Abstract
Post-injury remodeling is a complex process involving temporal specific cellular interactions in the injured tissue where the resident fibroblasts play multiple roles. Here, we performed single-cell and spatial transcriptome analysis in human and mouse infarcted hearts to dissect the molecular basis of these interactions. We identified a unique fibroblast subset with high CD248 expression, strongly associated with extracellular matrix remodeling. Genetic Cd248 deletion in fibroblasts mitigated cardiac fibrosis and dysfunction following ischemia/reperfusion. Mechanistically, CD248 stabilizes type I transforming growth factor beta receptor and thus upregulates fibroblast ACKR3 expression, leading to enhanced T cell retention. This CD248-mediated fibroblast–T cell interaction is required to sustain fibroblast activation and scar expansion. Disrupting this interaction using monoclonal antibody or chimeric antigen receptor T cell reduces T cell infiltration and consequently ameliorates cardiac fibrosis and dysfunction. Our findings reveal a CD248+ fibroblast subpopulation as a key regulator of immune–fibroblast cross–talk and a potential therapy to treat tissue fibrosis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All data are available in the main text or the Supplementary Information. The bulk RNA-seq raw data for Fig. 3a,b are deposited in the Gene Expression Omnibus (GEO) under accession no. GSE287283. Raw scRNA-seq data for infarcted mouse hearts (Fig. 1a–f), infarcted human hearts (Fig. 1g–i), conditional CD248 KO mouse hearts subjected to MI (Extended Data Fig. 4d–g) and spatial transcriptome data for infarcted mouse hearts (Figs. 1j,k and 3f–i) are deposited in the Genome Sequence Archive (GSA) and GSA-Human at the National Genomics Data Center, China National Center for Bioinformation, under accession numbers CRA022616, HRA010249, CRA022593 and CRA022597, respectively, and are publicly accessible at https://ngdc.cncb.ac.cn/gsa/ and https://ngdc.cncb.ac.cn/gsa-human/. Additionally, scRNA-seq data of human fibrotic lung samples are available from the GEO (GSE121611), and human fibrotic kidney scRNA-seq data and human heart spatial transcriptome data can be accessed via Zenodo (https://doi.org/10.5281/zenodo.4059315 and https://doi.org/10.5281/zenodo.6580069)8,19.
Code availability
The code used to process the data is publicly available on GitHub (https://github.com/lingjun66/CD248/).
References
Henderson, N. C., Rieder, F. & Wynn, T. A. Fibrosis: from mechanisms to medicines. Nature 587, 555–566 (2020).
Kong, P., Christia, P. & Frangogiannis, N. G. The pathogenesis of cardiac fibrosis. Cell. Mol. Life Sci. 71, 549–574 (2014).
Travers, J. G., Kamal, F. A., Robbins, J., Yutzey, K. E. & Blaxall, B. C. Cardiac fibrosis: the fibroblast awakens. Circ. Res. 118, 1021–1040 (2016).
Ottaviano, F. G. & Yee, K. O. Communication signals between cardiac fibroblasts and cardiac myocytes. J. Cardiovasc. Pharmacol. 57, 513–521 (2011).
Bhattacharya, M. & Ramachandran, P. Immunology of human fibrosis. Nat. Immunol. 24, 1423–1433 (2023).
Forte, E., Furtado, M. B. & Rosenthal, N. The interstitium in cardiac repair: role of the immune-stromal cell interplay. Nat. Rev. Cardiol. 15, 601–616 (2018).
Ramachandran, P. et al. Resolving the fibrotic niche of human liver cirrhosis at single-cell level. Nature 575, 512–518 (2019).
Kuppe, C. et al. Decoding myofibroblast origins in human kidney fibrosis. Nature 589, 281–286 (2021).
Bowers, S. L. K., Meng, Q. & Molkentin, J. D. Fibroblasts orchestrate cellular crosstalk in the heart through the ECM. Nat. Cardiovasc. Res. 1, 312–321 (2022).
Amrute, J. M. et al. Targeting immune-fibroblast crosstalk in myocardial infarction and cardiac fibrosis. Preprint at https://doi.org/10.21203/rs.3.rs-2402606/v1 (2023).
