample size. For consistency across analyses, the same median-based grouping was applied to both alpha and beta diversity measures. Throughout this study, ‘lower’ refers to mAge deviation values below the median threshold, indicating individuals whose biological age is markedly below their chronological age. Conversely, ‘higher’ designates values above the median, reflecting smaller or even positive deviations between biological and chronological age.
Statistical and machine learning analyses
Covariate screening and model adjustment
To ensure consistency and preserve statistical power in this relatively small cohort, a unified covariate screening strategy was applied. Only covariates significantly associated with each outcome were retained. Potential confounders, including age, sex, BMI, smoking status, ethnicity, education level, and sequencing depth (for non-rarefied analyses), were selected based on prior evidence that these factors shape the oral microbiome or cognition39,40,41,42 and were evaluated for their association with relevant outcome variables (e.g., alpha diversity metrics, beta diversity, MMSE scores) using test-appropriate univariate models. Data on periodontal status and oral-hygiene behaviour were not available, and alcohol-intake variables were excluded because they were near-monomorphic (>95% low or no use) with ≥20% missingness in at least one beverage category. Pearson correlation was applied for continuous variables, Spearman correlation for ordinal variables, and t tests or ANOVA for binary and categorical variables, respectively. Resulting p-values were adjusted using the Benjamini–Hochberg procedure (FDR < 0.05). No covariates were included for alpha diversity models (Supplementary Data 4); sex was retained for beta diversity (PERMANOVA, Supplementary Data 6); and education level, sex, smoking, and ethnicity were included for MMSE models.
Alpha diversity analyses
To assess microbial diversity within individuals in the cohort, alpha diversity metrics, including Shannon and Simpson indices (calculated as 1-D), were computed at ASV level. To further examine microbial richness and the evenness of taxonomic distributions, ASV richness and Pielou’s evenness were calculated. All metrics were obtained utilizing the Python package sci-kit bio (v0.6.2) and visualized using Seaborn (v0.13.2). They were averaged over manual 1000-fold rarefaction at a sequencing depth of 20651 to account for differences in library size43. Library size for rarefaction was set to balance sample inclusion and read detection, leading to the exclusion of 8 samples for alpha diversity calculations. Alpha diversity indices were compared between groups using Mann–Whitney U tests for dichotomously grouped variables (e.g., mAge deviation, chronological age, BADL), and Kruskal–Wallis tests for ordinally grouped outcomes (e.g., MMSE, SPPB, IADL), with a significance threshold of p < 0.05. Effect sizes were reported as Cliff’s delta (for Mann–Whitney) or η² (for Kruskal–Wallis). The association between alpha diversity indices and cognitive or physical assessments was analysed using the narrowest binning scheme that preserved at least two distinct groups, while avoiding overlapping bin edges and accounting for skewed distributions. Quartiles were used for MMSE and SPPB, tertiles for IADL due to limited score spread, and a binary split for BADL reflecting its strong skew. Given that BADL data collection occurred at significantly different times (see Data Curation), a generalized linear model (GLM) with a binomial family and logit link function was also applied to each alpha diversity index to account for potential temporal biases (statsmodels v0.14.2).
Beta diversity analyses
To quantify differences between mAge deviation groups, beta diversities were calculated using robust Aitchison’s and phylogenetic robust Aitchison’s distances (Gemelli v0.0.11)44,45 as well as Bray-Curtis dissimilarities on ASV level (vegan v2.6-8). In brief, the first two methods utilize a robust version of the centred log ratio transform, which calculates the geometric mean only on nonzero entries, avoiding unwanted library size dependence of the geometric mean induced by pseudo-value imputation in sparse matrices46. The phylogenetic version integrates Fast UniFrac and received additional input in the form of a phylogenetic tree created by FastTree (v2.1.11), built on sequence alignments obtained through MAFFT (v7.515). For both robust distance metrics three principal components were computed, two of which were then plotted. Since Bray–Curtis does not use a robust centred log ratio transform, 1000-fold rarefaction was again performed and dissimilarity matrices averaged over all iterations (vegan v2.6-8). Results were visualized using principal coordinates (ggplot2 v3.5.1). Individuals for all three beta diversities were coloured according to their mAge deviation group. Centroids and 95% confidence ellipses were calculated for each group. Associations between beta diversity and mAge deviation group or other outcome variables (MMSE, BADL, IADL, SPPB) were tested using PERMANOVA with 10,000 permutations, adjusting for sex following a significant result (vegan v2.6-8). Time interaction effects were added for both BADL and SPPB, based on earlier sensitivity analysis (see Data Curation). Subsequently, similarity percentages (SIMPER) were used to assess contributions of taxa to the significant PERMANOVA result obtained by Bray–Curtis using 999 permutations (vegan v2.6-8). Data were normalized by library size for each sample. Next, community compositions were visualized using stacked bar plots after normalizing data at species, genera and phyla level. Species beyond the cumulative sum of 75% relative abundance and genera beyond 90% relative abundance were grouped into ‘Other’.
Differential abundance analyses
Gene family profiles in the form of KEGG orthologous (KO) groups and pathway abundances were predicted using PICRUSt 2.0 (v1). Differences in sequencing depth between groups were verified to be non-significant using Mann-Whitney U at a p ≤ 0.05 significance level. Differential abundance analysis was then conducted using the package ALDEx2 (v1.36.0) in R on ASV, genus, KO groups, and KEGG pathway level for mAge deviation classes and chronological age via median split. Prior to analysis, prevalence filtering was applied to remove features present in fewer than 15% of samples. Data was clr-transformed through the internal ALDEx2 pipeline. Additionally, MaAsLin2 (v1.22.0) was applied using default preprocessing settings on the same prevalence-filtered feature tables. Standard covariate screening was not feasible for differential abundance tools, so a predefined subset of covariates (sex, BMI, and smoking) was used for adjustment. Ethnicity was omitted due to low variability and no prior association with either alpha or beta diversity in the cohort. Covariate adjustment was applied only in MaAsLin2 models, where unadjusted comparisons yielded significant results; no covariates were included in ALDEx2 analyses due to a lack of significant findings. Outputs were visualized using ggplot2 (v3.5.1), yielding four volcano plots. In preliminary analyses, comparisons restricted to samples in the outer quantiles of mAge deviation yielded no additional significant findings and are therefore not shown.