Hammerich, L. & Tacke, F. Hepatic inflammatory responses in liver fibrosis. Nat. Rev. Gastroenterol. Hepatol. 20, 633–646 (2023).
Bradshaw, A. D. & DeLeon-Pennell, K. Y. T-cell regulation of fibroblasts and cardiac fibrosis. Matrix Biol. 91-92, 167–175 (2020).
Farbehi, N. et al. Single-cell expression profiling reveals dynamic flux of cardiac stromal, vascular and immune cells in health and injury. Elife https://doi.org/10.7554/eLife.43882 (2019).
Forte, E. et al. Dynamic interstitial cell response during myocardial infarction predicts resilience to rupture in genetically diverse mice. Cell Rep. 30, 3149–3163 (2020).
Janbandhu, V. et al. Hif-1a suppresses ROS-induced proliferation of cardiac fibroblasts following myocardial infarction. Cell Stem Cell 29, 281–297 (2022).
Patrick, R. et al. Integration mapping of cardiac fibroblast single-cell transcriptomes elucidates cellular principles of fibrosis in diverse pathologies. Sci. Adv. 10, eadk8501 (2024).
Batlle, E. & Massague, J. Transforming growth factor-beta signaling in immunity and cancer. Immunity 50, 924–940 (2019).
Frangogiannis, N. G. Transforming growth factor-beta in myocardial disease. Nat. Rev. Cardiol. 19, 435–455 (2022).
Reyfman, P. A. et al. Single-cell transcriptomic analysis of human lung provides insights into the pathobiology of pulmonary fibrosis. Am. J. Respir. Crit. Care Med. 199, 1517–1536 (2019).
Tallquist, M. D. Cardiac fibroblast diversity. Annu. Rev. Physiol. 82, 63–78 (2020).
Miranda, A. M. A. et al. Single-cell transcriptomics for the assessment of cardiac disease. Nat. Rev. Cardiol. 20, 289–308 (2023).
Ruiz-Villalba, A. et al. Single-cell RNA sequencing analysis reveals a crucial role for CTHRC1 (collagen triple helix repeat containing 1) cardiac fibroblasts after myocardial infarction. Circulation 142, 1831–1847 (2020).
Gladka, M. M. et al. Single-cell sequencing of the healthy and diseased heart reveals cytoskeleton-associated protein 4 as a new modulator of fibroblasts activation. Circulation 138, 166–180 (2018).
Buechler, M. B., Fu, W. & Turley, S. J. Fibroblast-macrophage reciprocal interactions in health, fibrosis, and cancer. Immunity 54, 903–915 (2021).
Dolejsi, T. et al. Adult T-cells impair neonatal cardiac regeneration. Eur. Heart J. 43, 2698–2709 (2022).
Bayer, A. L. et al. T-cell MyD88 is a novel regulator of cardiac fibrosis through modulation of T-cell activation. Circ. Res. 133, 412–429 (2023).
Feng, G. et al. CCL17 aggravates myocardial injury by suppressing recruitment of regulatory T cells. Circulation 145, 765–782 (2022).
Davidson, S. et al. Fibroblasts as immune regulators in infection, inflammation and cancer. Nat. Rev. Immunol. 21, 704–717 (2021).
Komatsu, N. & Takayanagi, H. Mechanisms of joint destruction in rheumatoid arthritis - immune cell-fibroblast-bone interactions. Nat. Rev. Rheumatol. 18, 415–429 (2022).
Chhabra, Y. & Weeraratna, A. T. Fibroblasts in cancer: unity in heterogeneity. Cell 186, 1580–1609 (2023).
Arpinati, L. & Scherz-Shouval, R. From gatekeepers to providers: regulation of immune functions by cancer-associated fibroblasts. Trends Cancer 9, 421–443 (2023).
Valdez, Y., Maia, M. & Conway, E. M. CD248: reviewing its role in health and disease. Curr. Drug Targets 13, 432–439 (2012).
Maia, M. et al. CD248 and its cytoplasmic domain: a therapeutic target for arthritis. Arthritis Rheum. 62, 3595–3606 (2010).
Alampour-Rajabi, S. et al. MIF interacts with CXCR7 to promote receptor internalization, ERK1/2 and ZAP-70 signaling, and lymphocyte chemotaxis. FASEB J. 29, 4497–4511 (2015).
Gencer, S. et al. Endothelial ACKR3 drives atherosclerosis by promoting immune cell adhesion to vascular endothelium. Basic Res. Cardiol. 117, 30 (2022).
Li, S. et al. Activation of MAPK signaling by CXCR7 leads to enzalutamide resistance in prostate cancer. Cancer Res. 79, 2580–2592 (2019).
Aghajanian, H. et al. Targeting cardiac fibrosis with engineered T cells. Nature 573, 430–433 (2019).
Rurik, J. G. et al. CAR T cells produced in vivo to treat cardiac injury. Science 375, 91–96 (2022).
Kanisicak, O. et al. Genetic lineage tracing defines myofibroblast origin and function in the injured heart. Nat. Commun. 7, 12260 (2016).
Travers, J. G. et al. Pharmacological and activated fibroblast targeting of Gβγ-GRK2 after myocardial ischemia attenuates heart failure progression. J. Am. Coll. Cardiol. 70, 958–971 (2017).
Kuwabara, J. T. et al. Consequences of PDGFRα+ fibroblast reduction in adult murine hearts. Elife https://doi.org/10.7554/eLife.69854 (2022).
Chen H. et al. CD248-targeted BBIR-T cell therapy against late-activated fibroblasts in cardiac repair after myocardial infarction. Nat. Commun. https://doi.org/10.1038/s41467-025-56703-2 (2025).
Ash, S. L. et al. Targeting the activated microenvironment with endosialin (CD248)-directed CAR-T cells ablates perivascular cells to impair tumor growth and metastasis. J. Immunother. Cancer https://doi.org/10.1136/jitc-2023-008608 (2024).
Fierle, J. K. et al. Soluble trivalent engagers redirect cytolytic T cell activity toward tumor endothelial marker 1. Cell Rep. Med. 2, 100362 (2021).
Hardie, D. L. et al. The stromal cell antigen CD248 (endosialin) is expressed on naive CD8+ human T cells and regulates proliferation. Immunology 133, 288–295 (2011).
Parker, K. R. et al. Single-cell analyses identify brain mural cells expressing CD19 as potential off-tumor targets for CAR-T immunotherapies. Cell 183, 126–142 (2020).
Roybal, K. T. et al. Precision tumor recognition by T cells with combinatorial antigen-sensing circuits. Cell 164, 770–779 (2016).
Fedorov, V. D., Themeli, M. & Sadelain, M. PD-1- and CTLA-4-based inhibitory chimeric antigen receptors (iCARs) divert off-target immunotherapy responses. Sci. Transl. Med. 5, 215ra172 (2013).
Khalil, H. et al. Cell-specific ablation of Hsp47 defines the collagen-producing cells in the injured heart. JCI Insight 4, e128722 (2019).
Lindsey, M. L. et al. Guidelines for in vivo mouse models of myocardial infarction. Am. J. Physiol. Heart. Circ. Physiol. 321, H1056–H1073 (2021).
Respress, J. L. & Wehrens, X. H. Transthoracic echocardiography in mice. J. Vis. Exp. https://doi.org/10.3791/1738 (2010).
Zacchigna, S. et al. Towards standardization of echocardiography for the evaluation of left ventricular function in adult rodents: a position paper of the ESC Working Group on Myocardial Function. Cardiovasc. Res. 117, 43–59 (2021).
Koenig, A. L. et al. Single-cell transcriptomics reveals cell-type-specific diversification in human heart failure. Nat. Cardiovasc. Res. 1, 263–280 (2022).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–1792 (2022).
Moon, K. R. et al. Visualizing structure and transitions in high-dimensional biological data. Nat. Biotechnol. 37, 1482–1492 (2019).
Kuppe, C. et al. Spatial multi-omic map of human myocardial infarction. Nature 608, 766–777 (2022).
Zhao, A. et al. Rapid isolation of high-affinity human antibodies against the tumor vascular marker endosialin/TEM1, using a paired yeast-display/secretory scFv library platform. J. Immunol. Methods 363, 221–232 (2011).