Machine learning
To prepare count data for machine learning, genus-level raw abundances were subjected to a centered log-ratio transform. Pseudo values were drawn from a uniform distribution between 0.1 times the detection limit and the detection limit. This naïve imputation approach has been shown to perform well even compared to more sophisticated methods47. Machine learning classifiers were then fitted to the data using abundances on genus level as input and group labels for mAge deviation as output by means of Scikit-learn (v1.1.2) and xgboost (v1.7.2) packages. Model selection was performed using a pipeline comparing accuracies in nested cross-validation (n_outer = 5, n_inner = 3) between nine different models, obtained by mixing three feature selection techniques (none, L1 logistic regression, backwards sequential feature selection (SFS)) with three model types (random forest, XGBoost, stochastic gradient boosting with XGBoost). Feature selection was done once per outer fold split on the entire outer training data using 3-fold cross-validation. Bayesian hyperparameter selection was done afterwards using Optuna (v3.1.0) on inner folds (n_trials = 25). Refer to Supplementary Data 13 for more details on the chosen hyperparameter space. The aim of the pipeline was to select a model that would avoid overfitting, which was common on this dataset, while still being able to capture meaningful insights. The best performing model architecture, a stochastic gradient boosted XGBoost model with backwards sequential feature selection, was then also used to train a confounder model using data on age, sex, BMI, and smoking as input. Additionally, a random baseline was estimated by randomly shuffling the labels in the test set once per outer fold and reporting its performance. All models were judged on accuracy, ROC-AUC, and F1 score. Normal distribution of these performance metrics was guaranteed using Shapiro-Wilk at a p ≤ 0.05 cutoff before they were compared between models using a Welch’s t test at a p ≤ 0.05 threshold. SHAP feature importance values (v0.41.0) were obtained from the model trained on microbial genera. Additionally, distributions of the 10 most positive and negative features across all folds have been retrieved.
Network analyses
Finally, to visualize microbial community structure and its relationship with CpG sites, a cross-domain network using SPIEC-EASI was fitted (v1.1.3, lambda.min.ratio =1e−1, MB method)48. To ensure the robustness of edge selection and minimize overfitting to sample-specific noise, we used the StARS (Stability Approach to Regularization Selection) framework within SPIEC-EASI. This approach fits multiple networks on random subsamples of the data and selects the sparsest model whose overall edge variability remains below a predefined threshold (0.05 in our case, using 80% subsampling over 50 repetitions), favoring network structures that are consistently reproducible across resampled subsets of the cohort. CpG sites were selected based on the union of all sites used for the four epigenetic clocks used in this paper. Then, sites with more than twenty percent missing values across all samples were excluded, while the remainder was filled using median imputation. The resulting 1624 CpG sites were then deconstructed into principal components, of which the first one hundred were fed to the network. In terms of microbial taxa, only the 267 ASVs present in at least 15 percent of samples were used as input. The resulting network was then trimmed by removing taxa without any connections and CpG PC nodes that were only connected to other CpG PC nodes. Community membership was assigned using Louvain’s method. Given a normalized mutual information value of only around 0.7 after one hundred repeated runs of Louvain, indicating variability in community membership allocation, a community consensus membership approach using an iterative version of Louvain over 1000 iterations was chosen to improve stability49. Gene annotations related to CpG sites with high loadings in PCs connected to microbial taxa were extracted using the Ensembl database (hg38 build). For each relevant PC, we selected the ten CpGs with the highest positive and the ten with the highest negative loadings (twenty CpGs total per PC), thereby capturing the most influential features without introducing excessive noise. Gene annotations were then assigned based on the genomic coordinates of each CpG using Ensembl’s regulatory annotation. Tissue enrichment analysis of the resulting gene list was performed using the TISSUES Curated 2025 database via Enrichr50,51.
Cognition models
To examine the oral microbiome’s relationship with cognition, a generalized linear model using the identity link function and Gaussian family parameters as outcome was fitted (statsmodels v0.14.2). As inputs, ASVs connected to CpG PCs in the network were included, alongside abundance-filtered ASVs (mean relative abundance ≥ 0.005) belonging to families Veillonellaceae and Prevotellacae to capture all possible species that were previously identified to be either central to the network or associated with mAge deviation Alpha and Beta diversities. Covariates retained through screening for the cognition outcome—education level, sex, smoking status, and ethnicity—were included in the model. P-values were adjusted for multiple comparisons using the Benjamini–Hochberg method to maintain an FDR < 0.05.
Statistics and Reproducibility
Unless otherwise specified, analyses were performed using Python (v3.11.5) in a Conda environment (v23.7.4) and R (v4.4.1) with open-source, version-controlled packages. Statistical significance was evaluated at p < 0.05 unless stated otherwise and adjusted for multiple testing using the Benjamini–Hochberg false discovery rate procedure. Each participant represents an independent biological replicate (n = 311); no technical replicates were generated.
To evaluate the robustness of time-related effects, analyses were repeated across complete-case, multiple-imputed, and bounded-imputed datasets. Model assumptions were verified using diagnostic plots and normality tests where applicable. Machine learning models were trained using nested cross-validation with fixed random seeds and repeated feature selection to prevent overfitting. Network inference reproducibility was assessed through stability selection (StARS) with fixed random seeds and repeated subsampling.
Results
The 311 oldest-old individuals (Table 1) had a median age of 88.6 [87.3–90.4] years; 62.1% were female, 94.5% of Chinese ethnicity, and most were lifetime non-smokers (68.5%) with low education. Median mAge was 81.4 [77.4–84.5] years. Chronological age showed a modest but statistically significant positive association with mAge (Pearson r = 0.23, p = 4.35 × 10⁻⁵; Spearman ρ = 0.20, p = 3.50 × 10⁻⁴; Supplementary Fig. S2). PCHorvath and PCPhenoAge clocks showed lower mAge deviations than PCHannum and PCGrimAge, with a combined median of −9.86 years (Supplementary Fig. S3).