Yang, F. et al. Interaction with CD68 and regulation of GAS6 expression by endosialin in fibroblasts drives recruitment and polarization of macrophages in hepatocellular carcinoma. Cancer Res. 80, 3892–3905 (2020).
Wang, L. C. et al. Targeting fibroblast activation protein in tumor stroma with chimeric antigen receptor T cells can inhibit tumor growth and augment host immunity without severe toxicity. Cancer Immunol. Res. 2, 154–166 (2014).
Bansal, S. S. et al. Activated T lymphocytes are essential drivers of pathological remodeling in ischemic heart failure. Circ. Heart. Fail. 10, e003688 (2017).
Li, I. M. H., Horwell, A. L., Chu, G., de Crombrugghe, B. & Bou-Gharios, G. Characterization of mesenchymal-fibroblast cells using the Col1a2 promoter/enhancer. Methods Mol. Biol. 1627, 139–161 (2017).
Acknowledgements
This work was supported by grants from the Key Project of National Natural Science Foundation of China (no. 82430013 for X.H.), National Key Research and Development Program of China (no. 2023YFA1800700 for X.H.), National Science Foundation for Distinguished Young Scholars (no. 82225004 for X.H.), National Natural Science Foundation of China (no. U22A20267 and 82030014 for Jian’an Wang, U21A20338 and 82370256 for J.C., 82370240 for C.N., and 82400296 for G.L.) and received financial support from Binjiang Institute of Zhejiang University (ZY202205SMKY002 for X.H.) and Fundamental Research Funds for the Central Universities (226-2024-00224 for G.L. and K20240127 for B.W.).
Author information
Authors and Affiliations
Contributions
G.L., C.N., Jiacheng Wang and F.Z. performed mouse in vivo surgeries and downstream experiments. G.L. isolated NMCFs, and G.L., F.L. and S.Y. performed downstream in vitro experiments. G.L. isolated spleen T cells, and performed all CAR T cell therapy-associated experiments. G.L., H.L. and J.Z. performed Sirius red staining and analyzed images. Y.Z., J. Cai, Y.S. and Z.Z. performed immunofluorescence staining and analyzed images. Z.F., L.W., M.L. and S.S. performed scRNA-seq and spatial transcriptomic analysis. B.W., Y.L. and J.H. analyzed the data. X.X., Y.X., J. Chen, W.Z. and Y.W. contributed to the experimental design and manuscript modification. Jian’an Wang and X.H. designed the experiments, supervised the study and drafted and revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cardiovascular Research thanks Joel G. Rurik, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Global view of cardiac fibroblast single-cell RNA sequencing from human myocardial infarction heart.
a, UMAP dimensionality reduction plots of all 6989 sequenced cardiac fibroblasts from different cardiac areas of a MI patient. b, Enrichment of each fibroblast subset in human heart from infarct zone or remote area. c, Heatmap of top marker genes for each fibroblast subtype in UMAP. d, Venn diagram analysis of mouse cardiac F9 subset and human F5 subset. e, Quantification score of top 100 differentially expressed genes in mouse heart CD248+ fibroblast subset across different human heart fibroblast subsets. f, Immunofluorescence analysis of CD248 (red) expression in vimentin-marked cardiac fibroblasts from normal and infarcted human heart. g, Western-blot analysis of CD248 expression (by the Abcam antibody, no. 67273) in I/R-injured mouse hearts at different timepoints (sham, 3 days, 7 days, 14 days and 28 days post I/R). h, Different cell types were FACS-sorted from infarcted heart to analyze CD248 expression (by the Abcam antibody, no. 67273) by western-blot at 28 days post-infarction. i, Co-immunofluorescence staining of CD248 (red) with cardiomyocytes (Troponin I, green), endothelial cells (CD31, green), fibroblasts (Vimentin, green), T cells (CD3, green) and macrophage (CD68, green) in mouse heart 28 days post-infarction respectively. Scale bar = 20 μm.
Extended Data Fig. 2 Global knock-out of CD248 attenuates cardiac dysfunction in mice subjected to I/R or MI.