Higher alpha diversity associated with higher mAge deviation and physical function
Individuals with a higher mAge deviation showed significantly higher Simpson diversity (Fig. 1A), reflecting this index’s sensitivity to greater abundance of dominant taxa. Similar but non-significant trends were observed for ASV richness and Shannon (Fig. 1B, D), but not for Pielou’s evenness (Fig. 1C). Individuals with a lower mAge deviation skewed toward the lower quartile in all indices, with a greater proportion showing lower microbiome diversity overall. MMSE, SPPB, chronological age, and IADL showed no significant associations with alpha diversity (Supplementary Fig. S4-7), although individuals with the highest BADL scores exhibited significantly higher Shannon, and richness indices, persisting after time-adjusted analysis (Supplementary Fig. S8, Supplementary Data 5). Summary statistics for all outcomes and their associations with alpha diversity are presented in Table 2.
Simpson (A) and Shannon (B) alpha diversity indices, Pielou Evenness (C), and ASV Richness (D) index comparing both mAge deviation groups. In all violin plots, the centre line represents the median, box limits constitute upper and lower quartiles. Dots represent individual samples. “*” indicates significance at p-value ≤ 0.05. Statistical comparisons were performed using two-sided Mann–Whitney U tests (no multiple-comparison correction). All plots each capture 150 biologically independent individuals in the lower mAge deviation group and 153 biologically independent individuals in the higher mAge deviation group as determined by cohort composition after filtering out samples with low sequencing depth for rarefaction. ASV amplicon sequence variant, mAge methylation age.
Beta diversities consistently associated with physical function but not cognition, chronological age or physical performance
IADL scores were consistently associated with both the Bray–Curtis dissimilarity index and robust Aitchison’s distance (p-value = 0.021, R-squared = 0.005 and p-value = 0.017, R-squared = 0.013, respectively), even when being controlled for by sex, which was independently associated with beta diversity (see Supplementary Data 6 for full covariate testing). Associations for BADL and SPPB were present only before accounting for time effects, and MMSE as well as chronological age showed no association with any beta diversity metric (Table 3).
Trend in the shift of oral microbiome dominance structure associated with mAge deviation
PERMANOVA showed no association of mAge deviation groups with robust or phylogenetic Aitchison’s distance (Fig. 2A, B) but did reveal a significant link with Bray Curtis dissimilarity (Fig. 2C; p = 0.028, R² = 0.005), which lost significance after sex adjustment (Table 3). SIMPER analysis identified ASVs significantly contributing to Bray–Curtis dissimilarities between mAge deviation groups (Fig. 2D). The magnitude of their contributions was observed to scale with their abundance, meaning more dominant taxa added more information in differentiating between groups. Microbiome composition analysis (Fig. 3A) identified unclassified Prevotella_7 and Veillonella, both key contributors to SIMPER, to be among the three most abundant species and four most abundant genera annotations (Fig. 3A, B). The most contributing insignificant ASV, which belongs to the Streptococcus_pneumoniae species, also had the highest mean abundance of all ASVs (Fig. 2D). Moreover, the Streptococcus genus was the second most abundant overall, showing a non-significant trend toward higher relative abundance in individuals with greater mAge deviation (Fig. 3B). At the phylum level, more Bacteroidota (including Prevotella_7) and fewer Proteobacteria (notably Neisseria and uncultured Haemophilus, Fig. 2D) were observed in the group with lower mAge deviation (Fig. 3C).
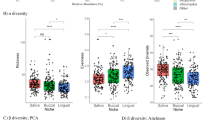
Robust principal component analysis (n = 311 biologically independent samples) (A), phylogenetic robust principal component analysis (n = 311) (B), and Bray–Curtis principal coordinate analysis (n = 303 after rarefaction) (C) with samples coloured according to their corresponding mAge deviation group. The percentage of variance explained in (A) and (B) is based on the proportion of the three computed eigenvalues, whereas in (C), it is compared to the total variance across all eigenvalues. In (D), contributions of ASVs to group differences based on Bray–Curtis distances are displayed (n = 303), calculated using SIMPER with 999 permutations. ASVs above the dashed line show significant contributions at unadjusted p ≤ 0.05. The six most abundant ASVs across the entire dataset are annotated at the species level. Mean abundance refers to the average abundance of a given ASV across mAge deviation groups. PC principal component, PCo principal coordinate.
Bar plots depicting species (A), genus (B), and phylum (C) composition of both groups. Species that do not contribute to the cumulative sum up to 0.75, and genera below the cumulative sum of 0.9, have been grouped under “Other”. Each plot includes n = 150 biologically independent participants in the lower mAge deviation group and n = 153 in the higher group. mAge methylation age.
Differential abundance analysis using MaAsLin2 on prevalence-filtered data identified two KEGG pathways (GLUCONEO.PWY, HEMESYN2.PWY) and two KO groups (K00059, K00086) that were significantly associated with mAge deviation (p = 0.0002–0.0003; q = 0.048, Supplementary Data 7), but these associations were not retained after adjusting for sex, BMI, and smoking (q > 0.97 for all, Supplementary Data 8). At the genus level, Prevotella_7 and Veillonella—both abundant taxa highlighted by SIMPER—were associated with mAge deviation in covariate-adjusted MaAsLin2 models only prior to multiple testing correction (p = 0.008 and 0.038; q = 0.38 and 0.50, respectively). Prevotella_7 showed no nominal associations with sex, BMI, or smoking, and Veillonella nominally associated only with sex (q > 0.05 for all). No ASVs or genera were significantly associated with mAge deviation after multiple testing correction. ALDEx2 identified no significant features at any taxonomic or functional level (Supplementary Fig. S9). Neither method yielded significant associations with chronological age (Supplementary Fig. S10, Supplementary Data 9).