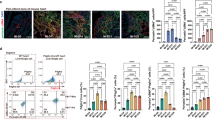
a, Genotyping of CD248 knock-out (KO) mice. b, Representative pulse-wave Doppler (left) and tissue Doppler (right) images of WT and CD248 KO mice. c, Quantification of baseline mitral E/e’ ratio, isovolumic relaxation time (IVRT), and left ventricular myocardial performance index (LV MPI) in WT and CD248 KO mice. d, e, Flow cytometry analysis of the proportion (d) and number (e) of PDGFRα+ fibroblasts in WT and CD248 KO mouse hearts. f, Biochemical analysis of hepatic function, renal function, and blood lipid levels by analyzing serum aspartate aminotransferase (AST), alanine aminotransferase (ALT), albumin (ALB), total protein (TP), total bilirubin (TBIL), uric acid (UA), urea nitrogen (BUN), creatinine (CREA), triglyceride (TG), cholesterol (CHOL), high density lipoprotein (HDL), and low density lipoprotein (LDL) in WT and CD248 KO mice. g, Schematic of experiment for I/R in CD248 KO mice. h, Representative M-mode echocardiogram images of mice from WT+Sham, CD248 KO+Sham, WT + I/R, CD248 KO + I/R at 4 days and 28 days post-I/R injury. i, LVESV and LVEDV measurement in animals of WT+Sham, CD248 KO+Sham, WT + I/R, and CD248 KO + I/R groups at 4 days and 28 days post I/R. n = 5 animals in CD248 KO group, and n = 6 animals in the other groups. j, Schematic of experiment for MI in CD248 KO mice. k. Representative M-mode echocardiogram images of mice from WT+Sham, CD248 KO+Sham, WT + I/R, CD248 KO + I/R at 4 days and 28 days post-MI injury. l, LVESV and LVEDV measurement in animals of WT+Sham (n = 6), CD248 KO+Sham (n = 6), WT + MI (n = 9), and CD248 KO + MI (n = 9) groups at 4 days and 28 days post MI. Data are shown as mean ± s.e.m.; unpaired two-tailed Student’s t-test (c-f) and two-way ANOVA (i and l).
Extended Data Fig. 3 Specific deletion of CD248 in cardiac fibroblasts improved cardiac function in mice subjected to I/R injury.
a, Genotyping of CD248flox-Postn-CreER mice. b, Scheme of in vivo I/R experiment conducted on CD248fl/fl and CD248Postn-creER mice. c, Western-blot analysis of CD248 deletion efficiency (by the Proteintech antibody, no. 60170) driven by Postn-CreER in mice with or without I/R injury. d, UMAP of fibroblast subpopulations of infarcted CD248Postn-CreER and CD248fl/fl mouse hearts 14 days post-myocardial infarction. e, Top expressed marker genes of each fibroblast subpopulation. f, Proportion of each fibroblast subpopulation in the infarcted CD248fl/fl and CD248Postn-CreER mouse hearts 14 days post-myocardial infarction. g, z-score of CD248 in UAMP. h, Representative M-mode echocardiogram images of animals from CD248fl/fl+Sham, CD248Postn-CreER+Sham, CD248fl/fl+I/R, CD248Postn-CreER+I/R groups at 4 days and 28 days post-I/R. i, LVESV and LVEDV measurement of animals from CD248fl/fl+Sham (n = 6), CD248Postn-CreER+Sham (n = 6), CD248fl/fl+I/R (n = 7), CD248Postn-CreER+I/R (n = 7) groups at 4 days and 28 days post I/R. Data are shown as mean ± s.e.m.; Statistical analysis was performed using two-way ANOVA.
Extended Data Fig. 4 CD248+ fibroblasts were involved in cardiac inflammation regulation post ischemia injuries, in which ACKR3 played an important role.