To gain further insights into taxa able to differentiate between mAge deviation groups, machine learning classifiers were fitted. Machine learning classifiers trained on microbial genera did not outperform random baselines in predicting mAge deviation (p < 0.05), whereas confounder-based models (age, sex, ethnicity, smoking, BMI) did (Supplementary Fig. S11A, B). SHAP analyses indicated reliance on only a subset of microbial genera (Supplementary Fig. S11C, D). However, SHAP values were too small (scale of 1e−19) to render them informative for model decision making and thus, further interpretation.
Three oral microbial subcommunities and seven microbial taxa associated with host methylation levels in aging-related CpG sites
A SPIEC-EASI network revealed seven distinct microbial subcommunities and seven direct associations between ASVs and principal components capturing methylation at age-related CpG sites (Fig. 4). While one subcommunity was disconnected, all other subcommunities cohesively shaped the oral microbiome. An unclassified Veillonella species was the most connected ASV, consistent with its significance in earlier beta diversity analyses (Fig. 2D). Overall, three out of the six most central nodes in the network were unclassified Veillonella species, present in two distinct subcommunities.
The network was inferred using SPIEC-EASI (n = 311 biologically independent participants). Round nodes represent microbial taxa, square nodes represent principal components of aging-related CpG sites. Microbial nodes have been scaled in size according to their network degree centrality. Colour of nodes represents community membership determined by iterative Louvain. Taxa (1–7) connected to CpG PCs (A–E) have been highlighted in red and annotated at the species level in the legend. PCs of CpG sites have been additionally annotated with the percentage of variation they explain. PC principal component.
All five CpG PCs each explained 0.31–1.06% of blood methylation variation (2.9% cumulatively) and mapped to three subcommunities, with two containing two PCs each. These PCs were linked to seven ASVs, generally within the same subcommunity, except for one moderately abundant Veillonella. The PC A and B subcommunity (combined 1.75% variation) connected to Selenomonas, Dialister, and Veillonella species, while PC C and E subcommunity (combined 0.82% variation) linked to taxa belonging to Capnocytophaga, Rothia aeria, and Abiotrophia. Finally, PC D (0.33% variation) formed a single link to a Prevotella_7 species. Because these PCs represent variation in methylation levels at aging-related CpG sites, gene annotations associated with their high-loading CpGs (top ten positive and top ten negative loadings) were further examined (Supplementary Data 10). PCs A and B both captured variation in CpG methylation levels of overlapping genes like ATP13A4 (cation-transporting ATPase), ZMIZ1 (zinc finger protein involved in transcriptional regulation and neural development), CALD1 (actin-myosin interaction regulator), and TRAF3 (ubiquitin ligase regulating i.e. mitogen-activated protein kinase (MAPK) and interferon regulatory factor (IRF) pathways), among others. Both principal components were classified into the same microbial-methylation subcommunity. Principal components C and E capture genes such as ICAM5 (facilitates intracellular adhesion in cells of the nervous system) and CACNA1A (voltage-sensitive calcium channels), respectively, without overlapping gene sets. Principal component D included CpG sites associated with genes such as SMPD3 (sphingomyelin hydrolysis), RBFOX1 (neuronal RNA-binding protein), NGEF (guanine exchange factor involved in axon guidance), and P2RY6 (microglial receptor). Enrichment analysis of all involved genes revealed that the top-ranked relevant tissues were brain, central nervous system, and neurons (adjusted p ≈ 0.056 for all), pointing toward a predominantly neurological functional profile (Supplementary Data 11).
Prevotella_7 species associated with better cognitive function
Given this largely neurological profile of connected genes, and the fact that some corresponding microbes metabolize lactate or produce short-chain fatty acids (SCFAs), both relevant to brain health52,53, the association between oral bacterial abundances of relevant taxa and MMSE scores was evaluated (Supplementary Data 12).
Three unclassified Prevotellaceae species were significantly related to cognition after adjusting for multiple testing (Benjamini-Hochberg) and accounting for education, sex, smoking habits, and ethnicity. Two Prevotella_7 species were positively associated with MMSE scores (Fig. 5A, B), whereas one Alloprevotella species was negatively associated with MMSE scores (Fig. 5C). One of the identified Prevotella_7 species (Fig. 5A) had the second highest mean abundance across the cohort and was most significantly associated with mAge deviation beta diversities (Fig. 2D). In order to put regression effect sizes into a context capturing clinical significance, the associations of MMSE scores with the 10th and the 90th percentile of relative abundance of the three identified species is displayed in Fig. 5D. Higher abundance of Prevotella_7 species is associated with higher MMSE scores between 0.89 and 1.57 points, whereas greater abundance of the identified Alloprevotella species is associated with 1.43-point lower MMSE scores.
Panels (A–C) show scatter plots with linear regression lines (generalized linear model, Gaussian family with identity link, two-sided test, n = 311 biologically independent participants) indicating the relationship between MMSE scores and the relative abundance of specific microbial species: s__Prevotella_7_unclassified (A, B) and s__Alloprevotella_unclassified (C). Red lines represent the fitted linear regression mean, with shaded 95% confidence intervals. Panel (D) depicts the change in MMSE scores (ΔMMSE) associated with moving from the 10th to the 90th percentile of microbe abundance for three bacterial taxa. Positive changes in MMSE scores are shown in green, while negative changes are shown in red. MMSE Mini-Mental State Examination.
Discussion
An association was observed between biological age measured by mAge, physical and cognitive function, and the oral microbiome in the oldest old Singaporeans. Individuals with a higher mAge deviation and those with better BADL scores exhibited a trend towards greater oral alpha diversity, significant for the Simpson index in the case of mAge deviation and for multiple indices in the case of BADL; while Bray–Curtis beta diversity was weakly associated with IADL scores and differed by mAge deviation before adjustment for sex. Bray–Curtis dissimilarities appeared to reflect compositional shifts involving dominant taxa such as Prevotella_7 and Veillonella. Both showed non-significant trends toward higher abundance in participants with lower mAge deviation. However, no individual taxa reached FDR significance in differential abundance testing. Both genera had taxa linked to aging-related CpG site methylation, many of which mapped to genes active in the nervous system. Three Prevotellaceae ASVs, notably a highly abundant Prevotella_7, showed significant associations with cognitive function.