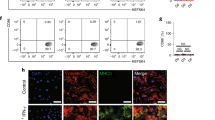
a, Heatmap of up-regulated and down-regulated genes related with inflammation between CD248+ and CD248- fibroblasts isolated from infarcted mouses heart at 14 days post-infarction. b-e, Immunofluorescence analysis of the infiltration of CD68+ macrophage (b) and CD3+ T cells (d) in hearts of CD248 KO and WT mice 14 days after I/R injury, with statistical analysis respectively shown in c and e. n = 5 in each group. Scale bar = 20 μm. f, g. Representative dot plots showing cardiac T cells in WT and CD248 KO mice at one week post-I/R (f) or MI (g) injuries. h, ACKR3 mRNA level detected by RT-PCR in NMCFs transfected with sh-CD248 and OE-CD248 lentivirus. n = 4 in each group. i, ACKR3 protein expression in NMCFs with sh-CD248 treatment compared with scramble treatment, among which CD248 expression analysis was performed using the Abcam antibody (no. 67273). Statistical analysis was plotted in j. n = 3 in each group. Data are shown as mean ± s.e.m.; unpaired two-tailed Student’s t-test (c, e and j) and one-way ANOVA (h).
Extended Data Fig. 5 Knocking-down CD248 reduced TGF-βRI expression and inhibited the activation of canonical and non-canonical signaling down-streaming of TGF-β receptor.
a, Western-blot analysis of canonical and non-canonical TGF-β signaling pathways (p-Smad2/Smad2, p-Smad3/Smad3, p-ERK/ERK, p-p38/p38) in WT and CD248 KO mouse hearts with or without I/R injury. Statistical analysis was shown in b. n = 6 in each group. c, Western-blot analysis of TGF-βRI and its downstream canonical and non-canonical signaling (p-Smad2/Smad2, p-Smad3/Smad3, p-ERK/ERK, p-p38/p38) in cardiac fibroblasts transfected with scramble or OE-CD248 lentivirus, and statistical analysis was shown in d. n = 3 in each group. e, RT-PCR analysis of TGF-βRI and TGF-βRII mRNA level in CD248-overexpression (OE-CD248) versus scramble-treated cardiac fibroblasts, n = 4 in each group. f, TGF-βRI protein level detected in cardiac fibroblasts treated with Scramble+DMSO, sh-CD248 + DMSO, sh-CD248+baflomycin and sh-CD248 + MG132 separately. Statistical analysis was shown in g. n = 6 in each group. h, Combination analysis of TGF-βRI and CD248 by co-immunoprecipitation in cultured cardiac fibroblasts. CD248 expression analysis in a and f was conducted using the Abcam antibody (no. 67273), while analysis in c and h was performed using the Proteintech antibody (no. 60170). Data are shown as mean ± s.e.m.; unpaired two-tailed Student’s t-test (d and e) and two-way ANOVA (b and g).
Extended Data Fig. 6 CD248 monoclonal antibody reduced cardiac T cell infiltration and attenuated cardiac dysfunction in mice subjected to I/R injury.
a, RT-PCR analysis of CD248 mRNA level in the mouse heart, lung, aorta, liver, kidney, spleen and intestinal tract at baseline and 3 days, 7 days, and 14 days post myocardial infarction, with CD248 mRNA level in sham-operated mouse hearts as normalization. n = 5-6 animals in each group. b, Binding affinity of IgG78 with CD248 protein as reflected by OD value at different concentration. c, Flow cytometry analysis of the binding specificity of IgG78 with CD248 using CD248 knockout and CD248 overexpressed adult mouse cardiac fibroblasts. d, Representative M-mode echocardiogram images of mice from sham+IgG, sham+IgG78, I/R+IgG, I/R + IgG78 at 4 days and 28 days post I/R injury. e, LVESV and LVEDV measurement in mice from Sham+IgG, Sham+IgG78, I/R+IgG, I/R + IgG78 groups at 4 days and 28 days post I/R injury. n = 6 animals in each group. f-i, Immunofluorescence analysis of CD3+ T cell (f) and CD68+ macrophage (h) infiltration in I/R-injured mouse hearts with IgG or IgG78 treatment; and statistical analysis were respectively shown in g and i. n = 5 animals in each group. Scale bar = 20 μm. j, Representative dot plots showing cardiac T cells infiltration in I/R-injured mouse hearts after three times of IgG or IgG78 treatment. k, Western-blot analysis of ACKR3 expression in TGF-β-stimulated cardiac fibroblasts with IgG or IgG78 treatment; and statistical analysis were shown in l. Data are shown as mean ± s.e.m.; unpaired two-tailed Student’s t-test (g and i) and two-way ANOVA (e and l).