Contrary to initial expectations, individuals with a higher mAge deviation displayed greater alpha diversity. Although no prior studies have investigated the association between mAge and oral alpha diversity, findings in 8224 middle-aged American adults showed that greater oral diversity is associated with lower all-cause mortality54. Since the epigenetic clocks used in this study are strongly associated with mortality risk8,9, individuals with lower mAge deviation were presumed to have higher alpha diversity. At the oldest old age however, the microbiome’s nonlinear changes over a lifetime22 may lead to lower alpha diversity alongside higher abundances of beneficial taxa. Nonetheless, greater Shannon diversity and richness were associated with better BADL, which is closely linked to frailty55, thus aligning with existing literature showing a higher Shannon diversity and richness to be inversely associated with frailty in 1357 Canadian adults aged 35–70 years15, a finding later corroborated in 679 British twins (median (IQR) age 66 (15) years)14. In both studies, beta diversity was associated with frailty, mirroring its association with IADL in this study. The lack of association of cognitive function with either alpha or beta diversity has previously been reported in 30 African Americans (mean (range) age 54.8 (41–73) years)17. Although the oral microbiome may influence physical performance via nitric oxide production25, no significant associations with alpha or beta diversity were observed. Overall, these findings in the oldest old mirror microbial patterns observed in younger cohorts, but their association with biological age stands out as both novel and counterintuitive. A possible explanation for these divergent associations is that diversity itself is an imperfect proxy for microbial health56. Greater diversity does not always imply better outcomes and has been observed in dysregulated states such as major depressive disorder57 and in formula-fed infants58. Microbial resilience, and thus host health59, may instead depend on community stability, which reflects the balance between cooperation and competition: extensive interdependence among many taxa can reduce stability, whereas competitive interactions tend to preserve it60. In this dataset, stability could arise through two distinct ecological routes: (i) increased dominance of competitive taxa, reflected by higher Simpson diversity and lower biological age deviation, or (ii) expansion of the overall pool of competing species, reflected by higher Shannon and richness indices and better physical function (BADL). Ultimately, whether higher diversity represents resilience, and thus beneficial health outcomes, or instability likely depends on the functional roles of the dominant or newly incorporated taxa.
Lower alpha diversity in individuals with lower mAge deviation, along with beta diversity associations, mainly stems from higher Prevotella_7 and Veillonella abundance. Veillonella also served as a highly connected hub in the oral microbial network and was associated with aging-related CpG site methylation. Veillonella and Prevotella species produce SCFAs like propionate during lactate and pyruvate metabolism, and fibre fermentation, respectively61,62. SCFAs have been associated with several age-related diseases and epigenetic alterations. Propionate supplementation at 10 g/day in overweight adults increased peptide YY (PYY) and glucagon-like peptide-1 (GLP-1) secretion, reducing energy intake, weight gain, and maintaining insulin sensitivity compared to an inulin-control group63. These findings are particularly relevant as obesity has been found to cause DNA methylation changes64. Additionally, propionic acid supplementation in a randomized, double-blinded, placebo-controlled human trial also decreased low-density lipoprotein (LDL), total, and non-high-density lipoprotein (HDL) cholesterol levels, along with reduced intestinal cholesterol absorption in mouse models65. In mice with lung cancer, oral propionate showed life extension benefits, mediated through chromatin remodelling66. Other research has reported histone acetylation as a key mechanism by which propionate and other SCFAs alter gene expression at both intra- and extraintestinal sites67. Given that chromatin remodelling and histone acetylation are closely linked to DNA methylation68, these data suggest that oral microbiota-derived metabolites may plausibly impact epigenetic regulation in the aging process. Moreover, many of the aging-related CpG sites whose methylation levels were associated with a Prevotella species were located on genes coding for proteins that are active in the nervous system. For example, RBFOX1 has been linked to brain amyloidosis and Alzheimer’s disease in humans69, and P2RY6 is implicated in Alzheimer’s and Parkinson’s diseases, neuroinflammation, and aging-induced synaptic loss in mouse models70. Given that lower gut abundance of Prevotellaceae has been associated with Parkinson’s disease71, and SCFAs have been shown to improve neural survival and stimulate neurogenesis in the enteric nervous system in mice models72, a plausible hypothesis is that oral microbial profiles enriched in SCFA-producing taxa may influence central nervous system function via metabolite-driven epigenetic pathways. A less diverse microbiome optimized for SCFA production could broadly modify DNA methylation, hence indirectly affecting biological age through alterations at loci critical for cognition and brain health. While this mechanistic pathway was not tested in the present study, it offers a conceptual framework for future research. However, functional microbiome data are necessary to elucidate the differential impacts of Alloprevotella and Prevotella species on cognition, as even closely related Prevotella species can exhibit markedly different metabolic capacities73.
Future research should hence integrate metagenomic, longitudinal metatranscriptomic, and salivary metabolomic analyses to better capture the full spectrum of oral microbial diversity, including low-abundance taxa such as viruses, fungi, archaea, and their respective functional activity. Another limitation of this research is the chosen study population. Focusing on the oldest-old in Singapore enhances insights into advanced age but introduces survivorship bias, as individuals harbouring harmful microbial profiles may be underrepresented. This may have limited statistical power, such that differential abundance signals observed at nominal significance in this cohort could reach FDR significance in more age-diverse populations. The composition of the oral microbiome may also differ in edentulous participants74, which could introduce a potential confounding variable. However, this factor was not considered in the current analysis due to the unavailability of relevant data. Periodontal-disease status and detailed dietary-pattern data were likewise not incorporated and therefore remain potential unmeasured confounders39,40. The unique demographic also constrains the generalizability of findings, while the study’s cross-sectional design precludes causal inferences and underscores the need for longitudinal or interventional approaches. In addition, biosample collection, including oral microbiome and epigenetic data, occurred several months to years after demographic and functional assessments (median lag 189–336 days). This temporal offset introduces uncertainty, as molecular profiles may not fully reflect the physiological state at testing. Although significant time interactions (BADL and SPPB) were accounted for, unmeasured changes in health or lifestyle could still have influenced microbiome composition, DNA methylation, and functional outcomes. Longitudinal studies with synchronized sampling and repeated measures will be needed to resolve temporal dynamics and causal directionality. Further complexity arises from potential epigenetic clock miscalibration in older Asian cohorts, due to them being mainly trained on younger Caucasian individuals4,5,6,7, which could hence obscure accurate biological age estimation. To evaluate the robustness of observed associations, future research may benefit from applying ethnicity-specific clocks, such as the iCAS-DNAmAge model trained on Chinese cohorts75. Expanding investigations with animal models or other mechanistic approaches would clarify how oral microbiota–derived metabolites, such as SCFAs, modulate DNA methylation and related epigenetic and nervous system processes. While individual taxa showed statistically significant associations with MMSE, effect sizes were modest (∼0.9–1.6 points). Prior work suggests a 2–4 point MMSE shift is typically needed to indicate clinically meaningful change beyond measurement error76. However, at age 84, MMSE declines by ~0.53 points annually77, suggesting that even small cross-sectional differences may reflect deviations from expected cognitive trajectories. Importantly, cumulative effects of broader oral community structure, rather than isolated taxa, are likely to underlie observed associations, consistent with the complexity of neurocognitive aging. Consistent with the modest effect sizes observed throughout this study, the exploratory machine learning model captured only weak global association signals and did not outperform random or confounder-only baselines, underscoring the need for larger, more diverse multi-omic cohorts and modelling strategies that account for community-level network structure and site-specific epigenetic effects.
In summary, among the oldest old Singaporeans lower mAge deviation was linked to lower oral microbial Simpson diversity, characterized by a trend towards greater abundances of Prevotella_7 and Veillonella species, both of which were associated with methylation changes at CpG sites implicated in aging and cognitive function. Moreover, several Prevotellaceae taxa are associated with cognitive measures, suggesting that the oral microbiome may play a more nuanced role in the aging process than previously known. These novel findings offer a promising framework for understanding the interplay between oral health and aging, and they lay the groundwork for future studies.
Data availability
The data used in this analysis are available upon reasonable request from the corresponding author (Prof Andrea Maier, a.b.maier@vu.nl). All data access requests will be reviewed by the data committee, and a response will be provided within approximately four weeks. A formal Research Collaboration Agreement (RCA) may be required prior to data sharing to define the scope of use, responsibilities of parties involved, and conditions for data protection and publication. Source data underlying all main figures are provided as Supplementary Data 14.
Code availability
All analysis scripts and workflows used for data processing, statistical analyses, and figure generation are available at https://github.com/Tim-DIVO/sg90-omicrobiome-dnam78. The repository includes version-controlled Python (v3.11.5) and R (v4.4.1) scripts with detailed documentation of key parameters and package dependencies. No access restrictions apply.
References
Campisi, J. et al. From discoveries in ageing research to therapeutics for healthy ageing. Nature 571, 183–192 (2019).
López-Otín, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. The hallmarks of aging. Cell 153, 1194–1217 (2013).
Zampieri, M. et al. Reconfiguration of DNA methylation in aging. Mech. Ageing Dev. 151, 60–70 (2015).
Lu, A. T. et al. DNA methylation GrimAge strongly predicts lifespan and healthspan. Aging 11, 303–327 (2019).
Levine, M. E. et al. An epigenetic biomarker of aging for lifespan and healthspan. Aging 10, 573–591 (2018).
Hannum, G. et al. Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol. Cell 49, 359–367 (2013).
Horvath, S. DNA methylation age of human tissues and cell types. Genome Biol. 14, R115 (2013).
Oblak, L., van der Zaag, J., Higgins-Chen, A. T., Levine, M. E. & Boks, M. P. A systematic review of biological, social and environmental factors associated with epigenetic clock acceleration. Ageing Res. Rev. 69, 101348 (2021).
Chervova, O. et al. Breaking new ground on human health and well-being with epigenetic clocks: a systematic review and meta-analysis of epigenetic age acceleration associations. Ageing Res. Rev. 102, 102552 (2024).
Huang, Y., Chen, X., Ye, J., Yi, H. & Zheng, X. Causal effect of gut microbiota on DNA methylation phenotypic age acceleration: a two-sample Mendelian randomization study. Sci. Rep. 13, 18830 (2023).
Gilbert, J. A. et al. Current understanding of the human microbiome. Nat. Med. 24, 392–400 (2018).
Chen, X. et al. Oral microbiota and risk for esophageal squamous cell carcinoma in a high-risk area of China. PLoS One 10, e0143603 (2015).
Zhang, W. et al. Salivary microbial dysbiosis is associated with systemic inflammatory markers and predicted oral metabolites in non-small cell lung cancer patients. J. Cancer 10, 1651–1662 (2019).
Wells, P. M. et al. Influential factors of saliva microbiota composition. Sci Rep 12 (2022).
DeClercq, V., Wright, R. J., Nearing, J. T. & Langille, M. G. I. Oral microbial signatures associated with age and frailty in Canadian adults. Sci. Rep. 14, 9685 (2024).
Vogtmann, E. et al. Oral microbial community composition is associated with pancreatic cancer: a case-control study in Iran. Cancer Med 9, 797–806 (2020).
Yang, I. et al. Oral microbiome and cognition among black cancer caregivers. Nurs. Res. 74, 47–55 (2025).
Marin, M.-F. et al. Chronic stress, cognitive functioning and mental health. Neurobiol. Learn Mem. 96, 583–595 (2011).
Hu, T. et al. Relation between emotion regulation and mental health: a meta-analysis review. Psychol. Rep. 114, 341–362 (2014).