Extended Data Fig. 7 CD248 CAR T construction strategy and in vivo detection.
a, Schematic diagram of Pan T isolation, anti-CD3/CD28 microbead activation, and retroviral vector transduction to construct CAR T cell. b, Construct maps encoding mouse CD248 CAR and mouse PSCA CAR. c, Flow cytometry analysis showing the expression level of CAR in PSCA CAR T or CD248 CAR T cells, with statistical analysis shown in d. e, CAR T cell subsets analysis by flow cytometry. f, g. Cytotoxic T cell activity as determined by CD248 expression (f) and CD248+ proportion (g) of fibroblasts using flow cytometry, in which the CD248-overexpressed fibroblasts as targets. T: E ratio, target-to-effector ratio. n = 3 independent experiments in each group. h, Representative CFSE images of the hearts from T cell or CD248 CAR T cell-transfused I/R-injured mice 48 hours after cell transfusion, and statistical analysis of fluorescence intensity were plotted in i. n = 3 animals in each group. j, Immunofluorescence analysis of PSCA CAR T and CD248 CAR T infiltration by GFP staining in I/R-injured mouse hearts 72 hours post-CAR T cell injection. k, Representative dot plots showing cardiac PSCA or CD248 CAR T cells infiltration in I/R-injured mouse hearts 72 hours after CAR T cell injection. l, Quantification of the number of PSCA or CD248 CAR T cells in the I/R-injured mouse hearts at 72 hours post-CAR T cell injection. m, Assessment of the activation status of cardiac infiltrated CAR T cells by CD69 detection using flow cytometry, with statistical analysis shown in n. o, p, Flow cytometry analysis of PSCA or CD248 CAR T cell infiltration (o) and activation (p) in the lung tissues of mice subjected to cardiac I/R injury at 72 hours post-CAR T cell injection. Data are shown as mean ± s.e.m.; unpaired two-tailed Student’s t-test (i, l, n, o and p), one-way ANOVA (d and e) and two-way ANOVA (g).
Extended Data Fig. 8 Assessment of cardiac function, cardiac T cell infiltration, and safety evaluation after CD248 CAR T cell therapy in mice.
a, Representative M-mode echocardiogram images of animals from Sham, I/R, I/R + PCSA CAR T, I/R + CD248 CAR T groups at 7 days and 28 days post I/R injury. b, LVESV and LVEDV measurement in animals from Sham, I/R, I/R + PCSA CAR T, I/R + CD248 CAR T groups at 7 days and 28 days post I/R. n = 8 animals in I/R + PSCA CAR T group, and n = 9 animals in the other groups. c, Representative dot plots showing cardiac T cells infiltration in I/R-injured mouse hearts one week after PSCA CAR T or CD248 CAR T treatment. d, Safety evaluation of CD248 CAR T cell therapy by serum detection of IL-6, IL-10, MCP-1, IFN-γ, TNF-α, IL-1β, IL-10, and IL12p7 using cytometric bead arrays in mice subjected to I/R. n = 6 in each group. e, Representative images of H&E staining of the liver, lung, spleen in animals from Sham, I/R and I/R + CD248 CAR T groups. Data are shown as mean ± s.e.m.; unpaired two-tailed Student’s t-test (d) and two-way ANOVA (b).
Extended Data Fig. 9 Analysis of CD248+ fibroblast in mouse pulmonary and renal fibrosis samples and single-cell RNA sequencing database on human pulmonary and renal fibrosis.
a, b, Immunofluorescence analysis of CD248 (red) positive fibroblasts (Vimentin, green) in mouse bleomycin-treated lungs (a) and unilateral ureteral obstruction-subjected kidneys (b). Scale bar = 20 μm. c, UMAP of pulmonary fibroblast subpopulation in patients with pulmonary fibrosis. d, CD248 expression analysis across all lung fibroblast subsets. e, f, GO (e) and KEGG (f) signaling enrichment analysis from CD248+ F5 lung fibroblast subset. g, UMAP of renal fibroblast subpopulation in patients with chronic kidney disease. h, CD248 expression analysis across all renal fibroblast subsets. i, j, GO (i) and KEGG (j) signaling enrichment analysis from CD248+ F6 kidney fibroblast subset.