Liu, Y. et al. A salivary microbiome-based auxiliary diagnostic model for type 2 diabetes mellitus. Arch. Oral. Biol. 126, 105118 (2021).
Murugesan, S., Elanbari, M., Bangarusamy, D. K., Terranegra, A. & Al Khodor, S. Can the salivary microbiome predict cardiovascular diseases? Lessons learned from the Qatari population. Front. Microbiol. 12 (2021).
Kazarina, A. et al. Oral microbiome variations related to ageing: possible implications beyond oral health. Arch. Microbiol. 205, 116 (2023).
Chen, Z. et al. The intersection between oral microbiota, host gene methylation and patient outcomes in head and neck squamous cell carcinoma. Cancers (Basel) 12, 3425 (2020).
Cai, L. et al. Integrative analysis reveals associations between oral microbiota dysbiosis and host genetic and epigenetic aberrations in oral cavity squamous cell carcinoma. NPJ Biofilms Microbiomes 10, 39 (2024).
Bryan, N. S., Burleigh, M. C. & Easton, C. The oral microbiome, nitric oxide and exercise performance. Nitric Oxide 125–126, 23–30 (2022).
Ng, T. P. et al. Socio-environmental, lifestyle, behavioural, and psychological determinants of biological ageing: the singapore longitudinal ageing study. Gerontology 66, 603–613 (2020).
Chua, K. Y. et al. Visceral fat area is the measure of obesity best associated with mobility disability in community dwelling oldest-old Chinese adults. BMC Geriatr. 21, 282 (2021).
Hankin, J. H. et al. Singapore Chinese Health Study: development, validation, and calibration of the quantitative food frequency questionnaire. Nutr. Cancer 39, 187–195 (2001).
Guan, L. et al. The SG90 cohort of the oldest-old in Singapore. Eur. J. Epidemiol. https://doi.org/10.1007/s10654-025-01275-0 (2025)
Feng, L., Chong, M. S., Lim, W. S. & Ng, T. P. The Modified Mini-Mental State Examination test: normative data for Singapore Chinese older adults and its performance in detecting early cognitive impairment. Singap. Med. J. 53, 458–462 (2012).
Freiberger, E. et al. Performance-based physical function in older community-dwelling persons: a systematic review of instruments. Age Ageing 41, 712–721 (2012).
Mahoney, F. I. & Barthel, D. W. Functional evaluation: the BARTHEL INDEX. Md. State Med. J. 14, 61–65 (1965).
Graf, C. The Lawton Instrumental Activities of Daily Living Scale. AJN, Am. J. Nurs. 108, 52–62 (2008).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2012).
Higgins-Chen, A. T. et al. A computational solution for bolstering reliability of epigenetic clocks: implications for clinical trials and longitudinal tracking. Nat. Aging 2, 644–661 (2022).
Ori, A. P. S., Lu, A. T., Horvath, S. & Ophoff, R. A. Significant variation in the performance of DNA methylation predictors across data preprocessing and normalization strategies. Genome Biol. 23, 225 (2022).
Sandalova, E. et al. Alpha-ketoglutarate supplementation and BiologicaL agE in middle-aged adults (ABLE)—intervention study protocol. Geroscience 45, 2897–2907 (2023).
Koncevičius, K. et al. Epigenetic age oscillates during the day. Aging Cell 23 (2024).
Nearing, J. T., DeClercq, V., Van Limbergen, J. & Langille, M. G. I. Assessing the variation within the oral microbiome of healthy adults. mSphere 5 (2020).
Chaturvedi, A. K. et al. Oral microbiome profile of the US population. JAMA Netw. Open 8, e258283 (2025).
Wells, P. M. et al. Influential factors of saliva microbiota composition. Sci. Rep. 12, 18894 (2022).
Crum, R. M., Anthony, J. C., Bassett, S. S. & Folstein, M. F. Population-based norms for the Mini-Mental State Examination by age and educational level. JAMA 269, 2386–2391 (1993).
Schloss, P. D. Rarefaction is currently the best approach to control for uneven sequencing effort in amplicon sequence analyses. mSphere 9 (2024).
Martino, C. et al. A novel sparse compositional technique reveals microbial perturbations. mSystems 4 (2019).
Martino, C. et al. Compositionally aware phylogenetic beta-diversity measures better resolve microbiomes associated with phenotype. mSystems 7 (2022).
te Beest, D. E., Nijhuis, E. H., Möhlmann, T. W. R. & ter Braak, C. J. F. Log-ratio analysis of microbiome data with many zeroes is library size dependent. Mol. Ecol. Resour. 21, 1866–1874 (2021).
Lubbe, S., Filzmoser, P. & Templ, M. Comparison of zero replacement strategies for compositional data with large numbers of zeros. Chemometrics Intell. Lab. Syst. 210, 104248 (2021).
Kurtz, Z. D. et al. Sparse and compositionally robust inference of microbial ecological networks. PLoS Comput. Biol. 11, e1004226 (2015).
Lancichinetti, A. & Fortunato, S. Consensus clustering in complex networks. Sci. Rep. 2, 336 (2012).
Palasca, O., Santos, A., Stolte, C., Gorodkin, J. & Jensen, L. J. TISSUES 2.0: an integrative web resource on mammalian tissue expression. Database 2018 (2018).
Xie, Z. et al. Gene set knowledge discovery with enrichr. Curr. Protoc. 1 (2021).
Mansuy-Aubert, V. & Ravussin, Y. Short chain fatty acids: the messengers from down below. Front. Neurosci. 17 (2023).
Ross, J. M. et al. High brain lactate is a hallmark of aging and caused by a shift in the lactate dehydrogenase A/B ratio. Proc. Natl. Acad. Sci. 107, 20087–20092 (2010).
Yang, Z. et al. Association of oral microbiome diversity and all-cause mortality in the general US population and in individuals with chronic diseases: a prospective cohort study. J. Clin. Periodontol. 51, 1490–1501 (2024).
Ji, D. et al. Screening for frailty and its association with activities of daily living, cognitive impairment, and falls among community-dwelling older adults in China. BMC Geriatr. 24, 576 (2024).
Johnson, K. V. A. & Burnet, P. W. J. Microbiome: Should we diversify from diversity? Gut Microbes 7, 455–458 (2016).
Jiang, H. et al. Altered fecal microbiota composition in patients with major depressive disorder. Brain Behav. Immun. 48, 186–194 (2015).
Ma, J. et al. Comparison of gut microbiota in exclusively breast-fed and formula-fed babies: a study of 91 term infants. Sci. Rep. 10 (2020).
Fassarella, M. et al. Gut microbiome stability and resilience: Elucidating the response to perturbations in order to modulate gut health. Gut 70, 595–605 (2021).
Coyte, K. Z., Schluter, J. & Foster, K. R. The ecology of the microbiome: Networks, competition, and stability. Science 350, 663–666 (2015).
Ng, S. K. C. & Hamilton, I. R. Carbon dioxide fixation by Veillonella parvula M 4 and its relation to propionic acid formation. Can. J. Microbiol. 19, 715–723 (1973).
Chen, T. et al. Fiber-utilizing capacity varies in Prevotella- versus Bacteroides-dominated gut microbiota. Sci. Rep. 7, 2594 (2017).
Chambers, E. S. et al. Effects of targeted delivery of propionate to the human colon on appetite regulation, body weight maintenance and adiposity in overweight adults. Gut 64, 1744–1754 (2015).
Wahl, S. et al. Epigenome-wide association study of body mass index, and the adverse outcomes of adiposity. Nature 541, 81–86 (2017).
Haghikia, A. et al. Propionate attenuates atherosclerosis by immune-dependent regulation of intestinal cholesterol metabolism. Eur. Heart J. 43, 518–533 (2022).
Ramesh, V. et al. Propionate reinforces epithelial identity and reduces aggressiveness of lung carcinoma. EMBO Mol. Med. 15 (2023).
Stein, R. A. & Riber, L. Epigenetic effects of short-chain fatty acids from the large intestine on host cells. microLife 4 (2023).
Wang, G. G., Allis, C. D. & Chi, P. Chromatin remodeling and cancer, part I: covalent histone modifications. Trends Mol. Med. 13, 363–372 (2007).
Raghavan, N. S. et al. Association between common variants in RBFOX1, an RNA-binding protein, and brain amyloidosis in early and preclinical alzheimer disease. JAMA Neurol. 77, 1288 (2020).
Dundee, J. M., Puigdellívol, M., Butler, R., Cockram, T. O. J. & Brown, G. C. P2Y6 receptor-dependent microglial phagocytosis of synapses mediates synaptic and memory loss in aging. Aging Cell 22 (2023).
Claudino dos Santos, J. C. et al. Role of enteric glia and microbiota-gut-brain axis in parkinson disease pathogenesis. Ageing Res. Rev. 84, 101812 (2023).
Vicentini, F. A. et al. Intestinal microbiota shapes gut physiology and regulates enteric neurons and glia. Microbiome 9, 210 (2021).
Purushe, J. et al. Comparative genome analysis of Prevotella Ruminicola and Prevotella bryantii: insights into their environmental niche. Microb. Ecol. 60, 721–729 (2010).
Gazdeck, R. K. et al. Diversity of the oral microbiome between dentate and edentulous individuals. Oral. Dis. 25, 911–918 (2019).
Zheng, Z. et al. DNA methylation clocks for estimating biological age in Chinese cohorts. Protein Cell 15, 575–593 (2024).
Hensel, A., Angermeyer, M. C. & Riedel-Heller, S. G. Measuring cognitive change in older adults: reliable change indices for the Mini-Mental State Examination. J. Neurol. Neurosurg. Psychiatry 78, 1298–1303 (2007).
Nagaratnam, J. M., Sharmin, S., Diker, A., Lim, W. K. & Maier, A. B. Trajectories of Mini-Mental State Examination Scores over the lifespan in general populations: a systematic review and meta-regression analysis. Clin. Gerontol. 45, 467–476 (2022).
Wehnes, T. Analysis Github Repository. https://doi.org/10.5281/zenodo.17541946 (2025).
Acknowledgements
The authors extend our gratitude to the SG90 research teams as well as study participants for their valuable contributions. We would like to acknowledge that computational work involved in this research work is partially supported by NUS IT’s Research Computing group using grant number NUSREC-HPC-00001. The SG90 study was supported by the NUHS-A*STAR Biology of Human Ageing Program seed funding. The SCHS-SG90 study was additionally supported by the Agency for Science, Technology and Research (A*STAR) under the A*STAR-AMED Joint Grant Call in “Biological and molecular determinants of human ageing (AMED2016- SG002)”, as well as the National Medical Research Council of Singapore grant (NMRC/TA/0053/2016 and NMRC/CSA/0009/2022).
Author information
Authors and Affiliations
Contributions
All authors contributed significantly to the manuscript. Conceptualization: T.W., W.W., T.P.N., W.P.K., Y.S.C., and A.B.M. Methodology: T.W. and W.W. Data collection and curation: T.W., J.H.T., L.G., C.M.R.H., T.P.N., L.F., B.K.K., W.P.K., Y.S.C., and A.B.M. Formal analysis and investigation: T.W. and J.H.T. Writing—original draft preparation: T.W. Writing—review and editing: T.W., W.W., A.V., L.F., A.B.M. Supervision: W.W. and A.B.M. Funding acquisition and resources: T.P.N., W.P.K., Y.S.C., and A.B.M.
Corresponding author
Ethics declarations
Competing interests
The authors declared no competing interests.
Peer review
Peer review information
Communications Medicine thanks Diptaraj Chaudhari and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wehnes, T., Wang, W., Tay, J.H. et al. Oral short-chain fatty acid-producing bacteria may be associated with biological age and cognition among the oldest old. Commun Med 6, 37 (2026). https://doi.org/10.1038/s43856-025-01288-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s43856-025-01288-6