Extended Data Fig. 10 Gating strategies used for flow cytometry analysis and cell sorting.
a, Gating strategy to sort cardiac fibroblasts (CD45-CD31-PDGFR-α+ ) from infarcted mouse hearts for single-cell RNA sequencing (Fig. 1a-f) and CD248 expression analysis (Fig. 1n-p). b, Gating strategy to sort cardiac fibroblasts (CD45-CD31-PDGFR-α+ ) from ischemia/reperfusion-injured mouse hearts for ACKR3 expression analysis (Fig. 4d, e). c, Gating strategy for flow cytometry analysis of the proportion of macrophages, neutrophils, T cells and B cells in ischemia/reperfusion-injured mouse hearts (Fig. 3c). d, Gating strategy for flow cytometry analysis of CD4+ /CD8+ and TH1/TH2/TH17 T cell number in the ischemic mouse hearts (Figs. 3d, e, 4n, 6h, and 7g).
Supplementary information
Supplementary Information (download PDF )
Supplementary Figures 1-3, and Supplementary Tables 1-6.
Source data
Source Data Fig. 1 (download XLSX )
Statistical Source Data.
Source Data Fig. 1 (download PDF )
Unprocessed western blots for Fig. 1.
Source Data Fig. 2 (download XLSX )
Statistical Source Data for Fig. 2.
Source Data Fig. 2 (download PDF )
Unprocessed western blots for Fig. 2.
Source Data Fig. 3 (download XLSX )
Statistical Source Data for Fig. 3.
Source Data Fig. 4 (download XLSX )
Statistical Source Data for Fig. 4.
Source Data Fig. 4 (download PDF )
Unprocessed western blots for Fig. 4.
Source Data Fig. 5 (download XLSX )
Statistical Source Data for Fig. 5.
Source Data Fig. 5 (download PDF )
Unprocessed western blots for Fig. 5.
Source Data Fig. 6 (download XLSX )
Statistical Source Data for Fig. 6.
Source Data Fig. 6 (download PDF )
Unprocessed western blots for Fig. 6.
Source Data Fig. 7 (download XLSX )
Statistical Source Data for Fig. 7.
Source Data Fig. 8 (download XLSX )
Statistical Source Data for Fig. 8.
Source Data Extended Data Fig. 1 (download PDF )
Unprocessed western blots for Extended Data Fig. 1.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical Source Data for Extended Data Fig. 2.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical Source Data for Extended Data Fig. 3.
Source Data Extended Data Fig. 3 (download PDF )
Unprocessed western blots for Extended Data Fig. 3.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical Source Data for Extended Data Fig. 4.
Source Data Extended Data Fig. 4 (download PDF )
Unprocessed western blots for Extended Data Fig. 4.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical Source Data for Extended Data Fig. 5.
Source Data Extended Data Fig. 5 (download PDF )
Unprocessed western blots for Extended Data Fig. 5.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical Source Data for Extended Data Fig. 6.
Source Data Extended Data Fig. 6 (download PDF )
Unprocessed western blots for Extended Data Fig. 6.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical Source Data for Extended Data Fig. 7.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical Source Data for Extended Data Fig. 8.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, G., Ni, C., Wang, J. et al. Dynamic molecular atlas of cardiac fibrosis at single-cell resolution shows CD248 in cardiac fibroblasts orchestrates interactions with immune cells. Nat Cardiovasc Res 4, 380–396 (2025). https://doi.org/10.1038/s44161-025-00617-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s44161-025-00617-1
This article is cited by
-
Mechanisms, precision therapies, and technological frontiers in coronary atherosclerosis: a comprehensive review
Acta Pharmacologica Sinica (2026)
-
CD248-targeted BBIR-T cell therapy against late-activated fibroblasts in cardiac repair after myocardial infarction
Nature Communications (2025)